Abstract
Purpose
Genetic testing has uncovered large numbers of variants in the BRCA2 gene for which the clinical significance is unclear. Cancer risk prediction of these variants of uncertain significance (VUS) can be improved by reliable assessment of the extent of impairment of the tumor suppressor function(s) of BRCA2.
Methods
Here, we evaluated the performance of the mouse embryonic stem cell (mESC)-based functional assay on an extensive set of BRCA2 missense variants.
Results
Whereas all 20 nonpathogenic (class 1/2) variants were able to complement the cell lethal phenotype induced by loss of endogenous mouse Brca2, only 1 out of 15 pathogenic (class 4/5) variants (p.Gly2609Asp) was able to do so. However, in this variant the major tumor suppressive activity of BRCA2, i.e., homology directed repair (HDR), was severely abrogated. Among 43 evaluated VUS (class 3), 7 were unable to complement the lethal phenotype of mouse Brca2 loss while 7 other variants displayed a more severe reduction of HDR activity than observed for class 1/ 2 variants.
Conclusion
The mESC-based BRCA2 functional assay can reliably determine the functional impact of VUS, distinguish between pathogenic and nonpathogenic variants, and may contribute to improved cancer risk estimation for BRCA2 VUS carriers.
Similar content being viewed by others
Introduction
Inherited pathogenic variants in BRCA2 confer an average cumulative risk by age 70 years of about 60% to develop breast cancer in combination with an increased ovarian cancer risk up to 18%.1,2 In addition, they may increase the risk to develop other cancer types including pancreatic and prostate cancer.3,4 Clinical management of carriers of pathogenic variants consists of intensified screening programs, the option to undergo prophylactic surgery, and presymptomatic genetic testing of family members.5 Furthermore, targeted treatment of BRCA1/2-associated tumors has recently become available following the registration of poly ADP ribose polymerase (PARP) inhibitors.6
Variants associated with high cancer risk typically disrupt BRCA2 function, but for many variants identified by genetic testing the functional impact cannot be inferred from sequence information alone. Those variants are therefore defined as variants of uncertain significance (VUS)7 and they represent a major challenge for genetic counselling and clinical management of the families in which they are identified. Depending on the inclusion criteria used for genetic testing these may comprise up to 20% of all identified variants.7,8,9 Worldwide, more than 2000 unique BRCA2 VUS have been identified, including missense and silent substitutions, small in-frame insertions and deletions, and intronic variants.10,11
To classify BRCA2 variants, a prior likelihood of pathogenicity, based on in silico analysis of the sequence alteration, can be combined with the available genetic and epidemiological data (such as family history of cancer, tumor histopathology, cosegregation with disease, and cooccurrence) to calculate the posterior probability that a variant is pathogenic, in a so-called multifactorial likelihood model.9 Based on these posterior probabilities a 5-tier classification system has been introduced in which each class is associated with specific recommendations for clinical management.12 Class 1/2 variants have very low posterior probabilities (<0.05) for pathogenicity and are probably benign, whereas class 4/5 variants have very high posterior probabilities (>0.95) for being associated with cancer risk equivalent to classical pathogenic variants that encode a truncated BRCA2 protein. By definition, a VUS falls in class 3. Unfortunately, as the occurrence of a VUS is usually rare, there are often insufficient clinical data to make clinically meaningful inferences about their associated cancer risks and therefore it has been difficult to move a VUS into either class 1/2 or class 4/5 on the basis of clinical data. Information on protein conformation or function can serve as independent classifiers of VUS.13,14,15,16,17 Quantitative functional data transformed into likelihood ratios might improve the performance of predictive models when used in combination with available genetic and epidemiological data. BRCA2 plays a major role in the DNA damage response (DDR), the network of interacting pathways that together execute the response upon DNA damage. Disruption of BRCA2 leads to a deficiency in homologous recombination and consequently sensitivity to DNA damaging agents that induce replication fork stalling and DNA double strand breaks (DSBs).18 Various assays have been developed to assess the functional impact of variants in BRCA219 which are able to accurately discriminate between nonpathogenic (class 1/2) and pathogenic variants (class 4/5).
