Abstract
In recent years, chromosomal microarray analysis has permitted the discovery of rearrangements underlying several neurodevelopmental disorders and still represents the first diagnostic test for unexplained neurodevelopmental disabilities. Here we report a family of consanguineous parents showing psychiatric disorders and their two sons both affected by intellectual disability, ataxia, and behavioral disorder. SNP/CGH array analysis in this family demonstrated in both siblings a biallelic duplication inherited from the heterozygous parents, disrupting the ADGRB3 gene. ADGRB3, also known as BAI3, belongs to the subfamily of adhesion G protein-coupled receptors (adhesion GPCRs) that regulate many aspects of the central nervous system, including axon guidance, myelination, and synapse formation. Single nucleotide polymorphisms and copy number variants involving ADGRB3 have recently been associated with psychiatric disorders. These findings further support this association and also suggest that biallelic variants affecting the function of the ADGRB3 gene may also cause cognitive impairments and ataxia.
Similar content being viewed by others
Introduction
Both gains and losses of dosage-sensitive genes, and also the genomic disruption of regulatory associated domains [1, 2] leading to a dysregulation of gene expression, represent the principal pathogenic mechanisms of disease-causing copy number variants (CNVs). While most of them behave as dominant alleles and have been identified in sporadic diseases, recessive CNVs have also been associated with several disorders, either in a homozygous state or as associated with a recessive variant in the other allele. Not surprisingly, heterozygous carriers of deletions involving recessive genes are frequent in the general population [3], and homozygous CNVs encompassing recessive disease genes have been reported in consanguineous pedigrees [4, 5].
Herein we report two affected siblings born to consanguineous parents sharing a similar phenotype, characterized by intellectual disability, cerebellar atrophy with predominantly vermis involvement, ataxia, strabismus, and behavioral disorder. Single nucleotide polymorphism/comparative genomic hybridization (SNP/CGH) array analysis demonstrated in both siblings a homozygous duplication of chromosome 6q12 involving the ADGRB3 (MIM 602684) gene, inherited from the heterozygous parents. A subsequent analysis of the breakpoint regions clearly indicated a direct tandem duplication disrupting the genomic structure of the gene.
ADGRB3 is a member of a subfamily of adhesion G protein-coupled receptors (adhesion GPCRs) that play a key role in the regulation of several aspects of the central nervous system, such as axon guidance, myelination, and synapse formation and function [6]. The subfamily ADGRB includes ADGRB1, ADGRB2, and ADGRB3 genes (also known as BAI1, BAI2, and BAI3, respectively), which are widely expressed in the postnatal and adult brain. ADGRB1 and ADGRB3 are enriched in the postsynaptic density while ADGRB3 is also highly expressed in the cerebellum [7, 8]. Interestingly, ADGRB3 has already been associated with schizophrenia, bipolar disorder, and drug addiction [9,10,11,12]. Here we report the first instance of a homozygous disruption of ADGRB3 and discuss the possible role of this gene in the pathogenesis of the disease in this family.
Subjects and methods
Family
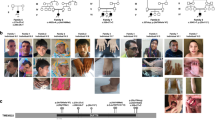
The study was conducted in accordance with the Declaration of Helsinki and national guidelines. A written informed consent was obtained from each subject or their guardian, and the study was approved by the local ethical committee. The parents of the affected probands were first cousins once removed. The extended pedigree is shown in Fig. 1, and clinical features of the affected individuals are summarized in Table 1. Rearrangements and phenotypic details were submitted to DECIPHER (https://decipher.sanger.ac.uk/).
Patient V:1 was a 36-year-old woman (Decipher ID 367222). She was born at term by cesarean delivery after a normal pregnancy; she had mild respiratory distress at birth. She walked at the age of 14 months with an unstable gait. At the age of 5 years, electromyography and muscular biopsy gave normal results. At the age of 6 years, strabismus and tremors were evident in her hands.
She was initially admitted to our hospital at age 16, when she presented with borderline intellectual functioning, memory difficulties, cerebellar ataxia, fatigue and affective emotional issues, such as anxiety and mood reduction.
