Abstract
Polypeptides containing periodic aromatic residues in their main chains were synthesized via papain-catalyzed chemoenzymatic polymerization of tripeptide ester monomers under moderate conditions in aqueous buffers. As the monomer, 4-aminobenzoic acid (Abz) was modified with Gly or Ala at the N- and C-termini to mitigate the poor substrate specificity of papain to unnatural amino acids. The tripeptide esters, namely, GlyAbzGly and AlaAbzAla ethyl esters, can be recognized and polymerized by papain, resulting in the polypeptides poly(GlyAbzGly) and poly(AlaAbzAla), respectively, with periodic sequences containing Abz units every three residues. Copolymerization of tripeptide esters with Gly or Ala ethyl ester also proceeded in the presence of papain. The secondary structures of the Abz-containing polypeptides were investigated by IR and wide-angle X-ray diffraction (WAXD) analysis. The WAXD profile of poly(GlyAbzGly) was similar to that of polyGly, whereas poly(AlaAbzAla) adopted a sheet-like structure similar to the β-sheet of polyAla. Thermal analysis of Abz-containing polypeptides revealed that the high thermal stability of the Abz-containing polypeptides is related to the distinct sequences that periodically include Abz residues.
Similar content being viewed by others
Introduction
Polypeptides are fascinating materials in various fields because of their structural variety arising from various amino acid sequences. The physical and biological properties of polypeptides are strongly related to higher-order structures assembled via intra-/interchain interactions between amino acid residues, such as hydrogen bonds. Precise amino acid sequences in the polypeptide backbone of natural proteins are essential to construct various secondary structures, which further assemble into more complicated architectures. In particular, structural proteins have highly repetitive periodic sequences and thereby form specific structures that impart an appropriate mechanical strength to living organisms or tissues. For example, collagen is dominantly composed of repetitive GlyXaaYaa (mainly Xaa=Pro; Yaa=Hpr: 4-hydroxyproline) motifs in its sequence [1, 2]. Collagen forms a left-handed triple helix driven by the periodic sequence and exhibits excellent hardness to support living tissues. Therefore, it is important to synthesize polypeptides with appropriate sequences to achieve desired physical properties.
Polypeptides are conventionally synthesized by condensation between the amino and carboxy groups of amino acids using condensing agents. Sequential control during polypeptide synthesis demands specialized techniques to achieve complicated sequences. Precise construction of functional polypeptide sequences has been achieved by solid-phase peptide synthesis (SPPS), which utilizes solid supports to realize sequential condensation and deprotection reactions [3]. The SPPS method is a powerful tool to synthesize complicated polypeptide sequences with specific characteristics [4, 5], but the available polypeptide length is limited within a relatively short chain range. However, a solution polymerization method has also been developed for polypeptide synthesis. Conventional polycondensation of amino acids, including their derivatives, generally suffers from side reactions resulting in the formation of undesired cyclic oligomers [6,7,8]. Ring-opening polymerization of N-carboxy amino acid anhydrides (NCAs) is the most successful method to synthesize polypeptides in solution; however, the sequence of the resulting polypeptides is limited to homo, random, or block copolymers [9,10,11,12,13]. Recently, some unique synthetic methods were developed to synthesize polypeptides composed of unique periodic sequences. Okamoto et al. demonstrated that polypeptides containing alternating leucine and aromatic amino acids were synthesized by multiple stepwise condensation reactions [14]. Koyama et al. developed a novel synthetic method for polypeptides based on the Ugi reaction to construct a polypeptide backbone from imine and isocyanide-modified carboxylate monomers, resulting in alternating polypeptides [15, 16]. This reaction is a tandem reaction, including rearrangement to form amide bonds, which allows broad structural variations of the N-substituent of the amide groups. As another example, chemoenzymatic synthesis using proteases is a green method to synthesize various polypeptide materials [17]. Aminolysis reactions of amino acid esters catalyzed by proteases proceed under kinetically controlled conditions, resulting in polypeptide formation with excellent regio- and stereoselectivity. Because oligopeptide esters can also be activated by proteases, various periodic sequences are easily obtained by chemoenzymatic polymerization. Gross et al. synthesized alternating polypeptides via protease-catalyzed polymerization of dipeptide esters, such as AlaGly and LysLeu ethyl esters [18, 19]. We also demonstrated that the papain-catalyzed copolymerization of the ValProGly tripeptide and ValGly dipeptide esters afforded poly(ValProGly-co-ValGly), which possesses a periodic sequence similar to that of elastin, a structural protein exhibiting excellent elasticity [20]. The resulting polypeptides showed a reversible structural transition in response to temperature change, which is an essential characteristic for elastin to form a cross-linked hierarchical structure.
The introduction of unnatural structures into a polypeptide backbone allows polypeptide materials to obtain novel functionalities and improved physical properties. In our previous study, we successfully introduced nylon units, a synthetic amino acid for commercial polyamides, into polypeptides via chemoenzymatic polymerization of leucine and nylon ethyl esters using papain [21, 22]. A melting point could be detected for the polypeptide with a random sequence of leucine and nylon units, whereas no such transition could be detected for polyleucine below its decomposition temperature. This result indicates that polypeptides containing nylon units can be subjected to thermal processing for practical use. In addition, di- and trifunctionalized synthetic substrates can also be involved in chemoenzymatic polymerization, resulting in the formation of special structures. The polymerization of l-phenylalanine or l-lysine ethyl ester in the presence of a triamine compound afforded star-shaped polypeptides [23, 24]. On the other hand, l-alanine ethyl ester was polymerized with a bis(amino acid ester) derivative using papain to give telechelic-type polypeptides [25, 26]. These uniquely shaped polypeptides exhibited unique physical properties allowing the polypeptides to be utilized as carriers for gene delivery or as fillers to reinforce structural materials. Despite the usefulness of unnatural amino acids, however, these nylon ester monomers suffer from low polymerization efficiency because of the low substrate specificity of papain. Recently, we demonstrated that the poor affinity of unnatural amino acids to proteases can be mitigated by modification with natural amino acids in chemoenzymatic polymerization. A tripeptide ester consisting of 2-aminoisobutyric acid (Aib) between two alanine residues was efficiently polymerized in the presence of papain, whereas the chemoenzymatic polymerization of Aib ester did not proceed because of low affinity to papain [27, 28]. The resulting polypeptide with an Aib-containing periodic sequence transitioned to an α-helix secondary structure, driven by the helix-inducing Aib units. This method of using tripeptide ester derivatives effectively introduces the unnatural Aib residues into the polypeptide backbone and can be applied to other types of unnatural monomers [29].
