Abstract
Hereditary hearing loss is a monogenic disease with high genetic heterogeneity. Variants in more than 100 deafness genes underlie the basis of its pathogenesis. The aim of this study was to assess the ratio of SNVs in known deafness genes contributing to the etiology of both sporadic and familial sensorineural hearing loss patients from China. DNA samples from 1127 individuals, including normal hearing controls (n = 616), sporadic SNHL patients (n = 433), and deaf individuals (n = 78) from 30 hearing loss pedigrees were collected. The NGS tests included analysis of sequence alterations in 129 genes. The variants were interpreted according to the ACMG/AMP guidelines for genetic hearing loss combined with NGS data from 616 ethnically matched normal hearing adult controls. We identified a positive molecular diagnosis in 226 patients with sporadic SNHL (52.19%) and in patients from 17 deafness pedigrees (56.67%). Ethnically matched MAF filtering reduced the variants of unknown significance by 8.7%, from 6216 to 5675. Some complexities that may restrict causative variant identification are discussed. This report highlight the clinical utility of NGS panels identifying disease-causing variants for the diagnosis of hearing loss and underlines the importance of a broad data of control and ACMG/AMP standards for accurate clinical delineation of VUS variants.
Similar content being viewed by others
Introduction
Hearing impairment is one of the most common human disabilities. According to the World Health Organization, 5% of the world’s population (~360 million people) suffer from disabling hearing. More people are affected by severe hearing loss than by epilepsy, multiple sclerosis, spinal injury, stroke, Huntington’s disease, and Parkinson’s disease combined [1]. Sensorineural hearing loss (SNHL) accounts for ~90% of all human hearing loss cases. Among these patients, genetic factors are estimated to be responsible for >60% of the cases [2, 3], and most of these cases are caused by a single nucleotide variants (SNVs), a small fraction by a small insertion–deletion (indel) variant or copy number variants (CNVs) [4, 5]. Depending on the gene involved, the hearing loss can either be syndromic or nonsyndromic. A genetic diagnosis is valuable for providing essential prognostic information needed for deciding optimal treatment/rehabilitation options, and is needed for genetic consulting to predict the risk of recurrence [6].
Molecular epidemiological studies have found several common deafness genes, such as GJB2, SLC26A4, and mitochondrial 12S rRNA, which appear to account for 30–50% of congenital hearing loss cases [7,8,9,10,11,12]. However, genetic variants responsible for a large number of cases of hereditary hearing loss remain unknown, especially in patients with sporadic hearing loss. Next-generation sequencing [2] has greatly increased efficiency in screening known deafness genes for diagnostic purposes and in identifying new deafness genes [5, 13,14,15]. However, the accuracy and clinical utility of the NGS approach has not been systematically evaluated in a large number of clinically diagnosed cases of sporadic SNHL and deafness pedigrees of Chinese ethnicity. This translational study employed NGS to screen 119 nuclear deafness genes and 10 mitochondria genes in patients with sporadic SNHL, individuals from deafness pedigrees, and ethnically matched normal hearing controls. The data were used to systematically assess the diagnostic rate of the deafness genes panel and the role of ethnically matched controls in interpreting variants to eliminate false positives.
Materials and methods
Collection of patient and control samples
This study was performed according to the protocol approved by the Ethics Committee of the Chinese People’s Liberation Army General Hospital. We collected DNA samples from 1127 individuals, including normal hearing controls (n = 616), sporadic patients clinically diagnosed SNHL (n = 433), and individuals from 30 deaf pedigrees including familial cases (n = 78). Ninety-nine family members with normal hearing from the 30 deaf pedigrees were enrolled for segregation analysis.
The sporadic cases were recruited from outpatients who visited the hospital during 2011–2013; only unique probands were included (each nuclear and/or extended family was represented only by the proband). The patients comprised those who claimed hearing loss and were verified by either subjective and/or objective hearing tests, those who failed the newborn hearing screening and were further diagnosed as deafness by objective hearing tests, and those who failed the newborn deafness gene screening carrying at least one variant of the nine including GJB2 c.35delG, c.176del16, c.235delC, c.299delAT (NM_004004.5), GJB3 c.538C>T (NM_001005752.1), SLC26A4 c.919-2A>G (NG_008489.1), c.2168A>G (NM_000441.1), mtDNA 12S rRNA, and were further referred to the otology clinic and diagnosed as hearing impairment by objective hearing tests. Chinese individuals with normal hearing (controls) were verified by pure-tone audiometry. Only those with a PTA (pure-tone average) threshold of 0–20 dB were enrolled. The average age of this population was 31.18 ± 10.18 years; no control individuals claimed hearing problems or family history, and all passed physical examinations. All participants were informed about the scope and requirements of the study, and all signed the consent forms approved by the Ethics Committees of the participating institutions.
