Abstract
Purpose
Variants in NUS1 are associated with a congenital disorder of glycosylation, developmental and epileptic encephalopathies, and are possible contributors to Parkinson disease pathogenesis. How the diverse functions of the NUS1-encoded Nogo B receptor (NgBR) relate to these different phenotypes is largely unknown. We present three patients with de novo heterozygous variants in NUS1 that cause a complex movement disorder, define pathogenic mechanisms in cells and zebrafish, and identify possible therapy.
Methods
Comprehensive functional studies were performed using patient fibroblasts, and a zebrafish model mimicking NUS1 haploinsufficiency.
Results
We show that de novo NUS1 variants reduce NgBR and Niemann–Pick type C2 (NPC2) protein amount, impair dolichol biosynthesis, and cause lysosomal cholesterol accumulation. Reducing nus1 expression 50% in zebrafish embryos causes abnormal swim behaviors, cholesterol accumulation in the nervous system, and impaired turnover of lysosomal membrane proteins. Reduction of cholesterol buildup with 2-hydroxypropyl-ß-cyclodextrin significantly alleviates lysosomal proteolysis and motility defects.
Conclusion
Our results demonstrate that these NUS1 variants cause multiple lysosomal phenotypes in cells. We show that the movement deficits associated with nus1 reduction in zebrafish arise in part from defective efflux of cholesterol from lysosomes, suggesting that treatments targeting cholesterol accumulation could be therapeutic.
Similar content being viewed by others
INTRODUCTION
NUS1, located on 6q22.1, encodes the Nogo B receptor (NgBR), a highly conserved multifunctional protein. NgBR associates with the DHDDS gene product (CIT; cis-isoprenyltransferase), to form a heterodimeric complex with cis-prenyltransferase activity.1,2,3,4 This complex is necessary for the biosynthesis of polyprenol and dolichol lipids, with the latter serving as the lipid carrier for precursors of the N-linked glycosylation pathway.5 The C-terminus of NgBR also interacts with and stabilizes NPC2, the soluble lysosomal protein required for proper efflux of cholesterol out of lysosomes to the ER.6 Lastly, NgBR is the receptor for Nogo B, a ligand that is highly expressed in the vasculature and facilitates the chemotaxis and adhesion of endothelial cells.7,8,9
The relevance of this gene to human health was recently highlighted by the identification of NUS1-CDG (OMIM 617082) patients bearing pathogenic homozygous variants in this gene.10 Fibroblasts from these patients displayed defects in N-glycosylation and intralysosomal cholesterol storage resembling pathogenic variants in the NPC1 and NPC2 genes. Individuals with de novo, heterozygous, loss-of-function variants in NUS1 have also been reported in association with autosomal dominant type 55 intellectual disability with seizures (OMIM 617831).11 NUS1 is also a proposed contributor to Parkinson disease (PD), as an increased frequency of missense variants within this gene has been observed in patients with early onset PD.12,13 More recent studies, however, draw this connection into question.12,14,15 Other genetic studies have found de novo NUS1 variants in patients with complex dystonia, ataxia, tremor, and epilepsy, reinforcing the biological importance of this gene in neuronal and/or neuromuscular systems.16,17,18,19 Despite the growing number of patients with movement phenotypes bearing NUS1 variants, the mechanistic basis for these phenotypes is largely lacking. Moreover, it is not known whether these reported heterozygous variants cause glycosylation and/or lysosomal defects in patient cells.
Here we report three patients bearing de novo, heterozygous NUS1 variants that present with a range of neurological manifestations and movement phenotypes. Functional studies in two of the patient cell lines revealed a spectrum of lysosomal-related defects consistent with the functions mediated by NgBR. Knockdown of nus1 in zebrafish caused altered swimming behaviors, neuronal cholesterol storage, and impaired turnover of a lysosomal integral membrane protein. Congruent with a role for intracellular cholesterol accumulation in the development of these phenotypes, treatment of nus1-depleted embryos with 2-hydroxypropyl-ß-cyclodextrin significantly improved the phenotypic and molecular consequences. These findings provide insight into the mechanistic basis for the movement phenotypes associated with heterozygous NUS1 variants. The implications of these results with regard to the mechanistic basis and clinical management of the neurological manifestations in these patients are discussed.
MATERIALS AND METHODS
The methods for cell-based analyses are found in the Supplementary files.
Biochemical experiments in zebrafish
For western blots, embryos were manually deyolked and harvested at the time points indicated. Protein concentration in detergent lysates was determined using the micro-BCA assay. The primary antibodies included anti-NgBR (Abcam, ab168351, 1:500), anti-NPC2 (1:1,000, R&D Systems, AF8644) and anti-Cherry (catalog number ab167453, Abcam). Western blots were developed using Clarity Western ECL Substrate (Bio-Rad) and analyzed on the Bio-Rad ChemiDoc Imaging System. For Western analyses of Lamp1 following heat shock, 4-dpf embryos were incubated in 39 °C embryo water for 30 minutes to induce expression. Embryos were subsequently harvested at the indicated time points post–heat shock. For filipin staining, 6–7 dpf embryos were fixed with 4% paraformaldehyde at 4 °C overnight. Fixed embryos were rinsed with phosphate buffered saline and stained with 0.5 mg/mL of filipin (Sigma, F4767-1MG) and 1% goat serum for 2.5 hours at room temperature. Stained embryos were mounted in 0.8% hanging agarose drops. Embryos were imaged with a 40× water immersion objective (N.A.1.15) on an Olympus FV3000 laser scanning confocal microscope. For treatment with 2-hydroxypropyl-beta-cyclodextrin (ßCD, Sigma, SLBX3717), 100 mM stock solution was generated in water. Embryos were treated at time points indicated by adding ßCD at a final concentration of 2.5 mM to their growth medium. Projections were generated using the ImageJ software (NIH) and processed using Adobe PhotoShop (CS6).
