Abstract
CYFIP2, encoding the evolutionary highly conserved cytoplasmic FMRP interacting protein 2, has previously been proposed as a candidate gene for intellectual disability and autism because of its important role linking FMRP-dependent transcription regulation and actin polymerization via the WAVE regulatory complex (WRC). Recently, de novo variants affecting the amino acid p.Arg87 of CYFIP2 were reported in four individuals with epileptic encephalopathy. We here report 12 independent patients harboring a variety of de novo variants in CYFIP2 broadening the molecular and clinical spectrum of a novel CYFIP2-related neurodevelopmental disorder. Using trio whole-exome or -genome sequencing, we identified 12 independent patients carrying a total of eight distinct de novo variants in CYFIP2 with a shared phenotype of intellectual disability, seizures, and muscular hypotonia. We detected seven different missense variants, of which two occurred recurrently (p.(Arg87Cys) and p.(Ile664Met)), and a splice donor variant in the last intron for which we showed exon skipping in the transcript. The latter is expected to escape nonsense-mediated mRNA decay resulting in a truncated protein. Despite the large spacing in the primary structure, the variants spatially cluster in the tertiary structure and are all predicted to weaken the interaction with WAVE1 or NCKAP1 of the actin polymerization regulating WRC-complex. Preliminary genotype–phenotype correlation indicates a profound phenotype in p.Arg87 substitutions and a more variable phenotype in other alterations. This study evidenced a variety of de novo variants in CYFIP2 as a novel cause of mostly severe intellectual disability with seizures and muscular hypotonia.
Similar content being viewed by others
Introduction
Intellectual disability (ID) is a genetically heterogeneous disorder with an estimated prevalence of 2–3%, with 0.3–0.5% severely affected [1]. Epilepsy is a frequent comorbidity [2, 3], which worsens its psychosocial outcome [4]. Genetic defects are thought to be a major cause of ID and epilepsy, but the identification of causative variants is complicated by tremendous genetic heterogeneity. Next-generation sequencing techniques, such as whole-exome sequencing (WES) and whole-genome sequencing (WGS), have been shown to be outstanding tools for the identification of causative variants in a large number of established and novel disease genes, especially when a patient-parent trio is available [5,6,7,8,9,10]. However, prioritization of candidate variants and genes can still be challenging. Aside from ranking the impact of the variants themselves (truncation, substitution, etc.), characteristics like expression pattern, biological function and processes, interaction partners, and molecular networks as well as phenotypes of knockout models are often used to underpin the potential role of genes in disease pathogenesis. Several ID and epilepsy genes converge into common networks and play key roles in neurogenesis, neuronal migration and synaptic functions [1, 11]. Hence, other involved players and protein families are suitable candidates for novel ID and epilepsy disorders.
The evolutionary highly conserved cytoplasmic FMRP interacting proteins (CYFIPs) represent one such candidate protein family for ID and epilepsy. Its two members in humans, CYFIP1 (MIM 606322) and CYFIP2 (MIM 606323) (also known as PIR121), interact with the fragile X mental retardation protein (FMRP), a RNA-binding protein with an important role in translational control, the absence of which leads to fragile X syndrome (MIM 300624) [12]. Additionally, CYFIP1 and CYFIP2 were detected at the synapse [12] and are members of the canonical WAVE regulatory complex (WRC). This complex is activated by interaction of CYFIP with the small Rho GTPase Rac1 [13,14,15,16] and dominant-negative and constitutively active RAC1 variants have been recently reported in ID (MIM 617751) [17]. The WRC is a key regulator of actin dynamics [18] and missense and/or loss-of-function variants in two non-muscle actins, ACTB and ACTG1, cause syndromic forms of ID (MIM 243310 and 614583) [19, 20]. Given their role as “link” between Rac1, the WAVE complex, and FMRP, the CYFIP proteins have already been proposed as good candidates for ID and autism [13]. However, the focus remained on the more extensively studied CYFIP1, which is also one of the four genes deleted in the 15q11.2 BP1–BP2 microdeletion syndrome located within the Prader-Willi/Angelman syndrome region [21]. Concerning CYFIP2, only recently three variants affecting the Arginine at position p.87 have been published as causative in four patients with early-onset epileptic encephalopathy [22].
Using WES or WGS in trios of an affected child and its healthy parents, we independently identified eight distinct, spatially clustering de novo variants in CYFIP2 in a total of 12 patients delineating the genetic and clinical spectrum associated with variants in CYFIP2.
Methods
Subjects and high-throughput sequencing
The twelve patients underwent clinical examination at centers in Switzerland, Germany, The United Kingdom, Estonia, the USA, and France. Trio WES or WGS was either performed on a routine diagnostic basis with subsequent consent for publication or as part of a research study approved by the local ethics commissions on human research and were in keeping with international standards. A detailed clinical description of the patients is provided in Table 1 and the case reports in the supplement.
