Abstract
The functional and metabolic characteristics of specific adipose tissue (AT) depots seem to be determined by intrinsic mechanisms. We performed a comprehensive transcriptome profiling of human AT from distinct fat depots to unravel their unique features potentially explaining molecular mechanisms underlying AT distribution and their contribution to health and disease. Post-mortem AT samples of five body donors from 15 anatomical locations were collected. Global mRNA expression was measured by Illumina® Human HT-12 v4 Expression BeadChips. Data were validated using qPCR and Western Blot in a subset of ATs from seven additional body donors. Buccal and heel AT clearly separated from the “classical” subcutaneous AT depots, and perirenal and epicardial AT were distinct from visceral depots. Gene-set enrichment analyses pointed to an inflammatory environment and insulin resistance particularly in the carotid sheath AT depot. Moreover, the epicardial fat transcriptome was enriched for genes involved in extracellular matrix remodeling, inflammation, immune signaling, coagulation, thrombosis, beigeing, and apoptosis. Interestingly, a striking downregulation of the expression of leptin receptor was found in AT from heel compared with all other AT depots. The distinct gene expression patterns are likely to define fat depot specific AT functions in metabolism, energy storage, immunity, body insulation or as cushions. Improved knowledge of the gene expression profiles of various fat depots may strongly benefit studies aimed at better understanding of the genetics and the pathophysiology of obesity and adverse body fat composition.
Similar content being viewed by others
Introduction
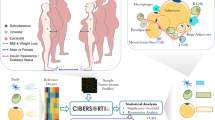
Adipose tissue (AT) is mainly composed of adipocytes and adipocyte precursor cells but also cells of connective tissue, vasculature, cells of the immune system, and nerve cells [1]. It does not only serve to store energy (mainly in white adipose tissue [WAT]) and to regulate body temperature, energy, and glucose balance (e.g., in brown adipose tissue [BAT], “beige” adipocytes in WAT) but also has mechanical functions (hand, heel, and renal capsule). Over the last decades AT has been treated as an endocrine organ [2, 3] supposed to secrete hundreds of bioactive factors contributing for example to the regulation of hunger and satiety, insulin sensitivity, inflammation, or ectopic lipid storage [4]. Fat distribution is controlled by genetic factors as suggested by numerous heritability studies [5]. However, the specific molecular mechanisms underlying AT distribution are still poorly understood. Extreme phenotypes of body fat distribution, i.e., steatopygia, lipodystrophies, or lipomatosis, are good models to address the genetic background of body fat distribution. In conditions of adverse AT distribution such as in lipodystrophies (e.g., familial partial lipodystrophy Dunnigan type (FPLD2, OMIM151660); congenital generalized lipodystrophy) certain AT depots are missing while others remain unaffected or even accumulate more fat [6]. For example in subjects with lipodystrophy type Dunnigan, the heel fat pad is the only location not affected by the extreme loss of subcutaneous adipose tissue (ScAT) which is observed at the limbs suggesting different pathways of lipid accumulation (personal observation, Fig. 1). Adipose progenitor cells and preadipocytes in different AT locations may derive from different cell lineages that determine their capacity for proliferation and differentiation and are influenced by the cell (micro-)environment, e.g., sex hormones or glucocorticoids, and the cellular composition of the depot as well as the extracellular matrix [7, 8]. As a result, rates of hypertrophy and hyperplasia during growth and remodeling vary among fat depots [7]. Lineage tracing studies raise evidence that white adipocytes may derive from different precursors such as mesenchymal progenitors (lateral plate mesoderm) and neural crest stem cells (ectoderm, suggested to be a source of ScAT) [9,10,11,12,13]. This is supported by transcriptional profiling studies that revealed characteristic expression pattern of developmental genes for subcutaneous and visceral WAT [7, 14, 15].
However, majority of these studies are restricted to the comparison of human AT depots which are accessible during surgery or biopsy in a defined body compartment. Body donors help to overcome this obstacle by providing a unique possibility to get access to adipose depots, which are regularly not available. In this study, we performed a comprehensive transcriptome profiling of human AT from 15 AT locations dispersed throughout the human body to unravel their unique features potentially explaining molecular mechanisms underlying AT distribution and their contribution to health and disease.
Subjects, methods, data processing and statistical analysis
Subjects
Human AT samples from 15 different fat depots (Fig. 2) were obtained from five body donors of the Institute of Anatomy at Leipzig University. Epicardial AT was available only for three body donors, and transverse colon was missing for one body donor. In total, 72 AT samples were initially available. During their lifetime, body donors gave their informed and written consent to the donation of their bodies for teaching and research purposes. As part of the body donor program regulated by the Saxonian Death and Funeral Act of 1994 (third section, paragraph 18 item 8), institutional approval for the use of the post-mortem tissues of human body donors was obtained from the Institute of Anatomy at Leipzig University. Samples were either snap frozen in liquid nitrogen for protein or RNA analyses or immediately fixed in 4% paraformaldehyde (PFA) and embedded in paraffin for histological evaluation. Additional characteristics including age, causes of death and post-mortem delay (pmd) are given in Table 1. For validation of microarrays RNA was extracted from seven further body donors (subcutaneous depots (periumbilical region, upper leg, and upper arm) as well as from heel fat pad; Table 1).
