Abstract
Genome-wide association studies (GWAS) of quantitative electrocardiographic (ECG) traits in large consortia have identified more than 130 loci associated with QT interval, QRS duration, PR interval, and heart rate (RR interval). In the current study, we meta-analyzed genome-wide association results from 30,000 mostly Dutch samples on four ECG traits: PR interval, QRS duration, QT interval, and RR interval. SNP genotype data was imputed using the Genome of the Netherlands reference panel encompassing 19 million SNPs, including millions of rare SNPs (minor allele frequency < 5%). In addition to many known loci, we identified seven novel locus-trait associations: KCND3, NR3C1, and PLN for PR interval, KCNE1, SGIP1, and NFKB1 for QT interval, and ATP2A2 for QRS duration, of which six were successfully replicated. At these seven loci, we performed conditional analyses and annotated significant SNPs (in exons and regulatory regions), demonstrating involvement of cardiac-related pathways and regulation of nearby genes.
Similar content being viewed by others
Introduction
Quantitative electrocardiographic (ECG) traits have been well studied in large consortia, identifying over 130 significant loci. Some loci were associated with multiple traits. Nevertheless, these loci collectively explain only a small portion of the genetic variation of these traits [1]. Large GWAS meta-analyses on PR interval [2, 3], RR interval/heart rate [4, 5], QRS duration [6, 7], and QT interval [8,9,10] were based on HapMap imputations [11]. Testing ~2.5 million SNPs, these studies provided good coverage of common variation in the genome. SNPs with lower allele frequencies (e.g., minor allele frequencies between 1 and 5%), however, are poorly covered [12, 13]. While HapMap included only 270 samples (30 trios and 90 unrelated samples) from three continental populations [11], the 1000 Genomes Project Phase 3 contains 2504 samples from 26 populations [14]. Larger reference panels cover a broader variety of haplotypes and, therefore, increase the quality of imputation in a GWAS sample. Moreover, the number of observed SNPs also increases, expanding the number available for imputation. This has led to novel findings in non-ECG related studies [15].
In the current study, we meta-analyzed genome-wide data on four ECG traits in 30,000 predominantly Dutch samples. We tested over 19 million SNPs for association, which were imputed using the Genome of the Netherlands (GoNL) reference panel [16]. This dataset contains whole-genome sequencing data at 12x coverage collected in 250 families (trios and parents with two offspring). Nearly all polymorphic sites with a population frequency of more than 0.5% are captured. This makes it one of the largest single population sequencing efforts worldwide and the trio design ensures very accurate haplotype phasing. These features and the good match with the predominantly Dutch cohorts, make this dataset well suited as a reference panel for imputation. Using this approach, we had two aims: (1) the discovery of novel loci associated with ECG traits, and (2) the fine-mapping and functional annotation of known regions associated with ECG traits. We increased our SNP density almost seven-fold compared to previous studies based on HapMap, enabling us to study key signals in much finer detail.
Methods
Individual cohort data
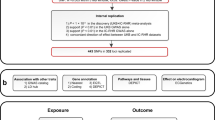
Eight cohorts were included in the discovery phase of this study, totaling approximately 30,000 samples (Supplementary Tables 1 and 2, Supplementary Notes). Most study participants were Dutch with the exception of most participants of PROSPER; this study included approximately 19% samples of Dutch origin, while the remaining samples were of other European descent. All cohorts performed stringent quality control to exclude low-quality samples and SNPs prior to imputation and also post-imputation. Imputation was performed using 998 phased haplotypes from the Genome of the Netherlands Project release 4 as the reference panel, encompassing 19,763,454 SNPs [16]. All genomic data in this manuscript is listed with respect to the hg19 (build37) reference genome.
