Abstract
Purpose
Pseudoxanthoma elasticum (PXE) is a heritable disorder affecting elastic fibers in the skin, eyes, and cardiovascular system. It is caused by biallelic pathogenic variants in the ABCC6 gene. To date, over 300 ABCC6 variants are associated with PXE, more than half being missense variants. Correct variant interpretation is essential for establishing a direct link between the variant and the patient’s phenotype and has important implications for diagnosis and treatment.
Methods
We used a systematic approach for interpretation of 271 previously reported and 15 novel ABCC6 missense variants, based on the semiquantitative classification system Sherloc.
Results
Only 35% of variants were very likely to contribute directly to disease, in contrast to reported interpretations in ClinVar, while 59% of variants are currently of uncertain significance (VUS). Subclasses were created to distinguish VUS that are leaning toward likely benign or pathogenic, increasing the number of (likely) pathogenic ABCC6 missense variants to 47%.
Conclusion
Besides highlighting discrepancies between the Sherloc, American College of Medical Genetics and Genomics and the Association for Molecular Pathology (ACMG-AMP), ClinVar, and Leiden Open Variation Database (LOVD) classification, our results emphasize the need for segregation analysis, functional assays, and detailed evidence sharing in variant databases to reach a confident interpretation of ABCC6 missense variants and subsequent appropriate genetic and preconceptual counseling.
Similar content being viewed by others
INTRODUCTION
Pseudoxanthoma elasticum (PXE, OMIM 264800) is an autosomal recessive (AR) disorder in which mineralization and fragmentation of elastic fibers lead to symptoms in the skin (papular skin lesions in flexural areas), eyes (angioid streaks, peau d’orange, subretinal neovascularization, and hemorrhage) and (cardio)vascular system (peripheral artery disease, gastrointestinal bleeding, ischemic stroke).1 Age of onset, disease severity, and natural evolution are highly variable between patients and families.2
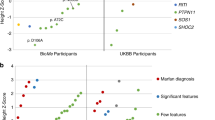
In 2000, pathogenic variants in the ABCC6 gene (OMIM *603234) were identified in PXE.3,4,5 The 31 exons of ABCC6 encode a 1503 amino acid–long adenosine triphosphate (ATP)-dependent transmembrane transporter that is mainly expressed at the basolateral membrane in liver and kidney cells (Fig. 1).6 ABCC6 has structural similarity with other ABC transporters of the subfamily C, with three transmembrane domains (TMDs) and two nucleotide binding domains (NBDs). Though ABCC6 was recently shown to be involved in the homeostasis of the calcification inhibitor inorganic pyrophosphate (PPi), its biological substrate(s) remain elusive.7
(a) The ABCC6 gene is located on 16p13.11 and is flanked by two pseudogenes ABCC6P1 and ABCC6P2 (genomic positions according to build GRCh38). The arrows denote the direction of gene transcription. The transcript of ABCC6 contains 31 exons. ABCC6P1 and ABCC6P2 are homologous with the region spanning from exon 1 to intron 9, and exon 1 to intron 4 respectively. (b) The ABCC6 transporter contains the following structures: 3 transmembrane domains (TMD0-2), 2 linker domains (L0-1), and 2 nucleotide binding domains (NBD1–2). Conserved sequences within the NBD domains are: A-loop (A), Walker A (WA), Q-loop (Q), LSGGQ-signature (C), Walker B (WB), D-loop (D), H-loop (H) and PDZ-like sequence. Frequently mutated regions are indicated in gray circles. EC extracellular, IC intracellular, P plasma membrane.
Variant detection in ABCC6 is complicated by the presence of two pseudogenes—ABCC6P1 and ABCC6P2—with high sequence homology with the first nine and first four exons respectively.8 To ensure that sequencing results are specific for ABCC6, distinct primers have been designed for exons 1 to 9 based on nucleotide differences between parental ABCC6 and its pseudogenes.8 Today, over 300 ABCC6 variants have been reported in the National Center for Biotechnology Information (NCBI) ClinVar database. These include 38 nonsense variants, 51 frameshift, 26 splice-site, and 240 missense variants. Because of their abundance, more than 50% of patients harbor at least one missense variant in their genotype.2,9 However, in contrast to variants leading to nonsense mediated decay or truncated protein formation, the impact of these missense variants on protein function is less predictable. They can disrupt protein function by altering the conformational stability, trafficking, and/or transport activity of the protein, though many will be very well tolerated. For this reason, criteria to classify missense substitutions received much attention. Different classification approaches however resulted in variable interpretations of the same variant.10 In 2015, the American College of Medical Genetics and Genomics and the Association for Molecular Pathology (ACMG-AMP) established guidelines for variant interpretation, in which modifiers such as “benign,” “likely benign,” “uncertain significance,” “likely pathogenic,” and “pathogenic” were used to stratify the pathogenicity gradient.11 Despite this important effort, it was disputed that certain criteria were ambiguous, too clinical, or lacked specificity.12,13 Recently, a refinement of these guidelines was proposed, coined Semiquantitative, Hierarchical Evidence-based Rules for LOCus interpretation (Sherloc).13 Sherloc offers a numerical score-based system in which evidence types are grouped and supported by hierarchical decision trees, producing reproducible variant interpretation results.
