Abstract
Horticultural crops provide humans with many valuable products. The improvement of the yield and quality of horticultural crops has been receiving increasing research attention. Given the development and advantages of genome-editing technologies, research that uses genome editing to improve horticultural crops has substantially increased in recent years. Here, we briefly review the different genome-editing systems used in horticultural research with a focus on clustered regularly interspaced palindromic repeats (CRISPR)/CRISPR-associated 9 (Cas9)-mediated genome editing. We also summarize recent progress in the application of genome editing for horticultural crop improvement. The combination of rapidly advancing genome-editing technology with breeding will greatly increase horticultural crop production and quality.
Similar content being viewed by others
Introduction
As an important branch of agriculture, horticulture originated thousands of years ago and has developed greatly during the course of human history. Horticultural crops are generally considered to include vegetable and fruit crops as well as floricultural and ornamental plants, which are cultivated for food, for nutritional and medical use, and for esthetic enjoyment1. Vegetable and fruit crops are low in calories but contain high levels of vitamins and minerals2, making them indispensable for balancing our daily diet. Although the supply of horticultural products is increasing, the diversity and nutritional value of the products are decreasing3. These decreases can be partially attributed to the narrow genetic diversity of horticultural crops resulting from domestication and breeding as well as reproductive barriers that inhibit genetic introgression from wild relatives. Therefore, the generation of genetic resources with diverse and desirable characteristics will be of great value for improving horticultural products.
Thousands of years ago, humans began to improve crops by introducing new traits from crossable relatives. The essential goal of this process was the transfer of desirable genetic variations. As late as 1930s, the available variations were generated solely through natural or spontaneous processes. Breeders subsequently learned to produce mutants by using chemical mutagens or radiation4. Both spontaneous and induced mutations have significantly increased crop yield and quality5. Given the rareness and randomness of these mutations, however, obtaining suitable materials for crop improvement has proven to be laborious and time consuming4.
With the rapid progress in molecular biology, DNA sequence-specific manipulation has become a powerful tool. In 1987, several animal scientists invented gene-targeting technology that relies on homologous recombination (HR). This innovative technology enabled researchers to precisely edit (though with a low frequency) an endogenous gene after introducing a donor template into mouse embryonic stem cells6,7. Similar progress was subsequently reported by plant researchers, but with an extremely low editing frequency of 0.5–7.2 × 10−4 8,9. DNA double-stranded breaks (DSBs), which commonly result in HR in meiotic chromosomes10, were later used to increase the HR frequency in gene targeting11. In addition to HR, DSBs can be repaired through the error-prone nonhomologous end-joining (NHEJ) pathway in somatic cells, which can generate mutations via the small deletions or insertions that occur at a break site12. Scientists have used the following kinds of engineered endonucleases to introduce site-specific DSBs: meganucleases (MNs), zinc finger nucleases (ZFNs), transcription activator-like effector nucleases (TALENs), clustered regularly interspaced short palindromic repeats (CRISPR)/CRISPR-associated 9 (Cas9), and CRISPR from Prevotella and Francisella 1 (CRISPR/Cpf1). These engineered endonucleases have enabled genome editing in various biological systems13,14,15,16.
With the advent of CRISPR/Cas9, the application of genome editing to horticultural crops has greatly advanced. In this review, we first introduce and compare the engineered nucleases that are used for genome editing. We then consider their current applications in horticulture. Finally, we discuss the implications and challenges of genome editing for the improvement of horticultural crops.
Genome-editing systems
Sequence-specific DNA binding, such as the interaction between a transcription factor and a promoter, is a common phenomenon. For genome editing, the previously mentioned nucleases can target specific sequences to generate DSBs under the guidance of protein–DNA interaction (for MNs, ZFNs, and TALENs) or RNA–DNA base-pairing (for CRISPR/Cas9 and CRISPR/Cpf1)16,17.
Meganucleases or homing nucleases
The first class of nucleases for genome editing, MNs or homing endonucleases, was discovered in the genomes of microorganisms or organelles. By recognizing DNA sequence elements ranging from 12 to 40 bp, these nucleases cut both strands of DNA in a site-specific manner (Fig. 1a)18. Among MNs, the I-CreI protein has received the most research attention and has been reported to be effective in maize19, but the rare occurrence of recognizable sites limits the ability of I-CreI and other MNs to edit desired target sites17. To broaden the application of MNs, researchers have used mutagenesis or combinatorial assembly to produce MN variants that target the desired DNA sequence20,21. Nevertheless, the overlapping recognition and catalytic domains of modified MNs cause difficulties and often compromise their catalytic activity15. For these reasons, MNs have not been widely used by plant scientists.
a A meganuclease can recognize a DNA sequence element of 12–40 bp and cut both strands at specific sites, forming sticky double-stranded breaks (DSBs). b In ZFNs, each zinc finger recognizes a 3-bp DNA sequence. Target specificity is achieved by arrays of several zinc fingers. Each DNA strand is bound by one zinc finger array linked with FokI, which in dimer form cuts DNA strands. c In TALENs, the central binding domain of each TALE consists of 13–28 repeats. Each repeat (a highly conserved sequence of 34 amino acids) can recognize and bind one nucleotide through the variable di-residues at the 12th and 13th positions. Paired TALENs lead to the dimerization of FokI, and the dimers cut the DNA stands, forming sticky DSBs at the target site. d In the CRISPR/Cas9 system, a single guide RNA (sgRNA) pairs with the target sequence upstream of a 5′-NGG-3′ PAM motif (N=A, T, C or G). The Cas9 endonuclease cuts the noncomplementary and complementary DNA strands at a location 3 nucleotides upstream of the PAM motif with RuvC and HNH domains, respectively. The cutting forms a blunt end DSB. e In the CRISPR/Cpf1 system, target specificity is achieved by the pairing of crRNA with the DNA strand downstream of a 5′-TTN-3′ PAM motif. The Cpf1 endonuclease uses the RuvC and Nuc domains to cut noncomplementary and complementary DNA strands at different positions, producing DSBs with sticky ends
ZFNs and TALENs
As suggested by their names, ZFNs or TALENs are generated by fusing the DNA cleavage domain of the endonuclease FokI with zinc fingers (ZFs) or with transcriptional activator-like effectors (TALEs). The FokI endonuclease domain mediates independent and nonspecific DNA cleavage upon dimerization and is not involved in any sequence recognition22. Therefore, a pair of ZFs or TALEs, each fused with a FokI endonuclease domain, is designed to achieve site-specific cleavage23,24,25. ZFs are found in transcription factors, with each finger domain recognizing three specific nucleotides. ZFNs typically exhibit an array of 3 or 4 finger domains, which can recognize 18–24 bp sequences when a ZFN occurs as a dimer23,25. Many studies have been conducted to improve ZFN applicability, efficiency, and precision26,27, but there are still concerns about interference from neighboring finger domains and the limited number of recognition sites (Fig. 1b)15.
In contrast to ZFNs, TALENs achieve sequence specificity via the customizable DNA-binding domains of TALEs, which are proteins excreted by the common bacterial plant pathogen Xanthomonas28. During pathogenesis, TALEs bind to a specific sequence of plant promoters to activate gene expression to facilitate infection28. The central binding domain of TALEs consists of 13–28 repeat sequences. Each repeat, which encodes a highly conserved sequence of 34 amino acids, can recognize and bind to one nucleotide through the variable di-residues at the 12th and 13th positions29,30,31. Such one-to-one pairing, together with the negligible context dependency on neighboring repeats, enables TALENs to target desired sequences (Fig. 1c)32,33. In general, TALENs outperform ZFNs in terms of precision and accessibility.