We evaluated four phenotypes on the electrocardiogram: RR interval, PR interval, QRS duration, and QT interval. Seven out of eight cohorts contributed data to all four phenotypes; NTR only had data on RR interval available. Samples of non-European descent and samples with missing data were excluded, as well as individuals that fulfilled any of the exclusion criteria listed in Supplementary Table 3. SNPs were individually tested for association with each trait using linear models. For all four phenotypes, we included age, sex, height, BMI, and study specific covariates (for instance to correct for study site, relatedness, or population stratification) as covariates. In addition, RR interval and hypertension (in those cohorts that had data available on this measure) were included as covariates for QT interval to reduce noise introduced by these factors. We chose these covariates to correspond with previously published GWAS on these four ECG traits.
Quality control and meta-analysis
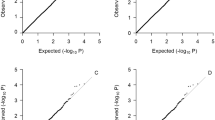
Association results from all cohorts were collected at a single site and underwent quality control. SNPs with extreme values of beta (>1000 or <−1000), standard error (SE) (>1000), or imputation quality (<0.1 or >1.1) were removed and distributions of beta, SE, and P-values were manually checked. We made QQ-plots to test P-value distributions, which were stratified by minor allele frequency and by imputation quality. Aberrant subsets of SNPs (usually with very low frequency) were removed from downstream analyses.
Inverse-variance fixed-effect model meta-analyses were conducted for all four traits using MANTEL [17]. For each individual GWAS, genomic inflation factors (lambda) were calculated and, during meta-analysis, standard errors were adjusted accordingly to correct for population structure and technical errors. We did not correct for genomic inflation after meta-analysis. SNP associations were considered significant if P ≤ 5 × 10–8.
Follow-up on known and novel loci
For each locus, we tested the number of independent signals using the LD structure from GoNL in GCTA-COJO, which was designed to allow conditional analyses based on summary-level data [18]. Secondary hits had to fulfill two criteria: (1) genome-wide significant in the GWAS, and (2) P < 1 × 10–5 after conditioning to correct for multiple testing of 4757 significant SNPs across all four traits. A novel locus for a trait was defined if the significant SNPs, or SNPs within a distance of 1 Mb upstream and downstream of the significant SNPs, had not been observed before in GWAS of the same trait. We performed a look-up of all novel loci in previous HapMap-based GWAS.
Replication of novel loci in CHARGE
We sought to replicate our findings in 13 independent cohorts taking part in the CHARGE consortium [19] (Supplementary Tables 1 and 2, Supplementary notes). Twelve studies (TwinsUK, CHS, ARIC, KORA F3, KORA S4, JHS, AGES, BRIGHT, YFS, INGI-FVG, and INGI-CARL) used 1000 Genomes Phase 1 as their imputation reference panel and a single study (Inter99) provided only genotyped data. All studies contained samples of European ancestry, except for JHS, which consisted only of African-American samples. The summary-level results for all novel SNPs determined in the discovery analysis were combined in inverse-variance fixed-effects meta-analyses. A two-sided P-value ≤ 0.05, in conjunction with a concordant effect direction, was considered significant.
In silico tests of possibly functional SNPs
We looked up the functional annotations for all SNPs that reached genome-wide significance in any of the four traits. First, we checked whether SNPs were potentially damaging to protein function, testing all non-synonymous SNPs in SIFT [20] and PolyPhen-2 [21]. Second, we used GREAT [22] to identify biological pathways in which regulatory SNPs are involved, testing the index SNPs for all locus-trait associations. Lastly, we tested all significant SNPs one by one for their possible effect on regulatory regions using RegulomeDB [23].