Tissue slices of 7 µm were stained using antibody against the adipocyte specific marker perilipin (green), the macrophage marker AIF1 (red), and the nuclei were stained with DAPI (blue). Scale bar in the heel image represents 50 µm. For a larger view of the tissue slices see Supplemental Fig. 1). MR imaging was done with a male volunteer and written consent for publication of MR images was obtained. Mean adipocyte diameters ± SD (min/max) were calculated from measures of four body donors (1–3, and 6) with a post-mortem delay ≤12 h.
Methods
Immunofluorescence staining of AT sections, measurement of adipocyte size, and Western Blot analysis
Immunofluorescence staining was done as previously described [16]. Mean adipocyte diameter has been assessed on H&E stained paraffin sections as described previously for the ATs of four body donors (donor 1–3, and 6; Table 1) [16, 17]. Western blot analysis was performed as described previously (Supplemental Material) [18].
RNA extraction and cDNA synthesis
AT samples (100–130 mg) were homogenized in ball mill tubes (ceramic balls, 1.4 mm) in 1 ml Trizol® (Invitrogen-TFS, Germany; Supplemental Material). RNA was extracted using the InviTrap® Spin Tissue RNA Mini Kit (Stratec Molecular GmbH, Berlin, Germany) according to the manufacturer’s protocol “RNA “clean up” from Trizol® aqueous phase”. RNA integrity (RIN) and concentration were examined using an Agilent 2100 Bioanalyzer (Agilent, Santa Clara, California, USA). 1 μg of RNA per sample was reverse transcribed (Promega, Mannheim, Germany).
Quantitative real-time RT-PCR analysis
For quantification of mRNA expression of the leptin receptor, quantitative real-time PCR was performed. Two different types of the leptin receptor were quantified: LEPR common (OB-R), detecting all forms of the leptin receptor, and LEPR full length (OB-Rfl), detecting only the long form (for primer sequences see Supplemental Material). The mRNA expression of the LEPR was determined relative to the 18S ribosomal RNA (18s) reference gene.
Gene expression analyses using gene chips
mRNA expression data were obtained by Human HT-12 v4 Expression BeadChips (Illumina®, San Diego, California, USA). For the first three body donors tissues were processed in duplicates (excluding the ascending colon AT sample of body donor 1 due to the lowest RIN), and for upper leg each in quadruplicate. AT samples from body donor 4 and 5 were processed once.
Data processing
Preprocessing
Raw data of 47,323 gene-expression probes and 887 control probes were extracted by Illumina GenomeStudio without additional background correction. The data were further processed within R/Bioconductor R [19]. Initially, six samples were excluded: (i) three samples (2.5%) because of low RNA quality, and (ii) three samples (2.5%) having an extreme number of expressed genes (defined as median ± 4 × interquartile ranges (IQR) of the cohort’s values). Transcripts not expressed at p = 0.05 (as defined by Illumina and implemented in the R/Bioconductor package lumi [20]) in at least 5% of all samples per tissue location were excluded from further analysis. 37,928 (80.1%) probes remained in the analysis in all subgroups. Expression values were quantile-normalized and log2-transformed [21]. For an extended description please see the Supplemental Material.
Outlier detection and batch-correction
We adapted an approach from Oldham et al. [22]. This was done separately for all subgroups and resulted in 11 outlier samples (9.1%). We used a linear approach for batch-correction. We found the pmd and the Sentrix barcode as best fitting batch effect model and corrected for these effects. For further outlier detection, we calculated the Euclidian distance between all samples of the same subgroup and the group-center. This was defined as the average of samples after removing 10% of samples manifesting largest distances from the group-center done separately for each subgroup (implemented in the R/Bioconductor package lumi [20]). The threshold for outlier detection was defined as two times the median distance to the center.
Gene mapping
Mapping of genes corresponding to expression probes and assignment of gene names was done using information of a remapping approach [23] applying gene-information of the NCBI database. This information was retrieved using the R add-on package from Bioconductor illuminaHumanv4.db_1.26.0 that relates to NCBI data dated on 2015-March-17. This resulted in a total of 28,430 valid gene-expression probes (=variables) corresponding to 20,213 unique genes available in all subgroups.
Visualization
We used the Qlucore Omics Explorer (QOE, Qlucore, Lund, Sweden) for visualization of the data and the statistical analyses. Predicted loci were checked in the actual version of the NCBI database [GRCh38, last access May 2019] and if available, the gene was assigned or the probe was withdrawn from the list count.
Data statement
The dataset used for the analyses with Qlucore was submitted to figshare (https://figshare.com/about) and is accessible via the DOI 10.6084/m9.figshare.11836032 or the following link:
Statistical analysis
Mean values were calculated for duplicates/quadruplicates. Finally, 54 samples were included in the analysis (N for each depot see Supplemental Material). Sigma-normalization was applied to the dataset. The dataset was not pre-collapsed due to the assignment of the same gene symbol to different transcripts represented by different probes. Principal component analyses and hierarchical clustering were performed to investigate the structure of the data. Multi-group and two-group comparisons between the AT depots were performed using the F-test (ANOVA) and Student’s t test (two-sided), respectively. The analyses were adjusted for age and gender. A p value <1.1 × 10−6 was defined to be significant. The DAVID 6.8 Functional Annotation Tool (https://david.ncifcrf.gov/) and the Gene Ontology enRIchment anaLysis and visuaLizAtion tool “GOrilla” (http://cbl-gorilla.cs.technion.ac.il/) were used for enrichment analysis and visualization [24, 25].
