Abstract
Rubus fruits are high-value crops that are sought after by consumers for their flavor, visual appeal, and health benefits. To meet this demand, production of red and black raspberries (R. idaeus L. and R. occidentalis L.), blackberries (R. subgenus Rubus), and hybrids, such as Boysenberry and marionberry, is growing worldwide. Rubus breeding programmes are continually striving to improve flavor, texture, machine harvestability, and yield, provide pest and disease resistance, improve storage and processing properties, and optimize fruits and plants for different production and harvest systems. Breeders face numerous challenges, such as polyploidy, the lack of genetic diversity in many of the elite cultivars, and until recently, the relative shortage of genetic and genomic resources available for Rubus. This review will highlight the development of continually improving genetic maps, the identification of Quantitative Trait Loci (QTL)s controlling key traits, draft genomes for red and black raspberry, and efforts to improve gene models. The development of genetic maps and markers, the molecular characterization of wild species and germplasm, and high-throughput genotyping platforms will expedite breeding of improved cultivars. Fully sequenced genomes and accurate gene models facilitate identification of genes underlying traits of interest and enable gene editing technologies such as CRISPR/Cas9.
Similar content being viewed by others

Introduction
Rubus is a large and diverse genus in the Rosoideae subfamily of Rosaceae, with over 740 species described worldwide1. Based on phenotypic diversity, it is thought that Rubus originated in southwestern China2. Pliny the Elder (AD 45) wrote about the people of Troy gathering “ida fruits” (red raspberries) at the base of Mount Ida in what is now Turkey. Although there are species native to most temperate regions, they are also found from the sub-tropics to Arctic regions and can grow from sea level to 4500 m3.
The genus is divided into 12 subgenera4. There is a wide spectrum of wild species, but the focus for domestication and breeding, and the most economically important crops are red and black raspberry (R. idaeus L. and R. occidentalis L., both in subgenus Idaeobatus), and blackberry (R. subgenus Rubus). Raspberries are diploid (2n = 2 × = 14), and blackberries range from diploid to 12 × (2n = 2 × = 14 to 2n = 12 × = 84). Red and black raspberries readily hybridize to produce purple raspberries. Generally, blackberry cultivars are not assigned a species, as there are several species in the ancestry of all the cultivars. Similarly, R. idaeus and several other species hybridize with the blackberry species to produce fertile accessions. Natural and human made hybrids are common within Rubus. This review will focus primarily on the past, present, and future of genetic and genomic tools to facilitate the improvement of raspberries and blackberries.
Prior to domestication, the primary use of Rubus, especially blackberry, was medicinal and they were foraged by indigenous communities. There are records of the root, leaves, stem, and fruits being used to treat a variety of ailments5. More recently, Rubus fruits were found to be very high in secondary metabolites, such as anthocyanins and phenolics, that provide antioxidant capacity, supporting their reputation as “superfoods”6,7,8,9,10. Blackberries are particularly high in dietary fiber, vitamins C and K, and manganese11.
Modern cultivars are bred for the fresh market and processing (e.g., freezing, drying, canning), and for the home garden. In 2017, world production of red and black raspberries reached 840,000 tonnes, with Europe and the Americas being the top producers12. A survey of global blackberry production in 2005 indicated that the two largest production regions were North America (59,123 tonnes) and Europe (43,000 tonnes)13. During the past 14 years, since the last worldwide survey was conducted, the global blackberry industry has experienced rapid change and growth. This growth has been driven by increased consumer demand, new cultivars, advanced production methods, and year-round product availability. Blackberries are currently the fourth most economically important U.S. berry crop, accounting for $549 million in sales during 201614. The Mexican fresh-market blackberry industry, in particular, has rapidly expanded, with almost 11,000 ha planted in 2015 and most fruit destined for export to the USA and other markets15,16. Fresh-market blackberry production is also growing in other regions including Southern Europe, Australia, and Central and South America.
There are a number of Rubus breeding programs worldwide. General breeding targets include high fruit quality and yield, extended cropping season, good storage and processing properties, disease and pest resistance, and adaptation to local growing environments. Other desired traits are specific for the fresh market, processing, and ornamental cultivars.
Rubus life cycle and physiology
The plants, commonly known as brambles or caneberries, generally grow as a deciduous shrub with biennial canes that are initiated from a perennial root system. The canes often have epidermal spines, ranging from hair-like to sharp thorns. Growth forms can be erect, trailing, or more vine-like. Rubus can propagate sexually, by apomixis, and vegetatively, enabling them to be highly invasive.
Most Rubus plants are biennial-fruiting (BF) (also called floricane-fruiting or summer-fruiting); these initiate axillary floral buds toward autumn of the first year of growth, but the buds do not develop into fruits until spring/summer of the following year. Annual-fruiting (AF) cultivars of both raspberry and blackberry (also called primocane-fruiting or autumn-fruiting) initiate flowers in late spring/early summer and these develop into fruits from summer until late autumn of the same year. In both types, flowering and fruiting initiates from the shoot tip and develops basipetally after vegetative growth has stopped. The key developmental difference between the two flowering phenologies is that AF floral buds are initiated earlier and progress directly to fruit set, whereas floral initiation is followed by dormancy in BF types17,18,19,20,21,22.
Based on a number of studies, floral induction in BF cultivars is triggered by a combination of decreased temperatures and shorter photoperiod18,19,21,23. While there is no absolute requirement for AF cultivars to experience chilling in the season prior to initiate flowering, as newly initiated canes can progress through to fruiting in a single season, the expression of AF in terms of floral consistency across canes and the total number of flowers is strongly influenced by chilling17,24. There is also evidence of a short juvenile phase of 15 or more vegetative nodes before plants are able to flower, even under inductive conditions19,21.
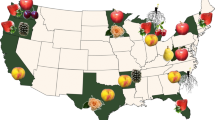
Rubus fruits are an aggregate of small fleshy drupelets, each containing a single seed derived from a fertilized ovary. The fruit takes 35–45 days to develop and can be germinated after scarification and a short period of stratification. At maturity raspberries detach from the receptacle, whereas blackberries and many hybrids do not, and the receptacle is picked with the fruit. Figure 1 shows a range of Rubus fruits.
