Abstract
Purpose
To provide a validated method to confidently identify exon-containing copy-number variants (CNVs), with a low false discovery rate (FDR), in targeted sequencing data from a clinical laboratory with particular focus on single-exon CNVs.
Methods
DNA sequence coverage data are normalized within each sample and subsequently exonic CNVs are identified in a batch of samples, when the target log2 ratio of the sample to the batch median exceeds defined thresholds. The quality of exonic CNV calls is assessed by C-scores (Z-like scores) using thresholds derived from gold standard samples and simulation studies. We integrate an ExonQC threshold to lower FDR and compare performance with alternate software (VisCap).
Results
Thirteen CNVs were used as a truth set to validate Atlas-CNV and compared with VisCap. We demonstrated FDR reduction in validation, simulation, and 10,926 eMERGESeq samples without sensitivity loss. Sixty-four multiexon and 29 single-exon CNVs with high C-scores were assessed by Multiplex Ligation-dependent Probe Amplification (MLPA).
Conclusion
Atlas-CNV is validated as a method to identify exonic CNVs in targeted sequencing data generated in the clinical laboratory. The ExonQC and C-score assignment can reduce FDR (identification of targets with high variance) and improve calling accuracy of single-exon CNVs respectively. We propose guidelines and criteria to identify high confidence single-exon CNVs.
Similar content being viewed by others
INTRODUCTION
Copy-number variation (CNV) is an important feature of the human genome and can confer disease susceptibility.1,2,3,4 The ability to detect CNVs accurately is critical for both genetic diagnostics and to advance understanding their impact on gene function. Next-generation sequencing (NGS) based targeted gene panels are commonly used in clinical genetic testing and various methods5,6,7,8,9,10,11,12,13 have been developed to identify exonic CNVs in gene panel sequence data. Gene panels afford a qualitatively different opportunity to assess small CNVs due to their typically deeper sequence coverage when compared with genome or exome sequencing (ES). Ideally, the methods would detect single-exon CNVs, but this is challenging because a single exon represents one data point, which must exhibit minimal noise and maximal signal compared with multiexonic CNVs, which have corroborating data points. Detecting and reporting CNVs in the clinical context is another challenge as false positives (FPs) must be minimized, while all true positives (TPs) are all identified. Achieving the high accuracy required for clinical applications invariably demands validation on alternative platforms, which bears its own set of technical challenges.
Existing CNV tools that have been validated for the clinical setting include VisCap,9 CoNVaDING,10 DeCON,11 ExomeDepth,12 and others.5,8,13,14 CoNVaDING and ExomeDepth are methods that are successful at detecting single-exon CNVs by evaluating individual exon suitability for variant detection and selecting highly correlated samples as reference controls. In these methods, however, each single-exon CNV is treated equally and there is no mechanism to evaluate the confidence of the call. VisCap has relied on human visual scoring to reduce FPs and evaluate small CNVs.
The Electronic Medical Records and Genomics (eMERGE) Network provides an opportunity to develop tools addressing these challenges. Briefly, eMERGE Phase III is the continuation of a National Institutes of Health (NIH) program that aims to incorporate genomic information into medical records (25,000 participants) by identifying rare genetic variants using eMERGESeq (a targeted gene panel designed by multiple eMERGE sites to identify impactful variants; https://emerge.mc.vanderbilt.edu/the-emergeseq-platform/ and https://doi.org/10.1101/457523), and their effects in 109 clinically relevant genes, including the American College of Medical Genetics and Genomics (ACMG)15 56 medically actionable genes. Here, we present Atlas-CNV, a method to identify CNVs even at the single-exon level based on the normalized coverage among samples but constrained to the same capture experiment. We incorporate standard deviation (StDev) thresholds to remove low quality exons and samples for controlling FPs. Atlas-CNV produces graphical gene and exon bar plots to allow for visualization by clinicians and diagnosticians. The leveraging of the C-scoring exons to prioritize high quality single-exon CNVs enables significant reduction of FPs and obviates much of the need for costly expert-based reviewing. Atlas-CNV is designed to analyze gene-targeted data from a set of samples without the need for additional sequencing controls. Instead, a “midpool” experiment is defined from a subset of the multiplexed samples within a single probe hybridization capture experiment undergoing identical experimental conditions. The typical eMERGESeq midpool is optimized to 45–48 samples. Overall, Atlas-CNV is a fast (<2 minute/midpool/CPU-core) and versatile caller that integrates easily into clinical pipelines.