Patient 1 (P1) was part of a study including 63 unrelated patients with early-onset epileptic encephalopathy (EE) and trio WES was carried out as described elsewhere [23, 24]. P2, P4, and P5 were part of eight cases with de novo CYFIP2 rare nonsynonymous variants identified in 2793 cases with ID/DD and seizures who underwent trio WES by the GeneDx Laboratory (Gaithersburg, MD, USA) as described elsewhere (supplement and Tanaka et al. [25]). The remaining five cases could not be included into this study due to lack of consent for publication. The variant reported in P3 was identified by trio WGS conducted at the HudsonAlpha Institute for Biotechnology (Huntsville, AL, USA) in a cohort of 437 patients with ID/DD or other rare undiagnosed conditions. Analysis was performed as described elsewhere [26]. P6 was one of 128 diagnostic Estonian patients with developmental delay (DD), dysmorphism and/or other developmental problems and the CYFIP2 variant was detected by trio WES performed by the Genomics Platform at the Broad Institute of MIT and Harvard (details see supplemental methods). P7 and P9 underwent trio WES as part of the Deciphering Developmental Disorders (DDD) project as previously described [6]. The trio exome data of P9 were re-examined as part of a local ‘solving the unsolved’ project in Manchester using a previously described pipeline [27]. P8 was one of 1312 patients with ID and epilepsy that underwent trio WES at Ambry Genetics (Aliso Viejo, CA, USA) as described elsewhere [28]. WES of P10 and P11 and their parents was performed as previously described (supplement and Kremer et al. [29]). The variant in P12 was detected by trio WES using a MedExome (Roche, Madison, WI, USA) and a NextSeq500 (Illumina, San Diego, CA, USA) at the NGS platform of the Lyon University Hospital.
Throughout the text, figures, and tables, all reported variants are designated for the CYFIP2 reference transcript NM_001291722.1, which encodes for NP_001278651.1, following the current HGVS nomenclature (varnomen.hgvs.org) with exons numbered from 1 to 32 consecutively, and NC_000005.10 was used as a genomic reference sequence for the intronic variant. All reported variants have been submitted to the Leiden Open Variation Database (www.LOVD.nl/CYFIP2, patient IDs 00180890–00180901).
RT-PCR on RNA
To confirm the predicted splice effect of the variant c.3669+1G>T detected in P2, RNA from the patient and six controls was extracted using the PAXgene System (PreAnalytiX, Hombrechtikon, Switzerland) and transcribed reversely using Superscript III (ThermoFisher Scientific, Waltham, MA, USA). RT-PCR was performed by using specific primers located in the exons 29 and 32 of CYFIP2 and products were analyzed by agarose gel electrophoresis. Exon skipping in the patient was verified by Sanger sequencing of amplified products.
Variant modelling
The effect of the variants was investigated on the basis of the crystal structure of the WAVE regulatory complex (PDB: 3P8C [14], 4N78 [30]). This complex was crystallized with CYFIP1, which was replaced by CYFIP2 (88% sequence identity to CYFIP1) in our model using Modeller 9.16 [31]. Chimera [32] was used for structure analysis and visualization. The effect of the variants on the CYFIP2-WAVE1 binding affinity were assessed using BindProfX [33].
Results
Genetic findings
Through our trio WES study, utilizing GeneMatcher [34], Decipher Database [35], and personal communications, we identified 12 patients with eight distinct de novo CYFIP2 variants (seven missense and one splice site variant): c.259C>T p.(Arg87Cys) in P1, P3, P4, and P8; c.322T>C p.(Tyr108His) in P12; c.1363G>C p.(Ala455Pro) in P10; c.1992C>G p.(Ile664Met) in P6 and P9; c.1993G>A p.(Glu665Lys) in P7; c.2170G>C p.(Asp724His) in P11; c.2174A>G p.(Gln725Arg) in P5; and c.3669+1G>T p.(?) in P2 (Fig. 1, Table 1, for alternative nomenclature see Supplemental Table S1). Notably, all the variants are absent from GnomAD [36] and all except p.(Tyr108His) are predicted unanimously to have a deleterious effect on protein function by automated online prediction tools (missense variant predictions: SIFT, PolyPhen2, MutationTaster; splice variant predictions: MaxEnt, NNSplice, Human Splicing Finder, GeneSplicer, SpliceSiteFinder-like; all predictions provided by Alamut Visual Version 2.10 July 2017 (Interactive Biosoftware, Rouen, France)). Furthermore, the CYFIP2 gene is indicated to be intolerant to both, missense (z = 6.15) and loss-of-function (LoF) (probability of LoF intolerance pLI = 1.00) variants in ExAC [36].
