Abstract
Purpose
Current interpretation guidelines for germline variants in high-risk cancer susceptibility genes consider predicted loss-of-function (LoF) variants, such as nonsense variants and variants in the canonical splice site sequences ofBRCA2, to be associated with high cancer risk. However, some variant alleles produce alternative transcripts that encode (partially) functional protein isoforms leading to possible incorrect risk estimations. For accurate classification of variants it is therefore essential that alternative transcripts are identified and functionally characterized.
Methods
We systematically evaluated a large panel of human BRCA2 variants for the production of alternative transcripts and assessed their capacity to exert BRCA2 protein functionality. Evaluated variants included all single-exon deletions, various multiple-exon deletions, intronic variants at the canonical splice donor and acceptor sequences, and variants that previously have been shown to affect messenger RNA (mRNA) splicing in carriers.
Results
Multiple alternative transcripts encoding (partially) functional protein isoforms were identified (e.g., ∆[E4–E7], ∆[E6–E7], ∆E[6q39_E8], ∆[E10], ∆[E12], ∆E[12–14]). Expression of these transcripts did attenuate the impact of predicted LoF variants such as the canonical splice site variants c.631+2T>G, c.517-2A>G, c.6842-2A>G, c.6937+1G>A, and nonsense variants c.491T>A, c.581G>A, and c.6901G>T.
Conclusion
These results allow refinement of variant interpretation guidelines for BRCA2 by providing insight into the functional consequences of naturally occurring and variant-related alternative splicing events.
Similar content being viewed by others
INTRODUCTION
Genetic testing of individuals with an enhanced risk of developing breast or ovarian cancer is routine clinical practice. Predicted loss-of-function (LoF) variants in BRCA1 and BRCA2, such as nonsense variants, frame-shifting indels, and variants at the canonical splice sites, are considered to be associated with high cancer risk and carriers and their family members are managed accordingly.
Recently, however, it was established that some naturally occurring alternative transcripts of BRCA1 and BRCA2 encode protein isoforms with residual tumor suppressive activity.1,2,3,4,5,6,7 As a consequence, the pathogenic potential of predicted LoF variants located in an exon absent in these alternative transcripts may be substantially smaller than assumed.
Current gene-specific variant classification guidelines by ENIGMA (https://enigmaconsortium.org/) as well as the generic guidelines published by the American College of Medical Genetics and Genomics (ACMG) and Association for Molecular Pathology (AMP)8 have therefore included a cautionary note. ENIGMA classification rules (https://enigmaconsortium.org/) state that variants found to produce messenger RNA (mRNA) transcript(s) predicted to encode isoforms that do not disrupt known clinically important functional domains should be considered class 3. The ACMG/AMP guidelines pose that the Pathogenic Very Strong (PVS1) code for predicted loss-of-function variants (nonsense, frameshift, canonical ±1 or 2 splice sites, initiation codon, single or multiexon deletion) may no longer be valid if a variant induces an in-frame deletion or insertion that leaves the functional domains of the protein intact.9 Furthermore, caution is warranted for a variant allele that produces multiple mRNA transcripts as both transcript ratios and the functional integrity of the isoforms can affect its clinical relevance. Although alternative transcripts have been described for both BRCA1 and BRCA2,10,11 a systematic analysis of the functionality of encoded protein isoforms has not been performed, which complicates the application of these variant classification guidelines.
For many BRCA1 and BRCA2 variants (both intronic and exonic) an effect on mRNA splicing has been reported using either patient RNA or minigene analysis.12,13,14,15,16,17,18,19,20,21 The analysis of patient RNA is however often hampered by the inability to determine allele-specific transcript expression. It then remains unclear if and to what extent wild type (WT) mRNA is still produced from the variant allele. To more directly assess the impact of an individual variant on both the nature and level of aberrant transcripts, minigene assays have been developed. These assays however lack the genomic context of the complete gene, limiting the detection of potential alternative transcripts. Jointly, the currently available approaches may provide evidence toward pathogenicity, but they all suffer from the same limitation: they do not provide insight into the in vivo functional consequences of variants that affect splicing, an important component of assessing variant pathogenicity. This shortcoming underscores the need for more detailed analyses per gene in which the presence and expression levels of alternative transcripts, either naturally occurring or induced by a variant, can be linked to protein function.
We recently validated a mouse embryonic stem cell (mESC)–based assay as a sensitive test for functional characterization of BRCA2 missense variants.22 As sequence alterations are introduced in the full-length (FL) human BRCA2 gene, the functional impact of all types of variants can be assessed including those that affect mRNA splicing. In addition, the presence of only a single human BRCA2 allele makes the mESC system eminently suited for alternative mRNA transcript analysis.
In the present study, we show that the nature and relative contribution of naturally occurring transcripts to the overall expression of human BRCA2 expressed in mESC is highly similar to those detected in various human tissues and cell lines. Furthermore, we systematically characterized a large panel of alternative transcripts for their ability to encode for (partially) functional BRCA2 protein.