Here, we focus on aromatic amino acids as unnatural structures to be inserted into the polypeptide backbone. Aromatic amino acid units can offer high thermal stability and high mechanical properties due to their rigid structures; hence, polyamides composed of aromatic monomers are widely used as high-performance engineering plastics. The addition of such aromatic units to the main chain of polypeptides can affect the polypeptide secondary structures due to the presence of rigid aromatic rings. Conventional polyamides consisting of aromatic monomers such as 4-aminobenzoic acid (Abz) possess a rigid, planar backbone that assembles into highly crystalline structures via π-π stacking and hydrogen bonding, which confers thermal stability and stiffness to practical materials. However, random or branched structures in aromatic polyamides favor amorphous structures over crystalline structures [30, 31]. Therefore, sequence control of aromatic monomer-containing polypeptides is essential for thermal and mechanical properties. There are few reports that introducing aromatic amino acids into peptide sequences can induce specific secondary structures [14, 32, 33]. To synthesize the polypeptides containing aromatic units, we designed novel tripeptide esters in which Abz is sandwiched between Gly and Ala units, namely, GlyAbzGly and AlaAbzAla esters. Chemoenzymatic polymerization of these tripeptide esters using papain successfully provided polypeptides that contain periodic Abz units. Structural characterization of the resulting Abz-containing polypeptides revealed that the periodic sequence is a key to maintaining specific secondary structures that exhibit high thermal properties.
Experimental procedure
Materials
Papain (EC No. 3.4.22.2) was purchased from Wako Pure Chemical Industries, Ltd. (Osaka, Japan) and used as received. The activity was ~0.5 U g−1, where one unit is defined as the amount of papain needed to hydrolyze 1 mmol of N-benzoyl-dl-arginine p-nitroanilide per minute at pH 7.5 and 25 °C. Amino acid derivatives and 1-ethyl-3-(3-dimethylaminopropyl)carbodiimide (EDC) HCl salt were purchased from Watanabe Chemical Industries, Ltd. (Hiroshima, Japan), and used as received. The other chemicals were purchased from Wako Pure Chemical Industries, Ltd., or Tokyo Chemical Industry Co., Ltd. (Tokyo, Japan), and used as received without purification unless otherwise noted.
Synthesis of AbzGly-OEt HCl salt
A solution of EDC HCl salt (9.59 g, 50 mmol) in chloroform was added to a solution of N-Boc-Abz-OH (11.86 g, 50 mmol), Gly-OEt HCl salt (6.98 g, 50 mmol), 1-hydroxybenzotriazole (HOBt) monohydrate (6.76 g, 50 mmol), and triethylamine (TEA) (7.0 mL, 50 mmol) in chloroform (50 mL) in a round-bottom flask equipped with an addition funnel at −10 °C under nitrogen. The resulting solution was stirred at 25 °C for 24 h. The mixture was washed with 5% NAHCO3 aq. and brine. Afterward, the organic layer was dried with MgSO4 and concentrated by a rotary evaporator to give Boc-AbzGly-OEt as a white solid. The crude product was then dissolved in dichloromethane (80 mL), and trifluoroacetic acid (TFA, 96 mL) was added to the solution. After stirring at 25 °C for 2 h, the solvent was removed by vacuum distillation. The solid was dissolved in dioxane/HCl (4 M, 10 mL). The solution was poured into diethyl ether, and the precipitate was filtered, washed with diethyl ether, and dried under vacuum to give AbzGly-OEt HCl salt as a white solid. The yield was 11.20 g (86.6%).
Synthesis of GlyAbzGly-OEt HCl salt
A solution of EDC HCl salt (3.89 g, 20.3 mmol) in chloroform (25 mL) was added dropwise to a solution of N-Boc-glycine (3.56 g, 20.3 mmol), AbzGly-OEt HCl salt (5.25 g, 20.3 mmol), HOBt monohydrate (3.11 g, 20.3 mmol), and TEA (2.83 mL, 20.3 mmol) in chloroform (40 mL) in a round-bottom flask equipped with an addition funnel at −10 °C under nitrogen. The resulting solution was stirred at 25 °C for 24 h. The mixture was washed with 5% NaHCO3 aq. and brine, and the organic layer was concentrated by a rotary evaporator. The crude product was dissolved in dichloromethane (80 mL), and TFA (96 mL) was added to the solution. After stirring at 25 °C for 2 h, the solvent was removed by vacuum distillation. The solid was dissolved in dioxane/HCl (4 M, 10 mL). The solution was poured into diethyl ether, and the precipitate was filtered, washed with diethyl ether, and dried under vacuum to give GlyAbzGly-OEt HCl salt as a white solid. The yield was 5.18 g (80.8%).