Genetic markers for faster breeding
Relative to other crops, there have been few genetic and genomic resources available for Rubus improvement until recently. For raspberry, the lack of genetic diversity found in most of the elite cultivars presents another challenge25. The development of high-density genetic maps and markers, the molecular characterization of more wild species and germplasm, and high-throughput genotyping platforms will expedite breeding of improved cultivars.
Early DNA markers, such as minisatellites, restriction fragment length polymorphisms (RFLPs), random amplified polymorphic DNA (RAPDs), amplified fragment length polymorphisms (AFLPs), internal transcribed spacer region (ITS), and their use in Rubus have been reviewed by others26,27. Newer markers such as simple sequence repeat (SSR), also known as microsatellites, and single nucleotide polymorphism (SNP) markers are available for Rubus. Microsatellite-enriched libraries were first used to develop a limited number of genomic SSR markers in R. alceifolius28, R. idaeus ‘Glen Moy’29,30,31 and ‘Meeker’32, R. hochstetterorum33, blackberry ‘Marion’32, R. glaucus34, and R. coreanus35.
Expressed sequence tags (ESTs) later provided a good source of EST-SSR markers and were developed from: red raspberry “Glen Moy”29, “Glen Ample”, “Latham”36, and “Heritage”;37 the black raspberry “Bristol”;38 and the blackberry “Merton Thornless”39. Next-generation sequencing was also used to develop SSR sequences from short reads of red and black raspberry40. Additional SSRs and SNP markers were also developed from transcriptome sequences of ‘Boysen’41 and “Loch Ness”42 blackberries and from candidate gene sequences43,44,45,46,47, most of which were identified from Bacterial Artificial Chromosome (BAC) sequences of “Glen Moy” red raspberry and used for linkage mapping. Microsatellite markers have been used for fingerprinting, cultivar identification, or pedigree confirmation25,32,48,49,50,51,52, assessment of genetic diversity25,53,54,55,56,57,58,59,60, in addition to linkage mapping and QTL analyses29,30,36,38,43,44,45,46,47,61,62,63,64,65,66,67,68,69,70.
QTLs/loci controlling traits of interest
Graham et al.29 created the first genetic linkage map from a “Glen Moy” × “Latham” red raspberry population, and this map has subsequently been improved using genotyping by sequencing (GBS)71. Analysis of this population has identified numerous QTLs affecting fruit ripening traits, such as fruit texture, color, anthocyanin accumulation, flavor volatiles and fruit softening29,30,43,44,47,63,64,65,72. Genes in the phenylpropanoid pathway and candidate genes in the biosynthesis of flavor volatiles have been mapped to these loci45,46,47. Two QTLs that are important for the genetic control of the crumbly fruit disorder were identified65. For a recent and comprehensive review of QTL and trait mapping, see McCallum et al.73.
The genetic regulation of annual vs biennial fruiting has been the subject of multiple studies62,74,75,76,77,78,79,80,81. Castro et al.62 reported mapping a single locus on LG7 conferring AF in blackberry. However, in this study LG7 was assigned by default and none of the LG7 markers mapped to the black raspberry genetic map of Bushakra et al.61. More recently, two additional loci conferring AF have been identified on LG3 and LG4 of red raspberry using a high-density map generated by GBS-based SNP markers (Jibran et al.89, paper submitted).
Several QTLs for cane splitting have been identified67, with two of these co-locating with a previously identified QTL for plant vigor and another associated with a QTL for resistance to root rot64. Gene H, which controls cane pubescence and is associated with resistance to cane botrytis and spur blight has been mapped to LG230. Three other loci were identified that are associated with resistance to rust and cane spot, and spine density30. Resistance to raspberry aphids has been mapped to two loci on LG3 and LG638,66. QTLs for root vigor and resistance to phytophthora root rot have been identified64 and markers from these deployed by the James Hutton Institute Raspberry Breeding Consortium. QTLs relating to physical traits that affect pest burden also have been identified63. Recently, attempts have been made to map QTLs from hyperspectral traits in an attempt to develop high-throughput phenotyping approaches82.
At least six different types of dwarf raspberry83,84 and one brachytic dwarf blackberry85 have been described. One dwarf raspberry locus has been mapped to LG666 and another to the bottom of LG2 (Jibran et al.89, unpublished data), suggesting that multiple loci can confer a dwarf habit.
The first tetraploid blackberry genetic linkage map was constructed from a full-sib family of “Prime-Jim” × “Arapaho” that was segregating for thornlessness and AF62. One hundred and nineteen SSR markers were used to create an integrated linkage map composed of seven linkage groups and to map loci for thornlessness and AF to LG4 and 7, respectively. A second tetraploid blackberry linkage map was constructed also using simplex markers consisting of restriction-site associated genomic DNA (RAD-Seq) in the tetraploid “Chester Thornless” × “Prime-Jim” population86. Parental haplotype maps for “Chester Thornless” and “Prime-Jim”, consisted of 29 linkage groups spanning 1059 cM and 31 linkage groups spanning 1025 cM, respectively; which provided supporting evidence for the position of the thornless locus on LG4. No broadly predictive markers were identified for thornlessness or AF in either of these mapping studies and molecular markers have yet to be used for any application other than parentage confirmation in applied blackberry breeding programs.
Genomes sequenced
Next-generation sequencing techniques in addition to long-read PacBio sequencing and Hi-C scaffolding have been used to generate a chromosome-scale genome assembly of a highly homozygous wild accession (ORUS 4115–3) of black raspberry87,88,89. The V3 reference genome has a contig N50 of 5.1 Mb, consists of 235 contigs that were anchored and oriented into seven chromosomes, and contains 47 Mb of new sequences including large pericentromeric regions and thousands of previously unannotated protein-coding genes.