As more than half of ABCC6 variants are missense substitutions, a uniform and reproducible approach for variant classification is essential for elucidating their causal nature, though this has never been done in a systematic way. We performed a systematic review and reassessed the pathogenicity of 271 previously reported and 15 newly identified ABCC6 missense variants using population, clinical, segregation, experimental, and in silico predictive data within the Sherloc classification framework. We compared results with the ACMG-AMP, ClinVar, and Leiden Open Variation Database (LOVD) classifications. Because a significant part of the missense variants were class 3 (uncertain significance), which remain difficult to use in a clinical setting, we made adjustments to the Sherloc framework to allow a more reliable prediction of which class 3 variants are still likely to be disease-causing.
MATERIALS AND METHODS
Ethics statement
Written, informed consent was obtained from all patients, and the Declaration of Helsinki protocols were followed. This study was approved by the local ethical committees (Ghent University Hospital).
Collection of ABCC6 missense variants and evidence
The 286 missense variants included in this study originate from ClinVar ((http://www.ncbi.nlm.nih.gov/clinvar/) and LOVD (https://www.LOVD.nl/ABCC6), our in-house database or systematic literature review (Figure S1). The clinical and functional evidence gathered for each variant are summarized in Table S1, along with their classification result. Genotypes of PXE patients from our in-house database are listed in Table S2. C-notations are based on NM_001171.6 and p-notations on NP_001162.5.
Sherloc adjustments and ABCC6-specific considerations
We used a scoring system based on Sherloc version 4.213 (Fig. 2). The rules and guidelines were adapted to fit the scope of evaluating ABCC6 missense variants, as shown in Figure S2. We made two adjustments to the framework: (1) because the Genome Aggregation Database (gnomAD) includes bigger population data sets than the Exome Aggregation Consortium (ExAC), the allele frequency (AF) threshold for “somewhat high” of “allele count of 8” was replaced by “0.01%,”14 and (2) we decided not to consider multiple unrelated observations as additive, as this would lead to overestimation of pathogenicity. Instead, we used the highest scoring observation in case of multiple observations, but an extra pathogenic point (1 P) was given when more than ten unrelated observations were found.
Missense variants were collected along with available clinical and functional evidence, which were scored using a classification approach based on Sherloc.13 Based on the total score, the variant was assigned to one of five classes (C1–C5). For class 3, variants were subdivided among three subclasses: those with a likely benign or pathogenic tendency and those that are truly uncertain. Sherloc-derived classification results were compared with American College of Medical Genetics and Genomics–Association for Molecular Pathology (ACMG-AMP)–derived result and with interpretations reported in ClinVar and/or Leiden Open Variation Database (LOVD) if available. AF allele frequency, B benign point, P pathogenic point, VUS variants of uncertain significance.
In our analysis, we considered the following ABCC6 functional domains: linker domain L0 (residues 189–302), NBD1 (residues 623–859), and NBD2 (residues 1254–1503) including the PDZ-like sequence (residues 1498–1503).15,16,17,18 Exon 24 was considered a mutational hotspot.2 Motifs residing in NBDs that are highly conserved among ABC transporters and are important for function are listed in Table S3 and were considered “critical” residues. Experimental evidence included results from validated, qualitative functional assays evaluating protein function and localization.18,19,20,21,22 Results from morpholino-based zebrafish messenger RNA (mRNA) rescue assays were excluded as a high risk for nonspecificity is reported.23 The following prediction tools were used to produce computational data: sequence and evolutionary conservation-based Align-GVGD24 and MutationAssessor,25 protein sequence and structure-based PolyPhen-2,26 and supervised learning algorithm MutationTaster27 (Table S4). A “deleterious” or “not deleterious” effect was achieved when ≥3 predictions tools reached a consensus.
