Abstract
Genotype-first combined with reverse phenotyping has shown to be a powerful tool in human genetics, especially in the era of next generation sequencing. This combines the identification of individuals with mutations in the same gene and linking these to consistent (endo)phenotypes to establish disease causality. We have performed a MIP (molecular inversion probe)-based targeted re-sequencing study in 3,275 individuals with intellectual disability (ID) to facilitate a genotype-first approach for 24 genes previously implicated in ID.
Combining our data with data from a publicly available database, we confirmed 11 of these 24 genes to be relevant for ID. Amongst these, PHIP was shown to have an enrichment of disruptive mutations in the individuals with ID (5 out of 3,275). Through international collaboration, we identified a total of 23 individuals with PHIP mutations and elucidated the associated phenotype. Remarkably, all 23 individuals had developmental delay/ID and the majority were overweight or obese. Other features comprised behavioral problems (hyperactivity, aggression, features of autism and/or mood disorder) and dysmorphisms (full eyebrows and/or synophrys, upturned nose, large ears and tapering fingers). Interestingly, PHIP encodes two protein-isoforms, PHIP/DCAF14 and NDRP, each involved in neurodevelopmental processes, including E3 ubiquitination and neuronal differentiation. Detailed genotype-phenotype analysis points towards haploinsufficiency of PHIP/DCAF14, and not NDRP, as the underlying cause of the phenotype.
Thus, we demonstrated the use of large scale re-sequencing by MIPs, followed by reverse phenotyping, as a constructive approach to verify candidate disease genes and identify novel syndromes, highlighted by PHIP haploinsufficiency causing an ID-overweight syndrome.
Similar content being viewed by others
Introduction
Intellectual disability (ID) affects approximately 1–2% of the population in Western countries and represents a significant health burden on affected individuals and families. The majority of cases with severe ID are caused by genetic mutations [1,2,3], but the diagnosis of individual cases remains challenging due to the large genotypic and phenotypic heterogeneity. Over the past years, whole exome and whole genome sequencing studies have proven to be valuable discovery tools to identify novel candidate genes in a broad spectrum of neurodevelopmental disorders [3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19]. However, finding sufficient evidence for causality after the initial candidate locus or gene discovery can prove difficult. In addition to functional analyses of mutations, increasing the number of individuals with a (de novo) mutation in the same gene and performing deep-phenotyping in the respective individuals is not only essential to establish causality, but also to gain insight into the phenotypic spectrum related to the mutated gene. This approach is called reverse phenotyping and has been shown to be a highly effective strategy [20,21,22,23]. Moreover, it allows for quantification of the phenotypic information which helps to identify novel disease genes [24]. To increase the number of individuals with a mutation in the same gene, fast and affordable sequencing of candidate genes in a large cohort is required. Recently, molecular inversion probes (MIPs) have been optimized to fulfill these criteria [25], and have been used to identify and confirm genes for their involvement in other similarly heterogeneous phenotypes, including autism spectrum disorder [25,26,27,28], epilepsy [29, 30] and oligodontia [31].
Here we applied a genotype-first approach, in which we screened a large cohort of 3,275 individuals with ID for mutations in the coding sequence of 24 previously reported candidate ID genes [5, 6]. These genes were previously denoted as candidate ID genes based on the identification of a de novo mutation in an individual with severe ID, for which mutation severity was predicted to be (likely) pathogenic [5, 6]. The goal of this study was to identify additional individuals with ID and a de novo mutation in these genes, thereby confirming their implication in disease, and to determine whether particular genotypes manifest as clinically recognizable phenotypic entities.
Materials and methods
We performed targeted re-sequencing of the coding sequence of 24 candidate ID genes (Table 1 ) as described previously [25, 26]. A set of 1,490 MIPs was designed and DNA from 3,275 individuals with ID was collected through an international collaboration (Nijmegen, The Netherlands; Troina, Italy, Seattle, United States). MIP libraries of a maximum of 192 samples each were pooled and sequenced on one lane of a HiSeq2000 instrument (Illumina), resulting in an average coverage of >200 fold per individual. Sequence reads were mapped to UCSC human reference genome assembly hg19 and variants were called as described previously [25]. Variant annotation was performed with an in-house pipeline [3], providing information of the variant effect at cDNA and protein level, evolutionary conservation scores, and frequency information based on > 15,000 in-house exomes and public databases such as ExAC and GoNL [32,33,34].