Antisense morpholino injections and Zebrabox
Morpholino knockdown of nus1 was performed using a previously validated translation-blocking morpholino (5′- ACACCATCTCATACAGCGAAGCCAT-3′) purchased from Gene Tools, LLC (Eugene, OR, USA). The degree of morpholino knockdown was assayed by western blot using the anti-NgBR antibody following injection of 0.3–1.0 µM reagent into 1-cell stage embryos; 4–5 dpf, larvae were placed one per well into 12-well plates containing 2 ml embryo medium. T Locomotor activity was monitored 5–8 dpf using the Zebrabox System (ViewPoint Inc., Toronto, Canada). Plate was placed into the sensory deprivation chamber, desensitized for 15 minutes, and behavior subsequently recorded at 10-minute intervals. The low detection threshold was set to 20. The large activity threshold was set to 8 and the inactive threshold was set to 4. Data was analyzed using GraphPad Prism.
RESULTS
Clinical description of patients
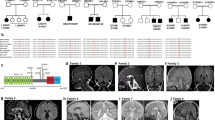
The clinical features and exome sequencing results of the patients are summarized in Supplemental Table 1. The technical details of exome sequencing analysis of the different variants for pathogenicity are provided in Supplemental Tables 2 and 3, respectively. The position of these variants within the protein and gene structures is depicted in Supplemental Fig. 1.
Patient 1
Patient 1 is a now 14-year-old female who presented for a genetics evaluation at age 12 due to concerns for possible spinocerebellar ataxia. She was noted to have a history of borderline intellectual disability, seizures, cerebellar intention tremor, ocular flutter, and mild gait ataxia. Early developmental milestones were normal. She has learning difficulties in school and is noted to be 3-4 grades behind her peers with a full-scale IQ of 76. She has had two major motor seizures along with occasional prolonged absence seizures. Her tremor was first noted at 3 years old and is stable. Ocular movements consist of her eyes moving horizontally back and forth with occasional upward movements. She had a fine tremor with mild gait ataxia and difficulty with tandem walking. Reflexes were normal. She has a normal brain magnetic resonance image (MRI). Electroencephalograms (EEGs) revealed mild generalized background slowing with frequent, often low voltage and poorly formed, bifrontal sharp wave discharges occurring in isolation or as runs of 2.5 Hz during sleep. She had normal autosomal dominant and autosomal recessive ataxia gene panels, as well as a normal serum transferrin test. Exome sequencing revealed a de novo heterozygous c.734G>T (p.Gly245Val) variant in the NUS1 gene.
Patient 2
Patient 2 is a 35-year-old female who was initially evaluated at 30 years old with a history of moderate intellectual disability, seizures, tremor, and slight gait ataxia. Early developmental milestones were delayed with her ability to say single words starting at 3 years old and phrases at 5 years old. She had an initial absence seizure at 4 years old in which she went limp and was unresponsive. She had no subsequent seizures until she had a grand mal seizure at 14 years old. She had three seizures between 14 and 30 years old, all related to medication changes. At 22 years old, she developed a tremor in her hands and feet. An EEG at 28 years old showed left cortical irritation with bursts of spikes, excessive slowing bilaterally and bursts of irregular delta slowing bilaterally. She had a normal brain MRI at 31 years old. Exome sequencing identified a de novo heterozygous c.752T>G (p.Leu251*) variant in the NUS1 gene.
Patient 3
Patient 3 is a now 5-year-old male who had a genetics evaluation at 3 years old for a history of developmental delay, seizures, and tremor. He has mild to moderate motor delays related to balance issues and moderate to severe speech delay. Seizure activity began at 2 years old with a generalized tonic–clonic seizure associated with fever. He has a history of myoclonic seizures and has more recently developed complex partial seizures that occur a few times a week to multiple times in a day, especially on warmer days. A tremor involving his upper extremities developed at 2 years old. The tremor is worse after seizure activity but improved after treatment with topiramate. He has not developed ataxia but was noted to have dysarthria. No evidence of hypotonia or other neurologic findings were identified. Subcortical parietal gliosis was noted on a brain MRI. EEG revealed abnormal bifrontal or generalized epileptiform discharges. Exome sequencing revealed a de novo heterozygous c.415 + 1G>A variant in the NUS1 gene.