The clinical history was obtained by investigators, with special emphasis on the onset age of hearing loss; family history of deafness; pregnancy and labor history; general health condition; potential environmental causes of hearing loss such as infections and trauma; and information on exposure to known or possible ototoxic drugs. Cases involving trauma and otitis media were excluded from this study. Conventional clinical examinations included pure-tone audiometry, acoustic immittance, and auditory brainstem responses (ABR). Distortion product otoacoustic emissions, ABR, and auditory steady state response tests were carried out in babies who failed the hearing screening and in children who were uncooperative during the subjective audiometry examination. The level of hearing loss was described in terms of PTA (calculated as the average of the threshold measured at 0.5, 1.0, 2.0, and 4.0 kHz) as follows: normal hearing, <20 dB; mild hearing impairment, 21–40 dB; moderate hearing impairment, 41–70 dB; severe hearing impairment, 71–90 dB; and profound hearing impairment, >91 dB. High-resolution computed tomography (CT) of the temporal bone was performed in the sporadic and pedigree SNHL cases to exclude middle ear pathological changes and to diagnose inner ear malformation. Magnetic resonance imaging was performed when the CT scan failed to reveal membranous labyrinth problems in a patient. Syndromic hearing loss was diagnosed based on a range of evidence, including self-reported symptoms, family history, and physical examination, with special attention to the external ears and neck, skin, hair, eyes, and digits. Data Management System for Clinical Otology software (Computer Software Copyright: 2008SR06229) was used to store and track the information.
DNA isolation and previous genetic test
The DNA was isolated from blood extracted from a peripheral vein by a registered nurse in the genetic testing center of the Chinese PLA General Hospital. Genomic DNA (gDNA) was extracted using gDNA blood extraction kits (Qiagen, Valencia, CA, USA) within 1 week of sample collection. The quality of the gDNA was examined by assessing the optical density ratio (260/280 ratio) and by gel electrophoresis imaging for the presence of a high-molecular-weight gDNA band. All of the 433 sporadic cases were sequenced 12S rRNA, the coding areas of GJB2 and SLC26A4 previously.
Library preparation and sequencing
The DA3 panel assay (Otogenetics Corporation, Atlanta, GA) includes 119 nuclear deafness genes and 10 mitochondria genes (Table S1). Library preparation and sequencing were described in our previous report [16].
Bioinformatics
After sequencing the target region, quality control was performed to ensure data accuracy. Low-quality data were filtered out to obtain clean sequencing data. Burrows–Wheeler alignment was used to align the clean sequence to the human reference genome hg19 nuclear genes or GRCh37 for mitochondria genes. GATK was used to detect SNP and Indel variants. VEP and dbNSFP databases were used to obtain the variants information, including minor allele frequency (MAF), variant consequence, altered protein function, gene information, and related disease information.
Statistical analysis
After raw variants were annotated, we performed further filtering to identify candidate variants. Variants meeting all the following requirements were included: (1) MAF of KG and Exome Aggregation Consortium (ExAC) database < 0.01; (2) 0.005 < MAF < 0.05; (3) variants affecting protein function, including loss-of-function and missense variants. After that, we obtained 142 variants from 433 hearing loss cases and 616 normal controls. We performed a case-control association study using Fisher’s exact test and adjusted p-values using Bonferroni multiple testing correction. Variants with p-values less than 3.521E−04 (0.05/142) were considered to be significantly associated with disease.
The variants interpreted by the ACMG/AMP guidelines for genetic hearing loss were revalued using allele frequencies from the ethnically matched controls. The variants were categorized as BA1 if the MAF thresholds of autosomal-recessive variants were >0.005 and autosomal-dominant variants were >0.001 in the controls [17, 18]. The above thresholds >0.005 of autosomal-recessive variants excluded specific variants in GJB2 and SLC26A4. By combining the clinical phenotype, inherited model, and related previous studies, we obtained a list of candidate variants. Sanger analysis was performed to verify the variant identified by NGS. The pipeline of bioinformatic analysis was shown in Fig. 1.
The FASTQ data and variant call format were permanently reserved on (CNSA). The pathogenic and likely pathogenic variants were submitted to ClinVar (ClinVarAccessions: SCV000902290~SCV000902403).
Results
The data mapped to the targeted region have a mean depth of 149.027 fold, and the coverage of 1X was 99.8%. The capture rate is nearly 100%.
The phenotypic information of all patients and controls were shown in Table 1.
