Abstract
HBV infection initiates hepatitis B and promotes liver cirrhosis and hepatocellular carcinoma. IFN-α is commonly used in hepatitis B therapy, but how it inhibits HBV is not fully understood. We screened 285 human interferon-stimulated genes (ISGs) for anti-HBV activity using a cell-based assay, which revealed several anti-HBV ISGs. Among these ISGs, SAMD4A was the strongest suppressor of HBV replication. We found the binding site of SAMD4A in HBV RNA, which was a previously unidentified Smaug recognition region (SRE) sequence conserved in HBV variants. SAMD4A binds to the SRE site in viral RNA to trigger its degradation. The SAM domain in SAMD4A is critical for RNA binding and the C-terminal domain of SAMD4A is required for SAMD4A anti-HBV function. Human SAMD4B is a homolog of human SAMD4A but is not an ISG, and the murine genome encodes SAMD4. All these SAMD4 proteins suppressed HBV replication when overexpressed in vitro and in vivo. We also showed that knocking out the Samd4 gene in hepatocytes led to a higher level of HBV replication in mice and AAV-delivered SAMD4A expression reduced the virus titer in HBV-producing transgenic mice. In addition, a database analysis revealed a negative correlation between the levels of SAMD4A/B and HBV in patients. Our data suggest that SAMD4A is an important anti-HBV ISG for use in IFN therapy of hepatitis B and that the levels of SAMD4A/B expression are related to HBV sensitivity in humans.
Similar content being viewed by others
Introduction
Human hepatitis B virus (HBV) infection, the major cause of hepatitis B disease, remains a global public health problem.1 HBV is a hepatotropic DNA virus with a 3.2 kb partially double-stranded genome. After the HBV receptor sodium taurocholate cotransporting polypeptide (NTCP)-mediated viral particle enters hepatocytes,2,3 the viral capsid is translocated to the nucleus. Then, the viral genomic relaxes and circular DNA (rcDNA) is released into the nucleus and converted into covalently closed circular DNA (cccDNA).4 The cccDNA functions as a minichromosome and four viral RNA species are transcribed: a 3.5 kb mRNA (pregenomic RNA), 2.4 kb mRNA (preS1 HBs RNA), 2.1 kb mRNA (preS2/S HBs RNA), and 0.7 kb mRNA (HBX RNA), which are initiated by different promoters but share the same poly-A tail. The viral proteins HBV e antigen (HBeAg), HBcAg, and P are encoded by the 3.5 kb RNA. A large surface protein is encoded by 2.4 kb RNA. Middle-sized and small surface proteins (HBsAg) are mainly encoded by the 2.1 kb RNA and HBX is encoded by the 0.7 kb RNA.5,6
Nucleoside/nucleotide analogs and interferon (IFN-α) are two major clinical therapies for HBV-infected patients. However, drug-resistant viruses often appear after long-term treatment by nucleoside/nucleotide analogs and fewer than 40% of patients benefit from IFN therapy.7,8 IFN-stimulated genes (ISGs) are believed to defend against extracellular pathogens.9 A few ISGs have been reported to limit HBV replication directly and the mechanism of IFN-α therapy is not fully understood.10 Hence, further identification of anti-HBV ISGs is important for understanding and improving IFN therapy.
To identify the ISGs that can effectively limit HBV replication, we screened 285 ISGs in an HBV-infectible HepG2 cell line. By examining the level of antigen to HBV in cells overexpressing ISGs, we identified sterile alpha motif domain-containing 4A (SAMD4A), the overexpression of which strikingly inhibited HBV replication. SAMD4A is a mammalian homolog of Drosophila Smaug, which was reported to be a posttranscriptional repressor regulating processes such as maternal RNA destabilization, the maternal-to-zygotic transition, and early embryo development.11 The sterile alpha motif (SAM) domain contained in SAMD4A is a highly conserved domain shown to bind directly to an RNA stem loop, which is also known as a Smaug recognition region (SRE).12 SAMD4A is involved in neuron RNA granule formation13 and translational repression in mammals.14,15 High levels of SAMD4A expression reduce the nuclear accumulation of CUGBP1 in myoblasts from DM1 patients.16 A screen of a mutant mouse library revealed that SAMD4A-defective mice have multiple developmental defects due to the dysregulation of mTOC1 signaling17 and mitogen-inducible gene 6 expression.18 However, the participation of SAMD4A in host defenses has not been reported. In this study, we demonstrated that SAMD4A and its homolog, SAMD4B, can inhibit HBV replication. This inhibition is mediated by the binding of SAMD4A to the SRE-like sequence located in viral RNA. We also showed the inhibition of HBV by SAMD4A/B and murine SAMD4 in mouse models, suggesting the in vivo relevance of the anti-HBV function of SAMD4 family members.
