Abstract
Purpose
Nontruncating variants in SMARCA2, encoding a catalytic subunit of SWI/SNF chromatin remodeling complex, cause Nicolaides–Baraitser syndrome (NCBRS), a condition with intellectual disability and multiple congenital anomalies. Other disorders due to SMARCA2 are unknown.
Methods
By next-generation sequencing, we identified candidate variants in SMARCA2 in 20 individuals from 18 families with a syndromic neurodevelopmental disorder not consistent with NCBRS. To stratify variant interpretation, we functionally analyzed SMARCA2 variants in yeasts and performed transcriptomic and genome methylation analyses on blood leukocytes.
Results
Of 20 individuals, 14 showed a recognizable phenotype with recurrent features including epicanthal folds, blepharophimosis, and downturned nasal tip along with variable degree of intellectual disability (or blepharophimosis intellectual disability syndrome [BIS]). In contrast to most NCBRS variants, all SMARCA2 variants associated with BIS are localized outside the helicase domains. Yeast phenotype assays differentiated NCBRS from non-NCBRS SMARCA2 variants. Transcriptomic and DNA methylation signatures differentiated NCBRS from BIS and those with nonspecific phenotype. In the remaining six individuals with nonspecific dysmorphic features, clinical and molecular data did not permit variant reclassification.
Conclusion
We identified a novel recognizable syndrome named BIS associated with clustered de novo SMARCA2 variants outside the helicase domains, phenotypically and molecularly distinct from NCBRS.
Similar content being viewed by others
INTRODUCTION
Next-generation sequencing offers the unprecedented opportunity for unbiased screening in diagnosis of Mendelian disorders and is rapidly changing the diagnostic work-up in clinical genetics. The genotype-first approach is becoming the method of choice overcoming traditional approaches based on recognition of specific patterns of clinical anomalies.1,2 Increasing use of exome sequencing (ES) for diagnosis of intellectual disability (ID) and neurodevelopmental disorders (NDD) has identified a large number of variants of unknown significance (VUS). When VUS are the sole candidates detected and arise de novo, they raise questions about extension of the known disease spectrum or about previously unrecognized new disorders.3 Reverse phenotyping becomes essential for interpretation of these variants.4,5
Nicolaides–Baraitser syndrome (NCBRS) is a rare ID/congenital malformation syndrome,6 whose main features include ID, speech delay, seizures, microcephaly, coarse facial features, and phalangeal abnormalities. NCBRS has a recognizable pattern of anomalies including ectodermal anomalies, facial coarsening with thick nares, broad philtrum, wide mouth and thin upper and thick lower vermillion, and limb anomalies with prominent interphalangeal joints, broad distal phalanges, and short metacarpals and/or metatarsals.7 More than 60 de novo heterozygous pathogenic variants in SMARCA2 gene have been reported in NCBRS.7,8,9 Genomic deletions of 9p24.3-p23 region encompassing the whole SMARCA2 gene are also causative but reported phenotypes are not consistent with NCBRS.10,11
SMARCA2 encodes one of the two helicase-related catalytic subunits of the superfamily II helicase group of the BRG1 and BRM-associated factors (BAF) complex, the mammalian homolog of switch/sucrose nonfermentable (SWI/SNF), a chromatin remodeling complex that regulates expression of several genes.8,12 SMARCA2 canonical domains involved in DNA binding and ATP hydrolysis are highly conserved from unicellular eukaryotes (yeast) to human. ATP hydrolysis provides energy to disrupt histone–DNA interactions and promotes nucleosome sliding and repositioning, resulting in increased accessibility of DNA for transcription and replication. Variants causing NCBRS are mainly missense and rarely in-frame exon deletions.7 They all affect the ATPase domain or nearby residues, in either Helicase ATP-binding or Helicase C-terminal subdomains.7,13,14,15,16,17,18,19,20,21,22,23 Disruption of BAF complex in NCBRS was recently found to affect DNA methylation.24 In most cases, SMARCA2 variants were identified by targeted gene sequencing following clinical suspicion of NCBRS. In this study, we identified missense variants in SMARCA2 in 20 individuals through pan-genomic sequencing without an a priori clinical diagnosis of NCBRS. By reverse phenotyping and functional analyses, we delineated a new recognizable syndrome in 14 of the 20 individuals who all harbored de novo pathogenic variants located outside the helicase domains.
MATERIALS AND METHODS
Enrollment of subjects
Subjects were recruited through international collaboration with the Matchmaker exchange initiative25 and the European Reference Network (ERN) ITHACA. All subjects were referred for clinical genetic evaluation of ID and/or behavioral problems associated with dysmorphic features and congenital anomalies. After a negative diagnostic work-up including array comparative genomic hybridization (aCGH)-based chromosomal studies, targeted sequencing, exome sequencing (ES), or genome sequencing (GS) was proposed as part of the diagnostic procedure. Targeted sequencing was performed in subjects 2, 9, and 17 whereas all remaining cases were analyzed by either TruSight One panel (Illumina, Inc.), exome or genome sequencing as trios. Paternity was confirmed in all cases except subject 18 (mother of subject 17). SMARCA2 variants were all confirmed by Sanger sequencing and parents were available in all cases except subject 18. American College of Medical Genetics and Genomics (ACMG) classification was used for variant interpretation.26
Ethics statement
The study was approved by Ethic committees at Federico II University Hospital (protocol number 48/16) and University Hospital Dijon Bourgogne (Comité de protection des personnes Est I, Centre Hospitalier La Chartreuse—protocol number 2016/38). Additional subjects were identified through the Deciphering Developmental Disorders project (Cambridge South REC reference 10/H0305/83, and Republic of Ireland REC GEN/284/12), the 100,000 Genomes Project (East of England—Cambridge South REC 14/EE/1112), and the BUILD Study (London–Camden & Kings Cross REC 17/LO/0981). All the participants or their families consented to participation in the study. Consent for photograph publication was obtained for all individuals shown in Fig. 1.