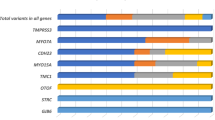
In the 119 nuclear deafness genes, variants in exon or splicing regions (totaling 6185) and intron regions (totaling 16521) were called after the preliminary bioinformatic analysis. We identified 73 (73/6185, 1.18%) pathogenic and 49 (49/6185, 0.79%) likely pathogenic variants (Table S2). In all, 1804 (1804/6185, 29.17%) variants were classified as benign, 919 (919/6185, 14.86%) as likely benign, and 3340 (3340/6185, 54.00%) variants were classified as uncertain significance. As for the definitely deafness-related mitochondrial variants, four homoplasmic m.1555A>G and one homoplasmic m.1494C>T carriers were found in the case group, and one homoplasmic m.1555A>G carrier was identified in the control group. Of the five patients carrying m.1555A>G or m.1494C>T, history of aminoglycoside antibiotics usage due to infection was clear.
Variants classification and evaluation
We chose 1000 Genomes, Exome Sequencing Project [19], ExAC, HapMap, and Wellderly plus five in-house databases for MAF filtering. A total of 8.75% (541/6184) of exon variants were filtered out by the NGS data from the ethnically matched controls. After Bonferroni correction, 6 variants (GJB2 c.235delC, c.299_300delAT, SLC26A4 c.919-2A>G, c.2168A>G, c.1174A>T, USH2A c.6998T>C) achieved a conservative significance threshold. Among the 541 variants, the 1000 Genomes, ESP, ExAC databases reported no MAF thresholds for autosomal-recessive variants >0.005 (excluding specific variants in GJB2 and SLC26A4) and autosomal-dominant variants >0.001; however, MAF > 0.005 was found in our control group in the recessive inheritance pattern and >0.001 in the dominant inheritance pattern and accordingly we categorized them to BA1. This result illustrated the power of this step and stressed the need for deeper and richer databases of multiple ethnicities to identify low-frequency variants specific to ancestry groups that are often unique to specific populations [20].
The allele frequency for GJB2 c.109G>A (p.Val37Ile) was 3.8% (33/866, 1 homozygote and 31 heterozygotes) and 3% (37/1232, 1 homozygote and 35 heterozygotes) in the case and control groups, respectively; the difference between the two groups was not statistically significant. The homozygous p.V37I variant in GJB2 is prevalent in East and Southeast Asians and may lead to a broad spectrum of hearing phenotypes from mild-to-moderate hearing loss with reduced penetrance to profound hearing loss. The homozygous p.V37I knockin mice developed progressive, mild-to-moderate hearing loss over the first 4–9 months which modeled the hearing phenotype of the human patients, and confocal immunostaining and electron microscopic scanning revealed minor loss of the outer hair cells [21]. Thus, we regard it as likely pathogenic variants. The fact that the majority of the case cohort showed severe to profound hearing loss may be the explanation for the lack of statistical significance. This result suggests that full-scale evaluations of the pathogenicity of a variant must include molecular epidemiology, functional experiments, and information from published studies.
Positive diagnostic rate for SNVs
In sporadic SNHL cases, variants in 24 genes were verified for genetic etiology in 52.19% (226/433) by Sanger sequencing and cosegregation analysis in their families (Table 2). The basic and diagnostic information for each sample were shown in Fig. 2. Considering the ratio of hereditary factors in hearing loss (60%), the positive diagnostic rate of the gene panel containing 129 genes in hereditary hearing loss might reach 86.98% (52.19%/60%). Detailed genotypes are shown in Table S3. Possible causative variants were found in 7.39% (32/433) of the cases; however, DNA samples and exact phenotypes of the family members were not available, and the genetic diagnosis could not be confirmed. Although 119 or more genes were included in clinical genetic testing, a distinct hotspot gene spectrum focused on several genes including GJB2, SLC26A4, 12S rRNA, and MYO15A. The allele and carrier frequencies for variants in common deafness gene are shown in Table S4.
The interaction graph of basic and diagnostic information for each sample. Positive diagnosis is influenced by ethnic, clinical, and phenotypic characteristics in sporadic hearing loss population. N for each combination of two reported characteristics for all combinations. Color/shading reflects the number of patients with the paired criteria, up to the maximum of n = 433
In deafness pedigrees, variants affecting protein function in 12 genes were verified with the positive diagnostic rate 56.67% (17/30) (Table 3). Detailed information on these deafness pedigrees is provided in our previous studies [22,23,24,25,26,27,28,29].
Discussion
Identification of the precise genetic cause of hearing loss can provide helpful information for treatment like cochlear implantation [30], and patient management such as hearing and speech rehabilitation, prediction of prognosis, genetic counseling, and precise genetic therapies [19]. Previously, no comprehensive diagnostic testing had been completed in a Chinese deafness cohort, and no related gene variant data had been obtained in normal hearing controls with PTA-verified tests. To determine the aggregate genetic contribution to SNHL, we performed clinical genetic testing on 119 nuclear deafness genes and 10 mitochondria genes in 433 sequentially recruited patients with sporadic hearing loss, 616 controls, and 30 deafness pedigrees. No sporadic patients were excluded based on phenotype, inheritance, or previous testing.
