Abstract
Purpose
We report for the first time, the use of clinical genome sequencing (GS) in an unbiased pediatric cohort. We describe the clinical validation, patient metrics, ordering patterns, results, reimbursement, and physician retrieval of results for the first consecutive 80 cases.
Methods
Clinical GS was performed for both inpatients and outpatients undergoing etiologic evaluations. Results were reported in the electronic medical record. Evidence of report retrieval by clinicians and whether interpretation was concordant with laboratory report was obtained through retrospective chart review.
Results
Twenty definitive diagnoses were made in 19 patients (24%; n = 80). Except for two partial gene deletions, all diagnostic variants would have been detectable by our exome methods. Surprisingly, there was no documentation of communication of results to the family in the medical record for 17.5% of patients, and in 7.5%, physician and laboratory interpretations were discordant. Average insurance reimbursement was 30.2%, with yield for commercial payers significantly higher, at 54.1%.
Conclusions
The detection rate of GS is equivalent and potentially superior to exome sequencing (ES). Reimbursement rates were variable but overall satisfactory for commercial insurers, and poor for government entities. In addition, we identify opportunities for improvement in the communication of results to families, likely translatable to other tests and other institutions.
Similar content being viewed by others
Introduction
Clinical exome sequencing (ES) has become a widely used tool for the diagnosis of rare genetic diseases, affording coverage of the coding region at a fraction of the cost of genome sequencing (GS). There are advantages and disadvantages to both methods. Exomes offer higher sequencing depth, albeit more biased coverage. Genomes have a greater sequencing and storage cost, but because they do not require enrichment the laboratory processing time is much shorter. Although noncoding regions are available with GS, these are largely not analyzed in the clinical setting. For these reasons, the use of WGS is generally limited to the research setting. In 2015, our center launched clinical GS (cGS) following extensive experience with research-based GS for the molecular diagnosis of pediatric patients with presumed rare Mendelian disorders. We introduced a mixed trio next-generation sequencing (NGS) test, in which GS is performed for the proband and ES is done for both parents, as available. This has the benefit of readily identifying de novo variants, phasing, validation of parentage, and improved variant interpretation. Here, we report for the first time the use of clinical GS in a pediatric hospital, including our process, ordering metrics, diagnostic yield, reimbursement, and clinician interpretation of the report to families.
Materials and methods
Patients and phenotyping
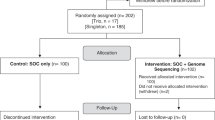
Eighty patients with suspected genetic disorders received cGS between 13 August 2015 and 24 April 2017 (Suppl. Table 1). The majority (57) underwent trio sequencing with both parent samples; 17 were run as parent/child duos, and only 4 as singletons. Two were run as a proband–parent–sibling trio. The average age of this cohort was 6.9 years. Etiologic testing was performed for a range of clinical concerns by a variety of subspecialists. The majority of cGS (47/80 or 59%) was ordered by geneticists, followed by neurologists (22 or 28%) and other subspecialists (Table 1) from both inpatient (21) and outpatient (59) services. Most patients (57/80 or 71%) were also tested by a genome-wide array methodology (oligo-array comparative genomic hybridization [CGH], single-nucleotide polymorphism [SNP] array, or exon array) and no diagnostic findings were reported (Suppl. Table 1.) Approval was obtained from the institutional review board at Children's Mercy Hospital.
GS testing process
Given the high cost of the cGS, insurance preauthorization was initiated for all outpatient orders. Following notice of preauthorization, the parents were consented by a genetic counselor. If testing was denied, financial assistance or other options more favorable to the family’s situation were explored. Inpatients were consented directly by a genetic counselor with no preauthorization required. GS was performed on each proband, and ES was completed on both parents, as available, for segregation analysis and verification of familial relationships. Alternative informative family members were also acceptable, dependent on family history. A report revealing the inheritance of each variant was issued for the proband, however no report was provided for parents or other relatives. Secondary findings were reported, if requested. The volume of this testing is naturally limited at our institution by genetic counselors’ availability.