(a) Pictures of subjects 1–13 at different ages are shown (pictures of subject 14 are not available). Note the striking similarities in facial appearance with blepharophimosis, epicanthal folds, highly arched and sparse eyebrows, sparse eyelashes, broad nasal bridge, and downturned nasal tip. (b) Facial features of four individuals with de novo SMARCA2 variants without a recurrent pattern of anomalies.
Yeast strains and assays
Yeast analyses were performed with strains derived from Saccharomyces cerevisiae haploid strain BY4741. Primers and double stranded DNA fragments, strains, and plasmids are listed in Supplementary Tables S1–S3. SNF2 gene (YOR290C) was edited at the genomic locus using markerless CRISPR/Cas9 genome engineering, as previously described27 and as reported in details in Supplementary Methods. Oligonucleotides, plasmids, and donor DNA used to generate each strain is summarized in Supplementary Table S4. Yeast growth assays are also described in Supplementary Methods.
RNA extraction, qPCR analysis, and RNA sequencing
RNA was extracted with PAXgene® blood RNA kit from fresh peripheral blood following manufacturer’s instructions. RNA concentration and purity were measured using NanoDrop ND1000 spectrometer. RNA integrity was evaluated by Agilent Bioanalyser. RNA with RNA Integrity Number above or equal to 7.0 were used for analysis. For RNA reverse transcription, first-strand complementary DNA (cDNA) was synthetized from 0.5 µg total RNA using iScript™ Reverse Transcription Supermix (Bio-Rad®). In a final volume of 20 µl the real-time polymerase chain reaction (PCR) mix contained 2 ng of reverse transcribed total RNA, 300 nM of forward and reverse primers, and iTaq™ SYBR® Green supermix_Bio-Rad®. Quantitative PCR (qPCR) was performed in triplicate in 96-well plates on CFX96™ thermocycler (Bio-Rad®) using HotStarTaq Plus DNA Polymerase (Qiagen®) according to manufacturer’s protocol. Sequences of primers are listed in Supplementary Table S1. SMARCA2 expression was normalized using Human Ribosomal Protein Large P0 (RPLP0). Methods for RNA-seq alignment and differential expression analysis, unsupervised clustering, and gene set enrichment analyses are included in Supplementary Methods.
DNA methylation analysis
Methods for methylation experiment and DNA methylation profiling and analysis are reported in Supplementary Methods.
RESULTS
Reverse phenotyping of individuals carrying SMARCA2 variants
By sequencing a panel of ID-related genes, ES, or GS we identified 20 subjects from 18 families carrying heterozygous missense variants in SMARCA2 with 19 being de novo (ACMG criterion PS2). All variants were absent in general population databases (ACMG criterion PM2), affected evolutionary conserved amino acids according to Genomic Evolutionary Rate Profiling (GERP), and were predicted to be damaging by multiple in silico tools (Supplementary Table S5). No additional candidate variants were detected in any of the subjects. For two amino acid changes (p.[Ile932Thr] and p.[Leu529Val]), variants of the same codon were reported in ClinVar as VUS, but clinical descriptions were unavailable. These variants were categorized as likely pathogenic based on PS2, PM2, and PP3 (computational evidence in support of deleterious effect on the gene and its product) criteria.26
We first compared phenotypes of the 20 individuals and clinicians specifically looked for NCBRS features.8 Sparse hair (6/20), coarse features (2/20), thick lower lip (6/20), or prominent interphalangeal joints (2/20) were infrequently observed (Supplementary Tables S6, S7). Although individuals presented with isolated anomalies, none of them cumulated enough findings supporting a diagnosis of NCBRS. Nonspecific clinical features were recurrent across all individuals. Hypotonia was detected in 13/20, developmental delay and ID of variable severity in most, postnatal onset microcephaly in 8/20, and behavioral problems in 13/20 individuals. Comparison of facial features revealed that 14 subjects shared a strikingly distinctive facial appearance (subjects 1–14) characterized by blepharophimosis (14/14), epicanthal folds (14/14), sparse eyebrows and lashes (11/14), highly arched eyebrows (10/14), broad nasal bridge (10/14), and downturned nasal tip (8/14) (Fig. 1a, Table 1 and Supplementary Table S6). Frontal bossing, hypertelorism, pinched nose, hypoplastic alae nasi, tented upper lip vermillion, exaggerated Cupid’s bow, and open mouth with U-shaped upper lip vermillion were also frequently observed (Fig. 1a, Table 1 and Supplementary Table S6). Besides blepharophimosis, features distinct from NCBRS were the shape of the nose with downturned tip in BIS rather than upturned and with large base as in NCBRS, and the absence of coarse features, thick lower lip, or prominent interphalangeal joints that are all typical NCBRS features. Moreover, abnormal dentition (widely spaced teeth, enamel hypoplasia, and premature tooth loss) were frequently detected (8/12). In addition, they frequently had limb anomalies such as joint contractures, and thin or tapered fingers. These facial, tooth, and limb anomalies were rarely found in the other six individuals (subjects 15–20, Fig. 1b, Table 1 and Supplementary Table S7). These 14 individuals (subjects 1–14) sharing similar dysmorphic features frequently had postnatal microcephaly (6/14), hypotonia (9/12), and moderate to severe ID was present in all cases. Speech abilities were severely delayed with seven individuals speaking few or no words. Motor delay was frequent and severe with walking age ranging from 22 months to 8 years, but six individuals never achieved independent ambulation and used wheelchair or assistive devices for ambulation. Two individuals had limb spasticity. Abnormality of vision with mostly refractive errors, respiratory problems with recurrent infections, or gastrointestinal problems comprising gastroesophageal reflux disease and feeding difficulties were frequent (Supplementary Table S6). Because of recurrent dysmorphic features and clinical findings in these 14 subjects, we hypothesized that they might be affected with a new disorder, which we proposed to name blepharophimosis intellectual disability syndrome (BIS). Face-to-Gene (FDNA Inc., Boston, Massachusetts; https://www.face2gene.com) analysis28 of facial features of BIS and the nonspecific phenotype confirmed the differences in facial features among these two groups and NCBRS (Supplementary Fig. S1).