The comprehensive testing resulted in the identification of gene sequence alterations associated with the underlying genetic cause for hearing loss in 226 patients (52.19%), higher than the 41% identified among cases with multiethnic background [15] and ~40% among Japanese patients [31]. Although Christina et al. [15] reported a diagnostic rate of 1% in patients with unilateral hearing loss (69 cases), no diagnostic positive result was found in the patient with unilateral hearing loss in our study. The negative result might be explained by the small sample of cases with unilateral hearing loss. In the 226 patients with positive diagnoses, the percentages of recessive, dominant, mitochondrial, and X-linked inheritance patterns were 94.24% (213/226), 2.21% (5/226), 2.21% (5/226), and 1.33% (3/226), respectively. Causative dominant variants were relatively less frequently identified in sporadic hearing loss. In our study, 7.39% (32/433) of our cases with likely causative variants could not be genetically diagnosed due to the lack of DNA samples or exact phenotypes from family members. In addition, some of our assumed de novo variants lacked confirmation of paternity and maternity.
Genetic spectrum in the Chinese population
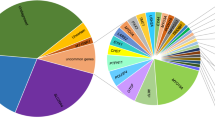
Despite the high genetic heterogeneity of hearing loss, most disease-causing variants are rarely recurrent [32] except those in GJB2 and SLC26A4. Our study showed that disease-causing variants in GJB2 and SLC26A4 were found in exceptionally high numbers, followed by variants in USH2A, MYO15A, MYO1F, and MYO7A. Variants affecting protein function in GJB2 and SLC26A4 were the most common cause of autosomal-recessive NSHL, accounting for 80.42% (184/226) of the genetic basis in all hereditary hearing loss patients. The genetic spectrum for autosomal-dominant NSHL seems to be wide and not focused on one or a few genes; this result differs from the findings of a study involving Japanese subjects, which reported KCNQ4 as the most frequent cause for dominant NSHL [31]. We made a comparison across multiple populations (Table 4) and found the genetic diagnostic rate varied from 12.7 to 66.7% which may due to the number of genes captured, variations in patient selection criteria, previous genetic testing and enrollment case numbers [33,34,35,36,37,38,39,40,41,42,43,44,45,46]. In the cases with Caucasian ethnicity, nearly three-fourths of all diagnoses were attributable to ten genes and the four genes most frequently implicated were GJB2 (22%), STRC (16%), SLC26A4 (7%), and TECTA (5%), which is different from those of our study except GJB2. The pathogenesis of 433 Chinese sporadic hearing loss patients was shown in Fig. 3a. The ratio of hereditary factors in congenital hearing loss is at least 60%, and the cause of the other 40% was considered to be environmental and unknown factors [2, 3]. After DA3 testing, there are 7.81% of the SNHL patients in this study assumed to be due to genetic factors yet to be discovered. A comparison of the positive diagnostic rate in the common deafness genes (GJB2, SLC26A4, and 12S rRNA) in this case cohort with that in a larger sample (16,456 cases) from our deafness genetic testing center revealed that our sample selection was representative of the Chinese deafness population (Fig. 3b). Although targeted genomic enrichment followed by NGS has been shown to be an efficient strategy for the clinical diagnosis of hearing loss, in the Chinese Han population, a first-line test for frequent genes with Sanger sequencing is advantageous for economic and time-saving considerations.
Etiology classification and the representation analysis of the studied sporadic patient cohort. a Pathogenesis of 433 Chinese sporadic hearing loss patients. b The representation comparison of the studied 433 cases with 16,456 patients from our Clinic. Salmon pink indicated the positive diagnostic ratio of deafness panel (including 119 genes and mitochrodrial genome) in 433 cases. Light green indicated the positive diagnostic ratio of Sanger sequencing for common deafness genes (GJB2, SLC26A4, and mit12S rRNA) in 16,456 cases. The first three pairs of columns showed there are no significant differences of the positive testing ratio on GJB2, SLC26A4, mit12S rRNA between the two patient cohorts, which indicated that the cases enrolled in this study could represent a larger deafness population in China. The largest ethnic group in both cohorts was Han Chinese, comprising up to 95% of the total sample
The importance of ethnically matched MAF filtering
Extensive optimization and evaluation procedures are required for all NGS platforms to ensure a reliable and routine application of NGS technologies in diagnostics. We included a filtering step based on NGS data of MAFs in 616 Chinese normal hearing controls to minimize false-positive results [20]. In aggregate, ethnicity-specific MAF filtering helped reduce the list of VUS variants from 6184 to 5643. This step improved the annotation of identified variants and allowed us to recategorize 541 variants as benign that were otherwise annotated as VUS, and recategorize several variants as VUS that were otherwise annotated as pathogenic by the 1000 Genomes, ESP, ExAC, HapMap, and Wellderly databases.
