Abstract
Purpose
Proline Rich 12 (PRR12) is a gene of unknown function with suspected DNA-binding activity, expressed in developing mice and human brains. Predicted loss-of-function variants in this gene are extremely rare, indicating high intolerance of haploinsufficiency.
Methods
Three individuals with intellectual disability and iris anomalies and truncating de novo PRR12 variants were described previously. We add 21 individuals with similar PRR12 variants identified via matchmaking platforms, bringing the total number to 24.
Results
We observed 12 frameshift, 6 nonsense, 1 splice-site, and 2 missense variants and one patient with a gross deletion involving PRR12. Three individuals had additional genetic findings, possibly confounding the phenotype. All patients had developmental impairment. Variable structural eye defects were observed in 12/24 individuals (50%) including anophthalmia, microphthalmia, colobomas, optic nerve and iris abnormalities. Additional common features included hypotonia (61%), heart defects (52%), growth failure (54%), and kidney anomalies (35%). PrediXcan analysis showed that phecodes most strongly associated with reduced predicted PRR12 expression were enriched for eye- (7/30) and kidney- (4/30) phenotypes, such as wet macular degeneration and chronic kidney disease.
Conclusion
These findings support PRR12 haploinsufficiency as a cause for a novel disorder with a wide clinical spectrum marked chiefly by neurodevelopmental and eye abnormalities.
Graphic Abstract

Similar content being viewed by others

INTRODUCTION
Exome sequencing (ES) has become a mainstay in clinical genetics as a comprehensive, unbiased method for diagnosing genetic disease.1,2 However, a clinically useful molecular diagnosis is made in only 30–40% of cases.3,4 Recently, global data sharing efforts through large-scale platforms such as Matchmaker Exchange or GeneMatcher have enabled the rapid delineation of novel rare genetic syndromes by matching patients with variants in the same candidate gene and overlapping clinical features.5,6,7,8 Nonetheless, implicating candidate genes can be difficult due to low numbers of patients, the possibility of other contributing or confounding variants in other genes, and the potential for wide phenotypic variability.5,8 Proline Rich 12 (PRR12) is such a novel candidate gene recently encountered in exome studies of individuals with multisystem developmental disorders.9
PRR12 encodes a 211-kDa nuclear protein with suspected DNA-binding activity that is highly expressed in mouse and human brains, particularly in early development,10,11 and in the mouse visual system.12 The coding sequence of PRR12 is well conserved among vertebrates.9 PRR12 seems to be highly intolerant of loss-of-function (LOF) changes given that predicted LOF variants are exceedingly rare in the Genome Aggregation Database (gnomAD) (v3 and v2.1.1).13 The first report implicating PRR12 as a disease gene described a female patient with a de novo t(10;19)(q22.3;q13.33) reciprocal translocation disrupting both PRR12 and ZMIZ1 who presented with intellectual disability and neuropsychiatric changes.10 Consecutively, heterozygous, de novo, apparent loss-of-function variants in PRR12 were identified in three unrelated individuals who presented with global developmental delay and iris abnormalities.9
We collated clinical information from 21 additional individuals presenting with overlapping developmental features and variable eye abnormalities, harboring heterozygous apparent loss-of-function variants in PRR12 (three of whom had additional genetic findings), along with in silico evidence supporting its pathogenicity. We confirm a role for PRR12 in human disease and explore its variable clinical phenotype.
MATERIALS AND METHODS
Patients
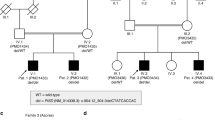
Patients 21–23 were described previously9 and the additional 21 patients were identified through international collaboration via GeneMatcher/Matchmaker Exchange.5,14 The whole cohort consists of 11 female and 13 male patients aged between 5 months and 36 years. All patients underwent chromosomal microarray testing and trio-based clinical ES, when possible, or alternatively, proband-based clinical ES with parental follow-up studies. Duo- and proband-based ES was performed on patients 3 and 12, respectively, due to parental unavailability. Patient 24 had chromosomal microarray analysis only and the deletion was a recombination product of maternal intrachromosomal insertion, as previously described.15 All PRR12 variants are reported on the NM_020719.3 (RefSeq)/ENST00000418929.7 (Ensembl) transcript. ES and analysis were performed at local commercial or research-based diagnostic laboratories.