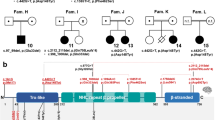
Most NCBRS individuals carry variants spanning from exons 15 to 25 with exon 25 being a hotspot.7,8 These exons encode SMARCA2 ATPase domain that is split into a helicase ATP-binding domain and a Helicase C-terminal domain11 (Fig. 2a). The 14 BIS individuals harbored variants clustered in exons 8 or 9 (p.[Arg505Gln], p.[Gly513Val], p.[Arg525His], p.[Arg525Cys], p.[Leu529Val], p.[Asp534Asn]) corresponding to the region between small helicase/SANT‐associated domain (HSA) and Helicase ATP-binding domain, or in exon 19 (p.[Glu929Val], p.[Arg937His], p.[Arg937Leu], p.[(Arg937Cys]) mapping to the linker region located between DExx Helicase ATP-binding and Helicase C-terminal domains. Variants affecting Arg525 and Arg937 residues were identified in four and five cases, respectively. Among other individuals with a nonspecific phenotype but still distinct from NCBRS, variants were spread throughout the gene in exons 8 (p.[His484Asn], p.[Asn486Lys]), 14 (p.[Leu719Pro]), 19 (p.[Ile932Thr]), and 21 (p.[Lys1014Glu]) (Fig. 2a, Supplementary Table S7).
(a) Blue dots indicate variant harbored by NCBRS individuals reported in previously published studies7,8,9 (Table 1, Supplementary Tables S6 and S7). Purple dots indicate variants of the 14 individuals with BIS (subjects 1–14, Table 1, Supplementary Table S6), yellow dots indicate variants in the nonspecific phenotype from this study (subjects 15–20, Supplementary Table S7), and white dots indicate cases reported elsewhere.7,35,39 (b) Organization of human SMARCA2 and yeast Snf2 proteins, highlighting the portion of Snf2 presented in the structural representation. (c,d) Structural representation of yeast Snf2 and its ATPase domain (orange) and Helicase C-terminal (red) bound to a nucleosome (gray) from the PDB file 5z3l. (e,f) Localization on the Snf2 structure of variants detected in NCBRS, BIS, and nonspecific phenotype. Localization of NCBRS residues (cyan) is restricted to the ATPase catalytic domain while BIS variants cluster on an ɑ-helix that defines an interaction surface with other subunits of the SWI/SNF complex. Snf2-Leu977 (human SMARCA2-Ile932) is positioned within the interface defined by BIS residues. The Snf2-Lys1057 mutant (human SMARCA2-Lys1014) is likely to disrupt its interaction with DNA. Snf2-Leu762 (human SMARCA2-Leu719) is localized at the periphery of the Snf2 structure.
Structural analysis of SMARCA2 variants
Alignment of human SMARCA2 and yeast Snf2 sequences revealed that 50 of 57 positions carrying SMARCA2 pathogenic variants are either identical or highly similar from human to yeast, respectively. Therefore, 87.7% of those residues are conserved (Supplementary Table S8). We located these residues on a molecular structure of yeast Snf2 bound to a nucleosome (PDB 5z3l)29 where we highlighted the helicase domain with its ATP-binding pocket and helicase C terminus (Fig. 2b–d). Amino acid residues affected in NCBRS and BIS are clearly located in different parts of the Snf2 helicase domain: NCBRS residues are within the core helicase domain whereas BIS residues are on an alpha-helix that is likely at the interface with other members of the SWI/SNF complex (Fig. 2e, f).30
Finally, residues affected in the nonspecific phenotype can be discriminated based on their localization (Fig. 2f). Snf2-Leu977 (SMARCA2-Ile932) is positioned within the interface defined by BIS residues, Snf2-Lys1057 (SMARCA2-Lys1014) is likely to disrupt its interaction with DNA, and Snf2-Ile762 (SMARCA2-Leu719) is localized at the periphery of Snf2. Snf2-His637 (SMARCA2-His484) and Snf2-His639 (SMARCA2-Asn486) are at the interface between the HSA domain and the actin-related protein (ARP) module.31,32
Functional studies of SMARCA2 variants in yeast
Among residues involved in the 12 SMARCA2 variants reported, 9 were conserved in S. cerevisiae Snf2 protein and 2 were similar (Supplementary Table S8). We selected SMARCA2-Asn486Lys, SMARCA2-Glu929Val and SMARCA2-Arg937Leu variants and generated the corresponding Snf2-Asn639Lys, Snf2-Glu974Val, and Snf2-Arg982Leu mutants. We also introduced the Snf2-Pro926Leu and Snf2-Pro824Arg mutations, corresponding to SMARCA2-Pro883Leu7 and SMARCA2-Pro781Arg (Supplementary Fig. S2) typical NCBRS variants, and the snf2∆ deletion lacking the complete SNF2 open reading frame (Fig. 3a).