The functional data presented here can be used to refine classification guidelines for variants in BRCA2 and improve the validity of PVS1 assignments for this gene. Moreover, alternative splicing is a general feature of many multiexon coding genes, and should be considered as a mechanism by which the assumed pathogenic potential of predicted LoF variants may be attenuated or even circumvented.
MATERIALS AND METHODS
Generation of exon-deletion variants
Thirty different exon-deletion variants (i.e., 25 single-exon deletions as well as five multiple-exon deletions) were generated in the full-length human BRCA2 gene located on a bacterial artificial chromosome (BAC) (clone RP11-777I19, BACPAC) as described previously23 (Tables S1, S5). Once the deletion was confirmed by Sanger sequencing BAC DNA was isolated according to manufacturer protocol (NucleoBond® Xtra Midi, Macherey-Nagel).
Selection and generation of BRCA2 variants
Single-nucleotide variants that are likely to affect BRCA2 mRNA splicing were selected from the ClinVar database (https://www.ncbi.nlm.nih.gov/clinvar) consisting of variants in the canonical ±1 or 2 splice sites (Table S2) or of the last nucleotide of an exon (Table S3). In addition, we included variants for which aberrant splicing patterns had been reported in the literature to assess whether BRCA2 variants expressed in mESC yield similar patterns of alternative transcripts as human cells (Table S3). Furthermore, from the ClinVar database, we selected nonsense variants located within exons that are absent from naturally occurring alternative transcripts (Ex3–7, Ex12, Ex18, and Ex19) or other alternative in-frame transcripts comprising a single-exon deletion (Ex10 and Ex26) (Table S2). Variants were generated in the complete human BRCA2 gene as described previously.22 Primer sequences are listed in Table S5.
mESC-based functional assay
The mESC-based functional assay involves the introduction of humanBRCA2 variants into a hemizygousBrca2 mESC line as described previously (Fig. S1).22 For the cell viability assay, 6 × 104 cells were seeded in triplo on 60-mm cell culture dishes and subsequently treated for 16 hours with 1.0 µM 4-Hydroxytamoxifen (4-OHT) (Sigma Aldrich). The next day, cells were washed with phosphate-buffered saline (PBS) and cultured for six days in the presence of hypoxanthine–aminopterin–thymidine (HAT) and subsequently five days in the presence of hypoxanthine–thymidine (HT). Thirteen days after 4-OHT treatment, one culture dish was used to visualize clonal survival by methylene blue staining. For each variant, the number of clones was compared with WT BRCA2 expressing cells and based on that categorized into one of three categories: full (similar numbers of clones as WTBRCA2), intermediate (fewer and smaller clones than WT BRCA2), and noncomplementing (absence of viable clones) variants (Fig. S2). Variants of the full and intermediate complementing categories were assessed in the homology directed repair (HDR) assay as described previously.22 A flowchart for the interpretation of functional data generated by the mESC assay is presented in Fig. S4.
Reverse Transcription-PCR (RT-PCR)
To study the effect of a variant on mRNA splicing, RNA was isolated using a trizol-based protocol and complementary DNA (cDNA) was synthesized using the ProtoScript II First Strand cDNA synthesis kit (NEB) according to manufacturer’s instructions. For variants that failed functional complementation in the cell viability assay, RNA was isolated prior to removal of the conditional mouse Brca2 (mBrca2) allele. Then, 2 µl of cDNA was amplified with GoTaq polymerase (Promega) and human BRCA2 exon-specific primer pairs (Table S5) under the following polymerase chain reaction (PCR) conditions; 95 °C for 5 minutes, followed by 28 cycles of 95 °C for 30 seconds, 55 °C for 30 seconds, 72 °C for 2 minutes, and a final step at 72 °C for 10 minutes. RT-PCR products were separated on 0.8–1.5% agarose gels stained with ethidium bromide and visualized by exposure to ultraviolet (UV) light. Individual bands were reamplified by band-stab PCR24 and purified PCR products were subjected to Sanger sequencing to identify which transcript they represented. Importantly, not every band on the gel reflected a unique mRNA transcript as some of the products represented single-stranded PCR products.
Quantitative analysis of naturally occurring alternative transcripts in mESC expressing WT BRCA2
Capillary electrophoresis (CE) analysis of alternative splicing has been extensively described previously.11 In brief, we used a panel of overlapping RT-PCR assays (combinations of forward and fluorescent-labeled reverse primers located in different exons) that allowed a comprehensive screening of BRCA2 splicing events by CE.
Analysis was performed on two technical replicas of RNA samples from mESC expressing WT BRCA2. RNA samples (approximately 1 μg) were subjected to cDNA synthesis using a PrimeScript RT reagent kit with random primers according to the manufacturer’s protocol (Takara Biotechnology). We performed 13 different RT-PCR assays spanning exons 1–4, 1–6, 3–8, 4–9, 7–10, 11–14, 11–15, 14–16, 16–19, 16–22, 19–22, 20–24, and 22–27 (sequences of all primers are available upon request). PCR products were analyzed by CE (50-cm capillary arrays) in a 3130 Genetic Analyzer (Applied Biosystems) with GeneScan 500-LIZ/1200-LIZ size standards (Applied Biosystems) as internal markers. Size calling was performed with GeneMapper v4.0 Software (Applied Biosystems). For comparison, RNA samples from lymphoblastoid cell lines (LCLs) were analyzed in parallel. By comparing the relative contribution of the same alternative transcripts between samples from mESC and human cells, the quantification is not influenced by overestimation of the expression of shorter transcripts as previously shown to occur using RT-PCR in combination with CE.2,25,26 Only fragments over 50 relative fluorescent units (RFUs) were considered to represent distinct transcripts.