Brain magnetic resonance imaging (MRI) documented moderate cerebral and cerebellar atrophy, with predominant vermis involvement (Fig. 2a, c).
Electroencephalogram (EEG), EMG, plasma amino acids, isoelectric focusing of transferrin, urinary organic acids, long-chain fatty acids, phytanic acid, plasma carnitine profile, molecular genetic screening for SCA 1, 2, 3, 6, 7, 8, and Friedreich ataxia were normal.
At age 26, tremor was reduced due to the treatment with levetiracetam. A further brain MRI scan at age 34 was similar to the previous one (Fig. 2d, f).
At the last neurological assessment she had scanning speech, mild muscular hypotonia in the upper limbs, mild diffuse muscular weakness, hyporeflexia in the upper limbs, hyperreflexia in the lower limbs, postural and intentional tremor in her head, trunk and upper limbs, dysmetria, dysdiadochokinesia, autonomous ambulation with ataxic gait, deficiency of motor and ocular coordination, lumbar hyperlordosis, mild lumbar scoliosis, and flat feet.
Audiological examination was normal.
An ophthalmological examination showed astigmatism and strabismus. The fundus oculi was normal. She showed a mild intellectual disability (IQ = 55, ICV = 88) on WAIS IV (Wechsler, 2008) [13]. In the psychopathological sphere, mood alterations and behavioral concerns giving rise to an anxious/depressive disorder were found. On the Vineland Adaptive Behavior Scale [14] her overall Adaptive Behavior Composite was 72 (normal mean: 100).
Patient V:2 (Decipher ID 367221) was a 32-year-old man born at term by eutocic delivery after a normal pregnancy, with no respiratory distress. At the age of 1 year, strabismus and tremor of the hands were noted. He started walking at the age of 18 months with an unstable gait and spoke his first words at 3 years of age.
Muscular biopsy, performed at the age of 3 years, showed myogenic changes. EEG, EMG, and neurometabolic screenings were reported normal.
At the age of 11 years, during measles infection, he had epileptic seizures with loss of consciousness, eye revulsion, and limb hypertonia for a few seconds.
He was initially admitted to our hospital at age 12, when he presented with dysarthric speech, intention tremor of the head, trunk and upper limbs, dysmetria, dysdiadochokinesia, autonomous ambulation with ataxic gait, and a deficiency of motor and ocular coordination. Brain MRI documented moderate cerebral and cerebellar atrophy, with predominant vermis involvement (Fig. 3a, c).
At age 19, soliloquy, inappropriate laughing, social closure, and hallucinations appeared, partially reduced by a treatment with clozapine. Thereafter, the gait and tremor worsened. EEG showed slow rhythms over frontal and central regions.
On neurological examination he presented psychomotor slowness, paralysis of orizzontal coniugate eye movements, dysarthric speech, hyporeflexia at the upper limbs, hypereflexia at the lower limbs, postural and intention tremor of the head, trunk and upper limbs, dysmetria, dysdiadochokinesia, autonomous ambulation with ataxic gait, and deficiency of motor and ocular coordination. Additionally, he had scapular winging, lumbar hyperlordosis, dorsal hyperkyphosis, and flat feet.
The ophthalmological examination revealed nystagmus, astigmatism, and strabismus. Audiological examination was normal while a further brain MRI scan was comparable to the previous one, showing a similar cerebral and cerebellar atrophy (Fig. 3d, f).
POLG gene analysis was negative.
A severe intellectual disability (IQ = 41) was diagnosed through the non-verbal intelligence scale Leiter 3 [15]. He presented with hypomodulated facial expression, and a slackened and restricted speech. Impairments due to an alteration in the emotional and instinctual sphere with an unusual behavior were observed. He developed soliloquy as well as nervousness and outbursts of anger, if contradicted. As for the adaptation sphere, he showed a need for constant support (Quotient at Vineland II = 20).