Sequencing, variant detection, QC, and annotation
Genomic DNA from blood (or in three cases, fetal tissue) was prepared using either Kapa Hyper or Illumina TruSeq HT library prep following mechanical shearing on the Covaris LE220. Libraries were analyzed on a TapeStation or Fragment Analyzer to verify fragment size and distribution before being sequenced to 90 Gb or greater in 2 × 125 paired reads on the Illumina HiSeq2500 or 2 × 150 paired reads on a HiSeq4000. Basecalling was performed and required a minimum of 500 K raw cluster density with 75% passing filter. In addition, the percent of reads at or above Q30 should be greater than 80%. If these quality control (QC) metrics were satisfied, samples were processed through an alignment and variant detection pipeline using DRAGEN 2.0.4-2.1.3; although some older samples were processed using Burrows–Wheeler Aligner 0.7.2 and Genome Analysis Toolkit 3.2-2. Postpipelining QC included a minimum of 85% of reads aligning to the human genome and a minimum of 85 Gb of data obtained after alignment is complete. Variant annotation and categorization was performed using Rapid Understanding of Nucleotide variant Effect Software (RUNES v.3.4.3 – v4.2.4) as previously described.1,2
Analysis, interpretation, reporting
For each patient, the phenotype was extracted from the electronic medical record and recorded using Human Phenotype Ontology (HPO) terms.3,4 This information was entered in SSAGA, a clinical correlation tool and database that maps the phenotypic features of genetic diseases with candidate genes, allowing for archiving patient phenotypes and the generation of gene lists for variant filtering.1,2,5,6 Furthermore, Phenomizer was used to generate additional lists of candidate genes,3 as needed. However, these phenotype filters were removed to allow for manual nomination of additional genes relevant to the phenotype. Variants were identified and filtered to 1% frequency and prioritized by type using VIKING software, as previously described.1,2,5,6 Intronic variants more than 20 base pairs from the exon boundary were not analyzed unless they were previously reported in the literature as pathogenic, there was a specific gene of interest, or a partial genotype was uncovered where a second pathogenic variant would be diagnostic. No copy-number variant (CNV) caller was used, however, manual inspection of alignments was performed as needed. Variants in genes potentially related to the phenotype were interpreted using the American College of Medical Genetics and Genomics (ACMG) guidelines.7 Pathogenic, likely pathogenic, and variants of unknown significance were reported clinically in addition to pathogenic and likely pathogenic variants in the ACMG-59 secondary finding gene list,8,9 if requested by the family. For each patient, it was recorded whether follow-up of result was documented in the medical record, and whether the laboratory interpretation concurred with what was conveyed in providers’ notes.
Clinical validation and QC
Building on a wealth of experience running research GS for diagnosis of pediatric patients, including rapid genomes in infants in the neonatal intensive care unit (NICU),2,5,6 we sought to validate GS clinically. A total of 179 GS samples had been run on a research basis, including affected children and their family members. In a highly selected population such as infants in the NICU, the diagnostic rate was as high as 57%, indicating high clinical utility.5 Diagnostic variants in these cases were confirmed by clinical Sanger sequencing with no discrepancies. For clinical validation, a control sample, NA12878, was run four times on two different instruments and compared with the GetRM dataset supplied by the Centers for Disease Control and Prevention. In addition, multiple replicates of two other controls, NA12753 and NA07019, were run and compared with data generated from the Omni5 SNP chip by the HapMap project and, a research sample, UDT_173, was run twice and subjected to the same comparison. Sensitivity and specificity were determined by defining true positives as variants present in both GS and SNP chip; false negatives were called if a variant was present in the SNP chip data alone; false positives were defined as variants called only by GS. True negatives were defined as negative in both NGS and SNP chip data. In addition, alignment statistics were provided. This analysis was extended to compare single read pools as well as a hybrid, single read, and paired end pool, with no differences observed. The following was determined based on averaging the calculations for individual runs: sensitivity, 98.7%; specificity, 99.2%; accuracy, 98.4%; precision, 99.8%. Rapid versus nonrapid: exomes and targeted panel runs were compared in v2 rapid chemistry versus v4 high output chemistry. NA12878 genomes were compared in v4 high output versus v1 ultra rapid chemistry; results were essentially identical. In addition, alignment rates as well as sensitivity/specificity were compared between rapid runs and nonrapid on the HiSeq2500 and found to be equivalent.