Western blot analysis
Western blot analysis was performed using NuPAGE™ Novex™ 3–8% Tris-Acetate Protein Gels (ThermoFisher Scientific). BRCA2 protein was detected with the rabbit polyclonal antibody (BETHYL, A303–434A-T) directed against a region between amino acids 450–500 in exon 10 of BRCA2. Protein signal was detected by electrochemiluminescence (Amersham ECL RPN2235 Biocompare). It is important to note that most in-frame protein isoforms cannot be distinguished by western blot analysis due to the small difference in size between the full-length BRCA2 protein (BRCA2 FL protein isoform, 3418 aa) and BRCA2 protein isoforms deleted for only one or a few small exons.
RESULTS
Naturally occurring alternative splicing of BRCA2 mRNA
The mESC-based functional assay allows evaluation of any type ofBRCA2 variant in its natural genomic context. Variants are introduced in a human BRCA2-containing BAC, transfected into mESC containing a single, conditional mBrca2 allele and assessed for their ability to rescue the cell lethality provoked by removal of endogenousmBrca2 (Fig. S1). Three phenotypes can be distinguished for variants, i.e., fully complementing (similar number of clones as WT BRCA2), intermediate (fewer and smaller clones than WT BRCA2), and noncomplementing (absence of viable clones) (Fig. S2). Subsequently, variants of the full and intermediate complementing categories can be tested for their ability to perform HDR, the most prominent tumor suppressor function of BRCA2. The assay was previously validated by functional assessment of a large series of classified BRCA2 missense variants and revealed a high sensitivity and specificity for variant classification.22 It is important to note that the complementation phenotype reflects the impact of variants on HDR as well as other BRCA2-associated cellular processes that play a role in the preservation of genome stability. Consequently, the correlation between complementation phenotype and HDR capacity is not absolute, but in general intermediate complementing variants display a severe reduction in repair capacity.
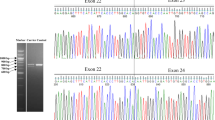
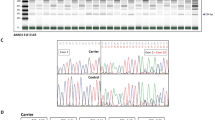
To determine whether processing of human BRCA2 mRNA by the murine spliceosome accurately reflects the splicing process in human cells, we determined the presence and quantity of the major naturally occurring isoforms that are produced from a genomic copy of the human BRCA2 gene in mESC. RNA analysis showed that the repertoire of the major naturally occurring mRNA transcripts (i.e., ∆[E3], ∆[E6q39_E7], ∆[E12], and ∆[E17–E18]) of mESC expressing WT BRCA2 closely resembled that of human LCLs both qualitatively (all predominant splicing events are detected, novel splicing events are not observed) and quantitatively (similar expression ratios relative to FL transcript) (Fig. 1).11 Up to this date, no tissue-specific transcripts have been observed in nonmalignant breast epithelia, ovarian epithelia, or ovarian fimbria.11,15
Functional characterization of exon deletions in BRCA2
Although splicing is a highly coordinated process, it is currently impossible to predict which alternative transcripts will be produced when the splice recognition site of a particular exon is destroyed or when a complete exon has been deleted. Furthermore, it is unclear when in-frame transcripts are produced whether these encode for protein isoforms that retain tumor suppressor activity. To systematically bridge this knowledge gap, we generated 30 different exon-deletion (DelEx) variants in the human BRCA2 gene, including all single-exon deletions and five in-frame multiple-exon deletions (Table S1) and analyzed the alternative transcripts these DelEx variants produced as well as their ability to preserve BRCA2 functionality.
After removal of the conditional mBrca2 allele, 17 DelEx variants failed to complement the cell lethal phenotype induced by loss of mBrca2. Eight DelEx variants displayed full complementation, while complementation was intermediate for five other DelEx variants (Table S1, Fig. S2).
