Abstract
Multidrug resistance represents a growing threat to human beings, and alternative antimicrobial regimens to conventional antibiotic paradigms are being extensively searched to fight against multidrug-resistant bacteria (MDRB). Although the antimicrobial potency of silver nanomaterials (AgNMs) has been previously elaborated, their efficacy against MDRB still remains to be strengthened. Here, our data revealed that small-sized silver nanoclusters (AgNCs) are superior to conventional silver nanoparticles (AgNPs) as robust antimicrobials against multidrug-resistant (MDR) Pseudomonas aeruginosa (P. aeruginosa). The core structure and surface ligands of AgNCs are crucial for the outstanding antibacterial activity of AgNCs. On the one hand, due to the presence of amphiphilic ligands, AgNCs are relatively prone to associate with the cell membrane and partake in endocytosis with targeted bacterial cells. Molecular dynamics simulations also corroborated this finding. On the other hand, the nanocluster structure of AgNCs led to strong peroxidase-like activity associated with massive production of reactive oxygen species (ROS), which contributes to their overall bactericidal potency. These outstanding features of AgNCs result in elevated bacterial killing efficacy by impairing the cell wall/membrane, promoting oxidative stress and attenuating pivotal cellular processes, e.g., ATP synthesis. Notably, AgNCs manifested great efficacy in treating P. aeruginosa-generated pneumonia in mice and increased the survival of infected animals, as well as exhibited excellent biocompatibility. Taken together, the results of this study pinpoint the great promise of AgNCs as new alternative therapeutics against MDR P. aeruginosa.
Similar content being viewed by others
Introduction
Multidrug-resistant bacteria (MDRB) are more pernicious than their precursory, drug-susceptible strains, as they can survive attack from most drugs relative to. Thus, it is clinically difficult to eradicate MDRB due to their resistance to a wide spectrum of antibiotics. According to the World Health Organization (WHO), the number of MDRB has been increasing quickly in recent years, incurring serious public health problems and bringing about substantial economic burden1,2,3. In this context, new drugs and innovative strategies are urgently needed to combat MDRB. Thus far, numerous new therapeutics have been explored to resist MDRB, e.g., nanomaterials. In fact, a wide range of nanomaterials, including metallic (e.g., silver4 and gold5 nanoparticles), nonmetallic (e.g., selenium nanoparticles6,7), metal oxide (e.g., titanium dioxide8 and iron oxide9), and carbonaceous nanomaterials (e.g., graphene10) have been demonstrated to possess multidrug-resistant (MDR) bactericidal effects. The following general mechanisms have been proposed for the ability of nanomaterials to overcome antimicrobial resistance. (i) Membrane damage: nanomaterials tend to rupture the cell structure and damage the cell membrane11; (ii) reactive oxygen species (ROS) production: many nanomaterials induce the production of ROS12, which damage DNA and cause metabolic disorders13; (iii) enzyme-mimicking performance: the peroxidase-like activity of nanomaterials such as palladium nanocrystals and iron oxide enhances their bactericidal properties14; (iv) ion dissolution: released ions, such as silver ions, directly attack biomolecules, e.g., thiol-rich proteins; and (v) eradication of the biofilm: since biofilm formation is an important reason for drug resistance development, some nanomaterials, such as metal oxide nanoparticles, could eradicate biofilms due to their strong redox reaction capabilities that perturb cell metabolism15. Among all types of nanomaterials, silver (Ag) nanomaterials represent a family of the most effective antimicrobial nanomedicines with exceptionally wide applications16,17,18,19. To date, the most prevalent form of Ag nanomaterials (AgNMs) is Ag nanoparticles (AgNPs)4,20. Although a wealth of past studies have evaluated the potency of AgNPs against MDRB, their antimicrobial efficacy remains to be strengthened, and drug resistance towards AgNPs could likely form after repeated treatments4,21,22,23,24,25,26. Thus, it is of great importance to augment Ag’s antimicrobial potency through various physicochemical improvements that would enable more active nano–bio interactions and elicit greater stress on pathogens. In this context, continuous efforts are being invested to look for more potent AgNMs.
Strikingly, metal nanoclusters are becoming more attractive because they exactly fill in the gap between small molecules (<1 nm) and conventional nanoparticles (>3 nm)27,28,29,30. Recently, AgNCs have attracted extensive e attention for various applications, such as in biomedicine31. In addition to the essential difference in size between AgNCs and AgNPs, the core is differently formed because AgNCs are formed with the precise control of their composition at the atomic level, which could not be achieved in conventional AgNPs32,33. To this end, AgNCs bear physicochemical properties superior to those of conventional AgNPs, including substantial surface reactivities towards exchange reactions, enabling versatile solubility in different media34. Therefore, advanced nanocluster chemistry would empower diverse manipulations of the type and composition of surface ligands on AgNCs, giving rise to distinct AgNCs with precise size, composition and surface properties33,35,36. In addition, diverse surface ligands on AgNCs warrant their desirable stability in various solutions and in biological settings34,37,38. Additionally, owing to their atomically precise molecular structure, AgNCs offer an ideal platform to understand the interaction between Ag and biological molecules34,36,38. In this respect, these outstanding characteristics, including amphiphilic modalities and active nano–bio interactions, might also be utilized to combat pathogens. Nonetheless, the antimicrobial potency of AgNCs has scarcely been determined to date.
Although the antimicrobial efficacy of AgNMs has been found to correlate to many physicochemical properties, e.g., size, core/shell material, and the nature of the surface ligands (which dictate solubility, charge and stability)4,24,25, most previous studies examined individual properties, and no conclusive understandings have been obtained regarding the preferential combination of these properties. However, no quantitative informatic tools are currently available to offer appreciable predictive power for the design of desirable module incorporation. To explore the parameters that exclusively dictate the antimicrobial efficacy of AgNMs, we sought to optimize the contributions of size, surface hydrophilicity and hydrophobicity, and core fraction to determine the most suitable integration of multiple physicochemical properties. We determined that small-sized amphiphilic silver clusters, namely, AgNCs, are superior to conventional AgNPs, showing active nano–bio interactions and outstanding activity in combating MDR P. aeruginosa both in vitro and in vivo.
Materials and methods
Chemicals and reagents
The chemicals and reagents used in the current study include silver nitrate (Sinopharm, Shanghai, China), triethylamine (Macklin, Shanghai, China), mercaptosuccinic acid (Alfa Aesar, Shanghai, Beijing), 1-admantanethiol (Sigma, St. Louis, USA), tetraphenylphosphonium bromide (J&K Scientific, Beijing, China), N,N-dimethylformamide (J&K Scientific, Beijing, China), sodium borohydride (Sinopharm, Shanghai, China), dichloromethane (Sinopharm, Shanghai, China), absolute methanol (Sinopharm, Shanghai, China), ammonia (Sinopharm, Shanghai, China), ethanol absolute (Sinopharm, Shanghai, China), sodium chloride (Sinopharm, Shanghai, China), triethylamine (Macklin, Shanghai, China), tryptone (Oxoid, Basingstoke, UK), yeast extract (Oxoid, Basingstoke, UK), agar power (Solarbio, Beijing, China), dialysis membranes (Solarbio, Beijing, China), DCFH-DA (Sigma, St. Louis, USA), Hoechst 33342/PI (Solarbio, Beijing, China), D2000 DNA marker (GenStar, Beijing, China), 2×Taq PCR StarMix with loading dye (GenStar, Beijing, China), and Agarose G-10 (IOWEST, Chicago, USA).
Workflow for literature searching and data mining
As shown in Supplementary Fig. S1, we collected research papers and extracted data according to the following steps:
-
a.