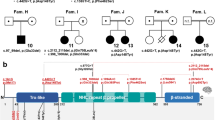
Gene and protein structure of CYFIP2 with the eight variants identified in twelve patients. a Schematic drawing of the CYFIP2 gene as well as CYFIP2 protein with its conserved domains. The position of the eight variants identified in our twelve patients (indicated in black) as well as the recently published variants (Nakashima et al. [22], indicated in grey and with †) are depicted. All variants occurred de novo and have been detected in heterozygous state (variant nomenclature and gene structure according to NM_001291722.1 with exons numbered from 1 to 32 consecutively and NC_000005.10 for intronic sequences, protein structure according to NP_001278651.1 and NCBI's conserved domain database [51]). b, c Splice variant c.3669+1G>T affecting the conserved splice donor site after exon 31 induces skipping of exon 31 in P2. RT-PCR on RNA from peripheral blood leukocytes with primers located in the exons 29 and 32 (indicated as arrows) resulted in an additional aberrant product of 381 bp in the patient (lane 1), whereas the amplification in six controls (lanes 2–7) resulted in the expected 529 bp fragment only (S, size standard; lane 8, genomic DNA control; lane 9, no template control). Exon skipping in the patient was verified by the sequencing of amplified products (Supplemental Figure S1) and is predicted to result in a frameshift followed by a premature stop codon after 3 altered amino acids (NM_001291722.1:r.3522_3669del, p.(Glu1174Aspfs*3))
The variant c.3669+1G>T is predicted to affect the conserved splice donor site in the last intron of CYFIP2, and RT-PCR performed using RNA from peripheral blood leukocytes of P2 confirmed skipping of exon 31 (NM_001291722.1:r.3522_3669del) (Fig. 1 and Supplemental Figure S1). Lack of exon 31, which is the second to last exon, is predicted to result in a frameshift followed by a premature stop codon (p.(Glu1174Aspfs*3)). Accordingly, this aberrant transcript is not expected to be subject to nonsense-mediated mRNA decay [37].
Variant modelling
The missense variants p.(Arg87Cys) and p.(Tyr108His) are located in the DUF1394 subdomain of CYFIP2, whereas the remaining missense variants (p.(Ala455Pro), p.(Ile664Met), p.(Glu665Lys), p.(Asp724His), and p.(Gln725Arg)) are located in the FragX_IP subdomain (Fig. 1). In the three-dimensional fold of CYFIP2, both the subdomains are tightly interlocked, placing Arg87 in close spatial proximity to Ile664, Glu665, Asp724, and Gln725. Moreover, these variants are located at the CYFIP2-WAVE1 interface (Fig. 2a, b). As noted by Nakashima et al. [22], the Arg87 sidechain forms tight interactions with two glutamate residues and intermolecular interactions to Tyr151 of WAVE1. Since these interactions are lost in the p.(Arg87Cys) variant, this exchange is predicted to cause structural instability around the variant site thereby leading to aberrant WAVE1 activation [22].
a Structure of the WAVE regulatory complex (WRC) indicating the sites of variants. CYFIP is shown in orange and the C-terminus, which is absent in the p.(Glu1174Aspfs*3) splice variant, is highlighted in red. The C-terminus forms tight interactions to the NCKAP1 protein (shown in cyan space-filled presentation). Residues affected by missense variants are shown in space-filled presentation and colored by atom types. Residues Arg87, Ile664, Glu665, Asp724 and Gln725 interact with WAVE1 (shown in green space-filled presentation) and Ala455 interacts with NCKAP1. Tyr108 is located in close vicinity to the CYFIP-WAVE1 interface. b Close-up view on the CYFIP2-WAVE1 interaction site disturbed by most of the variants observed. WAVE1 residues 151-173 comprising helix α6 (important for VCA domain stabilization) and residues 531-543 corresponding to the C-helix (part of VCA domain) are shown in light green and dark green, respectively. The interacting CYFIP residues Arg87, Ile664, Glu665, Asp724, and Gln725 are shown in space-filled presentation and colored by atom types. c Close-up view of the CYFIP2-WAVE1 interface in the vicinity of Tyr108. Residues 133-142 of WAVE1 are shown in green and residues 100-114 of CYFIP2 are shown in orange (ribbon representation). Key residues discussed in text are shown in space-filled presentation and Tyr108 is colored by atom type. d Close-up view on the CYFIP2-NCKAP1 interface in the vicinity of Ala455. NCKAP1 is shown in cyan and residues 444-469 of CYFIP2 are shown as orange ribbon with Ala455 in space-filled presentation
In contrast to Arg87, residues Ile664, Glu665, Asp724, and Gln725 form weaker intramolecular interactions suggesting that the variants found at these positions are not expected to significantly affect the stability of CYFIP2 itself. Nevertheless, all the five residues form direct interactions with WAVE1, which are predicted to be disturbed by these variants (Fig. 2a). A closer inspection reveals that interactions are formed with α6- and C-helices of WAVE1 (Fig. 2b). The C-helix (residues 531-543) is a part of the WAVE1 VCA-region and was shown to be critical for activation of the Arp2/3 complex [38]. The tight interaction between helices α6 and C plays an important role for stabilizing the VCA-region in an inactive conformation. Thus, these five missense variants are predicted to weaken the CYFIP2–WAVE1 interface at this functionally important site favoring release of the VCA-region leading to WAVE1 activation. To quantify the effect of the variants we calculated the changes in the free energy of the CYFIP2–WAVE1 interaction for each of the WAVE1 interface variants (free energy change in kcal/mol): p.(Arg87Cys) 1.75; p.(Ile664Met) 1.20; p.(Glu665Lys) 1.32; p.(Asp724His) 2.08; and p.(Gln725Arg) 1.09. Here we report changes in CYFIP2–WAVE1 affinity instead of CYFIP2 stability, which explains the difference in free energy reported in this manuscript for p.(Arg87Cys) (1.75 kcal/mol) compared with that reported by Nakashima et al. (~4.5 kcal/mol) [22].
