Abstract
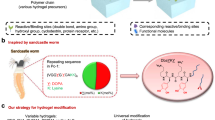
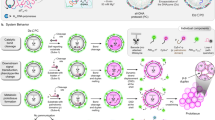
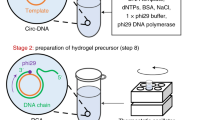
Advancements in DNA nanotechnology and nanobioengineering allowed the use of DNA as a generic material, as well as a genetic material. Among various DNA-based materials, DNA hydrogels achieve a DNA-based network at the bulk scale. Notably, a recently developed enzyme-based fabrication method established a route to synthesize physically crosslinked networks from DNA. This new class of DNA networks paved the way for creating different forms of DNA materials by utilizing the characteristics of a biofunctional polymer. This Focus Review provides a brief overview of the enzyme-based fabrication method of physical DNA hydrogels and discusses the latest developments in technology and their applications.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Seeman NC, Sleiman HF. DNA nanotechnology. Nat Rev Mater. 2018;3:17068. https://doi.org/10.1038/natrevmats.2017.68.
Hamada S, Murata S. Substrate-assisted assembly of interconnected single-duplex DNA nanostructures. Angew Chem Int Ed Engl. 2009;48:6820–3. https://doi.org/10.1002/anie.200902662.
Cutler JI, Auyeung E, Mirkin CA. Spherical nucleic acids. J Am Chem Soc. 2012;134:1376–91. https://doi.org/10.1021/ja209351u.
Hamada S, Tan SJ, Luo D. Nanoparticle crystallization: DNA-bonded ‘atoms’. Nat Mater 2014;13:121–2. https://doi.org/10.1038/nmat3876.
Tan SJ, Campolongo MJ, Luo D, Cheng W. Building plasmonic nanostructures with DNA. Nat Nanotechnol. 2011;6:268–76. https://doi.org/10.1038/nnano.2011.49.
Um SH, Lee JB, Kwon SY, Li Y, Luo D. Dendrimer-like DNA-based fluorescence nanobarcodes. Nat Protoc. 2006;1:995–1000. https://doi.org/10.1038/nprot.2006.141.
Li F, Tang J, Geng J, Luo D, Yang D. Polymeric DNA hydrogel: design, synthesis and applications. Prog Polym Sci. 2019;98:101163. https://doi.org/10.1016/j.progpolymsci.2019.101163.
Yang D, Hartman MR, Derrien TL, Hamada S, An D, Yancey KG, et al. DNA materials: bridging nanotechnology and biotechnology. Acc Chem Res. 2014;47:1902–11. https://doi.org/10.1021/ar5001082.
Wang D, Hu Y, Liu P, Luo D. Bioresponsive DNA hydrogels: beyond the conventional stimuli responsiveness. Acc Chem Res. 2017;50:733–9. https://doi.org/10.1021/acs.accounts.6b00581.
Um SH, Lee JB, Park N, Kwon SY, Umbach CC, Luo D. Enzyme-catalysed assembly of DNA hydrogel. Nat Mater. 2006;5:797–801. https://doi.org/10.1038/nmat1741.
Cheng W, Campolongo MJ, Cha JJ, Tan SJ, Umbach CC, Muller DA, et al. Free-standing nanoparticle superlattice sheets controlled by DNA. Nat Mater. 2009;8:519–25. https://doi.org/10.1038/nmat2440.
Cheng W, Park N, Walter MT, Hartman MR, Luo D. Nanopatterning self-assembled nanoparticle superlattices by moulding microdroplets. Nat Nanotechnol. 2008;3:682–90. https://doi.org/10.1038/nnano.2008.279.
Li Y, Cu YT, Luo D. Multiplexed detection of pathogen DNA with DNA-based fluorescence nanobarcodes. Nat Biotechnol. 2005;23:885–9. https://doi.org/10.1038/nbt1106.
Derrien TL, Hamada S, Zhou M, Smilgies D-M, Luo D. Three-dimensional nanoparticle assemblies with tunable plasmonics via a layer-by-layer process. Nano Today. 2020;30:100823. https://doi.org/10.1016/j.nantod.2019.100823.