The average depth of coverage for coding region bases was 37.71×, with 98.2% of bases covered at 10× and 88.5% covered at 20× (Suppl. Table 2). Median exonic coverage was 38.1×. Across the entire genome, average coverage was 40.7×, with 97.1% of all bases covered at 10× and 91.3% of bases covered at 20×. Mean number of variants was 4.81 M with 4.13 M SNV and 676,043 K small indel calls per patient. Of these, on average, 457,716 had a frequency less than 1% and 26,892 were coding. CNVs were not evaluated systematically by a software tool, but alignments were examined manually as needed. Array-CGH or SNP array was performed separately, as requested.
Results
Diagnostic yield
A total of 20 definitive diagnoses were made in 19 patients, yielding a diagnostic rate of 24% (n = 80) (Table 2). Of these, 1 patient had 2 diagnoses. In addition, a number of patients have potentially positive results requiring further follow-up with functional or other studies (Suppl. Table 1). Two cases had definitive diagnoses but had additional symptoms not explained by the diagnosis: for example, case 1 had a diagnosis of Mowat–Wilson syndrome, with additional findings including hypogammaglobulinemia, lymphopenia, and fever; and case 70 had an FBN1 variant consistent with Marfan syndrome but had an undiagnosed arrhythmia as well. The average number of variants reported per patient was 7.6 (6.5 for diagnostic group and 8 for nondiagnostic group; Table 1) Not surprisingly, patients who underwent trio sequencing had a higher rate of diagnoses (15/57, 26%) than other modalities (4/23, 17%; Table 1). Of the diagnostic group, 6 of 19 patients (32%) were deceased, including 3 fetal demises and one still birth, whereas only 4 of 61 in the undiagnosed group (7%) were deceased (P = 0.004; significant). The odds of receiving a diagnosis were 7.01 (95% confidence interval [CI] = 2.25, 21.82; p = 0.001) times higher in those ordered in an inpatient setting (52%) than those completed in outpatient settings (14%). However, there was no statistically significant difference in rate of diagnosis between testing ordered by genetics as compared with other specialties, nor was there a difference by age at the time of testing. Although every potential inheritance pattern was observed in this study, the most common mode was de novo (10/20 or 50%; Tables 1, 2). In one case, the inheritance pattern was unclear because the patient was compound heterozygous for two predicted loss of function variants in MYH3, a gene associated with autosomal dominant disease. The average turnaround time was 72.5 days; however, results were expedited in critical cases upon request.
Secondary findings
During the informed consent process, 73/80 (91.2%) families opted in to receive secondary findings. Secondary findings were reported in four cases, with variants in APOB, PMS2, RYR1, and TNNT2 (Suppl. Table 1).
Comparing ES vs. GS
To compare ES with GS, diagnostic variants from cGS were examined in a summary coverage report for an exome validation dataset of 814 samples. In all cases except two, the diagnostic genotype detected by GS would have been detected by ES given sufficient coverage of the nucleotide (Suppl. Table 2); the exceptions being two partial gene deletions. These cases illustrate the superiority of GS for the detection of deletions (Suppl. Data: Illustrative cases).