Materials and methods
Cells and cell lines
HepG2, Huh7, HEK293, and HEK293T cells were purchased from ATCC. For in vitro HBV infection, HepG2-2B1 (a doxycycline-controlled human NTCP-overexpressing HepG2 cell line) and HepaRG cells were cultured as previously described.19,20 HBV viral stocks for the cell-based infection were obtained from a concentrate of HepAD38 cell culture supernatants following the method of a previous study.19 SAMD4A- and SAMD4B-KO HepG2 and HepG2-2B1 cells were generated using the CRISPR/Cas9 gene-editing technique. The knockout (KO) cells were generated by sequencing targeted loci. The HepG2, HepG2-2B1, HEK293T, Huh7, and AD38 cell lines were maintained in Dulbecco’s modified Eagle’s medium and supplemented with 10% fetal bovine serum, 2 mM l-glutamine, nonessential amino acids to a final concentration of 0.1 mM, 100 IU penicillin, and 100 mg/mL streptomycin at 37 °C in a humidified incubator with 5% CO2. All the cell lines were authenticated by their morphology and DNA sequencing, and were repeatedly tested to ensure they were mycoplasma-free, as assessed by the MycoAlert mycoplasma detection kit (Lonza, LT-07).
Reagents and antibodies
Doxycycline hyclate (D9891) and actinomycin D (A9415) were purchased from Sigma. The HBeAg ELISA kit and HBsAg ELISA kit were purchased from Beijing Wantai Biological Pharmacy. Rabbit anti-SAMD4A antibody (1 : 500, 17387-1-AP) and rabbit anti-SAMD4B (1 : 500, 17723-1-AP) were purchased from Proteintech Group. Mouse anti-Flag (1 : 2000, F3165) and rabbit anti-myc (1 : 2000, A7470) antibodies were purchased from Sigma. Rabbit anti-HA (1 : 5000, 3724S) was purchased from CST. Mouse anti-GAPDH (AC002, 1 : 5000) was purchased from ABclonal Technology. Rabbit anti-S6 (1 : 1000, #9452), rabbit anti-phospho-S6 (Ser235/236) (1 : 2000, #4858), rabbit anti-phospho-4EBP1 (Thr70) (1 : 1000, #13996), rabbit anti-phospho-4EBP1 (Ser65) (1 : 1000, #9456), rabbit anti-phospho-4EBP1 (Thr37/46) (1 : 1000, #2855), and rabbit anti-4EBP1 antibodies (1 : 1000, #9452) were purchased from CST. Rabbit anti-HBcAg and mouse anti-HBsAg antibodies were developed by Professor Quan Yuan as previously described.21 Mouse anti-HSV-1 GD (1 : 2000, sc-21719) was purchased from Santa Cruz. Rabbit anti-VSVG was generated by Xiamen University Laboratory Animal Center.
Constructs
All ISGs were amplified from a human cDNA library and cloned into a pBOB-CMV vector. cDNA from SAMD4A, SAMD4B, and a variety of their mutations were cloned into a pBOB-CMV vector with or without the indicated tag. HBV A-type, C-type, and D-type replicon plasmids cloned in the pTSMP-CMV vector were developed by Professor Quan Yuan as previously described.22 The SRE-like1 or mutant-SRE-like1 sequence was inserted into the 5′ or 3′-untranslated region (UTR) of the pBOB-CMV-GFP plasmid or the coding sequence after the 172nd glutamic acid. All plasmids were verified by DNA sequencing. Details of the plasmid sequence are available upon request. The following HBV SRE-like sequences were inserted into green fluorescent protein (GFP): SRE-like1: 5′-CTGACGCAACCCCCACTGGCTGGGGCTTGGTCATGG-3′; Mutant-SRE-like1: 5′-CTGACGCAACCCCCAATCGCTGGGGCTTGGTCATGG-3′; and SRE-like2: 5′-GTTTTGCTCGCAGCAGGTCTGGAGCAAACA-3′.
Lentivirus preparation and infection
For lentivirus production, HEK293T cells were transfected with lentiviral vectors and virus-packing plasmids by calcium phosphate precipitation. The virus-containing medium was collected 36 h after the transfection and added to HepG2-2B1 cells as indicated with 10 μg/mL polybrene. The infectious medium was replaced with fresh medium after 12 h.