Variant classes and subclasses
Based on the total score, the variant is assigned to one of following classes: class 1 for benign variants, class 2 for likely benign variants, class 3 for variants of uncertain significance (VUS), class 4 for likely pathogenic variants, and class 5 for pathogenic variants (Fig. 2). To gain more insight into the possible pathogenic nature of class 3 variants, we created three subclasses: class 3 variants with a tendency toward likely pathogenic (class 3LP) or likely benign (class 3LB), and those that are truly uncertain (class 3U). Based on the consensus that genetic variation is mostly neutral and pathogenic variants are rare,13 the point thresholds for class 3LB and 3LP were kept asymmetrical (2 benign points and 3 pathogenic points respectively), similar to Sherloc point thresholds for class 2 and class 4 (Fig. 2).
Refinement of classification results
For each variant, the same evidence was used to classify them using the ACMG-AMP guidelines (Table S5). Sherloc results were also compared with the corresponding interpretation in ClinVar and in LOVD if available (Tables S1 and S8).
We used Site Directed Mutator (SDM)28 to predict protein stability of variants using a validated model for NBD1 of ABCC6 6BZS in the Research Collaboratory for Structural Bioinformatics Protein Data Bank (RCSB PDB) as input18 (Table S6).
RESULTS
Impact of the ABCC6 pseudogenes
If not careful, variants occurring in one of the pseudogenes can be mistaken for a potential disease-causing variant. Two approaches are commonly used to avoid the detection of pseudogenic variants: (1) using allele specific primers that are designed based on nucleotide differences between ABCC6 and its pseudogenes or (2) using long range polymerase chain reaction (PCR) followed by nested PCR.8 Some variants can however occur in both the parental and pseudogene, and are not necessarily neutral. For example, p.(Arg64Gln) (class 3U), p.(Ala78Thr) (class 3U), p.(Glu125Lys) (class 3U), and p.(Arg265Gly) (class 1) were found as pseudogenic variants, but the same variants were found in the parental gene in other studies (Table S7). Only variants that were proven to be ABCC6-specific were used for our analysis. Three variants (p.[Ala158Val], p.[Lys281Glu], and p.[Ile319Val]) were excluded from analysis because they were only reported in the pseudogene or it was not possible to confirm ABCC6-specificity (Table S7).
ABCC6 variants with founder effect
Because the AF of a variant is a strong indicator of its benign or pathogenic character, a founder effect—when a few individuals from an original population establish a new population that becomes isolated—resulting in elevated variant AF can lead to misclassification of variants. The most common disease-causing ABCC6 missense variants in the European-derived South Afrikaner PXE population (p.[Arg1138Gln] and p.[Arg1339Cys], both class 5) were shown to originate from common ancestors.21 Also p.(Arg518Gln) (class 5) was shown to be a founder variant in English speaking South African PXE patients.29 Nonetheless, the AF of these variants falls within pathogenic range.
However, for the variants p.(Glu125Lys) (class 3LB), p.(Arg887Cys) (class 3LB), and p.(Gly1327Glu) (class 2), the AF is elevated in the African population (0.3%, 0.3%, and 1.4% respectively). For p.(Ala950Thr) (class 3LB) the AF is higher in the Ashkenazi Jewish population (1%), and p.(Arg1141Gln) (class 3LB) is more common in the South Asian population (0.4%). By consequence, the total AF observed for these five variants falls outside the pathogenic range (>0.01%). Systematic literature review did not provide substantial evidence of a founder effect for these variants so that total AF was used to evaluate these variants. Their classification should however be interpreted with caution because of the observed AF.
Classification of ABCC6 missense variants with Sherloc
Evaluation of 286 ABCC6 missense variants with Sherloc revealed the majority to be class 3 variants (59%), followed by class 5 (23%), class 4 (12%), class 1 (4%), and class 2 (2%) (Fig. 3a, Table S1).
(a) Classification of ABCC6 missense variants into 5 classes (C1–C5). Class 3 variants were subdivided into those with likely benign tendency (C3LB), uncertain (C3U), and likely pathogenic tendency (C3LP). (b) Localization of variants within the ABCC6 protein structure. Light gray bars represent the total number of variants per functional domain. Dark gray bars represent the number of variants of each Sherloc-derived (sub)class per functional domain. L0 first linker domain, L1 second linker domain flanking NBD1, NBD nucleotide binding domain, TMD transmembrane domain.
Class 1 variants (n = 12) are supported by substantial benign clinical evidence including a high AF (>1% or > 0.3%) and abundant homozygous observations (ranging from 8 to 138,663) in gnomAD (Table S1). None of these variants are reported as founder or recurrent variant in affected individuals. In case of discrepancy, the clinical data (i.e., the variant is present in a large percentage of healthy individuals) are more persuasive than the functional data (i.e., the variant is present in a functional domain or is predicted to be not deleterious). All have been reported (likely) benign in ClinVar, except for p.(Arg265Gly), which has conflicting interpretations.