(a) Schematic diagram of the human SMARCA2 protein and its S. cerevisiae homolog Snf2 (Top) and alignment of SMARCA2 homologs from human, Mus musculus, Drosophila melanogaster, Caenorhabditis elegans, Schizosaccharomyces pombe, and S. cerevisiae (bottom). SMARCA2 variant positions and the corresponding residues in S. cerevisiae Snf2 used in this study are indicated by dots. (b) Yeast growth assay on solid media containing various fermentable (glucose, galactose, raffinose) and nonfermentable (glycerol, acetate) carbon sources, or containing 100 mM hydroxyurea (HU). Similar results were obtained for two additional biological replicates. (c) Yeast growth assay in liquid media containing glucose or raffinose, incubated at 30 °C. Optical density (OD) at 595 nm was monitored over time and the average of four biological replicates is plotted. (d) Growth rates were calculated from the plots; error bars indicate the standard deviation. ***Indicates a p value < 0.01 determined in a two-sided Welch t-test and a difference of at least 10% compared with the wild-type (WT) sample.
When grown on solid glucose medium, only snf2∆ strain was clearly impaired. Other more stringent conditions previously shown to affect snf2 mutant strains33,34 were investigated, including other fermentable (galactose, raffinose) and nonfermentable (glycerol, acetate) carbon sources or HU-containing medium, both in optimal growth temperature (30 °C) or under heat stress condition (37 °C). Under these conditions, snf2∆ and Snf2-Pro926Leu mutants showed very severe or severe growth defect respectively, while Snf2-Glu974Val, Snf2-Arg982Leu, and Snf2-Asn639Lys were growing like the wild-type strain (Fig. 3b). Results of quantitative growth assay in liquid medium were consistent with results on solid medium with Snf2-Glu974Val, Snf2-Arg982Leu, and Snf2-Asn639Lys showing wild-type phenotype, and Snf2-Pro926Leu and snf2∆ displaying growth defects (Fig. 3c, d). Overall, S. cerevisiae strains modeling typical NCBRS variants behaved macroscopically differently from strains modeling BIS or the nonspecific phenotype. However, this approach did not provide insights into the molecular mechanisms differentiating NCBRS from BIS or the nonspecific phenotype.
Transcriptomic signature differentiating SMARCA2 variants
To investigate the molecular mechanisms involved in phenotypic differences among healthy individuals and individuals with NCBRS, BIS, or the nonspecific phenotype, we performed total RNA-seq from blood-derived RNA. RNA-seq was performed on individuals with BIS (subjects 1, 9), nonspecific phenotype (subjects 15, 17(IC), and 18(M)), two NCBRS patients carrying pathogenic SMARCA2 variants (included as positive controls), and three healthy individuals (negative controls). First, we performed several unbiased RNA-seq clustering analyses to identify subgroups of signatures that would help classifying individuals according to their transcriptional profiles. Principal component analysis of RNA-seq data revealed that the first principal component (PC1), which represents 60% of total variation in the data, separated two groups of individuals. These two groups corresponded to (1) NCBRS and BIS individuals and (2) nonspecific phenotype and negative control samples. The second principal component (PC2), accounting for 13% of total variation, barely distinguished NCBRS from BIS individuals (Supplementary Fig. S3). These results were also corroborated by consensus clustering analysis approach (Fig. 4a). At cluster counts (k) ranging from 2 to 8, NCBRS and BIS samples could not be separated from each other without overfitting the model, indicating that these two subgroups show the closest expression profile compared with the remaining samples (Supplementary Fig. S4). Next, we performed differential expression analysis searching for a common molecular signature that would distinguish individuals carrying causative variants for NCBRS and BIS from the other samples. Interestingly, we observed that SMARCA2 expression was significantly downregulated compared with controls (log2FC = −0.74, adjusted P value = 0.0025) (Supplementary Table S9). We independently validated this finding by SMARCA2 real-time qPCR in the whole cohort of available samples, i.e., 20 samples including the previously described samples and 10 more samples comprising 7 additional controls, 2 additional BIS cases (variants p.Leu529Val, and p.Arg937His), and 1 additional nonspecific phenotype case (p.His484Asn). Again, SMARCA2 expression was downregulated in NCBRS and BIS samples compared with controls and nonspecific subjects carrying p.Ile932Thr and p.Lys1014Glu variants (Fig. 4b). Hierarchical clustering analysis of the top 5000 differentially expressed genes in the whole RNA-seq data set clustered samples into two main groups: NCBRS and BIS individuals into one cluster, and individuals with nonspecific phenotype and controls in the other cluster. This analysis also showed that NCBRS and BIS individuals could be subdivided into two subgroups, suggesting that each has a distinctive expression profile (Fig. 4c). Further analysis of differentially expressed genes compared with control samples (|log2FC| ≥ 0.58 and adjusted P value ≤ 0.01) (Fig. 4d, Supplementary Fig. S5) resulted in 3051 (NCBRS versus control samples) and 1832 (BIS versus control samples) differentially expressed genes (adjusted p value cutoff ≤ 0.01 and log fold changes ≥ 0.58). Among them, 1375 were shared between NCBRS and BIS, whereas 1676 and 457, respectively, were specific to each group (Fig. 4e and Supplementary Table S9). This observation indicates that differentially expressed genes can distinguish NCBRS and BIS. Interestingly, CAMERA analysis on Gene Ontologies (GO) associated with differentially expressed genes indicated that terms linked to transcriptional regulation and protein translation were significantly enriched in both classes of affected individuals (Supplementary Table S10), consistent with the role of SMARCA2 in regulating gene expression.