Functional studies on patient fibroblasts
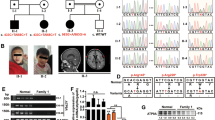
To define the molecular consequences of the NUS1 variants, two patient fibroblast lines (from P1 and P2) were used to investigate potential glycosylation defects, intralysosomal cholesterol storage, and the impact of variants on NgBR protein levels. Western blot analysis revealed a 50% reduction in the steady-state level of NgBR protein in both patient cell lines (Fig. 1a), supporting the likelihood these variants cause loss of function either through altered transcript abundance or decreased protein stability. Quantitative reverse transcription polymerase chain reaction (RT-PCR) of P1 cells showed a 30% reduction in NUS1 transcript abundance (Fig. 1b). Cloning and sequencing of amplicons indicate 64% of P1’s NUS1 messenger RNA (mRNA) is wild type (WT), while 36% contains the variant. In P2 cells no reduction in NUS1 transcript abundance was noted and 58% of the mRNA was WT. However, P2 generates a premature termination codon at position 251 and gel-based analyses of transcripts revealed a shorter species, indicating aberrant mRNA may be degraded by nonsense-mediated decay. A lower molecular weight band also observed in the NgBR western blot likely represents the truncated protein (Fig. 1a). The recently published crystal structures of the NUS1 and DHDDS gene products suggest the substitution of valine for glycine at position 245 in P1 likely disrupts the stability/folding of three ß-sheet structures that converge near that position (Supplemental Fig. 2).1,2
(a) Representative western blot for NgBR in wild type (WT) and patient fibroblasts and quantitation of NgBR levels relative to ß-actin (n = 3). Error = S.E.M. Statistical analysis was performed using a Dunnett’s test. (b) Quantitative polymerase chain reaction (PCR) analysis of NUS1 transcripts in WT and P1 fibroblasts (n = 3). The number of WT vs. variant alleles is shown. (c) Analysis of total dolichol and polyprenol levels in WT and patient fibroblasts. Analysis of the two patient lines (P1; n = 5 and P2; n = 3) were performed at different times using separate WT cells. Statistical analysis was performed using an unpaired Student’s t-test. (d) Representative western blots for ICAM1 and LAMP2 in WT and patient fibroblasts and quantitation of protein levels relative to HDAC1 (n = 3). Error = S.E.M. Statistical analysis was performed using a Dunnett’s test. (e) Representative western blot for NPC2 in WT and patient fibroblasts and quantitation of levels relative to ß-actin (n = 3). Error = S.E.M. Statistical analysis was performed using a Dunnett’s test. (f) Representative images of filipin-stained WT and patient fibroblasts and quantitation of pixel intensity in at least 40 different regions across 20 different cells. Error = S.E.M. Statistical analysis was performed using a Dunnett’s test. Scale bar = 10 µm. (g) Protein-normalized activity of acid-ß-glucosidase, ß-hexosaminidase, ß-glucuronidase, and ß-galactosidase in WT and patient fibroblasts. Arbitrary fluorescence units are plotted. Statistical analysis was performed using a Dunnett’s test. For all statistics *p < 0.05, **p < 0.01, ***p < 0.001. Red boxes indicate the additional correction (Dunnett’s test) was applied for comparison to a single control group.
To address the functional consequences of the NUS1 variants, we first asked whether the reduction in NgBR is sufficient to decrease polyprenol/dolichol abundance in patient cells. Lipid analysis was performed on control and patient fibroblasts. The levels of total polyprenol and dolichol lipids were diminished (Fig. 1c) in both patient fibroblast lines, suggesting that a 50% reduction in NgBR protein is sufficient to impact biosynthesis of these precursors. Next, possible glycosylation and lysosomal phenotypes were assessed. Western blot for several glycoprotein markers, including an integral lysosomal membrane protein (LAMP2) and the cell adhesion molecule (ICAM1), demonstrated either increased electrophoretic mobility or reduced abundance in the patient cells (Fig. 1d; Supplemental Fig. 3a). The lack of significant shift in electrophoretic mobility of cellular LAMP2 protein and normal serum transferrin profiles do not support biosynthetic glycosylation defects in these patients. The reduction in ICAM1 abundance in patient’s cells is interesting though it is not a specific finding and can be related to other NgBR-related mechanism. Furthermore, qPCR analyses also showed no increase in transcript abundance of multiple endoplasmic reticulum (ER) stress genes in patient cells, suggesting a stress response due to abundant protein underglycosylation is not present (Supplemental Fig. 3b).
Association between NgBR and the lysosomal protein NPC2 has been shown to stabilize NPC2, in turn modulating intracellular cholesterol transport and homeostasis.6 Western blot analysis showed that the steady-state levels of NPC2 were decreased in both patient lines compared with control fibroblasts, with P2 fibroblasts more strongly impacted (Fig. 1e). The reductions in NPC2 correspond with increased accumulation of lysosomal cholesterol, as assessed by filipin staining. Although the reduction in NPC2 is less pronounced in P1 cells, cholesterol storage was more apparent in these cells compared with P2 (Fig. 1f). Similar increases in cholesterol accumulation were also noted in NgBR +/− mouse embryonic fibroblasts.6 Together these findings reinforce the idea that cellular phenotypes occur in NUS1 heterozygotes due to haploinsufficiency.