RNA analysis revealed that various DelEx variants expressed multiple alternative mRNA transcripts (e.g., DelEx4 variant did not only produce ∆[E4] but also ∆[E4–E7] transcript) (Table S1). Evaluation of the transcripts generated by the DelEx variants displaying full complementation allowed us to identify several potential rescue transcripts, i.e., encoding (partially) functional BRCA2 protein isoforms (Table S1) that could be detected by western blot analysis (Fig. S3). The most potent in-frame rescue transcripts being ∆(E4–E7) (r.317_631del315), ∆(E6–E7) (r.476_631del156), ∆(E6q39_E8) (r.478_681del204), ∆(E10) (r.794_1909del1116), ∆(E12) (r.6842_6937del96) and ∆(E12–E14) (r.6842_7435del594) (summarized in Figs. 2b and 3, Table S1). BRCA2 transcripts expressed by variants displaying intermediate complementation encoded protein isoforms that were either truncated and/or reduced in quantity (DelEx5, DelEx14, DelEx15, DelEx16, DelEx18). In some cases (DelEx15, DelEx16, DelEx18) the nature of the rescue transcript remains elusive. DelEx variants that failed to complement cell lethality either produced no detectable transcript (DelEx2) or (a mixture of) out-of-frame transcripts (DelEx6, 9, 13,, 20, 21, 22, 23, 24, 25) and nonfunctional in-frame transcripts (DelEx3, 3–7, 11, 14–16, 17, 19, 26) (summarized in Figs. 2b and 3, Table S1). Congruently with their complementation phenotype, the fully complementing DelEx variants displayed HDR levels above 50%. In contrast, HDR activity of the five variants that showed intermediate complementation was severely diminished to a level previously defined for variants associated with enhanced breast cancer risk (HDR < 30%) (Fig. 4a).22
(a) BRCA2 reading frame and (b) the functionality conferred by alternative in-frame BRCA2 isoforms including single- and multiple-exon deletions. Figure adapted from Mesman et al.22 Green box = homology directed repair (HDR) capacity >50%. Red box = no complementation or HDR capacity ≤ 30%.
Splicing patterns of BRCA2 variants in or surrounding (a) Ex2–7, (b) Ex8–13, (c) Ex14–17, (d) Ex18–26. The location of exonic primers used for RT-PCR is indicated by arrows above the schematic representation of the analyzed cDNA region. Unique mRNA transcripts indicated in this figure by “other” are defined in Tables S1–S3. Asterisks denote nonsense variants. + transcript detected. FL full length, M marker (Smartladder 200 to 10000 bp from Eurogentec) WT wild type.
Homology directed repair (HDR) capacity measured in the Direct Repeat - Green Fluorescent Protein (DR-GFP) reporter assay for (a) DelEx variants, (b) PVS1 variants, and (c) potential spliceogenicBRCA2 variants outside the canonical splice sites. HDR capacity is expressed as the percentage GFP positive cells relative to the GFP positive population in wild type (WT) BRCA2 samples. Error bars indicate the SD of six independent GFP measurements per variant. The HDR capacity was measured for all variants that were able to complement the loss of cell viability following Cre-mediated deletion of the conditional mBrca2 allele. The HDR range of classified nonpathogenic (class 1/2) and pathogenic (class 4/5) BRCA2 missense variants is plotted at the right y-axis.22 Asterisks denote nonsense variants.
Based on the functionality of the transcripts lacking specific exons as summarized in Fig. 2b, it is concluded that exons 4, 5, 6, 7, 8, 10, 12, 13, and 14 do not encode essential parts of BRCA2 protein and that protein isoforms encoded by naturally occurring or variant-induced in-frame alternative transcripts lacking one or multiple of these exons may (partially) retain BRCA2’s functionality in HDR.
Functional characterization of BRCA2 PVS1 variants
Variant classification using ACMG/AMP guidelines involves several benign and pathogenic evidence criteria, including a pathogenic criterion (PVS1) for predicted LoF variants (nonsense, frameshift, canonical ±1 or 2 splice sites, initiation codon, single- or multiexon deletion and duplications).9 The results from our DelEx variant analyses suggest that nonsense, out-of-frame indels, and spliceogenic variants either located in or affecting splicing of exons that encode nonessential domains of the BRCA2 protein may not lead to complete LoF because of the production of rescue transcripts. To investigate this in more detail, we characterized a panel of 29 nonsense and spliceogenic PVS1 variants for their ability to produce BRCA2 isoforms that retain residual protein activity (Table S2).
Of the ten nonsense variants that were evaluated, one variant (c.6901G>T [located in Ex12]) displayed full complementation while three nonsense variants (c.491T>A [Ex6], c.581G>A [Ex7], and c.9572G>A [Ex26]) showed intermediate complementation. Variant c.6901G>T almost exclusively produced the ∆(E12) transcript (Fig. 3b). In line with the previously demonstrated functionality of variant DelEx12 (51% HDR), variant c.6901G>T revealed a moderate functional impact and retained 43% HDR capacity (Fig. 4b). In cells expressing either variant c.491T>A or c.581G>A, the expression level of the naturally occurring ∆(E4–E7) transcript was slightly enhanced compared with cells expressing WT BRCA2 (Fig. 3a). As shown for the DelEx4–7 variant this alternative transcript encodes a HDR-competent protein isoform. Nevertheless, the expression level of the ∆(E4–E7) transcript is apparently insufficient to retain full BRCA2 functionality as both nonsense variants display a severe impact on HDR (Fig. 4b). Variant c.9572G>A produced two transcripts: the FL transcript containing the stop codon and a ∆Ex26 transcript. As the DelEx26 variant failed to complement loss of endogenous BRCA2, the observed complementation of c.9572G>A is unlikely the consequence of increased Ex26 skipping. The severe reduction in HDR activity detected for c.9572G>A, only 25% activity compared with WT, most likely reflects some residual activity conferred by the truncated BRCA2 protein isoform (Fig. 4b).
