Abstract
A bud sport is a lateral shoot, inflorescence or single flower/fruit with a visibly different phenotype from the rest of the plant. The new phenotype is often caused by a stable somatic mutation in a single cell that is passed on to its clonal descendants and eventually populates part or all of a meristem. In many cases, a bud sport can be vegetatively propagated, thereby preserving the novel phenotype without sexual reproduction. Bud sports provide new characteristics while retaining the desirable qualities of the parent plant, which is why many bud sports have been developed into popular cultivars. We present an overview of the history of bud sports, the causes and methods of detecting somaclonal variation, and the types of mutant phenotypes that have arisen spontaneously. We focus on examples where the molecular or cytological changes causing the phenotype have been identified. Analysis of these sports has provided valuable insight into developmental processes, gene function and regulation, and in some cases has revealed new information about layer-specific roles of some genes. Examination of the molecular changes causing a phenotype and in some cases reversion back to the original state has contributed to our understanding of the mechanisms that drive genomic evolution.
Similar content being viewed by others
Introduction
The use of traditional breeding to improve the quality of perennial fruit, nut, or ornamental plants is hindered by several factors. Many perennial species have a long juvenile period and generation time, some are self-incompatible, and most are highly heterozygous, which means valuable qualities may be lost through sexual reproduction. For these reasons, many woody perennials are vegetatively propagated by cutting, grafting, and budding, which can preserve desirable genotypes over long periods of time. Indeed, there are examples of wine grapes that have been clonally propagated for centuries1,2.
Occasionally, a lateral shoot, inflorescence or single flower/fruit is discovered with a visibly different phenotype from the rest of the plant. These are called bud sports and are often caused by a stable somatic mutation in a single cell that is passed on to its clonal descendants and eventually populates part or all of a meristem, enabling vegetative propagation of the new mutant. Although these are relatively rare events, sport mutations often provide valuable new characteristics while retaining the desirable qualities of the parent plant. Therefore, somatic mutation represents a mechanism to generate new genetic variability which is especially important for species with low levels of variation. Many economically important perennial cultivars are bud sports3,4. By 1936 there were at least 1664 known fruit tree bud sports, representing 32% of the plant patents issued by the U.S. Patent Office at that time5. More recently, Okie6 reported that more than 170 commercialized cultivars of peach and nectarine are derived from bud sport mutations.
The identification of bud sports relies on astute observation by the grower, breeder, or gardener. While these have likely been observed since humans have been cultivating plants, the earliest report of a bud sport was published by the botanist Gaspard Bauhin in 1598 and describes the unusual leaf phenotype of a Chelidonium majus (celandine) plant found in a herb garden7. In 1644, gardener Pietro Nati noticed a shoot bearing unusual fruit (the “Bizarria” orange) growing from the graft junction of two types of citrus8. Charles Darwin was fascinated by “sporting plants” and published numerous reports of spontaneous mutants9,10,11,12. In his famous book, The Variation of Plants and Animals Under Domestication11, Darwin noted that “Many cases have been recorded of a whole plant, or a single branch, or bud suddenly producing flowers different from the proper type in colour, form, size, doubleness, or other character. Half the flower, or a smaller segment sometimes changes colour”. In some cases, the molecular or cytological change(s) causing the new phenotype have been identified, which has contributed to our knowledge of the mechanisms that lead to the formation of sports and provided novel information about gene function and regulation. This review will highlight some of these examples.
Meristems make the plant
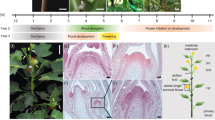
Most of the above-ground parts of the plant are produced by clusters of rapidly dividing cells in the apical and axillary meristems. Angiosperm meristems are organized into one or more outer layers, the tunica, and an inner layer or corpus which reflect stereotypical patterns of cell division gleaned from histological analysis and cell lineage studies13. Cells in the tunica tend to divide anticlinally (new cell walls formed perpendicular to the surface) such that they form one or more clonally distinct layers (Fig. 1). The number of tunica layers ranges from one to five with most species having two, the L1 and L2. The L1 gives rise to the epidermis, the L2 generates sub-epidermal layers and the germline. The corpus or L3 divides in all planes and gives rise to cells that become the core of lateral organs and the cortex of the stem.
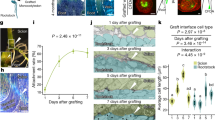
L1 and L2 indicate different layers of floral meristems, and R and r represent functional and non-functional alleles of the RIANT gene, respectively. White flower carrying two non-functional alleles of the RIANT gene (rr). a Pink flower derived from a periclinal chimera. b Pink flower with red somatic sectors derived from a mericlinal chimera. c White flower with pink somatic sectors derived from a mericlinal chimera. d White flower with red somatic sectors derived from a sectorial chimera. e Red flowering carrying one functional and one non-functional allele of the RIANT gene. Source: Cheng et al.61
Superimposed over the tunica/corpus organization are distinct zones that are defined by histological properties and cell division rates. At the very apex of each layer, there are a very small number of cells that are larger and stain less densely than surrounding cells13. These initial cells divide slowly and act as a reservoir of stem cells that replenish the supply of rapidly dividing cells on the flanks and central zone of the meristem that eventually become incorporated into lateral organs and the stem13.
The analysis of chimeric plants comprised of cells with distinct genotypes and phenotypes has provided valuable insight into meristem organization14. For example, a somatic mutation occurring in a cell located close to the apex often results in a mericlinal sector, which is visible in a portion of an organ (Fig. 1b, c, d). Occasionally, a mutant cell will populate an entire layer, creating a periclinal or layer chimera that can remain stable for long periods of time (Fig. 1a–c). Adventitious shoots originating at graft junctions can produce graft chimeras, which are generally periclinal chimeras comprised of two distinct genotypes or even different species15,16,17,18. Two excellent reviews provide in-depth information about the use of both spontaneous and induced chimeras as a research tool19 and as a source of valuable new horticultural cultivars20. Mutant L2 periclinal chimeras enter the germline, and thereafter can be sexually propagated, leading to the mutation becoming fixed in subsequent generations. Periclinal chimeras can also become homogenized when mutant cells divide into adjacent layers and eventually displace wild-type cells.
It has been assumed that meristematic cells accumulate more mutations because they have high rates of mitosis. However, it has recently been shown that shoot apical meristem cells have a relatively low mutation rate even in large perennial plants21,22, which may explain the low rates of somatic variation found in long-lived trees23.
Causes and identification of somaclonal variation
Mutations can be caused by changes to the DNA sequence or by epigenetic variations that modify DNA or histones and affect gene expression, but do not alter the sequence itself. DNA sequence errors may occur during replication, recombination, DNA damage repair during mitosis, and by transposable elements (TE). Additionally, epigenetic variations such as DNA methylation, histone modification, chromatin remodelling, and RNA silencing can cause stable changes in gene expression, generating sports.
The most frequent mutations caused by DNA polymerase errors are point mutations, tandem repeats, small insertions and deletions (indels), and base mismatches. Polymerase slippage is known to produce variability in simple sequence repeat (SSR) regions. For example, Aranzana and co-workers24 estimated a mutation rate of 1.1% of SSR alleles between peach sports. More recently, high-throughput next-generation sequencing (NGS) technologies have been used to estimate variability between bud mutants, providing more comprehensive information. Whole-genome sequences of “Fuji” apple and four bud mutant cultivars were compared to identify small polymorphisms, which revealed an average rate of eight single-nucleotide polymorphisms (SNPs) and 1.2 indels per Mb over the genome25. In grape, Carrier et al.26 compared whole-genome sequences of Pinot noir and three of its clones and found 1.6 SNPs and 5.1 indels per Mb.