To more broadly assess an impact on lysosomal function in patient cells, the activity of multiple glycosidases was profiled. Acid-ß-glucosidase and ß-hexosaminidase activity was reduced in both the P1 and P2 fibroblasts but the activity of ß-galactosidase and ß-glucuronidase was either unaffected or slightly elevated (Fig. 1g). It is not clear why acid-ß-glucosidase and ß-hexosaminidase activity is selectively reduced, but may reflect NgBR’s ability to interact with and stabilize additional lysosomal proteins besides NPC2. For acid-ß-glucosidase, this reduction may also relate to impaired trafficking to the lysosome via a pathway distinct from the other soluble hydrolases.20,21,22 Together, these data demonstrate that de novo loss-of-function variants in NUS1 cause multiple lysosomal defects in patient cells.
The observed/expected (o/e) constraint score for NUS1 generated by gnomAD is 0 with a 90% confidence interval of 0–0.25 indicating that haploinsufficiency of this gene is not tolerated.23 While this supports haploinsufficiency as a pathogenic mechanism, we next aimed to formally rule out dominant negative effects as the basis for the cellular phenotypes. WT NUS1 DNA or NUS1 DNA bearing the c.734G>T (p.Gly245Val) variant in P1 were overexpressed in HeLa cells and glycosylation and cholesterol accumulation were assessed (Supplemental Fig. 4). As a transfection control, a pcDNA3.1 construct expressing GFP was also utilized. Transfection of both NUS1 DNA vectors caused robust expression of NUS1 protein in the cells (Supplemental Fig. 4A). Although a decrease in the steady-state level of the NPC2 protein was noted in both the WT and Gly245Val overexpressing cells, this is likely a function of transfection, as the GFP construct produced comparable decreases. Notably, neither WT or p.Gly245Val NgBR overexpression in HeLa cells generated any intralysosomal cholesterol storage (Supplemental Fig. 4B). In addition, no detectable impact on glycoprotein glycosylation was observed in HeLa cells overexpressing WT and Gly245Val NgBR protein, as judged by ICAM1 and LAMP2 western blot in transfected cells (Supplemental Fig. 4C). Together, these data support intolerance of this gene to loss-of-function variants and haploinsufficiency as a pathogenic mechanism.
Zebrafish-based functional studies
Western blot analyses of NgBR in zebrafish embryos 1–5 days postfertilization (dpf) show the full length protein is abundant in early development (Fig. 2a). Several developmentally regulated lower molecular weight forms, possibly resulting from alternative protein processing, were noted. To ask whether NUS1 haploinsufficiency explains the clinical features of P1 and P2, we utilized an antisense morpholino to tune the knockdown of nus1 expression in developing embryos. Dose-dependent reductions were achieved using 0.3–1.0 µM morpholino, with 0.5 µM consistently reducing steady-state levels ~50% from 1 to 6 dpf (Fig. 2b,c). Although no overt physical phenotypes were noted in nus1 morphants (Fig. 2d), several differences in their motility and swimming behaviors emerged 4–5 dpf. Unlike WT controls, which swim continuously, nus1 morphants move erratically, stopping and starting often. To objectively measure differences between WT and morphant embryo swim behaviors we employed the Zebrabox, a behavioral analysis system that utilizes a high-speed camera housed in a sensory deprived chamber, to document and quantitate animal motility (Fig. 2e). Five dpf embryos were placed one per well in 12-well dishes and sensory deprived in the enclosed recording chamber for 15 minutes. Swim behavior was subsequently recorded in 10-minute intervals. Using this method, swim behaviors of WT and nus1 morphants were recorded once daily for 4 consecutive days. Traced paths of individual swim events revealed differences in swim speed, distance swam, the number of swim events initiated, and overall swim trajectory between WT and nus1 morphants. Most notably, nus1 morphants spent 40% more time swimming at high velocity (red paths) than WT embryos (Fig. 2e). As a result, morphant embryos swam up to 40% further than WT embryos at each age analyzed. Although they spend more time swimming at high speed, nus1 morphants also swim further at slow velocity (green paths). Importantly, in both categories, increased swim distance is associated with twice as many swim events. This indicates that nus1 morphants move by initiating numerous short bursts of high-speed swim events. In combination with the pattern of traced swim paths, these data suggest morphant motility is uncoordinated compared with WT controls. WT embryos consistently propagate a slow continuous path that traverses the well whereas ~25% of nus1-deficient animals swim in a circle around the periphery of the well.