CRISPR/Cas9 and CRISPR/Cpf1
Unlike ZFN and TALEN systems, which depend on protein–DNA binding specificity, the CRISPR system relies on RNA–DNA binding to achieve sequence specificity. During the functional elucidation of the CRISPR/Cas system, its involvement in bacterial resistance to viruses was experimentally demonstrated34, and several components, including crRNA, PAM motif, and tracrRNA, were discovered to be necessary for this system35,36,37. More interestingly, reconstructed key components of the CRISPR/Cas9 system can introduce DSBs in a site-specific way, suggesting the potential use of this programmable RNA-guided CRISPR/Cas9 system for genome editing in organisms other than bacteria38,39. This possibility was soon demonstrated in human and mouse cells40,41,42, zebrafish43, and plants44,45,46,47,48. In the system, site-specific binding to the target is achieved via RNA-DNA pairing of a 20-nt sequence in the chimeric single-guide RNA (sgRNA) with the target. The other crRNA- and tracrRNA-derived sequences also interact with the target to form an RNA:DNA heteroduplex that is recognized by the collective interactions of several Cas9 domains: PI, REC1, RuvC, and NUC. Thereafter, the RuvC and HNH domains cut the noncomplementary and complementary DNA strands at a location 3 nucleotides upstream of the PAM motif, respectively (Fig. 1d). The recognizable PAM motif of Cas9 is 5′-NGG-3′ (N=A, T, C, or G), and this G-rich feature prevents the design of sgRNAs in T-rich regions49.
Cpf1, another endonuclease in the class 2 Type V CRISPR system, has also been found to be efficient in plant genome editing50 and to present unique features51. First, Cpf1 does not require an additional tracrRNA to form a mature crRNA. Second, unlike Cas9, which recognizes G-rich PAM sequences, Cpf1 recognizes T-rich PAM sequences. Finally, whereas cutting by the Cas9 endonuclease produces blunt ends, cutting by the Cpf1 endonuclease produces cohesive ends (Fig. 1e). In addition to causing site-specific mutations, CRISPR genome-editing systems can be used to achieve gene regulation52,53 through the manipulation of the nuclease-inactivated Cas9 (dCas9).
Each of the endonucleases used for genome editing has unique properties because of differences in their underlying mechanisms (Fig. 1 and Table 1, Zhang et al.16,54; Knott and Doudna55). In addition to generating indel mutations at target sequences, CRISPR/Cas systems have been adapted for precise base editing56,57,58,59. Base editors usually consist of an sgRNA-guided Cas9 nickase (nCas9) fused with a deaminase that causes C to T or A to G base conversions. These resources greatly increase the versatility of the tools that can be used for precise manipulation of horticultural crops.
Current status of genome editing in horticultural crops
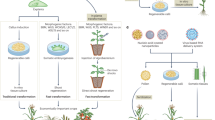
To obtain genetic resources with diverse characteristics for breeding, both spontaneous and induced mutations have been commonly used60. The rareness and uncertainty of these mutations have motivated scientists to find ways to introduce precise mutations at target sites15,17. Recently, most genome-editing studies on plants have been carried out in model systems and staple crops44,45,46, but the application of genome editing to horticultural crops is rapidly increasing61. In 2013, the first example of genome editing in a horticultural crop was achieved via a TALEN in Brassica oleracea62. In the following years, the number of studies involving genome editing in horticulture has exponentially increased (Fig. 2a, Table 2), and CRISPR-based systems now dominate. The functions of genes targeted by genome editing are very diverse, but researchers have focused most on targets affecting development, followed by targets affecting metabolism and stress responses. In addition, studies that focus on the improvement of the CRISPR/Cas9 system in horticultural crops frequently use marker/reporter genes as targets such as phytoene desaturase (PDS), whose mutation results in an albino phenotype (Fig. 2b). Among horticultural crops, tomato has received much more attention regarding genome editing than other crops: ~42% of genome-editing studies have involved tomato, whereas ~13% have involved potato. Although most (72%) genome editing with horticultural crops is performed in vegetables (Fig. 2c), some floral and medicinal plants have also been successfully manipulated by genome editing (Fig. 2c).
The information used in this figure was retrieved through May 31 of 2019. According to the information from https://aps.dac.gov.in/Public/Crops.pdf, horticultural crops include vegetables, fruits, florals, and medicinal plants. a The number of research articles involving the editing of horticultural crops with ZFNs, TALENs, and CRISPR/Cas9 from 2013 to 2019 (only the first 5 months). b The number of research articles in which the edited genes were mainly associated with development, metabolism, stress tolerance and other functions. c The number of research articles involving gene editing of different kinds of horticultural crops
In tomato, development-related genes have been edited to manipulate flowering patterns and fruit development. The tomato BLADE-ON-PETIOLE (BOP) genes, which encode transcriptional cofactors, can regulate inflorescence structure, and knock-out of SlBOP genes by gene editing reduces the number of flowers per inflorescence63. CRISPR/Cas9-induced mutations in the flowering repressor self-pruning 5G lead to rapid flowering and early harvest64. In addition, editing of the cis-regulatory region of SlCLV365 or the coding regions of SlDML266, SlORRM467 and the RIN locus68 alters fruit development and ripening. Interestingly, multiplex targeting of several genes that are important for tomato domestication was found to greatly alter the properties of the wild tomato relative Solanum pimpinellifolium such that the generated mutants were similar to cultivated tomato69,70. In potato, when the vacuolar invertase gene was disrupted by TALEN, the cold storage and processing of tubers were improved71. Another recent study in potato showed the possibility of overcoming self-incompatibility by editing the S-RNase gene, which would provide an alternative method of propagation through seeds72. In addition to tomato and potato, other horticultural crops have also been edited to obtain desirable traits. Genes related to resistance to plant pathogens such as Xanthomonas citri73,74 and Botrytis cinerea75 have been manipulated in citrus, apple, and grape. In oilseed crops, genes involved in fatty acid metabolism have been frequently targeted to improve oil quality76,77,78,79. The application of genome editing to improve crops is based on knowledge of the association between genes and their controlled traits. In the future, functional characterization of genes in different crops will help to identify valuable targets that could be edited for potential horticultural improvement, such as increased productivity, marketing quality, and nutritional value.
Possible implications of genome editing in horticulture
The goal of breeding is to harness genetic variations to introduce desirable traits. These genetic variations can arise in various ways, such as by spontaneous mutation, chemical mutagenesis, and physical mutagenesis. Gene editing could be regarded as biological mutagenesis. In comparison with other approaches, genome-editing technology is superior in terms of versatility, efficiency, and specificity. For instance, CRISPR-based genome editing can cause many types of mutations in target sequences, including small insertions/deletions, deletions of large fragments, gene replacement, and precise base substitutions16. In addition, genome-editing technology is continuously advancing: the endonuclease Cpf151 and newly discovered or designed Cas9 variants80,81 can recognize different PAM sequences, thereby broadening the genome-wide sites that can be targeted for editing.
Genome-edited plants are not considered genetically modified organisms (GMOs) in countries such as the U.S. and Japan but are still under strict GMO regulation in Europe. The largest difference between genome-edited plants and GMOs is that the genomes of edited plants can be free of exogenous DNA sequences. The exogenous DNA of the editing tools can be removed through genetic segregation82 or may never have to be introduced if CRISPR reagents are delivered as ribonucleoproteins83,84.