Class 2 variants are the least represented (n = 5; Table S1). All have an elevated AF (>0.01%) not consistent with disease, and were homozygously found in two or more individuals in gnomAD. The latter cannot be seen as an absolute criterion, because—although fully penetrant—the severity of the PXE phenotype is age dependent. Therefore, the possibility of seemingly unaffected individuals with a mild or late phenotype being included in gnomAD populations cannot be completely ruled out. The high AF, however, points toward a benign effect. Segregation analysis is needed to clarify the relationship between these variants and PXE. ClinVar interpretations varied from likely benign to pathogenic.
Class 3 variants—scoring not enough points in either direction—are most abundant (n = 169) and most difficult to interpret (Table S1). To gain more insight in the likelihood of their benign or pathogenic nature, we isolated class 3 variants with a tendency toward likely pathogenic (class 3LP) or likely benign (class 3LB) from the truly uncertain variants (class 3U). As such, 9% (15/169) of class 3 variants became class 3LB, 20% (34/169) class 3LP, and 71% (120/169) remained class 3U (Fig. 3a).
The class 3LB variants have elevated AFs (>0.01%) and lacked evidence from segregation analysis. Experimental evidence was available for p.(Glu125Lys) and p.(Val787Ile), the first showing no impact on membrane localization and the latter showing minimal impact on ABCC6 biosynthesis and folding respectively.18,21 The majority of class 3LB variants (13/15) were predicted to be not deleterious. Nine of 15 were interpreted as (likely) benign by ClinVar.
Class 3LP variants have an AF within pathogenic range or are absent in gnomAD. One novel variant was found: p.(Ala452Asp). Those that were reported in ClinVar (n = 28/34) were indicated as pathogenic.
Class 3U variants often did not receive any points from clinical observations (93/120) because they did not meet the requirements: information was lacking (65/93), variant was either heterozygous (17/93) or acting in cis (2/93), or the individual had an alternate cause of disease (7/93) or did not have the required phenotype (2/93). The 3U class includes ten novel variants and two variants that correspond to pseudogenic sequence differences (Table S7). One remarkable variant in this group is p.(Arg391Gly). This variant has been reported repeatedly in PXE patients and was shown to segregate with disease in families in a compound heterozygous state.9,30,31,32 Recently, the variant was observed in 2 unrelated β-thalassemia patients with PXE symptoms.33 Another study showed that this variant is associated with late-onset PXE, and suggests it acts as a hypomorphic variant.34 On the other hand, gnomAD AF for this variant is high (>0.3%) in nearly every population and includes five homozygous observations, supporting a benign classification. Taken together, this variant is supported by substantial pathogenic and benign evidence.
Class 4 variants (n = 34) often affect a residue involved in another (likely) pathogenic variant (5/34) (Table S1). This group includes three novel variants: p.(Trp218Arg), p.(Ser633Ile), and p.(Ala1385Pro). Seven class 4 variants were shown to interfere on the protein level. Variants p.(Leu677Pro), p.(Leu726Pro), p.(Asp778Asn), and p.(Ser1307Pro) affected expression, protein folding, or full length processing,18,22 Variants p.(Ser1121Trp), p.(Thr1301Ile) display partial intracellular retention which could be restored by the chemical chaperone 4-phenylbutyrate (4-PBA).35 The intracellularly retained variant p.(Gly1321Ser) abolished ABCC6 transport activity.19,20,35 Variant p.(Arg419Gln) shows proper membrane localization, while p.(Glu699Asp) showed minimal changes in protein folding and full length expression, suggesting that abnormal subcellular trafficking or protein synthesis are not the pathomechanisms associated with PXE for these variants.18,21
Class 5 variants (n = 66) are frequently reported in large screening studies of PXE patients and are usually well documented (Table S1). A substantial number (17/66) affect critical residues (Table S3). Most of the variants reported in ClinVar (52/66) were designated as pathogenic in ClinVar. Frequently reported variants in PXE patients are p.(Thr364Arg), p.(Arg518Gln), p.(Gly755Arg), p.(Arg765Gln), p.(Arg807Gln), p.(Arg1114Cys), p.(Arg1114His), p.(Thr1130Met), p.(Arg1138Trp), p.(Arg1138Gln), p.(Arg1164Gln), p.(Arg1221His), p.(Gly1302Arg), p.(Ala1303Pro), p.(Arg1314Trp), p.(Arg1314Gln), p.(Arg1339Cys), p.(Arg1339His), p.(Glu1400Lys), and p.(Arg1459His). One novel class 5 variant was found: p.(Ile808Thr). For several class 5 variants, experimental data has confirmed their pathogenicity: p.(Arg1114Pro), p.(Arg1138Gln), p.(Arg1314Trp), p.(Arg1339Cys), and p.(Gln1347His) were (partially) retained in the cytoplasm of mouse hepatocytes.21,35 Reduced protein solubility and/or reduced full length processing was observed for p.(Gly663Cys), p.(Leu673Pro), p.(Arg760Trp), p.(Arg765Gln), p.(Ala766Asp), p.(Asp777Asn), p.(Val810Met), p.(Thr811Met), p.(Ala820Pro), p.(Leu826Pro), p.(Gly1299Ser), p.(Leu1335Pro), and p.(Arg1339Cys), however, not for p.(Gln698Pro).18,22 In turn, transport assays showed diminished transport activity for synthetic substrates for p.(Val1298Phe) and p.(Gly1302Arg).19,20,35
Location analysis reveals that missense variants occur throughout the entire ABCC6 structure (Fig. 3b). Notably, nearly half of class 1 (6/12), class 3LP (14/34), and class 4 (15/34) variants reside in one of the NBDs. The NBDs also harbor most class 5 variants (46/66). The majority of class 2 variants (4/5) and class 3U variants (72/120) reside in the transmembrane domains (Fig. 3b). Moreover, one third of the variants in TMD2 are located in exon 24, which forms the eight cytoplasmatic loop (Table S1).