Literature search. All articles were searched by the keywords “silver nanoparticles”, “nanosilver”, or “nAg” and “antibacterial”, “antimicrobial” or “bactericidal” using the Web of Science (http://apps.webofknowledge.com) and PubMed (www.ncbi.nlm.nih.gov). A total of >1700 publications were retrieved.
-
b.
Inclusion and exclusion criteria. Inclusion criteria: (1) AgNMs were not doped; (2) the particle size was clearly measured and <100 nm; (3) the antibacterial effect evaluation was through conventional determination, such as minimum inhibitory concentration (MIC) value and antibacterial efficiency determination; (4) AgNMs was synthesized through chemical synthesis methods with definite surface modification; and (5) the bacterial strains and sources were clearly described. Exclusion criteria: (1) the development of nonsingle homogeneous systems, such as textiles, pharmaceutical devices, gels, alloys, plastics, and clothing, and (2) the use of green synthetic methods resulting in unclear surface modifications. After combining the inclusion and exclusion criteria, a total of 266 data sets from 36 research papers were collected.
-
c.
Data extraction: detailed information, including size, surface ligand, surface charge, gram feature, bacterial strain, exposure time, drug resistance and MIC values, was extracted from 36 research papers.
Calculation of the importance for various properties via random forest regression
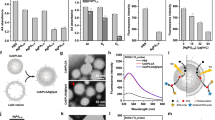
To quantify the importance of complex and difficult-to-normalize attributes (e.g., surface ligands, strains, drug resistance, etc.) affecting the MIC values, we employed an advanced machine learning approach: the random forest model (RF model)39,40. Here, the language package “randomForest” of “R” was used, and the number of repetitions was set at 200. The strategy of increasing the mean squared error (MSE) was carried out to directly measure the impact of each factor on the prediction accuracy of the RF model. The principle was the rearrangement of the order of these factors versus the reduction of the model accuracy. This method indicated that the accuracy of the model was not affected by less important factors, such as the gram feature, whereas the accuracy of the model was greatly affected by important indicators, such as size and surface ligands (Fig. 1a).
a Importance of seven factors in dictating the MIC values (MSE: means squared error). b Classes of size-determined effects on the survival of different bacterial strains as reflected by MIC values. Red represents a size <15 nm, green indicates 15 to 30 nm, blue denotes 30 to 50 nm, and black represents 50 to 100 nm. c A schematic depicting the morphology and surface properties of AgNPs (#1 and #2) and AgNCs, where –COOH represents mercaptosuccinic acid and -Adm denotes 1-admantanethiol. d Representative images of AgNPs and AgNCs dissolved in water and CH2Cl2, reflective of their hydrophilic and amphiphilic features. e Absorption profiles of AgNPs #1, AgNPs #2, and AgNCs.
The classification map (Fig. 1b) was generated with “R”, where Gram-positive and Gram-negative features served as the definition of two groups, the readout of bactericidal efficacy was the MIC value, and the color denoted the differential size range of AgNMs. Additionally, to recognize the correlation between particle size and MIC valued, a general linear regression model (Fig. 1b) was used to fit the scatter points.
Synthesis of AgNCs and AgNPs (#1 and #2) and characterization
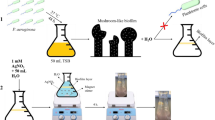
AgNCs were synthesized in two steps, as previously described33. First, the core of the AgNCs was synthesized through a biphasic reaction. Second, the amphiphilic shell of the AgNCs was subjected to ligand exchange with mercaptosuccinic acid (MSA) to obtain amphiphilic AgNCs. A detailed description is provided as follows. Step 1: AgNO3 (60 mg) was dissolved in a mixed solvent of dichloromethane and methanol in an ice bath. Thereafter, 1-adamantanethiol (S-Adm) (15 mg) and tetraphenylphosphonium bromide (12 mg) were added and vigorously stirred. After stirring for 20 min, 1 mL of an aqueous sodium borohydride (NaBH4) solution (45 mg/mL) and 50 μL of triethylamine were quickly added under vigorous stirring. Afterwards, the reaction was aged for 12 h at 4 °C. The aqueous phase was then gently removed, and the mixture in the organic phase was washed several times with water. Dark green rhombic crystals were crystallized from methylene chloride (CH2Cl2) in a week. Step 2: MSA (0.8 mg) was added into a 1 mL ethanol suspension of organophilic AgNCs (5 mg). After stirring for 3 h at room temperature, concentrated ammonia (6 μL) was added to precipitate AgNCs. Solid products were separated by centrifugation, washed three times with ethanol, and finally dissolved in deionized water, followed by characterization.
AgNPs #1 was made based on the following process. Briefly, AgNO3 (60 mg) was dissolved in 4 mL of N,N-dimethylformamide in an ice bath. Then, MSA (13.50 mg) and tetraphenylphosphonium bromide (12 mg) were added and vigorously stirred. After stirring for 20 min, 1 mL of an aqueous NaBH4 solution (45 mg/mL) was added into each reaction after 50 μL of triethylamine was quickly added under stirring conditions. The reaction was thereafter aged for 6 h at room temperature. AgNPs with a uniform size were separated by centrifugation, washed with water, and eventually dissolved in deionized (DI) water. For the synthesis of AgNCs #2, MSA (13.50 mg) was replaced by MSA (6.75 mg) and S-Adm (7.50 mg); otherwise, the other steps were the same as the synthesis for AgNPs #1.
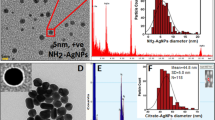
AgNCs and AgNPs were characterized by transmission electron microscopy (TEM) (JEM-2100F, Japan). Fourier transform infrared (FTIR) spectroscopy was performed on an iS50 FTIR spectrometer (Thermo Fisher, USA). The ultraviolet (UV)−visible (Vis) spectra were acquired on a UV–Vis spectrophotometer DU-800 (Backman, USA). The zeta potential in water and culture medium was measured with a dynamic light scattering (DLS) instrument (Malvern, UK). The elemental composition of TEM-analyzed specimens was assessed through EDS (OXFORD, London, UK).
Bacterial strains and growth determination
MDRB strains of P. aeruginosa, A. baumannii, and E. coli were obtained and maintained with approval from the inhouse ethics committee at the China-Japan Friendship Hospital41,42, and their corresponding regular drug-susceptible strains were commercial strains acquired from the ATCC. Bacterial cell growth and killing efficacy were determined through growth curves reflective of absorption at OD600 and colony forming unit (CFU) counting according to standard protocols, as described previously41,43. In the current work, we repeatedly thawed the stored bacteria from the parental lines to avoid secondary resistance.
Enzyme-like activity of AgNMs
For the TMB (3,3’,5,5’-tetramethylbenzidine) assay, an aqueous solution of AgNPs or AgNCs was diluted to a final concentration of 51.20 μg/mL with DI water, and then 100 μL of H2O2 (10 M) and 20 μL of TMB (100 mM dissolved in DMSO) were added. The volume of the final reaction system was adjusted to 1 mL with DI water. After 5 min, the absorbance curve was monitored by a UV–Vis spectrophotometer (UNICO, UV-2800). DI water was employed as the untreated control in TMB experiments.
For electron spin resonance (ESR) spectrum acquisition, a 50 μL system (5 μL of 5,5-dimethyl-1-pyrroline n-oxide (DMPO) (1 M), 5 μL of H2O2 (10 M), 5 μL of AgNPs or AgNCs (484 μg/mL), and 35 μL of DI water) were subjected to electron paramagnetic resonance (EPR, BRUKER, A300–10/12). DI water was employed as the untreated control in both the TMB and ESR experiments.