A mouse embryonic stem cell (mESC)-based assay was developed for the functional assessment of human BRCA2 variants.20,21 This mESC-based system involves the introduction of human BRCA2 into a hemizygous mouse Brca2 mESC line and allows evaluation of the functional consequences of any BRCA2 variant, including those located in either exonic or intronic sequences that may affect RNA splicing. Variants are being assessed for their ability to complement the loss of cell viability following Cre-mediated deletion of a conditional Brca2 allele. Nonfunctional protein variants are unable to overcome Brca2 loss while variants resulting in (partially) functional BRCA2 protein are able to complement cell lethality and can be assessed for their capacity to perform BRCA2 functions.20
An extensive validation was performed to determine the sensitivity and specificity of the assay. To this end, we evaluated its ability to correctly discriminate between pathogenic (class 4/5 [n = 15]) and nonpathogenic (class 1/2 [n = 20]) variants that had previously been classified on the basis of genetic and clinical data. In addition, we assessed its performance on variants characterized in previously published functional studies 14,17,20,22 (n = 20) and determined the functional impact of 23 VUS identified in the clinic.
Materials and methods
Selection of classified missense variants in BRCA2
To establish the sensitivity and specificity of the BRCA2 mESC-based assay, we selected all known pathogenic missense variants (class 4/5 [n = 15]) and a similarly sized set of nonpathogenic missense variants (class 1/2 [n = 20]) in BRCA2 for which the clinical significance has been established on the basis of clinical and genetic data (Table 1).19
Selection of VUS in BRCA2
Functional analysis was performed for a set of VUS (n = 43) consisting of 3 variants that previously have been tested in a mESC-based system20,22 and 17 variants characterized by a complementary DNA (cDNA)-based functional assay in V-C8 Chinese hamster cells.14,17
In addition, we included 23 variants that have been identified in counselees visiting one of the Dutch Clinical Genetic Centers and who were eligible for genetic testing according to the Dutch guidelines (Table 2). The majority of the variants were missense or single amino acid deletion variants. In silico splice site prediction analysis was performed for all variants using five different algorithms.23 With the exception of c.6935A>T (p.Asp2312Val), which is predicted to lead to reduced donor site recognition of exon 12 by two programs, an effect on RNA splicing for the other variants is unlikely.
Generation of variants in the human BRCA2 gene
A library of pUC19 plasmids each containing a single BRCA2 exon and about 100 nucleotides of intron sequence upstream and downstream of the exon was generated to enable introduction of variants (QuickChange site-directed mutagenesis from Stratagene). Following sequence confirmation, homology arms were added to the exon containing the variant by polymerase chain reaction (PCR). Subsequently, Red/ET recombineering was used in combination with a positive/negative selection procedure to introduce the variant into a bacterial artificial chromosome (BAC) (clone RP11-777I19, BACPAC) containing the full length human BRCA2 gene as described previously.21
Cell culture
mESC culture was performed as described previously.21 Prior to transfections, protein/RNA isolations, and biological assays, cells were cultured on gelatin-coated plates using buffalo rat liver cell (BRL)-conditioned mESC medium.
Insertion of the DR-GFP construct in mouse embryonic stem cells
Insertion of the pX59 direct repeat green fluorescent protein (DR-GFP) construct at the Pim1 locus in the Brca2 -/loxP mESC line21 was performed as described.24 Integration of the construct was confirmed by Southern blot analysis and via PCR using the following primers:
Pim1Ex1F: 5′-AAGATCAACTCCCTG GCCCACCTGCG-3′, Pim1Ex4R 5′-TGTTCTCGTCCTTGATGTCG-3, and Hyg3A 5′-CCGCTCGTCTGGCTA AGAT-3′ (Figure S1).
Generation of mouse embryonic stem cells expressing BRCA2 variants
BACs carrying BRCA2 variants were transfected into the Pl2F7 conditional Brca2 knockout mESC line containing the DR-GFP construct (Brca2-/loxP; Pim1DR-GFP/WT). This mESC line contains one conditional endogenous Brca2 allele with a loxP site on either side of the complete Brca2 locus and one disrupted endogenous Brca2 allele in which exon 11 has been disrupted (Figure S2). Per variant 1.5 × 106 cells were transfected in suspension with 4 µg BAC DNA using Lipofectamine 2000 (Invitrogen). Transfected cells were subsequently split over two 60-mm cell culture dishes and cultured in the presence of G418 (200 μg/ml), starting 24 h post transfection. Ten days post transfection G418 resistant clones of each 60-mm dish were pooled (at least 50 clones) providing two independent polyclonal cell populations (PCPs).
RT-PCR and western blot
BRCA2 expression was determined in two independent PCP per variant by reverse transcription PCR (RT-PCR) and western blot analysis (Figure S3).21 Western blot analysis was performed using NuPAGE™ Novex™ 3–8% Tris-Acetate Protein Gels (ThermoFisher Scientific). BRCA2 protein was detected with the rabbit polyclonal antibody (BETHYL, A303–434A-T-1) recognizing the region between amino acids 450–500 of human BRCA2. Protein signal was detected by electrochemiluminescence (Amersham ECL RPN2235 Biocompare) and quantified using ImageQuant TL software.