The reported GJB3 digenic variants c.580G>A and the dominant variant c.538C>T appeared to be nonpathogenic. In our study, six patients and six controls were GJB3 c.580G>A heterozygotes. None of the six deafness patients carried the pathogenic GJB2 variants reported by Liu et al. [47]. Three patients and one control were GJB3 c.538C>T heterozygotes. Considering the Sanger results and phenotype of family members, we recategorized c.580G>A (BA1) and c.538C>T (BS1 + BS4) as benign. Our study supports the findings that querified variants in GJB3 as a cause of deafness [17]. Further studies are required to verify whether GJB3 variants cause fully penetrant deafness.
Molecular diagnosis in patients with SNHL and inner ear malformations
The etiologies of inner ear malformations have not been completely clarified. Genotype and phenotype correlations were determined in EVA and IP-III. In this study, we observed at least seven types of inner ear malformations in 100 patients; the rate of inner ear malformations in SNHL was 23.09% (100/433). The most common type was EVA (89 cases), which is an autosomal-recessive disease. In our study, 94.3% (84/89) of EVA cases carried bi-allelic causative variants, 3.37% (3/89) had monoallelic, and 2.25% (2/89) had no variant in SLC26A4. Neither KCNJ10 nor FOXI I variants contributed to the molecular etiology of EVA among patients in this study. The relevance ratio of bi-allelic SLC26A4 variants is 36% in Caucasians, 66% in Japanese, and 81% in Koreans [48,49,50]. In this study, the second most common type was IP-III, an X-linked genetic disease in which the basal turn of the cochlea is placed directly at the end of the inner auditory canal (three cases); 100% of the male patients with this malformation carried hemizygous variants in POU3F4. Other inner ear malformations included cochlea aplasia (one case), cochlea hypoplasia (one case), isolated Mondini deformity (three cases), common cavity deformity (two cases), and narrow internal auditory canal (one case). No genetic pathogenesis was identified in the types of inner ear deformity other than EVA and IP-III, indicating that the etiology of inner ear malformation may be multifactorial.
Molecular diagnosis in patients with syndromic SNHL
The clinical diagnosis of syndromic SNHL helps direct the test panel choice, thereby saving time and cost. Three patients with Waardenburg syndrome (WS) were diagnosed clinically by the phenotype of hearing loss with dermal pigmentation and iris heterochrosis. In two of the WS cases, genetic variants in PAX3 and MITF were identified, respectively, whereas the other case showed no causative variant in WS-related genes (PAX3, MITF, SNAI2, and EDNRB). No Pendred syndrome was diagnosed in our patient cohort, which may be due to the fact that the average age of our patients with EVA was 9.36 years. The phenotype of goiter is not present at birth, developing in early puberty (40%) or adulthood (60%). In addition, it is possible that our sample size was not large enough. We identified ten cases with bi-allelic variants in the Usher syndrome-related genes, USH2A, MYO7A, CDH23, and ADGRV1. Usher syndrome is a clinically and genetically heterogeneous autosomal-recessive disorder characterized by sensorineural hearing deficiencies and later development of progressive retinitis pigmentosa. It is the most frequent cause of combined deafness and blindness in adults and affects 3–6% of children born with hearing impairment. None of the ten cases had vestibular dysfunction or hypoplastic, and vestibular and ophthalmologic examinations were not performed at the beginning of this study. The average age of these ten patients was 12.77 years. Hearing loss occurred postlingually in five cases, prelingually in four, and one with ambiguous onset. In consideration of the variable extent of vestibular involvement and the usual onset of retinitis pigmentosa symptoms in the second decade of life, vestibular evaluation and ophthalmologic follow-up will be performed in the future. In addition, there is the possibility that variants in MYO7A and CDH23 caused DFNB2 and DFNB12.
Conclusions
We demonstrated that the DA3 panel assay covering 129 genes had a positive diagnostic rate of 52.19% on sequence alterations for sporadic SNHL patients and 56.67% for deafness pedigrees. WES or WGS should be reserved only for negative cases as an opportunity to discover novel candidate genes. Ethnically matched MAF filtering changed the categorization of 8.75% of our list of variants from VUS to benign. This report highlight the clinical utility of NGS panels identifying disease-causing variants for the clinical diagnosis of deafness and underlines the importance of a broad data of control and the ACMG/AMP guidelines for accurate clinical delineation of VUS variants. The results provide more detailed information for genetic consulting to predict the risk of recurrence.
Data availability
All data are presented in the manuscript or additional files.
References
Hudspeth AJ. How hearing happens. Neuron. 1997;19:947–50.
Marazita ML, Ploughman LM, Rawlings B, Remington E, Arnos KS, Nance WE. Genetic epidemiological studies of early-onset deafness in the U.S. school-age population. Am J Med Genet. 1993;46:486–91.