We benchmarked and validated Atlas-CNV on known CNVs previously identified by exome sequencing (ES) and chip array, demonstrated the C-scoring feature through simulation on a subset of eMERGESeq samples (sequenced at the Baylor College of Medicine Human Genome Sequencing Center), and assessed its performance by verifying 64 and 29 multi- and single-exon CNVs respectively through Multiplex Ligation-dependent Probe Amplification (MLPA). Currently, while eMERGE reports CNVs spanning two or more contiguous exons, our findings support the feasibility of including single-exon CNVs into medical records with the potential for increased diagnostic yield.16
MATERIALS AND METHODS
Atlas-CNV (v.0) is available at http://github.com/theodorc/atlas-cnv (Perl 5.12.2, R 3.1.1). Three inputs are required: (1) Genome Analysis Toolkit (GATK) DoC interval summary files, (2), a panel design containing target exons, and (3) a sample file with gender and/or midpool groupings.
Clinical sequencing
Our clinical pipeline processes 45–48 samples per midpool experiment. Briefly, sample DNA is isolated, sheared, ligated to barcode adapters for multiplexing, then incubated with capture probes, and sequenced on Illumina HiSeq 2500 instruments with two midpools loaded on a single flow-cell lane. Paired-end reads are aligned to the hg19 reference using bwa-0.6.2 (ref. 17) with GATK-2.5.2 (ref. 18) for realignment, recalibration, and depth of coverage calculations (DoC).
RPKM normalization and sample quality
The read depth (RD) data is normalized at the individual sample level. GATK DoC is converted to average RD per target, and normalized as a fraction of the sample coverage with RPKM19 (reads per thousand bases per million reads sequenced) as illustrated in Fig. 1a. Essentially, this step converts the average RD per target to the equivalent number of reads (100 bp/read) and reports the proportion to the total number of mapped reads in the sample per million. At each exon, the median sample is selected as the reference after removing the 5% outliers (Z-score at 1.96). Then, log2 scores of the sample/median ratio are computed accordingly for all exons on all samples as shown in Fig. 1b. To assess sample quality, we define a SampleQC (Supplementary Figure S1) with two components: (1) a 0.2 threshold on the StDev of log2 scores for the sample, and (2) a one-way analysis of variance (ANOVA) F-test at 5% significance applied on the mean RPKM coverage between midpool samples. If either component is not satisfied, the sample is labeled “Fail.” The former is determined by computing the StDev of log2 scores in one theoretical diploid sample with noise randomly introduced (R function runif to generate random deviates) into the sample/median (S/M) ratio in 5% increments (Supplementary Figure S3). For a panel of 2000 targets, we show that StDev = 0.2 is equivalent to 25% overall sample variability, which means given an exon deletion, it is possible that noise could account for up to 75% (0.25/0.33) of the overall signal to reach the calling threshold for deletions (0.66 or log2 = −0.6), and vice versa for duplications (1.32 or log2 = 0.4). While this can vary and lead to FPs, bona fide signal may be present. Thus, we set 0.2 as the default upper bound.
Atlas-CNV method for calling exonic copy-number variants (CNVs) in a midpool of samples. (a) RPKM normalization is first performed on each sample with each exon assigned a single coverage value expressed as a proportion of reads per unit kilobase per sample reads in millions. (b) The median sample at each exon is selected as the reference after the 5% outliers are excluded. Log2 scores of sample/median are computed for each sample at that exon and the StDev of these scores is called the EStDev for that exon. Low quality exons and samples are filtered and flagged. (c) CNVs are called with thresholds at the exon level with visual bar plots at the sample gene level and at the batch exon level. ANOVA analysis of variance, DoC depth of coverage.
Exon quality and calling exonic CNVs
As illustrated in Fig. 1b, we also use log2 scores to assess exon data quality because having even-exon normalized coverage is essential to reduce FPs. Thus, we remove outliers where coverage exceeds a threshold based upon the StDev of log2 scores at a given exon (EStDev), which is calculated on midpool samples. We evaluate the overall distribution of all EStDevs and establish a data derived threshold termed ExonQC, which we define as the EStDev at 99.9% of the EStDev distribution (z = 3.921). An exon is labeled “Fail” if its EStDev exceeds ExonQC. To account for EStDev overinflation due to true exonic CNVs, we first exclude the 5% outliers (approximately two samples), to produce a tighter EStDev value that also allows for samples with identical CNVs (kinship). Numerous failed exons may indicate aberrant experimental steps, or systematic biases (Supplementary Figure S2). Typical exon fail rate per midpool is 0.5%.