Functional complementation of mouse embryonic stem cells expressing BRCA2 variants
The conditional Brca2 allele was removed following transient transfection with a Cre-expressing plasmid (pCAG-Cre:GFP from Addgene). Recombination between the loxP sites restores the HPRT1 minigene and allows cells to grow in the presence of hypoxanthine–aminopterin–thymidine (HAT).20 For each variant two independent PCPs of BAC clones were used for transfection. Per PCP 1.2 × 106 cells were transfected with 4 µg plasmid DNA in suspension using Lipofectamine 2000 (Invitrogen) and subsequently seeded in triplo on 90-mm cell culture dishes. The transfection efficiency was determined by monitoring GFP expression 16 h after transfection. Cells were cultured for 6 days in the presence of HAT and subsequently 5 days in the presence of hypoxanthine-thymidine (HT). Thirteen days after transfection one culture dish was used for visualization of clones by methylene blue staining. Clones arising on the remaining two culture dishes were pooled. This procedure allows downstream functional analysis of two independent PCPs per variant (Figure S4).
Cell cycle analysis
Per variant, two PCPs of HAT-resistant clones were subjected to functional analysis in the HDR assay and survival assays. In parallel 15 × 103 cells were seeded per well in 96-well plates to detect potential effects of the variant on cell cycle progression. Two days after seeding cells were fixed in ice cold ethanol (70%) and subsequently DNA was stained with propidium iodide (50 µg/ml) in the presence of RNaseA (0.1 mg/ml) and Triton X-100 (0.05%). The fluorescence intensity was measured by flow cytometry (Guava, Millipore) and used to determine cell cycle phase distribution (Flowing software 2). Cell cycle analysis was performed in duplo for each PCP (50,000 cells), resulting in four cell cycle measurements per variant.
HDR assay
BRCA2-expressing cells were seeded at a density of 15 × 103 cells per well on gelatin-coated 96-well plates in BRL-conditioned mES cell culture medium. On the next day, cells were transfected with an I-Sce1-mCherry plasmid26 using Lipofectamine 2000. Three days after transfection the fraction of mCherry/GFP double-positive cells was determined by flow cytometry (Guava, Millipore). For each variant the proportion of GFP-positive cells was calculated relative to wild type (WT) BRCA2. HDR measurements were performed in triplo for each independent PCP resulting in six data points per variant.
Sensitivity assays with cisplatin and PARP inhibitor treatment
BRCA2-expressing cells were seeded at a density of 30 × 103 cells per well on gelatin-coated 96-well plates to measure the sensitivity of BRCA2 variants to the DNA crosslinking agent cisplatin (Accord) and the PARP inhibitor KU-0058948 (Axon Medchem). Cells were treated in triplo with the following concentrations of cisplatin: 0 µM (1% phosphate-buffered saline [PBS]), 0.16 µM, 0.31 µM, 0.63 µM, 1.25 µM, and 2.5 µM. Applied concentrations of PARP inhibitor were 0 nM (1% dimethyl sulfoxide [DMSO]), 8 nM, 16 nM, 31 nM, 62.5 nM, and 125 nM. Cell counts were determined after 48 h of continuous exposure by flow cytometry (Guava, Millipore). Survival was calculated by dividing the number of surviving cells in treated samples by the number of untreated cells of the same variant. The IC50 values plotted for the cisplatin survival assays were calculated as previously described using the drc package in the R programming language.26,27
Results
The widespread use of BRCA1/2 pathogenic variant testing in individuals with a family history of breast or ovarian cancer has uncovered large numbers of VUS. Classification of a VUS as either class 1/2 or class 4/5 may be improved when the impact of a VUS on BRCA2 protein function can be taken into consideration. To this end, a semihigh-throughput assay with robust quantitative read-outs is needed. We therefore further optimized the mESC-based BRCA2 functional assay20,21 (Fig. S2). Here, we validated its performance on a large series of pathogenic and nonpathogenic missense variants. The specificity and sensitivity of the optimized mESC system was determined by assessing the functional impact of a panel of class 1/2 and class 4/5 missense variants associated with respectively a low and high posterior probability of pathogenicity (Table 1, Fig. 1). In addition, we included an extensive series of BRCA2 VUS to evaluate their effect on protein function (Table 2, Fig. 1).