Communication of results
Genomic reports are complex, and the communication of results to families is challenging. The amount of information in genomic reports may be difficult to prioritize, with an average of eight variants reported per patient in this cohort. Providers may have limited knowledge of variant interpretation categories as well as evidence-based guidelines used for variant classification. In addition, families may have limited understanding of the information conveyed. Given the importance of cGS results and their potential impact for clinical management, we sought documentation in all 80 cases of communication of results by the provider to the family in the electronic medical record (EMR) and whether this was in agreement with what was reported by the laboratory. Surprisingly, for 14 of 80 (17.5%) patients, we did not find documentation of communication of the results to the family (Suppl. Table 1). Of the cases not disclosed to families, most were nondiagnostic; however, case 24 had a diagnostic result as well as a reportable secondary finding. Seven of the 14 cases (50%) with no documentation were ordered as inpatients, despite representing only 26% of the overall cohort. Indeed, documentation of results communication was not found for 33% of inpatients compared with 12% of outpatients (odds ratio [OR] = 3.71, 95% CI = 1.12, 12.36; p = 0.032). This failure to document return of results in the EMR raises concern that results were not disclosed to families; of note in three cases results were not even viewed by treating providers. The odds of GS cases referred by Genetics not having documentation of result communication were 0.136 times lower (95% CI = 0.03, 0.54; p = 0.005) than that of the odds in cases referred by other subspecialties.
Of the remaining 66 cases with documented results disclosure, 6 (9%) were found to be discordant between the laboratory report and clinicians’ interpretations recorded in the EMR (Suppl. Table 3). From the laboratory perspective, five of these cases were considered nondiagnostic and reported with variants of uncertain significance (VUSs). In case 23, the physician mistakenly interpreted two VUS in two different autosomal recessive genes with similar names as being diagnostic, citing “two mutations in ATP13, consistent with DYT12.” In case 22, the lab reported a nondiagnostic genotype with compound heterozygous VUSs in TCIRG1, interpreted by the clinician as being diagnostic for autosomal recessive osteopetrosis, a transplantable condition. The patient was placed on the transplant list but has no matched donor and currently shows minimal biochemical evidence of disease. In case 26, a maternally inherited VUS in TSC2, c.1440_1441insGAG, also present in an affected sibling, was communicated as being a VUS by a clinical geneticist following examination specifically looking for stigmata of tuberous sclerosis complex (TSC). Although both siblings had imaging showing renal angiomyolipomas, in the absence of other findings (including normal brain magnetic resonance image [MRI]) there was insufficient criteria to clinically diagnose TSC. However, follow-up clinic notes in other subspecialties, including neurology and nephrology, recorded “genetically confirmed or likely TSC” and patient was transitioned to TSC clinic. The fourth discrepancy was in case 44, a stillborn female fetus with prenatal onset of arthrogryposis (reported as patient 4) (ref. 10) and a de novo BICD2 variant reported as pathogenic and diagnostic. Because BICD2 variants had not been associated with prenatal onset disease, the clinical team hedged on calling it diagnostic, emphasizing a potential role of VUSs in a second gene, AGRN. Since the time of reporting, at least four isolated patients with arthrogryposis multiplex congenita and hypotonia have been reported with de novo BICD2 missense variants.11,12 In case 45, the ordering provider concurred that the study was inconclusive, however, subsequent notes from a provider in the same clinic called the variant pathogenic and diagnostic. Finally, in case 55, while the cGS results were reviewed in the genetics follow-up note as being nondiagnostic VUSs, array-CGH was recommended and reported as a CNV, which was interpreted in a different clinic as a “known diagnosis.”
Cost and reimbursement
For this cohort, we obtained insurance preauthorization for 56 patients, and reimbursement data was available for 38 patients billed for GS. Cumulatively, the average total reimbursement was 30.2%, however government payers reimbursed at a lower rate overall than commercial payers, with the 17 claims filed to commercial insurance, yielding 54.1% reimbursement versus only 13.1% for the 21 claims filed to government payers. Some of this is explained by the capitation of one of the largest government payers in this area; however, even when taking this into consideration the reimbursement rate is lower than commercial payers. Another explanation is that commercial payers are more likely to require authorization while government payers often respond “Authorization not required.” Testing performed when authorization is neither required nor obtained is more likely to be denied after the test is performed. In addition, we note a potential trend of rejecting cGS in favor of NGS-based panels or ES. Indeed, GS is considered “experimental” and is not covered by one of our largest commercial payers.
