Abstract
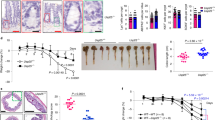
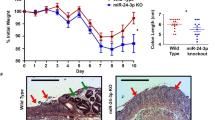
Citrobacter rodentium is a murine pathogen causing transmissible colonic hyperplasia and colitis with a pathogenic mechanism similar to foodborne enterohaemorrhagic Escherichia coli in humans. Mechanisms underlying intestinal responses to C. rodentium infection are incompletely understood. We identified 24 colonic microRNAs (miRNAs) as significantly deregulated in response to C. rodentium, including miR-7a, -17, -19a, -20a, -20b, -92a, -106a, -132, -200a, and -2137; most of these miRNAs belong to the oncogenic miR-17-92 clusters. Pathways involved in cell cycle, cancers, and immune responses were enriched among the predicted targets of these miRNAs. We further demonstrated that an apoptosis facilitator, Bim, is a candidate gene target of miRNA-mediated host response to the infection. These findings suggest that host miRNAs participate in C. rodentium pathogenesis and may represent novel treatment targets.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 digital issues and online access to articles
$119.00 per year
only $19.83 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Mayer CL, Leibowitz CS, Kurosawa S, Stearns-Kurosawa DJ. Shiga toxins and the pathophysiology of hemolytic uremic syndrome in humans and animals. Toxins. 2012;4:1261–87.
Collins JW, Keeney KM, Crepin VF, Rathinam VA, Fitzgerald KA, Finlay BB, et al. Citrobacter rodentium: infection, inflammation and the microbiota. Nat Rev Microbiol. 2014;12:612–23.
Borenshtein D, Fry RC, Groff EB, Nambiar PR, Carey VJ, Fox JG, et al. Diarrhea as a cause of mortality in a mouse model of infectious colitis. Genome Biol. 2008;9:R122.
Friedman RC, Farh KK, Burge CB, Bartel DP. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 2009;19:92–105.
Archambaud C, Sismeiro O, Toedling J, Soubigou G, Becavin C, Lechat P, et al. The intestinal microbiota interferes with the microRNA response upon oral Listeria infection. mBio. 2013;4:e00707–13.
Schulte LN, Eulalio A, Mollenkopf HJ, Reinhardt R, Vogel J. Analysis of the host microRNA response to Salmonella uncovers the control of major cytokines by the let-7 family. EMBO J. 2011;30:1977–89.
Kaakoush NO, Deshpande NP, Man SM, Burgos-Portugal JA, Khattak FA, Raftery MJ, et al. Transcriptomic and proteomic analyses reveal key innate immune signatures in the host response to the gastrointestinal pathogen Campylobacter concisus. Infect Immun. 2015;83:832–45.
Chandrakesan P, Jakkula LU, Ahmed I, Roy B, Anant S, Umar S. Differential effects of beta-catenin and NF-kappaB interplay in the regulation of cell proliferation, inflammation and tumorigenesis in response to bacterial infection. PLoS ONE. 2013;8:e79432.
Roy BC, Subramaniam D, Ahmed I, Jala VR, Hester CM, Greiner KA, et al. Role of bacterial infection in the epigenetic regulation of Wnt antagonist WIF1 by PRC2 protein EZH2. Oncogene. 2015;34:4519–30.
Brabletz S, Brabletz T. The ZEB/miR-200 feedback loop--a motor of cellular plasticity in development and cancer? EMBO Rep. 2010;11:670–7.
Matsushima K, Isomoto H, Inoue N, Nakayama T, Hayashi T, Nakayama M, et al. MicroRNA signatures in Helicobacter pylori-infected gastric mucosa. Int J Cancer. 2011;128:361–70.
Zhang T, Yu J, Zhang Y, Li L, Chen Y, Li D, et al. Salmonella enterica serovar enteritidis modulates intestinal epithelial miR-128 levels to decrease macrophage recruitment via macrophage colony-stimulating factor. J Infect Dis. 2014;209:2000–11.
Li L, Shi QG, Lin F, Liang YG, Sun LJ, Mu JS, et al. Cytokine IL-6 is required in Citrobacter rodentium infection-induced intestinal Th17 responses and promotes IL-22 expression in inflammatory bowel disease. Mol Med Rep. 2014;9:831–6.
Clare S, John V, Walker AW, Hill JL, Abreu-Goodger C, Hale C, et al. Enhanced susceptibility to Citrobacter rodentium infection in microRNA-155-deficient mice. Infect Immun. 2013;81:723–32.