Position of BRCA2 variants. Schematic representation of the position of class 1/2 variants (green dots), class 4/5 variants (red dots), and BRCA2 variants of uncertain significance (VUS) (gray dots) in the BRCA2 gene. Ta transcriptional activation domain, P phosphorylation site, H helical domain, DBD DNA binding domain, OB oligonucleotide binding fold, T Tower domain, TR2 C-terminal RAD51 binding site, NLS nuclear localization signal, aa amino acids19,39,40
Cell cycle analysis
HDR is selectively used for the repair of DSBs in the S and G2 phase of the cell cycle when a sister chromatid is available to serve as donor for homologous DNA sequences.28,29 To exclude the possibility that a reduction in HDR activity observed for a BRCA2 variant is the consequence of a smaller fraction of cycling cells, we performed cell cycle distribution analysis of all BRCA2 variants by propidium iodide staining (Fig. S5), and observed no significant effect on cell cycle phase distribution among the nondeleterious BRCA2 variants.
Survival and HDR activity of class 1/2 and class 4/5 BRCA2 variants
Functional BRCA2 is essential for cell survival and normal embryonic development in mice.30,31 We first tested the ability of 15 class 4/5 variants to rescue the lethal cell phenotype of loss of Brca2 function after Cre-mediated removal of the conditional Brca2 allele. Fourteen of these variants were unable to form HAT-resistant clones in the absence of functional Brca2 (Table 1, Fig. 2), indicating that BRCA2 function was severely affected. Only the class 4 variant p.Gly2609Asp allowed outgrowth of a reduced number of HAT-resistant clones compared with WT BRCA2-expressing cells (Fig. 2). In contrast, all 20 class 1/2 variants were able to fully complement loss of endogenous Brca2 (Table 1, Fig. 2).
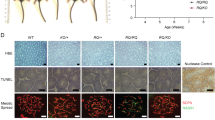
Representative images of complementation phenotypes and controls. Brca2-/loxPPim1DR-GFP/WT cells expressing WT BRCA2 or BRCA2 variants were transfected with a Cre-GFP expression plasmid to induce loss of the conditional Brca2 allele and restore the Hprt gene. Upon Cre-recombinase expression cells become Brca2 deficient, which is lethal unless complemented by the expression of a (partially) functional BRCA2 variant. Untransfected cells that still contain the conditional Brca2 allele lack Hprt expression and will subsequently not survive HAT selection as shown in the –Cre-recombinase control. Thirteen days post Cre-GFP transfection culture dishes were stained with methylene blue
Next, we assessed the capacity of surviving variants (i.e., all class 1/2 variants and class 4 variant p.Gly2609Asp) to use HDR for the repair of an I-Sce1-induced DSB in a DR-GFP reporter construct integrated at the Pim1 locus.25 HDR activity of class 1/2 variants relative to WT BRCA2 ranged between 50 and 120% (Fig. 3a), with the strongest reduction observed for variants p.Pro2589His, p.Arg2888Cys, and p.Arg3052Gln. In contrast, the class 4 variant p.Gly2609Asp displayed a more severe functional impairment of BRCA2 function with an HDR reduction of 72%. Based on the results obtained with the validation set, variants with an HDR level higher than 50% can be classified as class 1/2, whereas class 4/5 variants either are unable to rescue the lethal cell phenotype of Brca2 loss or display an HDR level below 30% of WT BRCA2. Accordingly, both the sensitivity and specificity of the assay were estimated at 100% (95% confidence interval 78–100 and 83–100% respectively).
Homology directed repair (HDR) activity of BRCA2 variants relative to wild type (WT) BRCA2 activity. GFP signal was measured in I-Sce1 expressing cells 2 days post transfection by flow cytometry for (a) classified BRCA2 missense variants and (b) BRCA2 variants of uncertain significance (VUS). Relative HDR activity is expressed as the ratio between the percentage of GFP-positive cells observed in BRCA2 variant expressing cells and the percentage of GFP-positive cells in WT BRCA2-expressing cells (green line). Brca2 (green bar) represents the conditional Brca2-/loxPPim1DR-GFP/WT cell line expressing endogenous Brca2. The upper gray box represents the HDR range of class 1/2 BRCA2 variants. The lower gray box represents the HDR range associated with >95% probability of pathogenicity as reported by Guidugli et al.17 (see also Fig. S6b). The black bars correspond to the mean HDR activity and error bars indicate the SD of at least six independent GFP measurements per variant as represented by the dots (purple = class 1/2, red = class 4/5)
Survival and HDR activity of BRCA2 VUS
Given the limited number of class 4/5 missense variants, we expanded our analysis with VUS that had previously displayed an impairment of BRCA2 function in other studies (n = 20) (refs. 17,19,22) including 2 VUS significantly associated with an intermediate risk of breast cancer (Fig. S8).32 In addition, we evaluated the functional consequences of 23 VUS (with focus on missense and single amino acid deletion variants) that had been identified by genetic screening (Table 2). The complementation phenotype of 7 of the 43 VUS tested, i.e., variants p.Gly2585Arg, p.Arg2625Ser, p.Tyr2660Asp, p.Leu2721His, p.Lys2750del, p.Glu3002Lys, and p.Asp3073Gly, resembled that of class 4/5 variants with respect to their inability to rescue the cell lethality imposed by Cre-mediated loss of Brca2 (Fig. 2, Table 2). In the case of variants p.Trp31Cys, p.Val2652Met, p.Arg2784Trp, and p.Arg2784Gln, only small numbers of HAT-resistant clones arose after removal of the conditional Brca2 allele indicating that BRCA2 function is compromised in these variants resulting in incomplete complementation (Fig. 2, Table 2).