The 70-year-old father (III:2, Decipher ID 367225) was born at term by normal delivery after a normal pregnancy. Psychomotor development was normal. At age 40, hallucinations, mood alterations, aggressive reactions, psychomotor agitation, and insomnia appeared, partially reduced by neuroleptic treatment. Neurological examination was normal and brain MRI showed white-matter signal abnormalities, moderate cerebral atrophy, and mild cerebellar vermis atrophy (Supplementary Figure 1A, B). The patient presented with normal cognitive functioning, as confirmed by the scores at WAIS IV test (IQ = 79, ICV = 104). The qualitative alterations found in the psychopathology profile were typical of a psychotic disorder. More specifically, he presented impairments in vigilance reaction times and communication, slow ideation, working-memory and processing-speed deficit, as well as mood-alteration episodes and aggressive reactions. He showed a score of 103 on the Vineland II scale and age-appropriate skills in communication, socialization, and daily-life activities.
The patient’s 65-year-old mother (IV:1; Decipher ID 367223) was born at term by eutocic delivery after a normal pregnancy. Psychomotor development was normal. She was initially admitted to our hospital at age 55. She presented a normal cognitive functioning, an anxious/depressive disorder, and restless legs syndrome, which improved by treatment with pramipexole.
Brain MRI showed some focal white-matter abnormal signs, moderate cerebral atrophy, and mild cerebellar vermis atrophy (Supplementary Figure 2A, B). She showed normal cognitive functioning (IQ = 91) and verbal comprehension index (ICV = 88) on WAIS IV. The psychopathological profile revealed symptoms consistent with a diagnosis of anxious/depressive disorder. She presented with mild mood alterations and alertness for her safety and that of her family. Significantly high scores were found for mood and anxiety/somatic concern (percentile score = 91°) on BAI (Beck Anxiety Inventory) and BDI (Beck Depression Inventory; score = 33) [16, 17]. Normal standard score levels were obtained in the adaptation areas (Quotient on Vineland II = 108). The spatial and temporal orientation was within the normal range; she was able to take care of herself and carry out routine activities of daily living.
At the last examination, clinical and neuroradiological (Supplementary Figure 2C, D) findings were unchanged.
No other members of the extended family were reported as having psychiatric or neurological illness, except individual III-1 who was reported suffering from depression. However, she refused both clinical evaluation and DNA testing. Individuals III:4, III:8, and IV:2 were apparently healthy and were willing to undergo genetic testing.
Characterization of the rearrangement
Genomic DNA was extracted by standard procedures. SNP/CGH array assays were carried out using the combined array CGH plus SNP from CytoSure ISCA UPD array 4 × 180 K containing 6186 unique SNPs and 137,100 oligonucleotide probes, with a backbone consisting of one oligonucleotide probe every 25 kb and higher oligonucleotide density in targeted regions determined by the International Standards for Cytogenomic Array consortium [18]. The arrays were performed according to the manufacturer’s protocol version 7.1, using appropriate Agilent Reference DNAs (Euro male and Euro female). The arrays were analyzed with the Agilent Microarray Scanner, Feature Extraction Software version 11.5, and Agilent Genomic Workbench 7.0. (for genomic coordinates refer to assembly GRCh37/hg19). NM_001704.2 was used as reference for ADGRB3 exon numbering, according to UCSC genome browser (http://genome.ucsc.edu/cgi-bin/hgTracks?db=hg19).
To better refine the rearrangement, a multiplex ligation-dependent probe amplification (MLPA) assay was designed and carried out on probands and their parents and in a group of ten unrelated control individuals (reagents and enzymes were from MRC-Holland, Amsterdam, The Netherlands). MLPA fluorescent amplicons were identified and quantified as previously described [19]. This allowed us to define a minimal breakpoint region at both sides of the rearrangement.
In order to characterize the rearrangement breakpoint at nucleotide-level resolution, long-range PCR and sequence analysis were performed. Genomic DNA was amplified using Expand Long Template PCR System (Roche Applied Science, Mannheim, Germany) according to the manufacturer’s instructions. Considering either a direct or inverted duplication, different primer pairs were designed in the minimal breakpoint regions, previously masked with Repeat Masker (www.repeatmasker.org).
Fluorescent in situ hybridization (FISH) experiment was performed on the metaphase chromosome by standard techniques using BAC insert clone RP11-980I14 located within the CNV region and RP11-661C14 as control clone for the 6p chromosome, both purchased from Empire Genomics (Buffalo, NY, USA).