Failures in the DNA replication process and damage caused by chemicals or radiation may produce DNA double-strand breaks (DSBs). Eukaryotic organisms have developed efficient mechanisms to repair DSBs through non-homologous end joining (NHEJ) and homologous recombination (HR) pathways. NHEJ is likely to occur more frequently than HR27 and does not require a homologous sequence to ligate the two DSB; it can result in small insertions or deletions of DNA at the break location and presumably contributes to microsatellite instability28. In contrast, the HR pathway requires an intact DNA molecule as template, such as the sister chromatid in a cell in S or G2 phases or the homologous chromosome. While recombination with the sister chromatid will not produce a mutation, recombination with the homologous chromosome may result in loss of heterozygosity or genome rearrangements29,30. Migliaro et al.31 characterized grape sports that were likely generated after independent DSBs and subsequent repair produced deletions ranging in size from a single bp to larger than Mbs along chromosome 2. More complex structural variations have been identified on three chromosomes of the grape sport Tempranillo blanco32, resembling a chromothripsis-like mechanism (clustered chromosomal rearrangements), which could have been generated by illegitimate re-joining of chromosome breaks in a unique event. Another example of structural variations that can alter gene expression and phenotype are copy number variants, which involve duplications or deletions of large segments of DNA33,34.
TEs contribute to genome plasticity and cause the majority of somatic variation in plants25,26,35,36,37. TEs can disrupt coding sequences, alter the expression of nearby genes and produce chromosome breakage leading to genome rearrangement and/or genome instability38. Retrotransposons are one class of TEs that are particularly disruptive because they replicate via reverse transcription of messenger RNAs and the duplicates integrate into other chromosomal locations. Related long terminal repeat (LTR) transposons are also sources of somatic variability as they can excise due to either homologous or illegitimate recombination between the terminal repeats, resulting in genomic loss or rearrangement. The activation and silencing of TEs is regulated by epigenetic mechanisms, therefore changes in such mechanisms may produce TE-driven somatic variability. Transcriptomic and epigenetic analyses have demonstrated variation in the transcription of genes associated with epigenetic mechanisms during bud dormancy release in fruit trees39,40, which may lead to the generation of sports. Stresses such as wounding, pruning, viral infection, and tissue culture are all known to induce the movement and/or activity of TEs41,42, as well as DNA damage43. McClintock44 first proposed that stress-induced activation of TEs could be a plant survival strategy to rapidly increase genotypic and phenotypic diversity in response to unfavourable conditions.
In most cases, bud sport mutations are genetically identical to their parent except for the new mutation; comparison of the parent and sport genotypes provides an opportunity to identify the molecular lesion responsible for the new phenotype. Several molecular and sequencing methods accompanied by bioinformatics pipelines have been developed to detect novel somatic changes. NGS technologies, especially those producing long, single-molecule reads using nanopore sequencing, have proven reliable in the detection of TEs that are transpositionally active in plants45, while new strategies are proposed to identify TEs active during plant development46. For a more precise estimation of somatic variability rates, such methods should consider the layer specificity of some mutations as in Marroni et al47.
Sports can arise from plants that are heterozygous for one or more loss-of-function mutation(s) that acquire independent new mutation(s) to the functional allele. Dominant gain-of-function mutations or a loss-of-function mutation to a haploinsufficient gene (one that requires two functional alleles to appear wild type) can produce a different phenotype with a single mutation event.
Lastly, not all bud sports are caused by genetic mutation. Hybridization between species and spontaneous changes in ploidy can introduce somatic variability leading to novel phenotypes48. Many sports originate as chimeras, comprised of cells from two distinct genotypes14,20. Most of the bud sports identified affect the fruit, probably because they are easy to observe, and many ornamental sports have altered floral or leaf phenotypes. In the sections that follow, we will group sports by their altered phenotype.
Floral and inflorescence morphology and/or colour
Humans have appreciated the beauty, colour and aromatic scent of flowers since at least 12,000 years ago49. Many ornamental plant cultivars originated as bud sports that change the appearance of flowers or inflorescences. Wild roses have a single whorl of five petals, whereas most cultivated roses (Rosa hybrida) have many petals. Dubois50 and co-workers demonstrated that the rose orthologue of AGAMOUS (RhAG) is expressed in whorls 3 and 4 of wild roses, consistent with its role as a C-function gene that has a key role in specifying stamen and carpel identity51. In double-flower roses, RhAG expression is restricted to a much smaller domain in the centre of the floral meristem and whorls of stamens are converted to petals. Analysis of sports that revert back to five petals also show expanded RhAG expression into whorl 350. The molecular lesion causing the misexpression of RhAG is unknown; identification of the mutation responsible may reveal information about the regulation of RhAG expression. The authors suggest that the boundary between A-function and C-function gene expression is very labile and might explain how double flower roses have arisen and been selected by humans more than twice.
Somatic grape mutants with altered flower and inflorescence development produce some of the most conspicuous phenotypes and illustrate the developmental plasticity of the tendril52. A somatic variant of Carignan has a reiterated reproductive meristem (RRM) phenotype, leading to large, highly branched fruit clusters and delayed anthesis. The tendrils of RRM mutants are also more indeterminate than wild type, displaying multiple branches or even conversion to a leafy shoot. The RRM phenotype is caused by insertion of a TE into the promoter of VvTFL1A, a close homologue of Arabidopsis TERMINAL FLOWER1 (TFL)35. Genetic and genomic analyses demonstrate that the insertion of the transposon is associated with upregulation of VvTFL1A. This is consistent with previous studies showing that TFL genes control the length of developmental phases and maintain indeterminate inflorescence growth53,54.
Mutants of Gamay, Morrastel, and Pinot initiate extra whorls of sepals and petals and are collectively known as multiple perianth whorls52. Stamen and carpel development is abnormal in variants of Bouchalès and Mourvèdre, the latter being completely sterile. Although the molecular lesion(s) causing these phenotypes is unknown, the MADS box ABC genes would be prime candidates55.
More than half of commercial varieties of azalea (Rhododendron simsii hybrids) are colour sports56. Solid colour and variegated sports are likely L1 mutants because the new phenotype is not transmitted to progeny57. The variegated sports are likely to be transposon-mediated changes to pigment genes, although no direct evidence exists. The “picotee” phenotype is characterized by petals with a coloured centre and white margins. In broad-margined “picotee” mutants, the margin cells were found to be tetraploid and the coloured cells diploid, suggesting that positionally determined polyploidisation underlies this pattern. In general, coloured azalea sports were hypermethylated relative to their parent57. In carnation (Dianthus caryophyllus), the L1 layers showed very different patterns of methylation to L2 and L3 layers, and these patterns differed widely between sports and their parents58.
Ornamental flowering peach trees can produce white, pink, and red flowers on the same tree59,60,61. In some cases, this variability has been attributed to an unstable TE in the W locus60 or differential expression of transcription factors and genes in the anthocyanin biosynthesis pathway between red and white flowers59, but genetic lesions responsible have not been identified. In the “Hongbaihuatao” (HBH) cultivar, Cheng61 and co-workers identified a small indel in RIANT, a gene encoding an anthocyanin transporter required for pigment accumulation. White flowers are homozygous for a 2-bp insertion which introduces a frameshift mutation and a premature stop codon. Red and pink flowers are heterozygous at the RIANT locus, with one non-functional allele and a second allele with either a 1-bp insertion or a 2-bp deletion that restores gene function. Periclinal, mericlinal, and sectorial chimeras with or without RIANT function produce white, pink, and red flowers (Fig. 1). These mutations are not in a microsatellite region, but are in a CG-rich region, which could increase the rate of small indels62.
Pollination, seedlessness, and fruit ripening
Self-incompatibility (SI) is a genetic mechanism that prevents inbreeding in some flowering plants. In most plants in which this SI mechanism operates, SI is controlled by a single, multi-allelic S-locus, which enables the pistil to reject pollen with the same S-allele63. Self-compatible sports have been identified in Japanese pear (Pyrus serotina)64 and almond (Prunus delcis)65. The pear sport is caused by genomic deletion of at least 4 kb that removes an S-RNase gene responsible for the SI reaction in the style. This mutation is only in the L1, providing evidence that the transmitting tissue in pear is L1-derived. In the self-compatible almond sport “Jeffries”, at least two mutations occurred, the deletion of one S haplotype and duplication of another, resulting in self-compatibility.