Results and discussion
First, we focused on all likely loss-of-function (LoF) variants, defined as stop-gain, frameshift and canonical splice site mutations, predicted to result in haploinsufficiency, as the majority of genes implicated in neurodevelopmental disorders (such as ID) known to date have haploinsufficiency as their underlying pathophysiological mechanism (Supplemental Table 1 ). We identified 40 individuals with ID who carried a LoF variant in 14 different candidate ID genes (Table 1, Supplemental Table 2). Notably, for 7 of these 14 genes, we observed more LoF variants in our cohort of individuals with ID/DD than expected when compared to population frequencies using ExAC (Table 1 ). Interestingly, for some genes, such as MIB1 we identified more LoF variants in our cohort than expected, whereas in fact population constraints metrics indicate that the gene is tolerant to such loss-of-function (pLI = 0; Table 1). In line with this observation, MIB1 LoF variants could be low-penetrance or oligogenic risk factors for nonsyndromic ID. One would hypothesize that phenotypic comparison of individuals with de novo LoF in these genes would, apart from the ID resulting from inclusion bias, not yield overlap. Indeed, when individuals with these LoF mutations were invited back into clinic, and a thorough clinical evaluation was performed, no consistent phenotype, other than ID, was observed. Moreover, the majority of the variants were inherited from a healthy parent (Supplemental Table 2 ). Of note, Individual 3 carried two de novo LoF mutations in both MIB1 and PHIP, without a remarkably different phenotype from other PHIP mutation carriers.
For 19 of 40 individuals with LoF variants in these 14 candidate ID genes, parental DNAs were available to establish the inheritance pattern and showed that 10 had occurred de novo and 9 were inherited through a parent (Table 1, Supplemental Table 2). We next compared the number of LoF de novo mutations identified in our cohort to the expected number based on the gene specific de novo mutation rate, as described before [10, 35]. These analyses showed that we identified an enrichment of de novo LoF mutations in MYT1L (p = 7.59e-03), TRIO (p = 2.63e-03), WAC (p = 4.71e-3) and PHIP (p = 0.043) in our cohort of 3,275 individuals with ID/DD (Table 1 ). To increase power, we selected for de novo mutations in individuals with a neurodevelopmental disorder (ID/DD; n = 5,164 individuals) from denovo-db [36] to have occurred in either of the 24 candidate ID genes, excluding those identified in the initial discovery cohort [6]. This selection yielded 98 de novo mutations (Supplemental Table 3 ). Assessment for enrichment of DNMs (de novo mutations) in our 24 candidate ID genes identified 13 genes (54%) with enrichment for DNMs, suggesting that these genes may be involved in the etiology of ID/DD. For 10 of these 13 genes previous detailed genotype-phenotype studies conclusively linked these to ID/DD pathology (Table 1) [28, 37,38,39,40,41,42,43,44,45,46,47,48,49,50]. Focusing on the identification of genes exerting their effects through haploinsufficiency, we next statistically combined the data obtained through MIPs and those published (Table 1 ). This analysis identified the same four genes, MYT1L, WAC, TRIO, and PHIP (Table 1 ), that showed enrichment for LoF mutations in the total cohort of 8,439 individuals with ID/DD as in our targeted assay. As genotype-phenotype studies for MYT1L, TRIO, and WAC had already been performed [45, 48,49,50], including the individuals described in Table 1, we therefore focused on PHIP.