Mutants generated via genome editing can be directly used for crop production or as prebreeding materials. Through genome editing, desirable traits can be directly introgressed into elite or heirloom lines without compromising other properties, and the resulting lines with targeted improvement will be ready for use in production. The wild relatives of cultivated varieties are also potential materials for genome editing because they generally present unique features in many important traits. For instance, wild species of cultivated tomato are more resistant to unfavorable environments than commercial cultivars85. Wild Solanum pimpinellifolium was recently domesticated by the editing of several important genes affecting plant architecture and fruit development, resulting in new tomato varieties with the desirable properties of cultivated tomato combined with the favorable traits of the wild species69,70. Mutations can generally be introduced in either the coding region or the cis-regulatory region of the targeted gene, and mutations in the cis-regulatory region could be used to generate quantitative variation for breeding selection. In tomato, for example, fruit locule number is determined by several naturally occurring mutations in the cis-regulatory regions of CLAVATA-WUSCHEL65. This finding motivated researchers to design a multiplexed CRISPR/Cas9 system targeting the CLAVATA-WUSCHEL promoters to generate tomato lines with a wide range of locule numbers. Quantitative variations have also been observed when the genes responsible for inflorescence and plant architecture are engineered65. In addition to regulating gene activity by editing the DNA sequence of the cis-regulatory region, gene activity can be regulated by the its epigenetic status of this region. By integrating genome editing (CRISPR/Cas9) with epigenetic regulation, researchers are able to target a gene of interest and modify its epigenetic status. For instance, an sgRNA-guided fusion protein between the dead Cas9 (dCas9) variant and the catalytic domain of the TEN-ELEVEN TRANSLOCATION1 (TET1cd) demethylase can remove 5mC at specific sites, thereby increasing gene expression86. An epigenetic mutant can also be crossed with the corresponding wild type to generate epigenetic recombinant inbred lines (epiRILs). Individuals from these populations are genetically identical but epigenetically distinct. Such populations have been constructed in Arabidopsis and exhibit considerable phenotypic variations87,88,89,90. These examples demonstrate that genome editing is an excellent tool for producing new alleles and epialleles, which are important sources of phenotypic variation for crop improvement.
Challenges and future perspectives for the improvement of horticultural crops through genome editing
Although genome editing has many advantages over conventional crop breeding, some challenges remain for its application to horticultural crops. In horticultural crops, molecular and genetic studies are difficult, which hinders the identification of genes responsible for desirable traits. Sequencing the genomes of horticultural crops of interest will be important for identifying genes associated with desirable traits. For crops lacking a reference genome, the target sequence could be cloned by using degenerate primers designed for conserved protein motifs with putative functions related to desirable traits. A good example is the mildew-resistance locus (MLO), which has been characterized in detail in barley91; the phylogenetically conservative nature of the MLO has facilitated the generation of powdery mildew-resistant plants in wheat, tomato, and strawberry92,93.
Once a gene to be edited has been identified, researchers must take into account the methods used to deliver editing reagents and the procedure for regenerating the edited mutants. To date, more than 25 horticultural plant species have been successfully edited (Table 2), usually with editing reagents delivered via Agrobacteria or virus systems, and the edited plants are regenerated via in vitro tissue culture. Although tissue culture-based transformation and regeneration is most widely used for genome editing, no well-established protocol for transformation and regeneration from tissue culture is available for many horticultural crops. In planta transformation, which is an alternative to in vitro tissue culture-based Agrobacterium transformation, refers to the infection of in vivo explants in which the targeted tissues are apical or auxiliary meristems, stigmas, pollens, or inflorescences94. This method has been successfully used to transform tomato95 and Brassica species96 and should be further explored for use in horticultural crops that are recalcitrant to traditional genetic transformation. Additionally, successful genetic transformation of horticultural crops requires the consideration of editing efficiency, which is affected by many factors, such as sgRNA number and GC content, the expression levels of sgRNA and Cas9, and the secondary structure of the paired sgRNA and target sequence97,98. In the future, the editing system should be further optimized in different crop species.
The elimination of foreign DNA fragments (transferred T-DNAs) to obtain transgene-free edited plants remains difficult in some highly heterozygous and clonally propagated horticultural species99, such as potato, sweet potato, and banana. One possibility is to generate many transformants, followed by high-throughput screening of transgene-free mutants100. This approach has been used to generate ~10% of mutants without foreign DNA100,101. Another approach for transgene-free genome editing is to deliver editing reagents as in vitro transcripts102 or ribonucleoproteins83,84.
In conclusion, mutagenesis via genome editing outperforms spontaneous and induced mutations in terms of precision and efficiency. Although this technology is being increasingly used in many crops, its widespread use in the breeding of horticultural crops will require three challenges to be surmounted. First, clear breeding traits of the horticultural crop in question should be identified via communication among consumers, breeders, and biologists. Second and third, suitable methods must be developed for delivering editing reagents and for subsequently regenerating mutants. Given the great potential of genome editing and the importance of horticultural crops, we expect that these challenges will be overcome in the near future.
References
Ravichandra, N. G. Horticulture and its role in the national economies. Horticultural Nematology (pp. 1–3. Springer, India, 2014).
Janick, J. Horticultural plant breeding: past accomplishments, future directions. Acta Hortic. 694, 61–65 (2005). https://doi.org/10.17660/ActaHortic.2005.694.6.
Khoury, C. K. et al. Increasing homogeneity in global food supplies and the implications for food security. Proc. Natl Acad. Sci. USA 111, 4001–4006 (2014).
Oladosu, Y. et al. Principle and application of plant mutagenesis in crop improvement: a review. Biotechnol. Biotechnol. Equip. 30, 1–16 (2016).
Ahloowalia, B. S. & Maluszynski, M. Induced mutations – A new paradigm in plant breeding. Euphytica 118, 167–173 (2001).
Thomas, K. R. & Capecchi, M. R. Site-directed mutagenesis by gene targeting in mouse embryo-derived stem cells. Cell 51, 503–512 (1987).
Doetschman, T. et al. Targetted correction of a mutant HPRT gene in mouse embryonic stem cells. Nature 330, 576–578 (1987).
Paszkowski, J., Baur, M., Bogucki, A. & Potrykus, I. Gene targeting in plants. EMBO J. 7, 4021–4026 (1988).
Hanin, M. et al. Gene targeting in Arabidopsis. Plant J. 28, 671–677 (2002).
Gothwal, S. K. et al. The double-strand break landscape of meiotic chromosomes is shaped by the Paf1 transcription elongation complex in Saccharomyces cerevisiae. Genetics 202, 497–512 (2016).
Puchta, H., Dujon, B. & Hohn, B. Homologous recombination in plant cells is enhanced by in vivo induction of double strand breaks into DNA by a site-specific endonuclease. Nucleic Acids Res. 21, 5034–5040 (1993).
Puchta, H. The repair of double-strand breaks in plants: mechanisms and consequences for genome evolution. J. Exp. Bot. 56, 1–14 (2005).
Zhu, C. et al. Characteristics of genome editing mutations in cereal crops. Trends Plant Sci. 22, 38–52 (2017).
Puchta, H. & Fauser, F. Gene targeting in plants: 25 years later. Int. J. Dev. Biol. 57, 629–637 (2013).
Voytas, D. F. Plant genome engineering with sequence-specific nucleases. Annu. Rev. Plant Biol. 64, 327–350 (2013).
Zhang, H. et al. Genome editing-principles and applications for functional genomics research and crop improvement. Crc. Crit. Rev. Plant Sci. 36, 291–309 (2017).
Rocha-Martins, M., Cavalheiro, G. R., Matos-Rodrigues, G. E. & Martins, R. A. P. From gene targeting to genome editing: Transgenic animals applications and beyond. Acad. Bras. Cienc. 87, 1323–1348 (2015).