Synthesis of AlaAbzAla-OEt HCl salt
A solution of EDC HCl salt (1.17 g, 6.09 mmol) in chloroform (10 mL) was added dropwise to a solution of N-Boc-l-alanine (1.15 g, 6.09 mmol), AbzAla-OEt HCl salt (1.66 g, 6.09 mmol), HOBt monohydrate (0.933 g, 6.09 mmol), and TEA (0.85 mL, 6.09 mmol) in chloroform (10 mL) in a round-bottom flask equipped with an addition funnel at −10 °C under nitrogen. The resulting solution was stirred at 25 °C for 24 h. Then, the mixture was washed with 5% NaHCO3 aq. and brine, and the organic layer was concentrated by a rotary evaporator. The crude product Boc-AlaAbzAla-OEt was dissolved in dichloromethane (8 mL) under nitrogen. After TFA (8 mL) was added, the mixture was stirred at 25 °C for 24 h. After the solvent was removed by vacuum distillation, the resulting solid was dissolved in dioxane/HCl (4 M, 10 mL). The solution was poured into diethyl ether, and the precipitate was filtered, washed with diethyl ether, and dried under vacuum to give AlaAbzAla-OEt HCl salt as a white solid. The yield was 1.423 g.
General procedure of chemoenzymatic polymerization of Abz-containing tripeptide monomers
To a 10-mL glass tube equipped with a stir bar, GlyAbzGly-OEt HCl (0.112 g, 0.354 mmol) and 1 M tris(hydroxymethyl)aminomethane (TRIS) buffer (0.75 mL, pH 8.0) were added, and the mixture was stirred at 40 °C until all substrates were completely dissolved. Then, a solution of papain (0.071 g) in phosphate buffer (0.25 mL) was added to one portion. The final concentrations of GlyAlaGly-OEt and papain were 0.25 M and 50 mg mL−1, respectively. The mixture was stirred at 40 °C and 800 rpm for 2 h using an EYELA ChemiStation (Tokyo Rikakikai Co. Ltd., Tokyo, Japan). After cooling to room temperature, the precipitate was collected by centrifugation at 9000 rpm and 4 °C for 15 min. The crude product was washed twice with deionized water and lyophilized to afford a white powder. The yield was 0.068 g (82.0%). Homopolymerization of AlaAbzAla-OEt and copolymerization with Gly-OEt or Ala-OEt were also carried out by using the same procedure described above.
Wide-angle X-ray diffraction (WAXD) measurements on polypeptides
The synchrotron WAXD measurements of the polypeptide powdery samples were performed on the BL45XU beamline (SPring-8, Harima, Japan) using X-ray energy of 12.4 keV (wavelength: 0.1 nm). The obtained two-dimensional (2D) diffraction patterns were converted to one-dimensional (1D) profiles by azimuthal integration using Fit2D [34]. The temperature dependence of the WAXD profiles was examined by measuring WAXD at elevated temperatures. The powdery samples were placed in a heating cell on the beamline and sealed with polyimide (Kapton) films on both sides. The WAXD measurement was performed at 25, 50, 75, 100, 125, 150, 175, 200, and 225 °C. The 1D WAXD profiles were obtained by Fit2D after a background pattern using only polyimide films was subtracted.
Optimization of chemical structures of polypeptides
The structures of poly(GlyAbzGly) and poly(AlaAbzAla) were geometrically optimized at the quantum mechanics (QM) level of theory. The initial model structures of the polypeptides poly(GlyAbzGly) in hexamer form and poly(AlaAbzAla) in trimeric form, both containing an ethyl ester cap, were constructed using the graphical interface of Gaussian, Gaussview6. Geometry optimization calculations were performed via density functional theory (DFT) with the B3LYP 6-31G basis set [35] in Gaussian 16 [36] until reaching a stationary point in the potential energy surface.
Analytical measurements
The IR spectra of the bulk samples were recorded on an IRPrestige-21 Fourier transform IR spectrophotometer (Shimadzu Corporation, Kyoto, Japan) with a MIRacle A single-reflection attenuated total reflectance (ATR) unit using a Ge prism. The 1H and 13C NMR spectra were recorded on a Varian NMR System 500 (Varian Medical Systems, Palo Alto, CA, USA) at 25 °C and frequencies of 500 and 125 MHz, respectively. Chloroform-d or deuterated dimethylsulfoxide (DMSO-d6)/trifluoroacetic acid (TFA-d) (5/1 volume ratio) was used as the solvent, and tetramethylsilane (TMS) served as an internal standard for polypeptides. Matrix-assisted laser desorption/ionization time-of-flight (MALDI TOF) mass spectrometric analysis was conducted using an ultrafleXtreme MALDI-TOF spectrophotometer (Bruker Daltonics, Billerica, MA, USA) operating in reflection mode at an accelerating voltage of 15 kV. The sample was dissolved in water/acetonitrile (0.8 mg mL−1) containing 0.1% TFA mixed with a solution of α-cyano-4-hydroxycinnamic acid (CHCA) in water/acetonitrile (10 mg mL−1) and deposited on an MTP 384 ground steel BC target plate. Thermogravimetric analysis (TGA) was performed on the polypeptide samples using a TGA/DSC2 (Mettler Toledo, Columbus, OH, USA). The polypeptide sample (~5 mg) was weighed on an aluminum pan and heated with an empty reference cell at a heating rate of 20 °C min−1 from 30 to 500 °C under nitrogen. Differential scanning calorimetry (DSC) measurements were performed on the polypeptide samples using a DSC 8500 (PerkinElmer, Waltham, MA, USA). The polypeptide sample (~5 mg) was weighed on an aluminum pan and subjected to heating/cooling cycles at a heating and cooling rate of 10 °C min−1 in a range from 30 to 500 °C under nitrogen. Circular dichroism (CD) spectroscopic analysis was conducted using a Jasco J-820 CD spectropolarimeter (JASCO, Tokyo, Japan). The polypeptides were dissolved in 2,2,2-trifluoroethanol (TFE) containing 1% TFA (1 μM) at 70 °C, and measurements were conducted on the polypeptide solutions in a 1 mm cuvette at 20 °C.