Intercellular ROS and extracellular MDA assays
Intracellular ROS production was measured in bacterial cells with a 2’, 7’-dichlorofluorescin diacetate (DCFH-DA) probe following an established protocol, as previously described44. Malondialdehyde (MDA), a product of lipid peroxidation, was assayed by the thiobarbituric acid (TBA) test, as previously described45, according to the manufacturer’s instructions (Solarbio, Beijing, China).
Cellular ATP level test
The level of cellular ATP in MDR P. aeruginosa was determined using a bioluminescence assay kit (Beyotime, Shanghai, China). Briefly, post-treatment, bacterial pellets were lysed with lysis buffer in Lysing Matrix B tubes (MPBio, CA, USA) and vortexed three times for 30 s with incubation on ice at intervals, followed by centrifugation at 12,000 × g for 5 min at 4 °C. Finally, 20 μL of supernatant dispersed in DI water was mixed with 80 μL of luciferase reagent, which catalyzed bioluminescence production from ATP and luciferin. The emitted bioluminescence linearly correlated to the ATP concentration, as measured using a microplate luminometer (Varioskan Flash, Thermo, USA).
TEM examination and propidium (PI) staining of bacterial cells
After exposure of AgNPs and AgNCs, P. aeruginosa cells were separated and then fixed by glutaraldehyde at 4 °C overnight, followed by TEM examination (JEM2010, Japan)46. Similarly, bacterial cells were stained with PI at 50 μg/mL for 20 min on ice and then visualized and examined on a TCS SP5 microscope (Leica, Germany).
Determination of cellular Ag uptake
The Ag mass in treated bacteria was analyzed with the inductively coupled plasma mass spectrometry (ICP-MS) method, as described previously47,48,49. In brief, an equal number of bacterial cells (determined by OD600 value) were collected and digested with a strong oxidation-acid solution (a 3:2 mixture of nitric acid and hydrogen peroxide) at room temperature. For the proportion of membrane-bound silver, the membrane was first extracted, followed by acid digestion. Eventually, the Ag content was measured on an ICP-MS instrument (NexION 300X, USA).
Molecular dynamics (MD) simulation
Coarse-grained molecular dynamics (CGMD) simulations were performed using the modified MARTINI force field, in which an average of four heavy atoms were represented by a single bead to increase the timescale and the space scale of the simulation system50. In our simulations, each bacterial outer membrane comprises two asymmetric leaflets. The outer leaflet consisted of 53 lipopolysaccharide (LPS) molecules and 13 dipalmitoyl-phosphatidylethanolamine (DPPE) molecules. The inner leaflet was composed of 175 DPPE molecules. GNOMM software was used to construct the initial structure of the bacterial outer membrane51. The topology of the coarse-grained (CG) LPS was obtained from the model, as proposed by Ma et al.52. The water model based on the Morse potential was used to replace the original MARTINI water to prevent unphysical water freezing on the membrane surface53,54. The CG structure of the AgNCs was derived from its atomic structure while reserving the position of the binding sites of the adamantine33. Here, each adamantine was coarse-grained into a trihedron structure with a mapping ratio of ~3:1, and each core Ag atom was mapped to one CG bead.
During the simulations of the interaction between AgNMs and the bacterial membrane, the system was equilibrated at zero tension using the NPT ensemble. The temperature boundary conditions were set using the Nose-Hoover algorithm, and the pressure boundary conditions were set using the Parrinello-Rahman pressure coupling method. For temperature coupling, the water, DPPE, and LPS molecules were independently coupled with relaxation times (\(\tau _T\)) of 0.20, 1.00, and 1.00 ps, respectively. The reference temperature was set at the body temperature of 310 K. The time step of all simulations was 10 fs. The nonbonding pair list was updated every 5 steps with a neighboring search cutoff of 1.60 nm. All simulations were performed using the GROMACS 4.5.5 package55.
Animal experimentation
Balb/c mice (7-week-old) were purchased from the Beijing Vital River Laboratory Animal Technology Co., Ltd. (Beijing, China). All animal experiments were approved by the Animal Ethics Committee of the Research Center for Eco-Environmental Sciences, Chinese Academy of Sciences. For the lung infection model, prior to lung infection, MDR P. aeruginosa 5835 cells were suspended in phosphate-buffered saline (PBS) to reach OD600 = 0.2. After an internasal drip for 30 min, AgNPs and AgNCs administration was performed on anesthetized animals (by 5% sodium pentobarbital) by instilling materials onto each nostril (in a total of 25 μL). Forty-eight hours later, mice were anesthetized, and bronchoalveolar tissue was flushed with prewarmed PBS to collect bronchoalveolar lavages. Afterwards, 100 μL of bronchoalveolar lavages from each mouse was subjected to culture on plates for 24 h. Blood was collected for serum total bilirubin measurement using a commercial kit (BioAssay Systems, California, USA). Serum IL-6 was determined by a commercial kit following the instructions from the manufacturer (NEOBIOSCIENCE, Beijing, China), and other organs were also collected for histological examination after H&E staining with respect to the standard protocol.
Identification of P. aeruginosa by endpoint polymer chain reaction (PCR)
Briefly, to extract bacterial DNA, a single colony from the plates in different groups was individually suspended in 20 μL of lysis buffer (0.25% sodium dodecyl sulfate and 0.05 M NaOH). After heating for 20 min at 95 °C in a dry bath (Thermo Fisher, USA), 180 μL of DI water was added. Then, isolated DNA was amplified in a 25 μL reaction volume, as described previously56, with the following conditions: each reaction contained 2 μL of DNA templates, 12.50 μL of StartMix, and 2 μL of mixed primer (PA-SS-F56: GGGGGATCTTCGGACCTCA; PA-SS-R56: TCCTTAGAGTGCCCACCCG) with additional DI water to reach a total volume of 25 μL. The PCR procedures are shown in Supplementary Table S1. After 25 cycles, ten microliter PCR products were subjected to 1% agarose gel electrophoresis (120 V/20 min), as described57.
Ag ion dissolution assessment
To determine Ag ion dissolution, AgNPs and AgNCs at 16 μg/mL in H2O were subjected to rotation at 180 rpm, and 1 mL of H2O was aliquoted at different time points and dialyzed with 8 kD dialysis bags in 100 mL of DI water for 2 h. Thereafter, dialyzed Ag ions into water were quantified through ICP-MS.
Statistical analysis
Experimental data are represented as the mean ± SD. The significant differences were analyzed with Student’s t-test using SPSS software. Statistical significance was determined at P < 0.05.