Remarkably, 5 of 19 canonical splice site variants tested were able to rescue cell lethality (Table S2). Enhanced expression of naturally occurring transcripts ∆(E4–E7) or ∆(E12) was detected for variants located in the canonical splice sites of Ex7 and Ex12 (i.e., c.517–2A>G [Ex7], c.631+2T>G ([Ex7], c.6842–2A>G [Ex12], c.6937+1G>A [Ex12]) and is likely responsible for their residual HDR activity (>50%, Fig. 4b). Variant c.7008–2A>T (Ex14) produced multiple alternative transcripts including three out-of-frame transcripts and one in-frame transcript containing a 246-bp (partial) deletion of Ex14 through an exon 14 cryptic acceptor site (Table S2, Fig. 3c). Although this variant was able to partially complement loss of endogenous Brca2, the level of HDR activity (35%) of this variant was severely impaired (Fig. 4b) and possibly results from the relatively low expression level of the potential rescue transcript ∆(E14p246). It should be noted that this variant has been observed in cis with c.631G>A variant for which an effect on RNA splicing (i.e., exon 7 skipping) has also been reported.27
Functional characterization of potential spliceogenic BRCA2 variants
For the vast majority of potential spliceogenic variants that are located outside the canonical splice sites of BRCA2 exons, it is unknown whether they truly affect splicing and if so, to what extent identified aberrant splicing events affect protein functionality. We selected 13 BRCA2 variants for which RNA analysis has been reported in human cells and determined their impact on both mRNA splicing and protein function in mESCs (Figs. 3 and 4c, Table S3).
Overall, human BRCA2 variants in mESCs rendered similar mRNA transcript profiles as previously detected in LCLs and minigene analysis with all major aberrant splicing events identified.12,28,29,30,31 The complementation phenotype of two variants, c.316+5G>C and c.7007G>A, resembled that of high-risk (class 4/5) variants with respect to their inability to rescue the cell lethality imposed by Cre-mediated loss of mBrca2 (Fig. S2). Variant c.316+5G>C only produced transcripts lacking exon 3, which, as discussed above, encodes a stable but nonfunctional protein isoform (Table S3, Figs. 3a and S3). Variant c.7007G>A (last nucleotide Ex13) expressed both two aberrant out-of-frame transcripts (∆[E13] and ∆[E12–E13]) and FL transcript (Fig. 3b). However, expression of the FL transcript was apparently too low to allow complementation (Fig. S3).
Variants c.8754+4A>G and c.9117G>A (last nucleotide Ex23) displayed full complementation of cell lethality but were severely impaired in their HDR capacity (Fig. 4c), in concordance with their recent classification as pathogenic variants.32 However, the nature of the transcript that is responsible for the rescue of cell viability remains elusive. Variant c.425G>T (last nucleotide Ex4) produced an out-of-frame transcript (∆[E4]) and a transcript (∆[E4–E7]) that preserves the reading frame, which is likely responsible for the residual 66% HDR capacity (Figs. 3a and 4c).
Also, deep intronic variants can impose aberrant splicing as previously reported for c.6937+594T>G.28,31 Due to the activation of a cryptic splice site an intronic fragment of 95 bases is inserted between exons 12 and 13 leading to an out-of-frame transcript. Molecular analysis revealed that although intron retention seems to be the predominant splicing event (Fig. 3b) for c.6937+594T>G, the variant allele produced sufficient FL transcript to rescue cell lethality and to retain residual HDR activity (46% compared with WT) (Fig. 4c). For the remaining seven potential spliceogenic variants mRNA splicing in five variants appeared not to be affected while in c.6853A>G and c.9501+3A>T sufficient quantities of FL transcript were produced to prevent substantial loss of BRCA2 function (Fig. 4c).
DISCUSSION
In classification guidelines documented by ACMG and AMP8 and ENIGMA (https://enigmaconsortium.org/), cautionary notes are included for variants that produce in-frame alternative gene transcripts that retain clinically important functional protein domains. Now that we have revealed various functionally redundant regions in the BRCA2 protein, it is possible to propose BRCA2-specific rules. Our results indicate that the majority of the presumed LoF variants will lead to inactivation of the BRCA2 protein, and hence, be associated with high cancer risk (Fig. 5). However, for a number of variants additional analyses will be required before they can be considered to represent pathogenic variants associated with high cancer risk. In particular, for variants in the canonical splice site regions of exons 4, 7, 8, 10, 12, and 14 caution is warranted since LoF may be prevented through elevated expression of in-frame rescue transcripts. Furthermore, nonsense variants, out-of-frame indels, and complete deletion of functionally redundant exons may for the same reasons retain (partial) functionality. Expression of ∆(E4–E7) or ∆(E12) transcripts in mESCs with BRCA2 nonsense variants in exons 6, 7, or 127 was sufficient to retain substantial BRCA2 protein functionality. These findings put into question whether the investigated splice site variants, nonsense variants, and complete exon deletions are associated with high cancer risk. As this model system provides an RNA splicing assay with a direct measure of protein function, the experimental data generated by this functional assay is eminently suited to be applied in variant interpretation. We would like to propose a refined provisional framework for functional evidence application in ACMG/AMP clinical variant interpretation guidelines.9,33,34 The decision tree shown in Fig. 5 may serve as a means to indicate those presumed LoF variants for which the PVS1 code might not be warranted.
