Abstract
Purpose
This study aims to provide a comprehensive description of the phenotypic and genotypic spectrum of SNAP25 developmental and epileptic encephalopathy (SNAP25-DEE) by reviewing newly identified and previously reported individuals.
Methods
Individuals harboring heterozygous missense or loss-of-function variants in SNAP25 were assembled through collaboration with international colleagues, matchmaking platforms, and literature review. For each individual, detailed phenotyping, classification, and structural modeling of the identified variant were performed.
Results
The cohort comprises 23 individuals with pathogenic or likely pathogenic de novo variants in SNAP25. Intellectual disability and early-onset epilepsy were identified as the core symptoms of SNAP25-DEE, with recurrent findings of movement disorders, cerebral visual impairment, and brain atrophy. Structural modeling for all variants predicted possible functional defects concerning SNAP25 or impaired interaction with other components of the SNARE complex.
Conclusion
We provide a comprehensive description of SNAP25-DEE with intellectual disability and early-onset epilepsy mostly occurring before the age of two years. These core symptoms and additional recurrent phenotypes show an overlap to genes encoding other components or associated proteins of the SNARE complex such as STX1B, STXBP1, or VAMP2. Thus, these findings advance the concept of a group of neurodevelopmental disorders that may be termed “SNAREopathies.”
Similar content being viewed by others
INTRODUCTION
The neuronal SNARE complex (soluble N-ethylmaleimide-sensitive-factor attachment receptor complex) plays a central role in the regulation of synaptic signaling by mediating membrane docking, priming, and fusion of synaptic vesicles with presynaptic membranes. This ultimately leads to the release of neurotransmitters into the synaptic cleft.1,2,3 During neuronal development, it promotes neurite outgrowth and the maturing process of synapses.4
The core SNARE complex comprises a four-helix bundle consisting of two SNAP25 helices (encoded by SNAP25), one syntaxin 1A helix (encoded by STX1A), and one synaptobrevin 2 helix (encoded by VAMP2).1,5
Pathogenic de novo variants disrupting SNARE proteins (VAMP2, MIM 618760) or SNARE complex associated proteins, such as STXBP1 (MIM 612164) and STX1B (MIM 616172), are a known cause for neurodevelopmental disorders consisting of an overlapping phenotype of developmental delay (DD), intellectual disability (ID), and epilepsy6,7,8,9,10 that were recently grouped as “SNAREopathies.”11
SNAP25 showed a significant enrichment for de novo variants in a cohort study of individuals with neurodevelopmental disorders with epilepsy.12 Six individuals with heterozygous variants in SNAP25, four of them of de novo origin, have been described in single case reports showing developmental delay, seizures, and variable neurological symptoms.13,14,15,16,17
The aim of this study is to establish a comprehensive description of the phenotypic spectrum of SNAP25 developmental and epileptic encephalopathy (SNAP25-DEE). We report 19 individuals harboring de novo variants in SNAP25 and review all four previously published individuals with de novo variants determined to be pathogenic or likely pathogenic. Through molecular modeling, we provide further insights into possible mechanisms through which the identified variants may disrupt SNAP25 and the SNARE complex.
MATERIALS AND METHODS
Research cohort and identification of variants
By using matchmaking platforms,18 personal communication with colleagues, and a literature review, 30 individuals harboring heterozygous missense or loss-of-function variants (LoF; nonsense, frameshift, and splice-site variants) in SNAP25 were assessed. No individuals with copy-number variants only encompassing SNAP25 were identified. After thorough evaluation, we included 23 individuals harboring de novo variants determined to be pathogenic or likely pathogenic, including 19 previously unreported individuals. The remaining seven individuals harbored variants of unknown significance (VUS; Supplemental Tables S2, S4.2, and S5.3) and were not included in the phenotypic description.
Phenotypic and genotypic information was obtained from the referring collaborators by using a standardized questionnaire to evaluate clinical, electroencephalography (EEG), and cranial magnetic resonance imaging (cMRI) findings as well as variant information. Variants were identified using trio exome sequencing (ES), trio genome sequencing (GS), or gene panel sequencing.