Smith RJ, Bale JF Jr., White KR. Sensorineural hearing loss in children. Lancet. 2005;365:879–90.
Hilgert N, Smith RJ, Van Camp G. Forty-six genes causing nonsyndromic hearing impairment: which ones should be analyzed in DNA diagnostics? Mutat Res. 2009;681:189–96.
Shearer AE, DeLuca AP, Hildebrand MS, Taylor KR, Gurrola J 2nd, et al. Comprehensive genetic testing for hereditary hearing loss using massively parallel sequencing. Proc Natl Acad Sci USA. 2010;107:21104–9.
Alford RL, Arnos KS, Fox M, Lin JW, Palmer CG, Pandya A, et al. American College of Medical Genetics and Genomics guideline for the clinical evaluation and etiologic diagnosis of hearing loss. Genet Med. 2014;16:347–55.
Brownstein Z, Avraham KB. Deafness genes in Israel: implications for diagnostics in the clinic. Pediatr Res. 2009;66:128–34.
Du W, Wang Q, Zhu Y, Wang Y, Guo Y. Associations between GJB2, mitochondrial 12S rRNA, SLC26A4 mutations, and hearing loss among three ethnicities. Biomed Res Int. 2014;2014:746838.
Ito T, Noguchi Y, Yashima T, Ohno K, Kitamura K. Hereditary hearing loss and deafness genes in Japan. J Med Dent Sci. 2010;57:1–10.
Mahdieh N, Rabbani B, Wiley S, Akbari MT, Zeinali S. Genetic causes of nonsyndromic hearing loss in Iran in comparison with other populations. J Hum Genet. 2010;55:639–48.
Shahin H, Walsh T, Rayyan AA, Lee MK, Higgins J, Dickel D, et al. Five novel loci for inherited hearing loss mapped by SNP-based homozygosity profiles in Palestinian families. Eur J Hum Genet. 2010;18:407–13.
Yuan Y, You Y, Huang D, Cui J, Wang Y, Wang Q, et al. Comprehensive molecular etiology analysis of nonsyndromic hearing impairment from typical areas in China. J Transl Med. 2009;7:79.
Lin X, Tang W, Ahmad S, Lu J, Colby CC, Zhu J, et al. Applications of targeted gene capture and next-generation sequencing technologies in studies of human deafness and other genetic disabilities. Hear Res. 2012;288:67–76.
Idan N, Brownstein Z, Shivatzki S, Avraham KB. Advances in genetic diagnostics for hereditary hearing loss. J Basic Clin Physiol Pharmacol. 2013;24:165–70.
Sloan-Heggen CM, Bierer AO, Shearer AE, Kolbe DL, Nishimura CJ, Frees KL, et al. Comprehensive genetic testing in the clinical evaluation of 1119 patients with hearing loss. Hum Genet. 2016;135:441–50.
Tang W, Qian D, Ahmad S, Mattox D, Todd NW, Han H, et al. A low-cost exon capture method suitable for large-scale screening of genetic deafness by the massively-parallel sequencing approach. Genet Test Mol Biomark. 2012;16:536–42.
Shearer AE, Eppsteiner RW, Booth KT, Ephraim SS, Gurrola J 2nd, Simpson A, et al. Utilizing ethnic-specific differences in minor allele frequency to recategorize reported pathogenic deafness variants. Am J Hum Genet. 2014;95:445–53.
Oza AM, DiStefano MT, Hemphill SE, Cushman BJ, Grant AR, Siegert RK, et al. Expert specification of the ACMG/AMP variant interpretation guidelines for genetic hearing loss. Hum Mutat. 2018;39:1593–613.
Muller U, Barr-Gillespie PG. New treatment options for hearing loss. Nat Rev Drug Discov. 2015;14:346–65.
Casals F, Bertranpetit J. Genetics. Human genetic variation, shared and private. Science. 2012;337:39–40.
Chen Y, Hu L, Wang X, Sun C, Lin X, Li L, et al. Characterization of a knock-in mouse model of the homozygous p.V37I variant in Gjb2. Sci Rep. 2016;6:33279.
Gao X, Huang SS, Yuan YY, Wang GJ, Xu JC, Ji YB, et al. Targeted gene capture and massively parallel sequencing identify TMC1 as the causative gene in a six-generation Chinese family with autosomal dominant hearing loss. Am J Med Genet A. 2015;167A:2357–65.
Gao X, Wang GJ, Yuan YY, Xin F, Han MY, Lu JQ, et al. Correction: novel compound heterozygous mutations in MYO7A associated with Usher syndrome 1 in a Chinese family. PLoS ONE. 2015;10:e0137883.