Results
Meta-analysis detects novel loci
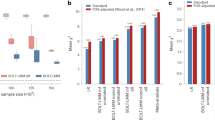
We conducted a GWAS meta-analysis comprising eight cohorts that together encompassed approximately 30,000 samples. Over 19 million SNPs, imputed using the GoNL reference panel, were assessed for association with four quantitative ECG traits: RR, PR, QRS, and QT. Considering all traits, we observed 52 locus-phenotype associations (17 for PR, 13 for QRS, 15 for QT, and 7 for RR; Supplementary Figures 1 and 2, Supplementary Table 4). A locus was defined as an associated region (containing one or more SNPs with P ≤ 5 × 10–8) that is located at least 1 Mb away from the next (i.e., if two associated SNPs are within 1 Mb, they belong to the same locus). Of these 52 loci, 45 have been observed before in large GWAS meta-analyses [2,3,4, 7,8,9] and seven are novel findings (Table 1). Box 1 shows regional association plots and provides additional information on the seven novel loci. Imputation qualities of the index SNPs were 0.60 and 0.84 for the relatively rare KCNE1 and KCDN1 variants, respectively, and >0.96 for the remaining common index SNPs. The variance explained by each of these variants ranges between 0.09 and 0.23%.
Novel loci associated with PR, QRS, and QT. KCND3, associated with PR interval (a, b). ARHGAP26 and NR3C1, associated with PR interval (c). SLC35F1 and PLN, associated with PR interval (d). ATP2A2, associated with QRS duration (e). SGIP1 and TCTEX1D1, associated with QT interval (f). NFKB1, associated with QT interval (g). KCNE1, associated with QT interval (h)
Fine-mapping of known loci
For each locus, we tested if more than one independent signal was present (Supplementary Table 4). Thirteen loci had suggestive evidence of having more than one independent signal; four locus-phenotype associations had five or more independent signals. The SCN5A/SCN10A locus was the most outstanding locus with eleven independent signals for PR, and six for QRS. NOS1AP for QT contained seven independent signals.
Replication in CHARGE
For six out of seven novel loci, we were able to conduct look-ups of the index SNP or a proxy SNP in strong LD (r2 ≥ 0.89) in previous large-scale HapMap-based GWAS. These GWAS contained over 70,000 samples each, and included many of the Dutch cohorts from our current study. All six loci were associated with their respective traits (P ≤ 0.004). Next, we tested the seven novel loci for replication in 13 studies from the CHARGE consortium. In contrast to the HapMap look-ups, this replication was independent from the Dutch discovery sample. Results are shown in Table 1. Allele frequencies were very similar to the discovery dataset, except for JHS, which consists of individuals of African-American descent. Effect directions for all seven SNPs were concordant between our primary findings and replication, with effect sizes between 0.2 and 1.5 times those of the betas in the discovery study. Six of seven loci were replicated with P < 0.05, three of which pass Bonferroni correction, accounting for seven tests.
Functional SNPs in genes and regulatory regions
All genome-wide significant SNPs were tested in silico for their potential effect on gene expression and protein structure. Ten loci contained, in total, 15 non-synonymous SNPs, which were tested using the prediction programs PolyPhen-2 and SIFT. According to PolyPhen-2, three SNPs were possibly damaging (rs1805128 in KCNE1 for QT, rs12666989 in UFSP1 for RR, and rs2070492 in SLC22A14 for PR). SIFT predicted only one SNP to be damaging to a protein (rs3746471 in KIAA1755 for RR).
We used GREAT to test all 100 index SNPs from the four ECG traits combined for their biological function in cis-regulatory regions. Significant GO-terms (molecular function, biological process, and cellular component), human phenotypes, and disease ontologies are shown in Supplementary Table 5a–d. In total, these index SNPs mapped to 103 genes.
Of 52 locus-phenotype associations, 34 contained significant SNPs that have a RegulomeDB score of 3 or better, meaning that they may affect protein binding (Supplementary Table 6). We observed 15 loci containing SNPs with scores of 1 (likely to affect binding and linked to the expression of a gene target), 15 loci containing SNPs with a maximum score of 2 (likely to affect binding), and four loci that have SNPs with a maximum score of 3 (less likely to affect binding). Eighteen loci contained only SNPs with scores from 4 to 6 (minimal binding evidence) and 7 (no data available).