Cell-based screening of anti-HBV ISGs
HBV viral stocks for the cell-based infection were obtained from a concentrate of HepAD38 cell culture supernatants following the method of a previous study.19 The expression of NTCP was induced by 1 μg/mL doxycycline administered to HepG2-2B1 cells, which were infected with lentivirus encoding different ISGs 2 days later and then challenged with HBV (multiplicity of infection = 30) after an additional 2 days. Sixteen hours after being infected with HBV, the cells were washed three times with phosphate-buffered saline (PBS) and then placed in fresh medium. The supernatants were collected every 2 days and subjected to an enzyme-linked immunosorbent assay (ELISA) using an HBeAg and surface antigen (HBsAg). Unless otherwise described, the protocol used to infect the HepG2-2B1 cells with HBV was also used for the other experiments described in this manuscript.
CRISPR/Cas9 gene-editing techniques
The gene-KO HepG2 and HepG2-2B1 cells were generated using the CRISPR/Cas9 gene-editing technique. The following initial targeting sequences were cloned in the gRNA vector:
SAMD4A: 5′-GCTCCCGGCGGCGAGCCTCG-3′;
SAMD4B: 5′-ACTGGAGGACCGCAACGCAC-3′;
4E-T: 5′-CGTAGCTTACCCGCAGAACT-3′; and
CNOT6: 5′-GTAGCTTACCCGCAGAACT-3′.
The plasmids carrying the gRNA and Cas9 genes were packed into lentiviruses and then used to infect HepG2-2B1 cells to mediate the gene editing. The KO cells were determined by sequencing targeted loci.
RNA interference
All lentivirus-shRNAs were inserted into a pLV-H1-EF1α-puro vector following the manufacturer’s instructions (Biosettia). The following shRNA target sequences were used:
SAMD4A: 5′-GCTCAGACTCTGTGGATTATG-3′;
SAMD4B: 5′-GGAGATGATGACACTGACTGA-3′;
4E-T-1: 5′-GCCTCTGTGAAGGAAGGTATA-3′;
4E-T-2: 5′-GCAAGTGGGACTTTGCCTTCT-3′;
CNOT6-1: 5′-GCATCTTTGTGGTCACTAA-3′; and
CNOT6-2: 5′-GCCATGATGCTTGTTTGCTTT-3′.
RNA–protein complex immunoprecipitation
HepG2 cells in 10 cm2 dishes were rinsed twice with ice-cold PBS before being collected in 2 mL of ice-cold PBS. The cell pellets were resuspended in 1 mL of RIP buffer (50 mM Tris pH 7.4 in Diethyl Pyrocarbonate-treated H2O, 150 mM NaCl, 4% Triton, 1 mM phenylmethylsulfonyl fluoride, and proteinase inhibitor cocktail, an RNase inhibitor) and placed on ice for 10 min to obtain cell extracts. For immunoprecipitation (IP), the cell extracts were incubated with 5 μg of Flag antibody for 2 h and incubated with A/G agarose beads for 1 h at 4 °C. The beads were washed five times with RIP buffer and resuspended in TRIzol (TaKaRa).
Quantitative PCR
Total RNA was extracted from cells with TRIzol according to the manufacturer’s instructions. cDNA was prepared with M-MLV reverse transcriptase and oligo-dT primers. Quantitative PCR analysis was performed using SYBR Green reagent along with gene-specific primers. All the results were analyzed by relative quantification and normalization to the RNA level of actin. The following primer sequences were used:
HBV DNA/total RNA RT-F: 5′-ACGTCCTTTGTTTACGTCCCGT-3′;
HBV DNA/total RNA RT-R: 5′-CCCAACTCCTCCCAGTCCTTAA-3′;
SAMD4A RT-F: 5′-ACACTTCCCCACAGAACATG-3′;
SAMD4A RT-R: 5′-TCATACTGAGGCACAACGAC-3′;
SAMD4B RT-F: 5′-GCACCTGGAGTCTCAGAAC-3′; and
SAMD4B RT-R: 5′-TGCAGAGCGTTTCGTAGG-3′.
Chromatin immunoprecipitation
Chromatin immunoprecipitation (ChIP) assays were performed following the standard protocol of the SimpleChIP Enzymatic Chromatin IP kit (Cell Signaling Technology; 9002S) using anti-RNA polymerase II antibody (Millipore, 05-952).