Jurica, M. S., Monnat, R. J. Jr. & Stoddard, B. L. DNA Recognition and Cleavage by the LAGLIDADG Homing Endonuclease I-Cre I. Mol. Cell 2, 469–476 (1998).
Gao, H. et al. Heritable targeted mutagenesis in maize using a designed endonuclease. Plant J. 61, 176–187 (2010).
Seligman, L. M. et al. Mutations altering the cleavage specificity of a homing endonuclease. Nucleic Acids Res. 30, 3870–3879 (2002).
Arnould, S. et al. Engineering of large numbers of highly specific homing endonucleases that induce recombination on novel DNA targets. J. Mol. Biol. 355, 443–458 (2006).
Durai, S. et al. Zinc finger nucleases: Custom-designed molecular scissors for genome engineering of plant and mammalian cells. Nucleic Acids Res. 33, 5978–5990 (2005).
Wright, D. A. et al. High-frequency homologous recombination in plants mediated by zinc-finger nucleases. Plant J. 44, 693–705 (2005).
Christian, M. et al. Targeting DNA double-strand breaks with TAL effector nucleases. Genetics 186, 756–761 (2010).
Bibikova, M. Enhancing gene targeting with designed zinc finger nucleases. Science 300, 764–764 (2003).
Sander, J. D. et al. Selection-free zinc-finger-nuclease engineering by context-dependent assembly (CoDA). Nat. Methods 8, 67–69 (2011).
Maeder, M. L. et al. Rapid "open-source" engineering of customized zinc-finger nucleases for highly efficient gene modification. Mol. Cell 31, 294–301 (2008).
Kay, S., Hahn, S., Marois, E., Hause, G. & Bonas, U. A bacterial effector acts as a plant transcription factor and induces a cell size regulator. Science 318, 648–651 (2007).
Moscou, M. J. & Bogdanove, A. J. A simple cipher governs DNA recognition by TAL effectors. Science 326, 1501 (2009).
Boch, J. et al. Breaking the code of DNA binding specificity of TAL-type III effectors. Science 326, 1509–1512 (2009).
Deng, D. et al. Structural basis for sequence-specific recognition of DNA by TAL effectors. Science 335, 720–723 (2012).
Cermak, T. et al. Efficient design and assembly of custom TALEN and other TAL effector-based constructs for DNA targeting. Nucleic Acids Res. 39, e82–e82 (2011).
Reyon, D. et al. FLASH assembly of TALENs for high-throughput genome editing. Nat. Biotechnol. 30, 460–465 (2012).
Barrangou, R. et al. CRISPR Provides Acquired Resistance Against Viruses in Prokaryotes. Science 315, 1709–1712 (2007).
Deveau, H. et al. Phage response to CRISPR-encoded resistance in Streptococcus thermophilus. J. Bacteriol. 190, 1390–1400 (2008).
Brouns, S. J. J. et al. Small CRISPR RNAs Guide Antiviral Defense in Prokaryotes. Science 321, 960–964 (2008).
Deltcheva, E. et al. CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature 471, 602–607 (2011).
Gasiunas, G., Barrangou, R., Horvath, P. & Siksnys, V. Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proc. Natl. Acad. Sci. U. S. A. 109, E2579–E2586 (2012).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Jinek, M. et al. RNA-programmed genome editing in human cells. Elife 2, e00471 (2013).
Mali, P. et al. RNA-Guided Human Genome Engineering via Cas9. Science 339, 823–826 (2013).
Cong, L. et al. Multiplex Genome Engineering Using CRISPR/Cas Systems. Science 339, 819–823 (2013).
Hwang, W. Y. et al. Efficient genome editing in zebrafish using a CRISPR-Cas system. Nat. Biotechnol. 31, 227–229 (2013).
Nekrasov, V., Staskawicz, B., Weigel, D., Jones, J. D. G. & Kamoun, S. Targeted mutagenesis in the model plant Nicotiana benthamiana using Cas9 RNA-guided endonuclease. Nat. Biotechnol. 31, 691–693 (2013).
Li, J.-F. et al. Multiplex and homologous recombination–mediated genome editing in Arabidopsis and Nicotiana benthamiana using guide RNA and Cas9. Nat. Biotechnol. 31, 688–691 (2013).
Shan, Q. et al. Targeted genome modification of crop plants using a CRISPR-Cas system. Nat. Biotechnol. 31, 686–688 (2013).
Mao, Y. et al. Application of the CRISPR–Cas system for efficient genome engineering in plants. Mol. Plant 6, 2008–2011 (2013).
Feng, Z. et al. Efficient genome editing in plants using a CRISPR/Cas system. Cell Res. 23, 1229–1232 (2013).
Nishimasu, H. et al. Crystal structure of Cas9 in complex with guide RNA and target DNA. Cell 156, 935–949 (2014).
Wang, M. et al. Multiplex gene editing in rice with simplified CRISPR-Cpf1 and CRISPR-Cas9 systems. J. Integr. Plant Biol. 60, 626–631 (2018).
Zetsche, B. et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR-Cas system. Cell 163, 759–771 (2015).
Qi, L. S. et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152, 1173–1183 (2013).
Gilbert, L. A. et al. XCRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell 154, 442–451 (2013).
Zhang, Y., Massel, K., Godwin, I. D. & Gao, C. Applications and potential of genome editing in crop improvement. Genome Biol. 19, 1–11 (2018).
Knott, G. J. & Doudna, J. A. CRISPR-Cas guides the future of genetic engineering. Science 361, 866–869 (2018).
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Kim, Y. B. et al. Increasing the genome-targeting scope and precision of base editing with engineered Cas9-cytidine deaminase fusions. Nat. Biotechnol. 35, 371–376 (2017).
Gaudelli, N. M. et al. Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017).
Hua, K., Tao, X., Yuan, F., Wang, D. & Zhu, J.-K. Precise A•T to G•C Base Editing in the Rice Genome. Mol. Plant 11, 627–630 (2018).
Boglioli, E. & Richard, M. Rewriting The Book Of Life: A New Era in Precision Gene Editing. (2015).
Karkute, S. G., Singh, A. K., Gupta, O. P., Singh, P. M. & Singh, B. CRISPR/Cas9 mediated genome engineering for improvement of horticultural crops. Front. Plant Sci. 8, 1–6 (2017).
Sun, Z. et al. Site-Specific Gene Targeting Using Transcription Activator-Like Effector (TALE)-Based Nuclease in Brassica oleracea. J. Integr. Plant Biol. 55, 1092–1103 (2013).
Xu, C., Park, S. J., Van Eck, J. & Lippman, Z. B. Control of inflorescence architecture in tomato by BTB/POZ transcriptional regulators. Genes Dev. 30, 2048–2061 (2016).
Soyk, S. et al. Variation in the flowering gene SELF PRUNING 5G promotes day-neutrality and early yield in tomato. Nat. Genet. 49, 162–168 (2017).
Rodríguez-Leal, D., Lemmon, Z. H., Man, J., Bartlett, M. E. & Lippman, Z. B. Engineering quantitative trait variation for crop improvement by genome editing. Cell 171, 470–480.e8 (2017).
Lang, Z. et al. Critical roles of DNA demethylation in the activation of ripening-induced genes and inhibition of ripening-repressed genes in tomato fruit. Proc. Natl Acad. Sci. USA 114, E4511–E4519 (2017).
Yang, Y. et al. The RNA editing factor SlORRM4 is required for normal fruit ripening in tomato. Plant Physiol. 175, pp.01265.2017 (2017).