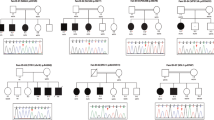
Our MIP screen identified five individuals with LoF mutations in PHIP, encoding Pleckstrin Homology Domain-Interacting Protein [MIM: 612870]. We used a previously published statistical model to determine whether the 5 initially identified individuals with LoF mutations in PHIP in the cohort consisting of 3,275 individuals with ID were phenotypically more similar to each other than could be expected for any 5 random individuals within the cohort [24]. Based on 26 commonly assessed phenotype features we found that the phenotypic similarity between the 5 individuals with a LoF mutation in PHIP was much larger than by random chance (p = 6.25e-05, permutation test, Supplementary Figure 1). In addition, we screened all private missense variants yielding one additional de novo mutation. Next we reached out to colleagues and data sharing resources to identify additional individuals with de novo mutations in PHIP. These efforts collectively identified 23 individuals with DD/ID and a pathogenic PHIP mutation (Fig. 1, Supplemental Table 4 ). For fourteen mutations, de novo origin could be confirmed by testing parental DNA samples. Interestingly, we identified 2 sib-pairs with PHIP mutations. For one of the sib-pairs (Individuals 18 and 19), we could prove inheritance of the mutation through their father, who himself also experienced learning difficulties. For the other sib-pair (Individuals 6 and 7) the mutation could not be detected in DNA from the mother and paternal DNA was unavailable for testing, leaving both paternal inheritance as well as germline mosaicism as possible explanations. Two individuals did not have pathogenic PHIP point mutations but gross chromosomal abnormalities, either disrupting the coding sequence of PHIP through translocation (Individual 13), or through total PHIP deletion (Individual 14; Supplemental Figure 2), and both had occurred de novo.
Photographs of individuals with a mutation in PHIP, overview of the PHIP gene with the 2 isoforms and location of mutations. a Photographs of 14 individuals with a mutation in PHIP of whom Individuals 18 and 19 were sisters. Shared facial features include a high forehead, full eyebrows/synophrys, characteristic upturned nose with thick alae nasi, a long philtrum and thin lips. Photographs are shown with informed consent of the individual or his/her parents. b Schematic representation of the PHIP gene (NM_017934.6) and the two protein isoforms PHIP/DCAF14 and NDRP. The point mutations are scattered throughout the coding sequence. The structural variants identified in Individuals 13 and 14 are depicted in detail in Supplemental Fig. 2. IRS, insulin receptor substrate domain; PH, pleckstrin homology domain
Detailed clinical information on the 23 individuals with a mutation in, or deletion of, PHIP is provided in Supplemental table 4 and the Supplemental Note Case reports. For all photographs shown (Fig. 1, Supplemental Fig. 3 ) informed consent was obtained. An overview of the main clinical features is shown in Table 2.
Most individuals had mild-severe ID whereas the individuals who did not have ID, had speech problems, global developmental delay in early childhood or learning difficulties. Behavioral problems were observed in 18 individuals (78%) and consisted of ADHD, features of autism, problems with impulse control, aggressive behavior, mood disorder, and anxiety. In addition to the DD/ID, the other most striking and overlapping feature in these individuals was overweight, which was observed in 17 individuals (74%), of whom 9 were obese.
Shared dysmorphic facial features comprised high forehead (67%), full eyebrows/synophrys (59%) with 15 individuals having a characteristic upturned nose with thick alae nasi (68%), a long philtrum (45%) and thin lips (36%) (Fig. 1 ). 14 individuals had large ears (64%) and 11 individuals had thick helices and earlobes (50%) (Supplemental Fig. 3 ). Most individuals (76%) had tapering fingers, bilateral clinodactyly of the fifth finger (64%) and some had bilateral cutaneous syndactyly of the second and third toes (30%) (Supplemental Fig. 3 ). Other shared features included urogenital problems (22%), vision problems (65%) and hypo/hyperpigmentation (35%).
In addition to our first report in 2012 [6], mutations in PHIP have recently also been reported in two individuals from an ASD cohort [28] and two individuals with ID, behavioral problems, overweight and characteristic facial features [51]. These reports confirm our hypothesis that mutations in PHIP lead to an ID-overweight syndrome and behavioral problems.
PHIP is translated into two different protein-isoforms: the longest isoform being Pleckstrin Homology Domain-Interacting Protein/DDB1- and CUL4-associated factor 14 (PHIP/DCAF14), and the shorter one being Neuronal Differentiation-Related Protein (NDRP), lacking the C-terminal part, when compared to the longest isoform (Fig. 1 ). The full length PHIP/DCAF14 protein contains (i) eight C-terminal WD40 repeats, forming an eight-bladed beta-propeller for protein-protein interactions, (ii) an insulin receptor substrate-1 (IRS-1) domain iii) two bromodomains, for protein-protein interaction and complex-assembly involved in transcriptional activation, and iv) a nuclear localization signal for import into the nucleus by nuclear transport. In addition to these well-defined domains, it also contains a region that is involved in pleckstrin homology (PH) domain-binding (Fig. 1 ).