Differences between ACMG-AMP and Sherloc
Classifying the same ABCC6 missense variants with the ACMG-AMP guidelines (Table S5) shows a 69% concordance between ACMG-AMP and Sherloc (Fig. 4a, b). Discordance was mainly caused by a conflict between VUS (class 3) and (likely) pathogenic (class 4/5) classification, and to a lesser extent by a confidence conflict between likely pathogenic (class 4) and pathogenic (class 5). If confidence conflicts are not considered as discordant, then concordance increases to 80%. Notably, no clinically significant conflict between (likely) benign (class 1/2) and (likely) pathogenic (class 4/5) was observed (Fig. 4c).
(a) Classification results are expressed in percentages of total number of variants (Sherloc and ACMG-AMP; n = 286) or total number of variants for which a ClinVar or LOVD interpretation was found (ClinVar: n = 237; LOVD: n = 38). The remaining 3% of ClinVar interpretations are conflicting. (b) Concordance (C) was the highest between Sherloc and ACMG-AMP results. Conversely, most discordance (D) was observed between Sherloc and ClinVar results. (c) For the discordant results the contribution of each type of conflict was expressed as a percentage of total discordant results. B benign, C1 class 1, C2 class 2, C3 class 3, C4 class 4, C5 class 5, LB likely benign, LP likely pathogenic, P pathogenic, VUS variant of uncertain significance.
Concordance with ClinVar and LOVD
Two hundred thirty-seven variants of 286 were reported in ClinVar. Only 36% of our Sherloc-derived classification results were concordant with ClinVar variant interpretation (Fig. 4b). Similar to ACMG-AMP, a conflict between VUS (class 3) and (likely) pathogenic (class 4/5) classification is the main reason for discordance (Fig. 4c). Concordance increases to 49% when confidence conflicts are not considered as discordant. Only two clinically significant conflicts were observed: the class 2 variant p.(Val514Ile) was submitted as pathogenic in ClinVar, but no assertion criteria were provided; and the class 5 variant p.(Arg1357Trp) was interpreted as benign, with criteria provided by a single submitter.
One hundred eighty-one variants of 286 were reported in LOVD, the majority with multiple entries per variant (143/181) (Table S8). To date, only a small number of variants (39/181) are assigned with a clinical classification result in LOVD. Comparing these classification results with those obtained with Sherloc revealed 44% concordance (Fig. 4b). Again, a conflict between VUS (class 3) and (likely) pathogenic (class 4/5) was most observed (Fig. 4c). When confidence conflicts are not considered, concordance increases to 62%. No clinically significant conflicts were observed.
DISCUSSION
In this study, we evaluated the pathogenic nature of reported and novel ABCC6 missense variants using the integrative Sherloc classifying system, taking into account the effects of the ABCC6 pseudogenes and described or suspected founder effects. Caution is required when interpreting variants that are detected within the pseudogenic region spanning from exon 1 to 9. A previously reported pseudogenic variant occurring in the parental gene should not be dismissed as benign and vice versa.