Ito, Y. et al. Re-evaluation of the rin mutation and the role of RIN in the induction of tomato ripening. Nat. Plants 3, 866–874 (2017).
Zsögön, A. et al. De novo domestication of wild tomato using genome editing. Nat. Biotechnol. 36, 1211–1216 (2018).
Li, T. et al. Domestication of wild tomato is accelerated by genome editing. Nat. Biotechnol. 36, 1160–1163 (2018).
Clasen, B. M. et al. Improving cold storage and processing traits in potato through targeted gene knockout. Plant Biotechnol. J. 14, 169–176 (2016).
Ye, M. et al. Generation of self-compatible diploid potato by knockout of S-RNase. Nat. Plants 4, 651–654 (2018).
Peng, A. et al. Engineering canker-resistant plants through CRISPR/Cas9-targeted editing of the susceptibility gene CsLOB1 promoter in citrus. Plant Biotechnol. J. 15, 1509–1519 (2017).
Jia, H. et al. Genome editing of the disease susceptibility gene CsLOB1 in citrus confers resistance to citrus canker. Plant Biotechnol. J. 15, 817–823 (2017).
Wang, X. et al. CRISPR/Cas9-mediated efficient targeted mutagenesis in grape in the first generation. Plant Biotechnol. J. 16, 844–855 (2018).
Okuzaki, A. et al. CRISPR/Cas9-mediated genome editing of the fatty acid desaturase 2 gene in Brassica napus. Plant Physiol. Biochem. 131, 63–69 (2018).
Ozseyhan, M. E., Kang, J., Mu, X. & Lu, C. Mutagenesis of the FAE1 genes significantly changes fatty acid composition in seeds of Camelina sativa. Plant Physiol. Biochem. 123, 1–7 (2018).
Morineau, C. et al. Selective gene dosage by CRISPR-Cas9 genome editing in hexaploid Camelina sativa. Plant Biotechnol. J. 15, 729–739 (2017).
Jiang, W. Z. et al. Significant enhancement of fatty acid composition in seeds of the allohexaploid, Camelina sativa, using CRISPR/Cas9 gene editing. Plant Biotechnol. J. 15, 648–657 (2017).
Hu, J. H. et al. Evolved Cas9 variants with broad PAM compatibility and high DNA specificity. Nature 556, 57–63 (2018).
Nishimasu, H. et al. Engineered CRISPR-Cas9 nuclease with expanded targeting space. Science 361, 1259–1262 (2018).
Schaeffer, S. M. & Nakata, P. A. CRISPR/Cas9-mediated genome editing and gene replacement in plants: Transitioning from lab to field. Plant Sci. 240, 130–142 (2015).
Woo, J. W. et al. DNA-free genome editing in plants with preassembled CRISPR-Cas9 ribonucleoproteins. Nat. Biotechnol. 33, 1162–1164 (2015).
Liang, Z. et al. Efficient DNA-free genome editing of bread wheat using CRISPR/Cas9 ribonucleoprotein complexes. Nat. Commun. 8, 14261 (2017).
Driedonks, N. et al. Exploring the natural variation for reproductive thermotolerance in wild tomato species. Euphytica 214, 67 (2018).
Gallego-Bartolomé, J. et al. Targeted DNA demethylation of the Arabidopsis genome using the human TET1 catalytic domain. Proc. Natl Acad. Sci. USA 115, E2125–E2134 (2018).
Cortijo, S. et al. Mapping the epigenetic basis of complex traits. Science 343, 1145–1148 (2014).
Kooke, R. et al. Epigenetic basis of morphological variation and phenotypic plasticity in Arabidopsis thaliana. Plant Cell 27, 337–348 (2015).
Johannes, F. et al. Assessing the impact of transgenerational epigenetic variation on complex traits. PLoS Genet. 5, e1000530 (2009).
Reinders, J. et al. Compromised stability of DNA methylation and transposon immobilization in mosaic Arabidopsis epigenomes. Genes Dev. 23, 939–950 (2009).
Büschges, R. et al. The barley Mlo gene: A novel control element of plant pathogen resistance. Cell 88, 695–705 (1997).
Wang, Y. et al. Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nat. Biotechnol. 32, 947–951 (2014).
Jiwan, D., Roalson, E. H., Main, D. & Dhingra, A. Antisense expression of peach mildew resistance locus O (PpMlo1) gene confers cross-species resistance to powdery mildew in Fragaria x ananassa. Transgenic Res. 22, 1119–1131 (2013).
Niazian, M., Sadatnoori, S. A., Galuszka, P. & Mortazavian, S. M. M. Tissue culture-based Agrobacterium-mediated and in Planta transformation methods. Czech J. Genet. Plant Breed. 53, 133–143 (2017).
Shah, S. H., Ali, S., Jan, S. A., Jalal-Ud-Din & Ali, G. M. Piercing and incubation method of in planta transformation producing stable transgenic plants by overexpressing DREB1A gene in tomato (Solanum lycopersicum Mill.). Plant Cell. Tissue Organ Cult. 120, 1139–1157 (2015).
Verma, S. S., Chinnusamy, V. & Bansal, K. C. A simplified floral dip method for transformation of Brassica napus and B. carinata. J. Plant Biochem. Biotechnol. 17, 197–200 (2008).
Hu, N. et al. Rapid and user-friendly open-source CRISPR/Cas9 system for single- or multi-site editing of tomato genome. Hortic. Res. 6, 7 (2019).
Kumlehn, J., Pietralla, J., Hensel, G., Pacher, M. & Puchta, H. The CRISPR/Cas revolution continues: From efficient gene editing for crop breeding to plant synthetic biology. J. Integr. Plant Biol. 60, 1127–1153 (2018).
Nadakuduti, S. S., Buell, C. R., Voytas, D. F., Starker, C. G. & Douches, D. S. Genome editing for crop improvement – applications in clonally propagated polyploids with a focus on potato (Solanum tuberosum L.). Front. Plant Sci. 9, 1607 (2018).
Chen, L. et al. A method for the production and expedient screening of CRISPR/Cas9-mediated non-transgenic mutant plants. Hortic. Res. 5, 13 (2018).
Veillet, F. et al. Transgene-free genome editing in tomato and potato plants using agrobacterium-mediated delivery of a CRISPR/Cas9 cytidine base editor. Int. J. Mol. Sci. 20, 402 (2019).
Zhang, Y. et al. Efficient and transgene-free genome editing in wheat through transient expression of CRISPR/Cas9 DNA or RNA. Nat. Commun. 7, 12617 (2016).
Danilo, B. et al. Efficient and transgene-free gene targeting using Agrobacterium-mediated delivery of the CRISPR/Cas9 system in tomato. Plant Cell Rep. 38, 459–462 (2019).
Ortigosa, A., Gimenez-Ibanez, S., Leonhardt, N. & Solano, R. Design of a bacterial speck resistant tomato by CRISPR/Cas9-mediated editing of SlJAZ2. Plant Biotechnol. J. 17, 665–673 (2019).
Wang, R. et al. Re-evaluation of transcription factor function in tomato fruit development and ripening with CRISPR/Cas9-mutagenesis. Sci. Rep. 9, 1696 (2019).
Wang, D. et al. Characterisation of CRISPR mutants targeting genes modulating pectin degradation in ripening tomato. Plant Physiol. pp.01187.2018, https://doi.org/10.1104/pp.18.01187 (2018).
Li, R. et al. CRISPR/Cas9-Mediated SlNPR1 mutagenesis reduces tomato plant drought tolerance. BMC Plant Biol. 19, 38 (2019).