In most species, fruit and seed development are linked; however, there are examples where fruit development occurs in the absence of seed development. Other forms of SI occur after pollen germination and affect fertilization or embryo development, often leading to the development of a seedless fruit. Numerous seedless citrus sports have been identified, some of which have become popular cultivars such as satsuma mandarin (Citrus reticulata) and seedless navel orange. The Zigui shatian pummelo (Citrus grandis Osbeck) sport produces self-pollinated fruit, but the seeds are sterile because of defective post-zygotic development66. The seedless mandarin sports Ougan67, Wuzishatangju68, and Huami Wugegonggan69 are caused by pollen abortion, blocked fertilization, and pollen sterility and embryo abortion, respectively.
Bud sports that ripen earlier or later than their parents have been identified in numerous species. While many aspects of fruit ripening are directly controlled by the hormone ethylene70,71, different genotypes vary in their sensitivity to ethylene72. Fruit types with ripening traits predominantly regulated by ethylene are often termed “climacteric”, while those that are predominantly regulated by other factors are labelled “non-climacteric”. Genomic sequencing of bud sports of the Japanese plum “Santa Rosa” revealed copy number variation in genes associated with ethylene perception and signal transduction73. The non-climacteric and supressed-climacteric mutants had significantly fewer copies of 1-aminocyclopropane-1-carboxylic acid (ACC) oxidase (the enzyme that catalyses the final step in ethylene synthesis) and the ethylene receptor gene ETHYLENE INSENSITIVE 1 (ETR1) relative to their climacteric parent. A number of frameshift mutations were also identified in genes involved with sugar transport and ethylene biosynthesis. Similarly, early-ripening “Beni Shogun” apples show increased expression of ethylene synthesis and signal transduction genes74. Late-ripening Tardivo mandarin sports are less sensitive to ethylene and have decreased expression of ETR1 and ETR275,76. Transcriptomic and proteomic analyses of several late-maturing bud sports of sweet orange have revealed differential expression of genes involved with abscisic acid (ABA), ethylene, and jasmonic acid (JA) synthesis and signal transduction, as well as sugar metabolism and carotenoid biosynthesis77,78,79.
Altered fruit colour
Humans have been propagating grapevine (Vitis vinifera L.) for fresh fruit and wine making for over 10,000 years and from the second half of the 20th century clonal selection for wine grape breeding has been intensively used, so it is not surprising that many somatic variants have been identified that affect berry quality traits (mainly colour) and that these sports have been developed into cultivars. Grape berry colour is caused by the accumulation of anthocyanins in the berry skin (L1) and flesh (L2)80. Two tandemly repeated Myb regulatory genes, VvMybA1 and VvMybA2, regulate red berry colour81,82,83. Mutations and instability affecting this locus are the molecular basis for the majority of grape colour sports.
White grapes are thought to have originated from a black-fruit ancestor via two independent mutations to the closely linked VvMybA1 and VvMybA2 genes. A Gret1 retrotransposon insertion into the promoter of VvMybA1 and a small indel causing a frameshift mutation in VvMybA2 inactivate both genes82,83. It is unclear exactly when the heterozygous red parent self-pollinated and gave rise to homozygous white progeny, but there is evidence that ancient Egyptians were making both red and white wine by 1332 BC84. Many of the white grape cultivars tested are homozygous for these same two mutations suggesting that most have a common origin82,83, 85,86,87. Loss of the Gret1 transposon in some sports restores VvMybA1 function and gives rise to coloured revertants82,86,88. Other loss-of-pigment sports have been shown to arise from insertion of other types of TEs near VvMybA26, short genomic insertions into the promoter or introns of VvMybA89, or large-scale genomic replacements and rearrangements near VvMybA32,90.
Dark skinned cultivars occasionally produce a bronze or pale coloured sport that eventually gives rise to white berries. Walker and co-workers demonstrated the molecular basis of two such examples91. Cabernet Sauvignon produces dark red berries, but is heterozygous for the mutant VvMybA genes described above. A new mutation causing a large deletion of the functional VvMybA genes occurred in the L2 of Malian, abolishing anthocyanin production in this layer and giving the berries a bronze colour. Malian is unstable and occasionally produces white grapes or white sectors following an invasion of mutant L2 cells into the L1 (Fig. 2). Pinot noir is another unstable red cultivar that gives rise to pale sports such as Pinot gris. The authors hypothesized that a separate deletion of the VVMybA locus occurred in the L2, and eventually invaded the L1 to generate Pinot blanc clones, a stable white cultivar.
a Bunches of Cabernet Sauvignon, Malian, and Shalistin. b A bunch bearing bronze berries (wild-type L1, mutant L2) on the original Cabernet Sauvignon plant. c A Malian vine with a white bunch (mutant L1 and L2). d A white sector on a Malian berry. Source: Walker et al.91
One of the most conspicuous fruit sports is the blood orange (Citrus sinensis L.), which requires exposure to cold to develop dark red fruit. The blood orange was first documented in Italy in 164692 and was propagated clonally and sexually for centuries. A second blood orange sport arose in China in the late 1800s or earlier93. Both sports are caused by insertion of a retroelement near Ruby, a Myb gene that regulates anthocyanin production in fruit94. Cold induces retroelement transcription which activates transcription of Ruby. Many of the blood oranges found throughout Europe are derived from recombination between the LTRs that were maintained as periclinal chimeras94.
Epigenetic changes to Myb loci can also produce sports with altered colouration. “Ralls” and “Zaosu Red” are red fruit sports in apple (Malus × domestica Bork.) and a hybrid of Asian and European pear (Pyrus pyrifolia and P. communis), respectively. Both are associated with decreased methylation of promoter regions of orthologous genes MdMYB1 and PyMYB1095,96. Conversely, “Blondee”, a yellow sport of a red parent, is associated with demethylation of the MdMYB10 promoter97. Some apple cultivars can produce fruit with both solid and striped pigmentation. The regions of red stripes are associated with increased expression of MdMYB10 and decreased methylation in the MdMYB10 promoter98.
Yellow flesh peaches (Prunus persica) arose from at least three independent loss-of-function mutations to the carotenoid cleavage dioxygenase4 (PpCCD4) gene99,100. These lesions are caused by an SNP introducing a premature stop, a dinucleotide (CT) insertion in a microsatellite region creating a frameshift mutation causing a premature stop, or the insertion of a retrotransposon into an intron. Genotyping yellow and white cultivars and somatic revertants demonstrated that yellow flesh peaches have two non-functional PpCCD4 alleles, whereas the white progenitors and revertants have at least one functional PpCCD4 allele, which is sufficient to block carotenoid accumulation in the flesh. Two white bud sport mutants, revertants from yellow cultivars, were used to validate the causal relationship between allele and phenotype. Both white sports have a yellow suture (an L1-derived tissue), indicating that these are periclinal chimeras comprised of a PpCCD4-deficient L1 and a revertant L2 that has restored one functional PpCCD4 allele99,100. In one of the sports, the insertion of another CT in the mutated microsatellite region restored the reading frame and removed the stop codon99. In the other sport, deletion of the retrotransposon led to restoration of gene function100.
Pink or red sports in orange101,102, pummelo103 (Citrus grandis Osbeck.), and grapefruit104 (Citrus paradisi Macf.) have greatly increased accumulation of lycopene in the fruit. The genetic lesions causing the increase in lycopene accumulation have not been identified; however, one report showed that a cluster of six candidate genes exhibited gene dosage variation and decreased transcription between the sport Hong Anliu and its parent105. A yellow citrus sport Pinalate has yellow fruit with high levels of carotenes and decreased ABA content106. Three citrus sports with brown skin have been reported to be caused by defective chlorophyll degradation107, altered carotenoid accumulation108, and defective synthesis or accumulation of β-citraurin108.
Altered fruit size or shape
After fruit colour, the most obvious type of sport mutant is one that has a different fruit size or shape. Final fruit size results from cell division and enlargement, and many fruit size sports are clearly affected in one of these processes. Increased DNA content caused by endoreduplication or chimeric polyploidization often results in larger cells and increased lateral organ and/or fruit size109,110. Interestingly, giant fruit sports in apple111 and pear112 both showed fruit-specific increases to DNA content and cell size, suggesting that fruit cell size is under separate regulation to other parts of the plant. Another giant pear sport had no change in ploidy, but showed increased expression of an actin-related protein that is involved with regulating cell proliferation in Arabidiopsis113,114.