Gao X, Yuan YY, Wang GJ, Xu JC, Su Y, Lin X, et al. Novel mutations and mutation combinations of TMPRSS3 cause various phenotypes in one Chinese family with autosomal recessive hearing impairment. Biomed Res Int. 2017;2017:4707315.
Huang A, Yuan Y, Liu Y, Zhu Q, Dai P. A novel EYA4 mutation causing hearing loss in a Chinese DFNA family and genotype-phenotype review of EYA4 in deafness. J Transl Med. 2015;13:154.
Huang B, Liu Y, Gao X, Xu J, Dai P, Zhu Q, et al. A novel pore-region mutation, c.887G > A (p.G296D) in KCNQ4, causing hearing loss in a Chinese family with autosomal dominant non-syndromic deafness 2. BMC Med Genet. 2017;18:36.
Su Y, Tang WX, Gao X, Yu F, Dai ZY, Zhao JD, et al. A novel mutation in the TECTA gene in a Chinese family with autosomal dominant nonsyndromic hearing loss. PLoS ONE. 2014;9:e89240.
Yuan Y, Gao X, Huang B, Lu J, Wang G, Lin X, et al. Phenotypic heterogeneity in a DFNA20/26 family segregating a novel ACTG1 mutation. BMC Genet. 2016;17:33.
Gao X, Xu JC, Wang WQ, Yuan YY, Bai D, Huang SS, et al. A missense mutation in POU4F3 causes midfrequency hearing loss in a Chinese ADNSHL family. Biomed Res Int. 2018;2018:5370802.
Eppsteiner RW, Shearer AE, Hildebrand MS, Deluca AP, Ji H, Dunn CC, et al. Prediction of cochlear implant performance by genetic mutation: the spiral ganglion hypothesis. Hear Res. 2012;292:51–8.
Moteki H, Azaiez H, Booth KT, Shearer AE, Sloan CM, Kolbe DL, et al. Comprehensive genetic testing with ethnic-specific filtering by allele frequency in a Japanese hearing-loss population. Clin Genet. 2016;89:466–72.
Diaz-Horta O, Duman D, Foster J 2nd, Sirmaci A, Gonzalez M, Mahdieh N, et al. Whole-exome sequencing efficiently detects rare mutations in autosomal recessive nonsyndromic hearing loss. PLoS ONE. 2012;7:e50628.
Shearer AE, Black-Ziegelbein EA, Hildebrand MS, Eppsteiner RW, Ravi H, Joshi S, et al. Advancing genetic testing for deafness with genomic technology. J Med Genet. 2013;50:627–34.
Shearer AE, Kolbe DL, Azaiez H, Sloan CM, Frees KL, Weaver AE, et al. Copy number variants are a common cause of non-syndromic hearing loss. Genome Med. 2014;6:37.
Shearer AE, Smith RJ. Massively parallel sequencing for genetic diagnosis of hearing loss: the new standard of care. Otolaryngol Head Neck Surg. 2015;153:175–82.
Wei Q, Zhu H, Qian X, Chen Z, Yao J, Lu Y, et al. Targeted genomic capture and massively parallel sequencing to identify novel variants causing Chinese hereditary hearing loss. J Transl Med. 2014;12:311.
Vona B, Muller T, Nanda I, Neuner C, Hofrichter MA, Schroder J, et al. Targeted next-generation sequencing of deafness genes in hearing-impaired individuals uncovers informative mutations. Genet Med. 2014;16:945–53.
Likar T, Hasanhodzic M, Teran N, Maver A, Peterlin B, Writzl K. Diagnostic outcomes of exome sequencing in patients with syndromic or non-syndromic hearing loss. PLoS ONE. 2018;13:e0188578.
Park JH, Kim NK, Kim AR, Rhee J, Oh SH, Koo JW, et al. Exploration of molecular genetic etiology for Korean cochlear implantees with severe to profound hearing loss and its implication. Orphanet J Rare Dis. 2014;9:167.
Zazo Seco C, Wesdorp M, Feenstra I, Pfundt R, Hehir-Kwa JY, Lelieveld SH, et al. The diagnostic yield of whole-exome sequencing targeting a gene panel for hearing impairment in The Netherlands. Eur J Hum Genet. 2017;25:308–14.
Mutai H, Suzuki N, Shimizu A, Torii C, Namba K, Morimoto N, et al. Diverse spectrum of rare deafness genes underlies early-childhood hearing loss in Japanese patients: a cross-sectional, multi-center next-generation sequencing study. Orphanet J Rare Dis. 2013;8:172.
Wu CC, Lin YH, Lu YC, Chen PJ, Yang WS, Hsu CJ, et al. Application of massively parallel sequencing to genetic diagnosis in multiplex families with idiopathic sensorineural hearing impairment. PLoS ONE. 2013;8:e57369.
Yang T, Wei X, Chai Y, Li L, Wu H. Genetic etiology study of the non-syndromic deafness in Chinese Hans by targeted next-generation sequencing. Orphanet J Rare Dis. 2013;8:85.