Discussion
Here we describe the first validation of cGS, and present the results, reimbursement rate, and follow-up from the first 80 cases reported clinically at a pediatric hospital. Our findings indicate that cGS is equivalent and potentially superior to ES, with a diagnostic rate of 24% in this cohort. As with any study, detection rates for both ES and GS range from 25 to 57% varying with the inclusion criteria.1,5,6,13,14,15,16 Unlike other studies of GS with highly selected patients based on degree of suspicion of a genetic disease, high rates of consanguinity,16,17,18 insistence on the availability of parent samples for trio analysis,16 or other inclusion criteria that could positively influence the diagnostic rate, this group is an unbiased cohort based on normal ordering patterns from a range of subspecialists in a pediatric setting. The average age of our patients was 6.9, and only 2 of 19 (10.5%) patients with a diagnosis were from consanguineous parents. In addition, this analysis was done using a conservative application of the ACMG guidelines for variant interpretation, unlike some studies that predated such guidelines.1,13 This study confirms the value of including parental samples in the analysis, as 95% of our diagnoses were patients with at least one parental sample run, and 79% were full trios. In addition, as other studies have reported,5,19 we confirm that patients in the diagnostic group have a significantly higher death rate (32% diagnosed vs. 7% undiagnosed; p = 0.025).
Negative results in the ~50–75% of patients undergoing ES or GS may be due to several factors, including the limited ability to accurately interpret variants, particularly in noncoding regions, current technical limitations, atypical patient phenotypes at the time of testing, and nongenetic or complex disease etiologies. In addition, there is still a large number of genes with limited or no known association with human diseases, with 20 or more new OMIM associations every month. However, even in well-characterized genes, variant interpretation remains challenging, with most interpreted as VUS. This highlights a tremendous need for additional studies to elucidate variant pathogenicity and further vet genes of unknown function. Although noncoding variants such as deep intronic variants affecting splicing or regulatory regions are detectable by GS, such variants, which far outnumber those in the coding regions, are not part of our current analysis protocol unless previously reported in the literature as pathogenic. The effects of such variants require functional characterization, and such investigation is outside the scope of clinical testing. However, patient variant files are stored for potential reanalysis when knowledge of this data matures. Additional explanations for undiagnosed patients include technical limitations associated with short read sequencing, which preclude the detection of triplet repeats, methylation defects, variants in homologous regions, and, in the absence of special software, many copy-number variants. While ordering physicians may submit a periodic reanalysis of cGS to interrogate newly discovered genes with human disease associations and updated variant annotation, automatic reanalysis is not performed as part of our procedure nor was it done for this study.
The advantages of cGS are many, both on the laboratory and analysis side. The workflow for laboratory staff is less labor intensive because there is no enrichment step. Other positive factors are evenness and overall less bias in coverage as compared with ES. Importantly, increased coverage results in a decrease in supplemental Sanger fill-ins for areas missed by ES, increased resolution of absence of heterozygosity for potential uniparental disomy, and exonic deletions, whole gene or multigene, are potentially detectable. Although a pipeline for detection of copy-number variants was not used, two partial gene deletions were found by manual inspection of alignments, leading to a faster diagnosis for critical patients in two cases (Suppl. data: Illustrative cases). Certainly the presence of a single VUS may have triggered additional testing such as exon-level array-CGH; however this would depend on the degree of interest in the candidate gene based on phenotypic fit, follow-up of the clinician, and regardless, would add to the time to report the final result. For both ES and GS, short read technology poses a challenge for the detection of variants complicated by pseudogenes or other homologous regions, and repetitive regions. However, advances in technology, both in software development and laboratory solutions, are beginning to offer feasible solutions for reliable CNV detection from GS and ES.
Costs and reimbursement
Despite the advantages, one cannot ignore the current expense of GS compared with ES, which is at least five times higher. Storage costs must also be considered, and GS generates 10 times the data of an exome.
Reimbursement for genetic testing remains challenging, and advocating for fair payment for these services is needed. Indeed, commercial payers with negotiated rates paid well; however, reimbursement by government payers was much lower and had a higher denial rate. Efforts to educate such entities on the importance and potential cost-effectiveness of this testing are needed. In a self-contained institution it is potentially worthwhile to offer genomic services regardless of reimbursement in certain patients if it lowers overall costs of hospitalization or other testing. However, this requires further study.