Results
Adipocyte size, viability of AT from body donors, and evaluation of RNA quality for microarray analysis
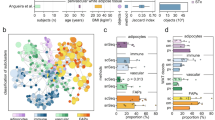
Adipocyte diameters were distributed according to the BMI of the donors (Fig. 3a). Interestingly, adipocytes from VAT and ScAT omentum majus show lower adipocyte diameters in comparison to all other investigated visceral AT (mesentery, colon AT samples) and ScAT depots respectively (Fig. 3b). AT sections of all 15 studied fat depots revealed a high degree of perilipin expressing adipocytes indicating high tissue viability (Fig. 2, Supplemental Fig. 1). AIF1 expressing AT macrophages were also present in most of the sections, but characteristic crown-like structures (typically found around dying adipocytes) were lacking (Fig. 2, Supplemental Fig. 1). We observed a broad spectrum of RINs for the tissues (RINs ranging from 2.2 to 8.8, median 6; Fig. 3c). After preprocessing of the data the following tissues were included in the analysis: all depots for body donors 1–3 except for VAT ascending colon of body donor 1; carotid sheath, VAT mesentery, perirenal AT, ScAT abdomen of body donor 4; and perirenal AT, ScAT abdomen, ScAT upper arm, ScAT upper leg, supraclavicular AT and buccal AT from body donor 5. In addition, the analysis of the mRNA expression of markers of autolysis (FASL, PTEN, CAPN1, and CASP3) did not reveal significant differences between the tissues (Fig. 3d).
a Mean adipocyte diameter in µm for four body donors with a pmd ≤ 12 h. b Mean adipocyte diameter in % of mean per individual. c RNA Integrity Number (RIN) plotted against post-mortem delay (pmd) for each body donor sampled on microarrays. RNA quality decreases with increasing pmd. Each column represents one body donor. d Marker for autolysis. Given are the mRNA expression values (log2 transformed) for each sample represented by a dot. The continuous line represents the mean overall samples. The data are adjusted for post-mortem delay, Sentrix barcode, age and sex. FASL-Fas ligand, PTEN-phosphatase and tensin homolog, CAPN1-calpain 1, CASP3-caspase 3.
Characterization of AT depot-specific gene expression signatures
To explore the overall gene expression differences between the 15 different AT locations, the preprocessed dataset was analyzed with F-test (ANOVA) using the location of the tissue samples as grouping variable. At p < 1.1 × 10−6 68% of the variance were explained by the first principal component with 165 variables (which can be collapsed to 132 validated genes) remaining in the analysis (Fig. 4a; Supplementary document – Table 1). ATs from the body cavities (except for kidney) present a clearly distinct expression pattern as compared with the AT from all other locations (Fig. 4b). Of note, the epicardial AT clusters with the visceral AT depots from the abdomen and the kidney AT clusters with the ScAT samples. Whereas the body cavity depots, except for kidney, manifest a homogeneous expression pattern, the expression pattern is more diverse in subcutaneous depots with distinct signatures for buccal, heel, and carotid sheath AT. In a separate analysis of gene expression of selected genes in tissues of two body donors with a pmd of 10 h, buccal, heel, and supraclavicular AT displayed higher expression of genes involved in adipogenesis and lipolysis compared with the other tissues (Supplementary Fig. 2). The samples of one tissue location were compared against all other tissue locations grouped together to look for gene sets specific for this fat tissue (differential expressed transcripts with p ≤ 0.001). We identified significant GO terms for most of the subcutaneous locations and the VAT omentum majus as well as VAT mesentery area suggesting that the expression pattern of the AT depot reflect their location (Supplementary Fig. 3; “Gene Lists for GOrilla” in the Variable_Lists_collection_V1-6_Supplemental document_1). However, one need to keep in mind that in the respective analyses only a maximum of five samples were compared against all other samples (N = 49) which clearly limited their statistical power. In addition, we investigated the AT depots of the body donors for signs of beiging/browning (Supplementary Results, Supplementary Fig. 4).
Discriminant analysis for identification of major determinants of variability
To identify major determinants of variability, AT depots (sub-locations) were organized in groups based on the anatomical location (Fig. 2) and based on the results of the multigroup comparison.