(a) Western blots of 1–5 dpf whole-embryo lysates show NgBR is expressed during early development. Full length (FL) protein is indicated by a red arrowhead, as is a smaller processed piece. n = 4 experimental replicates, 20 embryos per biological sample per experiment. (b,c) Western blot show injection of 0.5 and 1.0 nM nus1 morpholino reduces NgBR abundance ~50% in embryos 1–6 dpf. n = 4 experimental replicates, 20 embryos per biological sample per experiment. (d) Bright field images of wild type (WT) and nus1 morphant (MO) embryos 5 and 7 dpf show no outward physical defects. n = 100 embryos analyzed from three experiments. Scale bar = 100 µm. (e) Schematic describes workflow for Zebrabox analyses of embryo motility. Image of four representative wells are shown from one plate containing WT and nus1 morphants. Traced swim patterns indicate the total path swam by the embryo, as well as swim speed. Red lines indicate a high velocity (fast) swim, while green lines indicate lower velocity (slow) swim. Images show nus1 morphants spend more time swimming at higher velocity than WT, often swimming in a circle around the well. (f) Graphs summarizing the distance swam by an individual embryo (one dot) while at the fast or slow speed from 5 to 8 dpf. (g) Graphs illustrate the number of new swim events initiated by an individual fish (one dot) while at the slow or fast swim speed from 5 to 8 dpf. (h) Graphs illustrate the percent of total embryos in 4–5 experiments that exclusively swim in a circle at the edge of the well. n = 50–100 embryos. For all quantitation: n = 50–75 embryos over 4–6 independent experiments. Error = S.E.M., significance calculated by the Student’s t-test where *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
To ask if N-linked glycosylation is affected in nus1 morphants, we utilized a transgenic zebrafish line that expresses a LAMP1-Cherry fusion protein under the control of a heat shock promoter (Fig. 3a). WT and morphant transgenic embryos were heat shocked 4 dpf, harvested 0.5–10 hours postshock, and Lamp1-Cherry protein analyzed by western blot with an anti-Cherry antibody (Fig. 3b,c). These analyses show that while the same amount of Lamp1-Cherry protein is initially made (see 0.5 hours), by 3 hours postshock its abundance is significantly higher in morphant embryos. Increased Lamp1-Cherry persists 10 hours postshock. Accumulation of Lamp1-Cherry protein suggests lysosomal degradation may be impaired in nus1 morphants. Although lower molecular weight products were noted, no definitive reduction in N-glycan occupancy was observed. Based on findings from patient fibroblasts we also addressed whether reductions in Npc2 and excess cholesterol could explain impaired Lamp1 proteolysis. Western blots for Npc2 show its abundance is decreased in morphant embryos 1–4 dpf (Fig. 3d, e). Further, confocal analyses of embryos stained with filipin show reductions in Npc2 are associated with increased cholesterol (Fig. 3g, h). Cholesterol accumulation was particularly evident in the hindbrains, spinal cords, and motor neurons of nus1 morphants. Both motor neuron cell bodies and axonal projections were heavily stained by filipin in nus1 morphants but not WT controls.
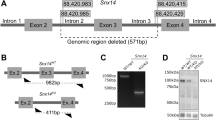
(a) Schematic illustrates transgene in zebrafish genome and the workflow for analyzing Lamp1 in wild type (WT) and nus1 morphant embryos. As shown, the transgene encodes a fusion between lamp1 and monomeric Cherry (mCherry); expression is controlled by the heat shock promoter (hsp701). (b,c) Western blot analysis and graphic quantitation of Lamp1 over a time course (0.5–10 hours) following heat shock induction of its expression show while similar amounts of Lamp1-mCherry are made post–heat shock, protein accumulates in nus1 morphants. (d,e) Western blot analysis and graphic quantitation of Npc2 abundance in WT and morphants (using either 0.5 or 1.0 µM morpholino) from 1 to 6 dpf. Red bracket indicate Npc2 doublet; both bands included in the quantitation of relative abundance. For all graphs: n = 3 experiments, with 25 embryo per sample per experiment. Error = S.E.M., significance calculated by the Student’s t-test where *p < 0.05, **p < 0.01. (f) Confocal analyses of filipin-stained WT and nus1 morphant (MO) embryos 6 dpf show cholesterol accumulation in the hindbrain, spinal cord (sc), and motor axons of morphant embryo tails. Yellow arrowheads highlight motor neuron cell bodies in ventral spinal cord and their associated axonal projections. Boxed region represents a region of interest (ROI) magnified in the right panel. Scale bars = 50 and 25 µM, respectively. (g) Graph represents quantitation of the average and maximum pixel intensities from the hindbrains, spinal cords, and axons of 10–15 embryos from n = 3 independent experiments. Error = S.E.M., significance calculated by the Student’s t-test where ***p < 0.001.
To ask whether cholesterol accumulation was associated with the movement phenotypes in nus1 morphants, embryos were treated 4 dpf with 2-hydroxypropyl-ß-cyclodextrin (ßCD). ßCD has been shown to facilitate cholesterol efflux from lysosomes reducing its accumulation in another Npc2-associated disorder, Niemann–Pick type C disease.24 Confocal analyses of 7-dpf filipin stained embryos (Fig. 4a, b) show treatment with ßCD reduces cholesterol accumulation in all three nus1-affected tissues, including hindbrain, spinal cord, and motor neurons. Western blot analysis of Lamp1-Cherry further demonstrates that reducing cholesterol storage via ßCD treatment improves lysosomal function in nus1 morphants (Fig. 4c–e). This is evidenced by decreased accumulation of Lamp1-Cherry in ßCD-treated morphants 5 and 10 hours postshock.