Tomlinson, L. et al. Using CRISPR/Cas9 genome editing in tomato to create a gibberellin-responsive dominant dwarf DELLA allele. Plant Biotechnol. J. 17, 132–140 (2019).
Ding, F., Wang, M. & Zhang, S. Sedoheptulose-1,7-Bisphosphatase is Involved in Methyl Jasmonate- and Dark-Induced Leaf Senescence in Tomato Plants. Int. J. Mol. Sci. 19, 3673 (2018).
Li, R. et al. Reduction of tomato-plant chilling tolerance by CRISPR-Cas9-mediated SlCBF1 mutagenesis. J. Agric. Food Chem. 66, 9042–9051 (2018).
Boase, M. R. et al. Gene editing of tomato via Agrobacterium-mediated transformation with CRISPR/Cas 9 constructs targeting cell wall genes. Vitr. Cell. Dev. Biol. 54, S98–S99 (2018).
D’Ambrosio, C., Giorio, G. & Stigliani, A. L. Knockout of NPTII marker gene in transgenic tomato plants using the CRISPR/Cas9 system. Vitr. Cell. Dev. Biol. 54, S96 (2018).
D’Ambrosio, C., Stigliani, A. L. & Giorio, G. CRISPR/Cas9 editing of carotenoid genes in tomato. Transgenic Res. 27, 367–378 (2018).
Hashimoto, R., Ueta, R., Abe, C., Osakabe, Y. & Osakabe, K. Efficient multiplex genome editing induces precise, and self-ligated type mutations in tomato plants. Front. Plant Sci. 9, 916 (2018).
Corem, S. et al. Redistribution of CHH Methylation and Small Interfering RNAs across the Genome of Tomato ddm1 Mutants. Plant Cell 30, 1628–1644 (2018).
Chen, L. et al. Evidence for a specific and critical role of mitogen-activated protein kinase 20 in uni-to-binucleate transition of microgametogenesis in tomato. New Phytol. 219, 176–194 (2018).
Dahan-Meir, T. et al. Efficient in planta gene targeting in tomato using geminiviral replicons and the CRISPR/Cas9 system. Plant J. 95, 5–16 (2018).
Prihatna, C., Barbetti, M. J. & Barker, S. J. A novel tomato fusarium wilt tolerance gene. Front. Microbiol. 9, 1226 (2018).
Kakeshpour, T., Wu, Q., Tamang, T., Park, J. & Park, S. Multiplex Genome Editing of Class II Glutaredoxins in Solanum lycopersicum Via a CRISPR/Cas9 System. Vitr. Cell. Dev. Biol. 54, S46–S47 (2018).
Li, R., Fu, D., Zhu, B., Luo, Y. & Zhu, H. CRISPR/Cas9-mediated mutagenesis of lncRNA1459 alters tomato fruit ripening. Plant J. 94, 513–524 (2018).
Li, X. et al. Lycopene is enriched in tomato fruit by CRISPR/Cas9-mediated multiplex genome editing. Front. Plant Sci. 9, 559 (2018).
Parkhi, V. et al. Demonstration of CRISPR-Cas9 mediated pds gene editing in a tomato hybrid parental line. Indian J. Genet. Plant Breed. 78, 132–137 (2018).
Li, R. et al. Multiplexed CRISPR/Cas9-mediated metabolic engineering of γ-aminobutyric acid levels in Solanum lycopersicum. Plant Biotechnol. J. 16, 415–427 (2018).
Deng, L. et al. Efficient generation of pink-fruited tomatoes using CRISPR/Cas9 system. J. Genet. Genomics 45, 51–54 (2018).
Tashkandi, M., Ali, Z., Aljedaani, F., Shami, A. & Mahfouz, M. M. Engineering resistance against Tomato yellow leaf curl virus via the CRISPR/Cas9 system in tomato. Plant Signal. Behav. 13, e1525996 (2018).
Jung, Y. J., Lee, G.-J., Bae, S. & Kang, K. K. Reduced Ethylene Production in Tomato Fruits upon CRSPR/Cas9-mediated LeMADS-RIN Mutagenesis. Hortic. Sci. 36, 396–405 (2018).
Yu, Q. H. et al. CRISPR/Cas9-induced Targeted Mutagenesis and Gene Replacement to Generate Long-shelf Life Tomato Lines. Sci. Rep. 7, 1–18 (2017).
Wang, L. et al. Reduced drought tolerance by CRISPR/Cas9-mediated SlMAPK3 mutagenesis in tomato plants. J. Agric. Food Chem. 65, 8674–8682 (2017).
Nonaka, S., Arai, C., Takayama, M., Matsukura, C. & Ezura, H. Efficient increase of ɣ-aminobutyric acid (GABA) content in tomato fruits by targeted mutagenesis. Sci. Rep. 7, 7057 (2017).
Roldan, M. V. G. et al. Natural and induced loss of function mutations in SlMBP21 MADS-box gene led to jointless-2 phenotype in tomato. Sci. Rep. 7, 4402 (2017).
Jacobs, T. B., Zhang, N., Patel, D. & Martin, G. B. Generation of a collection of mutant tomato lines using pooled CRISPR libraries. Plant Physiol. 174, 2023–2037 (2017).
Filler Hayut, S., Melamed Bessudo, C. & Levy, A. A. Targeted recombination between homologous chromosomes for precise breeding in tomato. Nat. Commun. 8, 15605 (2017).
Soyk, S. et al. Bypassing negative epistasis on yield in tomato imposed by a domestication gene. Cell 169, 1142–1155.e12 (2017).
Gago, C. et al. Targeted gene disruption coupled with metabolic screen approach to uncover the LEAFY COTYLEDON1-LIKE4 (L1L4) function in tomato fruit metabolism. Plant Cell Rep. 36, 1065–1082 (2017).
Shimatani, Z. et al. Targeted base editing in rice and tomato using a CRISPR-Cas9 cytidine deaminase fusion. Nat. Biotechnol. 35, 441–443 (2017).
Nekrasov, V. et al. Rapid generation of a transgene-free powdery mildew resistant tomato by genome deletion. Sci. Rep. 7, 482 (2017).
Ueta, R. et al. Rapid breeding of parthenocarpic tomato plants using CRISPR/Cas9. Sci. Rep. 7, 507 (2017).
Zsögön, A., Cermak, T., Voytas, D. & Peres, L. E. P. Genome editing as a tool to achieve the crop ideotype and de novo domestication of wild relatives: case study in tomato. Plant Sci. 256, 120–130 (2017).
Klap, C. et al. Tomato facultative parthenocarpy results from SlAGAMOUS-LIKE 6 loss of function. Plant Biotechnol. J. 15, 634–647 (2017).
Čermák, T. et al. A Multipurpose Toolkit to Enable Advanced Genome Engineering in Plants. Plant Cell 29, 1196–1217 (2017).
Hilioti, Z., Ganopoulos, I., Ajith, S., Bossis, I. & Tsaftaris, A. A novel arrangement of zinc finger nuclease system for in vivo targeted genome engineering: the tomato LEC1-LIKE4 gene case. Plant Cell Rep. 35, 2241–2255 (2016).
Pan, C. et al. CRISPR/Cas9-mediated efficient and heritable targeted mutagenesis in tomato plants in the first and later generations. Sci. Rep. 6, 24765 (2016).
Jacobs, T. B. & Martin, G. B. High-throughput CRISPR Vector Construction and Characterization of DNA Modifications by Generation of Tomato Hairy Roots. J. Vis. Exp. https://doi.org/10.3791/53843 (2016).
Čermák, T., Baltes, N. J., Čegan, R., Zhang, Y. & Voytas, D. F. High-frequency, precise modification of the tomato genome. Genome Biol. 16, 232 (2015).