Ndrp is expressed in neural tissues of mouse embryos suggesting an important role in normal neurodevelopment [52]. Depending on the developmental stage, it is either localized to the nucleus or cytosol [52]. However, there is paucity of studies on this particular isoform.
The presence of two different PHIP isoforms also leads to a speculation on likely genotype-phenotype correlations. Interestingly, insight into this matter is provided by Individual 13, who carried a translocation disrupting PHIP. Quantitative PCR in blood-derived RNA from this individual using two different primer sets, one targeting the exon-exon boundary of exons 10 and 11 (set 1), and one targeting the boundary of exons 35 and 36 (set 2), shows normal expression levels for set 1 when compared to unaffected controls, whereas set 2 shows reduced expression consistent with haploinsufficiency (Supplemental Fig. 2 ). This combination can only be explained by normal (and stable) expression of the shorter NDRP isoform, and abrogated expression of the full length PHIP/DCAF14. Given that Individual 13 is phenotypically indistinguishable from the individuals with disruptive mutations in the more C and N-terminal parts of the protein, it seems unlikely that NDRP is responsible for the phenotype observed, and suggests that disruption of the full length PHIP/DCAF14 is driving the observed phenotypic spectrum.
PHIP/DCAF14 has been shown to play an important role in several processes linked to neurodevelopment. First, PHIP/DCAF14 is one of the multiple substrate receptors of the proteolytic CUL4-DDB1, a cullin-RING finger ligase complex that binds target proteins for ubiquitination which results in degradation of the protein [53]. Mutations in the X-linked CUL4B component of the complex, lead to an ID syndrome in males, with macrocephaly, epilepsy and cerebral malformations [54,55,56]. Second, PHIP/DCAF14 is considered a regulator of the insulin and insulin-like growth factor signaling pathways via specific binding of the PH domain of the IRS-1 [57, 58]. Insulin receptor (IR) signaling plays an important role in the brain since IRs are found in synapses and dendrites of neurons and insulin stimulates the formation of dendritic spines and synapses via the PI3K/Akt/mTOR and Rac1 pathways [10, 59, 60]. Although the name and function might suggest a role in the development of diabetes, only 1 individual in our cohort (Individual 22) had type 2 diabetes, which could also be caused by her obesity.
Third, PHIP1/DCAF14 belongs to bromodomain family III which includes EP300 (E1A-binding protein p300 [MIM: 602700]), CREBBP (CREB-binding protein [MIM: 600140]) and BRWD3 (Bromodomain- and WD repeat-containing protein 3 [MIM: 300553]) [61]. Mutations in EP300 and CREBBP are known causes of Rubinstein-Taybi syndrome whereas BRWD3 mutations result in an X-linked form of ID (MRX93) [62]. Although individuals with a mutation in PHIP are facially distinct from Rubinstein-Taybi syndrome, their facial features do resemble the facial features of individuals with a mutation in BRWD3, such as the high forehead, synophrys and large ears. Notably, a substantial fraction of the individuals with mutations in either EP300, CREBBP [OMIM #180849, #613684], or BRWD3 are overweight [62,63,64]. PHIP itself seems to have a positive effect on cell-growth [65]. Mice deficient of Phip show postnatal growth retardation that is similar to IGF-1 null mice [66]. Although individuals with ID have a higher risk of being overweight in general [67], it is well known that some specific ID syndromes are associated with overweight/obesity [68] and based on our data, it appears that mutations in PHIP can be added to the list of ID-overweight syndromes.
In summary, we present a large genotype-first, targeted-re-sequencing effort, using MIPs for 24 candidate ID genes in >3,500 individuals with ID, aimed at the identification of additional mutations in these individuals. This screen, in combination with database resources and reverse phenotyping, verified the true nature of 11 candidate ID genes, and identified an ID-overweight syndrome caused by PHIP haploinsufficiency. The latter provides not only novel insights into the genetic basis and physiology of intellectual disability but overweight as well.
Web resources
ExAC Browser, http://exac.broadinstitute.org/
GeneMatcher, https://genematcher.org/
GoNL, http://www.nlgenome.nl/
OMIM, http://www.omim.org/
References
Veltman JA, Brunner HG. De novo mutations in human genetic disease. Nat Rev Genet 2012;13:565–75.