(a) Confocal analyses of filipin stained wild type (WT), nus1 morphant (MO), and ßCD-treated nus1 morphant embryos show reduced cholesterol accumulation in the hindbrain, spinal cord (sc), and motor axons of morphant embryos 7 dpf. Yellow arrowheads highlight axonal projections motor neurons in ventral spinal cords of nus1 morphants, which are not detected with filipin staining in either WT or ßCD-treated embryos. Scale bars = 50 µM. (b) Graph represents quantitation of pixel the average and maximum intensities from hindbrain, spinal cords, and axons of 15 embryos from n = 3 experiments. Error = S.E.M., significance calculated by the Student’s t-test where ***p < 0.001. (c) Schematic illustrates workflow for analysis of Lamp1 expression following ßCD treatment. (d,e) Western blot analysis and graphic quantitation of Lamp1 abundance (0.5–10 hours following heat shock induction of its expression) show Lamp1 accumulation is alleviated in nus1 morphants treated with 2.5 mM ßCD. n = 4 independent experiments with 25 embryos per sample per experiment. Error = S.E.M., significance calculated by the Student’s t-test where *p < 0.05, **p < 0.01.
Notably, nus1-depleted embryos showed significant improvement in several behavioral parameters following ßCD treatment (Fig. 5). This was particularly evident when animals were given two doses (first at 4 dpf and again at 6 dpf). Most notably, ßCD-treated nus1 morphants exhibit increased coordination, with fewer animals swimming in circles on the edge of the dish (Fig. 5b, c). Distance swam, swim speed, and numbers of swim events were also all significantly reduced in ßCD-treated morphants (Fig. 5b–d). Collectively these findings suggest that nus1-dependent cholesterol accumulation is at least partially responsible for the motility phenotypes associated with NUS1 haploinsufficiency.
(a) Schematic illustrates workflow for Zebrabox analyses of embryo motility following ßCD treatment. (b) Images show 4 representative wells from 12-well plates containing wild type (WT), nus1 morphant, or nus1 morphant embryos treated 1 or 2× with 2.5 mM ßCD. Traced swim patterns indicate the total path swam by the embryo, as well as swim speed. Red lines indicate a high velocity (fast) swim, while green lines indicate lower velocity (slow) swim. Images show treating nus1 morphants with ßCD reduces both their swim speed and the tendency to swim in a circle around the well. (c) Graph showing percent of total embryos swimming exclusively in a circle. (d) Graph showing distance swam by an individual embryo (black dot) at slow and fast swim speeds 5–8 dpf. Data show that treating nus1 morphants with ßCD twice significantly reduces swim distance, restoring WT-like distances. For all graphs: n = 3 experiments, with >25 embryos per “genotype” per condition. Error = S.E.M., significance calculated by the Dunnett’s test where correction for a single control sample was applied. *p < 0.05, **p < 0.01, ***p < 0.0001. Red boxes indicate an additional correction (Dunnett’s test) was applied for comparison to a single control group.
DISCUSSION
The present work expands the phenotypic consequences associated with NUS1 deficiency by demonstrating that de novo heterozygous loss-of-function variants in this gene are associated with seizures, ataxia, and movement phenotypes in affected individuals. We provide functional evidence that patient fibroblasts exhibit a spectrum of lysosomal defects, most clearly manifested by lower abundance of the lysosomal cholesterol efflux protein, NPC2, and accumulation of free cholesterol in this organelle. Furthermore, we establish in zebrafish that a comparable reduction in nus1 expression is sufficient to cause similar movement phenotypes that correlate with cholesterol accumulation in the brain and neuromuscular system. These phenotypes can be partially restored by treatment with 2-hydroxypropyl-ß-cyclodextrin, a chemical facilitator of cholesterol efflux from lysosomes. These findings indicate that at least some of the movement issues in human patients may be related to cholesterol accumulation in neuronal cells, and that therapies being developed for Niemann–Pick disease may be applicable to patients with NUS1 variants. Our results provide insight into the mechanistic basis for the movement phenotypes associated with loss-of-function alleles in this gene.
Our results in zebrafish embryos show abundant cholesterol storage in the brain and neuromuscular system of zebrafish embryos at half maximal levels of the nus1/NgBR protein. Biochemical analysis of the nus1-deficient embryos did not reveal any obvious defects in N-glycosylation occupancy on the LAMP1 protein but did show an increase in its steady-state amount. This is suggestive of impaired proteolysis and/or turnover within lysosomes, which could arise from altered activity of resident proteases due to the accumulation of free cholesterol.25,26 Recent studies suggest limited lysosomal proteolysis plays a role in the pathogenesis of PD.27 Loss of proteolytic capacity in lysosomes can impact processes such as mitophagy or other forms of autophagy, allowing failing organelles to accumulate and release toxic and reactive byproducts. While the movement phenotypes in our cohort do not overlap completely with PD, these data lend credence to the idea that neurons are particularly sensitive to disrupted lysosomal function.28 This is further supported by the observation that the most notable cholesterol accumulation in the embryos is within the central nervous and neuromuscular systems. Despite reduced proteolytic capacity in the nus1-depleted zebrafish embryos and the cholesterol storage noted in patient fibroblasts, we observed only selective impairment in the function of lysosomal glycosidases, in particular, reduced activity of acid-ß-glucosidase and ß-hexosaminidase. This specificity raises the possibility that NgBR may be responsible for stabilizing other lysosomal proteins besides NPC2. Likewise, decreased activity of these glycosidases may have secondary effects on the turnover of glycolipids and calcium homeostasis.29 Exploring this hypothesis, and the basis for the differences noted between the two patients, is a priority for future studies.