“Totsutanenashi” (TTN) is a small fruit sport in Japanese persimmon (Diospyros kaki Thumb.) that also increases sugar content in fruit and causes a more compact tree architecture115. Application of exogenous cytokinin restores normal fruit size116, suggesting that the mutant gene involves cytokinin biosynthesis or signal transduction. The progenitor of TTN is non-aploid (2n = 9×) and has produced many sports, raising the possibility that recent increases in ploidy have led to increased transposon movement and/or chromosomal rearrangements.
In grape, the fleshless berry (flb) mutation disrupts cell division and differentiation in the mesocarp of the fruit, resulting in a 10-fold reduction in berry weight117. Observations that the flb phenotype was unstable and that some progeny obtained from sexual propagation showed a new phenotype that failed to set fruit led to the discovery that the original flb mutant is an L2 chimera118. The flb mutation is caused by the insertion of a TE into the promoter of VvPI, resulting in ectopic expression of the grape homologue of Arabidopsis PISTILATA119,120. Analysis of plants that carry the mutation in the L1, L2, or in both layers demonstrate the differential effects of ectopic VvPI expression in each layer. VvPI expression in the L2 blocks flesh development by preventing mesocarp cells from differentiating, while VvPI expression in the L1 and L2 disrupts carpel development at fruit set119. This is an excellent example of how the study of a sport mutant has provided new insights into the layer-specific role of PI in floral and fleshy fruit development. Interestingly, both the TTN and flb L2 sports have increased sugar content and decreased phenolic compounds, which suggest a potential connection between fruit size and biochemical properties115,117.
Nectarines were first identified in China over 2000 years ago121. Genotypic analysis indicates that nectarines were introduced or arose in Europe multiple times, probably as bud sports122. Genomic data from five peach/nectarine accessions indicated that the insertion of a LTR retroelement in the coding sequence of PpeMYB25 is the likely cause of the hairless nectarine phenotype123. In Arabidopsis, loss-of-function mutations to the closely related MYB gene, GLABRA1, result in hairless leaves124. Nectarine sports of peach are frequently observed in orchards and some have been commercialized.
Flat fruit shape in peach is caused by a single semi-dominant locus (S), which may itself be a bud sport mutant that originated in China. Individuals heterozygous for S have flat fruit, while those homozygous for S show early abortion of the fruit125. Analysis of the flat fruit indicates they have fewer cells in the vertical axis due to earlier cessation of cell division relative to round fruit126. The flat fruit phenotype has been associated with a 10 kb deletion that removes the first 693 bp of a leucine-rich receptor-like kinase (LRR-RLK) gene (PRUPE.6G281100/ppa025511m), orthologous to the Arabidopsis BRASSINOSTEROID INSENSITIVE1 ASSOCIATED KINASE (BAK1) group125. BAK1-like genes regulate cell division and meristem size in Arabidopsis127,128, rice129, 130, and maize131. A bud sport mutant reverting from flat to round (Fig. 3) has been shown to carry a new mutation to PRUPE.6G281100 in the L2 layer, providing strong support that this gene is involved with the flat fruit phenotype. Guo et al.126 identified some flat peach accessions with two functional PRUPE.6G281100 alleles, suggesting that a second gene may also contribute to this phenotype. A polymorphism in PpCAD1/ppa003772m, annotated as CONSTITUTIVELY ACTIVATED CELL DEATH GENE1, was highly associated with the flat peach phenotype in a genome-wide association study132. However, the SNP is not predicted to alter gene function and the gene is not differentially expressed between flat and round fruit until the mature fruit stage.
Source: López-Girona et al.125
Changes to plant architecture
On rare occasions, a sport arises that drastically alters plant architecture. Pinot Meunier is a sport of Pinot noir that is characterized by having leaves and stems that are densely covered with trichomes and a conversion of tendrils into inflorescences133. Occasionally, leaf sectors lacking the hairy phenotype appear on Pinot Meunier suggesting that it is a periclinal chimera with a mutant L12,133. Indeed, plants regenerated from L1 or L2 layers indicate L1-derived plants are hairy, while L2-derived plants are hairless2. More striking is the dwarfed phenotype of L1-derived plants, caused by very short internodes (Fig. 4a, c). This dwarfed phenotype was not rescued by application of gibberellins (GAs), indicating that it is not a GA biosynthesis mutant134. The mutation is caused by a non-synonymous SNP in the highly conserved DELLA domain of VvGAI, which encodes a member of key GA-responsive proteins134. The conversion of tendrils to inflorescences supports the idea that the grape tendril is a modified inflorescence normally inhibited from floral development by GA. This is another excellent example of sport mutants providing new information about the layer-specific effects of a mutation.
In the early 1960s, apple grower Anthony Wijcik noticed an abnormal shoot on a “McIntosh” tree in his orchard135. The “Wijcik” sport has a “columnar” growth habit with a thick, stunted primary axis, very short internodes, and short lateral spurs rather than lateral branches (Fig. 5). This phenotype has generated considerable commercial interest since columnar trees could be planted at high density, require less pruning than standard types, and potentially enable mechanical harvesting. The columnar phenotype segregates as a single dominant allele136 that is associated with the integration of a retrotransposon into the genome36,137. Although there is some discrepancy in the size of the insertion, it is clear that the insertion does not disrupt any coding sequence, but does alter expression of nearby genes. Expression analysis of genes within 25 Kb of the insertion demonstrated that a gene encoding a putative 2OG-Fe (II) oxygenase was upregulated 14-fold over “McIntosh” in young axillary buds. Overexpression of this gene (MdCo31) in Arabidopsis resulted in very short inflorescences due to reduced internode lengths137. Members of this gene family are involved with the biosynthesis of ethylene, flavonoids, gibberellins, and defence against downy mildew. Several studies have used RNAseq to identify differentially expressed genes (DEGs) between columnar and standard trees36,138,139. Many of the DEG from shoot meristem tissue are involved with hormone metabolism and signalling140. Genes involved with lignin and terpene biosynthesis, and pathogen/pest attack response were highly upregulated in “Wijcik” leaves36. Two other genes near the insertion are upregulated in “Wijcik”. One encodes a helix-loop-helix transcription factor and the other Downy Mildew Resistant 6 (DMR6), a regulator of defence genes.
a Apple trees with a standard growth habit have long lateral branches and a wide branching angle relative to the primary shoot. b–c Columnar trees have a compact growth habit and produce mostly short fruit spurs with a narrow branch angle. Source: Peterson and Krost155
Somatic mutants of non-perennial species
Some annual crop plants are clonally propagated for a variety of reasons. Clonal propagation preserves the genotype and phenotype of the parent plant, which could be lost in sexual propagation. Prevention of sexual reproduction helps avoid inbreeding depression which is common in outcrossing species. In varieties selected for seedlessness, such as banana, sexual propagation may not be possible.
A large number of potato (Solanum tuberosum) cultivars are bud sports that alter the pigmentation of the tuber including the leading French fry processing cultivar “Russet Burbank“141,142,143,144,145. Many coloured potato sports are periclinal chimeras that show a range of pigmentation from white to purple143,144,145. Careful analysis of colouration patterns in these sports and their sexual offspring revealed that the L1 gives rise to most of the tuber skin except for a small patch below each eye, which is L2 derived143. The “Kostroma mutant” does not affect the tuber, but shows pronounced dissection of leaf blade and flower corolla146. Spontaneous mutations giving rise to coloured tubers and sectorial chimeras have also been developed into popular cultivars of sweetpotato (Ipomoea batatas)147.