A genome assembly of “Heritage” red raspberry has been reported90,91 but is not publicly available, while a fragmented short-read-derived draft assembly of “Glen Moy” and “Latham” was recently generated71. This draft assembly of 147,546 scaffolds covers 361,105,105 bp of the estimated 280 Mb genome. Comparison against plant near-universal single-copy orthologs using Benchmarking Universal Single-Copy Orthologs (BUSCO) indicated that over 90% of the 1440 BUSCO groups were present in the genome assembly. A new genome assembly of “Joan J” red raspberry has recently been generated using a combination of long single-molecular real-time (SMRT) PacBio reads and high coverage short reads92. This draft genome is 299 Mb, consists of 2145 scaffolds and has a BUSCO genome completeness of 95.3% and an N50 score of 638 Kb.
A team of researchers is developing reference genomes for two diploid blackberry accessions representing the sources of thornlessness (“Burbank Thornless”, R. ulmifolius inermis) and AF (“Hillquist”, R. argutus) in fresh-market blackberry breeding programs93.
High-throughput genotyping
At this time, the only high-throughput genotyping methods used in Rubus consist of reduced representation sequencing techniques such as target capture sequencing for phylogenetic analyses94, GBS for developing saturated linkage maps in black raspberry38 and in red raspberry71,90, and RAD-Seq for linkage mapping in blackberry95. Target capture was applied to 96 samples that included representatives from each of the 12 Rubus subgenera and from five known hybrids or economically important cultivars. The target capture baits included 926 single-copy loci from the R. occidentalis genome and 247 loci that are conserved among apple, peach and strawberry genomes94. Preliminary analyses indicated that the subgenus containing raspberries, Idaeobatus, was polyphylet with members occurring in five out of eight clades. The phylogenetic network identified a number of polyploid taxa as potential hybrids, indicated by their intermediate position between two major clades. “Marion”, “Logan”, and “Boysen”, which have multiple raspberry and blackberry species in their pedigrees, were located between the raspberry and blackberry clades94. The Elshire et al.96 method of digestion with ApeK1 was used by Bushakra et al.38 and Ward et al.90, while the two-enzyme method of Poland et al.97 was used by Hackett et al.71, and these resulted in highly saturated GBS-based linkage maps.
Anchoring physical and genomic maps, and synteny across Rosaceae
Linkage maps have been constructed for red raspberry29,30,36,43,44,45,46,47,63,64,65,66,68,69,70,71,90, black raspberry38, a cross between red and black raspberry61, and blackberry62,86,98. Bushakra et al.61 aligned the genetic map of the red raspberry parent, “Latham”, to BLAST-generated physical maps of F. vesca “Hawaii 4”99, Malus × domestica “Golden Delicious”100 and Prunus persica, “Lovell”101, and to the nine hypothetical Rosaceae ancestral chromosomes102, using sequence-based orthologous markers in common among them. The 1:1 collinearity of the seven “Latham” linkage groups to the seven Fragaria chromosomes led the authors to rename the groups to correspond to those used in Fragaria as Rubus Linkage Groups (RLGs) 1 through 7. This nomenclature was subsequently used in red raspberry90, in black raspberry38 and in blackberry62. Each of the seven linkage groups in Rubus were aligned to 1, 2 or 3 segments of the Malus and the Prunus genomes. Of these four Rosaceae genera, Prunus appears to show the fewest rearrangements from the proposed ancestral state.
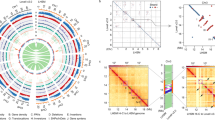
The pseudomolecules making up the V3 black raspberry genome assembly (Ro01-Ro07) were anchored to the seven haploid strawberry chromosomes (Fvb1–7) using markers from GBS-based genetic maps88 and confirmed a high degree of synteny across both genomes despite the 75 MY divergence. No major rearrangements were observed when comparing Ro01/Fvb1, Ro02/Fvb2, and Ro03/Fvb3, while the other four chromosome pairs had one or two major inversions. The black raspberry and F. vesca genomes had 15,727 syntenic gene pairs, and each genome had unique patterns of expansion/deletion based on gene-level microsynteny. VanBuren et al.88 suggested that differences in gene composition between these two species are due to a combination of tandem gene duplications, retrotransposon-mediated duplication/movement, fractionation/deletion, and mis-annotation. Most recently, genome-wide comparisons between R. idaeus, R. occidentalis, and nine other Rosaceae species have supported the high collinearity between raspberry and strawberry genomes92. Peach (Prunus persica) has slightly less collinearity with raspberry, although there are large conserved syntenic blocks (Fig. 2).
Circos plots displaying macrosynteny between the genomes of a Rubus idaeus and Rubus occidentalis, b R. idaeus and Fragaria vesca, and c R. idaeus and Prunus persica. For A to D, each connecting line represents an orthologous gene pair and the right half of each circle consists of the seven R. idaeus chromosomes colored by the spectral order in the rainbow. Reproduced with permission from Wight et al.92
Transcriptomic analysis
Eight RNAseq datasets (leaves, stems, canes, green fruit, red fruit, ripe fruit and root tissue harvested from Verticillium dahlia-treated and control plants two months after inoculation) of “Jewel” generated 28,005 protein-coding genes in the V1 black raspberry assembly87, while another eight RNAseq libraries (leaves, methyl jasmonate-treated leaves, flowers, canes, roots, green fruit, red fruit, and ripe fruit) from ORUS 4115–3 resulted in 34,545 high-confidence gene models in the V3 genome assembly88. The V3 annotation had 9301 new gene models that were improved or absent in the V1 assembly, while 4020 low-quality gene models from V1 were removed from V3 because of insufficient protein support of transposable element-related annotation. Surveying expression in ripening fruit of “Jewel” identified 4446 genes that were differentially expressed between green and ripe fruit, and 8376 genes between red and ripe fruit87. The genes that were upregulated during fruit ripening consisted of those involved in hydrolase activity, cell wall degradation, sugar transport, and anthocyanin accumulation and biosynthesis, among others. Genes involved in anthocyanin accumulation during fruit ripening were also previously identified in what was erroneously labelled R. coreanus103 and is, in fact, R. occidentalis104. A type I chalcone Isomerase, RcMCHI2 (Unigene 18325), from R. coreanus complemented an Arabidopsis testa 5–1 mutant and restored its ability to produce anthocyanin pigments in the cotyledon and hypocotyl and to accumulate delphinidin 3-O-rutinoside and cyaniding 3-O-rutinoside. Recently, differential expression was evaluated in fruit transcripts of R. coreanus from China across three developmental stages105. They identified 23 transcripts in the flavonoid biosynthesis pathways whose expression corresponded to metabolite accumulation during ripening. Seven representative genes were validated by sequencing after cloning, and their expression was confirmed by RT-qPCR. In that study, they also identified 119 nucleotide-binding site leucine-rich repeat (NBS-LRR) protein-coding genes. A red raspberry “fruit transcriptome” comprising a database of 56,000 unigenes has been established and mapped to the genome scaffolds of “Glen Moy” (Milne, personal communication).