We demonstrate that the majority of ABCC6 missense variants are class 3 variants (VUS). This is not surprising, considering that the classification is a snapshot of available evidence at a given time and most variants do not have enough evidence to support a classification of (likely) benign or (likely) pathogenic at this point. Indeed, segregation data providing information about zygosity and segregation are missing for many variants. Cosegregation with disease phenotype in one or more families supports the direct link between a variant and disease. In AR diseases, co-occurrence with a (likely) pathogenic variant in trans can be suggestive of a pathogenic nature. On the other hand, a variant co-occurring on the same allele with a (likely) pathogenic variant is less likely to be pathogenic, because there is no selective pressure against accumulation of additional variants in that allele. Twenty-five class 3 variants are reported as part of an apparent compound heterozygous genotype with a (likely) pathogenic variant. If the second variant is shown to act in trans, it would push the pathogenic score past the class 4 threshold for 16 of those variants. We thus decided to create subclasses within class 3 with point thresholds to highlight variants on the verge of class 4 (i.e., class 3LP) and demonstrate that 12% of all variants (20% of class 3) belong to this group. By contrast, variants on the verge of class 2 (class 3 LB) comprised only 5% of all variants (9% of class 3). Overall this implies that currently 47% of ABCC6 missense variants can be considered (likely) pathogenic with reasonable certainty. Nevertheless, classification results are dynamic and should be adjusted according to new evidence that emerges over time; this highlights the importance of follow-up of patients with class 3 variants.
Experimental studies clearly become increasingly important to evaluate class 3 variants. For our classification, we took into account in vitro and in vivo data revealing improper plasma membrane targeting or deficient transport activity of glutathione conjugates (N-ethylmaleimide S-glutathione and leukotriene C4) but such data are only available for a limited number of variants.19,20,21,22,35 An interesting model to study class 3 variants is zebrafish in which the ortholog abcc6a is targeted. However, in morpholino-induced knockdown zebrafish the results from rescue assays should be considered with caution.21,23,35,36,37 For this reason, we decided not to include such results. Alternatively, a promising CRISPR/Cas9-mediated knockout zebrafish model, showing dysregulates osteogenesis, could be used to functionally validate unclear variants.38 Such functional testing is, however, time-consuming and labor intensive. From a clinical point of view, the Sherloc classification system with the adaptations we have done for class 3 variants offers a pragmatic approach to evaluate known and novel ABCC6 variants.
In silico models showing the 3D protein structure can provide supporting evidence for pathogenicity and reveal the effect of variants on the protein’s structural properties. Fülöp et al. were the first to visualize missense variants on the interaction surfaces between both NBD domains (NBD–NBD interface) and NBD and the intracellular TMD (NBD–TMD interface).39 A significant number of class 3 LP, class 4, and class 5 variants (but also two class 1 variants), reside in the NBD–NBD interface, the NBD–TMD interface, or the predicted substrate binding positions. Recently, a detailed model of NBD1 (RCSB PDB: 6BZS) was published, in which various PXE-associated variants were positioned.18 Interestingly, the variants that resulted in misfolding of NBD1 showed little surface accessibility (RSA) and were buried within the NBD structure. We used the 6BZS model to calculate the RSA values of the variants occurring in NBD1 using the SDM webserver (Table S6).28 Fifty and sixty percent of class 4 and class 5 variants respectively residing in NBD1 were buried, as opposed to 32% of class 3 and 0% of class 1 variants in this domain. No class 2 variants were found in NBD1. Even though this is not direct evidence for pathogenicity, the results suggest that 3D models can indeed be useful for variant evaluation.
Comparison with the ACMG-AMP guidelines revealed a similar class distribution and a high—although not complete—concordance with Sherloc, which can be further improved by developing ABCC6-specific criteria within the ACMG-AMP framework. Even though Sherloc is based on the ACMG-AMP guidelines, it has implemented three useful concepts to evaluate variant data. First, it introduces the use of multiple fixed AF thresholds to capture the variable likelihood of pathogenicity, as opposed to disease-specific AF threshold, which requires knowledge about disease prevalence, penetrance, and genetic architecture.40 Second, the delineated criteria are grouped as clinical or functional evidence types, allowing to determine which data are more persuasive in case a conflict arises between clinical and functional observations. Third, criteria are assigned with a preset number of benign or pathogenic points that reflect the weight of the evidence type in the evaluation process.
Comparison of the Sherloc-derived classification results with the corresponding ClinVar variant interpretations revealed (1) a high discordance rate of 64% (51% when confidence conflicts are not considered) and (2) an overestimation of pathogenicity of class 3 and class 4 variants by ClinVar. It is important to note that less than half of variant reports in ClinVar (103/237) were supported by evidence (provided by a single submitter in 84 reports) that mostly lack detailed information about phenotype and zygosity, or were based on publications that were published before large population control databases became available. As a result, the reported clinical significance of certain variants in ClinVar may be outdated and need reviewing. A study evaluating performance of the same variant classification method between different laboratories found that reassessment of classification criteria and data sharing improves the discordance rate.12 Another important database that is widely consulted by geneticists is the LOVD. Probably the most important field of the LOVD database is the classification field. LOVD has two types of classifications: a functional classification indicating the effect of the variant on the function of the gene or protein, and a clinical classification indicating the effect of the variant on the individual’s health. Although each variant in LOVD is assigned with an effect on the function of the protein, we could usually not ascertain the experimental evidence on which this classification is based. Only few records (39 of 181 included in our analysis) have a clinical classification result. Comparing these classification results with those obtained with Sherloc showed 56% discordance (38% when confidence conflicts are not considered), although it should be noted that only the results of 14% (39/286) of the total number variants included in this analysis were eligible for comparison.