RESULTS
Most of the observed PRR12 variants are predicted to cause LOF
Twenty-one distinct variants were identified among the 23 patients with PRR12 sequence variants: 12 frameshift, 6 nonsense, 1 splice-site, and 2 missense variants were observed (Fig. 1a). In addition, patient 24 carried a 3.352-Mb 19q13.33-13.41 deletion that consisted of 146 annotated genes, including PRR12, with breakpoints outside of its coding sequence (Fig. 1c). Sequencing of biological parents, when possible, revealed that all PRR12 variants were de novo. The PRR12 variant was absent in the mother of patient 3 and maternal half-sister of patient 12 (Supplementary Table 1). The frameshift and nonsense variants occurred within exons 3–7, out of the 14 exons of this gene. Frameshift variants introduce a premature termination codon (PTC) after 10 to 148 residues. The de novo splice-site variant (c.4891-2A>G; patient 19) disrupts the canonical AG splice acceptor site upstream of exon 8 and if exon 8 is skipped, the reading frame for downstream translation will shift by −1 and introduce a PTC after 12 residues. Complete exclusion of exon 8 was previously reported in melanocytes (HsaEX0050423; VastDB).16 Since the PTCs introduced by the frameshift, nonsense, and splice-site variants occur prior to the penultimate exon, the cognate messenger RNA (mRNA) expressed from those alleles is expected to undergo nonsense-mediated decay (NMD) and the variants are thus predicted to cause LOF. Finally, the de novo missense variants observed in two patients (c. 3505C>T [p.Arg1169Trp] in patient 15 and c.5909T>C [p.Leu1970Pro] in patient 20) were both predicted to be “probably damaging” in silico (PolyPhen-2 scores of 0.998 and 1.000, respectively). They both result in nonconservative substitutions; the former in the AT-hook domain and the latter in an uncharacterized region of PRR12 near the C-terminus of the polypeptide chain (Fig. 1b). In addition, there are four affected individuals reported in the DECIPHER database with de novo PRR12 variants consisting of two LOF variants and two missense variants: c.4726G>T (p.Glu1576*) (280416), c.2585_2586insG (p.Ala863Glyfs*74) (277812), c.5383C>T (p.Pro1795Ser) (260525), and c.4387C>T (p.Pro1463Ser) (417908).17 The Pro1795Ser and Pro1463Ser missense variants are predicted to both be “possibly damaging” (PolyPhen-2 score of 0.661 and 0.945, respectively). There is also an entry in DECIPHER of a patient with a 2.04-Mb deletion (arr[hg19] 19q13.33-q13.41(50,086,504_52,125,032)x1) (251777).
(a) Exon diagram of the PRR12 coding sequence with variants of the coding sequence shown on the larger isoform of PRR12. A schematic diagram of the shorter isoform is given below (ENST00000615927.1). The three variants described by Leduc et al.9 are highlighted in blue and the three PRR12 variants reported in the DECIPHER database are highlighted in red. (b) PRR12 protein domains with predicted variants of the protein sequence. The corresponding patient number is given with each variant. Underlined variants are excluded in the shorter isoform. The three variants described by Leduc et al.9 are highlighted in blue and the three PRR12 variants reported in the DECIPHER database are highlighted in red. (c) Chromosomal region corresponding to the microdeletion reported in patient 24 shown with OMIM genes with probability of loss-of-function intolerance score (pLI) > 0.7.
PRR12 is highly intolerant to LOF variation
According to gnomAD, predicted LOF variants in PRR12 are exceedingly rare in this large data set of individuals without severe pediatric disease.13 Indeed, among 2,383 and 1,963 distinct PRR12 variants reported in gnomAD versions v2.1.1 and v3, 1 and 2 predicted LOF variants were reported, respectively, each with an allele count of 1. The former variant (c.2851delC) causes frameshift and a PTC after 91 residues, much like the variants reported in this study. However, the latter two variants, c.2503_2520del and c.2512_2520del, are expected to cause short in-frame deletions of a few amino acids and are not expected to cause NMD. There is a splice isoform of PRR12 produced by alternative promoter usage within exon 4 and splicing within the exon (Fig. 1). According to gnomAD, these deletions overlap the downstream intron–exon boundary unique to this isoform and remove a noncanonical splice acceptor site. Subsequently, the remainder of exon 4 may be skipped, but the reading frame would still not be shifted. Thus, it remains unclear if the latter two variants are cause of LOF. The probability of being LOF intolerant (pLI) and observed/expected (o/e) constraint scores for PRR12 are 1.0 and 0.0 (0–0.05; 90% confidence interval [CI]), respectively, with a LOF observed/expected upper bound fraction (LOEUF) score of 0.051, supporting a strongly deleterious role for LOF variants. Furthermore, the missense constraint Z-score is +2.98 indicating that PRR12 is also intolerant to missense variation.13 Of note, no additional missense variants at Arg1169, Pro1795 or Leu1970 are reported in gnomAD. There is however a Pro1463Leu missense variant. There are also five structural variants in gnomAD SVs v2.1, consisting of four overlapping deletions within intron 6 that do not involve splice sites and one large 4.98-Mb inversion encompassing 184 genes with breakpoints outside of the PRR12 gene. These variants are unlikely to affect PRR12 expression or function.
A broad spectrum of overlapping anomalies observed in individuals with PRR12 predicted LOF variants and deletion
Developmental impairment is a consistent finding
All patients had documented developmental impairment: 17 patients had a diagnosis of global developmental delay and 3 and 4 patients had isolated motor and speech–language developmental delay, respectively (Table 1; Supplementary Table 1). Mild to severe intellectual disability (ID) was documented for all patients over the age of 7 years (n = 11), where these data were available (Table 1). The four individuals with de novo PRR12 variants and the additional individual with the microdeletion involving PRR12 reported in DECIPHER also all show developmental impairment.