More recently, a red raspberry transcriptome was generated from 18 RNAseq libraries derived from fruit tissues at early stages of development92. A total of 35,566 protein-coding genes were annotated. Interestingly, 775 orthogroups were limited to the Rubus genera, and these were significantly enriched with genes involved in defense and gene regulation.
In the V1 gene models of black raspberry, 144 predicted genes contained motifs that were conserved among disease resistance R genes and 10 putative homologues to the tomato Ve1 gene that confers Verticillium wilt (VW) resistance87. Up to 147 genes, including eight with homology to genes associated with disease resistance and 12 with homology to transcription vectors, were differentially expressed between V. dahlia-inoculated and control “Jewel” roots38,87. More work is needed to identify the genes responsible for VW resistance.
Resources for blackberry
Genetic and genomic resources in blackberry have been delayed by challenges including polyploidy, multisomic inheritance, and heterozygosity. The development of a chromosome-level genome assembly in black raspberry and a new sequencing initiative with two diploid relatives in the subgenus Rubus will facilitate rapid advances in blackberry. Software has recently been developed for linkage and QTL mapping106,107, association analyses108 and genomic selection109 in multisomic polyploid species. These new software packages all require large datasets of SNP markers with accurate dosage calls, e.g. AAAT, AATT, ATTT, in heterozygous individuals. Software is available for estimating allelic dosage from a fixed genotyping platform, e.g., SNP chip110,111; however, there is currently no Rubus chip. Dosage estimation is significantly more difficult in GBS datasets because of problems of missing data and uneven depth of coverage. Fortunately, GBS strategies that severely restrict genome complexity112 and sequence capture methods113 have allowed the generation of large quantities of markers with sufficient read depth to estimate allele dosage in polyploid crop species lacking fixed SNP arrays. In the near future, these new Rubus genomic resources coupled with development of new software for analyzing genomes of polyploid plants will enable blackberry researchers to execute QTL mapping and genome-wide association studies (GWAS) for important breeding traits.
Future perspectives
Driven by consumer demand for high-health and delicious flavor, berryfruit sales have increased steadily in the past decade and are projected to grow in the future. There is also an increased demand from consumers for sustainably produced, pesticide-free and locally grown fruits. While in conflict with consumers’ desire for decreased plastic use, growers worldwide are increasingly moving towards production in containers under poly-tunnel houses to lengthen the production season, to reduce water and chemical inputs, and to protect from adverse weather. These new production systems bring new challenges that will change breeding targets. Climate change creates another set of problems for some cultivars, with many areas no longer receiving sufficient or predictable winter chill. Recent advances in genomic tools for Rubus will help accelerate breeding new cultivars optimized for the changing environment93,114.
New resources could be developed that would further fast-track both breeding and basic science discovery. The Rubus germplasm is incredibly variable and offers an excellent source of new traits and resistance to pests and diseases. Molecular and phenotypic characterization of existing and novel germplasm would have enormous potential. Tools such as genomic selection have greatly facilitated introducing desirable traits from wild species of apple115,116. By coordinating all sequencing data within the Rubus community, we could work towards developing a SNP chip for high-throughput genotyping.
Gene models created by computational prediction are often incorrect. Recently, the kiwifruit (Actinidia chinensis var. deliciosa) Red5 gene models were manually annotated by a consortium of kiwifruit researchers117. It was found that 91% of the previous, computationally predicted gene models were incorrect. Moreover, the manual annotation also revealed many translocation events that followed whole-genome duplication and enabled 164 Mb of previously unassigned sequence to be placed on chromosomes. A community annotation approach for raspberry would be relatively simple given the much smaller size of the raspberry genome. Improving the gene models will help with gene identification, allele mining, and with understanding the molecular basis of specific traits.
Rubus has many features that make it an excellent model system for Rosaceae. It has a very short juvenile phase, is easy to cross, the fruit develop quickly and produce many seeds. Being diploid, raspberry is well suited for genetic studies. Raspberry and blackberry are amenable to Agrobacterium-mediated transformation, enabling testing of gene function and gene editing using CRISPR/Cas9 systems118,119.
Although Rubus has somewhat lagged behind other crops in terms of having genetic and genomic tools, we can catch up rapidly by adopting the most successful strategies and by working together as a community.
References
Gu Y., Zhao C. M., Jun W. On Rubus resources in Hunan and Fujian provinces. in XXIII International Horticultural Congress. (ISHS, International Society for Horticultural Science, Wageningen, 1990).
Gu Y., Zhao C. M., Jin W., Li W. L. Evaluation of Rubus germplasm resources in China. Acta. Hortic. 317–324 (1993).
Hummer, K. E. Rubus diversity. Hortscience 31, 182 (1996).
Focke W. O. Species ruborum. Monographiae generis rubi prodromus, Vol. pt 1–3. E. (Schweizerbart, Stuttgart, 1910).
Verma, R., Gangrade, T., Punasiya, R. & Ghulaxe, C. Rubus fruticosus (blackberry) use as an herbal medicine. Pharmacogn. Rev. 8, 101–104 (2014).
Moyer, R. A., Hummer, K. E., Finn, C. E., Frei, B. & Wrolstad, R. E. Anthocyanins, phenolics, and antioxidant capacity in diverse small fruits: vaccinium, rubus, and ribes. J. Agric Food Chem. 50, 519–525 (2002).