The ability to perform HDR was evaluated for the 36 BRCA2 VUS that were able to complement loss of the conditional Brca2 allele (Fig. 3b). HDR levels varied widely between BRCA2 VUS and a subset of variants revealed a more severe reduction in HDR activity than observed among the class 1/2 variants. Variants p.Val2652Met, p.Arg2842Cys, p.Asp2913Gly, and p.Tyr3092Ser showed a more than 50% reduction in HDR capacity compared with WT BRCA2-expressing cells, while a 70–80% reduction was detected for variants p.Trp31Cys, p.Arg2784Trp, and p.Arg2784Gln.
We wondered to what extent our HDR measurements were similar to those previously obtained with a cDNA-based assay in Brca2- deficient V-C8 cells (Fig. S6).14,17 Linear regression analysis on HDR data of 24 viable variants analyzed with both methods (11 class 1/2 variants, 1 class 4/5 variant, and 12 VUS) indicated a high level of concordance (R2 = 0.71). Interestingly, using the linear equation from the regression analysis, extrapolation of the HDR activity for 19 BRCA2 variants (14 class 4/5 and 5 VUS) that were unable to complement loss of Brca2 showed it to range between 7 and 26% (Fig. S6b). Thus, a residual HDR activity of at least 25% of WT appears to be required to allow variants to overcome Brca2 loss-induced cell lethality in mESC.
Sensitivity to cisplatin and PARP inhibitor
Due to the pivotal role of BRCA2 in the repair of DSBs, the absence of functional BRCA2 protein will render cells vulnerable to compounds that introduce toxic DNA lesions that impede cellular processes such as transcription and replication.33 The crosslinking agent cisplatin, a clinically relevant chemotherapeutic for BRCA1/2-related tumors, was used for cell survival measurements in all variants. Analogous to the wide range of HDR activity detected, half-maximal inhibitory concentrations (IC50) of cisplatin varied considerably among class 1/2 variants (Fig. S7a, Table 1). Although IC50 values in eight class1/2 variants were lower than in WT BRCA2-expressing cells, class 4 variant p.Gly2609Asp displayed the largest increase in cisplatin sensitivity with a more than twofold reduced IC50 value. The low IC50 values calculated for variants p.Trp31Cys, p.Gly2508Ser, p.Val2652Met, p.Ser2704Phe, p.Arg2784Gln, p.Arg2842Leu, p.Pro3039Arg, p.Tyr3092Ser, and p.Lys3326Ter indicate hypersensitivity toward cisplatin treatment (Fig. S7b, Table 2).
The therapeutic strategy of PARP inhibition is a relatively novel approach to exploit the absence of efficient HDR. Clinical trials have revealed promising results for the use of PARP inhibitors in BRCA-associated breast and ovarian cancers.34,35,36 It is, however, unclear to what extent partial impairment of HDR activity sensitizes cells to inhibition of PARP. We treated cells expressing BRCA2 variants with different concentrations of a PARP inhibitor (KU-0058948) for 48 h and measured cell survival by flow cytometry. Neither the class 1/2 variants nor the class 4 variant p.Gly2609Asp displayed increased sensitivity towards PARP inhibitor treatment (Fig. S9). Apart from p.Trp31Cys and p.Arg2784Gln, none of the VUS, including variants with severely compromised HDR activity (>55% reduction), displayed increased sensitivity towards PARP inhibitor treatment (Fig. S9).
Discussion
In this study, we determined the sensitivity and specificity of the BRCA2 mESC-based assay using a validation panel of classified BRCA2 missense variants. The functional impact of variants associated with a high posterior probability (class 4/5) was distinctly different from that of variants with a low posterior probability (class 1/2) of pathogenicity. All class 1/2 variants (n = 20) complemented the lethal phenotype of Brca2 deficient mESC and displayed capacities to repair an I-Sce1 induced DSB via HDR between 50 and 120% of WT BRCA2 activity. In contrast, 14 of 15 class 4/5 variants were unable to rescue Brca2 loss-induced cell lethality. Although class 4 variant p.Gly2609Asp was able to complement loss of Brca2 this variant displayed a severe reduction in HDR activity (28% of WT). Variant classification on the basis of complementation phenotype and level of HDR activity allows a clear separation of class 1/2 (HDR >50%) and class 4/5 (no complementation or HDR <30%) variants with 100% sensitivity and specificity.