Discussion
We imputed over 19 million SNPs using GoNL as the reference panel, and tested these SNPs for association with four traits in eight predominantly Dutch cohorts comprising roughly 30,000 samples. We observed 52 locus-phenotype associations, seven of which were novel (Table 1, Box 1, Supplementary Table 4).
Discovery of loci associated with quantitative ECG traits
We detected seven novel loci, three for PR interval, three for QT interval, and one for QRS duration (Box 1). No novel loci were found for RR interval, accounting for loci previously associated with either RR interval [4] or heart rate [5]. We replicated six out of seven novel loci utilizing 13 independent studies from the CHARGE consortium. Interestingly, the only variant that does not replicate is rs74640693 for PR interval, located close to PLN (phospholamban). Variants in this gene have been consistently associated with various QRS measures [6] but not with PR interval. The gene transcribes the phospholamban protein, which is important in calcium signaling in cardiac muscle cells [24]. Although a Dutch-specific pathogenic mutation, p.Arg14del, in the PLN gene has been described [25], it is unlikely that this mutation drives the association signal in our study because the allele frequency of SNP rs74640693 is similar in our samples (4.9%) compared to other samples of European ancestry (4.6% in the 12 European CHARGE replication cohorts). Furthermore, the allele frequency of this SNP is ~5 times higher than that of the mutation and the SNP is located ~200 kb upstream of the PLN gene, so, therefore, not in LD with these mutations. In addition, a recent large study of PR interval used the Illumina exome chip to identify a common variant (rs74640693, allele frequency 47%) in this region [26], however, this variant is not in LD with the variant that we identified (r2 = 0.04). To confirm that the lack of association was not caused by strand issues (because rs74640693 is an A/T variant), we tested the nearby proxy SNP rs12210733 (which is an A/G variant, r2 = 0.89) in the CHARGE replication cohorts, and found it was also non-significant.
We looked up our top SNPs in previous, much larger, HapMap-based GWAS meta-analyses to determine why our SNPs were not identified in those studies (Table 1). Two loci contained rare SNPs with MAF < 5%. Low-frequency SNPs at KCND3 were not present in HapMap and could therefore not be tested. The functional SNP at KCNE1 was observed in a single cohort in a meta-analysis in 2009, but this result could neither be replicated in other cohorts [9], nor in later studies, because the imputation quality was too low.
For common SNPs (MAF > 5%), it is much more difficult to define why they were not previously observed at genome-wide significance. For many loci we may have better tags of the causal variants because our coverage is almost sevenfold greater. Indeed, the index SNPs at PLN (PR), NFKB1 (QT), and ATP2A2 (QRS) were not tested in previous studies. Nevertheless, for all SNPs, proxies with r2 > 0.9 were available in the respective studies (Table 1). Common SNPs at KCND3 (PR), NR3C1 (PR), and SGIP1 (QT) were present in HapMap. Both proxies and directly imputed SNPs were at least nominally significant in previous studies (P-values ranging from 10–3 to 10–6) with typically high imputation quality.
In addition to the “winner’s curse” effect, we expect that higher quality imputation due to the considerably larger haplotype panel (compared to HapMap) and the ancestry matching between GoNL and our Dutch cohorts will improve the power to detect a true association signal, if present. Although combining multiple reference panels for imputation is becoming the new standard [27], limitations to our study remain: (1) the GoNL reference panel may not contain sufficient information on rare SNPs; (2) the small sample size of individual cohorts may cause abnormal behavior of rare SNPs as a group, requiring us to remove that subset of SNPs; or (3) the sample size or power of the overall study is still limited to detect rare variant associations.
Fine-mapping of known loci
Although we did not sequence the loci containing the known and novel signals, we have a much denser interrogation of these regions compared to previous (HapMap-based) studies. In an attempt to fine map the significant loci, we annotated all significant SNPs with their predicted functional consequences.