According to data from gnomAD, SNAP25 (NM_130811.3) shows a reduced number of LoF and missense variants in controls: (1) a probability of loss-of-function intolerance (pLI) score = 0.99 and a loss-of-function observed/expected upper bound fraction (LOEUF) = 0.23 for LoF variants with one nonsense variant deposited at amino acid residue 204, two codons before the canonical stop; and (2) a z-score = 2.96 and LOEUF = 0.36 for missense variants. This indicates a selective constraint on these variant types in a control population not affected by early-onset neurodevelopmental disorders (NDD).19 Therefore, causality of both LoF and missense variants was assessed according to the guidelines of the American College of Medical Genetics and Genomics (ACMG),20 focusing on the following criteria: PS2 (de novo origin), PM2 (absent in population databases), and PP3 (multiple lines of computational evidence support a deleterious effect on the gene/gene product). For in silico prediction of missense variants, the following tools were used: CADD, REVEL, MutationTaster, M-CAP, PolyPhen-2, GERP++.21,22,23,24,25,26
Structural modeling
The structural effects of SNAP25 variants were modeled with SwissModel (Version 4.1.0)27 based on the experimental SNARE-αSNAP complex structures available (PDB codes: 3J96, 3J97, 3J98, 3J99, 6IP1, 6MDN).28,29,30 RasMol (Version 2.7.5)31 was used for structure analysis and visualization.
RESULTS
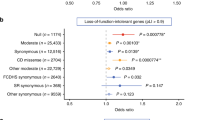
The key clinical findings in SNAP25-DEE comprise DD and/or ID, early-onset seizures, and variable neurological symptoms such as muscular hypotonia, movement disorders (ataxia, dystonia or tremor), cerebral visual impairment (CVI), and brain volume loss. An overview of the main clinical symptoms is presented in Table 1 and Fig. 1a.
(a) Upset-Plot40 of recurrent phenotypic combinations. The colored bars show the absolute number of observations of a certain phenotype in this cohort. The black bars indicate how many individuals presented with a certain phenotypic combination. (b) Location of missense and loss-of-function variants in SNAP25 with respect to domain structure (GenBank: NM_130811.3). Variants above protein scheme are de novo variants reported in this cohort with red circles indicating missense variants and orange squares indicating loss-of-function variants. Below the protein scheme are missense variants in gnomAD with allele count and the degree of coloring being proportionate to the allele count with the lightest gray indicating singletons. (c) Density plot of all missense variants (de novo pathogenic or likely pathogenic variants in red and variants present in gnomAD in blue).
Phenotypic spectrum
Global developmental delay/intellectual disability
All individuals presented with DD and a variable degree of ID, ranging between profound (4/20; 20%), severe (5/20; 25%), moderate (6/20; 30%), and mild (5/20; 25%).
Six individuals aged between 2 and 20 years remained nonverbal (6/17; 35%), with three being adolescent or adult. If language was acquired, individuals could either speak single words (2/17; 12%) or in sentences (9/17; 53%) with articulation noted to be poor or imprecise.
All individuals showed variable degrees of motor delay. Three individuals (3/15; 20%) aged older than three years were not able to walk, while four (4/15; 27%) needed assistance and eight individuals (8/15; 53%) were able to walk on their own.
Regression was reported in five individuals (5/17; 29%) with three of them showing signs of regression with the onset of seizures. This regression was primarily described as a loss of words previously learned.
Seizures
Seizures were reported in 17 individuals (17/23; 74%), while 6 individuals aged 2 months to 16 years had no history of seizures. The age of seizure onset ranged between the 7th day of life to 12 years, with a median age of 12 months. In all but three individuals, seizures started before age 2 years. A broad spectrum including epileptic spasms, generalized and focal seizures were reported with most individuals showing multiple seizure types over time. Primarily generalized or focal to bilateral tonic–clonic seizures were the most common seizure type having occurred in seven individuals (7/17; 41%). Further frequently observed seizure types include absence-like seizures (6/17; 35%) and epileptic spasms (5/17; 29%). Myoclonic, tonic, and atonic seizures were each reported for four individuals (4/17; 24%). Seizures reported in early childhood appeared to be more generalized in onset while older individuals aged 17 to 23 years predominantly exhibited (multi)focal epilepsies corresponding with respective EEG findings of multifocal epileptic discharges and generalized spike wave discharges. Seizure frequency ranged from numerous daily events (8/12; 67%) to isolated seizures (2/12; 11%). Response to antiepileptic drugs (AEDs) was inconsistent for individuals with recurrent seizures: 7/14 individuals (50%), were treated with more than three AEDs and still had frequent seizures.
Brain MRI findings
cMRI was performed on 21 individuals and was unremarkable in 15 individuals (15/21; 71%). Focal or generalized brain volume loss was noted in four individuals (4/21; 19%) aged 7 months to 23 years and signs of a leukoencephalopathy were present in two individuals (2/21; 10%).