Whole-exome sequencing
Whole-exome sequencing (WES) was performed by an external service provider (Biodiversa, Rovereto (TN), Italy). According to the provider’s description, whole-exome enrichment was carried out using the Agilent Sure Select Human All Exon V6 + UTRs kit (Agilent Technologies, Santa Clara, CA, USA) following the kit’s recommendations and sequenced with the Illumina Novaseq sequencer (Illumina, San Diego, CA, USA) to generate 150-bp-paired end reads that were aligned to the human genome (UCSC GRCh37/hg19), using BWA-MEM [20]. GATK tools (https://software.broadinstitute.org/gatk/) [21] including base quality score recalibration, indel realignment, duplicate removal, and SNP) and insertions/deletion (INDEL) identification were used according to the recommendations of GATK Best Practices [22]. Variants were annotated using the ANNOVAR software tool [23] and then filtered according to their predicted effects and allele frequencies in the available public databases (dbSNP http://www.ncbi.nlm.nih.gov/projects/SNP/, ExAC http://exac.broadinstitute.org/, 1000 Genomes http://www.1000genomes.org/, ESP6500 http://evs.gs.washington.edu/EVS/). Assuming an autosomal recessive inheritance of a rare variant from a common ancestor because of parental consanguinity, homozygous variants shared by both the affected siblings were selected and prioritized taking into account their predicted functional impact and the biological role of the protein. The same analysis was also performed for compound heterozygote variants common to both the siblings. Finally, we also hypothesized that a germline mosaicism for a dominant allele in a parent could have been responsible for the trait and searched for variants showing non-Mendelian inheritance.
Results
SNP/CGH array analysis
The analysis discovered in both the probands a 6q12 gain of about 300 kb with a log2 signal ratio (0.9–1.0) suggestive of triplication of the genomic segment (HGSV: g.?_?insNC_000006.11:(69575808_69592667)_(69897263_69922585);[2] ISCN 2016: arr[GRCh37] 6q12(69575808 × 2,69592667_69897263 × 4,69922585 × 2) while the same experiment identified in both the parents a duplication of the same region (Fig. 4a). The rearrangement, so far unreported in the Database of Genomic Variants (DGV, http://projects.tcag.ca/variation/project.html), was confirmed in probands and in their parents by an ad hoc designed MLPA assay, which did not identify any abnormality in individuals III:4, III:8, and IV:2 (data not shown). According to human reference sequence (NCBI Build GRCh37), the proximal and distal breakpoints of the rearrangement were located inside the ADGRB3 gene.
a A graphical overview of the CGH array analysis of the duplication (from top to bottom: father, brother, and daughter are represented). The duplication involves exons 417 of ADGRB3 gene symbolized by boxes. b Schematic location of primers (arrows) designed for long-range PCR. Only the hypothesis of direct tandem duplication is shown. Sanger sequencing of the breakpoint region aligned with the reference sequences is shown at the bottom of the figure. The box indicates the three additional bases not present in the reference sequences
FISH experiments
FISH analysis performed in the probands and in their mother was compatible with a tandem intrachromosomal duplication since the RP11-980I14 BAC probe hybridized to 6q chromosomes only. Of note, the FISH signals did not show any intensity variation on the 6q homologous chromosome in the probands while a clear intensity difference was observed in the heterozygous mother (Supplementary Figure 3).
SNP analysis
As expected, SNP analysis detected several regions of copy-neutral loss of heterozygosity in the probands (Supplementary Table 1), due to cosegregation of chromosomal regions identical by descent inherited from consanguineous parents (Supplementary Table 1). Interestingly, the probands shared only a 37-Mb region of homozygosity on chromosome 6q12–q14.16, containing the triplicated region and more than 170 Refseq genes (Supplementary Table 2).