Results and discussion
Definition of desirable parameters of AgNMs with superior antimicrobial capability
First, to define the potency of various physicochemical properties and experimental parameters affecting AgNM’s antimicrobial efficacy, quantitative calculation was carried out through meta-analysis by collecting 266 sets of data from 36 publications with rigorous inclusion/exclusion criteria. Hereby, the random forest regression model was used to quantify the contribution of seven indicators to dictate the MIC values, including three physicochemical properties (particle size, surface ligand and surface charge) and four experimental setups (gram feature, bacterial strain, exposure time and drug resistance). As shown in Fig. 1a, the physicochemical properties of AgNMs, i.e., the size and surface, bore much greater influence on their antibacterial efficacy than that of the experimental parameters, such as bacterial features and exposure time. Of note, the particle size was the most impactful attribute, with an increase in the mean squared error (MSE%) of ~20%, and further analysis manifested an inverse correlation between the particle size and antibacterial potency of AgNMs; that is, the smaller the size was, the greater the antimicrobial effect, with the greatest performance observed for sizes <15 nm (Fig. 1b). Since there is no sufficient literature to exhaustively analyze the effects of core fraction and hydrophilicity and hydrophobicity, we further integrated experimental data to look for optimal physicochemical property combinations. To this end, we deliberately synthesized relatively small-sized AgNCs and AgNPs with distinct hydrophilicity and hydrophobicity characteristics. As shown in Fig. 1c and Supplementary Fig. S2, AgNCs were synthesized from two types of AgNMs with different ligands, S-Adm and MSA, following our previous protocol with slight modifications2. TEM images displayed an average size of AgNCs of 2.80 nm (Supplementary Fig. S2c). Two types of AgNPs (#1 and #2) were fabricated with comparable diameters of ~3.80 nm and 3.20 nm, respectively (Supplementary Fig. S2a, b). For hydrophilicity/hydrophobicity control, the surface of AgNPs #1 harbored only MSA ligands, whereas the AgNPs #2 surface contained both S-Adm and MSA. As a result, AgNPs #1 could be dissolved only in water, while AgNPs #2 could be dissolved in water and partially dissolved in methylene chloride (CH2Cl2) (Fig. 1d). By contrast, AgNCs displayed good solubility in both water and CH2Cl2, suggesting that AgNCs are superior in ligand exchange reactions for hydrophilicity/hydrophobicity relative to AgNPs. Owing to the bulkiness of S-Adm, nanoclusters bear a low-surface thiolate coverage and, thus, manifest extremely high-surface reactivities towards exchange reactions with diverse ligands, including halides, phenylacetylene and thiols33. Moreover, these ligands, except halides, tend to replace [S-Adm] instead of halides due to the particular surface binding structure of the nanoclusters that are endowed by the bulky thiolates. In other words, some Ag–S bonds among these S atoms in the middle part of the cluster are longer than 2.70 Å, rendering the thiolates easy targets for attack and eventual replacement by these ligands33, as shown in Fig. 1c. Therefore, AgNCs could be made amphiphilic through metathesis with water-soluble ligands, yielding new functionalities for various bioapplications. As the water-soluble ligand MSA served as an exchange ligand, amphiphilic AgNCs were successfully obtained. The FTIR results showed that AgNCs had obvious absorption peaks at ~1704 and 1535 cm−1 (denoted by green arrows) (Supplementary Fig. S2d), corresponding to the C=O vibrations of MSA and N–H bending of ammonium acetate, respectively, as previously described58. In addition, AgNCs also contained S-Adm ligands, as indicated by the absorption peaks at 2885 and 2841 cm−1 (indicated by red arrows) (Supplementary Fig. S2d). Moreover, AgNCs exhibited a broad absorption peak at ~460 nm, which was mainly attributed to the core structure59, whereas AgNPs #1 and #2 displayed a significant absorption peak at ~410 nm (Fig. 1e). It should be noted that AgNPs #1 and #2 and AgNCs were all negatively charged in DI water (Supplementary Fig. S2e).
Antibacterial activity screening of AgNCs against MDR P. aeruginosa
P. aeruginosa is a leading pathogen of hospital-acquired pneumonia, often causing cystic fibrosis (CF)60,61, and MDRB strains cause even severe outcomes, such as increased mortality, prolonged hospital stays and even the necessity for surgery62. We therefore selected the P. aeruginosa MDRB strain (an isolate from a patent with serious infection, inhouse No. 5835) for detailed in vitro and in vivo antibacterial assessment. First, we screened the antibiotics towards the isolated P. aeruginosa strain in comparison to the corresponding regular strain. As shown in Supplementary Fig. S3, AgNCs suppressed both regular and MDR P. aeruginosa strains, and the MIC values on the susceptible and resistant P. aeruginosa strains indicated no selectivity of AgNCs towards the regular or MDRB strain, suggestive of wide-caliber targeting. Importantly, AgNCs revealed <2-fold greater efficacy than that of AgNPs (#1) in suppressing P. aeruginosa growth (Supplementary Fig. S3). Moreover, as shown in Supplementary Fig. S4, the screening data unveiled that AgNCs started to restrain MDR P. aeruginosa growth starting at very low concentrations, i.e., 1.60 μg/mL. A maximum inhibition (~90%) was observed at 6.40 μg/mL, in contrast to the 81% exhibited by AgNPs #1 and 62% by AgNPs #2 at the same concentration, as reflected by the measurement of the optical density at 600 nm (OD600) (P < 0.001). Subsequent growth curves demonstrated remarkable suppression of bacterial cell growth by AgNCs over time in contrast to mild inhibition by AgNPs #1 and #2 (Fig. 2a, P < 0.001). Moreover, the plate culture assessment corroborated the outstanding killing efficacy of AgNCs of ~72.3% and 99.80% at 3.20 and 6.40 μg/mL, in comparison to 59.50% and 95.50% for AgNPs #1 and 68.40% and 81.60% for AgNPs #2, respectively (Fig. 2b, c, P < 0.001). Additionally, we examined the sensitization effect of AgNPs and AgNCs in combination with antibiotics. As shown in Supplementary Fig. S5, AgNPs (#1) and AgNCs, especially the latter, greatly increased the sensitization of MDR P. aeruginosa to imipenem (IMP) and tobramycin (TOB). In this context, we revealed that AgNCs also acted to sensitize MDR P. aeruginosa to antibiotics. Together, these results unveiled that AgNCs bore pronounced bactericidal activity against MDR P. aeruginosa, and this bactericidal potency was superior to that of AgNPs.
a Cell growth curves were recorded over time starting from an OD600 value at 0.1, where P. aeruginosa MDR cells were exposed to AgNPs and AgNCs at 6.4 μg/mL. b The broth microdilution method42 was used to evaluate the antibacterial effect. CFU counting of P. aeruginosa MDR strain 5835 treated with AgNPs #1, AgNPs #2 and AgNCs at 3.2 and 6.4 μg/mL after culture on plates for 24 h, and quantified data are shown in c (n = 6). Pound sign (#) indicates P < 0.001.
Incredible capability of AgNCs to provoke free radicals
To interpret the enhanced cell death induced by AgNCs in comparison to that induced by AgNPs, we attempted to understand the plausible molecular basis underlying this difference. First, we determined the peroxidase-like activity of these AgNMs. The AgNCs manifested even greater peroxidase-like activity than AgNPs #1 and #2, as reflected by their reinforced capability to catalyze the conversion of the reporter molecule TMB (Fig. 3a, b). To further corroborate the remarkable peroxidase-like activity of AgNCs, the H2O2/Ag/DMPO spin trap system was used to detect the hydroxyl radical signals. As shown in Supplementary Fig. S6, the DMPO/•OH adduct signal intensity was enhanced in response to these nanomaterials (AgNPs #1 and AgNCs), and AgNCs promoted more •OH radical production by catalyzing the decomposition of H2O2, nearly 2-fold more than AgNPs (#1) catalysis. Altogether, these results essentially demonstrated that AgNCs showed more active peroxidase-like activity than AgNPs. Although other enzyme-mimicking activities have also been studied for diverse nanomaterials63,64, our finding on the peroxidase-like activity of AgNCs underpinned their antibacterial effects.
a and b Determination of peroxidase-like properties of AgNCs and AgNPs. The color change and the maximum absorbance values at 370 and 652 nm for the TMB reactions are shown in a and b, respectively. c Relative ROS production in P. aeruginosa cells upon exposure to AgNPs and AgNCs at 6.4 μg/mL over time (n = 3). The untreated control was set as 1.0. d The amount of total ATP production by P. aeruginosa cells after exposure to AgNPs and AgNCs at 6.4 μg/mL for 6 h (n = 3). e Relative ATP levels of P. aeruginosa cells upon AgNPs and AgNCs at 6.4 μg/mL for 6 h (n = 3) determined by normalizing to according OD600 values, and the untreated control was set as 1.0. An asterisk (*) denotes P < 0.05, and a pound sign (#) indicates P < 0.001.