Additional data and considerations are needed to determine the appropriate strength of the PS3/BS3 criteria as stated in Brnich et al.33HDR homology directed repair.
In the current multifactorial likelihood model (MLM), a prior probability of pathogenicity is combined with likelihood ratios estimated from clinical data resulting in a final posterior probability that assigns the variant to one of the five classes of the International Agency for Research on Cancer (IARC) classification system.21,35,36 The prior probability is an in silico prediction of the functional impact based on variant location and bioinformatic prediction of variant effect.37,38 Due to the high prior probability assigned to nonsense (0.99) and canonical splice site (0.97) variants the prior heavily impacts the final classification of a variant. However, a high prior might not be justified for presumed LoF variants in functionally redundant exons. For this reason, a reduced prior probability of 0.5 was proposed for variants in BRCA1 exons 9–10 or their proximal splice junction regions.38 Likewise, the prior probability of pathogenicity was set at 0.5 for variants in the splice acceptor and donor site of BRCA2 exon 12. Our results indicate that adjustment of the prior should be extended to other regions of BRCA2 in which presumed LoF variants still display considerable BRCA2 protein activity such as nonsense and splice site variants in exon 7 (Fig. S5). Furthermore, the design of the multifactorial likelihood model restricts its use to discrimination of variants that confer high cancer risk from those that do not. Recent data show that variants associated with reduced penetrance do exist in BRCA1 and BRCA2 and functional analysis might be required to identify these variants.39,40 Recently, Parsons et al.32 have performed multifactorial likelihood analyses for a large number of BRCA1 andBRCA2 variants, including 13 variants that were functionally characterized in this study (Table S4). For most variants, the IARC classification is in agreement with our functional data. However, two variants in respectively the splice acceptor site (c.517–2A>G) and donor site (c.631+2T>G) of exon 7 were classified as pathogenic based on multifactorial likelihood quantitative analysis, while in our analyses these variants show residual HDR capacity in the lower range of class 1/2 variants (Table S4). At this moment, the exact quantitative relationship between BRCA2 protein functionality and cancer risk is still unclear. Although HDR activity around 50% of WT activity was shown to correlate with an odds ratio of 2.5 for breast cancer,40 additional studies are required to define HDR activity ranges that allow assignment of variants to clinically relevant cancer risk categories (i.e., high, moderate, and low increased risk). The observation that presumed LoF variant alleles may retain (partial) functionality through the expression of alternative protein isoforms incites a shift in genetic diagnostics. These findings emphasize the need for inclusion of quantitative functional data to the MLM (as done in a qualitative way in ACMG/AMP guidelines) and specification of gene-specific classification guidelines.
Change history
24 June 2020
The original version of this Article omitted an essential reference (“Meulemans L, Mesman RLS, Caputo SM, et al. Skipping Nonsense to Maintain Function: The Paradigm of BRCA2 Exon 12. Cancer Res. 2020;80(7):1374-1386. DOI: 10.1158/0008-5472.CAN-19-2491”). This has now been added in both the PDF and HTML versions of the Article.
In addition, the original version of this Article contained a typo in the sentence "However, two variants in respectively the splice acceptor site (c.517–2A>G) and donor site (c.631+2T>G) of exon 7 were classified as pathogenic based...", and which was incorrectly given as "However, two variants in respectively the splice donor site (c.517–2A>G) and acceptor site (c.631+2T>G) of exon 7 were classified as pathogenic based...". Where donor must be acceptor and vice versa. This has now been corrected in both the PDF and HTML versions of the Article.
Finally, the original version of this Article contained an incorrect supplementary file. This has now been replaced in the HTML version of the Article.
References
Biswas K, Das R, Alter BP, et al. A comprehensive functional characterization of BRCA2 variants associated with Fanconi anemia using mouse ES cell-based assay. Blood. 2011;118:2430–2442.
de la Hoya M, Soukarieh O, Lopez-Perolio I, et al. Combined genetic and splicing analysis of BRCA1 c.[594-2A>C; 641A>G] highlights the relevance of naturally occurring in-frame transcripts for developing disease gene variant classification algorithms. Hum Mol Genet. 2016;25:2256–2268.
Li L, Biswas K, Habib LA, et al. Functional redundancy of exon 12 of BRCA2 revealed by a comprehensive analysis of the c.6853A>G (p.I2285V) variant. Hum Mutat. 2009;30:1543–1550.