(a) Representative clustering matrix on transcriptomic analysis of blood RNA. Nicolaides–Baraitser syndrome (NCBRS) and blepharophimosis intellectual disability syndrome (BIS) samples differentiate from control and nonspecific samples. (b) Blood SMARCA2 expression analysis measured by Reverse Transcription and quantitative PCR (RT-PCR). Each class was composed by n = 10 (control), 2 (NCBRS), 4 (BIS), and 4 (nonspecific) samples, respectively. (c) Heatmap representation of the expression level of the top 5000 most variable genes. (d) Global distribution of NCBRS (left panel) and BIS (right panel) differentially expressed (DE) genes compared with control samples (|log2FC | ≥ 0.58 and adjusted P value ≤ 0.01). Colored dots represent genes which are specifically deregulated in NCBRS (blue) and BIS (purple) samples. Uncoloured dots represent common differentially expressed genes in NCBRS and BIS (black area of the Venn diagram in (e)). (e) Venn diagram of differentially expressed genes. The overlap between NCBRS and BIS samples is shown in black. (f, g) Hierarchical clustering with heatmap using CpG probes differentially methylated in NCBRS and BIS. Rows indicate probes and columns indicate samples. Methylation levels are indicated using the color scale from blue to red, corresponding to a range of methylation levels from 0 to 1. The red top panes indicate the sample type. (f) Probes differentially methylated between NCBRS and controls generate a distinct pattern in BIS cases that was different from both groups. (g) Probes differentially methylated in BIS indicate a distinct methylation pattern between cases and controls and indicate a differential methylation profile between NCBRS and BIS.
Methylation signature differentiating SMARCA2 variants
Individuals with NCBRS were previously found to have aberrant genome methylation.24,35 Therefore, we performed genome-wide DNA methylation analysis in BIS individuals (subjects 7, 11, 12, and 13) using Illumina Infinium EPIC methylation arrays. We performed clustering analysis using probes for CpG sites as previously reported24 and compared NCBRS, BIS, and control samples. These probes completely separated NCBRS and BIS samples from controls. However, BIS cases also generated their own subcluster, distinct from NCBRS. This indicated that BIS genomes have a DNA methylation profile distinct form NCBRS (Fig. 4f). To identify the BIS DNA methylation episignature, we compared DNA methylation profiles of BIS cohort with 40 age- and gender-matched healthy controls (as described previously24) that identified a total of 163 CpG sites differentially methylated between the two groups (methylation difference >10%, multiple testing corrected P value < 0.01, Supplementary Table S11). When using this probe set for clustering analysis, BIS clearly separated from controls and NCBRS (Fig. 4g). These findings show that BIS and NCBRS have distinct DNA methylation signatures, supporting the clinical findings and indicating that these are separate conditions possibly due to different underlying disease mechanisms.
DISCUSSION
We identified 20 individuals without features of NCBRS but harboring SMARCA2 heterozygous missense changes that had been previously classified as either VUS or likely pathogenic according to ACMG criteria. By reverse phenotyping, we could clinically identify 14/20 subjects with a highly overlapping pattern of dysmorphic features suggesting the identification of a new recognizable syndrome that we named BIS. Functional yeast studies suggested a different molecular mechanism underlying NCBRS, BIS, and a nonspecific phenotype due to SMARCA2 variants. RNA-seq and genome methylation confirmed the clinical and molecular stratification of the three groups of individuals with SMARCA2 variants. Altogether, this study delineated at least a novel syndrome named as BIS and provides a more precise classification of SMARCA2 missense variants. However, further studies are needed to investigate the pathogenicity of the variants detected in individuals with the nonspecific phenotype (non-BIS and non-NCBRS). It is possible that SMARCA2 variants might result in a spectrum of phenotypes similar to ARID1B gene, also part of the SWI/SNF chromatin remodeling complex, that has been associated with multiple conditions raging from nonsyndromic ID to Coffin–Siris syndrome.36 If this is confirmed, additional diagnostic approaches will be required to correctly interpret the clinical significance of rare variants in SMARCA2, particularly in individuals presenting with nonspecific ID.
The increasing use of pan-genomic sequencing for the diagnosis of NDD is leading to the identification of a growing number of disease-causing genes. The de novo occurrence of a variant is a strong argument of pathogenicity.26 Nevertheless, despite being de novo, several missense variants remain of unknown significance.37 The descriptions of recurrent phenotypic similarities among individuals carrying de novo variants in the same gene is of utmost importance to support the existence of a new syndrome.5,38 Here, we describe 14 individuals with recurrent facial dysmorphisms consistent with a novel and recognizable phenotype. BIS individuals present with blepharophimosis, epicanthal folds, sparse eyebrows and eyelashes, broad nasal bridge, NDD that is most frequently severe, recurrent respiratory infections, and gastroesophageal reflux disease. Typical features of NCBRS especially facial dysmorphisms, ectodermal anomalies, and limb anomalies have been carefully searched for in individuals with BIS and the nonspecific phenotype but they were minimally detected in all 20 individuals and even more rarely in the 14 BIS individuals. Therefore, suspicion of mild NCBRS in these individuals was also ruled out. The lack of ptosis, epicanthus inversus, and limb abnormalities distinguishes BIS from other recognizable conditions with blepharophimosis, such as blepharophimosis–ptosis–epicanthus inversus.