Comparison of our HDR data with HDR measurements performed in Brca2- deficient V-C8 cells indicated them to be highly correlated (R2 = 0.71).14,17 In these studies an HDR cut-off of >95% probability of pathogenicity has been established at a level that is equivalent to a residual HDR activity of 30% in our measurements (Fig. S6b), in concordance with the HDR level (28%) that we obtained for class 4 variant p.Gly2609Asp.
Among the 23 VUS identified by genetic screening in the Netherlands, two variants (p.Arg2625Ser and p.Lys2750del,) did not rescue loss-of-Brca2-mediated cell lethality, strongly indicative for their pathogenicity. However, clinical and genetic information is required to confirm their association with high cancer risk. Based on the level of HDR activity, variant p.Asp2913Gly is characterized as a hypomorphic variant (HDR between 30 and 50%), while the other VUS displayed HDR activities (>50%) similar to nonpathogenic class 1/2 variants. The question remains whether hypomorphic variants (in this study defined by HDR activity between 30 and 50% of WT) are associated with a moderate risk to develop breast cancer. A recent case-control study32 showed an inverse correlation between breast cancer risk and HDR activity for four BRCA2 missense variants (Fig. S8).
The amino acid changes that either lead to the inability to complement Brca2 loss or that severely impair HDR activity cluster almost exclusively in the DNA binding domain of the protein (residues 2481–3186) (Fig. S10). The only exception is variant p.Trp31Cys, located in exon 3 encoding the PALB2 interaction domain, in agreement with previous studies.22,37
Defective HDR has been shown to sensitize cells to treatment with agents that induce DNA DSB. Indeed, in general attenuation of the HDR pathway sensitized cells to cisplatin-induced crosslinks in the DNA. Only variants p.Arg2842Cys and p.Asp2913Gly (>50% HDR reduction) showed no increased sensitivity to cisplatin. In contrast, impairment of HDR only occasionally resulted in sensitivity to treatment with a PARP inhibitor. Only variant p.Trp31Cys, and to a lesser extent p.Arg2784Gln, showed substantial sensitivity in this context.
In this study we performed experiments to assess the functional impact of BRCA2 variants on HDR, which is thought to be the most prominent tumor suppressor function of BRCA2. However, BRCA2 has been suggested to be involved in additional genome maintenance processes like centrosome amplification and replication fork stabilization, which might also be important for tumor suppression.18,38 For this reason we formally cannot exclude that carriers of variants that do not show an effect on HDR activity have no elevated cancer risk. Future studies have to demonstrate whether HDR independent functions of BRCA2 are relevant for tumor suppression.
In conclusion, the functional analysis described here provides insight into the functional consequences of BRCA2 variants and will be applied to a much broader spectrum of variants, including those that might influence RNA splicing. In the mESC assay, the functional impact of variant-associated alternative splicing can be analyzed in a monoallelic manner, which is currently impossible using other methods. Future work in collaborative studies within the Evidence-based Network for the Interpretation of Germline Mutant Alleles (ENIGMA) Consortium will be focused on the translation of functional results into associated cancer risk. The ultimate aim is to incorporate the functional read-outs into a multifactorial model in which the functional and clinical data on a specific VUS are merged to arrive at a decisive posterior probability of pathogenicity. These efforts will enable accurate cancer risk estimation and support clinical decision making in the absence of sufficient genetic data.
References
Mavaddat N, Peock S, Frost D, et al. Cancer risks for BRCA1 and BRCA2 mutation carriers: results from prospective analysis of EMBRACE. J Natl Cancer Inst. 2013;105:812–22.
Antoniou AC, Cunningham AP, Peto J, et al. The BOADICEA model of genetic susceptibility to breast and ovarian cancers: updates and extensions. Br J Cancer. 2008;98:1457–66.
Easton DF, Steele L, Fields P, et al. Cancer risks in two large breast cancer families linked to BRCA2 on chromosome 13q12-13. Am J Hum Genet. 1997;61:120–8.
van Asperen CJ, Brohet RM, Meijers-Heijboer EJ, et al. Cancer risks in BRCA2 families: estimates for sites other than breast and ovary. J Med Genet. 2005;42:711–9.
Lindor NM, Goldgar DE, Tavtigian SV, Plon SE, Couch FJ. BRCA1/2 sequence variants of uncertain significance: a primer for providers to assist in discussions and in medical management. Oncologist. 2013;18:518–24.