Siriwoharn, T., Wrolstad, R. E., Finn, C. E. & Pereira, C. B. Influence of cultivar, maturity, and sampling on blackberry (Rubus L. Hybrids) anthocyanins, polyphenolics, and antioxidant properties. J. Agric Food Chem. 52, 8021–8030 (2004).
Deighton, N., Brennan, R., Finn, C. & Davies, H. V. Antioxidant properties of domesticated and wild Rubus species. J. Sci. Food Agriculture 80, 1307–1313 (2000).
Kaume, L., Howard, L. R. & Devareddy, L. The blackberry fruit: a review on its composition and chemistry, metabolism and bioavailability, and health benefits. J. Agric. Food Chem. 60, 5716–5727 (2012). 2012/06/13.
R.D. H, S.M. H. Fruits and vegetables as functional foods for exercise and inflammation. in Bioactive foods and chronic disease states (ed Watson R. R.) (Elsevier, Oxford, 2013).
Nile, S. H. & Park, S. W. Edible berries: bioactive components and their effect on human health. Nutrition 30, 134–144 (2014).
FAOSTAT. http://www.fao.org/faostat/en/?#data/QC (2017).
Strik, B. C., Clark, J. R., Finn, C. E. & Bañados, M. P. Worldwide Blackberry Production. Food and Agriculture Organization of the United Nations. 17, 205 (2007).
California Strawberry Commission (2017). http://www.calstrawberry.com/Portals/0/Reports/Retail%20Reports/Retail%20Category%20Trends/Retail%20Category%20Trends%20-%20Total%20US.pdf?ver=2017-07-19-181746-493. [Accessed 03/03/2019].
Ahumada M. Blackberry production in Mexico. (Sun Belle Berries Cane Berry Day, San Luis Obispo, 2016). http://cesanluisobispo.ucanr.edu/files/239571.pdf.
Hall H. K. World blackberry production. in Blackberries and Their Hybrids. (eds Hall H. K., Funt R. C.) 308–314. (CABI Publishing, Boston, 2017).
Keep, E. Primocane (autumn)-fruiting raspberries: a review with particular reference to progress in breeding. J. Horticultural Sci. 63, 1–18 (1988). 1988/01/01.
Carew, J. G. et al. The control of the annual growth cycle in raspberry. J. Horticultural Sci. Biotechnol. 75, 495–503 (2000). 2000/01/01.
Sønsteby, A. & Heide, O. M. Effects of photoperiod and temperature on growth and flowering in the annual (primocane) fruiting raspberry (Rubus idaeus L.) cultivar ‘Polka’. J. Horticultural Sci. Biotechnol. 84, 439–446 (2009). 2009/01/01.
Heide O., Sønsteby A. Physiology of flowering and dormancy regulation in annual- and biennial-fruiting red raspberry (Rubus idaeus L.)—A review, vol. 86, (Journal of Horticultural Science and Biotechnology, 2011), https://doi.org/10.1080/14620316.2011.11512785.
Williams, I. H. Effects of environment on Rubus Idaeus L.: IV. Flower Initiation Dev. Inflorescence 34, 219–228pp (1959).
Williams, I. H. Effects of environment on Rubus Idaeus L. V. dormancy and flowering of the mature shoot. J. Horticultural Sci. 35, 214–220 (1960). 1960/01/01.
Hodnefjell, R., Heide, O. M., Rivero, R., Remberg, S. F. & Sønsteby, A. Control of growth cessation and floral initiation in red raspberry (Rubus idaeus L.) cultivars of diverse origin in controlled and natural environments. Sci. Horticulturae 233, 412–420 (2018). 2018/03/15/.
Gambardella M., Contreras E., Alcalde J., Neri D. Phenotyping primocane fruiting trait in raspberry (Rubus idaeus), 67–74. (International Society for Horticultural Science (ISHS), Leuven, 2016).
Dossett, M., Bassil, N. V., Lewers, K. S. & Finn, C. E. Genetic diversity in wild and cultivated black raspberry (Rubus occidentalis L.) evaluated by simple sequence repeat markers. Genet. Resour. Crop Evolution 59, 1849–1865 (2012).
Clark, J. R., Stafne, E. T., Hall H. K., Finn C. E. Blackberry breeding and genetics. In: Janick J (ed). Plant Breeding Reviews, vol. 29, pp 19–144. (John Wiley & Sons, Inc., 2007).
Antonius-Klemola, K. Molecular markers in Rubus (Rosaceae) research and breeding. J. Horticultural Sci. Biotechnol. 74, 149–160 (1999). 1999/01/01.
Amsellem, L., Noyer, J. L., Le Bourgeois, T. & Hossaert-McKey, M. Comparison of genetic diversity of the invasive weed Rubus alceifolius poir. (Rosaceae) in its native range and in areas of introduction, using amplified fragment length polymorphism (AFLP) markers. Mol. Ecol. 9, 443–455 (2000).
Graham, J. et al. The construction of a genetic linkage map of red raspberry (Rubus idaeus subsp. idaeus) based on AFLPs, genomic-SSR and EST-SSR markers. Theor. Appl. Genet 109, 740–749 (2004).
Graham, J., Smith, K., Tierney, I., MacKenzie, K. & Hackett, C. A. Mapping gene H controlling cane pubescence in raspberry and its association with resistance to cane botrytis and spur blight, rust and cane spot. Theor. Appl. Genet 112, 818–831 (2006).
Graham, J., Smith, K., Woodhead, M. & Russell, J. Development and use of simple sequence repeat SSR markers in Rubus species. Mol. Ecol. Notes 2, 250–252 (2002).
Castillo, N. R. F., Reed, B. M., Graham, J., Fernandez-Fernandez, F. & Bassil, N. V. Microsatellite markers for raspberry and blackberry. J. Am. Soc. Horticultural Sci. 135, 271–278 (2010). 2010.