Vissers LE, Gilissen C, Veltman JA. Genetic studies in intellectual disability and related disorders. Nat Rev Genet 2016;17:9–18.
Gilissen C, Hehir-Kwa JY, Thung DT, et al. Genome sequencing identifies major causes of severe intellectual disability. Nature 2014;511:344–7.
Rauch A, Wieczorek D, Graf E, et al. Range of genetic mutations associated with severe non-syndromic sporadic intellectual disability: an exome sequencing study. Lancet 2012;380:1674–82.
Vissers LE, de Ligt J, Gilissen C, et al. A de novo paradigm for mental retardation. Nat Genet 2010;42:1109–12.
de Ligt J, Willemsen MH, van Bon BW, et al. Diagnostic exome sequencing in persons with severe intellectual disability. N Engl J Med 2012;367:1921–9.
Grozeva D, Carss K, Spasic-Boskovic O, et al. Targeted next-generation sequencing analysis of 1,000 individuals with intellectual disability. Hum Mutat 2015;36:1197–204.
Study., D.D.D.. Large-scale discovery of novel genetic causes of developmental disorders. Nature 2015;519:223–8.
Hamdan FF, Srour M, Capo-Chichi JM, et al. De novo mutations in moderate or severe intellectual disability. PLoS Genet 2014;10:e1004772.
Lelieveld SH, Reijnders MR, Pfundt R, et al. Meta-analysis of 2,104 trios provides support for 10 new genes for intellectual disability. Nat Neurosci 2016;19:1194–6.
Neale BM, Kou Y, Liu L, et al. Patterns and rates of exonic de novo mutations in autism spectrum disorders. Nature 2012;485:242–5.
Iossifov I, O’Roak BJ, Sanders SJ, et al. The contribution of de novo coding mutations to autism spectrum disorder. Nature 2014;515:216–21.
EuroEPINOMICS-RES Consortium; Epilepsy Phenome/Genome Project; Epi4K Consortium. De novo mutations in synaptic transmission genes including DNM1 cause epileptic encephalopathies. Am J Hum Genet 2014;95:360–70.
Xu B, Ionita-Laza I, Roos JL, et al. De novo gene mutations highlight patterns of genetic and neural complexity in schizophrenia. Nat Genet 2012;44:1365–9.
Xu B, Roos JL, Dexheimer P, Boone B, Plummer B, Levy S, Gogos JA, Karayiorgou M. Exome sequencing supports a de novo mutational paradigm for schizophrenia. Nat Genet 2011;43:864–8.
McCarthy SE, Gillis J, Kramer M, et al. De novo mutations in schizophrenia implicate chromatin remodeling and support a genetic overlap with autism and intellectual disability. Mol Psychiatry 2014;19:652–8.
Gulsuner S, Walsh T, Watts AC, et al. Spatial and temporal mapping of de novo mutations in schizophrenia to a fetal prefrontal cortical network. Cell 2013;154:518–29.
Fromer M, Pocklington AJ, Kavanagh DH, et al. De novo mutations in schizophrenia implicate synaptic networks. Nature 2014;506:179–84.
McRae JF, Clayton S, Fitzgerald TW, et al. Prevalence, phenotype and architecture of developmental disorders caused by de novo mutation. bioRxiv 2016.
Stessman HA, Bernier R, Eichler EE. A genotype-first approach to defining the subtypes of a complex disease. Cell 2014;156:872–7.
de Goede C, Yue WW, Yan G, et al. Role of reverse phenotyping in interpretation of next generation sequencing data and a review of INPP5E related disorders. Eur J Paediatr Neurol 2016;20:286–295.
Nambot S, Masurel A, El Chehadeh S, Mosca-Boidron AL, Thauvin-Robinet C, Lefebvre M, Marle N. and Thevenon, J. 9q33.3q34.11 microdeletion: new contiguous gene syndrome encompassing STXBP1, LMX1B and ENG genes assessed using reverse phenotyping. Eur J Hum Genet 2016;24:830–837.
Stessman HA, Xiong B, Coe BP, et al. Targeted sequencing identifies 91 neurodevelopmental-disorder risk genes with autism and developmental-disability biases. Nat Genet 2017;49:515–26.
Vulto-van Silfhout AT, Gilissen C, Goeman JJ, et al. Quantification of phenotype information aids the identification of novel disease genes. Hum Mutat 2017;38:594–99.