To call CNVs, we apply two thresholds on the log2 scores (Fig. 1c): (1) a user-configurable hard limit of −0.6 and 0.4 (default) for losses and gains, and (2) a soft threshold derived from the data distribution using a Z-score cutoff at 99% (z = 2.576), intended as a boundary to threshold calls at the distribution tail. Any log2 scores exceeding both thresholds are called CNVs. Autosomes and sex chromosomes are analyzed separately with a sample file defining midpool and/or gender subgroupings.
Visualization and confidence score
For visualization, Atlas-CNV produces a sample gene plot with bars representing exons, and also sectioned exon bar plot(s) to display the context of the exonic CNV with all midpool samples. In the latter, the median sample is designated as blue, the test as red, and other samples in gray. Fig. 1c illustrates two deleted contiguous exons in CFTR.
We define a confidence score (C-score) assigned to each CNV exon (positive and negative scores for duplications and deletions) and propose three categories: “duplication or deletion,” “likely duplication or deletion,” and “uncertain duplication or deletion” with ranges to denote copy number (Supplementary Material Table 1–3). C-scores are somewhat analogous to Z-scores but rescale each exon to unit variance by dividing the individual log2 score by the EStDev, with the assumption that the mean log2 score for the given midpool is zero (diploid). C-scoring standardizes exons on the same comparative scale. For CNVs spanning multiple exons on a gene, individual C-scores are averaged on the gene plot report.
Performance measures and MLPA
To assess performance, we used CNV samples previously identified by ES, and the Illumina HumanExome-12v array as our gold standard (GS). We define sensitivity as the proportion of GS exons (true positives, TP) over the sum of TP and GS exons not called (false negatives, FN); specificity as the proportion of exons other than GS exons (true negatives, TN) over the sum TN and called exons that are not GS (false positives, FPs); precision as the proportion of TP over the sum of all positive calls (TP + FP); and false discovery rate (FDR) as the proportion of FP over all positive calls. We also define the reproducibility of a tool as the pairwise comparison of two identical runs (i, j) expressed as the proportion of common exons to the union of i and j exons. Finally, an estimated FDR (eFDR) is computed per sample using the Robust FDR procedure,20 which is based on p values obtained from C-scores.
MLPA (MRC Holland, www.mlpa.com) was used to confirm CNVs for available genes. Samples were processed according to vendor protocols using three controls: NA12878, NA12891, and NA12892.
RESULTS
Performance assessment and comparison
To assess Atlas-CNV performance and comparisons with VisCap, we selected 13 clinical samples with a heterozygous gene deletion previously identified by ES and the Illumina HumanExome-12v array as our gold standard (Table 1). Generally, the samples were sequenced in triplicates and divided into technical and biological replicates. The technical replicates used aliquots from the same midpool experiment for sequencing while the biological replicates (2 samples: 1–100155, 12–100189) are completely distinct experiments. To compute performance measures, we averaged the replicates of each sample’s comparison with the gold standard and report an overall mean of these 13 samples as bars in Fig. 2. We observed high sensitivity and specificity (>99%) across technical and biological replicates, but VisCap has lower precision (80%) and higher FDR (20%) than Atlas-CNV (95%, 4%), which can be attributed to filtering FPs by ExonQC. The legitimacy of filtered FPs was confirmed by their absence in ES data. We also observed that the average EStDev is nearly twice as large in FP calls (EStDev = 0.14 on 45 calls) compared with TP calls (EStDev = 0.079 on 1137 calls), and the average SampleQC StDev is 0.2 at z = 3 (99.9%).
Performance measures comparing Atlas-CNV and VisCap on 13 gold standard samples. Technical replicates were sequenced in triplicates for all samples. Biological replicates were performed on two samples (1 and 12). The performance measure for each sample is first obtained by averaging sample replicates in comparison with the gold standard (GS). Then, an overall mean of the 13 samples is plotted in each bar. Reproducibility is the pairwise comparison of two identical runs i, j expressed as the proportion of common exons to the union of exons in i and j. Sensitivity and specificity are the respective proportions of GS exons (true positives, TPs) or exons other than GS (true negatives, TNs) over the respective sum of these with FNs (false negatives; failed to call) or FPs (false positives; failed to reject). Precision and false discovery rate (FDR) are the respective proportions of TPs or FPs over all positives.