Comparison of visceral- vs. subcutaneous-like AT depots
The depots from the body cavities (VAT samples, epicardial AT) were compared with the “subcutaneous-like” depots (ScAT samples, in addition AT from buccal, heel, interscapular, carotid sheath, and including the kidney samples) in a two-group comparison. At p < 1.1 × 10−6 347 variables separate both groups corresponding to 275 genomic loci (collapsed for gene ID as a unique identifier, including confirmed predicted annotations, excluding non-specified variables; Supplementary document - Table 2). Twenty-one genes were higher expressed in the subcutaneous-like tissues, including developmental genes such as IRX3, TBX15, PRRX1, or SIX1 (Supplementary document - Table 2). Among the 271 transcripts (corresponding to 250 genes) found to be higher expressed in the visceral depots are mostly represented by genes that are described to play a role in the insulin pathway, in adipogenesis, and in obesity-associated inflammation (Supplementary document – Table 2). Mapping by DAVID indicated enrichment of genes involved in organ and tissue morphogenesis (Gene Ontology term for biological processes (BP)_FAT GO:0009887, GO:0048729; p < 1.9 × 10−6, False Discovery Rate (FDR) < 0.005; Supplementary document – Table 2.1).
Inter-depot comparison: visceral depots
In the next step, we compared gene expression patterns of different AT sub-depots of the intra-abdominal and intra-thoracic cavities (epicardial, ascending colon, descending colon, transverse colon, mesentery, and omentum majus).
At p < 1.1 × 10−6 only angiotensin II receptor type 1 (AGTR1) gene expression was lower in the VAT omentum majus samples compared with the other locations. Applying a p value of <0.001 and a FDR-threshold of 20% resulted in five loci of which four have been assigned to a gene (Fig. 5a). Interestingly, perirenal and epicardial AT are clearly distinct from visceral depot samples characterized e.g., by the expression of HOXC9 in perirenal, and GATA4 in epicardial AT (Fig. 5b, Supplementary document – Table 3).
Hierarchical clustered heat maps of (a) samples derived from the body cavity. F-test, p < 0.001 and FDR-threshold 0.2. b Samples from the body cavity including kidney. F-test, p < 0.001 and FDR-threshold 0.2. c Samples from all subcutaneous locations, and buccal, heel, carotis sheath and kidney. F-test, p < 1.1 × 10−6. For further data please see Supplementary document Table 3.
Inter-depot comparison: subcutaneous depots and perirenal AT
Comparisons of the subcutaneous ATs from upper leg, upper arm, abdomen, interscapular, and supraclavicular as well as the locations from buccal, heel, carotid sheath, and also kidney revealed that AT gene expression signatures from buccal, heel, and carotid sheath deviated significantly (p < 1.1 × 10−6) from all other ScAT depots (Fig. 5c, Supplementary document – Table 3). The tissues are characterized by higher expression of homeobox genes (Fig. 5c). After excluding buccal, heel, and carotid sheath AT from the analysis, remaining ScATs and perirenal AT were not significantly different in gene expression at genome-wide significance level. However, at p < 0.001 and FDR ≤ 20% 93 variables corresponding to 74 genes cluster the samples, separating kidney, interscapular, and supraclavicular AT from the ScAT depots of upper arm, upper leg, and abdomen (Supplementary document – Table 4).
Comparative analysis of AT depots from buccal, heel, carotid sheath, and epicardial AT
Subcutaneous depots from abdomen and extremities were merged to a group “subcutaneous”. To identify markers distinguishing the AT depot from buccal, heel, and carotid sheath from the subcutaneous depots two-group comparisons were performed (subcutaneous vs. buccal, heel or carotid sheath, respectively).
First, the subcutaneous depots were compared with the buccal samples. The observed differences, revealed 44 validated transcripts corresponding to 42 genes (p ≤ 7.7 × 10−5, FDR ≤ 5%) of which 22 are higher expressed in the buccal depot (Supplementary document – Table 5). Among these transcripts are homeobox genes and transcription factors that are described to be involved e.g., in tooth morphogenesis and development (LHX8, BARX1, and PAX9). Second, the depot from heel was compared with the subcutaneous samples. Three transcripts are significantly higher expressed in the AT samples from heel: HOXA13, LHX2, and ALX3. Applying a less stringent p value of ≤0.001 resulted in 74 differentially expressed validated transcripts, which correspond to 71 genomic loci (Supplementary document – Table 6) with gene enrichment in embryonic morphogenesis and limb development (Supplementary document – Table 6.1). Interestingly, in the heel depot the LEPR is expressed at very low levels. However, in this dataset FDR was high (33.3%). We confirmed the reduced LEPR expression in heel AT in comparison to subcutaneous abdominal and thigh AT by quantitative real-time PCR (p < 0.0001; <0.05, respectively; Fig. 6a, b).
a LEPR mRNA expression is significantly lower in heel AT compared with other subcutaneous AT locations. Ob-Rfl—LEPR full length, detecting only the long form of the receptor; Ob-R—all forms of the leptin receptor are detected. ****/*p < 0.0001/0.05. b Protein expression: LEPR is expressed in subcutaneous AT but not detectable in heel AT. Multiple bands occur due to different isoforms of the receptor.
Third, gene expression from the fat depot of the carotid sheath was compared with the subcutaneous depots. Three loci were higher expressed (p < 1.1 × 10−6), MAL, NTS, PKIA; and CLTA was lower expressed. Applying a FDR of ≤5% resulted in 266 differential expressed variables (p ≤ 4.65 × 10−4) corresponding to 241 valid transcripts and 234 genomic loci (Supplementary document – Table 7). This depot is characterized by the enrichment of genes involved in biological processes related to the immune system which points to the fact that this tissue may have had exhibited inflammatory milieu in the body donors (Supplementary document – Table 7.1).