Tyr108 is located close to the CYFIP2–WAVE1 interface and the adjacent residue Ile107 forms interactions with Tyr140 in the meander region of WAVE1 (Fig. 2c), which cooperatively stabilizes the VCA-element. These interactions are critically affected by alterations of the adjacent residues. For example, phosphorylation of Thr138 (WAVE1) by Cdk5 contributes to WRC activation and leads to altered cellular actin dynamics [14]. Thus, it appears feasible that replacement of Tyr108 (CYFIP2) by a charged histidine also perturbs the CYFIP–WAVE interaction at this functionally important site. Consequently, the variant p.(Tyr108His) may also lead to increased WAVE activation.
The splice variant c.3669+1G>T likely results in a C-terminally truncated CYFIP2 protein (p.(Glu1174Aspfs*3)). The C-terminus (red part illustrated in Fig. 2a) forms numerous contacts to NCKAP1, another component of the WRC, that are lost in the altered protein. A loss of interactions with NCKAP1 is also expected for the p.(Ala455Pro) missense variant, which is located in the CYFIP2-NCKAP1 interface (Fig. 2d). The calculated free energy change of 1.21 kcal/mol is in the same range as that observed for the variants in the CYFIP2-WAVE1 interface. Since Ala455 is located in an α-helix, an Ala455Pro exchange is additionally expected to disturb the structure of CYFIP2 itself, thus enhancing the damaging effect of this variant.
Clinical findings
Phenotype data are summarized in Table 1 and more detailed patient reports are provided in the supplement. The patients shared a phenotype of ID/DD (12/12) and epilepsy (11/12) except for P9, which was only reported to have had a single febrile seizure at the age of 6 months. In the majority of the cases (8/11), seizure onset occurred within the first year of life (range 5 weeks–4 years). In six patients seizures were intractable, whereas five children were reported to be currently seizure-free. All patients showed generalized or truncal hypotonia, in four cases combined with limb spasticity. Dysphagia with dependency on gastric tube feeding was present in five of 12 patients. Nine of 12 patients had visual impairment and/or strabismus. Eight patients developed absolute or relative microcephaly and in two patients the centile of head circumference decreased markedly. Common morphological features shared among patients in this cohort included long, tapered fingers, high forehead, narrow, mildly up-slanting palpebral fissures, apparent hypertelorism, bulbous nasal tip, full cheeks, everted lip vermillion, and retrognathia (Fig. 3). Brain MRI was performed in 11 of the patients and showed unspecific structural anomalies in six patients including cerebellar, white matter or diffuse atrophy in four patients, dysmorphic hemispheres and frontal lobe hypoplasia in one patient, and hypoplasia of the frontal lobe, the chiasma and retrochiasmatic region in one patient. Three patients had normal cerebral MRI scans at the age of 5 weeks (P3), 7 months and 1 year 1 month (P6), and 3 years 5 months (P7), respectively. EEG was available for all 11 patients with recurrent seizures and showed variable findings including focal, multifocal and generalized epileptiform discharges, and hypsarrhythmia. Developmental milestones were absent or delayed in all 12 patients. Only 6/12 patients achieved unassisted sitting and 2/12 were able to walk. Nine of 12 patients did not develop any verbal communication and one spoke only a few words at the age of 6 years. Psychomotor regression was reported in one patient (P2), who was standing and cruising and then regressed at the age of 4 years after a single tonic-clonic seizure event. All patients are alive and their age at last evaluation ranged from 1 year 4 months to 9 years.
Discussion
In this study we report 12 patients harboring eight de novo variants in CYFIP2 presenting with mostly severe but variable ID/DD, muscular hypotonia and, in all except one, a history of epilepsy with onset <4 years. Other commonly shared features included secondary microcephaly, limb spasticity, dysphagia, long tapering fingers, and similar facial traits (Table 1, Fig. 3).