For reproducibility, pairwise comparisons of replicates were first averaged per sample and then the 13 means were averaged and reported. For the two samples with additional distinct experiments, 3 representative pairwise comparisons were chosen beforehand as the biological replicates (first sample of the technical replicate set), while the remaining 12 comparisons were treated as technical replicates. We report a higher reproducibility in Atlas-CNV (92%) compared with VisCap (80%) indicating ExonQC may be filtering unrepeated calls in replicate runs.
We estimated an FDR based on p values from C-scores without prior knowledge of the truth set to confirm the Atlas-CNV FDR of 4%. Using a robust FDR routine under the assumption of a one-sided test,20 we computed the estimated FDR for the 13 gold standard samples in the range from 0.12% to 14% for technical replicates, and 0% to 15% for biological replicates at p value cutoffs between 0.009 and 0.01. Although the estimated range is broad, our reported FDR of 4% is within this range of 0–15% and suggests the utility of the procedure on prospective samples without orthogonal confirmation.
Analysis of eMERGE samples
We analyzed 10,926 eMERGESeq samples from 233 midpools (excluding PMS2 [ref. 21] due to highly homologous sequences and 4 midpools with >10% samples failed) with an average of 47 samples per midpool and average coverage of 252× per sample. First, we evaluated the ability of Atlas-CNV and VisCap to call CNVs with at least two contiguous exons. Both detected multiexonic CNVs in 2% of samples with 89% agreement (CNVs identified by both/all CNVs identified) at the sample and gene level (autosomes only). Atlas-CNV and VisCap identified 232 and 184 CNVs respectively with fewer samples failing SampleQC in Atlas-CNV than VisCap (90 vs. 208). Discordant calls (70) were largely made on samples failed by the other tool.
Second, we focused on single-exon detection and initially observed significant discrepancies in the number of these calls. First, Atlas-CNV called nearly five times fewer single-exon CNVs than VisCap (2240 vs. 10,417; dels = 861:5213, dups = 1379:5204); and second, CoNVaDING, a tool developed for single-exon detection, called even fewer than Atlas-CNV (685; 514 dels, 171 dups). Thus, to reduce the complexities of these comparisons and obtain an estimate of the FDR, we counted the number of single-exon CNVs present in >1% of samples, which we assume would likely be artifacts or common CNVs. We report 85% (8818/10,417) of VisCap calls, which are observed in only 10 exons, exist in >1% of the samples; 5% (114/2240) for Atlas-CNV (1 exon); and 0% (0/685) for CoNVaDING. This highlights the importance of having a mechanism to automatically filter low quality exons to reduce FPs (present in Atlas-CNV and CoNVaDING but not VisCap). In contrast, if we focus on calls in <1% of the samples (Atlas-CNV: 2126, VisCap: 1599, CoNVaDING: 685), which are more likely to be TPs, our results show 46% (or 741/1599) of VisCap and 42% (or 286/685) of CoNVaDING calls are in common with Atlas-CNV. While the concordance is low, closer examination revealed that missed calls were labeled as multiexon CNVs in Atlas-CNV or failed to meet the Atlas-CNV passing criteria for either an exon or sample (ExonQC or SampleQC). For example, 58% (or 395/685) of CoNVaDING calls either failed the Atlas-CNV ExonQC (6) or SampleQC (324), were labeled as multiexon CNVs (47), or flagged as FPs (18), leaving only 4 real missed calls. A similar outcome was observed in the VisCap comparison, but only 15% of calls failed an Atlas-CNV quality control (QC) (225) or were labeled as multiexon CNVs (8), leaving 39% of calls as truly missed (or 625/1599). Further examination of the common calls also showed CoNVaDING with a higher mean C-score (7.11 vs. 4.9) and lower mean EStDev (0.11 vs. 0.14) than VisCap (see summary in supplementary Table S4).