He L, Thomson JM, Hemann MT, Hernando-Monge E, Mu D, Goodson S, et al. A microRNA polycistron as a potential human oncogene. Nature. 2005;435:828–33.
Li Y, Lauriola M, Kim D, Francesconi M, D’Uva G, Shibata D, et al. Adenomatous polyposis coli (APC) regulates miR17-92 cluster through beta-catenin pathway in colorectal cancer. Oncogene. 2016;35:4558–68.
Tsuchida A, Ohno S, Wu W, Borjigin N, Fujita K, Aoki T, et al. miR-92 is a key oncogenic component of the miR-17-92 cluster in colon cancer. Cancer Sci. 2011;102:2264–71.
Zhang Z, Wang M, Eisel F, Tchatalbachev S, Chakraborty T, Meinhardt A, et al. Uropathogenic Escherichia coli epigenetically manipulate host cell death pathways. J Infect Dis. 2016;213:1198–207.
Liu SQ, Jiang S, Li C, Zhang B, Li QJ. miR-17-92 cluster targets phosphatase and tensin homology and Ikaros Family Zinc Finger 4 to promote TH17-mediated inflammation. J Biol Chem. 2014;289:12446–56.
Chen WX, Ren LH, Shi RH. Implication of miRNAs for inflammatory bowel disease treatment: systematic review. World J Gastrointest Pathophysiol. 2014;5:63–70.
Kim HY, Kwon HY, Ha Thi HT, Lee HJ, Kim GI, Hahm KB, et al. MicroRNA-132 and microRNA-223 control positive feedback circuit by regulating FOXO3a in inflammatory bowel disease. J Gastroenterol Hepatol. 2016;31:1727–35.
Rodrigues DM, Sousa AJ, Johnson-Henry KC, Sherman PM, Gareau MG. Probiotics are effective for the prevention and treatment of citrobacter rodentium-induced colitis in mice. J Infect Dis. 2012;206:99–109.
Bhinder G, Sham HP, Chan JM, Morampudi V, Jacobson K, Vallance BA. The Citrobacter rodentium mouse model: studying pathogen and host contributions to infectious colitis. J Vis Exp. 2013;72:e50222.
Glenn AJ, Fielding KA, Chen J, Comelli EM, Ward WE. Long-term vitamin D3 supplementation does not prevent colonic inflammation or modulate bone health in IL-10 knockout mice at young adulthood. Nutrients. 2014;6:3847–62.
Wine E, Shen-Tu G, Gareau MG, Goldberg HA, Licht C, Ngan B-Y, et al. Osteopontin mediates Citrobacter rodentium-induced colonic epithelial cell hyperplasia and attaching-effacing lesions. Am J Pathol. 2010;177:1320–32.
Kertesz M, Iovino N, Unnerstall U, Gaul U, Segal E. The role of site accessibility in microRNA target recognition. Nat Genet. 2007;39:1278–84.
Rahmati S, Abovsky M, Pastrello C, Jurisica I. pathDIP: an annotated resource for known and predicted human gene-pathway associations and pathway enrichment analysis. Nucleic Acids Res. 2017;45(D1):D419–d426.
Brown KR, Otasek D, Ali M, McGuffin MJ, Xie W, Devani B, et al. NAViGaTOR: network analysis, visualization and graphing toronto. Bioinformatics. 2009;25:3327–9.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-delta delta C(T)) method. Methods. 2001;25:402–8.
Acknowledgements
This work was supported by the Natural Sciences and Engineering Research Council of Canada (NSERC #356124) and the J. P. Bickell Foundation. Computational analysis was supported in part by NSERC (#203475), Ontario Research Fund (GL2-01-030), Canada Foundation for Innovation (CFI #225404, #30865), Canada Research Chair Program (CRC #225404), Ontario Research Fund (RE-03-020), and IBM. Elena Comelli holds the Lawson Family Chair in Microbiome Nutrition Research at the University of Toronto. Bijun Wen was partially supported by NSERC Alexander Graham Bell Canada Graduate Scholarship. NanoString service was provided by the Princess Margaret Genomics Centre, Toronto, Canada (www.pmgenomics.ca). The authors would like to thank the Division of Comparative Medicine staff at University of Toronto for help with animal care.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Wen, B., Tokar, T., Taibi, A. et al. Citrobacter rodentium alters the mouse colonic miRNome. Genes Immun 20, 207–213 (2019). https://doi.org/10.1038/s41435-018-0026-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41435-018-0026-z
This article is cited by
-
Microbiota medicine: towards clinical revolution
Journal of Translational Medicine (2022)