In conclusion, our evaluation of ABCC6 missense variants emphasizes the importance of a systematic and critical evaluation process while taking into account the variant’s genomic context. The high number of class 3 variants stresses the importance of segregation analysis in the family and confirms the need for detailed evidence sharing in variant databases and for functional testing to prove or refute their causality before returning them to patients.
References
Vanakker OM, et al. Novel clinico-molecular insights in pseudoxanthoma elasticum provide an efficient molecular screening method and a comprehensive diagnostic flowchart. Hum Mutat. 2008;29:205.
Pfendner EG, et al. Mutation detection in the ABCC6 gene and genotype-phenotype analysis in a large international case series affected by pseudoxanthoma elasticum. J Med Genet. 2007;44:621–628.
Bergen AAB, et al. Mutations in ABCC6 cause pseudoxanthoma elasticum. Nat Genet. 2000;25:228–231.
Le Saux O, et al. Mutations in a gene encoding an ABC transporter cause pseudoxanthoma elasticum. Nat Genet. 2000;25:223–227.
Ringpfeil F, Lebwohl MG, Christiano AM, Uitto J. Pseudoxanthoma elasticum: mutations in the MRP6 gene encoding a transmembrane ATP-binding cassette (ABC) transporter. Proc Natl Acad Sci U S A. 2000;97:6001–6006.
Scheffer GL, Hu X, Pijnenborg ACLM, Wijnholds J, Bergen AAB, Scheper RJ. MRP6 (ABCC6) detection in normal human tissues and tumors. Lab Invest. 2002;82:515–518.
Jansen RS, et al. ABCC6 prevents ectopic mineralization seen in pseudoxanthoma elasticum by inducing cellular nucleotide release. Proc Natl Acad Sci U S A. 2013;110:20206–20211.
Pulkkinen L, Nakano A, Ringpfeil F, Uitto J. Identification of ABCC6 pseudogenes on human chromosome 16p: implications for mutation detection in pseudoxanthoma elasticum. Hum Genet. 2001;109:356–365.
Miksch S, et al. Molecular genetics of pseudoxanthoma elasticum: type and frequency of mutations in ABCC6. Hum Mutat. 2005;26:235–248.
Hoskinson DC, Dubuc AM, Mason-Suares H. The current state of clinical interpretation of sequence variants. Curr Opin Genet Dev. 2017;42:33–39.
Richards S, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–424.
Amendola LM, et al. Erratum: Performance of ACMG-AMP variant-interpretation guidelines among nine laboratories in the Clinical Sequencing Exploratory Research Consortium. Am J Hum Genet. 2016 ;99:247.
Nykamp K, et al. Sherloc: a comprehensive refinement of the ACMG-AMP variant classification criteria. Genet Med. 2017;22:240.
Kobayashi Y, Yang S, Nykamp K, Garcia J, Lincoln SE, Topper SE. Pathogenic variant burden in the ExAC database: an empirical approach to evaluating population data for clinical variant interpretation. Genome Med. 2017;9:13.
Moody JE, Millen L, Binns D, Hunt JF, Thomas PJ. Cooperative, ATP-dependent association of the nucleotide binding cassettes during the catalytic cycle of ATP-binding cassette transporters. J Biol Chem. 2002;277:21111–21114.
Xue P, Crum CM, Thibodeau PH. Regulation of ABCC6 trafficking and stability by a conserved C-terminal PDZ-like sequence. PLoS One. 2014;9:e97360.
Miglionico R, et al. New insights into the roles of the N-terminal region of the ABCC6 transporter. J Bioenerg Biomembr. 2016;48:335.
Ran Y, Zheng A, Thibodeau PH. Structural analysis reveals pathomechanisms associated with pseudoxanthoma elasticum-causing mutations in the ABCC6 transporter. J Biol Chem. 2018;293:15855–15866.
Iliás A, et al. Loss of ATP-dependent transport activity in pseudoxanthoma elasticum-associated mutants of human ABCC6 (MRP6). J Biol Chem. 2002;277:16860–16867.