Utilization of genomic results
We have identified a significant problem of undisclosed or undocumented genomic results, unlikely limited to GS or our institution. This is concerning for any laboratory test, but is especially problematic for cGS/ES, the results of which often change the management of patients1,5,6,16,19 and is extremely costly to perform. It is certainly possible that verbal communication occurred and was not documented in the EMR; however in three cases there was a failure to view the cGS results by the ordering provider. The finding that 50% of the patients whose cGS was ordered during a hospitalization potentially did not receive results, indicates continuity of care following discharge is potentially problematic and provides an opportunity for intervention. While lab stewardship was found to be a problem in many subspecialties, it was relatively minimal in genetics, suggesting the involvement of a genetic counselor, for many reasons, is ideal. Of those results where communication was documented in the EMR, there was discordant interpretation between what the lab reported and what the provider communicated to the patient in six cases. The apparent reasons for this varied, but in half the cases was related to the patient phenotype, whether upgrading a VUS because it was felt to fit the phenotype, or de-emphasizing a variant because the phenotype was atypical. Variant interpretation guidelines do take phenotype into account but this is not considered strong criteria. The majority of cases with discordant interpretations between the laboratory and provider (5/6) were considered nondiagnostic by the laboratory, suggesting physicians may upgrade variants without the level of evidence required by the laboratory or may not understand the variant classification system used. Education of clinicians on methods for variant classification may be helpful. Interestingly, there was discordance in interpretations between different providers in three cases, which may reflect differing levels of genetics education. What the families are told by providers and what they understand adds an additional layer of complexity. This dataset is small but highlights a need for additional studies to identify the problem of failure to retrieve, disclose, or document genomic testing results, as well as variant reinterpretation by clinicians.
The increasing number of atypical or novel presentations catalogued as a broadened phenotypic spectrum for any particular genetic condition formerly ordered as a single-gene or panel-based test challenges the previous model of using serial genetic testing dictated by phenotype. With notable exceptions, most genetic conditions have enough heterogeneity and/or variability to warrant broad screening such as ES or GS. Many studies offer comparisons between “conventional” and genomic testing.1,5,6,13,16,19,20,21,22,23 At our institution, genomic medicine, including GS, has been used as conventional testing for several years, which benefits our patients by shortening their course to diagnosis and potential treatment.1,2,5,6 Here we show GS is equivalent or perhaps superior to ES from both the laboratory and diagnostic perspective. However, in all cases but two, where deletions where identified, ES would yield the same usable information for current clinical analysis. Therefore, in the absence of a low-cost high-throughput instrument, the cost of GS is currently difficult to justify, given the availability of ES. However, as the cost of sequencing decreases and the ability to interpret noncoding variants improves, an increasing diagnostic advantage of GS over ES will be realized.
References
Soden SE, Saunders CJ, Willig LK, et al. Effectiveness of exome and genome sequencing guided by acuity of illness for diagnosis of neurodevelopmental disorders. Sci Transl Med. 2014;6:265ra168.
Saunders CJ, Miller NA, Soden SE, et al. Rapid whole-genome sequencing for genetic disease diagnosis in neonatal intensive care units. Sci Transl Med. 2012;4:154ra135.
Kohler S, Vasilevsky NA, Engelstad M, et al. The Human Phenotype Ontology in 2017. Nucleic Acids Res. 2017;45(D1):D865–D876.
Shen F, Wang L, Liu H. Phenotypic analysis of clinical narratives using Human Phenotype Ontology. Stud Health Technol Inform. 2017;245:581–5.
Willig LK, Petrikin JE, Smith LD, et al. Whole-genome sequencing for identification of Mendelian disorders in critically ill infants: a retrospective analysis of diagnostic and clinical findings. Lancet Respir Med. 2015;3:377–87.
Petrikin JE, Cakici JA, Clark MM, et al. The NSIGHT1-randomized controlled trial: rapid whole-genome sequencing for accelerated etiologic diagnosis in critically ill infants. NPJ Genom Med. 2018;3:6.