Likewise, epicardial AT samples were compared with visceral depots (omentum majus, mesentery, ascending colon, descending colon, and transverse colon) clustered in the term “visceral” (two-group comparison). At p = 1.1 × 10−6 two transcripts were higher expressed in the epicardial depot compared with the visceral samples, COL4A4 and HBM. In a less stringent analysis using a p value of ≤0.001 and a FDR < 20% 99 differential expressed valid transcripts (corresponding to 98 genes) are displayed (Supplementary document – Table 8), which included genes involved in myocardial differentiation and function.
Discussion
Here, we compare AT gene expression signatures originating from 15 distinct fat depots of human body donors. It has been previously shown that the quality of RNA is highly tissue, donor, and pmd dependent [26, 27]. The influence of the pmd was also obvious in our AT samples as we had to exclude some AT samples from two body donors with pmd of 14 and 24 h, respectively. The multigroup comparisons of all 15 depots revealed that the main difference between the locations is derived from samples grouped in the general term “VAT” (including the epicardial AT) vs. those grouped as “ScAT” (including buccal, carotid sheath, heel, kidney, and interscapular AT). Interestingly, the kidney samples are clearly separated from the VAT samples and located in the ScAT group. In line with previous studies [28, 29], depot specific expression of developmental genes like the HOX family was observed in our study as well. Although the well acknowledged depot specificity of certain HOX genes is evident, the HOX expression pattern seems to be highly dependent on the specific tissues clustering and individual pair-wise comparisons (see Supplementary document Tables). Also consistent with previous reports, we confirmed differences between ScAT and VAT in gene expression patterns including not only developmental genes, but also genes potentially contributing to aberrant adipocyte function during obesity such as SIX1, which binds to adipogenic and brown marker genes and interacts with C/EBPα, C/EBPβ, and EBF2 [30].
Facets of epicardial and perirenal AT
There is strong evidence that epicardial AT plays an important role in heart disease and that there is a cross-talk between epicardial fat and the heart [31]. The epicardial fat transcriptome shows an enrichment of genes involved in extracellular matrix remodeling, inflammation, immune signaling, coagulation, thrombosis, beigeing, and apoptosis (compared with ScAT) [32]. Consistently, in epicardial AT we found significantly higher expression of genes representing these pathways (e.g., COL4A4, HBM, GATA4, amd CD68). Regulation of angiogenesis (GO:0045765) and regulation of vasculature development (GO: 901342) turned out to be the two major GO-terms. Comparing the epicardial AT mRNA expression with ScAT as previously reported by Gaborit et al. [32], we confirmed omentin (ITLN1) to be the strongest differentially expressed gene in the dataset (p = 9.78 × 10−9; data not shown). Omentin is an adipokine known to exert beneficial effects on metabolism [33] and to play a role in the pathogenesis of coronary artery disease [34].
Perirenal fat was described to be a predictor of early kidney damage in non-hypertensive and non-diabetic obese patients [35] and perirenal fat invasion seems to be a critical factor in renal cell carcinoma [36]. In our dataset, the kidney fat appears more distinguishable from the VAT than ScAT. Although the expression pattern in perirenal AT is more similar to the pattern in ScAT, it can still be separated according to differences in the expression of genes such as ankyrin repeat domain 20 family member A1 (ANKRD20A1), RNA binding motif single stranded interacting protein 1 (RBMS1), and DnaJ heat shock protein family (Hsp40) member B1 (DNAJB1) (data not shown). Of note, we did not detect any features of browning in perirenal AT samples but this might be due to the high individual variability in the tissue as described by Svensson et al. [37].
Facets of buccal, heel, interscapular, supraclavicular, and carotid sheath AT
The AT depots of buccal and heel were clearly separated from other the ScAT depots. Buccal AT is not only important for mastication and sucking in neonates and as cushion to protect neurovascular bundles but it also plays a role in oncological defects and in aesthetic facial surgery [38]. Interestingly, LHX8, which was previously described as a BAT-selective marker [39], was characteristic for the buccal AT depot. However, histologically it was described as pure white fat with less vesicles and smaller mitochondria indicative of a tissue manifesting lower metabolic activity [40]. LHX8 as well as other genes expressed in buccal AT (BARX1, PAX9) have been implicated to play a role in tooth morphogenesis in mice and men [41, 42]. The heel AT has been investigated for fatty acid composition compared with subcutaneous abdominal fat so far [43] but we are not aware of any gene expression study. Here, we found higher mRNA expression of genes that are involved in embryonic and limb morphogenesis and skeletal system development (HOXA13, LHX2, COBL, HOXA11, SALL1, TBX4, ALX3, PITX1, and BMP5). The heel fat pad is for example affected in conditions of rheumatoid arthritis or peripheral neuropathy [44, 45]. In congenital generalized lipodystrophy, mechanical AT depots like the heel fat pad remain unaffected [6] which might be due to a different metabolic program. For instance, the LEPR showed very low expression in heel AT and could not be detected on protein level. The samples from the carotid sheath showed an enriched expression of genes described to play a role in insulin resistance (PPP1R3E, PRKCQ, PRKCZ, CPT1B, MGEA5, RPS6KB2, OGT; KEGG_PATHWAY hsa04931:Insulin resistance). Furthermore, NTS was significantly higher expressed in the carotid sheath AT samples. NTS is distributed in the central nervous system, is involved in the maintenance of gut structure and function, and in the regulation of fat metabolism [46]. Besides the suggested involvement in cancer [47] NTS was recently described to play a role in diet-induced obesity [48]. In the Framingham Heart Study, proneurotensin concentrations were positively associated with the risk of incident cardiovascular events [49]. Neurotensin seems to affect the coronary vascular tone in the rat, guinea-pig and dog, and the gastric mucosal blood flow in humans [50] and the same might be true for the carotid artery.