Vozzi D, Morgan A, Vuckovic D, D’Eustacchio A, Abdulhadi K, Rubinato E, et al. Hereditary hearing loss: a 96 gene targeted sequencing protocol reveals novel alleles in a series of Italian and Qatari patients. Gene. 2014;542:209–16.
Brownstein Z, Friedman LM, Shahin H, Oron-Karni V, Kol N, Abu Rayyan A, et al. Targeted genomic capture and massively parallel sequencing to identify genes for hereditary hearing loss in Middle Eastern families. Genome Biol. 2011;12:R89.
Chen Y, Wang Z, Wang Z, Chen D, Chai Y, Pang X, et al. Targeted next-generation sequencing in Uyghur families with non-syndromic sensorineural hearing loss. PLoS ONE. 2015;10:e0127879.
Liu XZ, Yuan Y, Yan D, Ding EH, Ouyang XM, Fei Y, et al. Digenic inheritance of non-syndromic deafness caused by mutations at the gap junction proteins Cx26 and Cx31. Hum Genet. 2009;125:53–62.
Albert S, Blons H, Jonard L, Feldmann D, Chauvin P, Loundon N, et al. SLC26A4 gene is frequently involved in nonsyndromic hearing impairment with enlarged vestibular aqueduct in Caucasian populations. Eur J Hum Genet. 2006;14:773–9.
Tsukamoto K, Suzuki H, Harada D, Namba A, Abe S, Usami S. Distribution and frequencies of PDS (SLC26A4) mutations in Pendred syndrome and nonsyndromic hearing loss associated with enlarged vestibular aqueduct: a unique spectrum of mutations in Japanese. Eur J Hum Genet. 2003;11:916–22.
Campbell C, Cucci RA, Prasad S, Green GE, Edeal JB, Galer CE, et al. Pendred syndrome, DFNB4, and PDS/SLC26A4 identification of eight novel mutations and possible genotype-phenotype correlations. Hum Mutat. 2001;17:403–11.
Gu X, Guo L, Ji H, Sun S, Chai R, Wang L, Li H. Genetic testing for sporadic hearing loss using targeted massively parallel sequencing identifies 10 novel mutations. Clin Genet. 2015;87:588–93.
Acknowledgements
We sincerely thank all the family members for their participation and cooperation in this study.
Funding
This study was supported by grants from National Key Research and Development Project of China (2016YFC1000706) and National Natural Science Foundation of China (81873704) and Fostering Funds of Chinese PLA General Hospital for National Distinguished Young Scholar Science Fund (2017-JQPY-001) to YY, National Natural Science Foundation of China (81570929) to XG, National Key Research and Development Project (2016YFC1000700, 2016YFC1000704) and National Natural Science Foundation of China (81730029) and Natural Science Foundation Key Project of Beijing (7191011) to PD, National Institute on Deafness and other Communication Disorders (RO1 DC006483 and RO1 DC014496) to XL. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the paper.
Author contributions
YYY, QL, and SSH carried out the molecular genetic studies and Sanger sequence alignment in sporadic hearing loss patients and normal hearing controls. YS, XG, and DYK carried out the molecular genetic studies and Sanger sequence alignment in hearing loss pedigrees. JGZ, QFL, and HKL carried out the bioinformatic analysis of the sequencing data. YYY and JGZ drafted the paper. NWT and DM analyzed the genotype–phenotype correlations in all patients. XL and PD conceived the study, participated in its design and coordination. All authors have read and approved the final paper.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval
This study was approved by the Ethics Committee of the Chinese People’s Liberation Army General Hospital (reference number S2016-120-02).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yuan, Y., Li, Q., Su, Y. et al. Comprehensive genetic testing of Chinese SNHL patients and variants interpretation using ACMG guidelines and ethnically matched normal controls. Eur J Hum Genet 28, 231–243 (2020). https://doi.org/10.1038/s41431-019-0510-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41431-019-0510-6
This article is cited by
-
Quantitative thresholds for variant enrichment in 13,845 cases: improving pathogenicity classification in genetic hearing loss
Genome Medicine (2023)
-
Preimplantation genetic testing for hereditary hearing loss in Chinese population
Journal of Assisted Reproduction and Genetics (2023)
-
Addition of an affected family member to a previously ascertained autosomal recessive nonsyndromic hearing loss pedigree and systematic phenotype-genotype analysis of splice-site variants in MYO15A
BMC Medical Genomics (2022)
-
Variant analysis of 92 Chinese Han families with hearing loss
BMC Medical Genomics (2022)
-
A novel splice site variant c.1183 + 1 G > C in DFNA5 causing autosomal dominant nonsyndromic hearing loss in a Chinese family
BMC Medical Genomics (2022)