Results and discussion
Chemoenzymatic synthesis of polypeptides containing aromatic residues
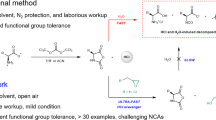
Unnatural amino acids are generally unfavorable substrates for chemoenzymatic polymerization using proteases due to their substrate specificity. Indeed, no polymer was obtained for the papain-catalyzed polymerization of the aromatic amino acid monomer 4-aminobenzoic acid ethyl ester (Abz-OEt), resulting in approximately complete recovery of the monomer, as observed in the case of other unnatural amino acids such as 2-aminoisobutyric acid and nylon esters (Scheme 1) [21, 27]. To overcome the poor specificity of the Abz unit to papain, we modified the Abz unit with Gly or Ala at both termini to obtain tripeptide ethyl esters via a solution-based condensation method. The resulting tripeptide esters, GlyAbzGly-OEt and AlaAbzAla-OEt, are expected to mitigate the mismatch between the Abz unit and papain because Gly and Ala show a high affinity to papain. We tried to polymerize these tripeptide esters by papain in an aqueous buffer to obtain polypeptides containing aromatic units (Scheme 1). The papain-catalyzed polymerization was carried out at 40 °C for 2 h with varying monomer concentrations and buffer solutions (salt concentration: 1 M, pH 8.0). As the polymerization proceeded, polypeptides were generated as a water-insoluble precipitate. The polymerization results are summarized in Table 1.
In the case of GlyAbzGly-OEt, no polymer was obtained in PBS because of poor solubility of GlyAbzGly-OEt in a phosphate buffer. The yield of the polypeptide poly(GlyAbzGly) depends on the buffer solution used for polymerization. When the polymerization was carried out using tris(hydroxymethyl)aminomethane (TRIS) buffer solution, the resulting yield increased up to 82.0%. The reason TRIS buffer gave the highest yield is probably due to the good solubility of the monomers but low solubility of the polypeptides, which resulted in efficient precipitation in TRIS buffer. The concentration of tripeptide monomers also affects the yield of polypeptides. The yield obtained by the polymerization in each buffer solution reached a maximum at a monomer concentration ranging from 0.1 to 0.25 M. At these concentrations, the concentration of papain also increased, which is assumed to accelerate the chemoenzymatic polymerization according to a previous study [37]. The best result for the synthesis of poly(GlyAbzGly) was achieved at a monomer concentration of 0.1 M using TRIS buffer. A similar tendency was found for the polymerization of AlaAbzAla-OEt using papain. The polymerization in TRIS buffer afforded a good yield, with poly(AlaAbzAla) obtained in an almost quantitative yield at a monomer concentration of 0.1 M. The chemical structures of the polypeptides were characterized by MALDI-TOF MS spectrometry and 1H NMR spectroscopy. The MALDI-TOF mass spectra of the obtained polypeptides showed a series of peaks with a peak interval of 233.08 or 261.11 m/z (Fig. S1), indicating that the polypeptides consist of repeating units of GlyAbzGly or AlaAbzAla, respectively. In addition to the peaks derived from poly(GlyAbzGly) and poly(AlaAbzAla) in the MALDI-TOF mass spectra, minor peaks corresponding to the m/z values of poly(GlyAbzGly) and poly(AlaAbzAla) with one or two amino acid defects were detected. These defective polypeptides were probably produced by transamidation side reactions during chemoenzymatic polymerization, as discussed in a previous study [37]. The number average molecular weight of the resulting Abz-containing polypeptides was ~1000–2000 regardless of the polymerization conditions and yield. In the 1H NMR spectra of the resulting polypeptides (Fig. 1), signals assignable to the aromatic protons of Abz units were clearly observed at 7.6–8.0 ppm, confirming the successful introduction of Abz units in the polypeptide backbone. Furthermore, the integral ratio of aromatic protons to Gly or Ala units was approximately equivalent to the theoretical values, namely, 33.3 mol% in molar ratios. This indicates that Abz units were introduced periodically into the polypeptides with a GlyAbzGly or AlaAbzAla repeating sequence, and the content of undesired defective products was small, although minor peaks related to transamidation products were detected in MALDI-TOF MS spectra.
The chemoenzymatic copolymerization of GlyAbzGly and AlaAbzAla ethyl esters with Gly-OEt and Ala-OEt, respectively, was also performed using papain under the optimized condition ([Monomer] = 0.25 M, 1 M TRIS buffer). The results of copolymerization are summarized in Table 2. The monomer feed ratio of Gly to GlyAbzGly or Ala to AlaAbzAla was varied from 20/80 to 80/20 in the copolymerization, and copolymers with different Abz contents in a random sequence were obtained. In the copolymerization of Gly and GlyAbzGly monomers, the Abz contents in the obtained polypeptides were slightly higher than those in the feed compositions. This is probably because the sequence randomness of Gly/GlyAbzGly copolymers increased the solubility of these copolymers in water due to the presence of hydrophilic Gly residues. Indeed, as the Gly content in the feeds increased, the yield of the polypeptide obtained as the water-insoluble part gradually decreased because the copolymer was assumed to show a lower ability to precipitate. We investigated the water-soluble part by MALDI-TOF MS, but only the monomer units with ester and hydrolyzed C-termini were observed. This means that the water-soluble fraction was easily subjected to hydrolysis by papain in the polymerization solution. On the other hand, the copolymerization of Ala and AlaAbzAla monomers afforded copolymers with Abz contents similar to those of the feed compositions. Based on the results, a wide range of Abz contents can be achieved in the polypeptide sequences.