Molecular characterization of the rearrangement
MLPA assay containing several probes across the breakpoint regions as defined by SNP/CGH array allowed us to narrow down these intervals to less than 4kb and to design PCR primer combinations under the assumption of tandem duplication. A unique long-range PCR amplicon of about 4kb was generated in both parents and affected brothers but not in a group of ten normal controls (i.e. specific of the rearrangement). Sanger sequencing of the junction fragment did not reveal any homology between the proximal and distal breakpoint-flanking sequences using BLAST2 tool from NCBI (https://www.ncbi.nlm.nih.gov/), but showed at the breakpoint junction three extra nucleotides (GTA) not present in the consensus sequences (Fig. 4b). These features are compatible with a classical non-homologous end joining mechanism [24]. The breakpoints of the tandem duplicated region were mapped in intron 3 (NC_000006.11, bp69588797) and intron 17 (NC_000006.11, bp69902710), respectively. The rearrangement resulted in a direct, intragenic duplication involving exons 4–17 of the ADGRB3 gene (g.69902710_69902711ins[GTA; 69588797_69902710]) (Fig. 4b).
Since ADGRB3 is almost exclusively expressed in the brain, we were not able to characterize the abnormal transcript, which however is predicted to be out-of-frame, leading to a premature stop codon and possibly to nonsense-mediated decay degradation of the aberrant mRNA.
Whole-exome sequencing analysis
WES analysis was performed on genomic DNA of the affected siblings and their parents. The minimum average coverage of the Refseq coding regions among experiments was 129 x , with 93% of the bases covered at least 20 xand 98% of the bases covered at least 10 × . Each individual had on an average 2,7817 variants in the coding regions and in the intron/exon junctions, including 2173 rare variants with minor allele frequencies < 1% (806 synonymous, 1309 missense, 40 nonsense, 18 frameshift, and 21 splicing variants). In order to investigate a possible autosomal recessive condition, rare homozygous and compound heterozygous variants shared by both siblings were filtered taking into account their predicted functional impact, allowing us to identify six genes with biallelic variants, including the OMIM disease gene PLEC (MIM: 601282), involved in several forms of epidermolysis bullosa and in autosomal recessive limb-girdle muscular dystrophy. However, neither this gene nor the remaining five non-OMIM disease genes showed a consistent relationship with the clinical features of the family. Finally, we did not identify any de novo variant in the probands that might have suggested a parental germinal mosaicism.
Discussion
Here we report two siblings from a consanguineous family with intellectual disability, cerebellar atrophy, and behavioral disorder, carrying a homozygous duplication inherited from their heterozygous parents who both had psychiatric symptoms. A further molecular characterization of the rearrangement demonstrated a disrupting intragenic duplication involving several exons of the ADGRB3 gene, coding for a member of the adhesion-GPCRs family, which includes proteins having a pivotal role in the development and maintenance of the central nervous system.
Not surprisingly, due to their functions, several ADRG genes have been linked to different human diseases [25] such as neural tube defects (OMIM 182940; CELSR1), bilateral frontoparietal polymicrogyria (OMIM 606854; ADGRG1) and bilateral perisylvian polymicrogyria (OMIM 615752; ADGRG1), lethal congenital contracture syndrome 9 (OMIM 616503; ADGRG6), and Usher syndrome type IIC (OMIM 605472; ADGRV1).
Recently, receptors of ADGRB subfamily of adhesion-GPCRs have emerged as central regulators of synapse development and plasticity. Different studies have shown that ADGRB proteins interact with the members of the C1q family of secreted complement-like proteins, which are highly expressed in the central nervous system [26, 27]. ADGRB3 is almost exclusively expressed in the brain, from where it localizes to excitatory synapses [7]. ADGRB3 has been shown to regulate excitatory synapse connectivity and formation during mouse cerebellum development and adulthood by binding with its ligand C1ql1, which is secreted by the excitatory climbing fibers (CFs) of inferior olivary neurons and acts as an anterograde signal to determine the postnatal pruning of supernumerary synapses and maintain a single-winner CF to Purkinje cells (PCs). A similar crucial role of ADGRB3 in PC synaptogenesis has also been demonstrated for parallel fibers arising from axons of granule cells throughout development and adulthood [7, 27].
Surprisingly, mice in which the ADGRB3 gene was specifically disrupted in PCs had normal cerebellum structure and no ataxia, showing normal motor coordination although they displayed fewer CF synapses and a highly impaired motor learning [28]. On the contrary, our patients present with cerebellar atrophy, ambulation with ataxic gait, and deficiency of motor and ocular coordination. It remains unclear why mice with a conditional loss of ADGRB3 in PCs did not show the full spectrum of cerebellar symptoms or whether the expression patterns and the exact functional roles diverge between human ADGRB3 and its mouse ortholog, leading to different phenotypic effects.