Neurological findings
The most common neurological finding was muscular hypotonia (12/19; 63%), with severe hypotonia leading to feeding difficulties being observed in four individuals (4/19; 21%). Spasticity was noted in four individuals (4/21; 19%). Further recurrent findings include movement disorders such as ataxia (7/21; 33%), dystonia (4/21; 19%) and tremor (2/21; 10%). CVI was described in six individuals (6/21; 29%).
Behavior
Most individuals were reported to have no behavioral issues. However, three individuals showed signs of an autism spectrum disorder (3/18; 17%) and four individuals presented with repetitive mannerisms such as hand flapping (4/18; 22%).
Additional findings
Musculoskeletal findings include bilateral clubfeet (5/21; 24%), joint hypermobility (4/21; 19%) and hip dysplasia (2/21; 10%). Most individuals had no or only minor dysmorphic features, with epicanthus being reported for three individuals (3/21; 14%). Further findings included a high-arched palate (4/21; 19%) with abnormal dentition or dental crowding (3/21; 14%; Supplemental Table S1 contains details on all phenotypic findings and Supplemental Table S3.1 lists all observed phenotypes ranked by frequency in standardized terminology according to the Human Phenotype Ontology).
Variant analysis
Of the 23 enrolled individuals with de novo variants, 15 different missense variants were identified, in addition to 4 LoF variants (2 nonsense and 2 splice donor variants).
All 19 variants were absent from the gnomAD database (last accessed July 2020). All pathogenic or likely pathogenic missense variants affected highly conserved amino acid residues (mean GERP++: 5.9) of the t-SNARE coiled-coil homology domain 1 (amino acid position 14–81) and t-SNARE coiled-coil homology domain 2 (amino acid position 135–202; Fig. 1b, c).32 All de novo missense variants were predicted to be deleterious by multiple in silico prediction programs (mean CADD: 29.6; for a complete overview of in silico analysis see Supplemental Tables S5.1 and S5.2).
Recurrent variants comprise the missense variant p.(Gly43Arg) identified in three individuals, the missense variant p.(Met71Thr) identified in two individuals, and the nonsense variant p.(Gln174*) identified in two individuals. Different missense variants affecting the same amino acid residue were also observed including p.(Asp166Gly) and p.(Asp166Tyr) as well as p.(Ala199Gly) and p.(Ala199Val).
In silico structural modeling
SNAP25 is part of the neuronal SNARE complex. The structure of this complex is shown in Fig. 2a,b indicating the positions of the variants detected in the present study. The sidechains of most residues affected are oriented toward the core of the SNARE complex, whereas few are oriented to the outside and interact with αSNAP, a protein that is involved in both SNARE assembly and disassembly after the completion of synaptic vesicle exocytosis.33
(a) Top view of the SNARE-αSNAP complex. The individual proteins are shown in ribbon presentation and in different colors: SNAP25 (blue), Syntaxin (green), VAMP2 (cyan), αSNAP (yellow, orange, red, purple). Residues, for which variants were detected, are shown in space-filled presentation and colored according to their atom type (cpk coloring). (b) Side view of the complex shown in (a). (c) Interactions of Gly43 in the wild-type. Gly43 (gray) is located at a sterically demanding position of the four-helix bundle close to Leu160 of SNAP25 and Phe216 of syntaxin. (d) The longer sidechain of the p.(Gly43Arg) variant forms steric clashes (red dotted circles) with the Leu160 and Phe216 sidechain thereby destabilizing the helix bundle. (e) Interactions of Ala199 in the wild-type. Ala199 (gray) is located in spatial proximity to Phe77 of VAMP2. (f) The longer sidechain of the p.(Ala199Val) variant forms steric clashes (red dotted circle) with the Phe77 sidechain thereby destabilizing the SNAP25-VAMP2 interaction.
To better understand the effects of the identified variants, molecular modeling was performed. This analysis indicated that all pathogenic or likely pathogenic variants were predicted to destabilize the structure of SNAP25 by causing steric clashes (p.[Gly43Arg], p.[Leu57Arg], p.[Gln66Pro], p.[Gln174Pro]), by weakening intramolecular interactions (p.[Leu50Ser], p.[Lys40Glu], p.[Ile192Tyr]), or by enhancing backbone flexibility (p.[Asp166Gly], p.[Ala199Gly], p.[Gln197*]). These structural effects are listed in more detail in the Supplemental Table S3.1. Other variants were predicted to predominantly destabilize the interactions with other components of the SNARE complex, namely STX1A: p.(Ile67Asn), p.(Met71Thr) or VAMP2: p.(Ala199Val). A third group of variants were predicted to result in disturbed interactions with αSNAP by causing steric clashes: p.(Val48Phe), p.(Asp166Tyr). It is important to note that some variants may both disturb SNAP25 structure and interactions, e.g., p.(Gly43Arg) or p.(Met71Thr). The structural effects of the p.(Gly43Arg) and p.(Ala199Val) variants are shown in detail in Fig. 2c–f. Taken together, despite differences in the proposed effects, all variants are expected to destabilize the SNARE complex itself or to affect its interactions with αSNAP (Supplemental Table S3.1).