First, we used SIFT and PolyPhen-2 to predict the effect of 15 non-synonymous SNPs that were associated with one of the ECG traits at genome-wide significance. PolyPhen-2 classified three SNPs as possibly damaging and SIFT predicted only one SNP to be damaging. These were non-overlapping, raising questions as to the actual effect of these SNPs on their respective genes. Functional studies should be pursued to test the direct effect of these SNPs on protein structure.
Combining all index SNPs, we tested the function of those SNPs located in cis-regulatory regions using GREAT [22]. We identified 100 independent SNP-trait associations, which mapped to 103 genes. Interestingly, we find hundreds of significant nodes, of which the vast majority is related to cardiac functioning and heart disease (Supplementary Table 5a–e). This shows that, indeed, many SNPs are located in cis-regulatory regions of genes that are critical in the functioning of the human heart, which implicates a regulatory function of these loci rather than a structural function changing the protein directly. One example is shown in Supplementary Figure 3; this figure contains all significant GO molecular function nodes. Most of these nodes are in the group of transporter activity, which includes all transmembrane channels that are known to be important for cardiac function.
Because the GREAT pathways show that many SNPs probably have their effect on the trait due to gene regulation, we extracted all significant SNPs from RegulomeDB to check which variants would likely affect binding in regulatory regions. A majority of loci contained at least one SNP that was expected to affect transcription factor binding (Supplementary Table 6). The score that is provided by RegulomeDB indicates that a SNP is likely (or less likely) located in a binding site. Interestingly, there are strong differences between loci in terms of the number of SNPs that may have a regulatory effect, and percentage of loci per trait that have a high score. For instance, seven out of 15 QT interval loci contains SNPs with a score of 1, while only a single PR interval locus contains a SNP with this score. The SCN5A/SCN10A locus is strongly associated with PR interval (best SNP P = 4.9 × 10–107) and contains over 450 significant SNPs. Nevertheless, only six SNPs have a score of 2 or 3, and none of the significant SNPs have a score of 1. However, many binding sites are tissue specific, and, therefore, testing SNPs with high scores systematically for their role in cardiac tissue could lead to more knowledge about their biological function.
Conclusions
Using the Genome of the Netherlands as imputation reference panel, we identified seven novel loci for quantitative ECG traits. Higher SNP density and higher imputation quality enabled us to annotate known loci, facilitating future studies to understand the biological effects of causal variants at many loci. Ultimately, combining imputation reference panels and increasing sample size for GWAS meta-analyses will continue to increase power for genetic discovery and novel target identification. With many sequencing efforts ongoing and large population-based cohorts being genotyped (such as UK Biobank, of which the first release data showed 46 novel loci for RR interval [28]), we can expect hundreds of novel loci for ECG phenotypes in the near future.
References
Silva CT, Kors JA, Amin N, Dehghan A, Witteman JC, Willemsen R, et al. Heritabilities, proportions of heritabilities explained by GWAS findings, and implications of cross-phenotype effects on PR interval. Hum Genet. 2015;134:1211–9.
Pfeufer A, van Noord C, Marciante KD, Arking DE, Larson MG, Smith AV, et al. Genome-wide association study of PR interval. Nat Genet. 2010;42:153–9.
van Setten J, Brody JA, Jamshidi Y, Swenson BR, Butler AM, Campbell H, et al. PR interval genome-wide association metaanalysis identifies 50 loci associated with atrial and atrioventricular electrical activity. Nat Commun. 2018;9:2904.
Eijgelsheim M, Newton-Cheh C, Sotoodehnia N, de Bakker PI, Muller M, Morrison AC, et al. Genome-wide association analysis identifies multiple loci related to resting heart rate. Hum Mol Genet. 2010;19:3885–94.
den Hoed M, Eijgelsheim M, Esko T, Brundel BJ, Peal DS, Evans DM, et al. Identification of heart rate-associated loci and their effects on cardiac conduction and rhythm disorders. Nat Genet. 2013;45:621–31.
van der Harst P, van Setten J, Verweij N, Vogler G, Franke L, Maurano MT, et al. 52 Genetic Loci Influencing Myocardial Mass. J Am Coll Cardiol. 2016;68:1435–48.