ADGRB3 is also highly expressed in hippocampal neurons, acting as a regulator of synapse density through its interaction with C1q3, another brain-specific secreted protein [7]. Indeed, ADGRB3 knockdown in hippocampal neurons highly impairs dendrite morphogenesis in mice [28]. The clinical spectrum of our patients also includes neuropsychiatric symptoms such as mood instability, anxiety, hallucinations, and social closure, which may be associated with hippocampus dysfunctions [29,30,31].
Interestingly, recent reports have linked the members of the ADGRB subfamily to neuropsychiatric diseases; ADGRB1 germline variants have been identified in autistic patients [32] while ADGRB3 SNPs and CNVs have been associated, in literatures, with schizophrenia, bipolar disorder, and drug addiction [10,11,12,13]. Liao et al. [11] reported on a family segregating a 4.4-Mb deletion at chromosome 6q12–13 involving ADGRB3, associated in almost all carrier individuals with psychiatric symptoms including schizophrenia and schizoaffective disorder. Although other genes were affected by the deletion, the authors suggested that ADGRB3 haploinsufficiency might be related to the psychiatric disease in that family. Interestingly, in our pedigree, heterozygous parents share psychiatric impairments such as psychotic disorder and anxious/depressive disorder. Taken together, these findings support the hypothesis that ADGRB3 haploinsufficiency may lead to psychiatric disorders in heterozygous individuals.
Although the functional role of ADGRB3 may explain the observed clinical picture, we are aware that extreme caution should be taken before claiming an unknown CNV as causative factor. As pointed out by different authors [33, 34], this is especially true in case of consanguineous families where other unmasked homozygous variants may be relevant for the disease. Indeed, some of the main clinical features of our patients, such as intellectual disability, cerebellar atrophy with vermis involvement, ataxia, and strabismus are also present in cerebellar ataxia, mental retardation, and dysequilibrium syndrome, a genetically heterogeneous disorder associated with recessive alleles in four genes (VLDLR, MIM: 192977; WDR81, MIM:614218; CA8 MIM:114815; ATP8A2, MIM:605870). In order to investigate the hypothesis that recessive variants may be, at least partly, responsible for the trait, we carried out WES assay in all the families. Although the analysis did not yield any putative causative variant, it remains possible that other genetic factors not covered by the WES assay may concur with the disorder.
Finally, it is important to point out that this study has some limitations that prevent drawing a firm conclusion concerning the role of ADGRB3 gene in the disease. Indeed, in spite of the potentially inactivating alteration of this gene, we were not able to analyze its impact at mRNA level. Moreover, no other patients with biallelic alterations of the ADRGB3 gene have been reported yet. Clearly, more studies are still needed to assess the role of ADGRB3 gene in brain diseases.
References
Girirajan S, Rosenfeld JA, Coe BP, Parikh S, Friedman N, Goldstein A, et al. Phenotypic heterogeneity of genomic disorders and rare copy-number variants. N Engl J Med. 2012;367:1321–31.
Lupiáñez DG, Kraft K, Heinrich V, Krawitz P, Brancati F, Klopocki E, et al. Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions. Cell. 2015;161:1012–5.
Boone PM, Campbell IM, Baggett BC, Soens ZT, Rao MM, Hixson PM, et al. Deletions of recessive disease genes: CNV contribution to carrier states and disease-causing alleles. Genome Res. 2013;23:1383–94.
Houcinat N, Llanas B, Moutton S, Toutain J, Cailley D, Arveiler B, et al. Homozygous 16p13.11 duplication associated with mild intellectual disability and urinary tract malformations in two siblings born from consanguineous parents. Am J Med Genet A. 2015;167A:2714–9.
Okamoto Y, Goksungur MT, Pehlivan D, Beck CR, Gonzaga-Jauregui C, Muzny DM, et al. Exonic duplication CNV of NDRG1 associated with autosomal-recessive HMSN-Lom/CMT4D. Genet Med. 2014;16:386–94.