To investigate genotype–phenotype correlation, we scrutinized variant clustering in BIS. Interestingly, BIS variants were found to affect the region between HSA and ATPase domain, or to occur inside the P loop containing nucleoside triphosphate hydrolase domain, located between helicase ATP-binding and helicase C-terminal domains. Although few NCBRS variants in the P loop7 and two variants close to the ATPase domain13,15 have been reported, variants outside the ATPase domain have not been described in NCBRS. In contrast, most individuals with BIS harbored SMARCA2 variants outside the ATPase. A case carrying a p.Trp1283Cys variant, located between the ATPase domain and the Bromodomain, was previously reported as a mild NCBRS but convincing facial dysmorphism was lacking, and only partial NCBRS DNA methylation signature was detected.35 In the same study, five individuals lacking the NCBRS phenotype but harboring benign or VUS SMARCA2 variants proximal to the ATPase domain were described.35 Unfortunately, detailed phenotypic information was not provided, and thus comparison with BIS individuals is not possible. Conversely, a subject carrying a p.Gly1420Arg variant in the bromodomain of SMARCA2, had facial features that appear to at least partially overlap with BIS.7 Finally, an individual without NCBRS features was also previously reported to carry a SMARCA2 variant in exon 9 [p.(Asp534Tyr)].39 Although pictures of this latter individual were not provided, features were reportedly suggestive of Cornelia de Lange or Hallerman–Streiff syndrome.
SMARCA2 directly binds ADNP (MIM 611386), the gene responsible for Helsmoortel–van der Aa syndrome.40 Interestingly, four individuals bearing variants within the bipartite nuclear localization signal domain of ADNP did not show the typical phenotype of Helsmoortel–van der Aa syndrome, but instead presented with blepharophimosis and epicanthal folds as the most striking facial features.41 This raises the hypothesis that clustered variants in ADNP and SMARCA2 cause an overlapping blepharophimosis–ID phenotype that might be the dysmorphology signature of their molecular interaction. Moreover, ADNP was found to interact with several major proteins of the SWI/SNF complex.42 It is tempting to speculate that pathogenic variants of SMARCA2 or ADNP result in partially shared phenotype because of similar loss of function at the chromatin level. RNA-seq and DNA methylation studies would be very attractive to investigate whether Helsmoortel–van der Aa syndrome with ADNP variants have a transcriptome and genome methylation signatures overlapping with BIS.
In our study, yeast experiments could not differentiate BIS variants from nonspecific individuals and controls. This result might reflect that yeast does not reach the complexity of human organisms despite high conservation of domain organization, sequence, and architecture between SMARCA2 human protein and S. cerevisiae homolog Snf2.32 As genome-wide transcriptomic analyses were not available, it is not possible to rule out that yeast mutants have differences at the transcriptional level despite the lack of overt phenotypic difference on the growth assays.
SMARCA2 encodes the core catalytic unit of SWI/SNF complex involved in regulation of gene expression.8 Because of the consistent clinical phenotype across the 14 individuals with BIS, we questioned about distinctive transcriptome signatures to stratify individuals with de novo SMARCA2 variants. Although the number of RNA samples was small, results suggest strong and reproducible consequences of BIS-causing SMARCA2 variants on transcriptome different from other SMARCA2 variants. Targeted quantitative PCR showed that BIS variants affect SMARCA2 messenger RNA (mRNA) levels, like NCBRS-causing variants. A shared pattern of expression among NCBRS and BIS on differential gene expression analysis is another evidence of pathogenicity. Altogether, qPCR and gene expression analysis enable the clustering of controls and nonspecific individuals on one side, and NCBRS plus BIS individuals on the other.
Previous studies showed that DNA methylation on peripheral blood cells identifies differentially methylated regions (DMRs) corresponding to SMARCA2 dysfunction in NCBRS versus other BAFopathies-associated genes.24 DMRs were effective to reclassify SMARCA2 VUS in atypical cases.35 Consistent with these studies, a specific epigenetic signature separated NCBRS from BIS cases, suggesting that this method can be used as an additional diagnostic assay for classification of VUS. Together with transcriptome signatures, these data support clinical separation of NCBRS from BIS.
Apart from the 14 BIS subjects, 6 remaining individuals did not show a recognizable phenotype (nonspecific phenotype). Although some common facial features could be observed and some of them were overlapping with BIS or NCBRS, their phenotypes were neither suggestive of NCBRS nor of BIS. Moreover, variants were spread throughout the gene, and RNA-seq did not distinguished these individuals from controls. Therefore, pathogenicity of SMARCA2 variants associated with the nonspecific phenotype remains uncertain. As additional patients with NDD and VUS in SMARC2 are identified and undergo in-depth phenotyping, it is possible that another, yet undescribed clinical entity could be associated with SMARCA2 variants.
In conclusion, we propose that SMARCA2 missense variants are responsible for at least two clinical entities: (1) NCBRS and (2) a recognizable condition with blepharophimosis, epicanthal folds, ID, and tooth anomalies herein described and named BIS.
Change history
07 October 2020
This Article was originally published without the accompanying Supplementary Material in the Word format. This file is now available in the HTML version of the Article; the PDF was correct from the time of publication.