Ito, Y., Nishizawa-Yokoi, A., Endo, M., Mikami, M. & Toki, S. CRISPR/Cas9-mediated mutagenesis of the RIN locus that regulates tomato fruit ripening. Biochem. Biophys. Res. Commun. 467, 76–82 (2015).
Lor, V. S., Starker, C. G., Voytas, D. F., Weiss, D. & Olszewski, N. E. Targeted mutagenesis of the tomato PROCERA gene using transcription activator-like effector nucleases. Plant Physiol. 166, 1288–1291 (2014).
Brooks, C., Nekrasov, V., Lippman, Z. B. & Van Eck, J. Efficient gene editing in tomato in the first generation using the clustered regularly interspaced short palindromic repeats/CRISPR-associated9 system. Plant Physiol. 166, 1292–1297 (2014).
Nakayasu, M. et al. Generation of α-solanine-free hairy roots of potato by CRISPR/Cas9 mediated genome editing of the St16DOX gene. Plant Physiol. Biochem. 131, 70–77 (2018).
Andersson, M. et al. Genome editing in potato via CRISPR-Cas9 ribonucleoprotein delivery. Physiol. Plant. 164, 378–384 (2018).
Enciso-Rodriguez, F. et al. Overcoming self-incompatibility in diploid potato using CRISPR-Cas9. Front. Plant Sci. 10, 376 (2019). https://doi.org/10.3389/fpls.2019.00376.
Makhotenko, A. V. et al. Functional Analysis of Coilin in Virus Resistance and Stress Tolerance of Potato Solanum tuberosum using CRISPR-Cas9 Editing. Dokl. Biochem. Biophys. 484, 88–91 (2019).
Kusano, H. et al. Establishment of a modified CRISPR/Cas9 system with increased mutagenesis frequency using the translational enhancer dMac3 and multiple guide RNAs in potato. Sci. Rep. 8, 13753 (2018).
Khromov, A. et al. Efficient DNA-free nanoparticle mediated genome editing of potato using CRISPR-Cas9 RNP complex. FEBS Open Bio 8, 167 (2018).
Ma, J. et al. Genome editing in potato plants by agrobacterium-mediated transient expression of transcription activator-like effector nucleases. Plant Biotechnol. Rep. 11, 249–258 (2017).
Zhou, X. et al. StMYB44 negatively regulates phosphate transport by suppressing expression of PHOSPHATE1 in potato. J. Exp. Bot. 68, 1265–1281 (2017).
Andersson, M. et al. Efficient targeted multiallelic mutagenesis in tetraploid potato (Solanum tuberosum) by transient CRISPR-Cas9 expression in protoplasts. Plant Cell Rep. 36, 117–128 (2017).
Forsyth, A., Weeks, T., Richael, C. & Duan, H. Transcription activator-like effector nucleases (TALEN)-mediated targeted DNA insertion in potato plants. Front. Plant Sci. 7, 1572 (2016).
Butler, N. M., Baltes, N. J., Voytas, D. F. & Douches, D. S. Geminivirus-mediated genome editing in potato (Solanum tuberosum L.) using sequence-specific nucleases. Front. Plant Sci. 7, 1045 (2016).
Butler, N. M., Atkins, P. A., Voytas, D. F. & Douches, D. S. Generation and Inheritance of Targeted Mutations in Potato (Solanum tuberosum L.) Using the CRISPR/Cas System. PLoS ONE 10, e0144591 (2015).
Nicolia, A. et al. Targeted gene mutation in tetraploid potato through transient TALEN expression in protoplasts. J. Biotechnol. 204, 17–24 (2015).
Wang, S. et al. Efficient targeted mutagenesis in potato by the CRISPR/Cas9 system. Plant Cell Rep. 34, 1473–1476 (2015).
Lawrenson, T. et al. Induction of targeted, heritable mutations in barley and Brassica oleracea using RNA-guided Cas9 nuclease. Genome Biol. 16, 258 (2015).
Ma, C. et al. CRISPR/Cas9-mediated multiple gene editing in Brassica oleracea var. capitata using the endogenous tRNA-processing system. Hortic. Res. 6, 20 (2019).
Hu, L. et al. Promoter variations in a homeobox gene, BnA10.LMI1, determine lobed leaves in rapeseed (Brassica napus L.). Theor. Appl. Genet. 131, 2699–2708 (2018).
Murovec, J., Guček, K., Bohanec, B., Avbelj, M. & Jerala, R. DNA-Free Genome Editing of Brassica oleracea and B. rapa Protoplasts Using CRISPR-Cas9 Ribonucleoprotein Complexes. Front. Plant Sci. 9, 1594 (2018).
Kirchner, T. W. et al. Molecular background of Pi deficiency-induced root hair growth in Brassica carinata -a fasciclin-like arabinogalactan protein is involved. Front. Plant Sci. 9, 1372 (2018).
Sun, Q. et al. CRISPR/Cas9-mediated multiplex genome editing of the BnWRKY11 and BnWRKY70 genes in Brassica napus L. Int. J. Mol. Sci. 19, 2716 (2018).
Jiang, L. et al. Histone lysine methyltransferases BnaSDG8.A and BnaSDG8.C are involved in the floral transition in Brassica napus. Plant J. 95, 672–685 (2018).
Yang, Y. et al. Precise editing of CLAVATA genes in Brassica napus L. regulates multilocular silique development. Plant Biotechnol. J. 16, 1322–1335 (2018).
Zhang, Y. et al. Defective APETALA2 genes lead to sepal modification in Brassica Crops. Front. Plant Sci. 9, 367 (2018).
Yang, H., Wu, J.-J., Tang, T., Liu, K.-D. & Dai, C. CRISPR/Cas9-mediated genome editing efficiently creates specific mutations at multiple loci using one sgRNA in Brassica napus. Sci. Rep. 7, 7489 (2017).
Kirchner, T. W., Niehaus, M., Debener, T., Schenk, M. K. & Herde, M. Efficient generation of mutations mediated by CRISPR/Cas9 in the hairy root transformation system of Brassica carinata. PLoS ONE 12, e0185429 (2017).
Braatz, J. et al. CRISPR-Cas9 targeted mutagenesis leads to simultaneous modification of different homoeologous gene copies in polyploid oilseed rape (Brassica napus). Plant Physiol. 174, 935–942 (2017).
Kui, L. et al. Building a genetic manipulation tool box for orchid biology: identification of constitutive promoters and application of CRISPR/Cas9 in the Orchid, Dendrobium officinale. Front. Plant Sci. 7, 2036 (2016).
Bertier, L. D. et al. High-resolution analysis of the efficiency, heritability, and editing outcomes of CRISPR/Cas9-induced modifications of NCED4 in Lettuce (Lactuca sativa). G3 Genes, Genomes, Genet. 8, 1513–1521 (2018).
Chandrasekaran, J. et al. Development of broad virus resistance in non-transgenic cucumber using CRISPR/Cas9 technology. Mol. Plant Pathol. 17, 1140–1153 (2016).
Hu, B. et al. Engineering non-transgenic gynoecious cucumber using an improved transformation protocol and optimized CRISPR/Cas9 system. Mol. Plant 10, 1575–1578 (2017).
Tripathi, J. N. et al. CRISPR/Cas9 editing of endogenous banana streak virus in the B genome of Musa spp. overcomes a major challenge in banana breeding. Commun. Biol. 2, 46 (2019).
Kaur, N. et al. CRISPR/Cas9-mediated efficient editing in phytoene desaturase (PDS) demonstrates precise manipulation in banana cv. Rasthali genome. Funct. Integr. Genomics 18, 89–99 (2018).