Hamann J, Aust G, Arac D, Engel FB, Formstone C, Fredriksson R, et al. International Union of Basic and Clinical Pharmacology. XCIV. Adhesion G protein-coupled receptors. Pharmacol Rev. 2015;67:338–67.
Sigoillot S, Iyer K, Binda F, González-Calvo I, Talleur M, Vodjdani G, et al. The secreted protein C1QL1 and its receptor BAI3 control the synaptic connectivity of excitatory inputs converging on cerebellar purkinje cells. Cell Rep. 2015;10:820–32.
Kakegawa W, Mitakidis N, Miura E, Abe M, Matsuda K, Takeo YH, et al. Anterograde C1ql1 signaling is required in order to determine and maintain a single-winner climbing fiber in the mouse cerebellum. Neuron. 2015;85:316–30.
De Rosse P, Lencz T, Burdick KE, Siris SG, Kane JM, Malhotra AK. “The genetics of symptom-based phenotypes: toward a molecular classification of schizophrenia. Schizophr Bull. 2008;34:1047–53.
Liao HM, Chao YL, Huang AL, Cheng MC, Chen YJ, Lee KF, et al. Identification and characterization of three inherited genomic copy number variations associated with familial schizophrenia. Schizophr Res. 2012;139:229–36.
McCarthy MJ, Nievergelt CM, Kelsoe JR, Welsh DK. A survey of genomic studies supports association of circadian clock genes with bipolar disorder spectrum illnesses and lithium response. PLoS ONE. 2012;7:e32091.
Liu QR, Drgon T, Johnson C, Walther D, Hess J, Uhl GR. Addiction molecular genetics: 639,401 SNP whole genome association identifies many ‘cell adhesion’ genes. Am J Med Genet, Part B: Neuropsychiatr Genet. 2006;141:918–25.
Wechsler, D Wechsler. Adult Intelligence Scale. 4th ed. San Antonio, TX: Pearson; 2008.
Sparrow SS, Cicchetti DV, Balla DA. Vineland adaptive behavior scales. 2nd ed. Circle Pines, MN: American Guidance Service; 2005.
Cornoldi C, Giofrè D, Belacchi C. Procedure di somministrazione e scoring dei subtest. In: di Roid GH, Miller LJ, Pomplun M, Koch C, editors. Leiter-3 Leiter International Scale. Firenze: Giunti O.S. Organizzazioni Speciali s.r.l. Adattamento italiano; 2016. p. 23–85.
Beck AT, Steer RA. Manual for the Beck Anxiety Inventory. San Antonio,TX: The Psychological Corporation; 1990.
Beck AT, Ward CH, Mendelson M, Mock J, Erbaugh J. An inventory for measuring depression. Arch Gen Psychiatry. 1961;4:561–71.
Baldwin EL, Lee JY, Blake DM, Alexander CR, Kogan AL, Ledbetter DH, et al. Enhanced detection of clinically relevant genomic imbalances using a targeted plus whole genome oligonucleotide microarray. Genet Med. 2008;10:415–29.
Lo-Castro A, Giana G, Fichera M, Castiglia L, Grillo L, Musumeci SA, et al. Deletion 2p25.2: A cryptic chromosome abnormality in a patient with autism and mental retardation detected using aCGH. Eur J Med Genet. 2009;52:67–70.
Li H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv . 2013;1303:3997.
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data 2010. Genome Res. 2010;20:1297–303.
Van der Auwera GA, Carneiro M, Hartl C, Poplin R, Del Angel G, Levy-Moonshine A, et al. From FastQ data to high-confidence variant calls: the Genome Analysis Toolkit best practices pipeline 2013. Curr Protoc Bioinformatics. 2013;43:11.10.1–11.10.33
Yang H, Wang K. Genomic variant annotation and prioritization with ANNOVAR and wANNOVAR. Nat Protoc. 2015;10:1556–66.
Vissers LE, Bhatt SS, Janssen IM, Xia Z, Lalani SR, Pfundt R, et al. Rare pathogenic microdeletions and tandem duplications are microhomology-mediated and stimulated by local genomic architecture. Hum Mol Genet. 2009;18:3579–93.