The secondary structure of the polypeptides
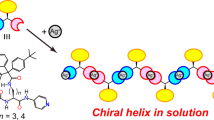
The secondary structures of the obtained polypeptides containing Abz units were characterized using IR spectrometry and WAXD analysis to investigate the effect of the introduction of the aromatic Abz units on the secondary structures. The IR spectra of all the polypeptides are shown in Fig. 2. The strong peak in the amide I region (1600–1700 cm−1) is derived from a stretching vibration mode of the amide carbonyl group in polypeptides and shows a significant shift associated with the secondary structures of polypeptides in which a coupled vibration occurs via inter- and intrachain hydrogen bonds [38, 39]. A sharp amide I peak was observed at 1645 and 1628 cm−1 for polyGly and polyAla and assigned as typical helix and β-sheet structures, respectively (Fig. 2, black lines) [38]. As the content of Abz units in the polypeptides increased, the amide I peak gradually split into multiple peaks. For poly(Gly-r-GlyAbzGly), the major peak split into two peaks at 1651 and 1631 cm−1, with a shoulder appearing in the range from 1700 to 1660 cm−1 that was assignable to random coil or turn structures. One split peak at 1651 cm−1 is still assignable to the helix conformation, whereas the peak at 1631 cm−1 is attributed to intermolecular hydrogen bonds in a more sheet-like structure. This tendency to adopt a sheet structure is probably due to the introduction of planar aromatic rings in the main chains. On the other hand, the amide I peak shifted slightly to a higher wavenumber for poly(Ala-r-AlaAbzAla). Both copolypeptides showed a new small peak at 1608 cm−1 in the amide I region derived from C=C stretching of aromatic rings. The amide II peak at 1550–1530 cm−1, mainly attributed to C–N stretching and N–H bending modes in amide bonds [40], showed a slight shift to a lower wavenumber. These results indicate that the bulky aromatic units in the polypeptide backbone hamper the hydrogen bonds coupled in helix and sheet structures to some extent. Therefore, the introduction of the Abz unit into polyGly or polyAla resulted in more random and complicated secondary structures. New peaks derived from the aromatic ring of Abz units also appeared at ~1500 and 850 cm−1 and were assignable to the stretching and bending vibration modes of the C–H bond in the para-substituted aromatic rings, respectively. These spectral differences clearly indicate that aromatic Abz units were introduced into the polypeptide backbone, and the secondary structure of the polypeptides was differentiated from the helical and β-sheet structures of polyGly and polyAla, respectively. A strong Cotton effect was observed in the circular dichroism (CD) spectrum of poly(AlaAbzAla) (Fig. S2), which indicates the existence of a specific secondary structure with a large chirality induced by l-Ala residues. On the other hand, poly(GlyAbzGly) showed no Cotton effect because of its achiral backbone.
IR spectra of Abz-containing polypeptides obtained by the copolymerization of a Gly and GlyAbzGly esters and b Ala and AlaAbzAla esters with different feed ratios. A broken line indicates the amide I peak of polyGly or polyAla assignable to α-helix or β-sheet structure, respectively. Arrows indicate the peaks derived from C=C stretching and C–H bending vibration modes of aromatic rings
WAXD measurements on the polypeptide powdery samples were carried out for further investigation of the secondary structures of Abz-containing polypeptides. The 1D WAXD profiles of poly(GlyAbzGly) and poly(AlaAbzAla) are shown in Fig. 3 with their backbone polypeptides, polyGly and polyAla, respectively. The PolyGly WAXD pattern was characteristic of the polyglycine II structure and showed a strong peak with a d-spacing of 4.15 Å corresponding to the length of hexagonal packing of helical polyGly chains with a threefold screw axis [41, 42]. On the other hand, the typical WAXD pattern of the antiparallel β-sheet structure was detected for polyAla [43,44,45]. Strong peaks with d-spacings of 5.26, 4.35, 4.13, and 3.68 Å were assigned to the (020), (210), (021), and (211) reflections, respectively. Poly(GlyAbzGly) showed a WAXD profile similar to that of polyGly, with broadening of the strong signal. This indicates that the introduction of bulky aromatic rings into the polyGly backbone did not alter the helical conformation but probably distorted the hexagonal packing of helices. In the case of poly(AlaAbzAla), the WAXD pattern was similar to the profile of an antiparallel β-sheet structure but showed slight peak shifts and peak broadening, indicating that poly(AlaAbzAla) adopts a sheet-like structure similar to the antiparallel β-sheet (β-pleated sheet) structure. The peak assignable to the (020) reflection for poly(AlaAbzAla) shifted to a higher q value (d = 5.04 Å) than that of polyAla, whereas other peaks assignable to the (210), (021), and (211) reflections shifted to lower q-values (d = 4.49, 4.19, and 3.77 Å, respectively). According to the crystal structure of polyAla, the (020) reflection represents the interlayer lattice between β-sheets. Therefore, the decrease in the d-spacing of the (020) reflection is attributed to a decrease in the distance between the sheets of poly(AlaAbzAla). This is assumed to be due to the π-stacking of aromatic rings, the planes of which are aligned parallel to the poly(AlaAbzAla) sheets. However, the introduction of bulky Abz units into polyAla probably expands or distorts the β-sheet structure of the polyAla backbone, resulting in an increase in the other d-spacings and peak broadening. Plausible structures of Abz-containing polypeptides optimized by DFT calculations revealed that both polypeptides adopted planar structures, which is most likely attributed to the introduction of aromatic rings. The optimized poly(GlyAbzGly) adopted a slightly twisted structure, whereas a sheet-like structure with a zig-zag conformation was obtained for poly(AlaAbzAla), as shown in Fig. S3.