Cassava (Manihot esculenta Crantz) is an important staple crop in Sub-Saharan Africa, the tropics, sub-tropics, and South Pacific islands. Cassava sports with altered leaf morphology and increased vigour are associated with spontaneous changes in ploidy, probably caused by the fertilization of unreduced gametes148,149,150. Many of the crops grown in Oceania originated from a very limited number of introduced genotypes and have been mostly or exclusively vegetatively propagated. Somaclonal variation has been a valuable source of phenotypic diversity for farmers growing taro (Colocasia esculenta Schott) and yam (Dioscorea alata L.)151, bananas and plantains (Musa spp.)152,153, and kava (Piper methysticum Forst f.)154.
Conclusions
Bud sport mutants introduce new genetic variability, which is crucial for species that lack variability or cannot be sexually reproduced. Somatic mutations can also provide valuable new characteristics while retaining the desirable qualities of the parent plant, which is why many popular cultivars have originated from sports. In some cases, the analysis of sports has revealed new information about layer-specific roles of some genes or has provided valuable insight into gene function during specific developmental processes. As sequencing technologies improve, sports also provide a means to examine mechanisms that drive genomic evolution.
References
This, P., Lacombe, T. & Thomas, M. R. Historical origins and genetic diversity of wine grapes. Trends Genet. 22, 511–519 (2006).
Franks, T., Botta, R., Thomas, M. R. & Franks, J. Chimerism in grapevines: implications for cultivar identity, ancestry and genetic improvement. Theor. Appl. Genet. 104, 192–199 (2002).
Granhall, I. Spontaneous and induced bud mutations in fruit trees. Acta Agric. Scand. 4, 594–600 (1954).
Van Harten, A. M. in Breeding for Ornamentals: Classical and Molecular Approaches (ed Vainstein, A.) 105–127 (Kluwer Academic Publishers, Dordrecht, the Netherlands, 2002).
Shamel, A. D. & Pomeroy, C. S. Bud mutations in horticultural crops. J. Hered. 27, 487–494 (1936).
Okie, W. R. Handbook of Peach and Nectarine Varieties: Performance in the Southeastern United States and Index of Names (U.S. Dept. of Agriculture, Agricultural Research Service; National Technical Information Service, distributor, Washington, DC; Springfield, VA, 1998).
Stubbe, H. Kurze Geschichte der Genetik bis zur Wiederentdeckung der Vererbungsregeln Gregor Mendels. (VEB Gustav Fischer Verlag, Jena, 1963).
Nati, P. Florentina phytologica observatio de malo Limonia citrata-aurantia, Florentiae vulgo la bizzaria, (Florence, Italy, 1674).
Darwin, C. R. Double flowers-their origin. Gard. Chron. Agric. Gaz. 36, 628 (1843).
Darwin, C. R. Variegated leaves. Gard. Chron. Agric. Gaz. 37, 621 (1844).
Darwin, C. R. The Variation of Animals and Plants under Domestication. (John Murray, London, 1858).
Darwin, C. R. On the Origin of Species. (John Murray, London, 1859).
Steeves, T. A. & Sussex, I. M. Patterns in Plant Development. (Cambridge University Press, New York, NY, 1989).
Tilney-Bassett, R. A. E. Plant Chimeras. (Edward Arnold Ltd., London, 1986).
Winkler, H. Über Pfropfbastarde und pflanliche Chimären. Ber. Der Dtsch. Bot. Ges. 25, 568–576 (1907).
Neilson-Jones, W. Plant Chimeras. (Methuen, London, 1969).
Stewart, R. N., Meyer, F. G. & Dermen, H. Camellia+‘Daisy Eagleson’, a graft chimera of Camellia sasanqua and C. japonia. Am. J. Bot. 59, 515–524 (1972).
Jwamasa, M.N.M., Okudal, N., & Ishiuchi, D. in International Citrus Congress, Florida, USA, 571–574 (FL, 1977).
Frank, M. H. & Chitwood, D. H. Plant chimeras: the good, the bad, and the ‘Bizzaria’. Dev. Biol. 419, 41–53 (2016).
Burge, G. K., Morgan, E. R. & Seelye, J. F. Opportunities for synthetic plant chimeral breeding: past and future. Plant Cell Tissue Organ Cult. 70, 13–21 (2002).
Watson, J. M. et al. Germline replications and somatic mutation accumulation are independent of vegetative life span in Arabidopsis. Proc. Natl Acad. Sci. USA 113, 12226–12231 (2016).
Lanfear, R. et al. Taller plants have lower rates of molecular evolution. Nat. Commun. 4, 1879 (2013).
Schmid-Siegert, E. et al. Low number of fixed somatic mutations in a long-lived oak tree. Nat. Plants 3, 926–929 (2017).
Aranzana, M. J., Carbo, J. & Arus, P. Microsatellite variability in peach [Prunus persica (L.) Batsch]: cultivar identification, marker mutation, pedigree inferences and population structure. Theor. Appl. Genet. 106, 1341–1352 (2003).
Lee, H. S. et al. Analysis of ‘Fuji’ apple somatic variants from next-generation sequencing. Genet. Mol. Res. 15: gmr.15038185 (2016).
Carrier, G. et al. Transposable elements are a major cause of somatic polymorphism in Vitis vinifera L. PLoS ONE 7, e32973 (2012).
Boyko, A., Hudson, D., Bhomkar, P., Kathiria, P. & Kovalchuk, I. Increase of homologous recombination frequency in vascular tissue of Arabidopsis plants exposed to salt stress. Plant Cell Physiol. 47, 736–742 (2006).
Golubov, A. et al. Microsatellite instability in Arabidopsis increases with plant development. Plant Physiol. 154, 1415–1427 (2010).
Pardo, B., Gomez-Gonzalez, B. & Aguilera, A. DNA repair in mammalian cells: DNA double-strand break repair: how to fix a broken relationship. Cell Mol. Life Sci. 66, 1039–1056 (2009).
LaFave, M. C. & Sekelsky, J. Mitotic recombination: Why? When? How? Where? PLoS Genet. 5, e1000411 (2009).
Migliaro, D. et al. Structural dynamics at the berry colour locus in Vitis vinifera L. somatic variants. Aust. J. Grape Wine Res. 20, 485–495 (2014).
Carbonell-Bejerano, P. et al. Catastrophic unbalanced genome rearrangements cause somatic loss of berry color in grapevine. Plant Physiol. 175, 786–801 (2017).
Samelak-Czajka, A. et al. MLPA-based analysis of copy number variation in plant populations. Front. Plant Sci. 8, 222 (2017).
Dolatabadian, A., Patel, D. A., Edwards, D. & Batley, J. Copy number variation and disease resistance in plants. Theor. Appl. Genet. 130, 2479–2490 (2017).
Fernandez, L., Torregrosa, L., Segura, V., Bouquet, A. & Martinez-Zapater, J. M. Transposon-induced gene activation as a mechanism generating cluster shape somatic variation in grapevine. Plant J. 61, 545–557 (2010).
Otto, D., Petersen, R., Brauksiepe, B., Braun, P. & Schmidt, E. R. The columnar mutation (“Co gene”) of apple (Malus×domestica) is associated with an integration of a Gypsy-like retrotransposon. Mol. Breed. 33, 863–880 (2014).
Lijavetzky, D., Cabezas, J., Ibanez, A., Rodriguez, V. & Martinez-Zapater, J. High throughput SNP discovery and genotyping in grapevine (Vitis vinifera L.) by combining a re-sequencing approach and SNPlex technology. BMC Genomics 8, 424 (2007).
Ikeda, Y. & Nishimura, T. in Nuclear Functions in Plant Transcription, Signaling and Development (eds Pontes, O. & Jin, H.) 13–29 (Springer New York, New York, NY, 2015).
Ríos, G., Leida, C., Conejero, A. & Badenes, M. L. Epigenetic regulation of bud dormancy events in perennial plants. Front. Plant Sci. 5, 247 (2014).
Bai, S. et al. Transcriptome analysis of Japanese pear (Pyrus pyrifolia Nakai) flower buds transitioning through endodormancy. Plant Cell Physiol. 54, 1132–1151 (2013).
Asíns, M. J., Monforte, A. J., Mestre, P. F. & Carbonell, E. A. Citrus and Prunuscopia-like retrotransposons. Theor. Appl. Genet. 99, 503–510 (1999).