Ledermann J, Harter P, Gourley C, et al. Olaparib maintenance therapy in patients with platinum-sensitive relapsed serous ovarian cancer: a preplanned retrospective analysis of outcomes by BRCA status in a randomised phase 2 trial. Lancet Oncol. 2014;15:852–61.
Eccles DM, Mitchell G, Monteiro AN, et al. BRCA1 and BRCA2 genetic testing-pitfalls and recommendations for managing variants of uncertain clinical significance. Ann Oncol. 2015;26:2057–65.
Eggington JM, Bowles KR, Moyes K, et al. A comprehensive laboratory-based program for classification of variants of uncertain significance in hereditary cancer genes. Clin Genet. 2014;86:229–37.
Lindor NM, Guidugli L, Wang X, et al. A review of a multifactorial probability-based model for classification of BRCA1 and BRCA2 variants of uncertain significance (VUS). Hum Mutat. 2012;33:8–21.
Spurdle AB, Healey S, Devereau A, et al. ENIGMA—evidence-based network for the interpretation of germline mutant alleles: an international initiative to evaluate risk and clinical significance associated with sequence variation in BRCA1 and BRCA2 genes. Hum Mutat. 2012;33:2–7.
NCBI ClinVar database https://www.ncbi.nlm.nih.gov/clinvar/ BRCA2 database https://www.ncbi.nlm.nih.gov/clinvar/?term=brca2%5Bgene%5D. Accessed 28 August 2017.
Plon SE, Eccles DM, Easton D, et al. Sequence variant classification and reporting: recommendations for improving the interpretation of cancer susceptibility genetic test results. Hum Mutat. 2008;29:1282–91.
Iversen ES Jr., Couch FJ, Goldgar DE, Tavtigian SV, Monteiro AN. A computational method to classify variants of uncertain significance using functional assay data with application to BRCA1. Cancer Epidemiol Biomarkers Prev. 2011;20:1078–88.
Guidugli L, Pankratz VS, Singh N, et al. A classification model for BRCA2 DNA binding domain missense variants based on homology-directed repair activity. Cancer Res. 2013;73:265–75.
Thouvenot P, Ben Yamin B, Fourriere L, et al. Functional assessment of genetic variants with outcomes adapted to clinical decision-making. PLoS Genet. 2016;12:e1006096.
Woods NT, Baskin R, Golubeva V, et al. Functional assays provide a robust tool for the clinical annotation of genetic variants of uncertain significance. NPJ Genom Med. 2016;1. https://doi.org/10.1038/npjgenmed.2016.1
Guidugli L, Shimelis H, Masica DL, et al. Assessment of the clinical relevance of BRCA2 missense variants by functional and computational approaches. Am J Hum Genet. 2018;102:233–48.
Fradet-Turcotte A, Sitz J, Grapton D, Orthwein A. BRCA2 functions: from DNA repair to replication fork stabilization. Endocr Relat Cancer. 2016;23:T1–17.
Guidugli L, Carreira A, Caputo SM, et al. Functional assays for analysis of variants of uncertain significance in BRCA2. Hum Mutat. 2014;35:151–64.
Kuznetsov SG, Liu P, Sharan SK. Mouse embryonic stem cell-based functional assay to evaluate mutations in BRCA2. Nat Med. 2008;14:875–81.
Hendriks G, Morolli B, Calleja FM, et al. An efficient pipeline for the generation and functional analysis of human BRCA2 variants of uncertain significance. Hum Mutat. 2014;35:1382–91.
Biswas K, Das R, Eggington JM, et al. Functional evaluation of BRCA2 variants mapping to the PALB2-binding and C-terminal DNA-binding domains using a mouse ES cell-based assay. Hum Mol Genet. 2012;21:3993–4006.
Alamut® Software Suite by Interactive Biosoftware http://www.interactive-biosoftware.com/. AlamutVisual-2.10. Accessed 28 August 2017.
Kass EM, Helgadottir HR, Chen CC, et al. Double-strand break repair by homologous recombination in primary mouse somatic cells requires BRCA1 but not the ATM kinase. Proc Natl Acad Sci USA. 2013;110:5564–9.
Moynahan ME, Pierce AJ, Jasin M. BRCA2 is required for homology-directed repair of chromosomal breaks. Mol Cell. 2001;7:263–72.
Bouwman P, van der Gulden H, van der Heijden I, et al. A high-throughput functional complementation assay for classification of BRCA1 missense variants. Cancer Discov. 2013;3:1142–55.
Ritz C, Baty F, Streibig JC, Gerhard D. Dose-response analysis using R. PLoS ONE. 2015;10:e0146021.