The secondary structures of random copolymers poly(Gly-r-GlyAbzGly) and poly(Ala-r-AlaAbzAla) were also investigated by WAXD measurement (Figs. S4 and S5). The WAXD profiles gradually changed from the secondary structure of the polyGly or polyAla backbone to that of poly(GlyAbzGly) or poly(AlaAbzAla) by increasing the content of Abz units in the polypeptides. The peaks observed for poly(Ala-r-AlaAbzAla) were relatively broadened in comparison with those of polyAla and poly(AlaAbzAla), indicating that the random insertion of Abz units impedes the formation of β-sheet crystals.
Effect of aromatic units on thermal properties
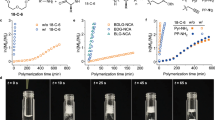
The effect of aromatic Abz units on the thermal properties of the polypeptides was investigated by thermogravimetric analysis (TGA) and differential scanning calorimetry (DSC). TGA curves and the thermal properties of (co)polypeptides containing Abz units are summarized in Fig. 4 and Table S1. All the polypeptides showed a weight loss (1.3–2.2% for polyGly and GlyAbzGly polypeptides and 2.6–4.9% for polyAla and AlaAbzAla polypeptides, as shown in Table S1) derived from water desorption below 100 °C. The thermal decomposition of polyGly occurred rapidly above 300 °C, and the 5% decomposition temperature (Td5) of polyGly was 302 °C (Fig. 4a). Poly(GlyAbzGly) showed Td5 at 301 °C, similar to that of polyGly, whereas poly(Gly-r-GlyAbzGly)s exhibited lower Td5 by ~10–20 °C. The Abz-containing polypeptides showed different decomposition behavior from that of polyGly: a gradual weight loss slower than that of polyGly was observed for Abz-containing polypeptides, resulting in a higher 10% decomposition temperature (Td10) (Table S1).
On the other hand, the same but the more emphasized trend was found for poly(AlaAbzAla) and the copolypeptides in comparison with polyAla (Fig. 4b). The randomness of the sequence significantly deteriorated the thermal stability of poly(AlaAbzAla). The Td5 of poly(Ala-r-AlaAbzAla) copolymers decreased substantially to 202–266 °C, and only poly(AlaAbzAla) showed high a Td5 (291 °C) comparable to that of polyAla (298 °C). The results of the thermal analysis indicate that periodic sequences containing Abz units adopt distinct secondary structures, as observed by the WAXD measurements, which resulted in high thermal stability. Otherwise, the random introduction of Abz units into polypeptides resulted in deterioration of the thermal properties because of the relatively random structures. The char yield at 500 °C for poly(AlaAbzAla) was higher than that of polyAla because of the high thermal stability of aromatic Abz units [46].
Among the copolypeptides containing Abz units, only poly(GlyAbzGly) and poly(AlaAbzAla) showed high Td5 values comparable to those of the backbone polypeptides polyGly and polyAla, respectively. These two polypeptides consist of a periodic sequence containing Abz units, which is assumed to form a distinct secondary structure compared to random copolymers, judging from the sharper peaks in the WAXD profiles for poly(GlyAbzGly) and poly(AlaAbzAla) than for poly(Gly-r-GlyAbzGly) and poly(Ala-r-AlaAbzAla) in Figs. S4 and S5. Therefore, the thermal analysis of poly(GlyAbzGly) and poly(AlaAbzAla) revealed that the copolymerization of Abz with Gly and Ala resulting in a distinct periodic sequence led to slightly higher decomposition temperatures than those of the backbone polypeptides, polyGly and polyAla, respectively. Copolypeptides generally show inferior thermal stability to homopolypeptides because of their sequence randomness. In a previous study, it was found that the copolypeptides of Gly and Ala showed lower thermal degradation temperatures than polyGly and polyAla homopolypeptides [37]. Rigid aromatic units in the polypeptide backbones probably contributed to the high thermal stability of poly(GlyAbzGly) and poly(AlaAbzAla) in addition to the effect of the secondary structures constructed by periodic sequences.
Then, the Abz-containing periodic peptides, namely, poly(GlyAbzGly) and poly(AlaAbzAla), were subjected to DSC measurements (Fig. 5). The DSC profile of poly(GlyAbzGly) showed a small peak at ~175 °C, which is probably due to the glass transition. Poly(AlaAbzAla) also showed a transition at 150 °C in the second heating scan. No melting point was observed for either polypeptide before degradation. Because polyGly and polyAla show no transition below the decomposition temperature, the aromatic rings in the polypeptide backbone induced glass transitions to the Abz-containing polypeptides. The transition was also confirmed by WAXD analysis during elevation of the temperature. The WAXD measurement was performed on the polypeptide samples every 25 °C up to 225 °C while heating the sample to investigate the transition of their secondary structures (Fig. 6). The WAXD pattern of poly(GlyAbzGly), where a strong peak at q = 15.2 nm−1 appears, was unchanged below 150 °C (Fig. 6a). Then, the peak was gradually split and sharpened above 150 °C, which almost corresponds to the transition temperature observed in the DSC measurement. We also investigated the full width at half maximum (FWHM) of the peak at 15.2 nm−1. Peak separation of the peak at 15.2 nm−1 provides two major peaks at 14.8 and 15.3 nm−1, and we evaluated the FWHM values of these peaks at each temperature. The results are summarized in Table S2. The FWHM of the peak at 15.3 nm−1 was ~0.20–0.22 nm−1 and remained constant up to 225 °C. In contrast, the FWHM of the peak at 14.8 nm−1 was 0.21 nm−1 up to 150 °C and decreased to 0.16 nm−1 above 150 °C, which caused the apparent peak splitting. This result suggests that a glass transition occurs at 175 °C for poly(GlyAbzGly). On the other hand, the peak shoulders at q = 13.2 and 15.2 nm−1 gradually decreased in the WAXD profiles of poly(AlaAbzAla) above 125 °C (Fig. 6b), and the WAXD profile similar to the β-sheet structure of polyAla was more emphasized at higher temperatures.