Our findings are in line with those recently reported by Nakashima et al. [22], who identified concurrently three de novo variants in patients with epileptic encephalopathy, all affecting the Arginine at position 87 in CYFIP2. All four of their patients had early-onset epileptic encephalopathy classified as Ohtahara or West syndrome, whereas we report a broader clinical spectrum ranging from a similarly severe phenotype including intractable epilepsy and profound ID from mild to moderate cognitive impairment (P7 & P9) without epilepsy (P9) (Table 1). The p.(Arg87Cys) variant recurred in four cases of this study, and was also reported twice in the previous cohort [22]. All six patients carrying this recurrent variant, as well as two further patients [22] with other substitutions of the p.Arg87 had profound ID, early-onset epilepsy and hypotonia. Seizures were intractable in six of these eight patients while two are currently seizure-free. Six of these eight patients had secondary microcephaly. MRI findings ranged from unremarkable to hypomyelination and atrophy. All four patients reported here carrying the p.(Arg87Cys) variant showed sleep disturbances, while this feature was not addressed by the previous report. Of note, this variant was also reported as a candidate variant in a patient with West syndrome who in addition harbored further de novo variants including a variant classified as “likely pathogenic” in the Kabuki syndrome (MIM 147920)-associated KMT2D gene [39]. We identified a further recurrent variant, p.(Ile664Met), present in two of our patients (P6 and P9) and associated with a more variable phenotype consisting of either severe developmental and epileptic encephalopathy or moderate non-epileptic ID, respectively. Therefore, currently available data indicate a profound phenotype in p.Arg87 substitutions, but a more variable phenotype in other variants.
The CYFIP2 gene is located in the chromosomal region 5q33.3 and encodes the cytoplasmic FMRP interacting protein 2 (CYFIP2). Only large segmental chromosomal aneusomies involving this region have been clinically described in two patients with ID and seizures, with a vast number of genes affected and unclear deletion borders in one patient [40, 41]. With regards to single gene defects, the study by Nakashima et al. [22] and this study are the first reports to establish defects in CYFIP2 as the underlying cause of severe neurodevelopmental disorders. Since neither elsewhere nor in our series small copy number or truncating variants triggering nonsense-mediated mRNA decay have been observed, haploinsufficiency seems unlikely as disease mechanism in the observed severe phenotype.
Both members of the highly conserved CYFIP protein family in humans, CYFIP1 and CYFIP2, interact with FMRP and are similarly co-localized with ribosomes and FMRP and were also detected at the synapse of mouse brain [12, 42]. However, despite their high homology it was shown that CYFIP1 interacts exclusively with FMRP, while CYFIP2 also interacts with the FMRP-related proteins FXR1P/2P [12]. Interestingly, it has been shown that regardless of unchanged RNA levels, the protein levels of CYFIP2 are increased in the blood of Fragile X patients [43]. This is in line with the fact that Cyfip2 mRNA (but not Cyfip1 mRNA) is ranked as a FMRP target in mouse brain [44] and the conception that FMRP acts as a suppressor of CYFIP2 translation by sequestration of its mRNA.
CYFIP1/2, along with the WAVE proteins (also known as WASF proteins), NCKAP1 (NAP1 or HEM1 in hematopoietic cells), ABI2 (or its paralogous proteins, ABI1 or NESH), and HSPC300 (BRK1), are members of the canonical WAVE regulatory complex (WRC), a key regulator of actin dynamics (Fig. 4) [13, 14, 16]. By the interaction of Rac1-GTP with CYFIP, the intrinsically inactivated complex is split into one subcomplex including CYFIP1/2, which is now capable to interact with other proteins, and another subcomplex including WAVE, which interacts with Arp2/3 triggering actin polymerization [13,14,15,16]. Here, the Arp2/3 complex is activated by binding the conserved VCA (verprolin homology, central and acidic regions) domain of WAVE, which is sequestrated in the inactivated complex by CYFIP, inhibiting WAVE activity [14, 15]. Actin filaments are important cytoskeletal structures in neurons with critical dynamics and especially the Arp2/3 complex is essential at multiple stages of neural development [45, 46].
Schematic structural organization of the WAVE regulatory complex (WRC) with its members WAVE1, CYFIP2, NCKAP1, ABI2, and HSPC300 in inactivated condition (left part) and after activation of WAVE function by RAC1-GTP binding (upper right part) or aberrant WAVE activation by CYFIP2 mutation as suggested by our data (lower right part)
Accordingly, in vitro and in vivo animal models of CYFIP2 (or its orthologs) showed phenotypes consistent with its discussed role as FMRP interactor and component of a key regulator of actin cytoskeleton, such as abnormalities of dendritic spines and axons [12, 13, 42, 47,48,49,50]. Of note, while homozygous null mice were lethal, indicating the necessity of CYFIP2 in early development, a heterozygous knock-out mouse showed abnormal behavior and cortical dendritic spines similar to that observed in Fmr1-null mice [47]. Given these findings and the high pLI score of 1.0 for CYFIP2, we assume that haploinsufficiency in humans may also cause a developmental phenotype. However, the apparently milder phenotype in the heterozygous Cyfip2 knock-out mouse model would be in line with the assumption that the observed severe human CYFIP2 phenotype is not caused by haploinsufficiency.