Study limitations
We have to acknowledge several limitations of the study: the sample size is small, whole AT samples including several cell types have been used for the analysis of the transcriptome, the age range of the donors was between 64 and 98, and comorbidities and causes of death differed between the subjects, which might have influenced the expression profiles. Nevertheless, studies offering the possibility of intra-individual comparisons of 15 AT depots derived from the same body donor are lacking, which makes the here presented design unique.
Conclusion
The expression pattern of the AT depots strongly reflects their location. The study provides a comprehensive view on the human AT depots to improve our knowledge of the gene expression profiles of various fat depots, which may strongly benefit studies aimed at better understanding of the genetics and the pathophysiology of adverse body fat composition.
References
Compher C, Badellino KO. Obesity and inflammation: lessons from bariatric surgery. J Parenter Enter Nutr. 2008;32:645–7.
Frühbeck G, Gómez-Ambrosi J, Muruzábal FJ, Burrell MA. The adipocyte: a model for integration of endocrine and metabolic signaling in energy metabolism regulation. Am J Physiol Endocrinol Metab. 2001;280:E827–47.
Villarroya F, Cereijo R, Villarroya J, Giralt M. Brown adipose tissue as a secretory organ. Nat Rev Endocrinol. 2017;13:26–35.
Lehr S, Hartwig S, Sell H. Adipokines: a treasure trove for the discovery of biomarkers for metabolic disorders. Proteom Clin Appl. 2012;6:91–101.
Schleinitz D, Böttcher Y, Blüher M, Kovacs P. The genetics of fat distribution. Diabetologia 2014;57:1276–86.
Garg A, Agarwal AK. Lipodystrophies: disorders of adipose tissue biology. Biochim Biophys Acta. 2009;1791:507–13.
Fried SK, Lee MJ, Karastergiou K. Shaping fat distribution: new insights into the molecular determinants of depot- and sex-dependent adipose biology. Obes (Silver Spring). 2015;23:1345–52.
Cawthorn WP, Scheller EL, MacDougald OA. Adipose tissue stem cells meet preadipocyte commitment: going back to the future. J Lipid Res. 2012;53:227–46.
Tang W, Zeve D, Suh JM, Bosnakovski D, Kyba M, Hammer RE, et al. White fat progenitor cells reside in the adipose vasculature. Science 2008;322:583–6.
Hudak CS, Gulyaeva O, Wang Y, Park SM, Lee L, Kang C, et al. Pref-1 marks very early mesenchymal precursors required for adipose tissue development and expansion. Cell Rep. 2014;8:678–87.
Chau YY, Bandiera R, Serrels A, Martínez-Estrada OM, Qing W, Lee M, et al. Visceral and subcutaneous fat have different origins and evidence supports a mesothelial source. Nat Cell Biol. 2014;16:367–75.
Billon N, Iannarelli P, Monteiro MC, Glavieux-Pardanaud C, Richardson WD, Kessaris N, et al. The generation of adipocytes by the neural crest. Development 2007;134:2283–92.
Le Lièvre CS, Le Douarin NM. Mesenchymal derivatives of the neural crest: analysis of chimaeric quail and chick embryos. J Embryol Exp Morphol. 1975;34:125–54.
White UA, Tchoukalova YD. Sex dimorphism and depot differences in adipose tissue function. Biochim Biophys Acta. 2014;1842:377–92.
Hilton C, Karpe F, Pinnick KE. Role of developmental transcription factors in white, brown and beige adipose tissues. Biochim Biophys Acta. 2015;1851:686–96.
du Plessis J, van Pelt J, Korf H, Mathieu C, van der Schueren B, Lannoo M, et al. Association of adipose tissue inflammation with histologic severity of nonalcoholic fatty liver disease. Gastroenterology 2015;149:635–.e14.
Braune J, Weyer U, Matz-Soja M, Hobusch C, Kern M, Kunath A, et al. Hedgehog signalling in myeloid cells impacts on body weight, adipose tissue inflammation and glucose metabolism. Diabetologia 2017;60:889–99.
Braune J, Weyer U, Hobusch C, Mauer J, Brüning JC, Bechmann I, et al. IL-6 Regulates M2 polarization and local proliferation of adipose tissue macrophages in obesity. J Immunol. 2017;198:2927–34.