Langenhan T, Piao X, Monk KR. Adhesion G protein-coupled receptors in nervous system development and disease. Nat Rev Neurosci. 2016;17:550–61.
Cbln YuzakiM. and C1q family proteins: new transneuronal cytokines. Cell Mol Life Sci. 2008;65:1698–705.
Bolliger MarcF, Martinelli DavidC, Thomas, Südhof C. The cell-adhesion G protein-coupled receptor BAI3 is a high-affinity receptor for C1q-like proteins. Proc Natl Acad Sci USA. 2011;108:2534–9.
Kakegawa W, Mitakidis N, Miura E, Abe M, Matsuda K, Takeo YH, et al. Anterograde C1ql1 signaling is required in order to determine and maintain a single-winner climbing fiber in the mouse cerebellum. Neuron. 2015;85:316–29.
Lanoue V, Usardi A, Sigoillot SM, Talleur M, Iyer K, Mariani J, et al. The adhesion-GPCR BAI3, a gene linked to psychiatric disorders, regulates dendrite morphogenesis in neurons. Mol Psychiatry. 2013;18:943–50.
Rubin RD, Watson PD, Duff MC, Cohen NJ. The role of the hippocampus in flexible cognition and social behavior. Front Hum Neurosci. 2014;8:742.
Montagrin A, Saiote C, Schiller D. The social hippocampus. Hippocampus. 2017. https://doi.org/10.1002/hipo.22797.
Michaelson JJ, Shi Y, Gujral M, Zheng H, Malhotra D, Jin X, et al. Whole-genome sequencing in autism identifies hot spots for de novo germline mutation. Cell. 2012;151:1431–42.
Vissers LE, de Ligt J, Gilissen C, Janssen I, Steehouwer M, de Vries P, et al. A de novo paradigm for mental retardation. Nat Genet. 2010;42:1109–12.
Giorgio E, Ciolfi A, Biamino E, Caputo V, Di Gregorio E, Belligni EF, et al. Whole exome sequencing is necessary to clarify ID/DD cases with de novo copy number variants of uncertain significance: Two proof-of-concept examples. Am J Med Genet A. 2016;170:1772–9.
Acknowledgements
We thank the probands and their family for their participation in this study. For technical support, we acknowledge the participation of Drs. Silvestra Amata, Angela Spalletta, Maurizio Sturnio, Angelo Gloria, Franco Scillato, and Pietro Schinocca from the Laboratory of Medical Genetics, IRCCS Associazione Oasi M SS., Troina (Italy). We also thank Dr. Anthony Charles for English language editing. MF was funded by grant RF-2011-02350693 from the Italian Ministry of Health.
Author contributions
Corresponding author MF had full access to all of the data in the study and took responsibility for the integrity of the data, accuracy of data analysis, and final responsibility for the decision to submit for publication. CS, CR, and MF were responsible for study supervision. MF and CS drafted the manuscript and interpreted the data. LS and MS performed WES analysis. CS, LS and MV equally contributed to the manuscript. MV performed MLPA assay, long-range PCR and breakpoint analysis. LC carried out SNP/CGH array analysis. OG performed FISH analysis. CS was in charge of the patients and contributed relevant clinical data for phenotypic characterization. SC performed psychological and cognitive assessment of the patients. TM, CR, EB, and LC critically revised the manuscript. All authors contributed to the study concept and design. All authors were responsible for drafting the manuscript, contributed to the acquisition, analysis, and interpretation of data, and read and approved the final manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethics approval and consent to participate
The research design, in accordance with the tenets of the Declaration of Helsinki and their reviews, was approved by the Ethics Committee. The patients included in the study signed a written informed consent to participate in the study.
Consent for publication
The participants included in the study signed a written informed consent to publish their data.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Scuderi, C., Saccuzzo, L., Vinci, M. et al. Biallelic intragenic duplication in ADGRB3 (BAI3) gene associated with intellectual disability, cerebellar atrophy, and behavioral disorder. Eur J Hum Genet 27, 594–602 (2019). https://doi.org/10.1038/s41431-018-0321-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41431-018-0321-1