Despite the large spacing in the primary structure, the residues affected by the missense variants p.(Arg87Cys), p.(Ile664Met), p.(Glu665Lys), p.(Asp724His), and p.(Gln725Arg) in our patients are located in close spatial proximity at the CYFIP2–WAVE1 interface, where they are predicted to impair binding of the WAVE VCA-region (Fig. 2b), therefore leading to increased WAVE activation and consecutively to increased Arp2/3 mediated actin polymerization. A similar effect is also expected for the p.(Tyr108His) variant, although caused by a slightly different structural mechanism. Of note, an enhanced WAVE net activity was already suggested after loss of Cyfip2 in a heterozygous mouse model [47] or dCyfip in Drosophila [49]. Increased WAVE activity was also suggested for the p.Arg87 variants observed by Nakashima et al. [22] supported by their finding of speckled co-localization of filamentous actin and mutant CYFIP2 as well as significantly increased aberrant filamentous actin accumulation in B16F1 cells transfected wildtype or mutant CYFIP2. However, they could not detect abnormal WAVE1 binding of the Arg87 mutants by co-immunoprecipitation in transfected HEK293T cells, but showed a consistently weaker interaction with the VCA domain in a GST pull-down assay.
The splice variant identified in P2 is predicted to result in a truncated protein (p.(Glu1174Aspfs*3)) due to exon skipping of the second to last exon. This transcript likely escapes nonsense-mediated decay. Based on our structural modelling the missing C-terminus is expected to lead to a loss of numerous contacts with NCKAP1 (also known as NAP1) within the WRC (Fig. 2) and therefore also to a disruption of the WRC and consecutive WAVE activation. A similar WRC disruption caused by loss of interactions with NCKAP1 is also expected for the p.(Ala455Pro) missense variant, which may be enhanced by decreased stability of CYFIP2.
Therefore, we speculate that the mechanism of aberrant WAVE activation caused by the different variants could be explained by activation of the Arp2/3-binding VCA domain either by (1) increased dissociation of the WRC and/or (2) disturbed interactions at the CYFIP2–WAVE1 interface favoring an exposed VCA-helix (Fig. 4). Given the variant type, the finding of unchanged CYFIP2 protein levels in a cell line of a p.(Arg87Leu) patient [22] and the results of our structural modelling, we assume that the detected variants act as partial loss-of-function variants in terms of their ability to stabilize the WRC and/or the inactive conformation of the VCA region, but eventually result in gain of function considering WAVE activation.
In conclusion, we provide evidence that a variety of spatially clustered de novo variants in CYFIP2 are causative for a new neurodevelopmental disorder characterized by intellectual disability and seizures. Our findings suggest that all identified de novo variants lead to increased WAVE activity by weakening the interaction with other components of the WAVE regulatory complex, a key regulator of the actin cytoskeleton.
References
van Bokhoven H. Genetic and epigenetic networks in intellectual disabilities. Annu Rev Genet. 2011;45:81–104.
Bowley C, Kerr M. Epilepsy and intellectual disability. J Intellect Disabil Res. 2000;44(Pt 5):529–43.
Reilly C, Atkinson P, Das KB, et al. Neurobehavioral comorbidities in children with active epilepsy: a population-based study. Pediatrics. 2014;133:e1586–93.
Chin RF, Cumberland PM, Pujar SS, Peckham C, Ross EM, Scott RC. Outcomes of childhood epilepsy at age 33 years: a population-based birth-cohort study. Epilepsia. 2011;52:1513–21.
Deciphering Developmental Disorders Study. Large-scale discovery of novel genetic causes of developmental disorders. Nature. 2015;519:223–8.
Deciphering Developmental Disorders Study. Prevalence and architecture of de novo mutations in developmental disorders. Nature. 2017;542:433–8.
Epi KC, Epilepsy Phenome/Genome P, Allen AS, et al. De novo mutations in epileptic encephalopathies. Nature. 2013;501:217–21.
Rauch A, Wieczorek D, Graf E, et al. Range of genetic mutations associated with severe non-syndromic sporadic intellectual disability: an exome sequencing study. Lancet. 2012;380:1674–82.
Yang Y, Muzny DM, Xia F, et al. Molecular findings among patients referred for clinical whole-exome sequencing. JAMA. 2014;312:1870–9.
Gilissen C, Hehir-Kwa JY, Thung DT, et al. Genome sequencing identifies major causes of severe intellectual disability. Nature. 2014;511:344–7.
Kochinke K, Zweier C, Nijhof B, et al. Systematic phenomics analysis deconvolutes genes mutated in intellectual disability into biologically coherent modules. Am J Hum Genet. 2016;98:149–64.
Schenck A, Bardoni B, Moro A, Bagni C, Mandel JL. A highly conserved protein family interacting with the fragile X mental retardation protein (FMRP) and displaying selective interactions with FMRP-related proteins FXR1P and FXR2P. Proc Natl Acad Sci Usa. 2001;98:8844–9.
Abekhoukh S, Bardoni B. CYFIP family proteins between autism and intellectual disability: links with Fragile X syndrome. Front Cell Neurosci. 2014;8:81.
Chen Z, Borek D, Padrick SB, et al. Structure and control of the actin regulatory WAVE complex. Nature. 2010;468:533–8.