Sotoodehnia N, Isaacs A, de Bakker PI, Dorr M, Newton-Cheh C, Nolte IM, et al. Common variants in 22 loci are associated with QRS duration and cardiac ventricular conduction. Nat Genet. 2010;42:1068–76.
Newton-Cheh C, Eijgelsheim M, Rice KM, de Bakker PI, Yin X, Estrada K, et al. Common variants at ten loci influence QT interval duration in the QTGEN Study. Nat Genet. 2009;41:399–406.
Pfeufer A, Sanna S, Arking DE, Muller M, Gateva V, Fuchsberger C, et al. Common variants at ten loci modulate the QT interval duration in the QTSCD Study. Nat Genet. 2009;41:407–14.
Arking DE, Pulit SL, Crotti L, van der Harst P, Munroe PB, Koopmann TT, et al. Genetic association study of QT interval highlights role for calcium signaling pathways in myocardial repolarization. Nat Genet. 2014;46:826–36.
Frazer KA, Ballinger DG, Cox DR, Hinds DA, Stuve LL, Gibbs RA, et al. A second generation human haplotype map of over 3.1 million SNPs. Nature. 2007;449:851–61.
Barrett JC, Cardon LR. Evaluating coverage of genome-wide association studies. Nat Genet. 2006;38:659–62.
Pe'er I, de Bakker PI, Maller J, Yelensky R, Altshuler D, Daly MJ. Evaluating and improving power in whole-genome association studies using fixed marker sets. Nat Genet. 2006;38:663–7.
Genomes Project C, Auton A, Brooks LD, Durbin RM, Garrison EP, Kang HM, et al. A global reference for human genetic variation. Nature. 2015;526:68–74.
de Vries PS, Sabater-Lleal M, Chasman DI, Trompet S, Ahluwalia TS, Teumer A, et al. Comparison of HapMap and 1000 Genomes Reference Panels in a Large-Scale Genome-Wide Association Study. PLoS ONE. 2017;12:e0167742.
Genome of the Netherlands C. Whole-genome sequence variation, population structure and demographic history of the Dutch population. Nat Genet. 2014;46:818–25.
de Bakker PI, Ferreira MA, Jia X, Neale BM, Raychaudhuri S, Voight BF. Practical aspects of imputation-driven meta-analysis of genome-wide association studies. Hum Mol Genet. 2008;17:R122–128.
Yang J, Ferreira T, Morris AP, Medland SE, Genetic Investigation of ATC, Replication DIG et al. Conditional and joint multiple-SNP analysis of GWAS summary statistics identifies additional variants influencing complex traits. Nat Genet. 2012;44:369–75. S361–363.
Psaty BM, O'Donnell CJ, Gudnason V, Lunetta KL, Folsom AR, Rotter JI, et al. Cohorts for Heart and Aging Research in Genomic Epidemiology (CHARGE) Consortium: Design of prospective meta-analyses of genome-wide association studies from 5 cohorts. Circ Cardiovasc Genet. 2009;2:73–80.
Kumar P, Henikoff S, Ng PC. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat Protoc. 2009;4:1073–81.
Adzhubei IA, Schmidt S, Peshkin L, Ramensky VE, Gerasimova A, Bork P, et al. A method and server for predicting damaging missense mutations. Nat Methods. 2010;7:248–9.
McLean CY, Bristor D, Hiller M, Clarke SL, Schaar BT, Lowe CB, et al. GREAT improves functional interpretation of cisregulatory regions. Nat Biotechnol. 2010;28:495–501.
Boyle AP, Hong EL, Hariharan M, Cheng Y, Schaub MA, Kasowski M, et al. Annotation of functional variation in personal genomes using RegulomeDB. Genome Res. 2012;22:1790–7.