Figure 3 summarizes the overall Atlas-CNV analysis. It includes results for PMS2 and X chromosome genes, the former of which may require further analyses.21 In total, 345 samples were identified (172 losses, 173 gains) for multiexonic CNVs, which represents an overall frequency of 3.2% (1.57% losses, 1.58% gains), or 0.03 CNVs/sample. Adding high confidence single-exon CNVs (abs[C-score] ≥8, 109 losses, 22 gains) increases the frequency to 4.36% (2.57% losses, 1.78% gains). As a relative comparison, the CNV frequency in ExAC22 is 1.43%. Interestingly, we detected CNVs in 41 of the 58 ACMG genes (excluding ATP7B) with the highest occurrence in OTC (24) and GLA (24) while CNVs were observed in 38 of 51 non-ACMG eMERGE genes with KCNE1 (18) and SLC25A40 (11) as the top genes. Finally, 13 exons with high frequency of failing ExonQC were identified in >25% of eMERGESeq midpools (Supplementary Table S5).
Distribution of copy-number variants (CNVs) in 10,926 eMERGE samples: deletions (a) and duplications (b). Values in bars represent the sample count broken down by CNV type (single-exon, exactly two exons, three or more, or full gene) with total counts and overall frequency listed in the legend [square brackets]. American College of Medical Genetics and Genomics (ACMG) genes are grouped alphabetically in the left dotted box and eMERGE-specific genes in the right. Gene names labeled in blue were identified in both gains and losses.
Evaluation of C-score threshold for CNV calling
Across all samples, a total of 2475 exons (on average 11 exons/midpool) were removed by exon filtering with an average EStDev = 0.36 compared with the 2240 passing exons with a much lower average EStDev = 0.13. We report an average eFDR of 3.14% across all samples, which coincides with the FDR (4%) reported in the performance assessment study (Fig. 2). However, even with low EStDev and FDR, there is still a tenfold enrichment of single over multiexon CNVs (2240/232). Therefore, we set out to determine whether C-scores could be used as a secondary assessment of confidence of single-exon CNVs primarily on the basis that standalone log2 scores do not account for the exon variability whereas C-scores are standardized by this exact variance. First, we computed the expected EStDev in a simulation of log2 scores by adding 5% increments of variability into the S/M ratio using the R function runif under a uniform distribution (Supplementary Figure S3). We determined EStDev at 0.08, 0.13, and 0.16 correspond respectively to 10%, 15%, and 20% variability. Therefore, for both eMERGESeq (mean EStDev = 0.92) and gold standard assessed samples (mean EStDev = 0.079), the multiexonic CNVs fall in the 10% noise range. However, eMERGE single-exon CNVs (2240) with nearly twice the mean EStDev (0.14), fall over the >15% noise range (Supplementary Figure S4). Therefore, to control single-exon FPs, both the log2 scores and EStDev should be utilized.
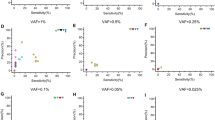
We performed a simulation of single-exon deletions to obtain C-score thresholds for optimal sensitivity. Briefly, 100 random samples were chosen from 59% (6413/10,926) of eMERGESeq samples with no previous single-exon deletion. Each sample was randomly assigned a single-exon deletion by artificially downsizing the read depth by 5% increments from 30% to 50%. The mean C-score and sensitivity were calculated at each coverage increment and plotted on a curve (Supplementary Figure S5). We iterated this analysis using three calling thresholds of −0.4, −0.5, and −0.6 (default), and for all instances in the simulation study, we noted the following: (1) EStDev range 0.08–0.09, (2) eFDR range 3–3.2%, (3) specificity >99%, and most importantly, (4) sensitivity >90% on C-scores >10 where read depth is reduced by >40%. We conclude that these observed ranges for C-score and EStDev are conservative for calling confident single-exon CNVs. When we applied a C-score ≥8 and EStDev ≤ 0.1 criteria on eMERGE samples, we identified 79 candidate single-exon CNVs (candidates for MLPA validation), which represents a 28-fold decrease from the total of 2240 identified single-exon CNVs. Notably, VisCap and CoNVaDING called 62 and 69 respectively out of these 79.