Lopes, M. S., Maciel, G. B., Mendonça, D., Gil, F. S. & Machado, A. Da. C. Isolation and characterization of simple sequence repeat loci in Rubus hochstetterorum and their use in other species from the Rosaceae family. Mol. Ecol. Notes 6, 750–752 (2006).
Marulanda, M., Lopez, A. M. & Uribe, M. Molecular characterization of the Andean blackberry, Rubus glaucus, using SSR markers. Genet Mol. Res 11, 322–331 (2012).
Lee, G. A. et al. Novel microsatellite markers acquired from Rubus coreanus Miq. and cross-amplification in other Rubus species. Molecules 20, 6432–6442 (2015).
Woodhead, M., McCallum, S., Smith, K., Cardle, L., Mazzitelli, L. & Graham, J. Identification, characterisation and mapping of simple sequence repeat (SSR) markers from raspberry root and bud ESTs22, pp. 555–563 (The James Hutton Institute, 2008).
Bushakra J. M., Lewers K. S., Staton M. E., Zhebentyayeva T., Saski C. A. Developing expressed sequence tag libraries and the discovery of simple sequence repeat markers for two species of raspberry (Rubus L.). BMC Plant Biol. 15, 1–11 (2015).
Bushakra, J. M. et al. A genetic linkage map of black raspberry (Rubus occidentalis) and the mapping of Ag4conferring resistance to the aphid Amphorophora agathonica. Theor. Appl. Genet. 128, 1631–1646 (2015).
Lewers, K. S. et al. A blackberry (Rubus L.) expressed sequence tag library for the development of simple sequence repeat markers. BMC plant Biol. 8, 69–69 (2008).
Dossett, M. et al. Development and transferability of black and red raspberry microsatellite markers from short-read sequences. J. Am. Soc. Horticultural Sci. 140, 243–252 (2015). 2015.
Li, W. L., Zhang, C. H., Yang, H. Y., Wu, W. L. & Lyu, L. F. SSR marker information mining in blackberry from transcriptome sequences. Acta Horticulturae 1133, 97–101 (2016).
Garcia-Seco, D., Zhang, Y., Gutierrez-Manero, F. J., Martin, C. & Ramos-Solano, B. RNA-Seq analysis and transcriptome assembly for blackberry (Rubus sp. Var. Lochness) fruit. BMC Genomics 16, 5 (2015).
Graham, J. et al. Mapping QTLs for developmental traits in raspberry from bud break to ripe fruit. Theor. Appl. Genet. 118, 1143–1155 (2009). 2009.
Kassim, A. et al. Environmental and seasonal influences on red raspberry anthocyanin antioxidant contents and identification of quantitative traits loci (QTL). Mol. Nutr. Food Res. 53, 625–634 (2009).
McCallum, S. et al. Genetic and environmental effects influencing fruit colour and QTL analysis in raspberry. Theor. Appl Genet 121, 611–627 (2010).
Woodhead, M. et al. Functional markers for red raspberry. J. Am. Soc. Horticultural Sci. 135, 418–427 (2010). 2010.
Paterson, A. et al. Environmental and seasonal influences on red raspberry flavour volatiles and identification of quantitative trait loci (QTL) and candidate genes. Theor. Appl. Genet. 126, 33–48 (2013).
Bassil N. V., Muminova M., Njuguna W. Microsatellite-based fingerprinting of western blackberries fromplants, iqf berries and puree 73–80 (International Society for Horticultural Science (ISHS), Leuven, 2010).
Zurn, J. D. et al. Validating blackberry seedling pedigrees and developing an improved multiplexed microsatellite fingerprinting set. J. Am. Soc. Horticultural Sci. 143, 381–390 (2018).
Girichev V., Hanke M.-V., Peil A., Flachowsky H. SSR fingerprinting of a German Rubus collection and pedigree based evaluation on trueness-to type. Genet. Resour. Crop Evol. 64, 189–203 (2015).
Bassil N. V., Nyberg A., Hummer K. E., Graham J., Dossett M., Finn C. E. A universal fingerprinting set for red raspberry. p. 83–87. (International Society for Horticultural Science (ISHS), Leuven, 2012).
Fernández-Fernández, F., Antanaviciute, L., Govan, C. L. & Sargent, D. J. Development of a multiplexed microsatellite set for fingerprinting red raspberry (Rubus idaeus) germplasm and its transferability to other Rubus species. J. Berry Res. 1, 177–187 (2011).
Bradish C. M., Overbaugh E., Ballington J., Fernandez G. E., Bassil N. V. Comparative diversity analysis of southeastern Rubus germplasm through molecular and pedigree techniques. p. 157–162. (International Society for Horticultural Science (ISHS), Leuven, 2016).
Ataei-e Jaliseh, S., Mehregan, I., Tarang, A. & Nejadsattari, T. A taxonomic review of Rubus L. (Rosaceae) in the Northern Iran based on the analysis of quantitative morphological characters. J. Biodivers. Environ. Sci. 6, 113–120 (2015).
Lee, K. J. et al. Genetic diversity and population structure of rubus accessions using simple sequence repeat markers. Plant Breed. Biotechnol. 4, 345–351 (2016).
Marulanda, M., Lopez, A. & Uribe, M. Molecular characterization of the Andean blackberry, Rubus glaucus, using SSR markers. Genet Mol. Res 11, 322–331 (2012).
Ochieng J. A. Assessment of diversity of blackberry (rubus l. subgenus rubus watson) accessions in Kenya using morphological and ssr markers. (Egerton University, 2018).
Kostamo, K., Toljamo, A., Antonius, K., Kokko, H. & Kärenlampi, S. Morphological and molecular identification to secure cultivar maintenance and management of self-sterile Rubus arcticus. Ann. Bot. 111, 713–721 (2013).
Rusu A. R., Graham J., Pamfil D., Vidican R. Identification of Rubus accessions in Romania and a comparison of their relatedness to European and North American cultivars. ProEnvironment Promediu 9, 46–52 (2016).
Wang Y., He W., Zhang J., Chen T., Chen Q., Tang H.-R. et al. Genetic Relationship between Rubus Parvifolius and R. Coreanus (Rubus, Rosaceae) based on Simple Sequence Repeat Markers. in Proc. 2017 2nd International Conference on Biological Sciences and Technology (BST 2017) (Atlantis Press, 2018).