Naim, F. et al. Gene editing the phytoene desaturase alleles of Cavendish banana using CRISPR/Cas9. Transgenic Res. 27, 451–460 (2018).
Wang, Z. et al. Optimized paired-sgRNA/Cas9 cloning and expression cassette triggers high-efficiency multiplex genome editing in kiwifruit. Plant Biotechnol. J. 16, 1424–1433 (2018).
Ren, F. et al. Efficiency optimization of CRISPR/Cas9-mediated targeted mutagenesis in Grape. Front. Plant Sci. 10, 612 (2019).
Osakabe, Y. et al. CRISPR–Cas9-mediated genome editing in apple and grapevine. Nat. Protoc. 13, 2844–2863 (2018).
Nakajima, I. et al. CRISPR/Cas9-mediated targeted mutagenesis in grape. PLoS ONE 12, e0177966 (2017).
Malnoy, M. et al. DNA-free genetically edited grapevine and apple protoplast using CRISPR/Cas9 Ribonucleoproteins. Front. Plant Sci. 7, 1904 (2016).
Ren, C. et al. CRISPR/Cas9-mediated efficient targeted mutagenesis in Chardonnay (Vitis vinifera L.). Sci. Rep. 6, 32289 (2016).
Zhang, S. et al. Regulation of citrus DMR6 via RNA interference and CRISPR/Cas9-mediated gene editing to improve Huanglongbing tolerance. Phytopathology 108, 13 (2018).
Wang, Y. Non-transgenic gene editing of Citrus sinensis using CRISPR/Cas9 ribonucleoprotein complexes. Phytopathology 108, 14 (2018).
Jia, H., Xu, J., Orbović, V., Zhang, Y. & Wang, N. Editing citrus genome via SaCas9/sgRNA system. Front. Plant Sci. 8, 2135 (2017).
Zhang, F., LeBlanc, C., Irish, V. F. & Jacob, Y. Rapid and efficient CRISPR/Cas9 gene editing in Citrus using the YAO promoter. Plant Cell Rep. 36, 1883–1887 (2017).
Jia, H. & Wang, N. Targeted genome editing of sweet orange using Cas9/sgRNA. PLoS ONE 9, e93806 (2014).
Kishi-Kaboshi, M., Aida, R. & Sasaki, K. Generation of gene-edited Chrysanthemum morifolium using multi-copy transgenes as targets and markers. Plant Cell Physiol. 58, pcw222 (2017).
Watanabe, K. et al. CRISPR/Cas9-mediated mutagenesis of the dihydroflavonol-4-reductase-B (DFR-B) locus in the Japanese morning glory Ipomoea (Pharbitis) nil. Sci. Rep. 7, 10028 (2017).
Watanabe, K., Oda-Yamamizo, C., Sage-Ono, K., Ohmiya, A. & Ono, M. Alteration of flower colour in Ipomoea nil through CRISPR/Cas9-mediated mutagenesis of carotenoid cleavage dioxygenase 4. Transgenic Res. https://doi.org/10.1007/s11248-017-0051-0 (2017).
Sun, L. & Kao, T.-H. CRISPR/Cas9-mediated knockout of PiSSK1 reveals essential role of S-locus F-box protein-containing SCF complexes in recognition of non-self S-RNases during cross-compatible pollination in self-incompatible Petunia inflata. Plant Reprod. https://doi.org/10.1007/s00497-017-0314-1 (2017).
Zhang, B., Yang, X., Yang, C., Li, M. & Guo, Y. Exploiting the CRISPR/Cas9 system for targeted genome mutagenesis in Petunia. Sci. Rep. 6, 20315 (2016).
Tian, S. et al. Engineering herbicide-resistant watermelon variety through CRISPR/Cas9-mediated base-editing. Plant Cell Rep. 37, 1353–1356 (2018).
Tian, S. et al. Efficient CRISPR/Cas9-based gene knockout in watermelon. Plant Cell Rep. 36, 399–406 (2017).
Li, B. et al. Targeted mutagenesis in the medicinal plant Salvia miltiorrhiza. Sci. Rep. 7, 43320 (2017).
Aznar-Moreno, J. A. & Durrett, T. P. Simultaneous targeting of multiple gene homeologs to alter seed oil production in Camelina sativa. Plant Cell Physiol. 58, 1260–1267 (2017).
Charrier, A. et al. Efficient targeted mutagenesis in apple and first time edition of pear using the CRISPR-Cas9 system. Front. Plant Sci. 10, 40 (2019).
Nishitani, C. et al. Efficient genome editing in apple using a CRISPR/Cas9 system. Sci. Rep. 6, 31481 (2016).
Peer, R. et al. Targeted mutagenesis using zinc-finger nucleases in perennial fruit trees. Planta 241, 941–951 (2015).
Xu, Z.-S., Feng, K. & Xiong, A.-S. CRISPR/Cas9-mediated multiply targeted mutagenesis in orange and purple carrot plants. Mol. Biotechnol. 61, 191–199 (2019).
Klimek-Chodacka, M., Oleszkiewicz, T., Lowder, L. G., Qi, Y. & Baranski, R. Efficient CRISPR/Cas9-based genome editing in carrot cells. Plant Cell Rep. 37, 575–586 (2018).
Nishihara, M., Higuchi, A., Watanabe, A. & Tasaki, K. Application of the CRISPR/Cas9 system for modification of flower color in Torenia fournieri. BMC Plant Biol. 18, 331 (2018).
Zhou, J., Wang, G. & Liu, Z. Efficient genome editing of wild strawberry genes, vector development and validation. Plant Biotechnol. J. 16, 1868–1877 (2018).
Xing, S. et al. CRISPR/Cas9-introduced single and multiple mutagenesis in strawberry. J. Genet. Genomics 45, 685–687 (2018).
Martín-Pizarro, C., Triviño, J. C. & Posé, D. Functional analysis of the TM6 MADS-box gene in the octoploid strawberry by CRISPR/Cas9-directed mutagenesis. J. Exp. Bot. 70, 885–895 (2019).
Wilson, F. M., Harrison, K., Armitage, A. D., Simkin, A. J. & Harrison, R. J. CRISPR/Cas9-mediated mutagenesis of phytoene desaturase in diploid and octoploid strawberry. Plant Methods 15, 45 (2019).
Ren, X. et al. Enhanced specificity and efficiency of the CRISPR/Cas9 system with optimized sgRNA parameters in Drosophila. Cell Rep. 9, 1151–1162 (2014).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Xu, J., Hua, K. & Lang, Z. Genome editing for horticultural crop improvement. Hortic Res 6, 113 (2019). https://doi.org/10.1038/s41438-019-0196-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41438-019-0196-5
This article is cited by
-
Gene Editing: Paving the Way for Enhancing Plant Tolerance to Abiotic Stresses-Mechanisms, Breakthroughs, and Future Prospects
Journal of Plant Growth Regulation (2024)
-
Fusion of a rice endogenous N-methylpurine DNA glycosylase to a plant adenine base transition editor ABE8e enables A-to-K base editing in rice plants
aBIOTECH (2024)
-
Applications and challenges of harnessing genome editing in oilseed crops
Journal of Plant Biochemistry and Biotechnology (2023)
-
CRISPR/Cas9-mediated targeted mutagenesis of two homoeoalleles in tobacco confers resistance to powdery mildew
Euphytica (2023)
-
Functional diversification and molecular mechanisms of FLOWERING LOCUS T/TERMINAL FLOWER 1 family genes in horticultural plants
Molecular Horticulture (2022)