Wessler, S. R. Plant retrotransposons: turned on by stress. Curr. Biol. 6, 959–961 (1996).
Manova, V. & Gruszka, D. DNA damage and repair in plants: from models to crops. Front. Plant Sci. 6, 885 (2015).
McClintock, B. The significance of responses of the genome to challenge. Science 226, 792–801 (1984).
Debladis, E., Llauro, C., Carpentier, M.-C., Mirouze, M. & Panaud, O. Detection of active transposable elements in Arabidopsis thaliana using Oxford Nanopore Sequencing technology. BMC Genomics 18, 537 (2017).
Lanciano, S. et al. Sequencing the extrachromosomal circular mobilome reveals retrotransposon activity in plants. PLoS Genet. 13, e1006630 (2017).
Marroni, F. et al. Reduction of heterozygosity (ROH) as a method to detect mosaic structural variation. Plant Biotechnol. J. 15, 791–793 (2017).
Springer, N. M. & Schmitz, R. J. Exploiting induced and natural epigenetic variation for crop improvement. Nat. Rev. Genet. 18, 563 (2017).
Nadel, D. et al. Earliest floral grave lining from 13,700–11,700-year-old Natufian burials at Raqefet Cave, Mt. Carmel, Israel. Proc. Natl Acad. Sci. USA 110, 11774–11778 (2013).
Dubois, A. et al. Tinkering with the C-function: a molecular frame for the selection of double flowers in cultivated roses. PLoS ONE 5, e9288 (2010).
Smyth, D. R., Bowman, J. L. & Meyerowitz, E. M. Early flower development in Arabidopsis. Plant Cell 2, 755 (1990).
Chatelet, P. et al. Characterization of Vitis vinifera L. somatic variants exhibiting abnormal flower development patterns. J. Exp. Bot. 58, 4107–4118 (2007).
Blázquez, M., Koornneef, M. & Putterill, J. Flowering on time: genes that regulate the floral transition: workshop on the molecular basis of flowering time control. EMBO Rep. 2, 1078–1082 (2001).
Bowman, J. L., Alvarez, J., Weigel, D., Meyerowitz, E. M. & Smyth, D. R. Control of flower development in Arabidopsis thaliana by APETALA1 and interacting genes. Development 119, 721 (1993).
Bowman, J. L., Smyth, D. R. & Meyerowitz, E. M. The ABC model of flower development: then and now. Development 139, 4095–4098 (2012).
De Schepper, S., Debergh, P., Van Bockstaele, E. & De Loose, M. 143–150 Molecular characterisation of flower colour genes in azalea sports (rhododendron simsii hybrids (International Society for Horticultural Science (ISHS), Leuven, 2001).
De Schepper, S. et al. Genetic and epigenetic aspects of somaclonal variation: flower colour bud sports in azalea, a case study. South Afr. J. Bot. 69, 117–128 (2003).
Yoshida, H. et al. Alteration of methylation profiles in distinct cell lineages of the layers during vegetative propagation in carnations (Dianthus caryophyllus). Euphytica 135, 247 (2004).
Chen, Y. et al. Transcriptome analysis of differentially expressed genes relevant to variegation in peach flowers. PLoS ONE 9, e90842 (2014).
Chaparro, J. X., Werner, D. J., Whetten, R. W. & O’Malley, D. M. Characterization of an unstable anthocyanin phenotype and estimation of somatic mutation rates in peach. J. Hered. 86, 186–193 (1995).
Cheng, J. et al. A small indel mutation in an anthocyanin transporter causes variegated colouration of peach flowers. J. Exp. Bot. 66, 7227–7239 (2015).
Tanay, A. & Siggia, E. D. Sequence context affects the rate of short insertions and deletions in flies and primates. Genome Biol. 9, R37 (2008).
Kao, T.-h & Tsukamoto, T. The molecular and genetic bases of S-RNase-based self-incompatibility. Plant Cell 16, S72–S83 (2004).
Sassa, H., Hirano, H., Nishio, T. & Koba, T. Style-specific self-compatible mutation caused by deletion of the S-RNase gene in Japanese pear (Pyrus serotina). Plant J. 12, 223–227 (1997).
Ushijima, K. et al. Characterization of the S-locus region of almond (Prunus dulcis): analysis of a somaclonal mutant and a cosmid contig for an S haplotype. Genetics 158, 379–386 (2001).
Chai, L., Ge, X., Biswas, M. K., Xu, Q. & Deng, X. Self-sterility in the mutant ‘Zigui shatian’ pummelo (Citrus grandis Osbeck) is due to abnormal post-zygotic embryo development and not self-incompatibility. Plant Cell Tissue Organ Cult. 104, 1–11 (2011).
Hu, Z. et al. Abnormal microspore development leads to pollen abortion in a seedless mutant of ‘Ougan’ Mandarin (Citrus suavissima Hort. ex Tanaka). J. Am. Soc. Hortic. Sci. 132, 777–782 (2007).
Ye, W. et al. Seedless mechanism of a new mandarin cultivar ‘Wuzishatangju’ (Citrus reticulata Blanco). Plant Sci. 177, 19–27 (2009).
Yonghua, Q., Xu, C., Ye, Z., Teixeira da Silva, J. & Hu, G. Seedless mechanism of a new citrus cultivar ‘Huami Wuhegonggan’ (Citrus sinensis × C. reticulata), Pak. J. Bot. 46, 2369-2378 (2015).
Klee, H. J. & Giovannoni, J. J. Genetics and control of tomato fruit ripening and quality attributes. Annu. Rev. Genet. 45, 41–59 (2011).
Liu, M., Pirrello, J., CHERVIN, C., Roustan, J.-P. & Bouzayen, M. Ethylene control of fruit ripening: revisiting the complex network of transcriptional regulation. Plant Physiol. 169, 2380–2390 (2015).
Paul, V., Pandey, R. & Srivastava, G. C. The fading distinctions between classical patterns of ripening in climacteric and non-climacteric fruit and the ubiquity of ethylene—an overview. J. Food Sci. Technol. 49, 1–21 (2012).
Fernandez i Marti, A., Saski, C. A., Manganaris, G. A., Gasic, K. & Crisosto, C. H. Genomic sequencing of Japanese plum (Prunus salicina Lindl.) mutants provides a new model for Rosaceae fruit ripening studies. Front. Plant Sci. 9, 21 (2018).
Dong, Q. L., Yan, Z. Y., Liu, Z. & Yao, Y. X. Early ripening events caused by bud mutation in Beni Shogun apple. Russ. J. Plant Physiol. 58, 439 (2011).
Distefano, G. et al. Physiological and molecular analysis of the maturation process in fruits of clementine mandarin and one of its late-ripening mutants. J. Agric. Food Chem. 57, 7974–7982 (2009).
Alós, E., Distefano, G., Rodrigo, M. J., Gentile, A. & Zacarías, L. Altered sensitivity to ethylene in ‘Tardivo’, a late-ripening mutant of Clementine mandarin. Physiol. Plant 151, 507–521 (2014).
Romero, P. et al. Unravelling molecular responses to moderate dehydration in harvested fruit of sweet orange (Citrus sinensis L. Osbeck) using a fruit-specific ABA-deficient mutant. J. Exp. Bot. 63, 2753–2767 (2012).
Wu, J., Xu, Z., Zhang, Y., Chai, L. & Yi, H. An integrative analysis of the transcriptome and proteome of the pulp of a spontaneous late-ripening sweet orange mutant and its wild type improves our understanding of fruit ripening in citrus. J. Exp. Bot. 65, 1651–1671 (2014).
Zhang, Y.-J. et al. Comparative transcriptome analyses between a spontaneous late-ripening sweet orange mutant and its wild type suggest the functions of ABA, sucrose and JA during citrus fruit ripening. PLoS ONE 9, e116056 (2015).
Boss, P. K., Davies, C. & Robinson, S. P. Expression of anthocyanin biosynthesis pathway genes in red and white grapes. Plant Mol. Biol. 32, 565–569 (1996).