Gentleman RC, Carey VJ, Bates DM, Bolstad B, Dettling M, Dudoit S, et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 2004;5:R80.
Du P, Kibbe WA, Lin SM. lumi: a pipeline for processing Illumina microarray. Bioinformatics 2008;24:1547–8.
Schmid R, Baum P, Ittrich C, Fundel-Clemens K, Huber W, Brors B, et al. Comparison of normalization methods for Illumina BeadChip HumanHT-12 v3. BMC Genom. 2010;11:349.
Oldham MC, Langfelder P, Horvath S. Network methods for describing sample relationships in genomic datasets: application to Huntington’s disease. BMC Syst Biol. 2012;6:63.
Barbosa-Morais NL, Dunning MJ, Samarajiwa SA, Darot JF, Ritchie ME, Lynch AG, et al. A re-annotation pipeline for Illumina BeadArrays: improving the interpretation of gene expression data. Nucleic Acids Res. 2010;38:e17.
Huang da W, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4:44–57.
Eden E, Navon R, Steinfeld I, Lipson D, Yakhini Z. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinforma. 2009;10:48.
Walker DG, Whetzel AM, Serrano G, Sue LI, Lue LF, Beach TG. Characterization of RNA isolated from eighteen different human tissues: results from a rapid human autopsy program. Cell Tissue Bank. 2016;17:361–75.
Gupta S, Halushka MK, Hilton GM, Arking DE. Postmortem cardiac tissue maintains gene expression profile even after late harvesting. BMC Genom. 2012;13:26.
Cantile M, Procino A, D’Armiento M, Cindolo L, Cillo C. HOX gene network is involved in the transcriptional regulation of in vivo human adipogenesis. J Cell Physiol. 2003;194:225–36.
Karastergiou K, Fried SK, Xie H, Lee MJ, Divoux A, Rosencrantz MA, et al. Distinct developmental signatures of human abdominal and gluteal subcutaneous adipose tissue depots. J Clin Endocrinol Metab. 2013;98:362–71.
Brunmeir R, Wu J, Peng X, Kim SY, Julien SG, Zhang Q, et al. Comparative transcriptomic and epigenomic analyses reveal new regulators of murine brown adipogenesis. PLoS Genet. 2016;12:e1006474.
Gaborit B, Sengenes C, Ancel P, Jacquier A, Dutour A. Role of epicardial adipose tissue in health and disease: a matter of fat? Compr Physiol. 2017;7:1051–82.
Gaborit B, Venteclef N, Ancel P, Pelloux V, Gariboldi V, Leprince P, et al. Human epicardial adipose tissue has a specific transcriptomic signature depending on its anatomical peri-atrial, peri-ventricular, or peri-coronary location. Cardiovasc Res. 2015;108:62–73.
Watanabe T, Watanabe-Kominato K, Takahashi Y, Kojima M, Watanabe R. Adipose tissue-derived Omentin-1 function and regulation. Compr Physiol. 2017;7:765–81.
Harada K, Shibata R, Ouchi N, Tokuda Y, Funakubo H, Suzuki M, et al. Increased expression of the adipocytokine omentin in the epicardial adipose tissue of coronary artery disease patients. Atherosclerosis 2016;251:299–304.
Sun X, Han F, Miao W, Hou N, Cao Z, Zhang G. Sonographic evaluation of para- and perirenal fat thickness is an independent predictor of early kidney damage in obese patients. Int Urol Nephrol. 2013;45:1589–95.
Jeon HG, Jeong IG, Kwak C, Kim HH, Lee SE, Lee E. Reevaluation of renal cell carcinoma and perirenal fat invasion only. J Urol. 2009;182:2137–43.
Svensson PA, Lindberg K, Hoffmann JM, Taube M, Pereira MJ, Mohsen-Kanson T, et al. Characterization of brown adipose tissue in the human perirenal depot. Obes (Silver Spring). 2014;22:1830–7.
Yousuf S, Tubbs RS, Wartmann CT, Kapos T, Cohen-Gadol AA, Loukas M. A review of the gross anatomy, functions, pathology, and clinical uses of the buccal fat pad. Surg Radio Anat. 2010;32:427–36.
Jespersen NZ, Larsen TJ, Peijs L, Daugaard S, Homøe P, Loft A, et al. A classical brown adipose tissue mRNA signature partly overlaps with brite in the supraclavicular region of adult humans. Cell Metab. 2013;17:798–805.
Kahn JL, Wolfram-Gabel R, Bourjat P. Anatomy and imaging of the deep fat of the face. Clin Anat. 2000;13:373–82.
Zheng L, Warotayanont R, Stahl J, Kunimatsu R, Klein O, DenBesten PK, et al. Inductive ability of human developing and differentiated dental mesenchyme. Cells Tissues Organs. 2013;198:99–110.
Stockton DW, Das P, Goldenberg M, D’Souza RN, Patel PI. Mutation of PAX9 is associated with oligodontia. Nat Genet. 2000;24:18–9.
Buschmann WR, Hudgins LC, Kummer F, Desai P, Jahss MH. Fatty acid composition of normal and atrophied heel fat pad. Foot Ankle 1993;14:389–94.