Le Saux O, et al. Expression and in vivo rescue of human ABCC6 disease-causing mutants in mouse liver. PLoS One. 2011;6:e24738.
Jin L, et al. Genetic heterogeneity of pseudoxanthoma elasticum: the Chinese signature profile of ABCC6 and ENPP1 mutations. J Invest Dermatol. 2015;135:2338.
Ran Y, Thibodeau PH. Stabilization of nucleotide binding domain dimers rescues ABCC6 mutants associated with pseudoxanthoma elasticum. J Biol Chem. 2017;292:1559–1572.
Kok FO, et al. Reverse genetic screening reveals poor correlation between morpholino-induced and mutant phenotypes in zebrafish. Dev Cell. 2015;32:97–108.
Mathe E, Olivier M, Kato S, Ishioka C, Hainaut P, Tavtigian SV. Computational approaches for predicting the biological effect of p53 missense mutations: a comparison of three sequence analysis based methods. Nucleic Acids Res. 2006;34:1317–1325.
Reva B, Antipin Y, Sander C. Determinants of protein function revealed by combinatorial entropy optimization. Genome Biol. 2007;8:R232.
Adzhubei I, Jordan DM, Sunyaev SR. Predicting functional effect of human missense mutations using PolyPhen-2. Curr Protoc Hum Genet. 2013;Unit 7:20.
Schwarz JM, Cooper DN, Schuelke M, Seelow D. Mutationtaster2: mutation prediction for the deep-sequencing age. Nat Methods. 2014;11:361–362.
Pandurangan AP, Ochoa-Montaño B, Ascher DB, Blundell TL. SDM: a server for predicting effects of mutations on protein stability. Nucleic Acids Res. 2017;45:W229–W235.
Ramsay M, et al. Spectrum of genetic variation at the ABCC6 locus in South Africans: pseudoxanthoma elasticum patients and healthy individuals. J Dermatol Sci. 2009;54:198–204.
Chassaing N, et al. Novel ABCC6 mutations in pseudoxanthoma elasticum. J Invest Dermatol. 2004;122:608–613.
Ringpfeil F, et al. Pseudoxanthoma elasticum is a recessive disease characterized by compound heterozygosity. J Invest Dermatol. 2006;126:782–786.
Nitschke Y, et al. Generalized arterial calcification of infancy and pseudoxanthoma elasticum can be caused by mutations in either ENPP1 or ABCC6. Am J Hum Genet. 2012;90:25–39.
Boraldi F, Lofaro FD, Costa S, Moscarelli P, Quaglino D. Rare co-occurrence of beta-thalassemia and pseudoxanthoma elasticum: novel biomolecular findings. Front Med (Lausanne). 2019;6:322.
Issa PC, Tysoe C, Caswell R. Late-onset pseudoxanthoma elasticum associated with a hypomorphic ABCC6 variant. Am J Ophthalmol. 20 May 2020; https://doi.org/10.1016/j.ajo.2020.05.009 [Epub ahead of print].
Pomozi V, et al. Analysis of pseudoxanthoma elasticum-causing missense mutants of ABCC6 in vivo; pharmacological correction of the mislocalized proteins. J Invest Dermatol. 2014;134:946–953.
Robu ME, et al. p53 activation by knockdown technologies. PLoS Genet. 2007;3:787–801.
Li Q, et al. The abcc6a gene expression is required for normal zebrafish development. J Invest Dermatol. 2010;130:2561–2568.
Van Gils M, Willaert A, De Vilder EYG, Coucke PJ, Vanakker OM. Generation and validation of a complete knockout model of abcc6a in zebrafish. J Invest Dermatol. 2018;138:2333–2342.
Fülöp K, Barna L, Symmons O, Závodszky P, Váradi A. Clustering of disease-causing mutations on the domain-domain interfaces of ABCC6. Biochem Biophys Res Commun. 2009;379:706–709.
Whiffin N, et al. Using high-resolution variant frequencies to empower clinical genome interpretation. Genet Med. 2017;19:1151–1158.
Acknowledgements
This study was supported by a Methusalem grant (BOFMET2015000401) from Ghent University. O.M.V. is a Senior Clinical Investigator of the research Foundation–Flanders (Belgium).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosure
The authors declare no conflicts of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Verschuere, S., Navassiolava, N., Martin, L. et al. Reassessment of causality of ABCC6 missense variants associated with pseudoxanthoma elasticum based on Sherloc. Genet Med 23, 131–139 (2021). https://doi.org/10.1038/s41436-020-00945-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-020-00945-6
Keywords
This article is cited by
-
A new enzymatic assay to quantify inorganic pyrophosphate in plasma
Analytical and Bioanalytical Chemistry (2023)