Kobayashi, S., Ishimaru, M., Hiraoka, K. & Honda, C. Myb-related genes of the Kyoho grape (Vitis labruscana) regulate anthocyanin biosynthesis. Planta 215, 924–933 (2002).
Kobayashi, S., Goto-Yamamoto, N. & Hirochika, H. Retrotransposon-induced mutations in grape skin color. Science 304, 982 (2004).
Walker, A. R. et al. White grapes arose through the mutation of two similar and adjacent regulatory genes. Plant J. 49, 772–785 (2007).
Guasch-Jané, M. R., Andrés-Lacueva, C., Jáuregui, O. & Lamuela-Raventós, R. M. First evidence of white wine in ancient Egypt from Tutankhamun’s tomb. J. Archaeol. Sci. 33, 1075–1080 (2006).
This, P., Lacombe, T., Cadle-Davidson, M. & Owens, C. L. Wine grape (Vitis vinifera L.) color associates with allelic variation in the domestication gene VvmybA1. Theor. Appl. Genet. 114, 723–730 (2007).
Lijavetzky, D. et al. Molecular genetics of berry colour variation in table grape. Mol. Genet. Genom. 276, 427–435 (2006).
Fournier-Level, A. et al. Quantitative genetic bases of anthocyanin variation in grape (Vitis vinifera L. ssp. sativa) berry: a quantitative trait locus to quantitative trait nucleotide integrated study. Genetics 183, 1127–1139 (2009).
Azuma, A. et al. Color recovery in berries of grape (Vitis vinifera L.) ‘Benitaka’, a bud sport of ‘Italia’, is caused by a novel allele at the VvmybA1 locus. Plant Sci. 176, 470–478 (2009).
Shimazaki, M., Fujita, K., Kobayashi, H. & Suzuki, S. Pink-colored grape berry is the result of short insertion in intron of color regulatory gene. PLoS ONE 6, e21308 (2011).
Pelsy, F., Dumas, V., Bévilacqua, L., Hocquigny, S. & Merdinoglu, D. Chromosome replacement and deletion lead to clonal polymorphism of berry color in grapevine. PLoS Genet. 11, e1005081 (2015).
Walker, A. R., Lee, E. & Robinson, S. P. Two new grape cultivars, bud sports of Cabernet Sauvignon bearing pale-coloured berries, are the result of deletion of two regulatory genes of the berry colour locus. Plant Mol. Biol. 62, 623–635 (2006).
Ferrari, G. B. Hesperides Sive Malorum Aureorum Cultura et Usu (Sumptibus Hermanni Scheus, Romae, 1646).
Yuan F., Long G., & Deng, Z. The 11th International Citrus Congress, Proceedings of the International Society of Citriculture, 70–72 (China Agriculture Press, Beijing, 2008).
Butelli, E. et al. Retrotransposons control fruit-specific, cold-dependent accumulation of anthocyanins in blood oranges. Plant Cell 24, 1242–1255 (2012).
Xu, Q. et al. Comparative transcripts profiling reveals new insight into molecular processes regulating lycopene accumulation in a sweet orange (Citrus sinensis) red-flesh mutant. Genetics and Molecular Research 15: gmr.15038185 (2009).
Qian, M., Sun, Y., Allan, A. C., Teng, Y. & Zhang, D. The red sport of ‘Zaosu’ pear and its red-striped pigmentation pattern are associated with demethylation of the PyMYB10 promoter. Phytochemistry 107, 16–23 (2014).
El-Sharkawy, I., Liang, D. & Xu, K. Transcriptome analysis of an apple (Malus×domestica) yellow fruit somatic mutation identifies a gene network module highly associated with anthocyanin and epigenetic regulation. J. Exp. Bot. 66, 7359–7376 (2015).
Telias, A. et al. Apple skin patterning is associated with differential expression of MYB10. BMC Plant Biol. 11, 93–93 (2011).
Falchi, R. et al. Three distinct mutational mechanisms acting on a single gene underpin the origin of yellow flesh in peach. Plant J. 76, 175–187 (2013).
Adami, M. et al. Identifying a Carotenoid Cleavage Dioxygenase (CCD4) gene controlling yellow/white fruit flesh color of peach. Plant Mol. Biol. Report 31, 1166–1175 (2013).
Liu, Q. et al. A novel bud mutation that confers abnormal patterns of lycopene accumulation in sweet orange fruit (Citrus sinensis L. Osbeck). J. Exp. Bot. 58, 4161–4171 (2007).
Alquezar, B., Rodrigo, M. J. & Zacarias, L. Regulation of carotenoid biosynthesis during fruit maturation in the red-fleshed orange mutant Cara Cara. Phytochemistry 69, 1997–2007 (2008).
Liu, W. et al. A spontaneous bud mutant that causes lycopene and β-carotene accumulation in the juice sacs of the parental Guanxi pummelo fruits (Citrus grandis (L.) Osbeck). Sci. Hortic. 198, 379–384 (2016).
Monselise, S. P. & Halevy, A. H. Detection of lycopene in pink orange fruit. Science 133, 1478 (1961).
Pan, Z., An, J., Zeng, W., Xiao, S. & Deng, X. Array-comparative genome hybridization reveals genome variations between a citrus bud mutant and its parental cultivar. Tree Genet. Genomes 8, 1379–1387 (2012).
Rodrigo, M. J., Marcos, J. F., Alferez, F., Mallent, M. D. & Zacarias, L. Characterization of Pinalate, a novel Citrus sinensis mutant with a fruit-specific alteration that results in yellow pigmentation and decreased ABA content. J. Exp. Bot. 54, 727–738 (2003).
Alós, E. et al. An evaluation of the basis and consequences of a stay-green mutation in the navel negra Citrus mutant using transcriptomic and proteomic profiling and metabolite analysis. Plant Physiol. 147, 1300 (2008).
Ríos, G. et al. Identification of a GCC transcription factor responding to fruit colour change events in citrus through the transcriptomic analyses of two mutants. BMC Plant Biol. 10, 276–276 (2010).
Stebbins, G. L. Chromosome Evolution in Higher Plants (Arnold, London, 1972).
Tal, M. in Polyploidy-Biological Relevance (ed Lewis, W.) 61–75 (Plenum Press, New York, 1980).
Malladi, A. & Hirst, P. M. Increase in fruit size of a spontaneous mutant of ‘Gala’ apple (Malus×domestica Borkh.) is facilitated by altered cell production and enhanced cell size. J. Exp. Bot. 61, 3003–3013 (2010).
Isuzugawa, K., Murayama, H. & Nishio, T. Characterization of a giant-fruit mutant exhibiting fruit-limited polyploidization in pear (Pyrus communis L.). Sci. Hortic. 170, 196–202 (2014).
Zhang, S. J. et al. Identification of differentially expressed genes in a spontaneous mutant of ‘Nanguoli’ pear (Pyrus ussuriensis Maxim) with large fruit. J. Hortic. Sci. Biotechnol. 86, 595–602 (2011).
Deal, R. B., Kandasamy, M. K., McKinney, E. C. & Meagher, R. B. The nuclear actin-related protein ARP6 is a pleiotropic developmental regulator required for the maintenance of flowering locus C expression and repression of flowering in Arabidopsis. Plant Cell 17, 2633–2646 (2005).
Yamane, H. et al. Growth characteristics of a small-fruit dwarf mutant arising from bud sport mutation in Japanese Persimmon (Diospyros kaki Thunb.). HortScience 43, 1726–1730 (2008).
Habu, T. et al. Differences in physiological characteristics and gene expression levels in fruits between Japanese Persimmon (Diospyros kaki Thunb.) ‘Hiratanenashi’ and its small fruit mutant ‘Totsutanenashi’. Hort. J. 85, 306–314 (2016).
Fernandez, L. et al. The grapevine fleshless berry mutation. A unique genotype to investigate differences between fleshy and nonfleshy fruit. Plant Physiol. 140, 537–547 (2006).
Fernandez, L. et al. Somatic chimerism, genetic inheritance, and mapping of the fleshless berry (flb) mutation in grapevine (Vitis vinifera L.). Genome 49, 721–728 (2006).