DISCUSSION
We present a sizable cohort of individuals with pathogenic or likely pathogenic de novo variants in SNAP25 and provide a comprehensive evaluation of an early-onset developmental and epileptic encephalopathy.
Moderate to profound DD and/or ID and early-onset seizures were identified as the core symptoms of SNAP25-DEE. In addition, ataxia, dystonia, CVI, brain volume loss, muscular hypotonia, and spasticity were identified as recurrent clinical symptoms. Individuals with the most severe course of SNAP25-DEE exhibited profound DD, onset of seizures in the first two months of life, spasticity, CVI, and brain volume loss.
Seizure semiology with a differentiation of focal or generalized onset can guide treatment options in individuals with epilepsy. An in-depth age-specific evaluation of seizure semiology was not possible in this comparatively small cohort, yet the adult individuals 5 and 12 exhibited mainly (multi)focal epilepsies. Video EEG data of individual 5 aged 20 years documented a focal epilepsy with parasagittal epileptic discharges corresponding with focal motor seizures (for the full video EEG report see Supplemental Table 1). This could indicate a possible evolution of seizures toward focal to bilateral tonic–clonic seizures in older individuals, but this will require further analysis of age-specific data on seizure semiology as well as their treatment to further elucidate SNAP25-DEE.
Of special interest for future clinical predictions is comparing the clinical course of individuals with recurrent variants or variants affecting the same amino acid residue. The de novo missense variant p.(Gly43Arg) was identified in three individuals. It is remarkable that all individuals showed a rather mild phenotype comprising mild to moderate ID, isolated generalized seizures that did not require treatment (individual 4), or showed good response to AEDs (individual 2) as well as ataxia and tremor. These finding suggests that this specific disruption of amino acid residue 43 could present with a milder variant-specific clinical course within SNAP25-DEE, while other individuals with pathogenic or likely pathogenic variants nearby in residues 40 and 48 presented with a more severe clinical course. Two de novo variants affecting the amino acid residue 166, p.(Asp166Gly) and p.(Asp166Tyr) were identified in individuals 12 and 13 aged 17 and 23 years, respectively. They presented with mild to moderate ID, focal and generalized seizures, and did not show additional neurological symptoms. A different picture is seen for the recurrent de novo variant p.(Met71Thr) that was identified in individuals 10 and 11. Individual 10 was able to speak in sentences at the age of eight years and had no history of seizures while individual 11 was only able to speak single words at age seven years and had daily seizures since the age of two years and six months.
All pathogenic or likely pathogenic missense variants are located in the coiled-coil homology domains 1 and 2 of SNAP25, representing potential hotspots, also considering that little variation is observed in population databases in these domains (see Fig. 1). The variants of four individuals aged 2 to 23 years with notable brain volume loss or signs of a leukoencephalopathy were located in the t-SNARE coiled-coil homology domain 2 (amino acid position 140–202), indicating a possible location-specific clinical observation (4/7; 57% of individuals with variants in this domain).