Motivated by the greater peroxidase activity of AgNCs in cell-free conditions, we endeavored to probe the free-radical generation promoted by AgNCs in cells, which may (at least in part) account for cellular injuries. As shown in Fig. 3c, while AgNPs #1 and #2 only triggered mild induction of ROS over time relative to that in untreated cells, AgNCs dramatically promoted ROS production at 6 h, with an almost threefold increase in ROS compared to those in the untreated control (P < 0.001). Similarly, the total amount of ATP declined by 16.90%, 39.50% and 72.80% in P. aeruginosa cells responding to AgNPs #1 and #2 and AgNCs after 6 h of exposure at 6.4 μg/mL, respectively, compared to that in the untreated control (Fig. 3d, P < 0.001). After normalization to the cell numbers, as reflected by the OD600 values, the relative cellular ATP production was also reduced by 51.10% in P. aeruginosa cells in response to AgNCs relative to untreated cells (Fig. 3e, P < 0.001), suggesting a great attenuation of electron transfer and respiratory chain function in living bacterial cells. In contrast, no significant decline in relative ATP production was revealed in AgNPs-treated cells (Fig. 3e). These findings defined a myriad of free radicals induced by AgNCs upon association with bacterial cells and unveiled significant impairments of various cellular machineries and processes through oxidative stress by AgNCs. Furthermore, the above results revealed a mild difference between AgNPs #1 and AgNPs #2. Therefore, only AgNPs #1 (termed AgNPs in the following sections) was compared to AgNCs for better mechanistic understanding in the following investigations.
AgNCs tend to associate with bacterial cells and cause wall/membrane damage
Afterwards, we inspected the molecular mechanisms underlying the robust antibacterial activity of AgNCs. Since AgNCs showed excellent amphiphilic modality due to the copresence of both hydrophilic (–COO–) and organophilic (S-Adm) groups on their surface, AgNCs should associate with bacterial cells better than AgNPs. To test this hypothesis, TEM analysis was conducted to shed light on the interface between AgNCs and bacterial cells. As shown in Fig. 4a, after exposing MDR P. aeruginosa 5835 to AgNCs for 6 h, a large proportion (>30.0%) of bacterial cells appeared in screw-like morphologies with a large amount of surrounding debris (indicative of previously dead cells) in comparison to the intact morphology of untreated cells, and this proportion was much lower in AgNPs-treated cells (~10%) (Fig. 4a and Supplementary Fig. S7, P < 0.001), indicating severe damage of P. aeruginosa bacterial cells upon introducing AgNCs. Enlarged images of the interactions between AgNCs and bacterial cells showed massive AgNCs localization in the form of agglomeration together with cell debris (shown by orange arrowheads) and inside cells (denoted by red arrowheads) (Fig. 4a). In contrast, scarce AgNPs adsorption could be visualized on the cellular wall/membrane or inside the cell, and most AgNPs self-aggregated outside the bacterial cells, as indicated by blue arrowheads (Fig. 4a). Notably, energy-dispersive X-ray spectroscopy (EDS) confirmed the nature of AgNMs in association with bacterial cells (Supplementary Fig. S8). These observations, thus, confirmed the active binding and internalization of AgNCs towards P. aeruginosa bacterial cells.
a Representative TEM images showing the interaction of AgNCs and AgNPs with P. aeruginosa cells after 6 h of exposure. Blue arrowheads indicate agglomerated particles outside cells, red arrowheads denote AgNCs inside cells, and orange arrows point at cell debris. The enlarged images show the detailed interplay between AgNCs, AgNPs, and bacteria. b Evaluation of cell wall/membrane damage using PI staining (in red) in MDR P. aeruginosa cells upon AgNPs and AgNCs at 6.4 μg/mL for 6 h. Images were visualized through confocal microscopy (×600, original magnification). c Quantitative data of PI-positive cells (n = 3). d Relative extracellular MDA levels in culture media from P. aeruginosa MDR cells treated with AgNPs and AgNCs at 6.4 μg/mL for 6 h (n = 3). The level in the untreated control was normalized to 1.0. A pound sign (#) denotes P < 0.05 for AgNCs compared to AgNPs. e and f Cellular uptake of AgNPs and AgNCs, as reflected by Ag mass, in P. aeruginosa cells after treatment for 6 h (n = 3). The e panel indicates the total silver mass, and the f panel denotes the silver in association with the plasma membrane. g Calculation of the ratio between the mass on the membrane and the total amount for AgNCs and AgNPs, respectively (n = 3).
To explore the nano–bio interaction between AgNPs or AgNCs and bacterial cells, we further investigated cellular wall/membrane injuries, as reflected by propidium (PI) staining and MDA leakage determination. As shown in Fig. 4b, c, >43% of PI-positive cells were visualized upon AgNCs treatment at 6.40 μg/mL for 6 h in comparison to the 1.5% observed in the untreated control, and the proportion of PI-positive cells was >2 times greater than that in cells treated with AgNPs (P < 0.001), which was in agreement with the above TEM results (Fig. 4a and Supplementary Fig. S7). Analogously, membrane damage led to elevated release of MDA into the culture medium, especially in AgNCs-treated cells (Fig. 4d, P < 0.001). Therefore, these results showed active binding and uptake of AgNCs by P. aeruginosa cells, coupled to severe membrane damage, giving rise to a reinforced likelihood of interaction between AgNCs and biomolecules within bacterial cells.
To further prove that severe membrane damage resulted from increased Ag binding, which was indicated by TEM (Fig. 4a), we employed ICP-MS to determine the Ag uptake of AgNPs and AgNCs of both total and membrane-binding Ag. As shown in Fig. 4e, f, both the total Ag uptake and the membrane-binding Ag increased by ~2-fold in AgNCs-treated cells relative to that in AgNPs-treated cells (P < 0.001). Moreover, Fig. 4g shows a greater proportion of membrane-bound Ag of the total Ag in AgNCs-treated cells than that in AgNPs-treated cells. Previous studies demonstrated that Ag ions released from particles also contributed to the overall toxicity in diverse organisms47,65,66. Therefore, we measured the percentage of ion dissolution from AgNCs and AgNPs. Our data showed limited release of Ag ions from AgNCs and AgNPs over time, and only ~1.50% of Ag ions were identified coming out of AgNCs and AgNPs after 24 h (Supplementary Fig. S9a). Nevertheless, no toxicity was found in P. aeruginosa cells at the concentration with a maximum dissolution rate (Supplementary Fig. S9b), ruling out the implication of Ag ions eliciting toxicity to P. aeruginosa. Therefore, these findings further supported our finding that enhanced uptake and binding, at least in part, accounted for severe membrane damage of bacterial cells upon AgNCs.