A variety of structural eye defects are observed
We observed a striking variety of structural eye abnormalities that affected 50% (12/24) of patients (Table 1, Fig. 3a). The most severe eye defects were anophthalmia/microphthalmia, which was observed in patients 1, 11, 13, and 17. Patient 1, in particular, had bilateral anophthalmia with the absence of optic nerves, optic tracts, and the optic chiasm on magnetic resonance imaging (Supplementary Table 1, Fig. 2a). DECIPHER patient 280416 is a female with aplasia/hypoplasia of the optic nerve, iris coloboma, and unilateral microphthalmia. Interestingly, globe defects (anophthalmia or microphthalmia) were observed only among female patients although males and females were close to equally represented in this cohort. Also common was coloboma of the eye: 29% (7/24) of patients had one or more colobomas, which most commonly affected the iris. Other areas affected were the optic nerve, macula, choroid retina, and lens. Two patients (patients 13 and 21) had bilateral colobomas. In contrast to the three previously described individuals with PRR12 variants (patients 21–23) who all had iris abnormalities, these were much less common in this larger cohort: 3 of the 21 new patients reported in this study had iris colobomas and 1 had stellate irides. Other structural eye defects included retinal dysplasia, persistent pupillary membrane, complex Rieger anomaly, bilateral oblong optic nerves, optic nerve hypoplasia, and congenital hypertrophy of retinal pigment epithelium. Patients 3, 15, 16, and 24 had not received a complete ophthalmological assessment; hence less obvious eye defects, such as those affecting the posterior chamber, may have been missed on physical examination. In addition to structural defects, many patients had visual impairment (77%; 17/22) and strabismus (36%; 8/22), including intermittent types. There were 12 individuals (50%) with no documented structural eye abnormalities, in keeping with the variability of this phenotype in this cohort (Fig. 3a).
Dysmorphic features are highly variable among the individuals with available facial photographs and do not seem to confer a recognizable pattern. Globe defects, in forms of bilateral anophthalmia in patient 1 (a) and microphthalmia in patient 11 (e) are depicted. Common distinctive features including wide-set eyes, epicanthal folds, low-set ears, upturned tip of the nose, and thin vermilion of the lips are observed. (a) Patient 1. (b) Patient 3. (c) Patient 8. (d) Patient 10. (e) Patient 11. (f) Patient 12. (g) Patient 14. (h) Patient 16. (i) Patient 18. (j) Patient 19. (k) Patient 20. (l) Patient 24.
A filled square indicates the presence of the listed clinical feature and a blank square indicates absence. An X denotes that the presence of the listed feature has not been ascertained. (a) Summary of eye phenotypes within our cohort organized into four distinct categories, including those with no apparent defects. Listed above are the patient numbers corresponding to Table 1. The “iris abnormality” feature excludes iris coloboma. * No obvious abnormalities on physical examination; incomplete ophthalmological assessment. † Individuals with multiple genetic diagnoses. (b) Comparison of clinical features among individuals with PRR12 variants affecting the long isoform (patients 1, 2, 4, 5, 21, and 22) or both splice isoforms. This graph excludes individuals with multiple genetic diagnoses (patients 6, 7, 9, and 24). Rows depict the prevalence of each listed feature in each subset. FTT failure to thrive.
Additional common clinical features include anomalies from various systems
Commonly observed systemic abnormalities were congenital heart (52%; 12/23) and kidney (35%; 8/23) defects (Table 1). Among the 12 patients with congenital heart defects, 6 had atrial septal defects, 2 had ventricular septal defects, and 3 had pulmonary stenosis (patient 4 had two defects; Supplementary Table 1). Congenital kidney anomalies were relatively minor and included hydronephrosis, duplicated ureters, and vesicoureteral reflux. Cryptorchidism (unilateral or bilateral) was common among male patients (38%; 5/13). Growth phenotypes were also observed relatively commonly; history of failure to thrive was documented in 54% (13/24) of patients and 29% (7/24) had microcephaly. Although a recognizable facial pattern could not be discerned from patient photos (Fig. 2), some nonspecific dysmorphic facial features that were commonly observed (seen in ≥6/24, 25%) included wide-set eyes, epicanthal folds, low-set ears, upturned nasal tip, and thin vermillion of the lip(s) (Table 1, Supplementary Table 1). Interestingly, cleft palate was observed in four individuals (17%). Rare gastrointestinal abnormalities were also identified; intestinal malrotation in 2 (8%) and Meckel’s diverticulum in 1 (4%) individuals, respectively. A history of hypotonia during or beyond the neonatal period was commonly observed (61%; 14/23).