Future studies with additional individuals with SNAP25-DEE will shed more light onto the underlying clinical course. Recent modeling suggests an incidence of de novo variants in SNAP25 in live births of approximately 1:100.000 for missense variants and 0.1:100.000 for nonsense variants.34
The underlying disease mechanisms for “SNAREopathies” have recently been summarized as very diverse, including many examples of haploinsufficiency due to LoF and missense variants, as well as instances of a dominant negative mechanism.11 Data on functional analyses of variants in SNAP25 is scarce as of now and is only available for p.(Ile67Asn).14 Cotransfection of chromaffin cells with mutant complementary DNA (cDNA) or with wild-type plus mutant cDNA both suppressed vesicle fusion to the same extent, indicating a dominant negative mechanism rather than haploinsufficiency for this variant. In the absence of comprehensive functional analyses, structural modeling of missense variants using published crystal structures can help in predicting the underlying mechanisms by which variants disrupt either SNAP25 or its interaction with other proteins of the SNARE complex. In some instances, individuals harboring variants for which similar structural changes were predicted showed a similar course of disease. The variants identified in individuals 10, 11 (p.[Met71Thr]) and 15 (p.[Ile192Thr]) are both predicted to cause a reduced packing with each other. All three individuals presented with moderate to severe ID while motor development seemed to be only mildly affected with all of them being able to walk on their own. In addition, individuals 11 and 15 presented with a rather late onset of seizures at 2,5 years and 12 years respectively, while individual 10 did not have a history of seizures. Structural modeling of the likely pathogenic variants p.(Gln174Pro) and p.(Arg198Pro) indicated a disruption of the helix of SNAP25 for both variants. Individuals 14 (p.[Gln174Pro]) and 16 (p.[Arg198Pro]) presented with a severe course of disease with a seizure onset at age 2 months, severe to profound DD, CVI, and spasticity. Of further interest are two variants affecting the amino acid residue 199, p.(Ala199Gly) (individual 17) and p.(Ala199Val) (individual 18) with regard to a possible interaction with the SNARE complex protein VAMP2. The variant p.(Ala199Val) is predicted to cause a steric clash with the amino acid residue 77 of VAMP2. In a recent study, three individuals with variants affecting the amino acid positions 75, 77, and 78 of VAMP2 were described.7 All individuals showed overlapping clinical symptoms to the two individuals of the current SNAP25 cohort comprising moderate to severe ID, onset of seizures in the first year of life, muscular hypotonia, ASD, stereotypic hand movements, CVI, absent speech, and unremarkable brain imaging.7 These findings indicate that the disruption of these structural domains in either SNAP25 or VAMP2 may cause a similar downstream functional effect and result in a similar clinical course. Putting these clinical observations in SNAP25-DEE in context to other known disease genes of the neuronal SNARE complex, it becomes clear that a shared clinical course has been repeatedly described, suggesting a shared phenotypic spectrum of the neuronal “SNAREopathies” (see Supplemental Table S6 and Fig. S1)7,10,35
The diverging clinical presentations of individuals with LoF variants allow some hypotheses to be drawn concerning the underlying molecular disease mechanism. Individuals with nonsense variants located in the last and next to last exon of SNAP25 show a more severe clinical course that contrast with the rather mildly affected individuals with splice variants located in the second and third exon. The nonsense variant p.(Gln174*) is located 32 base pairs from the end of exon seven of eight, so nonsense-mediated messenger RNA (mRNA) decay (NMD) cannot be readily assumed, and the variant more likely leads to the translation of a truncated protein.36 The same could be assumed for p.(Gln197*). Supporting causality for both nonsense variants is the fact that they are predicted to truncate SNAP25 by 33 and 10 highly conserved amino acids, respectively. In addition, multiple likely pathogenic missense variants occur downstream of both positions. Whether mechanisms other than haploinsufficiency could be involved in altered protein function in association with these variants remains to be investigated, but a dominant negative mechanism is possible. The two canonical splice-site variants c.72+1G>A, p.(?) and c.114+2T>G, p.(?) are located at donor site of in-frame exons 2 and 3. Both variants could lead to an in-frame exon skipping possibly resulting in a nonfunctional gene product on RNA or protein level that is quickly degraded resulting in haploinsufficiency. (for a report on in silico splice prediction see Supplemental Table S5.2). Another mildly affected individual with mild ID and no seizures inherited the frameshift variant c.464delG p.(Gly155Alafs*84) from his unaffected mother but with a maternal family history of learning difficulties (see individual V6, Supplemental Table 2). This variant was classified as a VUS and although it also likely escapes NMD, the substantial change in the amino acid sequence more likely results in a nonfunctional gene product that is quickly degraded and thereby acting via haploinsufficiency. Mouse models support the potential role of haploinsufficiency in the origin of SNAP25-DEE as SNAP25(+/-) mice display a susceptibility to induced seizures, EEG abnormalities, and cognitive deficits.37 Taken together, haploinsufficiency of SNAP25 may therefore lead to a rather mild phenotype but the preliminary findings reported in this cohort must be complemented by future analyses. Similar observations have been made concerning STX1B, where LoF variants leading to haploinsufficiency cause mild ID with epilepsy whereas missense variants cause a more severe form of DEE.35 This phenomenon of LoF variants resulting in a rather mild course of disease is also known for multiple other NDD genes encoding ion channels, e.g., GRIN2A or KCNQ2.38,39
In summary, de novo variants in SNAP25 cause an early-onset developmental and epileptic encephalopathy mainly characterized by ID and epilepsy, demonstrating a strong phenotypic overlap with disorders caused by the disruption of other components or associated proteins of the SNARE complex, including movement disorders, cerebral visual impairment, and brain atrophy. These findings add to the delineation of a group of disorders that may be called “SNAREopathies.”