O’Roak BJ, Vives L, Fu W, et al. Multiplex targeted sequencing identifies recurrently mutated genes in autism spectrum disorders. Science 2012;338:1619–22.
Coe BP, Witherspoon K, Rosenfeld JA, et al. Refining analyses of copy number variation identifies specific genes associated with developmental delay. Nat Genet 2014;46:1063–71.
O’Roak BJ, Stessman HA, Boyle EA, et al. Recurrent de novo mutations implicate novel genes underlying simplex autism risk. Nature Commun 2014;5:5595.
Wang T, Guo H, Xiong B, et al. De novo genic mutations among a Chinese autism spectrum disorder cohort. Nature Commun 2016;7:13316.
Carvill GL, Heavin SB, Yendle SC, et al. Targeted resequencing in epileptic encephalopathies identifies de novo mutations in CHD2 and SYNGAP1. Nat Genet 2013;45:825–30.
Hildebrand MS, Myers CT, Carvill GL, et al. A targeted resequencing gene panel for focal epilepsy. Neurology 2016;86:1605–12.
Ockeloen CW, Khandelwal KD, Dreesen K, et al. Novel mutations in LRP6 highlight the role of WNT signaling in tooth agenesis. Genet Med 2016;18:1158–62.
Lek M, Karczewski KJ, Minikel EV, et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 2016;536:285–91.
Genome of the Netherlands Consortium. Whole-genome sequence variation, population structure and demographic history of the Dutch population. Nat Genet 2014;46:818–25.
Francioli LC, Polak PP, Koren A, et al. Genome-wide patterns and properties of de novo mutations in humans. Nat Genet 2015;47:822–6.
Samocha KE, Robinson EB, Sanders SJ, et al. A framework for the interpretation of de novo mutation in human disease. Nat Genet 2014;46:944–50.
Turner TN, Yi Q, Krumm N, Huddleston J, Hoekzema K, HA FS, Doebley AL, Bernier RA, Nickerson DA, Eichler EE. denovo-db: a compendium of human de novo variants. Nucleic Acids Res 2017;45:D804–d811.
Deciphering Developmental Disorders Study. Large-scale discovery of novel genetic causes of developmental disorders. Nature 2015;519:223–8.
Vulto-van Silfhout AT, Rajamanickam S, Jensik PJ, et al. Mutations affecting the SAND domain of DEAF1 cause intellectual disability with severe speech impairment and behavioral problems. Am J Hum Genet 2014;94:649–61.
Willemsen MH, Vissers LE, Willemsen MA, et al. Mutations in DYNC1H1 cause severe intellectual disability with neuronal migration defects. J Med Genet 2012;49:179–83.
Willemsen MH, Ba W, Wissink-Lindhout WM, de Brouwer AP, et al. Involvement of the kinesin family members KIF4A and KIF5C in intellectual disability and synaptic function. J Med Genet 2014;51:487–94.
Poirier K, Lebrun N, Broix L, et al. Mutations in TUBG1, DYNC1H1, KIF5C and KIF2A cause malformations of cortical development and microcephaly. Nat Genet 2013;45:639–47.
Inui T, Kobayashi S, Ashikari Y, et al. Two cases of early-onset myoclonic seizures with continuous parietal delta activity caused by EEF1A2 mutations. Brain Dev 2016;38:520–4.
Nakajima J, Okamoto N, Tohyama J, et al. De novo EEF1A2 mutations in patients with characteristic facial features, intellectual disability, autistic behaviors and epilepsy. Clin Genet 2015;87:356–61.
Cavallin M, Hubert L, Cantagrel V, et al. Recurrent KIF5C mutation leading to frontal pachygyria without microcephaly. Neurogenetics 2016;17:79–82.
De Rocker N, Vergult S, Koolen D, et al. Refinement of the critical 2p25.3 deletion region: the role of MYT1L in intellectual disability and obesity. Genet Med 2015;17:460–6.
Houge G, Haesen D, Vissers LE, et al. B56delta-related protein phosphatase 2A dysfunction identified in patients with intellectual disability. J Clin Invest 2015;125:3051–62.