Coexisting variants in other disease genes were reported in some individuals in our cohort (Table 1, Supplementary Table 2). Patient 9 had macrocephaly in contrast to the smaller head size generally observed in the rest of the cohort (Supplementary Table 1). This patient carried a pathogenic PIK3CA variant known to cause an overgrowth disorder hallmarked by megalencephaly18 (Supplementary Table 2). Patient 6 was recently described in a report delineating the KDM6B-related disorder.19 Finally, patient 7 carried a likely pathogenic variant in LZTR1 (Noonan syndrome 10, OMIM 616564), providing a possible alternative explanation to history of failure to thrive, developmental delay, and dysmorphic features.
Reduced predicted expression of PRR12 is associated with acquired eye- and kidney-related disease
To assess the potential clinical consequences of PRR12 dysfunction, we retrieved data on PRR12 from a transcriptome-wide association study performed previously by Unlu et al.20 using PrediXcan analysis to identify clinical phenotypes associated with changes in predicted PRR12 expression, specifically when reduced. Cross-tissue analysis of the BioVU biobank (which consists of 25,000 SNP-typed European Americans with linked electronic health records) revealed several clinical phenotypes associated with whole-body reduction of predicted PRR12 expression (representative of constitutive de novo LOF variants) that were enriched for acquired eye- and kidney-related diseases (Table 2). Of the 30 phenotypes associated with reduced expression (as defined by an odds ratio [OR] per unit of standard deviation [SD] less than 1.00), 7 and 4 pertained to the visual or renal system, respectively. In alignment with the pLI, o/e, and LOEUF constraint scores reported in gnomAD, fewer associations with increased predicted PRR12 expression were noted, suggesting greater intolerance to loss of PRR12 function. PrediXcan analysis was used previously to link GRIK5 loss-of-function to eye and peripheral vascular disease.20 Similar to GRIK5, the 7 significant associations of eye phenotypes to reduced predicted PRR12 expression were also unlikely to have occurred by chance. Thus, there is a potential role for loss of PRR12 expression in acquired eye and kidney pathology, common findings in our cohort with predicted constitutional loss of PRR12 function.
DISCUSSION
A review of available clinical features in the total cohort of 24 individuals (and the 3 reported in DECIPHER), consisting of one deletion variant and almost exclusively predicted LOF variants in PRR12, a gene highly intolerant of such changes, revealed a consistent presentation of developmental delay/intellectual disability, and eye abnormalities. We noted marked variability in the type and severity of eye phenotypes with increased patient numbers, expanding the phenotype of consistent iris abnormalities reported previously in three patients.9 The larger cohort also allowed for the delineation of additional common systemic features, including congenital heart and kidney defects, hypotonia, failure to thrive, and microcephaly. A very recent report documents four additional truncating PRR12 variants in a cohort of individuals with microphthalmia/anophthalmia/coloboma, further supporting the impact of PRR12 loss in eye development.21
The microdeletion observed in patient 24 resulted in the heterozygous loss of 146 annotated genes, 86 of which are listed in OMIM, and 15 of which have associated disease phenotypes. However, only 2 of these 15 genes are seemingly highly intolerant of loss-of-function changes: NUP62 (pLI: 0.95) and PPP2R1A (pLI: 0.98). Biallelic variants in NUP62 are associated with autosomal recessive infantile striatonigral degeneration (OMIM 271930), thus heterozygous loss of this gene is likely noncontributory to this patient’s phenotype. PPP2R1A is associated with autosomal dominant intellectual disability (OMIM 616362) and its loss may have contributed to the developmental impairment observed in patient 24. None of 15 morbid genes are currently associated with eye, heart, or kidney abnormalities. When looking only at genes in this region that are somewhat intolerant of loss-of-function changes (pLI ≥ 0.76; the pLI of the remaining genes is ≤0.40) (n = 12) (Fig. 1c), PRR12 is one of the top candidates for this particular patient, especially given the presence of bilateral iris colobomata, which was commonly observed in individuals with sequence variants (Table 1). Altogether, the consistent clinical features among individuals with PRR12 sequence variants and a microdeletion including PRR12 strongly support the pathogenicity of PRR12 haploinsufficiency.
The high conservation of its amino acid sequence9 and intolerance to LOF and missense variants suggest that PRR12 likely serves an important, conserved biological function, but the gene remains poorly characterized. Specific functional studies are currently limited to expression analysis in the initial patient with the de novo translocation, and in fetal and adult brains in mice.10 However, functional information gleaned from publicly available data sets strongly supports the potential role of PRR12 in human disease.
PRR12 (also known as KIAA1205) is a ubiquitously expressed gene with highest levels of expression in the brain (particularly in the cerebellum and pituitary gland), the thyroid gland, and the female reproductive system (GTEx). It was shown previously that expression of the 212-kDa PRR12 protein product was restricted to the nucleus and was strongest in fetal E15 mice brains compared with adult brains.10 Similarly, across multiple brain structures, PRR12 RNA expression is elevated in fetal human samples compared with adult samples.11 The role of PRR12 in early neurodevelopment is further supported by its association with 4 of 5 promoter/enhancer regions (GeneHancer scores between 2.0 and 2.7) that exist in a poised chromatin state in several human embryonic and induced pluripotent stem cell lines and in neural progenitors cells.22,23 A poised state is thought to permit precise spatiotemporal regulation of genes during cell differentiation, especially during embryonic development, and has classically been associated with developmental genes, such as mFgf8 and mProk1.24,25 These findings suggest an important role for PRR12 in neural progenitor cells and early neurodevelopment.