Fernandez, L., Chaïb, J., Martinez-Zapater, J.-M., Thomas, M. R. & Torregrosa, L. Mis-expression of a PISTILLATA-like MADS box gene prevents fruit development in grapevine. Plant J. 73, 918–928 (2013).
Sreekantan, L., Torregrosa, L., Fernandez, L. & Thomas, M. R. VvMADS9, a class B MADS-box gene involved in grapevine flowering, shows different expression patterns in mutants with abnormal petal and stamen structures. Funct. Plant Biol. 33, 877–886 (2006).
Yoon, J. et al. Genetic diversity and ecogeographical phylogenetic relationships among peach and nectarine cultivars based on simple sequence repeat (SSR) markers. J. Am. Soc. Hortic. Sci. 131, 513–521 (2006).
Marchese, A., Tobutt, K. R. & Caruso, T. Molecular characterisation of Sicilian Prunus persica cultivars using microsatellites. J. Hortic. Sci. Biotechnol. 80, 121–129 (2005).
Vendramin, E. et al. A unique mutation in a MYB gene cosegregates with the nectarine phenotype in peach. PLoS ONE 9, e90574 (2014).
Hauser, M.-T., Harr, B. & Schlötterer, C. Trichome distribution in Arabidopsis thaliana and its close relative Arabidopsis lyrata: molecular analysis of the candidate gene GLABROUS1. Mol. Biol. Evol. 18, 1754–1763 (2001).
López-Girona, E. et al. A deletion affecting an LRR-RLK gene co-segregates with the fruit flat shape trait in peach. Sci. Rep. 7, 6714 (2017).
Guo, J. et al. Comparative transcriptome and microscopy analyses provide insights into flat shape formation in peach (Prunus persica). Front. Plant Sci. 8, 2215 (2017).
Mandel, T. et al. The ERECTA receptor kinase regulates Arabidopsis shoot apical meristem size, phyllotaxy and floral meristem identity. Development 141, 830–841 (2014).
Mandel, T. et al. Differential regulation of meristem size, morphology and organization by the ERECTA, CLAVATA and class III HD-ZIP pathways. Development 143, 1612–1622 (2016).
Suzaki, T. et al. The gene FLORAL ORGAN NUMBER1 regulates floral meristem size in rice and encodes a leucine-rich repeat receptor kinase orthologous to Arabidopsis CLAVATA1. Development 131, 5649–5657 (2004).
Zhang, C., Bai, M. Y. & Chong, K. Brassinosteroid-mediated regulation of agronomic traits in rice. Plant Cell Rep. 33, 683–696 (2014).
Bommert, P. et al. Thick tassel dwarf1 encodes a putative maize ortholog of the Arabidopsis CLAVATA1 leucine-rich repeat receptor-like kinase. Development 132, 1235–1245 (2005).
Cao, K. et al. Genome-wide association study of 12 agronomic traits in peach. Nat. Commun. 7, 13246 (2016).
Skene, K. G. M. & Barlass, M. Studies on the fragmented shoot apex of GrapevineIV. Separation of phenotypes in a periclinal chimera in vitro. J. Exp. Bot. 34, 1271–1280 (1983).
Boss, P. K. & Thomas, M. R. Association of dwarfism and floral induction with a grape ‘green revolution’ mutation. Nature 416, 847–850 (2002).
Fisher, D. V. in British Columbia Fruit Growers Association Quarterly Report, 3–10 (BC, Canada, 1969).
Lapins, K. O. Inheritance of compact growth type in apple. J. Am. Soc. Hortic. Sci. 101, 133–135 (1976).
Wolters, P. J., Schouten, H. J., Velasco, R., Si-Ammour, A. & Baldi, P. Evidence for regulation of columnar habit in apple by a putative 2OG-Fe(II) oxygenase. New Phytol. 200, 993–999 (2013).
Zhang, Y., Zhu, J. & Dai, H. Characterization of transcriptional differences between columnar and standard apple trees using RNA-Seq. Plant Mol. Biol. Report 30, 957–965 (2012).
Krost, C. et al. Evaluation of the hormonal state of columnar apple trees (Malus x domestica) based on high throughput gene expression studies. Plant Mol. Biol. 81, 211–220 (2013).
Krost, C., Petersen, R. & Schmidt, E. R. The transcriptomes of columnar and standard type apple trees (Malus x domestica)—a comparative study. Gene 498, 223–230 (2012).
Nassar, A. M. K., Ortiz-Medina, E., Leclerc, Y. & Donnelly, D. J. Periclinal chimeral status of New Brunswick ‘Russet Burbank’ potato. Am. J. Potato Res. 85, 432 (2008).
Bethke, P. C. et al. History and origin of russet Burbank (Netted Gem) a sport of Burbank. Am. J. Potato Res. 91, 594–609 (2014).
Dorst, J. C. Two remarkable bud-sports in the potato variety rode star. Euphytica 1, 184–186 (1952).
Howard, H. W. A full analysis of a potato chimera. Genetica 40, 233–241 (1969).
Howard, H. W. The stability of L1-mutant periclinal potato chimeras. Potato Res. 14, 91–93 (1971).
Asseyeva, T. Bud mutations in the potato and their chimerical nature. J. Genet. 19, 1–26 (1927).
Hernandez, T. P., Hernandez, T. & Miller, J. C. in Proc Am Soc Hortic Sci, 430–434 (1964).
Sardos, J. et al. Evidence for spontaneous polyploidization in cassava Manihot esculenta Crantz. Plant Syst. Evol. 283, 203 (2009).
Sang, Ki,H., Vijaya, K. B. & Robert, A. Spontaneous somatic tetraploids in cassava. Jpn. J. Breed. 42, 303–308 (1992).
Carvalhol, R., Guerra, M. & Carvalho, P. Occurrence of spontaneous triploidy in Manihot esculenta Crantz. Cytologia 64, 137–140 (1999).
Vandenbroucke, H. et al. Somaclonal variants of taro (Colocasia esculenta Schott) and yam (Dioscorea alata L.) are incorporated into farmers’ varietal portfolios in Vanuatu. Genet. Resour. Crop Evol. 63, 495–511 (2016).
Lebot, V., Aradhya, K. M., Manshardt, R. & Meilleur, B. Genetic relationships among cultivated bananas and plantains from Asia and the Pacific. Euphytica 67, 163–175 (1993).
Karamura, D. Somatic mutations and their implications to the conservation strategies of the East African highland bananas (Musa spp.). Acta Hortic. 879, 615 (2010).
Lebot, V. & Levesque, J. Genetic control of kavalactone chemotypes in Piper methysticum cultivars. Phytochemistry 43, 397–403 (1996).
Petersen, R. & Krost, C. Tracing a key player in the regulation of plant architecture: the columnar growth habit of apple trees (Malus×domestica). Planta 238, 1–22 (2013).
Acknowledgements
The authors thank Bruce Veit, Robert Schaffer, Mandy Walker, Ed Morgan and Ruth Williams for helpful comments on the manuscript. This work was funded by the New Zealand Ministry of Business, Innovation and Employment, contract C11X1310 and internal funding by The New Zealand Institute for Plant and Food Research Limited.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Foster, T.M., Aranzana, M.J. Attention sports fans! The far-reaching contributions of bud sport mutants to horticulture and plant biology. Hortic Res 5, 44 (2018). https://doi.org/10.1038/s41438-018-0062-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41438-018-0062-x
This article is cited by
-
From buds to shoots: insights into grapevine development from the Witch’s Broom bud sport
BMC Plant Biology (2024)
-
The vast majority of somatic mutations in plants are layer-specific
Genome Biology (2024)
-
Potential of wild, underutilized Actinidia callosa and Actinidia strigosa from Northeast India for kiwifruit breeding
Genetic Resources and Crop Evolution (2024)
-
Transcriptomic and metabolomic analyses reveal CmMYB308 as a key regulator in the pink flower color variation of ‘Dante Purple’ chrysanthemum
Plant Cell Reports (2024)
-
High level of somatic mutations detected in a diploid banana wild relative Musa basjoo
Molecular Genetics and Genomics (2023)