Luo W, Grupp IL, Harrer J, Ponniah S, Grupp G, Duffy JJ, et al. Targeted ablation of the phospholamban gene is associated with markedly enhanced myocardial contractility and loss of beta-agonist stimulation. Circ Res. 1994;75:401–9.
van der Zwaag PA, van Rijsingen IA, de Ruiter R, Nannenberg EA, Groeneweg JA, Post JG, et al. Recurrent and founder mutations in the Netherlands-Phospholamban p.Arg14del mutation causes arrhythmogenic cardiomyopathy. Neth Heart J. 2013;21:286–93.
Lin H, van Setten J, Smith AV, Bihlmeyer NA, Warren HR, Brody JA, et al. Common and Rare Coding Genetic Variation Underlying the Electrocardiographic PR Interval. Circ Genom Precis Med. 2018;11:e002037.
McCarthy S, Das S, Kretzschmar W, Delaneau O, Wood AR, Teumer A, et al. A reference panel of 64,976 haplotypes for genotype imputation. Nat Genet. 2016;48:1279–83.
Eppinga RN, Hagemeijer Y, Burgess S, Hinds DA, Stefansson K, Gudbjartsson DF, et al. Identification of genomic loci associated with resting heart rate and shared genetic predictors with all-cause mortality. Nat Genet. 2016;48:1557–63.
Verweij N, Mateo Leach I, Isaacs A, Arking DE, Bis JC, Pers TH, et al. Twenty-eight genetic loci associated with ST-T-wave amplitudes of the electrocardiogram. Hum Mol Genet. 2016;25:2093–103.
Low SK, Takahashi A, Ebana Y, Ozaki K, Christophersen IE, Ellinor PT, et al. Identification of six new genetic loci associated with atrial fibrillation in the Japanese population. Nat Genet. 2017;49:953–8.
Lundby A, Olesen SP. KCNE3 is an inhibitory subunit of the Kv4.3 potassium channel. Biochem Biophys Res Commun. 2006;346:958–67.
Delpon E, Cordeiro JM, Nunez L, Thomsen PE, Guerchicoff A, Pollevick GD, et al. Functional effects of KCNE3 mutation and its role in the development of Brugada syndrome. Circ Arrhythm Electrophysiol. 2008;1:209–18.
Lee YC, Durr A, Majczenko K, Huang YH, Liu YC, Lien CC, et al. Mutations in KCND3 cause spinocerebellar ataxia type 22. Ann Neurol. 2012;72:859–69.
Kadmiel M, Cidlowski JA. Glucocorticoid receptor signaling in health and disease. Trends Pharmacol Sci. 2013;34:518–30.
Rog-Zielinska EA, Thomson A, Kenyon CJ, Brownstein DG, Moran CM, Szumska D, et al. Glucocorticoid receptor is required for foetal heart maturation. Hum Mol Genet. 2013;22:3269–82.
Oakley RH, Cidlowski JA. The biology of the glucocorticoid receptor: new signaling mechanisms in health and disease. J Allergy Clin Immunol. 2013;132:1033–44.
Lundmark R, Doherty GJ, Howes MT, Cortese K, Vallis Y, Parton RG, et al. The GTPase-activating protein GRAF1 regulates the CLIC/GEEC endocytic pathway. Curr Biol. 2008;18:1802–8.
Doherty JT, Lenhart KC, Cameron MV, Mack CP, Conlon FL, Taylor JM. Skeletal muscle differentiation and fusion are regulated by the BAR-containing Rho-GTPase-activating protein (Rho-GAP), GRAF1. J Biol Chem. 2011;286:25903–21.
Borkhardt A, Bojesen S, Haas OA, Fuchs U, Bartelheimer D, Loncarevic IF, et al. The human GRAF gene is fused to MLL in a unique t(5;11)(q31; q23) and both alleles are disrupted in three cases of myelodysplastic syndrome/acute myeloid leukemia with a deletion 5q. Proc Natl Acad Sci USA. 2000;97:9168–73.