Seo A, Steinberg-Shemer O, Unal S, et al. Mechanism for survival of homozygous nonsense mutations in the tumor suppressor gene BRCA1. Proc Natl Acad Sci U S A. 2018;115:5241–5246.
Thirthagiri E, Klarmann KD, Shukla AK, et al. BRCA2 minor transcript lacking exons 4-7 supports viability in mice and may account for survival of humans with a pathogenic biallelic mutation. Hum Mol Genet. 2016;25:1934–1945.
Wang Y, Bernhardy AJ, Cruz C, et al. The BRCA1-Delta11q alternative splice isoform bypasses germline mutations and promotes therapeutic resistance to PARP inhibition and cisplatin. Cancer Res. 2016;76:2778–2790.
Meulemans L, Mesman RLS, Caputo SM, et al. Skipping Nonsense to Maintain Function: The Paradigm of BRCA2 Exon 12. Cancer Res. 2020;80:1374–1386.
Richards S, Aziz N, Bale S, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–424.
Abou Tayoun AN, Pesaran T, DiStefano MT, et al. Recommendations for interpreting the loss of function PVS1 ACMG/AMP variant criterion. Hum Mutat. 2018;39:1517–1524.
Colombo M, Blok MJ, Whiley P, et al. Comprehensive annotation of splice junctions supports pervasive alternative splicing at the BRCA1 locus: a report from the ENIGMA consortium. Hum Mol Genet. 2014;23:3666–3680.
Fackenthal JD, Yoshimatsu T, Zhang B, et al. Naturally occurring BRCA2 alternative mRNA splicing events in clinically relevant samples. J Med Genet. 2016;53:548–558.
Acedo A, Sanz DJ, Duran M, et al. Comprehensive splicing functional analysis of DNA variants of the BRCA2 gene by hybrid minigenes. Breast Cancer Res. 2012;14:R87.
Borg A, Haile RW, Malone KE, et al. Characterization of BRCA1 and BRCA2 deleterious mutations and variants of unknown clinical significance in unilateral and bilateral breast cancer: the WECARE study. Hum Mutat. 2010;31:E1200–E1240.
Brandao RD, van Roozendaal K, Tserpelis D, Gomez Garcia E, Blok MJ. Characterisation of unclassified variants in the BRCA1/2 genes with a putative effect on splicing. Breast Cancer Res Treat. 2011;129:971–982.
Brandao RD, Mensaert K, Lopez-Perolio I, et al. Targeted RNA-seq successfully identifies normal and pathogenic splicing events in breast/ovarian cancer susceptibility and Lynch syndrome genes. Int J Cancer. 2019;145:401–414.
Chen X, Truong TT, Weaver J, et al. Intronic alterations in BRCA1 and BRCA2: effect on mRNA splicing fidelity and expression. Hum Mutat. 2006;27:427–435.
Claes K, Poppe B, Machackova E, et al. Differentiating pathogenic mutations from polymorphic alterations in the splice sites of BRCA1 and BRCA2. Genes Chromosomes Cancer. 2003;37:314–320.
Colombo M, De Vecchi G, Caleca L, et al. Comparative in vitro and in silico analyses of variants in splicing regions of BRCA1 and BRCA2 genes and characterization of novel pathogenic mutations. PLoS One. 2013;8:e57173.
Fraile-Bethencourt E, Valenzuela-Palomo A, Diez-Gomez B, Caloca MJ, Gomez-Barrero S, Velasco EA. Minigene splicing assays identify 12 spliceogenic variants of BRCA2 exons 14 and 15. Front Genet. 2019;10:503.
Sanz DJ, Acedo A, Infante M, et al. A high proportion of DNA variants of BRCA1 and BRCA2 is associated with aberrant splicing in breast/ovarian cancer patients. Clin Cancer Res. 2010;16:1957–1967.
Thomassen M, Blanco A, Montagna M, et al. Characterization of BRCA1 and BRCA2 splicing variants: a collaborative report by ENIGMA consortium members. Breast Cancer Res Treat. 2012;132:1009–1023.
Mesman RLS, Calleja F, Hendriks G, et al. The functional impact of variants of uncertain significance in BRCA2. Genet Med. 2019;21:293–302.
Hendriks G, Morolli B, Calleja FM, et al. An efficient pipeline for the generation and functional analysis of human BRCA2 variants of uncertain significance. Hum Mutat. 2014;35:1382–1391.
Bjourson AJ, Cooper JE. Band-stab PCR: a simple technique for the purification of individual PCR products. Nucleic Acids Res. 1992;20:4675.
Colombo M, Lopez-Perolio I, Meeks HD, et al. The BRCA2 c.68-7T>A variant is not pathogenic: a model for clinical calibration of spliceogenicity. Hum Mutat. 2018;39:729–741.
Whiley PJ, de la Hoya M, Thomassen M, et al. Comparison of mRNA splicing assay protocols across multiple laboratories: recommendations for best practice in standardized clinical testing. Clin Chem. 2014;60:341–352.
Gaildrat P, Krieger S, Di Giacomo D, et al. Multiple sequence variants of BRCA2 exon 7 alter splicing regulation. J Med Genet. 2012;49:609–617.