MLPA confirmation of CNVs
Sixty-four multiexon CNVs (34 losses, 30 gains; mean C-score = 9.4, EStDev = 0.087) called by Atlas-CNV were selected for MLPA confirmation (Supplementary Table S6). Although MLPA has its own technical limitations, using it here as the truth set, we confirmed 55 CNVs with Atlas-CNV having higher sensitivity (88.8%) and lower FDR (25.0%) compared with VisCap (86.8% and 33.6% respectively). Notably, two confirmed CNVs (Vanderbilt-23 and Columbia-29) were missed by VisCap. The 9 unconfirmed CNVs were compared with the 55 confirmed CNVs and found to have significantly (1) deflated C-scores (5.4, P = 8.7e-05), (2) elevated EStDev (0.13, P = 2.2e-16), and (3) high CNV genes per sample (>6, P = 3.3e-16). A three-pronged criteria (C-score >8, EStDev < 0.1, CNV genes <3) could easily remove these 9 CNVs and cut the FDR in half to 12.7%. The examination of the actual remaining false positives and negatives indicate missed CNV exons were due to borderline signals on either the gene panel or MLPA platform. Finally, in a further separate analysis, three (28-Northwestern, 30-Columbia, 36-Mayo) samples of the confirmed CNVs (MYH7, LDLR) were also validated using a second gene panel in routine use. Overall, these multiexon CNV data demonstrate Atlas-CNV performed best with 86% (55/64) confirmed samples and 14% (9/64) failed.
MLPA was also used to evaluate the single-exon CNVs detected by Atlas-CNV (Table 2). Initially, 29 single-exon CNVs were selected for testing from the 79 candidates described above, consisting of 23 high confidence CNVs (22 losses, 1 gain; mean C-score = 12.3, EStDev = 0.081) and 6 borderline confidence CNVs (C-score <8 or EStDev > 0.1). However, exact probe reagents for the exon of interest were available for only 14 CNVs (samples 1–14), from which MLPA assays confirmed 10 CNVs (71.4% or 10/14) and were negative for the other 4 (28.6% or 4/14) although 3 of the failed cases had >3 CNV genes per sample. Thus, our overall validation confirmed 90.9% (10/11) of single-exon CNVs (samples 2–11) with C-score >8. One additional single-exon CNV was confirmed from the 15 CNVs (samples 15–29) without exon-specific probes when MLPA assays were carried out using nearby or flanking probes. This results in 14 inconclusive cases for which future investigation is needed using alternate validation methods.
DISCUSSION
Prior studies have demonstrated successful CNV detection in DNA sequence data generated from gene panels, but the burden of analyzing single-exon CNVs is a challenge because of high false-positive rates.8,9 Recently developed tools such as CoNVaDING10 and modified versions of ExomeDepth11,12 have been designed to identify single-exon CNVs, but lack quality metrics that enable differentiation of different levels of confidence. As a result, clinical laboratories may often ignore single-exon CNVs. Here, we introduced Atlas-CNV, a fast and accurate CNV calling method based on read depth that reports confidence scores for each CNV event that are used to reduce FPs. Previously established methods often failed to account for target variability arising from extremes of DNA sequence coverage, while Atlas-CNV implements multiple strategies to cope with these variabilities. Thus, Atlas-CNV’s advantage over other similar methods is the ability to screen or prioritize for high confidence single-exon CNVs. Atlas-CNV also overcomes the limitation of using log2 scores as the sole criteria to detect calls. Furthermore, while existing methods9,10 have also shown high sensitivity and specificity for detecting single exons in targeted sequence data, our method does not require additional control samples, prior knowledge of model parameters, or empirical assessments of specific panel designs.11,13 These require additional calibration, effort and cost.
Because Atlas-CNV and VisCap are integral components of the eMERGESeq program, we compared the two and our results showed 90% agreement for large or multiexonic variants. However, given VisCap’s lack of exon filtering, it is difficult to interpret the 39% (or 625/1599) of calls missed by Atlas-CNV for single-exon variants. Furthermore, visual inspection (VisCap) would be a cumbersome task. Therefore, we evaluated the single-exon CNVs in comparison with CoNVaDING calls and conclude Atlas-CNV only missed 4 calls (0.6% or 4/685). We report favorable mean Atlas-CNV C-scores (7.11) and EStDev (0.11) for the 286 common calls even though 57% (164/286) fail the abs(C-score) >8, EStDev ≤ 0.1 criteria. Interestingly, CoNVaDING missed 10 calls that were called by Atlas-CNV (met the abs[C-score] >8, EStDev ≤ 0.1). We also compared Atlas-CNV with exome CNV caller CoNIFER,23 which utilizes similar approaches, but includes a broad secondary normalization (SVD), which appears to overcorrect and smooth individual target signal when applied to eMERGESeq data. While advantageous for capture assays with high numbers of targets (exome), it loses detection power at the discrete single-exon level in smaller target panels, due in part to the lack of mechanisms to assess and score single exons. Thus, taken together these results highlight Atlas-CNV's performance and ability to prioritize CNV calls to control the FDR and reduce expensive and labor-intensive secondary tests.