MD simulations reveal greater intrusion of AgNCs than AgNPs into the bacterial bilayer membrane
To substantiate the above findings, we further used coarse-grained MD (CGMD) simulations to specifically study the difference in the process of AgNPs and AgNCs penetrating the bacterial bilayer membrane. Considering the asymmetry of the bacterial outer membrane (the outer layer mainly consists of LPS molecules, and the inner layer is composed of only DPPE molecules), we compared the interaction energies of the AgNMs with those of DPPE and LPS molecules by adding an external force to introduce AgNPs or AgNCs to enter the interior of the membrane (Fig. 5a). The interaction energy diagram in Fig. 5b shows that the van der Waals (VDW) interaction energy between the AgNMs and the LPS dominated the translocation of the AgNMs. Owing to the hydrophilicity of AgNPs, a weaker VDW interaction with the hydrophobic tails of DPPE and a much stronger VDW interaction with the polar heads of LPS were revealed compared to those of the amphiphilic AgNCs. A larger negative value of the total interaction energy indicated a stronger binding between AgNMs and molecules. Therefore, the AgNPs had to overcome more energy to penetrate the membrane. It should be noted that although the AgNPs could be more strongly repulsed by the negatively charged heads of the LPS than the AgNCs, the Coulomb interaction was relatively unimportant relative to the VDW interaction. Therefore, the amphiphilic property of AgNCs with two ligands could obviously reduce the interaction energy with the LPS of the outer membrane, therefore, intruding the outer membrane more readily.
a Initial interaction structure of the AgNPs, AgNCs, and the bacterial membrane. The head groups of the LPS and DPPE are shown as green sticks; the tails of the LPS and DPPE are shown as blue sticks. The carboxyl groups and adamantanes are shown as red and cyan sticks. The core silver and sulfur atoms are shown as yellow and orange beads. b Interaction energy between AgNPs/AgNCs and the LPS/DPPE molecules. The negative value represents the attraction interaction, and the positive value indicates the repulsion interaction. Coul and VDW represent Coulomb interactions and van der Waals interactions, respectively.
Antibacterial activity of AgNCs against MDR P. aeruginosa in mice with pulmonary infection
Next, we assessed the performance of AgNCs to combat MDR P. aeruginosa in a pulmonary infection mouse model, as depicted in Fig. 6a, where AgNCs or AgNPs were intranasally administered into mice 30 min after P. aeruginosa 5835 infection. First, we examined the survival of mice postinfection. In the untreated group, 4 out of 8 mice died within 48 h, in contrast to 3 out of 8 in the AgNPs-treated group and 1 out of 7 in the AgNCs-treated group (Fig. 6b), suggesting reinforced tolerance of mice against P. aeruginosa intrusion upon treatment, particularly that with AgNCs. The propagation of P. aeruginosa in the lungs was further determined by cultivating bronchoalveolar lavages from mice 2 days post administration. As shown in Fig. 6c, CFU counting showed a substantial reduction in P. aeruginosa cells in the lungs of the AgNCs-treated mice, with an average of 1 CFU/plate, in contrast to 53 CFU/plate in the AgNPs-treated group and 175 CFU/plate in the untreated group (P < 0.001), indicating the outstanding killing efficacy of AgNCs against P. aeruginosa in the lung. Through polymerase chain reaction (PCR) analysis of the evolutionally conserved sequence in P. aeruginosa56, our data corroborated the identity of P. aeruginosa from these clones (Supplementary Fig. S10).
a A schematic delineating the experimental procedure of AgNPsAgNPs and AgNCs administration in mice with lung infection of P. aeruginosa MDR strain 5835. b The survival rate of mice with or without treatment for 48 h. c CFU assessment of bronchoalveolar lavages from mice after 48 h of treatment with AgNPs and AgNCs at 0.8 mg/kg body weight. The average CFU data are shown in the right panel (n = 8). d H&E staining of lung sections from mice after treatment with AgNPs and AgNCs for 7 days. Black arrowheads indicate collapsed alveoli, green arrowheads denote inflammatory cells, blue arrowheads show compressed interstitial space, and yellow arrowheads represent hemolysis areas. The inserts show the enlarged view of the lung tissue (×600, original magnification). e Total bilirubin in sera from the above treated mice (n = 8). Pound sign (#) indicates P < 0.001, compared to untreated mice.
To further recognize the protective role of AgNCs in the lungs from P. aeruginosa infection, we examined lung histology. As shown in Fig. 6d, relative to that of the blank control mice, histological alterations featured very severe pneumonia, as characterized by extensive infiltration of inflammatory cells (indicated by green arrowheads), collapsed alveoli (indicated by black arrowheads) and compressed interstitial space (denoted by blue arrowheads). In stark contrast, AgNCs administration significantly ameliorated the pneumonic phenotypes, as these morphological injuries nearly disappeared in AgNCs-treated mice after treatment for 48 h and 7 days (Supplementary Fig. S11 and the bottom panel in Fig. 6d), confirming the remarkable killing efficacy of AgNCs against MDR P. aeruginosa 5835 in the lung. Moreover, marked hemolysis was observed in the interstitial space of untreated mice (denoted by yellow arrowheads). However, nearly no hemolysis was found in the lungs of AgNCs-treated mice (Fig. 6d). In support of these observations, the total serum bilirubin concentration was elevated ~4 times in untreated mice compared to that in the mock control (Fig. 6e, P < 0.001). In comparison, the bilirubin level dropped to baseline in AgNCs-treated mice (Fig. 6e, P < 0.001), indicative of the disappearance of hemolysis. Moreover, AgNCs tremendously abolished systemic inflammation, as evidenced by a > 50.0% reduction in the serum IL-6 content in mice after AgNCs treatment for 48 h (P < 0.05), in contrast to the small change in mice upon AgNPs treatment compared to the untreated control (Supplementary Fig. S12). Furthermore, AgNPs could also mitigate these pneumonic phenotypes but to a much lesser extent than AgNCs (Fig. 6d, e), consistent with the in vitro observations (Fig. 2). Consistently, the complete blood count (CBC) data suggested that systemic inflammation was relieved in infected mice upon treatment, especially upon AgNCs, as evidenced by the reduction in white blood cell counts (data not shown). Additionally, to probe the antibacterial activity of AgNCs against other MDRB, we also worked on clinical isolates that were deliberately purified from patients with serious infections. Additionally, four paired regular and corresponding MDRB strains, including Acinetobacter baumannii (A. baumannii) and Escherichia coli (E. coli), were subjected to treatment with AgNPs and AgNCs. As shown in Supplementary Fig. S13, AgNCs displayed a remarkable capability to repress the growth of all these strains at 3.20 and 6.40 μg/mL, showing much greater potency than that of AgNPs (P < 0.05). Altogether, these data substantiated the outstanding antibacterial activity of AgNCs against various MDRB.
Considering the biosafety and biocompatibility of AgNPs and AgNCs, we thoroughly assessed the toxicity of AgNPs and AgNCs in vivo and in vitro. As shown in Supplementary Fig. S14, the animal experimental results indicated that neither AgNPs nor AgNCs caused significant toxicity to diverse tissues, including heart, liver, spleen, lung, kidney and brain tissues, as characterized by histological examination based on H&E staining. Moreover, CBC was determined after treatment with AgNMs. As shown in Supplementary Table S2, no significant change in CBC indexes was found in mice upon treatment with AgNMs, confirming the excellent biocompatibility of these AgNMs. In addition, we also determined the cytotoxicity of AgNPs and AgNCs at various concentrations in different cell lines. As shown in Supplementary Fig. S15, the results indicated that AgNCs and AgNPs did not exhibit significant toxicity even at higher concentrations towards A549 (pulmonary epithelial cells), HL-7702 (normal human hepatocytes), and C2C12 (mouse myoblast cells) cells. With the combined results from both in vitro and in vivo experiments, we deduced that the AgNPs and AgNCs were biocompatible at the tested concentrations.