References
Menke LA, study DDD, Gardeitchik T, et al. Further delineation of an entity caused by CREBBP and EP300 mutations but not resembling Rubinstein-Taybi syndrome. Am J Med Genet A. 2018;176:862–876.
Hansen AW, Murugan M, Li H, et al. A genocentric approach to discovery of Mendelian disorders. Am J Hum Genet. 2019;105:974–986.
Karaca E, Posey JE, Coban Akdemir Z, et al. Phenotypic expansion illuminates multilocus pathogenic variation. Genet Med. 2018;20:1528–1537.
Schulze TG, McMahon FJ. Defining the phenotype in human genetic studies: forward genetics and reverse phenotyping. Hum Hered. 2004;58:131–138.
Hennekam RC, Biesecker LG. Next-generation sequencing demands next-generation phenotyping. Hum Mutat. 2012;33:884–886.
Nicolaides P, Baraitser M. An unusual syndrome with mental retardation and sparse hair. Clin Dysmorphol. 1993;2:232–236.
Sousa SB, Hennekam RC, Nicolaides-Baraitser Syndrome International Consortium. Phenotype and genotype in Nicolaides–Baraitser syndrome. Am J Med Genet C Semin Med Genet. 2014;166C:302–314.
Van Houdt JK, Nowakowska BA, Sousa SB, et al. Heterozygous missense mutations in SMARCA2 cause Nicolaides–Baraitser syndrome. Nat Genet. 2012;44:445–9, S441.
Wolff D, Endele S, Azzarello-Burri S, et al. In-frame deletion and missense mutations of the C-Terminal Helicase Domain of SMARCA2 in three patients with Nicolaides–Baraitser syndrome. Mol Syndromol. 2012;2:237–244.
Borlot F, Regan BM, Bassett AS, Stavropoulos DJ, Andrade DM. Prevalence of pathogenic copy number variation in adults with pediatric-onset epilepsy and intellectual disability. JAMA Neurol. 2017;74:1301–1311.
Sekiguchi F, Tsurusaki Y, Okamoto N, et al. Genetic abnormalities in a large cohort of Coffin–Siris syndrome patients. J Hum Genet. 2019;64:1173–1186.
Santen GW, Kriek M, van Attikum H. SWI/SNF complex in disorder: switching from malignancies to intellectual disability. Epigenetics. 2012;7:1219–1224.
Bramswig NC, Ludecke HJ, Alanay Y, et al. Exome sequencing unravels unexpected differential diagnoses in individuals with the tentative diagnosis of Coffin–Siris and Nicolaides–Baraitser syndromes. Hum Genet. 2015;134:553–568.
Ejaz R, Babul-Hirji R, Chitayat D. The evolving features of Nicolaides–Baraitser syndrome—a clinical report of a 20-year follow-up. Clin Case Rep. 2016;4:351–355.
Tang S, Hughes E, Lascelles K, Euro ERESmaewg, Simpson MA, Pal DK. New SMARCA2 mutation in a patient with Nicolaides–Baraitser syndrome and myoclonic astatic epilepsy. Am J Med Genet A. 2017;173:195–199.
Gripp KW, Baker L, Telegrafi A, Monaghan KG. The role of objective facial analysis using FDNA in making diagnoses following whole exome analysis. Report of two patients with mutations in the BAF complex genes. Am J Med Genet A. 2016;170:1754–1762.
Wieczorek D, Bogershausen N, Beleggia F, et al. A comprehensive molecular study on Coffin–Siris and Nicolaides–Baraitser syndromes identifies a broad molecular and clinical spectrum converging on altered chromatin remodeling. Hum Mol Genet. 2013;22:5121–5135.
Sanchez AI, Rojas JA. A SMARCA2 mutation in the first case report of Nicolaides–Baraitser syndrome in Latin America: genotype-phenotype correlation. Case Rep Genet. 2017;2017:8639617.
Tsurusaki Y, Okamoto N, Ohashi H, et al. Mutations affecting components of the SWI/SNF complex cause Coffin–Siris syndrome. Nat Genet. 2012;44:376–378.
Santen GW, Aten E, Vulto-van Silfhout AT, et al. Coffin–Siris syndrome and the BAF complex: genotype–phenotype study in 63 patients. Hum Mutat. 2013;34:1519–1528.
Yamamoto T, Imaizumi T, Yamamoto-Shimojima K, et al. Genomic backgrounds of Japanese patients with undiagnosed neurodevelopmental disorders. Brain Dev. 2019;41:776–782.
Mari F, Marozza A, Mencarelli MA, et al. Coffin–Siris and Nicolaides–Baraitser syndromes are a common well recognizable cause of intellectual disability. Brain Dev. 2015;37:527–536.
Kosho T, Okamoto N, Ohashi H, et al. Clinical correlations of mutations affecting six components of the SWI/SNF complex: detailed description of 21 patients and a review of the literature. Am J Med Genet A. 2013;161A:1221–1237.
Aref-Eshghi E, Bend EG, Hood RL, et al. BAFopathies’ DNA methylation epi-signatures demonstrate diagnostic utility and functional continuum of Coffin–Siris and Nicolaides–Baraitser syndromes. Nat Commun. 2018;9:4885.
Sobreira N, Schiettecatte F, Valle D, Hamosh A. GeneMatcher: a matching tool for connecting investigators with an interest in the same gene. Hum Mutat. 2015;36:928–930.
Richards S, Aziz N, Bale S, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–424.
Shaw WM, Yamauchi H, Mead J, et al. Engineering a model cell for rational tuning of GPCR signaling. Cell. 2019;177:782–96. e727.