Loveday C, Tatton-Brown K, Clarke M, et al. Mutations in the PP2A regulatory subunit B family genes PPP2R5B, PPP2R5C and PPP2R5D cause human overgrowth. Hum Mol Genet 2015;24:4775–9.
Ba W, Yan Y, Reijnders MR, et al. TRIO loss of function is associated with mild intellectual disability and affects dendritic branching and synapse function. Hum Mol Genet 2016;25:892–902.
DeSanto C, D’Aco K, Araujo GC, et al. WAC loss-of-function mutations cause a recognisable syndrome characterised by dysmorphic features, developmental delay and hypotonia and recapitulate 10p11.23 microdeletion syndrome. J Med Genet 2015;52:754–61.
Lugtenberg D, Reijnders MR, Fenckova M, et al. De novo loss-of-function mutations in WAC cause a recognizable intellectual disability syndrome and learning deficits in Drosophila. Eur J Hum Genet 2016;24:1145–53.
Webster E, Cho MT, Alexander N, et al. De novo PHIP-predicted deleterious variants are associated with developmental delay, intellectual disability, obesity, and dysmorphic features. Cold Spring Harb Mol Case Stud 2016;2:a001172.
Kato H, Chen S, Kiyama H, Ikeda K, Kimura N, Nakashima K, Taga T. Identification of a novel WD repeat-containing gene predominantly expressed in developing and regenerating neurons. J Biochem 2000;128:923–32.
Lee J, Zhou P. DCAFs, the missing link of the CUL4-DDB1 ubiquitin ligase. Mol Cell 2007;26:775–80.
Vulto-van Silfhout AT, Nakagawa T, Bahi-Buisson N, et al. Variants in CUL4B are associated with cerebral malformations. Hum Mutat 2015;36:106–17.
Tarpey PS, Raymond FL, O’Meara S, et al. Mutations in CUL4B, which encodes a ubiquitin E3 ligase subunit, cause an X-linked mental retardation syndrome associated with aggressive outbursts, seizures, relative macrocephaly, central obesity, hypogonadism, pes cavus, and tremor. Am J Hum Genet 2007;80:345–52.
Zou Y, Liu Q, Chen B, et al. Mutation in CUL4B, which encodes a member of cullin-RING ubiquitin ligase complex, causes X-linked mental retardation. Am J Hum Genet 2007;80:561–6.
Farhang-Fallah J, Yin X, Trentin G, Cheng AM, Rozakis-Adcock M. Cloning and characterization of PHIP, a novel insulin receptor substrate-1 pleckstrin homology domain interacting protein. J Biol Chem 2000;275:40492–7.
Farhang-Fallah J, Randhawa VK, Nimnual A, Klip A, Bar-Sagi D, Rozakis-Adcock M. The pleckstrin homology (PH) domain-interacting protein couples the insulin receptor substrate 1 PH domain to insulin signaling pathways leading to mitogenesis and GLUT4 translocation. Mol Cell Biol 2002;22:7325–36.
Lee CC, Huang CC, Hsu KS. Insulin promotes dendritic spine and synapse formation by the PI3K/Akt/mTOR and Rac1 signaling pathways. Neuropharmacology 2011;61:867–79.
Wang L, Zhou K, Fu Z, Yu D, Huang H, Zang X, Mo X. Brain development and akt signaling: the crossroads of signaling pathway and neurodevelopmental diseases. J Mol Neurosci 2016;61:379–84.
Filippakopoulos P, Picaud S, Mangos M, et al. Histone recognition and large-scale structural analysis of the human bromodomain family. Cell 2012;149:214–31.
Field M, Tarpey PS, Smith R, et al. Mutations in the BRWD3 gene cause X-linked mental retardation associated with macrocephaly. Am J Hum Genet 2007;81:367–74.
Fergelot P, Van Belzen M, Van Gils J, et al. Phenotype and genotype in 52 patients with Rubinstein-Taybi syndrome caused by EP300 mutations. Am J Med Genet A 2016;170:3069–82.
Grotto S, Drouin-Garraud V, Ounap K, et al. Clinical assessment of five patients with BRWD3 mutation at Xq21.1 gives further evidence for mild to moderate intellectual disability and macrocephaly. Eur J Med Genet 2014;57:200–6.