In light of the multiple functions carried out by NgBR, future studies will need to focus on defining whether and how the different functions of this protein contribute to the phenotypes seen in patients. Although alleviating cholesterol storage improved movement phenotypes of nus1-depleted embryos, it is unclear whether additional nus1 functions (including its role at the Nogo B receptor) also contribute. Since dolichol-based precursors are needed for other processes of other glycoconjugates such as the synthesis of GPI anchors, it is possible that impairment of these pathways is relevant. The sensitivity of the nervous system to alterations in many forms of glycosylation is well established, and could play a role in one or more of the phenotypes. Indeed, many proteins involved in neuronal development and function are heavily glycosylated, including voltage-gated ion channels, and can be impacted by limitations in dolichol-dependent precursors. In the event that deficits in glycosylation are more broadly implicated, increasing lipid flux away from de novo cholesterol biosynthesis and toward polyprenol/dolichol synthesis may be warranted. This can be done using inhibitors of squalene synthase, such as zaragozic acid, which would represent another possible therapeutic approach.30
Web Resources
Provean (protein variation effect analyser), http://provean.jcvi.org/index.php. PolyPhen2, http://genetics.bwh.harvard.edu/pph2/. MutationTaster, http://mutationtaster.org/. dbSNP, https://www.ncbi.nlm.nih.gov/snp/. gnomAD, http://gnomad.broadinstitute.org/. Mutalyzer, https://mutalyzer.nl/batch-jobs. ClinVar, https://www.ncbi.nlm.nih.gov/clinvar/.
Data and code availability
All variants described in this study have been deposited in ClinVar; accession numbers are VCV000981036, VCV000981034, VCV000981035 (Submission ID: SUB8124960; Organization ID: 1019).
References
Bar-El, M. L. et al. Structural basis of heterotetrameric assembly and disease mutations in the human cis-prenyltransferase complex. Nat. Commun. 11, 5273 (2020).
Edani, B. H. et al. Structural elucidation of the cis-prenyltransferase NgBR/DHDDS complex reveals insights in regulation of protein glycosylation. Proc. Natl. Acad. Sci. U. S. A. 117, 20794–20802 (2020).
Grabinska, K. A., Edani, B. H., Park, E. J., Kraehling, J. R. & Sessa, W. C. A conserved C-terminal RXG motif in the NgBR subunit of cis-prenyltransferase is critical for prenyltransferase activity. J. Biol. Chem. 292, 17351–17361 (2017).
Grabinska, K. A., Park, E. J. & Sessa, W. C. cis-Prenyltransferase: new insights into protein glycosylation, rubber synthesis, and human diseases. J. Biol. Chem. 291, 18582–18590 (2016).
Harrison, K. D. et al. Nogo-B receptor is necessary for cellular dolichol biosynthesis and protein N-glycosylation. EMBO J. 30, 2490–2500 (2011).
Harrison, K. D. et al. Nogo-B receptor stabilizes Niemann–Pick type C2 protein and regulates intracellular cholesterol trafficking. Cell Metab. 10, 208–218 (2009).
Park, E. J., Grabinska, K. A., Guan, Z. & Sessa, W. C. NgBR is essential for endothelial cell glycosylation and vascular development. EMBO Rep. 17, 167–177 (2016).
Miao, R. Q. et al. Identification of a receptor necessary for Nogo-B stimulated chemotaxis and morphogenesis of endothelial cells. Proc. Natl. Acad. Sci. U. S. A. 103, 10997–11002 (2006).
Teng, R. J. et al. Nogo-B receptor modulates angiogenesis response of pulmonary artery endothelial cells through eNOS coupling. Am. J. Respir. Cell Mol. Biol. 51, 169–177 (2014).
Park, E. J. et al. Mutation of Nogo-B receptor, a subunit of cis-prenyltransferase, causes a congenital disorder of glycosylation. Cell Metab. 20, 448–457 (2014).
Hamdan, F. F. et al. High rate of recurrent de novo mutations in developmental and epileptic encephalopathies. Am. J. Hum. Genet. 101, 664–685 (2017).
Chen, X. et al. Genetic analysis of NUS1 in Chinese patients with Parkinson’s disease. Neurobiol. Aging. 86, 202 e205–202 e206 (2020).
Guo, J. F. et al. Coding mutations in NUS1 contribute to Parkinson’s disease. Proc. Natl. Acad. Sci. U. S. A. 115, 11567–11572 (2018).
Bustos, B. I. et al. Replication assessment of NUS1 variants in Parkinson’s disease. Neurobiol. Aging. S0197-4580, 30389–4 (2020).
Yuan, L. et al. Extended study of NUS1 gene variants in Parkinson’s disease. Front. Neurol. 11, 583182 (2020).
Araki, K. et al. NUS1 mutation in a family with epilepsy, cerebellar ataxia, and tremor. Epilepsy Res. 164, 106371 (2020).