Richards S, Aziz N, Bale S, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–24.
Green RC, Berg JS, Grody WW, et al. ACMG recommendations for reporting of incidental findings in clinical exome and genome sequencing. Genet Med. 2013;15:565–74.
Kalia SS, Adelman K, Bale SJ, et al. Recommendations for reporting of secondary findings in clinical exome and genome sequencing, 2016 update (ACMG SFv2.0): a policy statement of the American College of Medical Genetics and Genomics. Genet Med. 2017;19:249–55.
Ahmed AA, Skaria P, Safina NP, et al. Arthrogryposis and pterygia as lethal end manifestations of genetically defined congenital myopathies. Am J Med Genet A. 2018;176:359–67.
Ravenscroft G, Di Donato N, Hahn G, et al. Recurrent de novo BICD2 mutation associated with arthrogryposis multiplex congenita and bilateral perisylvian polymicrogyria. Neuromuscul Disord. 2016;26:744–8.
Storbeck M, Horsberg Eriksen B, Unger A, et al. Phenotypic extremes of BICD2-opathies: from lethal, congenital muscular atrophy with arthrogryposis to asymptomatic with subclinical features. Eur J Hum Genet . 2017;25:1040–8.
Yang Y, Muzny DM, Reid JG, et al. Clinical whole-exome sequencing for the diagnosis of mendelian disorders. N Engl J Med. 2013;369:1502–11.
Lee H, Deignan JL, Dorrani N, et al. Clinical exome sequencing for genetic identification of rare Mendelian disorders. JAMA. 2014;312:1880–7.
Valencia CA, Husami A, Holle J, et al. Clinical impact and cost-effectiveness of whole exome sequencing as a diagnostic tool: a pediatric center’s experience. Front Pediatr. 2015;3:67.
Lionel AC, Costain G, Monfared N, et al. Improved diagnostic yield compared with targeted gene sequencing panels suggests a role for whole-genome sequencing as a first-tier genetic test. Genet Med. 2017;20:435–43.
Powis Z, Farwell KD, Alamillo CL, et al. Diagnostic exome sequencing for patients with a family history of consanguinity: over 38% of positive results are not autosomal recessive pattern. J Hum Genet. 2016;61:173–5.
Charng WL, Karaca E, Coban Akdemir Z, et al. Exome sequencing in mostly consanguineous Arab families with neurologic disease provides a high potential molecular diagnosis rate. BMC Med Genomics. 2016;9:42.
Meng L, Pammi M, Saronwala A, et al. Use of exome sequencing for infants in intensive care units: ascertainment of severe single-gene disorders and effect on medical management. JAMA Pediatr. 2017;171:e173438.
Bourchany A, Thauvin-Robinet C, Lehalle D, et al. Reducing diagnostic turnaround times of exome sequencing for families requiring timely diagnoses. Eur J Med Genet. 2017;60:595–604.
Costain G, Jobling R, Walker S, et al. Periodic reanalysis of whole-genome sequencing data enhances the diagnostic advantage over standard clinical genetic testing. Eur J Hum Genet . 2018;26:740–4.
Tan TY, Dillon OJ, Stark Z, et al. Diagnostic impact and cost-effectiveness of whole-exome sequencing for ambulant children with suspected monogenic conditions. JAMA Pediatr. 2017;171:855–62.
van Diemen CC, Kerstjens-Frederikse WS, Bergman KA, et al. Rapid targeted genomics in critically ill newborns. Pediatrics. 2017;140:1–11.
Acknowledgements
We gratefully acknowledge the patients and their families, as well as the referring physicians and thank our colleagues in the Center for Pediatric Genomic Medicine.
Disclosure
The authors declare no conflicts of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, and provide a link to the Creative Commons license. You do not have permission under this license to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Thiffault, I., Farrow, E., Zellmer, L. et al. Clinical genome sequencing in an unbiased pediatric cohort. Genet Med 21, 303–310 (2019). https://doi.org/10.1038/s41436-018-0075-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-018-0075-8