Lee JB, Peng S, Yang D, Roh YH, Funabashi H, Park N, et al. A mechanical metamaterial made from a DNA hydrogel. Nat Nanotechnol. 2012;7:816–20. https://doi.org/10.1038/nnano.2012.211.
Gilbert W, Dressler D. DNA replication: the rolling circle model. Cold Spring Harb Symp Quant Biol. 1968;33:473–84. https://doi.org/10.1101/sqb.1968.033.01.055.
Demidov VV. Introduction: 20+ years of rolling the DNA minicircles—state of the art in the RCA-based nucleic acid diagnostics and therapeutics. In: Demidov VV, editor. Rolling Circle Amplification (RCA). Springer; Cham, Switzerland, 2016. p. 1–7. http://www.springer.com/gp/book/9783319422244.
Johne R, Muller H, Rector A, van Ranst M, Stevens H. Rolling-circle amplification of viral DNA genomes using phi29 polymerase. Trends Microbiol. 2009;17:205–11. https://doi.org/10.1016/j.tim.2009.02.004.
Mohsen MG, Kool ET. The discovery of rolling circle amplification and rolling circle transcription. Acc Chem Res. 2016;49:2540–50. https://doi.org/10.1021/acs.accounts.6b00417.
Xu X, Jagota A, Peng S, Luo D, Wu M, Hui CY. Gravity and surface tension effects on the shape change of soft materials. Langmuir. 2013;29:8665–74. https://doi.org/10.1021/la400921h.
Park N, Um SH, Funabashi H, Xu J, Luo D. A cell-free protein-producing gel. Nat Mater. 2009;8:432–7. https://doi.org/10.1038/nmat2419.
Kahn JS, Ruiz RC, Sureka S, Peng S, Derrien TL, An D, et al. DNA microgels as a platform for cell-free protein expression and display. Biomacromolecules. 2016;17:2019–26. https://doi.org/10.1021/acs.biomac.6b00183.
Park N, Kahn JS, Rice EJ, Hartman MR, Funabashi H, Xu J, et al. High-yield cell-free protein production from P-gel. Nat Protoc. 2009;4:1759–70. https://doi.org/10.1038/nprot.2009.174.
Lee HY, Jeong H, Jung IY, Jang B, Seo YC, Lee H, et al. DhITACT: DNA hydrogel formation by isothermal amplification of complementary target in fluidic channels. Adv Mater. 2015;27:3513–7. https://doi.org/10.1002/adma.201500414.
Lin C, Wang X, Liu Y, Seeman NC, Yan H. Rolling circle enzymatic replication of a complex multi-crossover DNA nanostructure. J Am Chem Soc. 2007;129:14475–81. https://doi.org/10.1021/ja0760980.
Hamblin GD, Carneiro KM, Fakhoury JF, Bujold KE, Sleiman HF. Rolling circle amplification-templated DNA nanotubes show increased stability and cell penetration ability. J Am Chem Soc. 2012;134:2888–91. https://doi.org/10.1021/ja2107492.
Wilner OI, Willner I. Functionalized DNA nanostructures. Chem Rev. 2012;112:2528–56. https://doi.org/10.1021/cr200104q.
Wilner OI, Orbach R, Henning A, Teller C, Yehezkeli O, Mertig M, et al. Self-assembly of DNA nanotubes with controllable diameters. Nat Commun. 2011;2:540. https://doi.org/10.1038/ncomms1535.
Jang B, Kim B, Kim H, Kwon H, Kim M, Seo Y, et al. Enzymatic synthesis of self-assembled dicer substrate rna nanostructures for programmable gene silencing. Nano Lett. 2018;18:4279–84. https://doi.org/10.1021/acs.nanolett.8b01267.
Valero J, Pal N, Dhakal S, Walter NG, Famulok M. A bio-hybrid DNA rotor-stator nanoengine that moves along predefined tracks. Nat Nanotechnol. 2018;13:496–503. https://doi.org/10.1038/s41565-018-0109-z.
Ouyang X, Li J, Liu H, Zhao B, Yan J, Ma Y, et al. Rolling circle amplification-based DNA origami nanostructrures for intracellular delivery of immunostimulatory drugs. Small. 2013;9:3082–7. https://doi.org/10.1002/smll.201300458.