Bushakra, J. M. et al. Construction of black (Rubus occidentalis) and red (R. idaeus) raspberry linkage maps and their comparison to the genomes of strawberry, apple, and peach. Theor. Appl. Genet. 125, 311–327 (2012). 2012.
Castro, P., Stafne, E. T., Clark, J. R. & Lewers, K. S. Genetic map of the primocane-fruiting and thornless traits of tetraploid blackberry. Theor. Appl. Genet. 126, 2521–2532 (2013).
Graham, J. et al. Genetic and environmental regulation of plant architectural traits and opportunities for pest control in raspberry. Ann. Appl. Biol. 165, 318–328 (2014). 2014.
Graham, J. et al. Towards an understanding of the nature of resistance to Phytophthora root rot in red raspberry. Theor. Appl. Genet. 123, 585–601 (2011).
Graham, J. et al. Towards an understanding of the control of ‘crumbly’ fruit in red raspberry. Springerplus 4, 223 (2015).
Sargent, D. J. et al. Mapping of A1 conferring resistance to the aphid Amphorophora idaei and dw (dwarfing habit) in red raspberry (Rubus idaeus L.) using AFLP and microsatellite markers. BMC Plant Biol. 7, 15 (2007).
Woodhead, M. et al. Identification of quantitative trait loci for cane splitting in red raspberry (Rubus idaeus). Mol. Breed. 31, 111–122 (2013).
Pattison, J. A., Samuelian, S. K. & Weber, C. A. Inheritance of Phytophthora root rot resistance in red raspberry determined by generation means and molecular linkage analysis. Theor. Appl. Genet. 115, 225–236 (2007).
Molina-Bravo, R., Fernandez, G. & Sosinski, B. Quantitative trait locus analysis of tolerance to temperature fluctuations in winter, fruit characteristics, flower color, and prickle-free canes in raspberry. Mol. Breed. 33, 267–280 (2014).
Simpson, C. G. et al. Mapping and expression of genes associated with raspberry fruit ripening and softening. Theor. Appl. Genet. 130, 557–572 (2017).
Hackett, C. A. et al. Enhancement of Glen Moy x Latham raspberry linkage map using GbS to further understand control of developmental processes leading to fruit ripening. BMC Genet. 19, 59 (2018).
Dobson, P. et al. Over-seasons analysis of quantitative trait loci affecting phenolic content and antioxidant capacity in raspberry. J. Agric. food Chem. 60, 5360–5366 (2012).
McCallum S., Simpson C., Graham J. QTL Mapping and Marker Assisted Breeding in Rubus spp. In: Graham J., Brennan R. (eds). Raspberry: Breeding, Challenges and Advances. pp 121–144. (Springer International Publishing: Cham, 2018).
Haskell, G. The raspberry wild in Britain. Watsonia 4, 238–255 (1960).
Slate G. L. Breeding autumn-fruiting raspberries. in Proc. Proceedings of the American Society for Horticultural Science; 1940. p. 574–578. (1940).
Oberle G., Moore R. Transmission of the autumn-fruiting character in crosses of red and black raspberries. in Proc. American Society for Horticultural Science; 1952 . p. 235–237 (American Society for Horticultural Science, Alexandria, 1952).
Ourecky, D. K. Fall-bearing red raspberries, their future and potential. Symp . Breed. Mach. Harvesting Rubus Ribes 60 1976, 135–144 (1976).
Fejer, S. Inheritance of yield, yield components, and fall-fruiting habit in red raspberry diallel crosses. Can. J. Genet. Cytol. 19, 1–13 (1977).
Waldo, G. F. & Darrow, G. M. Breeding autumn-fruiting raspberries under Oregon conditions. Proc. Am. Soc. Horticultural Sci. 1941, 263–267 (1941).
Barrientos F., Rodriguez J. A. Transgressive segregation for winter chilling requirement in the red raspberry cultivar Malling Exploit. in Symposium on Breeding and Machine Harvesting of Rubus, Vol 112, 21–24 (1980).
Lewis, D. Genetical studies in cultivated raspberries. J. Genet. 38, 367–379 (1939).
Williams D., Aitkenhead M., Karley A., Graham J., Jones H. Use of Imaging Technologies for High Throughput Phenotyping: Breeding, Challenges and Advances. pp 145–158 (2018).
Jennings, D. L Raspberries and blackberries: their breeding, diseases and growth. (Academic Press, London, 1988).
Keep, E. Dwarfing in the raspberry, Rubus idaeus L. Euphytica 18, 256–276 (1968).
Worthington MLaJRC. Development of blackberry cultivars with novel plant architecture. Acta Horticulturae. (2019) (In Press).
Weber D. Linkage mapping in tetraploid blackberry (Rubus spp.) using high throughput genomic sequencing and restriction site associated DNA sequencing (RAD-SEQ). (University of Illinois at Urbana-Champaign, 2014).
VanBuren, R. et al. The genome of black raspberry (Rubus occidentalis). Plant J. 87, 535–547 (2016).
VanBuren R. et al. A near complete, chromosome-scale assembly of the black raspberry (Rubus occidentalis) genome. Gigascience 7, 1–9 (2018).
Jibran, R. et al. Chromosome-scale scaffolding of the black raspberry (Rubus occidentalis L.) genome based on chromatin interaction data. Hortic. Res. 5, 8 (2018).
Ward J. A. et al. Saturated linkage map construction in Rubus idaeus usinggenotyping by sequencing and genome-independent imputation. BMC Genomics. 14, 1–14 (2013).
Longhi, S. et al. Molecular genetics and genomics of the Rosoideae: state of the art and future perspectives. Hortic. Res. 1, 1 (2014).
Wight H., Zhou J., Li M., Hannenhalli S., Mount S., Liu Z. Draft genome assembly and annotation of red raspberry Rubus Idaeus. bioRxiv. 1–22 (2019).
Worthington M. L. et al. Development of new genomic resources and tools for molecular breeding in blackberry. Acta Horticulturae (2019) (in press).