Change history
08 March 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41436-020-01090-w
References
Rizo, J. & Xu, J. The synaptic vesicle release machinery. Annu. Rev. Biophys. 44, 339–367, https://doi.org/10.1146/annurev-biophys-060414-034057 (2015).
Jahn, R. & Fasshauer, D. Molecular machines governing exocytosis of synaptic vesicles. Nature. 490, 201–207, https://doi.org/10.1038/nature11320 (2012).
Ramakrishnan, N. A., Drescher, M. J. & Drescher, D. G. The SNARE complex in neuronal and sensory cells. Mol. Cell. Neurosci. 50, 58–69, https://doi.org/10.1016/j.mcn.2012.03.009 (2012).
Hepp, R. & Langley, K. SNAREs during development. Cell Tissue Res. 305, 247–253, https://doi.org/10.1007/s004410100359 (2001).
Han, J., Pluhackova, K. & Böckmann, R. A. The multifaceted role of SNARE proteins in membrane fusion. Front. Physiol. 8, 5, https://doi.org/10.3389/fphys.2017.00005 (2017).
Saitsu, H., Kato, M. & Mizuguchi, T. et al. De novo mutations in the gene encoding STXBP1 (MUNC18-1) cause early infantile epileptic encephalopathy. Nat. Genet. 40, 782–788, https://doi.org/10.1038/ng.150 (2008).
Salpietro, V., Malintan, N. T. & Llano-Rivas, I. et al. Mutations in the neuronal vesicular SNARE VAMP2 affect synaptic membrane fusion and impair human neurodevelopment. Am. J. Hum. Genet. 104, 721–730, https://doi.org/10.1016/j.ajhg.2019.02.016 (2019).
Schubert, J., Siekierska, A. & Langlois, M. et al. Mutations in STX1B, encoding a presynaptic protein, cause fever-associated epilepsy syndromes. Nat. Genet. 46, 1327–1332, https://doi.org/10.1038/ng.3130 (2014).
Redler, S., Strom, T. M. & Wieland, T. et al. Variants in CPLX1 in two families with autosomal-recessive severe infantile myoclonic epilepsy and ID. Eur. J. Hum. Genet. 25, 889–893, https://doi.org/10.1038/ejhg.2017.52 (2017).
Baker, K., Gordon, S. L. & Melland, H. et al. SYT1-associated neurodevelopmental disorder: a case series. Brain J Neurol 141, 2576–2591, https://doi.org/10.1093/brain/awy209 (2018).
Verhage, M. & Sørensen, J. B. SNAREopathies: diversity in mechanisms and symptoms. Neuron. 107, 22–37, https://doi.org/10.1016/j.neuron.2020.05.036 (2020).
Heyne, H. O., Singh, T. & Stamberger, H. et al. De novo variants in neurodevelopmental disorders with epilepsy. Nat. Genet. 50, 1048–1053, https://doi.org/10.1038/s41588-018-0143-7 (2018).
Rohena, L., Neidich, J. & Truitt Cho, M. et al. Mutation in SNAP25 as a novel genetic cause of epilepsy and intellectual disability. Rare Dis. 1, e26314, https://doi.org/10.4161/rdis.26314 (2013).
Shen, X.-M., Selcen, D., Brengman, J. & Engel, A. G. Mutant SNAP25B causes myasthenia, cortical hyperexcitability, ataxia, and intellectual disability. Neurology. 83, 2247–2255, https://doi.org/10.1212/WNL.0000000000001079 (2014).
Liang, J.-S., Wang, J.-S., & Lin L.-J., et al. Genetic diagnosis in children with epilepsy and developmental delay/mental retardation using targeted gene panel analysis. Neuropsychiatry 8, https://doi.org/10.4172/Neuropsychiatry.1000494 (2018).
Hamdan, F. F., Myers, C. T. & Cossette, P. et al. High rate of recurrent de novo mutations in developmental and epileptic encephalopathies. Am. J. Hum. Genet. 101, 664–685, https://doi.org/10.1016/j.ajhg.2017.09.008 (2017).
Fukuda, H., Imagawa, E. & Hamanaka, K. et al. A novel missense SNAP25b mutation in two affected siblings from an Israeli family showing seizures and cerebellar ataxia. J. Hum. Genet. 63, 673–676, https://doi.org/10.1038/s10038-018-0421-3 (2018).