Our results also show that optimal C-scores >10 can produce >90% sensitivity for single-exon deletions. We therefore propose the following usage guidelines to assist in selecting high confidence CNVs: (1) abs(C-score) >8, (2) EStDev < 0.1, (3) exon size >50 bp, (4) mappability >0.8, (5) CNV genes <3 per sample. The first two parameters were derived from multiexonic CNVs, the third is specific to small targets with potential coverage bias, and the last two are considerations to be vetted accordingly because samples with many CNVs could indicate either high FPs or the need for further study. Applying a criterion of only C-score ≥8 and EStDev ≤ 0.1, we prioritized 79 significant candidates from the initial 2240 single-exon calls. Of 11 cases with definitive MLPA assays we confirmed 10 with examples in CFTR, MLH1, and other genes like PKP2 and DSP in which known pathogenic variants can increase the risk of arrhythmogenic right ventricular cardiomyopathy, a leading cause of sudden heart failure in young people. Additional frameshift indels were also discovered in the DSP samples, raising the important role single-exon deletions play in compound heterozygotes for autosomal recessive disorders.
This work advances the confident identification of exonic CNVs, especially in clinical programs deployed at scale. Applying the Atlas-CNV approach to call single-exon variants should improve current variant calling standards and identify noncallable exons requiring alternate methods. We also expect that our knowledge of disease genes will increase as new single-exon CNVs are uncovered, catalogued in public databases, and reliably reported. Clinical sites receiving such reports will also benefit patients, who will obtain better diagnosis and treatment. In conclusion, we have demonstrated Atlas-CNV as a validated approach for clinical laboratories to screen the full spectrum of exonic CNVs in gene panels, with particular focus on single-exon CNVs.
References
Nowakowska B. Clinical interpretation of copy number variants in the human genome. J Appl Genet. 2017;58:449–457.
Zhang F, Gu W, Hurles ME, Lupski JR. Copy number variation in human health, disease, and evolution. Annu Rev Genomics Hum Genet. 2009;10:451–481.
Conrad DF, Pinto D, Redon R, et al. Origins and functional impact of copy number variation in the human genome. Nature. 2010;464:704–712.
Zarrei M, MacDonald JR, Merico D, Scherer SW. A copy number variation map of the human genome. Nat Rev Genet. 2015;16:172–183.
Nord AS, Lee M, King M-C, Walsh T. Accurate and exact CNV identification from targeted high-throughput sequence data. BMC Genomics. 2011;12:184.
Li J, Lupat R, Amarasinghe KC, et al. CONTRA: copy number analysis for targeted resequencing. Bioinformatics. 2012;28:1307–1313.
Bellos E, Kumar V, Lin C, et al. cnvCapSeq: detecting copy number variation in long-range targeted resequencing data. Nucleic Acids Res. 2014;42:e158.
Feng Y, Chen D, Wang GL, Zhang VW, Wong LJC. Improved molecular diagnosis by the detection of exonic deletions with target gene capture and deep sequencing. Genet Med. 2015;17:99–107.
Pugh TJ, Amr SS, Bowser MJ, et al. VisCap: inference and visualization of germ-line copy-number variants from targeted clinical sequencing data. Genet Med. 2016;18:712–719.
Johansson LF, van Dijk F, de Boer EN, et al. CoNVaDING: single exon variation detection in targeted NGS data. Hum Mutat. 2016;37:457–464.
Fowler A, Mahamdallie S, Ruark E, et al. Accurate clinical detection of exon copy number variants in a targeted NGS panel using DECoN. Wellcome Open Res. 2016;1:20.
Ellingford JM, Campbell C, Barton S, et al. Validation of copy number variation analysis for next-generation sequencing diagnostics. Eur J Hum Genet. 2017;25:719–724.
Kerkhof J, Schenkel LC, Reilly J, et al. Clinical validation of copy number variant detection from targeted next-generation sequencing panels. J Mol Diagn. 2017;19:905–920.
Gambin T, Akdemir ZC, Yuan B, et al. Homozygous and hemizygous CNV detection from exome sequencing data in a Mendelian disease cohort. Nucleic Acids Res. 2016;45:1633–1648.