Podcheko A, Northcott P, Bikopoulos G, Lee A, Bommareddi SR, Kushner JA, Farhang-Fallah J, Rozakis-Adcock M. Identification of a WD40 repeat-containing isoform of PHIP as a novel regulator of beta-cell growth and survival. Mol Cell Biol 2007;27:6484–96.
Li S, Francisco AB, Han C, et al. The full-length isoform of the mouse pleckstrin homology domain-interacting protein (PHIP) is required for postnatal growth. FEBS Lett 2010;584:4121–7.
Hsieh K, Rimmer JH, Heller T. Obesity and associated factors in adults with intellectual disability. J Intellect Disabil Res 2014;58:851–63.
Sabin MA, Werther GA, Kiess W. Genetics of obesity and overgrowth syndromes. Best Pract Res Clin Endocrinol Metab 2011;25:207–20.
Acknowledgements
We are grateful to all individuals and their parents for participating in this study.
We are also grateful to the Radboud Genomics Technology Center for performing Sanger sequencing. We also want to thank Choli Lee for technical assistance with HiSeq sequencing. We thank Boris Keren, AP-HP, Groupe Hospitalier Pitié-Salpêtrière, Département de Génétique, Paris, France, for interpretation of the exome data of Individual 15.
We thank Ineke van der Burgt, Human Genetics, Radboudumc, Nijmegen, The Netherlands and Christina Ringmann Fagerberg, Department of Clinical Genetics, Odense University Hospital, Denmark for clinical support.
We are grateful to all of the families at the participating Simons Simplex Collection (SSC) sites, as well as the principal investigators (A. Beaudet, R. Bernier, J. Constantino, E. Cook, E. Fombonne, D. Geschwind, R. Goin-Kochel, E. Hanson, D. Grice, A. Klin, D. Ledbetter, C. Lord, C. Martin, D. Martin, R. Maxim, J. Miles, O. Ousley, K. Pelphrey, B. Peterson, J. Piggot, C. Saulnier, M. State, W. Stone, J. Sutcliffe, C. Walsh, Z. Warren, E. Wijsman). We appreciate obtaining access to phenotypic data on SFARI Base.
This work was financially supported by grants from the Netherlands Organisation for Health Research and Development (917-86-319 to B.B.A.d.V., 912-12-109 to B.B.A.d.V. and J.A.V., 907-00-365 to T.K., and 918-15-667 to J.A.V) and the European Research Council (ERC; starting grant DENOVO 281964 to J.A.V.). This research was supported, in part, by the following: Simons Foundation Autism Research Initiative (SFARI 303241) to E.E.E., National Institutes of Health (R01MH101221 to E.E.E. and R01MH100047 to R.A.B.). E.E.E. is an investigator of the Howard Hughes Medical Institute. J.G. is supported by NHMRC research fellowship 1041920 and Channel 7 Children’s Research Foundation. This work was supported by the Italian Ministry of Health and “5 per mille” funding.
The authors would like to thank the Exome Aggregation Consortium and the groups that provided exome variant data for comparison. A full list of contributing groups can be found at http://exac.broadinstitute.org/about.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
GLC is a member of the scientific advisory board of Ambry Genetics.
Additional information
Sandra Jansen, Alexander Hoischen, Evan E. Eichler and Bert B. A. de Vries contributed equally to this work
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Jansen, S., Hoischen, A., Coe, B.P. et al. A genotype-first approach identifies an intellectual disability-overweight syndrome caused by PHIP haploinsufficiency. Eur J Hum Genet 26, 54–63 (2018). https://doi.org/10.1038/s41431-017-0039-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41431-017-0039-5
This article is cited by
-
A Comprehensive Review of Syndromic Forms of Obesity: Genetic Etiology, Clinical Features and Molecular Diagnosis
Current Obesity Reports (2024)
-
Novel loss-of-function variants in TRIO are associated with neurodevelopmental disorder: case report
BMC Medical Genetics (2020)
-
Quantitative genome-wide association analyses of receptive language in the Danish High Risk and Resilience Study
BMC Neuroscience (2020)
-
Genetic Variation in RIN3 in the Belgian Population Supports Its Involvement in the Pathogenesis of Paget’s Disease of Bone and Modifies the Age of Onset
Calcified Tissue International (2019)
-
Comprehensive genetic analysis of 57 families with clinically suspected Cornelia de Lange syndrome
Journal of Human Genetics (2019)