Chen B, Chou HT, Brautigam CA, et al. Rac1 GTPase activates the WAVE regulatory complex through two distinct binding sites. ELife 2017; 6:1–22.
Derivery E, Lombard B, Loew D, Gautreau A. The Wave complex is intrinsically inactive. Cell Motil Cytoskelet. 2009;66:777–90.
Reijnders MRF, Ansor NM, Kousi M. et al. RAC1 missense mutations in developmental disorders with diverse phenotypes. Am J Hum Genet. 2017;101:466–77.
Lee Y, Kim D, Ryu JR, et al. Phosphorylation of CYFIP2, a component of the WAVE-regulatory complex, regulates dendritic spine density and neurite outgrowth in cultured hippocampal neurons potentially by affecting the complex assembly. Neuroreport. 2017;28:749–54.
Riviere JB, van Bon BW, Hoischen A, et al. De novo mutations in the actin genes ACTB and ACTG1 cause Baraitser-Winter syndrome. Nat Genet. 2012;44:440–4. S441-442
Cuvertino S, Stuart HM, Chandler KE, et al. ACTB loss-of-function mutations result in a pleiotropic developmental disorder. Am J Hum Genet. 2017;101:1021–33.
Cox DM, Butler MG. The 15q11.2 BP1-BP2 microdeletion syndrome: a review. Int J Mol Sci. 2015;16:4068–82.
Nakashima M, Kato M, Aoto K, et al. De Novo hotspot variants in CYFIP2 cause early-onset epileptic encephalopathy. Ann Neurol. 2018;83:794–806. https://doi.org/10.1002/ana.25208.
Plecko B, Zweier M, Begemann A, et al. Confirmation of mutations in PROSC as a novel cause of vitamin B 6 -dependent epilepsy. J Med Genet. 2017;54:809–14.
Papuc SM, Abela L, Steindl K et al. The role of recessive inheritance in early-onset epileptic encephalopathies: a combined whole-exome sequencing and copy number study. Eur J Hum Genet. submitted. 2018. https://doi.org/10.1038/s41431-018-0299-8.
Tanaka AJ, Cho MT, Millan F, et al. Mutations in SPATA5 are associated with microcephaly, intellectual disability, seizures, and hearing loss. Am J Hum Genet. 2015;97:457–64.
Bowling KM, Thompson ML, Amaral MD, et al. Genomic diagnosis for children with intellectual disability and/or developmental delay. Genome Med. 2017;9:43.
Faundes V, Newman WG, Bernardini L. et al. Histone lysine methylases and demethylases in the landscape of human developmental disorders. Am J Hum Genet. 2018;102:175–87.
Farwell KD, Shahmirzadi L, El-Khechen D, et al. Enhanced utility of family-centered diagnostic exome sequencing with inheritance model-based analysis: results from 500 unselected families with undiagnosed genetic conditions. Genet Med. 2015;17:578–86.
Kremer LS, Bader DM, Mertes C, et al. Genetic diagnosis of Mendelian disorders via RNA sequencing. Nat Commun. 2017;8:15824.
Chen B, Brinkmann K, Chen Z, et al. The WAVE regulatory complex links diverse receptors to the actin cytoskeleton. Cell . 2014;156:195–207.
Webb B, Sali A. Protein structure modeling with MODELLER. Methods Mol Biol. 2017;1654:39–54.
Pettersen EF, Goddard TD, Huang CC, et al. UCSF Chimera:a visualization system for exploratory research and analysis. J Comput Chem. 2004;25:1605–12.
Xiong P, Zhang C, Zheng W, Zhang Y. BindProfX: assessing mutation-induced binding affinity change by protein interface profiles with pseudo-counts. J Mol Biol. 2017;429:426–34.
Sobreira N, Schiettecatte F, Valle D, Hamosh A. GeneMatcher: a matching tool for connecting investigators with an interest in the same gene. Hum Mutat. 2015;36:928–30.
Firth HV, Richards SM, Bevan AP, et al. DECIPHER: database of chromosomal imbalance and phenotype in humans using ensembl resources. Am J Hum Genet. 2009;84:524–33.
Lek M, Karczewski KJ, Minikel EV, et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature. 2016;536:285–91.
Khajavi M, Inoue K, Lupski JR. Nonsense-mediated mRNA decay modulates clinical outcome of genetic disease. Eur J Hum Genet. 2006;14:1074–81.
Panchal SC, Kaiser DA, Torres E, Pollard TD, Rosen MK. A conserved amphipathic helix in WASP/Scar proteins is essential for activation of Arp2/3 complex. Nat Struct Biol. 2003;10:591–8.
Peng J, Wang Y, He F, et al. Novel West syndrome candidate genes in a Chinese cohort. CNS Neurosci Ther. 2018;24:1196–1206. https://doi.org/10.1111/cns.12860.
Lee JH, Kim HJ, Yoon JM, et al. Interstitial deletion of 5q33.3q35.1 in a boy with severe mental retardation. Korean J Pediatr. 2016;59:S19–S24.