Kalia SS, Adelman K, Bale SJ, et al. Recommendations for reporting of secondary findings in clinical exome and genome sequencing, 2016 update (ACMG SF v2.0): a policy statement of the American College of Medical Genetics and Genomics. Genet Med. 2017;19:249–255.
Connolly JJ, Glessner JT, Almoguera B, et al. Copy number variation analysis in the context of electronic medical records and large-scale genomics consortium efforts. Front Genet. 2014;5:1–8.
Li H, Durbin R. Fast accurate long-read alignment Burrows-Wheeler transform. Bioinformatics. 2010;26:589–595.
McKenna A, Hanna M, Banks E, et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010;20:1297–1303.
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods. 2008;5:621–628.
Pounds S, Cheng C. Robust estimation of the false discovery rate. Bioinformatics. 2006;22:1979–1987.
Li J, Dai H, Feng Y, et al. A comprehensive strategy for accurate mutation detection of the highly homologous PMS2. J Mol Diagn. 2015;17:545–553.
Ruderfer DM, Hamamsy T, Lek M, et al. Patterns of genic intolerance of rare copy number variation in 59,898 human exomes. Nat Genet. 2016;48:1107–1111.
Krumm N, Sudmant PH, Ko A, et al. Copy number variation detection and genotyping from exome sequence data. Genome Res. 2012;22:1525–1532.
Acknowledgements
The eMERGE Network phase III work was funded through the following grants: U01HG8657 (Kaiser Permanente Washington, formerly Group Health Cooperative/University of Washington, Seattle); U01HG8685 (Brigham and Women’s Hospital); U01HG8672 (Vanderbilt University Medical Center); U01HG8666 (Cincinnati Children’s Hospital Medical Center); U01HG6379 (Mayo Clinic); U01HG8679 (Geisinger Clinic); U01HG8680 (Columbia University Health Sciences); U01HG8684 (Children’s Hospital of Philadelphia); U01HG8673 (Northwestern University); U01HG8701 (Vanderbilt University Medical Center, serving as the Coordinating Center); U01HG8676 (Partners Healthcare/Broad Institute); and U01HG8664 (Baylor College of Medicine).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosure
This work was funded by internal operating funds of the Baylor College of Medicine Human Genome Sequencing Center (HGSC), and by the NIH eMERGE program Phase III: U01HG8657 (Kaiser Permanente Washington/University of Washington); U01HG8685 (Brigham and Women’s Hospital); U01HG8672 (Vanderbilt University Medical Center); U01HG8666 (Cincinnati Children’s Hospital Medical Center); U01HG6379 (Mayo Clinic); U01HG8679 (Geisinger Clinic); U01HG8680 (Columbia University Health Sciences); U01HG8684 (Children’s Hospital of Philadelphia); U01HG8673 (Northwestern University); U01HG8701 (Vanderbilt University Medical Center serving as the Coordinating Center); U01HG8676 (Partners Healthcare/Broad Institute); and U01HG8664 (Baylor College of Medicine). The HGSC is a one of the two Sequencing Centers for the eMERGE III. The Electronic Medical Records and Genomics (eMERGE) Network is a National Human Genome Research Institute (NHGRI)-funded consortium tasked with developing methods and best practices for utilization of the electronic medical record (EMR) as a tool for genomic research. All authors are members of the eMERGE network and declare no conflicts of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. If you remix, transform, or build upon this article or a part thereof, you must distribute your contributions under the same license as the original. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/.
About this article
Cite this article
Chiang, T., Liu, X., Wu, TJ. et al. Atlas-CNV: a validated approach to call single-exon CNVs in the eMERGESeq gene panel. Genet Med 21, 2135–2144 (2019). https://doi.org/10.1038/s41436-019-0475-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-019-0475-4
Keywords
This article is cited by
-
Next-generation sequencing and bioinformatics in rare movement disorders
Nature Reviews Neurology (2024)
-
Prevalence of germline variants in Brazilian pancreatic carcinoma patients
Scientific Reports (2024)
-
Detecting copy number variation in next generation sequencing data from diagnostic gene panels
BMC Medical Genomics (2021)
-
Genetic profiling of primary and secondary tumors from patients with lung adenocarcinoma and bone metastases reveals targeted therapy options
Molecular Medicine (2020)
-
Deficiency of Adenosine Deaminase 2 (DADA2): Hidden Variants, Reduced Penetrance, and Unusual Inheritance
Journal of Clinical Immunology (2020)