Escribano-Diaz C, Orthwein A, Fradet-Turcotte A, et al. A cell cycle-dependent regulatory circuit composed of 53BP1-RIF1 and BRCA1-CtIP controls DNA repair pathway choice. Mol Cell. 2013;49:872–83.
Ira G, Pellicioli A, Balijja A, et al. DNA end resection, homologous recombination and DNA damage checkpoint activation require CDK1. Nature. 2004;431:1011–7.
Hakem R, de la Pompa JL, Mak TW. Developmental studies of Brca1 and Brca2 knock-out mice. J Mammary Gland Biol Neoplasia. 1998;3:431–45.
Sharan SK, Morimatsu M, Albrecht U, et al. Embryonic lethality and radiation hypersensitivity mediated by Rad51 in mice lacking Brca2. Nature. 1997;386:804–10.
Shimelis H, Mesman RLS, Von Nicolai C, et al. BRCA2 hypomorphic missense variants confer moderate risks of breast cancer. Cancer Res. 2017;77:2789–99.
Lord CJ, Ashworth A. BRCAness revisited. Nat Rev Cancer. 2016;16:110–20.
Livraghi L, Garber JE. PARP inhibitors in the management of breast cancer: current data and future prospects. BMC Med. 2015;13:188.
Cerrato A, Morra F, Celetti A. Use of poly ADP-ribose polymerase [PARP] inhibitors in cancer cells bearing DDR defects: the rationale for their inclusion in the clinic. J Exp Clin Cancer Res. 2016;35:179.
Yuan Z, Chen J, Li W, et al. PARP inhibitors as antitumor agents: a patent update (2013–2015). Expert Opin Ther Pat. 2017;27:363–82.
Xia B, Sheng Q, Nakanishi K, et al. Control of BRCA2 cellular and clinical functions by a nuclear partner, PALB2. Mol Cell. 2006;22:719–29.
Venkitaraman AR. Cancer susceptibility and the functions of BRCA1 and BRCA2. Cell. 2002;108:171–82.
Roy R, Chun J, Powell SN. BRCA1 and BRCA2: different roles in a common pathway of genome protection. Nat Rev Cancer. 2011;12:68–78.
Gudmundsdottir K, Ashworth A. The roles of BRCA1 and BRCA2 and associated proteins in the maintenance of genomic stability. Oncogene. 2006;25:5864–74.
Acknowledgements
The authors thank J. Jonkers and P. Bouwman (Netherlands Cancer Institute, Amsterdam, the Netherlands) for the I-SceI-mCherry plasmid; S.K. Sharan (National Cancer Institute at Frederick, Frederick, MD, USA) for the Pl2F7 conditional Brca2 knockout mES cell line;20 M. Jasin (Memorial Sloan-Kettering Cancer Center, New York, USA) for the DR-GFP reporter plasmid.25 The Dutch Clinical Genetic Centers are acknowledged for submission of VUS for functional analysis. The Dutch/Belgium VUS workgroup and the ENIGMA members are acknowledged for discussions regarding implementation of functional assays into clinical decision making. This work was financially supported by the Dutch Cancer Society KWF (UL2012-5649).
Author contributions
MV and HV designed and supervised the project. RM, FC, and BMo were responsible for data acquisition and RM analysed the results. HV, MV, PD, CA, and RM were involved in data interpretation. BMi performed the bioinformatics analysis. RM, MV, and HV wrote the manuscript and CA, GH, and PD revised the manuscript. All authors read and approved the final manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
DISCLOSURE
The authors declare no conflicts of interest.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, and provide a link to the Creative Commons license. You do not have permission under this license to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Mesman, R.L.S., Calléja, F.M.G.R., Hendriks, G. et al. The functional impact of variants of uncertain significance in BRCA2. Genet Med 21, 293–302 (2019). https://doi.org/10.1038/s41436-018-0052-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41436-018-0052-2
Keywords
This article is cited by
-
EMQN best practice guidelines for genetic testing in hereditary breast and ovarian cancer
European Journal of Human Genetics (2024)
-
Molecular profiling of BRCA1 and BRCA2 genes in Turkish patients with early-onset breast cancer
Egyptian Journal of Medical Human Genetics (2023)
-
Characterization of BRCA2 R3052Q variant in mice supports its functional impact as a low-risk variant
Cell Death & Disease (2023)
-
How does re-classification of variants of unknown significance (VUS) impact the management of patients at risk for hereditary breast cancer?
BMC Medical Genomics (2022)
-
Olaparib for Chinese metastatic castration-resistant prostate cancer: A real-world study of efficacy and gene predictive analysis
Medical Oncology (2022)