Brittsan AG, Kranias EG. Phospholamban and cardiac contractile function. J Mol Cell Cardiol. 2000;32:2131–9.
Nishimura M, Suzuki S, Satoh T, Naito S. Tissue-specific mRNA expression profiles of human solute carrier 35 transporters. Drug Metab Pharmacokinet. 2009;24:91–99.
Kranias EG, Hajjar RJ. Modulation of cardiac contractility by the phospholamban/SERCA2a regulatome. Circ Res. 2012;110:1646–60.
Trevaskis J, Walder K, Foletta V, Kerr-Bayles L, McMillan J, Cooper A, et al. Src homology 3-domain growth factor receptorbound 2-like (endophilin) interacting protein 1, a novel neuronal protein that regulates energy balance. Endocrinology. 2005;146:3757–64.
Uezu A, Horiuchi A, Kanda K, Kikuchi N, Umeda K, Tsujita K, et al. SGIP1alpha is an endocytic protein that directly interacts with phospholipids and Eps15. J Biol Chem. 2007;282:26481–9.
Cummings N, Shields KA, Curran JE, Bozaoglu K, Trevaskis J, Gluschenko K, et al. Genetic variation in SH3-domain GRB2-like (endophilin)-interacting protein 1 has a major impact on fat mass. Int J Obes (Lond). 2012;36:201–6.
Vogel U, Jensen MK, Due KM, Rimm EB, Wallin H, Nielsen MR, et al. The NFKB1 ATTG ins/del polymorphism and risk of coronary heart disease in three independent populations. Atherosclerosis. 2011;219:200–4.
Zhou B, Rao L, Peng Y, Wang Y, Li Y, Gao L, et al. Functional polymorphism of the NFKB1 gene promoter is related to the risk of dilated cardiomyopathy. BMC Med Genet. 2009;10:47.
Karban AS, Okazaki T, Panhuysen CI, Gallegos T, Potter JJ, Bailey-Wilson JE, et al. Functional annotation of a novel NFKB1 promoter polymorphism that increases risk for ulcerative colitis. Hum Mol Genet. 2004;13:35–45.
Tang T, Cui S, Deng X, Gong Z, Jiang G, Wang P, et al. Insertion/deletion polymorphism in the promoter region of NFKB1 gene increases susceptibility for superficial bladder cancer in Chinese. DNA Cell Biol. 2010;29:9–12.
Paulussen AD, Gilissen RA, Armstrong M, Doevendans PA, Verhasselt P, Smeets HJ, et al. Genetic variations of KCNQ1, KCNH2, SCN5A, KCNE1, and KCNE2 in drug-induced long QT syndrome patients. J Mol Med (Berl). 2004;82:182–8.
Funding
Folkert W. Asselbergs is supported by UCL Hospitals NIHR Biomedical Research Centre.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
de Bakker is currently an employee of and owns equity in Vertex Pharmaceuticals. M.J. Caulfield is Chief Scientist for Genomics England a UK Government company. The remaining authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
van Setten, J., Verweij, N., Mbarek, H. et al. Genome-wide association meta-analysis of 30,000 samples identifies seven novel loci for quantitative ECG traits. Eur J Hum Genet 27, 952–962 (2019). https://doi.org/10.1038/s41431-018-0295-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41431-018-0295-z
This article is cited by
-
Phenotypic and Genetic Factors Associated with Absence of Cardiomyopathy Symptoms in PLN:c.40_42delAGA Carriers
Journal of Cardiovascular Translational Research (2023)
-
The genetic case for cardiorespiratory fitness as a clinical vital sign and the routine prescription of physical activity in healthcare
Genome Medicine (2021)
-
Translating GWAS-identified loci for cardiac rhythm and rate using an in vivo image- and CRISPR/Cas9-based approach
Scientific Reports (2020)
-
TCTEX1D1 is a genetic modifier of disease progression in Duchenne muscular dystrophy
European Journal of Human Genetics (2020)