Sobreira, N., Schiettecatte, F., Valle, D. & Hamosh, A. GeneMatcher: a matching tool for connecting investigators with an interest in the same gene. Hum. Mutat. 36, 928–930, https://doi.org/10.1002/humu.22844 (2015).
Lek, M., Karczewski, K. J. & Minikel, E. V. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature. 536, 285–291, https://doi.org/10.1038/nature19057 (2016).
Richards, S., Aziz, N. & Bale, S. et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 17, 405–424, https://doi.org/10.1038/gim.2015.30 (2015).
Rentzsch, P., Witten, D. & Cooper, G. M. et al. CADD: predicting the deleteriousness of variants throughout the human genome. Nucleic Acids Res. 47, D886–D894, https://doi.org/10.1093/nar/gky1016 (2019).
Ioannidis, N. M., Rothstein, J. H. & Pejaver, V. et al. REVEL: an ensemble method for predicting the pathogenicity of rare missense variants. Am. J. Hum. Genet. 99, 877–885, https://doi.org/10.1016/j.ajhg.2016.08.016 (2016).
Schwarz, J. M., Rödelsperger, C., Schuelke, M. & Seelow, D. MutationTaster evaluates disease-causing potential of sequence alterations. Nat. Methods 7, 575–576, https://doi.org/10.1038/nmeth0810-575 (2010).
Jagadeesh, K. A., Wenger, A. M. & Berger, M. J. et al. M-CAP eliminates a majority of variants of uncertain significance in clinical exomes at high sensitivity. Nat. Genet. 48, 1581–1586, https://doi.org/10.1038/ng.3703 (2016).
Adzhubei, I., Jordan, D. M. & Sunyaev, S. R. Predicting functional effect of human missense mutations using PolyPhen-2. Curr. Protoc. Hum.Genet. Chapter 7, Unit7.20, https://doi.org/10.1002/0471142905.hg0720s76 (2013).
Cooper, G. M., Stone, E. A. & Asimenos, G. et al. Distribution and intensity of constraint in mammalian genomic sequence. Genome Res. 15, 901–913, https://doi.org/10.1101/gr.3577405 (2005).
Guex, N. & Peitsch, M. C. SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis. 18, 2714–2723, https://doi.org/10.1002/elps.1150181505 (1997).
Zhao, M., Wu, S. & Zhou, Q. et al. Mechanistic insights into the recycling machine of the SNARE complex. Nature. 518, 61–67, https://doi.org/10.1038/nature14148 (2015).
Huang, X., Sun, S. & Wang, X. et al. Mechanistic insights into the SNARE complex disassembly. Sci. Adv. 5, eaau8164, https://doi.org/10.1126/sciadv.aau8164 (2019).
White, K.I., Zhao, M., & Choi U.B., et al. Structural principles of SNARE complex recognition by the AAA+ protein NSF. eLife 7, https://doi.org/10.7554/eLife.38888 (2018).
Sayle, R. A. & Milner-White, E. J. RASMOL: biomolecular graphics for all. Trends Biochem. Sci. 20, 374, https://doi.org/10.1016/s0968-0004(00)89080-5 (1995).
The UniProt Consortium. UniProt: a worldwide hub of protein knowledge. Nucleic Acids Res. 47, D506–D515, https://doi.org/10.1093/nar/gky1049 (2019).
Ma, L., Kang, Y. & Jiao, J. et al. α-SNAP enhances SNARE zippering by stabilizing the SNARE four-helix bundle. Cell Rep. 15, 531–539, https://doi.org/10.1016/j.celrep.2016.03.050 (2016).
López-Rivera, J. A., Pérez-Palma, E. & Symonds, J. et al. A catalogue of new incidence estimates of monogenic neurodevelopmental disorders caused by de novo variants. Brain J Neurol 143, 1099–1105, https://doi.org/10.1093/brain/awaa051 (2020).
Wolking, S., May, P. & Mei, D. et al. Clinical spectrum of STX1B-related epileptic disorders. Neurology. 92, e1238–e1249, https://doi.org/10.1212/WNL.0000000000007089 (2019).
Popp, M. W.-L. & Maquat, L. E. Organizing principles of mammalian nonsense-mediated mRNA decay. Annu. Rev. Genet. 47, 139–165, https://doi.org/10.1146/annurev-genet-111212-133424 (2013).
Corradini, I., Donzelli, A. & Antonucci, F. et al. Epileptiform activity and cognitive deficits in SNAP-25(+/-) mice are normalized by antiepileptic drugs. Cereb. Cortex 24, 364–376, https://doi.org/10.1093/cercor/bhs316 (2014).