Carter K. et al. Increased phylogenetic resolution using target enrichment in Rubus. (American Society for Horticultural Science, Waikoloa, 2017).
Weber D. Linkage mapping in tetraploid blackberry (Rubus spp.) using high-throughput genomic sequencing and restriction-site associated DNA sequencing (RAD-seq). (University of Illinois at Urbana-Champaign, 2014).
Elshire, R. J. et al. A robust, simple genotyping-by-sequencing (GBS) approach for high diversity species. Plos ONE 6, e19379 (2011).
Poland, J. A., Brown, P. J., Sorrells, M. E. & Jannink, J.-L. Development of high-density genetic maps for barley and wheat using a novel two-enzyme genotyping-by-sequencing approach. PloS ONE 7, e32253 (2012).
Castro, P., Stafne, E. T., Clark, J. R. & Lewers, K. S. Genetic map of the primocane-fruiting and thornless traits of tetraploid blackberry. Theor. Appl. Genet. 126, 2521–2532 (2013).
Shulaev, V. et al. The genome of woodland strawberry (Fragaria vesca). Nat. Genet 43, 109–116 (2011).
Velasco, R. et al. The genome of the domesticated apple (Malus x domestica Borkh). Nat. Genet 42, 833–839 (2010).
Sosinski B., Verde I., Morgante M., Rokhsar D. The international peach genome initiative. A first draft of the peach genome sequence and its use for genetic diversity analysis in peach. in Proc. 5th International Rosaceae genomics conference; 2010 (2010).
Illa, E. et al. Comparative analysis of rosaceous genomes and the reconstruction of a putative ancestral genome for the family. BMC Evolut. Biol. 11, 9 (2011).
Hyun, T. K. et al. De-novo RNA sequencing and metabolite profiling to identify genes involved in anthocyanin biosynthesis in Korean black raspberry (Rubus coreanus Miquel). PLoS ONE 9, e88292 (2014).
Lee, J., Dossett, M. & Finn, C. Mistaken identity: clarification of Rubus coreanus Miquel (Bokbunja). Molecules 19, 10524–10533 (2014).
Chen, Q. et al. Transcriptomic profiling of fruit development in black raspberry Rubus coreanus. Int. J. genomics 2018, 8084032 (2018).
Hackett, C. A., Boskamp, B., Vogogias, A., Preedy, K. F. & Milne, I. TetraploidSNPMap: software for linkage analysis and QTL mapping in autotetraploid populations using SNP dosage data. J. Heredity 108, 438–442 (2017).
Bourke, P. M. et al. polymapR—linkage analysis and genetic map construction from F1 populations of outcrossing polyploids. Bioinformatics 34, 3496–3502 (2018).
Rosyara U. R., De Jong W. S., Douches D. S., Endelman J. B. Software for Genome-Wide Association Studies in Autopolyploids and Its Application to Potato. Plant Genome 9, 1–10 (2016).
Endelman, J. B. et al. Genetic variance partitioning and genome-wide prediction with allele dosage information in autotetraploid potato. Genetics 209, 77 (2018).
Schmitz Carley, C. A. et al. Automated tetraploid genotype calling by hierarchical clustering. Theor. Appl. Genet. 130, 717–726 (2017).
Voorrips, R. E., Gort, G. & Vosman, B. Genotype calling in tetraploid species from bi-allelic marker data using mixture models. BMC Bioinforma. 12, 172 (2011).
Wadl, P. A. et al. Genetic diversity and population structure of the USDA sweetpotato (Ipomoea batatas) germplasm collections using GBSpoly. Front Plant Sci. 9, 1166–1166 (2018).
JGAML, Uitdewilligen et al. A next-generation sequencing method for genotyping-by-sequencing of highly heterozygous autotetraploid potato. PLOS ONE 8, e62355 (2013).
Jennings S. N. Advances in Rubus Breeding. in Raspberry: Breeding, Challenges and Advances (eds Graham J., Brennan R.) pp 17–28. (Springer International Publishing, Cham, 2018).
Kumar, S. et al. Genomic selection for fruit quality traits in apple (Malus×domestica Borkh.). PLOS ONE 7, e36674 (2012).
Kumar, S., Bink, M. C. A. M., Volz, R. K., VGM, Bus & Chagne, D. Towards genomic selection in apple (Malus x domestica Borkh.) breeding programmes: prospects, challenges and strategies. Tree Genet. Genomes 8, 1–14 (2012).
Pilkington, S. M. et al. A manually annotated Actinidia chinensis var. chinensis (kiwifruit) genome highlights the challenges associated with draft genomes and gene prediction in plants. BMC Genomics 19, 257 (2018).
Mathews, H., Wagoner, W., Cohen, C., Kellogg, J. & Bestwick, R. Efficient genetic transformation of red raspberry, Rubus ideaus L. Plant Cell Rep. 14, 471–476 (1995).
Ma, X. et al. A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol. Plant 8, 1274–1284 (2015).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Foster, T.M., Bassil, N.V., Dossett, M. et al. Genetic and genomic resources for Rubus breeding: a roadmap for the future. Hortic Res 6, 116 (2019). https://doi.org/10.1038/s41438-019-0199-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41438-019-0199-2
This article is cited by
-
Genome-Wide Characterization and Expression of the bZIP Family in Black Raspberry
Journal of Plant Growth Regulation (2024)
-
Genetic diversity and population relationships in wild Korean black raspberry (Rubus coreanus Miq.) based on microsatellite markers: establishing a fruit tree breeding strategy
Horticulture, Environment, and Biotechnology (2024)
-
Untapped potential of non-conventional rubus species: bioactivity, nutrition, and livelihood opportunities
Plant Methods (2023)
-
Construction and genetic characterization of an interspecific raspberry hybrids panel aiming resistance to late leaf rust and adaptation to tropical regions
Scientific Reports (2023)
-
In vitro plant regeneration of ‘Prelude’ red raspberry (Rubus idaeus L.)
In Vitro Cellular & Developmental Biology - Plant (2023)