Anczukow O, Buisson M, Leone M, et al. BRCA2 deep intronic mutation causing activation of a cryptic exon: opening toward a new preventive therapeutic strategy. Clin Cancer Res. 2012;18:4903–4909.
Bonatti F, Pepe C, Tancredi M, et al. RNA-based analysis of BRCA1 and BRCA2 gene alterations. Cancer Genet Cytogenet. 2006;170:93–101.
Houdayer C, Caux-Moncoutier V, Krieger S, et al. Guidelines for splicing analysis in molecular diagnosis derived from a set of 327 combined in silico/in vitro studies on BRCA1 and BRCA2 variants. Hum Mutat. 2012;33:1228–1238.
Montalban G, Bonache S, Moles-Fernandez A, et al. Screening of BRCA1/2 deep intronic regions by targeted gene sequencing identifies the first germline BRCA1 variant causing pseudoexon activation in a patient with breast/ovarian cancer. J Med Genet. 2019;56:63–74.
Parsons MT, Tudini E, Li H, et al. Large scale multifactorial likelihood quantitative analysis of BRCA1 and BRCA2 variants: An ENIGMA resource to support clinical variant classification. Hum Mutat. 2019;40:1557–1578.
Brnich SE, Abou Tayoun AN, Couch FJ, et al. Recommendations for application of the functional evidence PS3/BS3 criterion using the ACMG/AMP sequence variant interpretation framework. Genome Med. 2019;12:3.
Whiley PJ, Guidugli L, Walker LC, et al. Splicing and multifactorial analysis of intronic BRCA1 and BRCA2 sequence variants identifies clinically significant splicing aberrations up to 12 nucleotides from the intron/exon boundary. Hum Mutat. 2011;32:678–687.
Lindor NM, Guidugli L, Wang X, et al. A review of a multifactorial probability-based model for classification of BRCA1 and BRCA2 variants of uncertain significance (VUS). Hum Mutat. 2012;33:8–21.
Plon SE, Eccles DM, Easton D, et al. Sequence variant classification and reporting: recommendations for improving the interpretation of cancer susceptibility genetic test results. Hum Mutat. 2008;29:1282–1291.
Tavtigian SV, Greenblatt MS, Lesueur F, Byrnes GB, IARC Unclassified Genetic Variants Working Group. In silico analysis of missense substitutions using sequence-alignment based methods. Hum Mutat. 2008;29:1327–1336.
Vallee MP, Di Sera TL, Nix DA, et al. Adding in silico assessment of potential splice aberration to the integrated evaluation of BRCA gene unclassified variants. Hum Mutat. 2016;37:627–639.
Moghadasi S, Meeks HD, Vreeswijk MP, et al. The BRCA1 c. 5096G>A p.Arg1699Gln (R1699Q) intermediate risk variant: breast and ovarian cancer risk estimation and recommendations for clinical management from the ENIGMA consortium. J Med Genet. 2018;55:15–20.
Shimelis H, Mesman RLS, Von Nicolai C, et al. BRCA2 hypomorphic missense variants confer moderate risks of breast cancer. Cancer Res. 2017;77:2789–2799.
Acknowledgements
The authors thank J. Jonkers and P. Bouwman (Netherlands Cancer Institute, Amsterdam, the Netherlands) for the I-SceI mCherry plasmid; S.K. Sharan (National Cancer Institute, Frederick, MD, USA) for the Pl2F7 conditional Brca2 knockout mESC line, M. Jasin (Memorial Sloan-Kettering Cancer Center, New York, USA) for the DR-GFP reporter plasmid, and Mandy Spurdle (QIMR Berghofer Medical Research Institute, Brisbane, Queensland, Australia) for critical reading of the manuscript and suggestions for clarifications. The Dutch/Belgium VUS workgroup and the ENIGMA members are acknowledged for discussions regarding implementation of functional assays into clinical decision making.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosure
The authors declare no conflicts of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, and provide a link to the Creative Commons license. You do not have permission under this license to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Mesman, R.L.S., Calléja, F.M.G.R., de la Hoya, M. et al. Alternative mRNA splicing can attenuate the pathogenicity of presumed loss-of-function variants in BRCA2. Genet Med 22, 1355–1365 (2020). https://doi.org/10.1038/s41436-020-0814-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-020-0814-5
Keywords
This article is cited by
-
Mendelian inheritance revisited: dominance and recessiveness in medical genetics
Nature Reviews Genetics (2023)
-
SpliceAI-visual: a free online tool to improve SpliceAI splicing variant interpretation
Human Genomics (2023)
-
Consistency of variant interpretations among bioinformaticians and clinical geneticists in hereditary cancer panels
European Journal of Human Genetics (2022)
-
Interpretation of BRCA2 Splicing Variants: A Case Series of Challenging Variant Interpretations and the Importance of Functional RNA Analysis
Familial Cancer (2022)
-
Recommendations for clinical interpretation of variants found in non-coding regions of the genome
Genome Medicine (2022)