In addition to the regular transcript, PRR12 also produces a shorter 130-kDa (1,215-aa) isoform via alternative splicing (ENST00000615927.1) that lacks exons 1–3 and the majority of exon 4 (Fig. 1). In contrast to the larger isoform, its expression seems to be elevated in adult brains compared with fetal brains. The complementary DNA (cDNA) of the short human isoform was also isolated from the brain.26 Given the differential subcellular localization and expression pattern between these isoforms in the brain, we can hypothesize that the short isoform has other neuronal functions in the adult brain. Alternative splicing is predicted to remove the mutated site for eight patients (patients 1, 2, 4, 5, 6, 7, 21, and 22) and to maintain expression of the short isoform from the mutant allele. Clinical features between this group and the remaining individuals whose variants are predicted to truncate both isoforms were not found to be significantly different (Fig. 3b). Growth phenotypes (failure to thrive and microcephaly) appeared more commonly, while kidney and heart defects appeared less commonly in the former group. However, there are too few individuals to make a meaningful conclusion. This finding supports that disruption of the nuclear function of the larger isoform is likely the basis of this developmental disorder, as suggested previously.9
The role of PRR12 in neural and eye development may involve its predicted ability to bind USP7,27 SOX2,28 and ESR2,29 as shown in publicly available protein interaction databases. Haploinsufficiency of USP7 (OMIM 616863) was recently shown to cause syndromic intellectual disability and developmental delay.30 Variants in SOX2 are a well-established cause of syndromic microphthalmia/clinical anophthalmia with variable defects of the optic nerve and/or central nervous system (OMIM 206900).31 ESR2 encodes a known estrogen receptor but is expressed in the developing eye of human embryos.32 While these predicted interactions remain to be experimentally proven, and the exact functional consequences are unknown, we speculate that substochiometric binding between PRR12 and these proteins, especially SOX2, may be a possible explanation for the occurrence of eye abnormalities. Further functional studies in animal models targeting this group of genes and their interactors would provide evidence into the pathogenesis of this novel disorder.
The significant phenotypic variability in neurodevelopmental and ophthalmological features within our cohort may be explained by other patient-, test-, and gene-related factors. Multiple molecular diagnoses, mostly identified with genome-wide sequencing, have been shown to occur in 3–5% of patients receiving a molecular diagnosis by these methods33,34 and was observed in three individuals in this cohort. The presence of these additional variants in genes associated with other multisystem disorders, as well as potential contributions from noncoding variants, limit our ability to ascribe phenotypes specifically to PRR12 loss of function. Also, patient 15 has a brother with a similar phenotype who does not share the de novo missense PRR12 variant that he harbors. As encountered with most of the recently identified disease genes, by taking the genotype-first approach based on ES (which has become increasingly available to a larger number of patients worldwide), we may have already captured milder forms of PRR12-related disorder in our large cohort. There are several well known single-gene disorders affecting both the eyes and kidneys, some of which are caused by defects in transcription factors with important roles in development, such as CHD7 in CHARGE syndrome (OMIM 214800), PAX2 in oculorenal syndrome (OMIM 120330), TBX22 in CHARGE-like syndrome (OMIM 302905), and SALL4 in acroreno-ocular syndrome (OMIM 607323). The clinical features in these syndromes are also notoriously variable, mostly evidenced by the huge clinical spectrum of CHARGE syndrome,35 as well as the variable eye findings in PAX2- and SALL4-related disorders, where presentations range from microphthalmia, to many types of colobomas, dysplastic optic discs, or other ocular features.36,37
PRR12, similar to the above genes, likely has roles in early development and remains a reasonable target to study its role in genetic, and perhaps epigenetic, regulation. PRR12 has suspected DNA-binding activity owing to its two predicted AT-hook domains and it may participate in gene regulation given that it is a nuclear-restricted protein.10 Interestingly, the top 100 genes that coexpress with PRR12 in humans are enriched for genes involved in transcription and its regulation (p < 10−18), chromatin regulators (p = 1.5 × 10−7), and SET domain-containing proteins (p = 9.7 × 10−7) (COXPRESdb7; DAVID).38,39 Moreover, similar terms are enriched among the top 200 genes that coexpress with PRR12 across different brain tissues from 8 weeks postconception to 40 years of age (r ≥ 0.748) and also include zinc finger proteins (p = 7.0 × 10−19), BAH domain-containing proteins (p = 2.1 × 10−6), and bromodomain-containing proteins (p = 1.0 × 10−6). These classes of proteins include transcription factors and chromatin regulators and may function in a complex network involving PRR12 to establish specific transcriptional programs. Of interest are the SET domain-containing proteins (which methylate histone lysine residues) and the bromodomain-containing proteins (which recognize acetylated lysine residues, including those on histones), which suggest a potential mechanism involving histone modifications and downstream chromatin regulation. Curiously, Lysine-402 of PRR12 is also acetylated and may be bound by coexpressed bromodomain proteins to modulate or mediate its function(s).9,40 The exact role of PRR12 in transcriptional regulation remains unclear and will require further functional studies to elucidate. While we can only speculate about the interactions between PRR12 and coexpressed genes, these associations are suggestive of a role for PRR12 in widespread gene regulation via epigenetic mechanisms.