Den, K. et al. Recurrent NUS1 canonical splice donor site mutation in two unrelated individuals with epilepsy, myoclonus, ataxia and scoliosis—a case report. BMC Neurol. 19, 253 (2019).
Szafranski, P. et al. 6q22.1 microdeletion and susceptibility to pediatric epilepsy. Eur. J. Hum. Genet. 23, 173–179 (2015).
Wirth, T. et al. Increased diagnostic yield in complex dystonia through exome sequencing. Parkinsonism Related Disord. 74, 50–56 (2020).
Reczek, D. et al. LIMP-2 is a receptor for lysosomal mannose-6-phosphate-independent targeting of beta-glucocerebrosidase. Cell. 131, 770–783 (2007).
Zhao, Y., Ren, J., Padilla-Parra, S., Fry, E. E. & Stuart, D. I. Lysosome sorting of beta-glucocerebrosidase by LIMP-2 is targeted by the mannose 6-phosphate receptor. Nat. Commun. 5, 4321 (2014).
Zunke, F. et al. Characterization of the complex formed by beta-glucocerebrosidase and the lysosomal integral membrane protein type-2. Proc. Natl. Acad. Sci. U. S. A. 113, 3791–3796 (2016).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature. 536, 285–291 (2016).
Ory, D. S. et al. Intrathecal 2-hydroxypropyl-beta-cyclodextrin decreases neurological disease progression in Niemann–Pick disease, type C1: a non-randomised, open-label, phase 1-2 trial. Lancet. 390, 1758–1768 (2017).
Elrick, M. J. & Lieberman, A. P. Autophagic dysfunction in a lysosomal storage disorder due to impaired proteolysis. Autophagy. 9, 234–235 (2013).
Elrick, M. J., Yu, T., Chung, C. & Lieberman, A. P. Impaired proteolysis underlies autophagic dysfunction in Niemann–Pick type C disease. Hum. Mol. Genet. 21, 4876–4887 (2012).
Nguyen, M., Wong, Y. C., Ysselstein, D., Severino, A. & Krainc, D. Synaptic, mitochondrial, and lysosomal dysfunction in Parkinson’s disease. Trends Neurosci. 42, 140–149 (2019).
Wong, Y. C. et al. Neuronal vulnerability in Parkinson disease: should the focus be on axons and synaptic terminals? Mov. Disord. 34, 1406–1422 (2019).
Lloyd-Evans, E. et al. Niemann–Pick disease type C1 is a sphingosine storage disease that causes deregulation of lysosomal calcium. Nat. Med. 14, 1247–1255 (2008).
Haeuptle, M. A. et al. Improvement of dolichol-linked oligosaccharide biosynthesis by the squalene synthase inhibitor zaragozic acid. J. Biol. Chem. 286, 6085–6091 (2011).
Acknowledgements
We acknowledge the patients and their families for their willingness to participate in this study. This work was supported by the Greenwood Genetic Center and grants from the National Institutes of Health (5R01-GM086524-11 to R.S. and H.F-S; AI108819 to M.B.C). We acknowledge the support of the Hazel and Bill Allin Aquaculture Facility housed at the Greenwood Genetic Center and thank the facility staff for their excellent animal care.
Author information
Authors and Affiliations
Contributions
Conceptualization: R.S., H.F.-S., M.J.L. Data curation: S.Y., T.W., E.F.M., M.B.C., R.J.L., H.F.-S., R.S. Formal analysis: S.Y., T.W., R.J.L., H.F.-S., R.S. Funding acquisition: R.S., H.F.-S., M.B.C. Investigation: S.Y., T.W., K.W., R.J.L., E.F.M., H.F.-S. Methodology: S.Y., T.W., M.B.C., H.F.-S., R.S. Project administration: R.S., H.F.-S., M.J.L. Resources: C.S., K.M., P.G., N.R., D.C., M.J.L. Supervision: R.S., H.F.-S. Validation: T.W., H.F.-S. Visualization: S.Y., T.W., H.F.-S. Writing—original draft: R.S., H.F.-S., M.J.L., R.J.L. Writing—review & editing: R.S., H.F.-S.
Corresponding author
Ethics declarations
Ethics declaration
Informed consents were signed by the parents of the proband and other patients prior to participation in the research. All procedures were employed after being reviewed and approved by the Institutional Review Board, and compliant with practices, at the Greenwood Genetic Center (GGC). Handling and euthanasia of fish complied with policies of the GGC, as approved by the GGC’s Institutional Animal Care and Use Committee (permit #A2019-01-003-A1).
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Yu, SH., Wang, T., Wiggins, K. et al. Lysosomal cholesterol accumulation contributes to the movement phenotypes associated with NUS1 haploinsufficiency. Genet Med 23, 1305–1314 (2021). https://doi.org/10.1038/s41436-021-01137-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-021-01137-6
This article is cited by
-
Genomic insights into the comorbidity between type 2 diabetes and schizophrenia
Schizophrenia (2024)
-
NUS1 Variants Cause Lennox-Gastaut Syndrome Related to Unfolded Protein Reaction Activation
Molecular Neurobiology (2024)
-
Novel NUS1 variant in a Chinese patient with progressive myoclonus epilepsy: a case report and systematic review
Neurological Sciences (2023)