Conclusions
To summarize, our experimental data together with a meta-analysis revealed that small AgNCs were superior to conventional AgNPs against MDR P. aeruginosa. Specifically, our findings unearthed the great importance of the size, core structure and surface ligands in dictating the antibacterial activity of AgNCs. Our experimental data together with MD simulation findings unveiled that AgNCs with amphiphilic ligands showed strong nano–bio interactions towards bacterial cells and thereby underwent active endocytosis by the targeted cells. Moreover, AgNCs harbored robust peroxidase-like activity and the strong capability to promote ROS generation. To this end, AgNCs manifested great bacteria-killing efficacy in vitro against MDR P. aeruginosa and a therapeutic outcome in treating P. aeruginosa-induced pneumonia in mice. AgNCs also bore excellent cyto-/biocompatibility. This study opens a new avenue for AgNCs as promising therapeutics to combat MDRB.
References
Willyard, C. The drug-resistant bacteria that pose the greatest health threats. Nature 543, 15 (2017).
Magiorakos, A.-P. et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Infect. 18, 268–281 (2012).
Potron, A., Poirel, L. & Nordmann, P. Emerging broad-spectrum resistance in Pseudomonas aeruginosa and Acinetobacter baumannii: mechanisms and epidemiology. Int. J. Antimicrob. Ag. 45, 568–585 (2015).
Le Ouay, B. & Stellacci, F. Antibacterial activity of silver nanoparticles: a surface science insight. Nano Today 10, 339–354 (2015).
Cui, Y. et al. The molecular mechanism of action of bactericidal gold nanoparticles on Escherichia coli. Biomaterials 33, 2327–2333 (2012).
Lin, A. et al. Bacteria-responsive biomimetic selenium nanosystem for multidrug-resistant bacterial infection detection and inhibition. ACS Nano 13, 13965–13984 (2019).
Huang, N., Chen, X., Zhu, X., Xu, M. & Liu, J. Ruthenium complexes/polypeptide self-assembled nanoparticles for identification of bacterial infection and targeted antibacterial research. Biomaterials 141, 296–313 (2017).
Jesline, A., John, N. P., Narayanan, P., Vani, C. & Murugan, S. Antimicrobial activity of zinc and titanium dioxide nanoparticles against biofilm-producing methicillin-resistant Staphylococcus aureus. Appl. Nanosci. 5, 157–162 (2015).
Masadeh, M. M. et al. Cerium oxide and iron oxide nanoparticles abolish the antibacterial activity of ciprofloxacin against gram positive and gram negative biofilm bacteria. Cytotechnology 67, 427–435 (2015).
Zheng, H. et al. Engineered graphene oxide nanocomposite capable of preventing the evolution of antimicrobial resistance. ACS Nano 13, 11488–11499 (2019).
Chen, J. et al. Graphene oxide exhibits broad-spectrum antimicrobial activity against bacterial phytopathogens and fungal conidia by intertwining and membrane perturbation. Nanoscale 6, 1879–1889 (2014).
Vranic, S. et al. Live imaging of label-free graphene oxide reveals critical factors causing oxidative-stress-mediated cellular responses. ACS Nano 12, 1373–1389 (2017).
Das, B. et al. Green synthesized silver nanoparticles destroy multidrug resistant bacteria via reactive oxygen species mediated membrane damage. Arab. J. Chem. 10, 862–876 (2017).
Cai, S. et al. Porous Pt/Ag nanoparticles with excellent multifunctional enzyme mimic activities and antibacterial effects. Nano Res. 10, 2056–2069 (2017).
Wang, L.-S., Gupta, A. & Rotello, V. M. Nanomaterials for the treatment of bacterial biofilms. ACS Infect. Dis. 2, 3–4 (2016).
Li, X. et al. Functional gold nanoparticles as potent antimicrobial agents against multi-drug-resistant bacteria. ACS Nano 8, 10682–10686 (2014).
Courtney, C. M. et al. Photoexcited quantum dots for killing multidrug-resistant bacteria. Nat. Mater. 15, 529–534 (2016).
Singh, R., Smitha, M. S. & Singh, S. P. The role of nanotechnology in combating multi-drug resistant bacteria. J. Nanosci. Nanotechnol. 14, 4745–4756 (2014).
O’Connell, K. M. G. et al. Combating multidrug-resistant bacteria: current strategies for the discovery of novel antibacterials. Angew. Chem. Int. Ed. 52, 10706–10733 (2013).
Rai, M. K., Deshmukh, S. D., Ingle, A. P. & Gade, A. K. Silver nanoparticles: the powerful nanoweapon against multidrug-resistant bacteria. J. Appl. Microbiol. 112, 841–852 (2012).
Franci, G. et al. Silver nanoparticles as potential antibacterial agents. Molecules 20, 8856–8874 (2015).
Agnihotri, S., Mukherji, S. & Mukherji, S. Size-controlled silver nanoparticles synthesized over the range 5-100 nm using the same protocol and their antibacterial efficacy. RSC Adv. 4, 3974–3983 (2014).
Sotiriou, G. A. & Pratsinis, S. E. Antibacterial activity of nanosilver ions and particles. Environ. Sci. Technol. 44, 5649–5654 (2010).
Chernousova, S. & Epple, M. Silver as antibacterial agent: ion, nanoparticle, and metal. Angew. Chem. Int. Ed. 52, 1636–1653 (2013).
Panáček, A. et al. Silver colloid nanoparticles: synthesis, characterization, and their antibacterial activity. J. Phys. Chem. B 110, 16248–16253 (2006).
Chaloupka, K., Malam, Y. & Seifalian, A. M. Nanosilver as a new generation of nanoproduct in biomedical applications. Trends Biotechnol. 28, 580–588 (2010).
Valden, M., Lai, X. & Goodman, D. W. Onset of catalytic activity of gold clusters on titania with the appearance of nonmetallic properties. Science 281, 1647–1650 (1998).
Suber, L. et al. Nanocluster superstructures or nanoparticles? The self-consuming scaffold decides. Nanoscale 10, 7472–7483 (2018).
Yau, S. H., Varnavski, O. & Goodson, T. An ultrafast look at Au nanoclusters. Acc. Chem. Res 46, 1506–1516 (2013).
Turner, M. et al. Selective oxidation with dioxygen by gold nanoparticle catalysts derived from 55-atom clusters. Nature 454, 981–983 (2008).
Tian, X., Kong, X.-J., Zhu, Z.-M., Chen, T.-T. & Chu, X. A new label-free and turn-on strategy for endonuclease detection using a DNA–silver nanocluster probe. Talanta 131, 116–120 (2015).
Yao, C. H. et al. Adding two active silver atoms on Au-25 nanoparticle. Nano Lett. 15, 1281–1287 (2015).
Ren, L. T. et al. Bulky surface ligands promote surface reactivities of [Ag141X12(S-Adm)(40)](3+) (X=Cl, Br, I) nanoclusters: models for multiple-twinned nanoparticles. J. Am. Chem. Soc. 139, 13288–13291 (2017).
Sun, Z. et al. Silver nanoclusters with specific ion recognition modulated by ligand passivation toward fluorimetric and colorimetric copper analysis and biological imaging. Sci. Rep. 6, 20553 (2016).
Cathcart, N. & Kitaev, V. Silver nanoclusters: single-stage scaleable synthesis of monodisperse species and their chirooptical properties. J. Phys. Chem. C. 114, 16010–16017 (2010).
Diez, I. & Ras, R. H. A. Fluorescent silver nanoclusters. Nanoscale 3, 1963–1970 (2011).
Dhanya, S., Saumya, V. & Rao, T. P. Synthesis of silver nanoclusters, characterization and application to trace level sensing of nitrate in aqueous media. Electrochim. Acta 102, 299–305 (2013).