We provide strong genetic evidence to indicate that haploinsufficiency of PRR12, a gene with potential roles in neurodevelopment and gene regulation, causes a neurodevelopmental disorder with variable features, including eye, kidney, and heart anomalies, and growth failure. Further studies are needed to establish the exact genotype–phenotype correlation and potential effects of other modifiers. These may include analyses of epigenetic modifications in these patients to help identify an epigenomic signature to aid in molecular diagnosis and variant interpretation. Functional studies in animal models, such as mice and zebrafish, will be invaluable in elucidating the molecular function of PRR12 and the exact pathogenetic mechanisms of PRR12 loss-of-function in human disease.
Data availability
Any materials, data, and data sets produced from or used for this study will be made available upon request to the authors. The PRR12 variants reported by GeneDx have been submitted to ClinVar. SUB numbers with corresponding patient numbers are: Patient 1: SUB9233172; Patient 2: SUB9233619; Patient 6 SUB9246261; Patient 7: SUB9246275; Patient 9: SUB9246289; Patient 12: SUB9246294; Patient 14: SUB9246299; Patient 17: SUB9246311.
References
Boycott, K. M. et al. International cooperation to enable the diagnosis of all rare genetic diseases. Am. J. Hum. Genet. 100, 695–705, https://doi.org/10.1016/j.ajhg.2017.04.003 (2017).
Retterer, K. et al. Clinical application of whole-exome sequencing across clinical indications. Genet. Med. 18, 696–704, https://doi.org/10.1038/gim.2015.148 (2016).
Trujillano, D. et al. Clinical exome sequencing: results from 2819 samples reflecting 1000 families. Eur. J. Hum. Genet. 25, 176–182, https://doi.org/10.1038/ejhg.2016.146 (2017).
Dillon, O. J. et al. Exome sequencing has higher diagnostic yield compared to simulated disease-specific panels in children with suspected monogenic disorders. Eur. J. Hum. Genet. 26, 644–651, https://doi.org/10.1038/s41431-018-0099-1 (2018).
Sobreira, N. L. M. et al. Matchmaker Exchange. Curr. Protoc. Hum. Genet. 95, 9.31.1–9.31.15, https://doi.org/10.1002/cphg.50 (2017).
Au, P. Y. B. et al. GeneMatcher aids in the identification of a new malformation syndrome with intellectual disability, unique facial dysmorphisms, and skeletal and connective tissue abnormalities caused by de novo variants in HNRNPK. Hum. Mutat. 36, 1009–1014, https://doi.org/10.1002/humu.22837 (2015).
O’Donnell-Luria, A. H. et al. Heterozygous variants in KMT2E cause a spectrum of neurodevelopmental disorders and epilepsy. Am. J. Hum. Genet. 104, 1210–1222, https://doi.org/10.1016/j.ajhg.2019.03.021 (2019).
Bruel, A.-L. et al. 2.5 years’ experience of GeneMatcher data-sharing: a powerful tool for identifying new genes responsible for rare diseases. Genet. Med. 21, 1657–1661, https://doi.org/10.1038/s41436-018-0383-z (2019).
Leduc, M. S. et al. De novo apparent loss-of-function mutations in PRR12 in three patients with intellectual disability and iris abnormalities. Hum. Genet. 137, 257–264, https://doi.org/10.1007/s00439-018-1877-0 (2018).
Córdova-Fletes, C. et al. A de novo t(10;19)(q22.3;q13.33) leads to ZMIZ1/PRR12 reciprocal fusion transcripts in a girl with intellectual disability and neuropsychiatric alterations. Neurogenetics. 16, 287–298, https://doi.org/10.1007/s10048-015-0452-2 (2015).
Miller, J. A. et al. Transcriptional landscape of the prenatal human brain. Nature. 508, 199–206, https://doi.org/10.1038/nature13185 (2014).
Bult, C. J., Blake, J. A., Smith, C. L., Kadin, J. A., Richardson, J. E. & Mouse Genome Database Group. Mouse Genome Database (MGD) 2019. Nucleic Acids Res. 47, D801–D806, https://doi.org/10.1093/nar/gky1056 (2019).
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature. 581, 434–443, https://doi.org/10.1038/s41586-020-2308-7 (2020).
Sobreira, N., Schiettecatte, F., Valle, D. & Hamosh, A. GeneMatcher: a matching tool for connecting investigators with an interest in the same gene. Hum. Mutat. 36, 1928–1930, https://doi.org/10.1002/humu.22844 (2015).