Resnick RB, Hudgins LC, Buschmann WR, Kummer FJ, Jahss MH. Analysis of the heel pad fat in rheumatoid arthritis. Foot Ankle Int. 1999;20:481–4.
Falsetti P, Frediani B, Acciai C, Baldi F, Filippou G, Marcolongo R. Heel fat pad involvement in rheumatoid arthritis and in spondyloarthropathies: an ultrasonographic study. Scand J Rheumatol. 2004;33:327–31.
Blackburn AM. Neurotensin. J Clin Pathol Suppl (Assoc Clin Pathol). 1978;8:12–6.
Ouyang Q, Zhou J, Yang W, Cui H, Xu M, Yi L. Oncogenic role of neurotensin and neurotensin receptors in various cancers. Clin Exp Pharm Physiol. 2017;44:841–6.
Li J, Song J, Zaytseva YY, Liu Y, Rychahou P, Jiang K, et al. An obligatory role for neurotensin in high-fat-diet-induced obesity. Nature 2016;533:411–5.
Januzzi JL Jr, Lyass A, Liu Y, Gaggin H, Trebnick A, Maisel AS, et al. Circulating proneurotensin concentrations and cardiovascular disease events in the community: the framingham heart study. Arterioscler Thromb Vasc Biol. 2016;36:1692–7.
Osadchii OE. Emerging role of neurotensin in regulation of the cardiovascular system. Eur J Pharm. 2015;762:184–92.
Acknowledgements
We would like to acknowledge the Core Unit DNA-Technologies headed by PD Dr Knut Krohn from the Medical Faculty/University of Leipzig. We thank Prof Markus Scholz and Dr Holger Kirsten from the Institute for Medical Informatics, Statistics and Epidemiology (IMISE)/University of Leipzig for bioinformatics support. We are grateful for excellent technical assistance by Petra Süptitz and Beate Gutsmann. Open Access Funding provided by Projekt DEAL.
Funding
This study was supported by grants to AT and PK from the Deutsche Forschungsgemeinschaft (DFG, SFB 1052/1, C01, B03, B09 and SPP 1629 TO 718/2-1), from the German Diabetes Association (DDG) and by the Integrated Research and Treatment Center (IFB) AdiposityDiseases (K403, K737, AD2-7123). DS is funded by the Integrated Research and Treatment Center (IFB) AdiposityDiseases (K4x/1). IFB is supported by the Federal Ministry of Education and Research (BMBF), Germany FKZ: 01EO1501 (research projects AD2-060E, AD2-06E95 and AD2-06E99 to PK).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
41431_2020_681_MOESM2_ESM.jpg
Supplemental Figure 1: Viablility of AT from body donors. Tissue slices of 7 µm were stained using antibody against the adipocyte specific marker perilipin (green), the macrophage marker AIF1 (red), and the nuclei were stained with DAPI (blue). Scale bar represents 50 µm.
41431_2020_681_MOESM3_ESM.png
Supplemental Figure 2: Proportion of gene expression between the tissues of selected biological functions and pathways. Calculation in percent based on the mean tissue expression of two body donors with a post mortem delay of 10 hours. White-blue-red coding=0-100%. (For readability, please open the figure with other software than Wndows photo.)
41431_2020_681_MOESM4_ESM.jpg
Supplemental Figure 3: Top enriched GO term for process (1), function (2) & component (3). Boxes for the respective tissue display top enriched GO term with p<0.05 and false discovery rate (FDR) q-value<0.2. 1) process (p-value, FDR q-value, enrichment) and/or 2) function (p-value, FDR q-value, enrichment) and/or 3) component (p-value, FDR q-value, enrichment). Gene lists from the single tissue comparison against all other tissues from the visceral area including epicardial AT were also combined for GOrilla as the single information for these tissues would not withstand statistical significance with the respective cut off for the false discovery rate (dashed line box). Analyzed transcripts for this figure are given in “Gene Lists for GOrilla” in the Variable Lists collection V1-6 Supplemental document 1.
41431_2020_681_MOESM5_ESM.docx
Supplemental Figure 4: Markers for beiging/browning. Given are the mRNA expression values for each sample represented by a dot for the following markers: BMP7, CIDEA, EBF2, FGF21, LHX8, PPARG, PRDM16, TBX1, TMEM26, UCP1. The continuous line (red) represents the mean over all samples. The small lines (green) represent the mean for all samples from one tissue location. The data displayed here are pre-processed, log2 transformed and are adjusted for post mortem delay, Sentrix barcode, age and sex. Of note, the Y-axes have different scaling. Further investigated marker with similar results (data not shown): ADAM17, BMP4, CAR4, TNFRSF9, CITED1, COX4, HOXC8/9, HSPB7, MIR133B, miR26-family, PPARGC1A, ZIC1.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Schleinitz, D., Krause, K., Wohland, T. et al. Identification of distinct transcriptome signatures of human adipose tissue from fifteen depots. Eur J Hum Genet 28, 1714–1725 (2020). https://doi.org/10.1038/s41431-020-0681-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41431-020-0681-1