Feng, L. P. et al. Silver nanoclusters with enhanced fluorescence and specific ion recognition capability triggered by alcohol solvents: a highly selective fluorimetric strategy for detecting iodide ions in urine. Chem. Commun. 53, 9466–9469 (2017).
Oh, E. et al. Meta-analysis of cellular toxicity for cadmium-containing quantum dots. Nat. Nanotechnol. 11, 479 (2016).
Breiman, L. Random forests. Mach. Learn. 45, 5–32 (2001).
Yao, B. et al. Clinical and molecular characteristics of multi-clone carbapenem-resistant hypervirulent (hypermucoviscous) Klebsiella pneumoniae isolates in a tertiary hospital in Beijing, China. Int. J. Infect. Dis. 37, 107–112 (2015).
Wiegand, I., Hilpert, K. & Hancock, R. E. W. Agar and broth dilution methods to determine the minimal inhibitory concentration (MIC) of antimicrobial substances. Nat. Protoc. 3, 163 (2008).
Zhang, Y. W. et al. Emergence of a hypervirulent carbapenem-resistant Klebsiella pneumoniae isolate from clinical infections in China. J. Infect. 71, 553–560 (2015).
Zheng, K. Y., Setyawati, M. I., Lim, T. P., Leong, D. T. & Xie, J. P. Antimicrobial cluster bombs: silver nanoclusters packed with daptomycin. ACS Nano 10, 7934–7942 (2016).
Gutteridge, J. M. C. & Halliwell, B. The measurement and mechanism of lipid-peroxidation in biological-systems. Trends Biochem. Sci. 15, 129–135 (1990).
Zhang, J., Wang, S. H., Gao, M., Li, R. B. & Liu, S. J. Multihierarchically profiling the biological effects of various metal-based nanoparticles in macrophages under low exposure doses. ACS Sustain. Chem. Eng. 6, 10374–10384 (2018).
Wang, Z., Xia, T. & Liu, S. J. Mechanisms of nanosilver-induced toxicological effects: more attention should be paid to its sublethal effects. Nanoscale 7, 7470–7481 (2015).
Qian, Y. et al. Silver nanoparticle-induced hemoglobin decrease involves alteration of histone 3 methylation status. Biomaterials 70, 12–22 (2015).
Chen, Y. et al. Nanosilver incurs an adaptive shunt of energy metabolism mode to glycolysis in tumor and nontumor cells. ACS Nano 8, 5813–5825 (2014).
Marrink, S. J., Risselada, H. J., Yefimov, S., Tieleman, D. P. & de Vries, A. H. The MARTINI force field: coarse grained model for biomolecular simulations. J. Phys. Chem. B 111, 7812–7824 (2007).
Baltoumas, F. A., Hamodrakas, S. J. & Iconomidou, V. A. The gram-negative outer membrane modeler: automated building of lipopolysaccharide-rich bacterial outer membranes in four force fields. J. Comput. Chem. 40, 1727–1734 (2019).
Ma, H., Irudayanathan, F. J., Jiang, W. & Nangia, S. Simulating Gram-negative bacterial outer membrane: a coarse grain model. J. Phys. Chem. B 119, 14668–14682 (2015).
Chiu, S. W., Scott, H. L. & Jakobsson, E. A coarse-grained model based on morse potential for water and n-alkanes. J. Chem. Theory Comput. 6, 851–863 (2010).
Bai, X., Xu, L., Tang, J. Y., Zuo, Y. Y. & Hu, G. Adsorption of phospholipids at the air-water surface. Biophys. J. 117, 1224–1233 (2019).
Hess, B., Kutzner, C., van der Spoel, D. & Lindahl, E. GROMACS 4: algorithms for highly efficient, load-balanced, and scalable molecular simulation. J. Chem. Theory Comput. 4, 435–447 (2008).
Spilker, T., Coenye, T., Vandamme, P. & LiPuma, J. J. PCR-based assay for differentiation of Pseudomonas aeruginosa from other Pseudomonas species recovered from cystic fibrosis patients. J. Clin. Microbiol. 42, 2074–2079 (2004).
Wang, Y. Y., Wang, F. B., Wang, H. L. & Song, M. Y. Graphene oxide enhances the specificity of the polymerase chain reaction by modifying primer-template matching. Sci. Rep. 7, ARTN 16510 (2017).
Kumar Trivedi, M. Fourier transform infrared and ultraviolet-visible spectroscopic characterization of ammonium acetate and ammonium chloride: an impact of biofield treatment. Mod. Chem. Appl. 03, https://doi.org/10.4172/2329-6798.1000163 (2015).
Takele, H., Greve, H., Pochstein, C., Zaporojtchenko, V. & Faupel, F. Plasmonic properties of Ag nanoclusters in various polymer matrices. Nanotechnology 17, 3499–3505 (2006).
Gellatly, S. L. & Hancock, R. E. W. Pseudomonas aeruginosa: new insights into pathogenesis and host defenses. Pathog. Dis. 67, 159–173 (2013).
Fujitani, S., Sun, H. Y., Yu, V. L. & Weingarten, J. A. Pneumonia due to Pseudomonas aeruginosa part I: epidemiology, clinical diagnosis, and source. Chest 139, 909–919 (2011).
Aloush, V., Navon-Venezia, S., Seigman-Igra, Y., Cabili, S. & Carmeli, Y. Multidrug-resistant Pseudomonas aeruginosa: risk factors and clinical impact. Antimicrob. Agents Chemother. 50, 43–48 (2006).
Gao, M. et al. Two‐dimensional Tin selenide (SnSe) nanosheets capable of mimicking key dehydrogenases in cellular metabolism. Angew. Chem. 132, 3647–3652 (2020).
Chong, Y. et al. Palladium concave nanocrystals with high-index facets accelerate ascorbate oxidation in cancer treatment. Nat. Commun. 9, 1–10 (2018).
Xiu, Z. M., Zhang, Q. B., Puppala, H. L., Colvin, V. L. & Alvarez, P. J. J. Negligible particle-specific antibacterial activity of silver nanoparticles. Nano Lett. 12, 4271–4275 (2012).
Kedziora, A. et al. Similarities and differences between silver ions and silver in nanoforms as antibacterial agents. Int. J. Mol. Sci. 19, ARTN 444 (2018).
Acknowledgements
This work was supported under grants from the National Natural Science Foundation of China (grant numbers: 21637004, 21920102007, 21922611 and 21707161), the Beijing Natural Science Foundation (grant number: 8191002) and the international collaboration key grant from the Chinese Academy of Sciences (grant number: 121311KYSB20190010).
Author information
Authors and Affiliations
Contributions
N.F.Z., G.Q.Z., and S.J.L. conceived and designed the study. Y.J.C., L.T.R., and L.X.S. carried out the experiments and analyzed the data. X.B. and G.Q.H. performed the MD simulations. L.T.R., G.Q.Z., L.X.S., and B.C. contributed reagents and materials. L.X.S. and B.C. provided the bacterial strains. All authors revised the manuscript and approved the submission.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, Y., Ren, L., Sun, L. et al. Amphiphilic silver nanoclusters show active nano–bio interaction with compelling antibacterial activity against multidrug-resistant bacteria. NPG Asia Mater 12, 56 (2020). https://doi.org/10.1038/s41427-020-00239-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41427-020-00239-y
This article is cited by
-
Size and charge effects of metal nanoclusters on antibacterial mechanisms
Journal of Nanobiotechnology (2023)
-
Antimicrobial potential and osteoblastic cell growth on electrochemically modified titanium surfaces with nanotubes and selenium or silver incorporation
Scientific Reports (2022)