Gu, S. et al. Mechanisms for complex chromosomal insertions. PLoS Genet. 12, e1006446, https://doi.org/10.1371/journal.pgen.1006446 (2016).
Tapial, J. et al. An atlas of alternative splicing profiles and functional associations reveals new regulatory programs and genes that simultaneously express multiple major isoforms. Genome Res. 27, 1759–1768, https://doi.org/10.1101/gr.220962.117 (2017).
Firth, H. V. et al. DECIPHER: Database of Chromosomal Imbalance and Phenotype in Humans Using Ensembl Resources. Am. J. Hum. Genet. 84, 524–533, https://doi.org/10.1016/j.ajhg.2009.03.010 (2009).
Mirzaa, G. et al. PIK3CA-associated developmental disorders exhibit distinct classes of mutations with variable expression and tissue distribution. JCI Insight. 1, e87623 (2016).
Stolerman, E. S. et al. Genetic variants in the KDM6B gene are associated with neurodevelopmental delays and dysmorphic features. Am. J. Med. Genet. A. 179, 1276–1286, https://doi.org/10.1002/ajmg.a.61173 (2019).
Unlu, G. et al. GRIK5 genetically regulated expression associated with eye and vascular phenomes: discovery through iteration among biobanks, electronic health records, and zebrafish. Am. J. Hum. Genet. 104, 503–519, https://doi.org/10.1016/j.ajhg.2019.01.017 (2019).
Reis, L. M. et al. Dominant variants in PRR12 result in unilateral or bilateral complex microphthalmia. Clin. Genet. https://doi.org/10.1111/cge.13897 (2020).
Fishilevich, S. et al. GeneHancer: genome-wide integration of enhancers and target genes in GeneCards. Database (Oxford). 2017, bax028 (2017).
Zerbino, D. R. et al. Ensembl 2018. Nucleic Acids Res. 46, D754–D761, https://doi.org/10.1093/nar/gkx1098 (2018).
Bernstein, B. E. et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell. 125, 315–326, https://doi.org/10.1016/j.cell.2006.02.041 (2006).
Harikumar, A. & Meshorer, E. Chromatin remodeling and bivalent histone modifications in embryonic stem cells. EMBO Rep. 16, 1609–1619, https://doi.org/10.15252/embr.201541011 (2015).
Nagase, T. et al. Prediction of the coding sequences of unidentified human genes. XV. The complete sequences of 100 new cDNA clones from brain which code for large proteins in vitro. DNA Res. 6, 337–345, https://doi.org/10.1093/dnares/6.5.337 (1999).
Havugimana, P. C. et al. A census of human soluble protein complexes. Cell. 150, 1068–1081, https://doi.org/10.1016/j.cell.2012.08.011 (2012).
Kim, B. R. et al. Identification of the SOX2 interactome by BioID reveals EP300 as a mediator of SOX2-dependent squamous differentiation and lung squamous cell carcinoma growth. Mol. Cell. Proteomics. 16, 1864–1888, https://doi.org/10.1074/mcp.M116.064451 (2017).
Giurato, G. et al. Quantitative mapping of RNA-mediated nuclear estrogen receptor β interactome in human breast cancer cells. Sci Data. 5, 180031, https://doi.org/10.1038/sdata.2018.31 (2018).
Fountain, M. D. et al. Pathogenic variants in USP7 cause a neurodevelopmental disorder with speech delays, altered behavior, and neurologic anomalies. Genet. Med. 21, 1797–1807 (2019).
Ragge, N. K. et al. SOX2 anophthalmia syndrome. Am. J. Med. Genet. A. 135, 1–7, https://doi.org/10.1002/ajmg.a.30642 (2005). discussion 8.
Baetens, D. et al. Biallelic and monoallelic ESR2 variants associated with 46,XY disorders of sex development. Genet. Med. 20, 717–727, https://doi.org/10.1038/gim.2017.163 (2018).
Balci, T. B. et al. Debunking Occam’s razor: diagnosing multiple genetic diseases in families by whole-exome sequencing. Clin. Genet. 92, 281–289, https://doi.org/10.1111/cge.12987 (2017).
Posey, J. E. et al. Resolution of disease phenotypes resulting from multilocus genomic variation. N. Engl. J. Med. 376, 21–31, https://doi.org/10.1056/NEJMoa1516767 (2017).
de Geus, C. M. et al. Guidelines in CHARGE syndrome and the missing link: cranial imaging. Am. J. Med. Genet. C Semin. Med. Genet. 175, 450–464, https://doi.org/10.1002/ajmg.c.31593 (2017).
Andreou, A. M. et al. TBX22 missense mutations found in patients with X-linked cleft palate affect DNA binding, sumoylation, and transcriptional repression. Am. J. Hum. Genet. 81, 700–712, https://doi.org/10.1086/521033 (2007).
Al-Baradie, R. et al. Duane radial ray syndrome (Okihiro syndrome) maps to 20q13 and results from mutations in SALL4, a new member of the SAL family. Am. J. Hum. Genet. 71, 1195–1199, https://doi.org/10.1086/343821 (2002).