Kageyama R, Kawamata I, Tanabe K, Suzuki Y, Nomura SM, Murata S. Construction of T-motif-based DNA nanostructures through enzymatic reactions. Chembiochem 2018;19:873–6. https://doi.org/10.1002/cbic.201700682.
Zhu G, Hu R, Zhao Z, Chen Z, Zhang X, Tan W. Noncanonical self-assembly of multifunctional DNA nanoflowers for biomedical applications. J Am Chem Soc. 2013;135:16438–45. https://doi.org/10.1021/ja406115e.
Lee JB, Hong J, Bonner DK, Poon Z, Hammond PT. Self-assembled RNA interference microsponges for efficient siRNA delivery. Nat Mater. 2012;11:316–22. https://doi.org/10.1038/nmat3253.
Hamada S, Yancey KG, Pardo Y, Gan M, Vanatta M, An D, et al. Dynamic DNA material with emergent locomotion behavior powered by artificial metabolism. Sci Robot. 2019;4:eaaw3512. https://doi.org/10.1126/scirobotics.aaw3512.
Lv Y, Hu R, Zhu G, Zhang X, Mei L, Liu Q, et al. Preparation and biomedical applications of programmable and multifunctional DNA nanoflowers. Nat Protoc. 2015;10:1508–24. https://doi.org/10.1038/nprot.2015.078.
Shopsowitz KE, Roh YH, Deng ZJ, Morton SW, Hammond PT. RNAi-microsponges form through self-assembly of the organic and inorganic products of transcription. Small. 2014;10:1623–33. https://doi.org/10.1002/smll.201302676.
Han D, Park Y, Kim H, Lee JB. Self-assembly of free-standing RNA membranes. Nat Commun. 2014;5:4367. https://doi.org/10.1038/ncomms5367.
Ali MM, Li F, Zhang Z, Zhang K, Kang DK, Ankrum JA, et al. Rolling circle amplification: a versatile tool for chemical biology, materials science and medicine. Chem Soc Rev. 2014;43:3324–41. https://doi.org/10.1039/c3cs60439j.
Gu L, Yan W, Liu L, Wang S, Zhang X, Lyu M. Research progress on rolling circle amplification (RCA)-based biomedical sensing. Pharmaceuticals 2018;11:35. https://doi.org/10.3390/ph11020035.
Hu R, Zhang X, Zhao Z, Zhu G, Chen T, Fu T, et al. DNA nanoflowers for multiplexed cellular imaging and traceable targeted drug delivery. Angew Chem Int Ed Engl. 2014;53:5821–6. https://doi.org/10.1002/anie.201400323.
Zhang L, Zhu G, Mei L, Wu C, Qiu L, Cui C, et al. Self-assembled DNA immunonanoflowers as multivalent CpG nanoagents. ACS Appl Mater Interfaces. 2015;7:24069–74. https://doi.org/10.1021/acsami.5b06987.
Han S, Lee JS, Lee JB. Synthesis of a multi-functional DNA nanosphere barcode system for direct cell detection. Nanoscale. 2017;9:14094–102. https://doi.org/10.1039/c7nr03615a.
Rosenwasser D, Hamada S, Luo D, Sabin JE. PolyBrick 3.0: live signatures through DNA hydrogels and digital ceramics. Int J Rapid Manuf. 2018;7:203. https://doi.org/10.1504/ijrapidm.2018.092909.
Acknowledgements
This work is supported by the U.S. National Science Foundation (NSF) (RoL: EAGER: DESYN-C3: 1844310 and SNM-1530522).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Hamada, S., Luo, D. Enzyme-based fabrication of physical DNA hydrogels: new materials and applications. Polym J 52, 891–898 (2020). https://doi.org/10.1038/s41428-020-0340-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41428-020-0340-y
This article is cited by
-
Phosphorylase-catalyzed synthesis and self-assembled structures of cellulose oligomers in the presence of protein denaturants
Polymer Journal (2022)
-
Nucleic acid-based fluorescent sensor systems: a review
Polymer Journal (2022)
-
Construction of a reduction-responsive oligonucleotide via a post-modification approach utilizing 4-nitrophenyl diazomethane
Polymer Journal (2021)