Gurovich Y, Hanani Y, Bar O, et al. Identifying facial phenotypes of genetic disorders using deep learning. Nat Med. 2019;25:60–64.
Li M, Xia X, Tian Y, et al. Mechanism of DNA translocation underlying chromatin remodelling by Snf2. Nature. 2019;567:409–413.
Mashtalir N, D’Avino AR, Michel BC, et al. Modular organization and assembly of SWI/SNF family chromatin remodeling complexes. Cell. 2018;175:1272–88. e1220.
Schubert HL, Wittmeyer J, Kasten MM, et al. Structure of an actin-related subcomplex of the SWI/SNF chromatin remodeler. Proc Natl Acad Sci U S A. 2013;110:3345–3350.
Han Y, Reyes AA, Malik S, He Y. Cryo-EM structure of SWI/SNF complex bound to a nucleosome. Nature. 2020;579:452–455.
Neigeborn L, Carlson M. Genes affecting the regulation of SUC2 gene expression by glucose repression in Saccharomyces cerevisiae. Genetics. 1984;108:845–858.
Minard LV, Lin LJ, Schultz MC. SWI/SNF and Asf1 independently promote derepression of the DNA damage response genes under conditions of replication stress. PLoS ONE. 2011;6:e21633.
Chater-Diehl E, Ejaz R, Cytrynbaum C, et al. New insights into DNA methylation signatures: SMARCA2 variants in Nicolaides–Baraitser syndrome. BMC Med Genomics. 2019;12:105.
van der Sluijs PJ, Jansen S, Vergano SA, et al. The ARID1B spectrum in 143 patients: from nonsyndromic intellectual disability to Coffin–Siris syndrome. Genet Med. 2019;21:1295–1307.
Krawitz P, Buske O, Zhu N, Brudno M, Robinson PN. The genomic birthday paradox: how much is enough? Hum Mutat. 2015;36:989–997.
Hoischen A, Krumm N, Eichler EE. Prioritization of neurodevelopmental disease genes by discovery of new mutations. Nat Neurosci. 2014;17:764–772.
Lee H, Deignan JL, Dorrani N, et al. Clinical exome sequencing for genetic identification of rare Mendelian disorders. JAMA. 2014;312:1880–1887.
Vandeweyer G, Helsmoortel C, Van Dijck A, et al. The transcriptional regulator ADNP links the BAF (SWI/SNF) complexes with autism. Am J Med Genet C Semin Med Genet. 2014;166C:315–326.
Takenouchi T, Miwa T, Sakamoto Y, et al. Further evidence that a blepharophimosis syndrome phenotype is associated with a specific class of mutation in the ADNP gene. Am J Med Genet A. 2017;173:1631–1634.
Mandel S, Gozes I. Activity-dependent neuroprotective protein constitutes a novel element in the SWI/SNF chromatin remodeling complex. J Biol Chem. 2007;282:34448–34456.
Acknowledgements
We thank parents of our patients and particularly Anna Caccavo and Charlotte Bull. We thank the Centre de Calcul de l’Université de Bourgogne (CCUB, https://haydn2005.u-bourgogne.fr/dsi-ccub/) for technical support and management of information technology platform. This work was supported by Telethon Foundation, Telethon Undiagnosed Diseases Program (TUDP, GSP15001), FEDER, and the Czech Ministry of Health (17-29423A and 00064203 to MH and MH). This work was in part generated within the European Reference Network ITHACA. C.S. is supported by French Medical Foundation (FRM), S.E.K. by French Medical Foundation and French National Research Agency (ANR-10-LABX-49-01 GRAL), J.G. by Finovi and French National Research Agency (ANR-18-CE18-007), and C.D. by Wellcome Trust (209568/Z/17/Z). Patients identified through the Deciphering Developmental Disorders (DDD) Study are reported. The DDD Study presents independent research commissioned by the Health Innovation Challenge Fund (grant number HICF-1009-003), a parallel funding partnership between Wellcome Trust and Department of Health, and the Wellcome Trust Sanger Institute (grant number WT098051). Views expressed in this publication are those of the author(s) and not necessarily of Wellcome Trust or the Department of Health. The study has UK Research Ethics Committee approval (10/H0305/83, granted by the Cambridge South REC, and GEN/284/12 granted by the Republic of Ireland REC). Part of the data presented here was provided through access to data and findings generated by the 100,000 Genomes Project, funded by National Institute for Health Research (NIHR) and NHS England (full acknowledgement on https://www.genomicsengland.co.uk/about-gecip/publications/).
Author information
Authors and Affiliations
Consortia
Corresponding authors
Ethics declarations
Disclosure
The authors declare no conflicts of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Cappuccio, G., Sayou, C., Tanno, P.L. et al. De novo SMARCA2 variants clustered outside the helicase domain cause a new recognizable syndrome with intellectual disability and blepharophimosis distinct from Nicolaides–Baraitser syndrome. Genet Med 22, 1838–1850 (2020). https://doi.org/10.1038/s41436-020-0898-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-020-0898-y
Keywords
This article is cited by
-
Use of dual genomic sequencing to screen mitochondrial diseases in pediatrics: a retrospective analysis
Scientific Reports (2023)
-
Landscape of mSWI/SNF chromatin remodeling complex perturbations in neurodevelopmental disorders
Nature Genetics (2023)
-
Clinical epigenomics: genome-wide DNA methylation analysis for the diagnosis of Mendelian disorders
Genetics in Medicine (2021)