Obayashi, T., Kagaya, Y., Aoki, Y., Tadaka, S. & Kinoshita, K. COXPRESdb v7: a gene coexpression database for 11 animal species supported by 23 coexpression platforms for technical evaluation and evolutionary inference. Nucleic Acids Res. 47, D55–D62, https://doi.org/10.1093/nar/gky1155 (2019).
Jiao, X. et al. DAVID-WS: a stateful web service to facilitate gene/protein list analysis. Bioinformatics. 28, 1805–1806, https://doi.org/10.1093/bioinformatics/bts251 (2012).
Choudhary, C. et al. Lysine acetylation targets protein complexes and co-regulates major cellular functions. Science. 325, 834–840, https://doi.org/10.1126/science.1175371 (2009).
Acknowledgements
We sincerely thank the patients and their families for their participation in this study. The authors thank the Genome Aggregation Database (gnomAD, https://gnomad.broadinstitute.org/about), DECIPHER (http://decipher.sanger.ac.uk), GTEx (https://www.gtexportal.org/home/index.html), BrainSpan atlas (https://www.brainspan.org/), GeneHancer (http://www.genecards.org/), and BioGRID (https://thebiogrid.org/), which provided valuable open-source genomic, expression, and proteomic data. F.C. is supported by the Schulich Research Opportunities Program and Department of Paediatrics Summer Studentship from the Schulich School of Medicine and Dentistry, London, Ontario, Canada. Analysis of patient 13 was supported by funding appointed to Department of Medical Sciences from the Italian Ministry for Education, University and Research (Ministero dell’Istruzione, dell’Università e della Ricerca–MIUR) under the program “Dipartimenti di Eccellenza 2018–2022”; Project code D15D18000410001. ES was performed as part of the Autism Sequencing Consortium and was supported by the National Institute of Mental Health (MH111661). Special thanks to Evelise Riberi and Giovanni Battista Ferrero for their analysis in this patient. Analysis of patient 16 was supported by grants 17-29423A and LM2018132 from the Czech Ministries of Health and Education. Analyses for patients 4 and 5 were partly supported by Initiative on Rare and Undiagnosed Diseases in Pediatrics (IRUD-P) (16ek0109166h0002, 17ek0109151s1) (TK) from the Japanese Agency for Medical Research and Development (AMED) and JSPS KAKENHI Grant Number JP18K07863 (TK) from Japan Society for the Promotion of Science. Special thanks to Kumiko Yanagi for their assistance in the analysis of these patients. K. Kawakami is supported by grants J-RDMM JP19ek0109288 and JP20ek0109484 from AMED.
Author information
Authors and Affiliations
Contributions
Conceptualization: T.B.B., W.B., V.M.S. Data curation: F.C., L.W., M.A.-R., D.J.A., A. Baxova, S.B., E.B., A. Brusco, O.C., T.F., M.G.-A., M.H., D.H., S.H., G.H., T.K., B.K., K. Kosaki, K. Kubota, J.M.L., M.A.M., P.R.M., M.T.M., S.M., G.M.M., H.O., N.O., D.R-B., P.R., Z.S., K.S., H.S., T.U., J.S.W., P.G.W., A.W., C.Z., I.M.W., S.R.L., V.M.S., T.B.B. Formal analysis: N.C., F.C. Funding acquisition: W.B., S.R.L., F.C., T.B.B, E.R., A.B., T.K., K. Kawakami. Investigation: N.C. Resources: T.B.B., I.M.W., W.B. Supervision: T.B.B. Visualization: F.C., L.W.; Writing—original draft: F.C. Writing—review & editing: M.A.-R., D.J.A., A. Brusco, M.G.-A., S.H., P.M., Z.S., W.B., T.B.B.
Corresponding authors
Ethics declarations
Ethics declaration
Written informed consent was obtained from all patients in accordance with protocols approved by the appropriate human subject ethics committees: through Baylor College of Medicine for patients 7, 8, 12, 17, 18, 19, 21, 22, 23, and 24 and through their respective academic/health sciences center for the rest of the cohort. Consents for publication of photographs were attained from parents/legal guardians of patients 1, 3, 8, 10, 11, 12, 14, 16, 18, 19, 20, and 24. Detailed clinical information was submitted by each patient’s clinical genetics team via a clinical questionnaire. The complete set of clinical information is provided in Supplementary Table 1. This study was approved by the Institutional Review Board at Baylor College of Medicine.
Competing interests
W.B. and L.W. are employees of Baylor Miraca Genetics Laboratories, BMGL. I.M.W. is an employee of GeneDx, Inc. The other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
About this article
Cite this article
Chowdhury, F., Wang, L., Al-Raqad, M. et al. Haploinsufficiency of PRR12 causes a spectrum of neurodevelopmental, eye, and multisystem abnormalities. Genet Med 23, 1234–1245 (2021). https://doi.org/10.1038/s41436-021-01129-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-021-01129-6