Conclusion
Polypeptides containing the aromatic amino acid Abz were successfully synthesized via papain-catalyzed polymerization of tripeptide ester monomers, GlyAbzGly-OEt and AlaAbzAla-OEt, under moderate conditions (aqueous buffer and 40 °C). The modification of the Abz unit with a natural amino acid, Gly or Ala, at both termini can mitigate the substrate mismatch in the catalytic pocket of papain to provide the Abz-containing polypeptides poly(GlyAbzGly) and poly(AlaAbzAla), whereas the unnatural Abz ester monomer is unreacted in the presence of papain. The resulting polypeptides contained periodic Abz units every three residues, whereas the Abz units were randomly introduced into the polypeptides synthesized via copolymerization of the tripeptide esters with Gly or Ala ethyl ester monomers. Structural analysis using IR and WAXD measurements revealed that the periodic polypeptide containing Abz units showed specific secondary structures similar to that of polyGly or polyAla, a backbone polypeptide. The thermal stability of poly(GlyAbzGly) and poly(AlaAbzAla) was comparable to those of polyGly and polyAla or slightly higher, but the random introduction of Abz units substantially deteriorates the thermal stability. Therefore, the periodic introduction of aromatic units is essential for achieving the excellent thermal properties of Abz-containing polypeptides, which are highly associated with their secondary structures. The method to synthesize the aromatic-containing polypeptides can be applied to develop novel eco-friendly materials with high thermal stability and high biodegradability due to their polypeptide backbone.
References
Wess TJ. Collagen Fibril Form and Function. In: Parry DAD, Squire JM, editors. Adv. Protein Chem. Amsterdam: Academic Press; 2005. 341–374.
Brodsky B, Persikov AV. Molecular Structure of the Collagen Triple Helix. In: Parry DAD, Squire JM, editors. Adv. Protein Chem. Amsterdam: Academic Press; 2005. 301–39.
Merrifield B. Solid phase synthesis. Science. 1986;232:341–7.
Suzuki Y, Shindo H. Binding sites and structure of peptides bound to SiO2 nanoparticles studied by solution NMR spectroscopy. Polym J. 2018;50:989–96.
Sawada T. Filamentous virus-based soft materials based on controlled assembly through liquid crystalline formation. Polym J. 2017;49:639.
Kricheldorf HR, Mang T. Stereospecificity of peptide synthesis by means of phosphorus derivatives: a model of peptide synthesis in molecular evolution. Int J Biol Macromol. 1983;5:258–66.
Higashi F, Sano K, Kakinoki H. High-molecular-weight poly(amino acid)s by the direct polycondensation reaction promoted by triphenyl phosphite and LiCl in the presence of polyvinylpyrrolidone. J Polym Sci Polym Chem 1980;18:1841–6.
Nooner DW, Oró J. Direct synthesis of polypeptides. J Mol Evol 1974;3:79–88.
Mizuno Y, Furuya H. Volume shrinkage of polypeptide hybrid xerogels induced by a helix-sense inversion. Polym J. 2019;51:337–44.
Harada A, Kataoka K. Polyion complex micelle formation from double-hydrophilic block copolymers composed of charged and non-charged segments in aqueous media. Polym J. 2017;50:95.
Habraken GJM, Heise A, Thornton PD. Block copolypeptides prepared by N-carboxyanhydride ring-opening polymerization. Macromol Rapid Commun. 2012;33:272–86.
Deming TJ. Synthetic polypeptides for biomedical applications. Prog Polym Sci. 2007;32:858–75.
Kricheldorf HR. Polypeptides and 100 years of chemistry of α-amino acid N-carboxyanhydrides. Angew Chem Int Ed. 2006;45:5752–84.
Okamura T, Seno S. Strategic construction of chiral helices: expanded poly(l-leucine) Containing p-phenylene moieties. Macromolecules. 2017;50:3500–9.
Koyama Y, Ihsan AB, Taira T, Imura T. Fluorinated polymer surfactants bearing an alternating peptide skeleton prepared by three-component polycondensation. RSC Adv. 2018;8:7509–13.
Koyama Y, Gudeangadi PG. One-pot synthesis of alternating peptides exploiting a new polymerization technique based on Ugi’s 4CC reaction. Chem Commun. 2017;53:3846–9.
Tsuchiya K, Numata K. Chemoenzymatic synthesis of polypeptides for use as functional and structural materials. Macromol Biosci. 2017;17:1700177.
Qin X, Xie W, Tian S, Cai J, Yuan H, Yu Z, et al. Enzyme-triggered hydrogelation via self-assembly of alternating peptides. Chem Commun. 2013;49:4839–41.
Qin X, Khuong AC, Yu Z, Du W, Decatur J, Gross RA. Simplifying alternating peptide synthesis by protease-catalyzed dipeptide oligomerization. Chem Commun. 2013;49:385–7.
Gudeangadi PG, Tsuchiya K, Sakai T, Numata K. Chemoenzymatic synthesis of polypeptides consisting of periodic di- and tri-peptide motifs similar to elastin. Polym Chem. 2018;9:2336–44.
Yazawa K, Gimenez-Dejoz J, Masunaga H, Hikima T, Numata K. Chemoenzymatic synthesis of a peptide containing nylon monomer units for thermally processable peptide material application. Polym Chem. 2017;8:4172–6.
Yazawa K, Numata K. Papain-catalyzed synthesis of polyglutamate containing a nylon monomer unit. Polymers. 2016;8:194.
Ageitos JM, Chuah J-A, Numata K. Chemo-enzymatic synthesis of linear and branched cationic peptides: evaluation as gene carriers. Macromol Biosci. 2015;15:990–1003.