Strehlow, V., Heyne, H. O. & Vlaskamp, D. R. M. et al. GRIN2A-related disorders: genotype and functional consequence predict phenotype. Brain J. Neurol. 142, 80–92, https://doi.org/10.1093/brain/awy304 (2019).
Weckhuysen, S., Mandelstam, S. & Suls, A. et al. KCNQ2 encephalopathy: emerging phenotype of a neonatal epileptic encephalopathy. Ann. Neurol. 71, 15–25, https://doi.org/10.1002/ana.22644 (2012).
Lex, A., Gehlenborg, N. & Strobelt, H. et al. UpSet: visualization of intersecting sets. IEEE Trans. Vis. Comput. Graph. 20, 1983–1992, https://doi.org/10.1109/TVCG.2014.2346248 (2014).
Acknowledgements
We thank the patients and their families for their participation and support of this study. We especially thank Elizabeth Dellureficio for her continuous efforts to bring affected families together. Work on individual 5 was supported in part by grants from SFARI and the JPB Foundation. We thank the clinicians involved with the care of these patients including Stephen Nirmal, Alasdair Parker, and Manali Chitre, UK. A.M., K.B., and M.A.K. are funded by the National Institute for Health Research (NIHR) GOSH BRC. The views expressed are those of the author(s) and not necessarily those of the NHS, the NIHR or the Department of Health. Individual 7 was enrolled in the NIHR BioResource research study, supported by the Cambridge Biomedical Research Centre and the NIHR for the NIHR BioResource (grant number RG65966). Individual 17 was ascertained in the Duke Genome Sequencing Clinic. Funding for the Duke Genome Sequencing Clinic which is supported by the Duke University Health System. Individual V6 was enrolled in Care4Rare Canada Consortium, funded by Genome Canada, Ontario Genomics Institute (OGI-147), Canadian Institutes of Health Research, Ontario Research Fund, Genome Alberta, Genome British Columbia, BC Children’s Hospital Foundation, BC Children’s Hospital Research Institute, BC Provincial Health Services Authority, Genome Quebec, and Children’s Hospital of Eastern Ontario Foundation. This study makes use of data generated by the DECIPHER Consortium. A full list of centers who contributed to the generation of the data is available from https://decipher.sanger.ac.uk/ and via email from decipher@sanger.ac.uk. Funding for the project was provided by the Wellcome Trust. R.S.M. was supported by a grant from the Lundbeck Foundation (R277-2018-802). Funding for the Duke Genome Sequencing Clinic was provided by Duke University Health System. Research reported in this publication was supported by the National Human Genome Research Institute of the National Institutes of Health under award number U01HG009599. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. We thank Katherine R. Chao for her help with the exome data analysis. The work in C.G. Bönnemann’s laboratory is supported by intramural funds from the NIH National Institute of Neurological Disorders and Stroke. Sequencing and analysis were provided by the Broad Institute of MIT and Harvard Center for Mendelian Genomics (Broad CMG) and was funded by the National Human Genome Research Institute, the National Eye Institute, and the National Heart, Lung and Blood Institute grant UM1 HG008900 to Daniel MacArthur and Heidi Rehm.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Competing interests
J.K.R. is an employee of GeneDx, Inc. N.S. is an employee and holds equity in Bristol Myers Squibb. H.M.M. is an employee and shareholder at Invitae Corporation. The other authors declare no competing interests.
Ethics declaration
All examined individuals or their legal guardians provided informed written consent for testing and publication. In some cases, testing was done as part of routine clinical care and therefore institutional ethics approval was not required. If done in a research setting, testing was approved by the ethics committees of the respective centers: the University of Leipzig (approval code: 402/16-ek), the Columbia Institutional Review Board, the National Institute of Neurological Disorders and Stroke, National Institutes of Health (protocol number 12-N-0095), the Duke Genome Sequencing Clinic (protocol number 00032301), the University of British Columbia institutional ethics review board (H10-03215), the Technical University Munich (#5360/12S), the East of the England Cambridge South national institutional review board (13/EE/0325), and the London Bloomsbury Research Ethics Committee.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised: Unfortunately, in the first sentence in the abstract, a spelling mistake was introduced during the production process after our proofreading. The wording was changed from “aims” to “aimsed.” In addition, Heather M. McLaughlin was not listed among the authors.
Supplementary information
Rights and permissions
About this article
Cite this article
Klöckner, C., Sticht, H., Zacher, P. et al. De novo variants in SNAP25 cause an early-onset developmental and epileptic encephalopathy. Genet Med 23, 653–660 (2021). https://doi.org/10.1038/s41436-020-01020-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-020-01020-w