Spranger S, Rommel B, Jauch A, Bodammer R, Mehl B, Bullerdiek J. Interstitial deletion of 5q33.3q35.1 in a girl with mild mental retardation. Am J Med Genet. 2000;93:107–9.
Pathania M, Davenport EC, Muir J, Sheehan DF, Lopez-Domenech G, Kittler JT. The autism and schizophrenia associated gene CYFIP1 is critical for the maintenance of dendritic complexity and the stabilization of mature spines. Transl Psychiatry. 2014;4:e374.
Hoeffer CA, Sanchez E, Hagerman RJ, et al. Altered mTOR signaling and enhanced CYFIP2 expression levels in subjects with fragile X syndrome. Genes Brain Behav. 2012;11:332–41.
Darnell JC, Van Driesche SJ, Zhang C, et al. FMRP stalls ribosomal translocation on mRNAs linked to synaptic function and autism. Cell . 2011;146:247–61.
Lei W, Omotade OF, Myers KR, Zheng JQ. Actin cytoskeleton in dendritic spine development and plasticity. Curr Opin Neurobiol. 2016;39:86–92.
Chou FS, Wang PS. The Arp2/3 complex is essential at multiple stages of neural development. Neurogenesis (Austin). 2016;3:e1261653.
Han K, Chen H, Gennarino VA, Richman R, Lu HC, Zoghbi HY. Fragile X-like behaviors and abnormal cortical dendritic spines in cytoplasmic FMR1-interacting protein 2-mutant mice. Hum Mol Genet. 2015;24:1813–23.
Pittman AJ, Gaynes JA, Chien CB. nev (cyfip2) is required for retinal lamination and axon guidance in the zebrafish retinotectal system. Dev Biol. 2010;344:784–94.
Zhao L, Wang D, Wang Q, Rodal AA, Zhang YQ. Drosophila cyfip regulates synaptic development and endocytosis by suppressing filamentous actin assembly. PLoS Genet. 2013;9:e1003450.
Tiwari SS, Mizuno K, Ghosh A, et al. Alzheimer-related decrease in CYFIP2 links amyloid production to tau hyperphosphorylation and memory loss. Brain. 2016;139:2751–65.
Marchler-Bauer A, Bo Y, Han L, et al. CDD/SPARCLE: functional classification of proteins via subfamily domain architectures. Nucleic Acids Res. 2017;45:D200–D203.
Acknowledgements
The authors are grateful to the participating individuals and their families. We thank Christian Söldner and Michel Albert for their support with preparation of figures. AR, LA, and BP were supported by the University of Zurich research priority program radiz (rare disease initiative Zurich). AR was further supported by the Swiss National Science Foundation (SNSF) grant 320030_179547. The Broad Center for Mendelian Genomics (UM1 HG008900) is funded by the National Human Genome Research Institute with supplemental funding provided by the National Heart, Lung, and Blood Institute under the Trans-Omics for Precision Medicine (TOPMed) program and the National Eye Institute. Support also provided by NIH T32 GM007748 (M.H.W.). The DDD Study (Cambridge South REC approval 10/H0305/83 and the Republic of Ireland REC GEN/284/12) presents independent research commissioned by the Health Innovation Challenge Fund (grant number HICF-1009-003), a parallel funding partnership between the Wellcome and the Department of Health, and the Wellcome Sanger Institute (grant number WT098051). The research team acknowledges the support of the National Institute for Health Research, through the Comprehensive Clinical Research Network. This study makes use of DECIPHER (http://decipher.sanger.ac.uk), which is funded by the Wellcome. The views expressed in this publication are those of the author(s) and not necessarily those of the Wellcome or the Department of Health. AB was supported by the Forschungskredit UZH and the Josef Huwyler Ruth Bernet-Engeli Stiftung.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Conflict of interest
Kirsty McWalter, Megan T. Cho, Maria J. Guillen Sacoto, and Kristin G. Monaghan are employees of GeneDx, Inc., a wholly owned subsidiary of OPKO Health, Inc. The remaining authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Zweier, M., Begemann, A., McWalter, K. et al. Spatially clustering de novo variants in CYFIP2, encoding the cytoplasmic FMRP interacting protein 2, cause intellectual disability and seizures. Eur J Hum Genet 27, 747–759 (2019). https://doi.org/10.1038/s41431-018-0331-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41431-018-0331-z
This article is cited by
-
Transcriptomic alterations in cortical astrocytes following the development of post-traumatic epilepsy
Scientific Reports (2024)
-
Orchestration of synaptic functions by WAVE regulatory complex-mediated actin reorganization
Experimental & Molecular Medicine (2023)
-
Structures reveal a key mechanism of WAVE regulatory complex activation by Rac1 GTPase
Nature Communications (2022)
-
Cytoplasmic FMR1 interacting protein (CYFIP) family members and their function in neural development and disorders
Molecular Biology Reports (2021)
-
Artificial intelligence enables comprehensive genome interpretation and nomination of candidate diagnoses for rare genetic diseases
Genome Medicine (2021)