Ageitos JM, Baker PJ, Sugahara M, Numata K. Proteinase K-catalyzed synthesis of linear and star oligo(L-phenylalanine) conjugates. Biomacromolecules. 2013;14:3635–42.
Tsuchiya K, Masunaga H, Numata K. Tensile reinforcement of silk films by the addition of telechelic-type polyalanine. Biomacromolecules. 2017;18:1002–9.
Tsuchiya K, Numata K. Papain-catalyzed chemoenzymatic synthesis of telechelic polypeptides using bis(leucine ethyl ester) initiator. Macromol Biosci. 2016;16:1001–8.
Tsuchiya K, Numata K. Chemoenzymatic synthesis of polypeptides containing the unnatural amino acid 2-aminoisobutyric acid. Chem Commun. 2017;53:7318–21.
Numata K. Poly(amino acid)s/polypeptides as potential functional and structural materials. Polym J. 2015;47:537–45.
Tsuchiya K, Numata K. Protease-Catalyzed Polymerization of Tripeptide Esters Containing Unnatural Amino Acids: α,α-Disubstituted and N-Alkylated Amino Acids. In, editors. Green Polymer Chemistry: New Products, Processes, and Applications: American Chemical Society; 2018. p. 95–105.
Yang G, Jikei M, Kakimoto M. Successful thermal self-polycondensation of AB2 monomer to form hyperbranched aromatic polyamide. Macromolecules. 1998;31:5964–6.
Ellis TS. Miscibility in blends of aliphatic polyamides and an aromatic polyamide, nylon 3Me6T. Polymer. 1988;29:2015–26.
Ishido Y, Kanbayashi N, Okamura T, Onitsuka K. Synthesis of nonnatural helical polypeptide via asymmetric polymerization and reductive cleavage of N–O bond. Macromolecules. 2017;50:5301–7.
Nowick JS, Lam KS, Khasanova TV, Kemnitzer WE, Maitra S, Mee HT, et al. An unnatural amino acid that induces β-sheet folding and interaction in peptides. J Am Chem Soc 2002;124:4972–3.
Numata K, Masunaga H, Hikima T, Sasaki S, Sekiyama K, Takata M. Use of extension-deformation-based crystallisation of silk fibres to differentiate their functions in nature. Soft Matter. 2015;11:6335–42.
Petersson GA, Bennett A, Tensfeldt TG, Al‐Laham MA, Shirley WA, Mantzaris J. A complete basis set model chemistry. I. The total energies of closed‐shell atoms and hydrides of the first‐row elements. J Chem Phys. 1988;89:2193–218.
Gaussian 16 Rev. A.03 (Wallingford, CT, 2016).
Ageitos JM, Yazawa K, Tateishi A, Tsuchiya K, Numata K. The benzyl ester group of amino acid monomers enhances substrate affinity and broadens the substrate specificity of the enzyme catalyst in chemoenzymatic copolymerization. Biomacromolecules. 2016;17:314–23.
Huang W, Krishnaji S, Tokareva OR, Kaplan D, Cebe P. Influence of water on protein transitions: morphology and secondary structure. Macromolecules. 2014;47:8107–14.
Rabotyagova OS, Cebe P, Kaplan DL. Role of polyalanine domains in β-sheet formation in spider silk block copolymers. Macromol Biosci. 2010;10:49–59.
Hu X, Kaplan D, Cebe P. Dynamic protein−water relationships during β-sheet formation. Macromolecules. 2008;41:3939–48.
Crick FHC, Rich A. Structure of polyglycine II. Nature. 1955;176:780.
Bamford CH, Brown L, Elliott A, Hanby WE, Trotter IF. β-Forms of fibrous proteins and synthetic polypeptides. Nature. 1953;171:1149.
Hamley IW, Dehsorkhi A, Castelletto V, Seitsonen J, Ruokolainen J, Iatrou H. Self-assembly of a model amphiphilic oligopeptide incorporating an arginine headgroup. Soft Matter. 2013;9:4794–801.
Riekel C, Vollrath F. Spider silk fibre extrusion: combined wide- and small-angle X-ray microdiffraction experiments. Int J Biol Macromol. 2001;29:203–10.
Arnott S, Dover SD, Elliott A. Structure of β-poly-l-alanine: refined atomic co-ordinates for an anti-parallel beta-pleated sheet. J Mol Biol. 1967;30:201–8.
Imai Y. Recent advances in synthesis of high-temperature aromatic polymers. React Funct Polym. 1996;30:3–15.
Acknowledgements
The authors acknowledge Dr. Takaaki Hikima for his assistance and discussion on the WAXD experiments at BL45XU SPring-8, Harima, Japan. This work was financially supported by the RIKEN Engineering Network Project, Impulsing Paradigm Change through the Disruptive Technologies Program (ImPACT) and JSPS KAKENHI (Grant Number JP17K18361).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tsuchiya, K., Kurokawa, N., Gimenez-Dejoz, J. et al. Periodic introduction of aromatic units in polypeptides via chemoenzymatic polymerization to yield specific secondary structures with high thermal stability. Polym J 51, 1287–1298 (2019). https://doi.org/10.1038/s41428-019-0242-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41428-019-0242-z
This article is cited by
-
Construction methodologies and sequence-oriented properties of sequence-controlled oligomers/polymers generated via radical polymerization
Polymer Journal (2021)
-
Aqueous spinning system with a citrate buffer for highly extensible silk fibers
Polymer Journal (2021)
-
How to define and study structural proteins as biopolymer materials
Polymer Journal (2020)
-
Self-assembly and hydrogel formation ability of Fmoc-dipeptides comprising α-methyl-L-phenylalanine
Polymer Journal (2020)