Enzyme-linked immunosorbent assays
The levels of HBsAg and HBeAg proteins in the culture supernatants of the HBV-infected HepG2-2B1 cells were measured by using HBsAg and HBeAg ELISA kits (Beijing Wantai) following the manufacturer’s protocol.
Southern and northern blotting
Total RNA was extracted from cells with TRIzol (TaKaRa) according to the manufacturer’s instructions. Contaminating DNA was removed by RNase-free DNase (Promega, M6101). RNAs were separated on a 1.2% agarose gel in the presence of formaldehyde and then blotted onto positively charged nylon membranes in 20× SSC (1× SSC is 0.15 M NaCl plus 0.015 M sodium citrate) buffer. The membranes were then probed with a Digoxigenin (DIG)-labeled HBV DNA probe. The secondary antibody was horseradish peroxidase-labeled goat anti-DIG and hybridization signals were visualized using X-ray exposure and an ECL western blotting substrate kit (Lulong Biotech).
Immunofluorescence imaging
Cells were fixed with freshly prepared 4% paraformaldehyde in PBS. The fixed cells were then permeabilized in 0.2% Triton X-100/PBS, blocked with 3% bovine serum albumin in PBS, stained with anti-HBcAg (mouse, 1 : 500) and labeled with goat anti-mouse Alexa Fluor 594 (1 : 1 000, Invitrogen) and Oregon green phalloidin (Invitrogen). All images were captured and processed using identical settings by a Zeiss LSM 780 laser scanning confocal microscope with a 100×/1.49 numerical aperture oil objective. Duplicate cultures were examined and similar results were obtained in at least three independent experiments.
Immunoprecipitation and western blotting
IP was performed using anti-Flag M2 beads as described.23 Western blotting of the cell lysates and IP were performed using anti-Flag and anti-HA as indicated. Three independent experiments were performed to assess reproducibility.
Mice
HBV transgenic mice were kindly provided by Professor Pei-Jer Chen (NTU, Taiwan).22 To generate Samd4f/f mice by CRISPR/Cas9 and a haploid embryonic stem cell system, the target sites of the guide RNA for Samd4 were designed as 5′-TCCATAGCTTGCCCGGATCT-3′ and 5′-GGTAAACTTTAGGGCCCAGT-3′. The haploid embryonic stem cells were a kind gift from Jin-Song LI (SIBCB, CAS). Additional information is provided upon request. All mice used in these studies shared a common genetic C57BL/6 background. All experiments were conducted in compliance with the regulations of Xiamen University.
Immunohistochemistry
During the mouse autopsy, dissected mouse tissues were harvested fresh and fixed in 4% formalin for 24 h. The fixed tissues were then embedded in paraffin and sliced in 5-µm thick sections with a microtome (Leica RM2016, Germany). Immunohistochemistry was performed using antibodies against rabbit HBcAg (1 : 1000) and mouse HBsAg (1 : 1000). The sections were counterstained with hematoxylin and then differentiated with 0.1% acid alcohol followed by bluing and a final dehydration.
Recombinant AAV preparation and injection
A recombinant AAV8-vector and pAAV8-SAMD4A virus were purchased from OBiO Technology (Shanghai). Eight-week-old male HBV transgenic mice were divided into three groups based on treatment received by intravenous injection: a normal control (n = 6), AAV8-vector (n = 6), or AAV8-SAMD4A (n = 6). Each group of mice was injected with 200 μl of saline containing 1 × 1012 v.g. of virus.
Statistical analysis
The data are presented as the means ± SD. The significance was evaluated with Student’s t-tests, two-way analyses of variance (ANOVAs) with Tukey’s post hoc tests, or repeated-measures ANOVAs for the time-course experiments. Statistical analyses were performed using GraphPad Prism, version 6.07 for Windows (GraphPad Software, Inc., La Jolla, CA).
For further details regarding the materials and methods used in this study, the reader is directed to the Supplementary Information.
Results
Identification of anti-HBV ISGs by cell-based screening
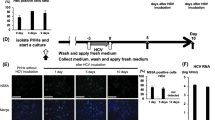
To identify the ISGs that may play roles in HBV propagation, we generated a cell line by stably transfecting a doxycycline-controlled NTCP-expressing plasmid into the HepG2 hepatocellular carcinoma cell line, enabling a cell-based screen to be performed (Supplementary Fig. 1A). A single clone that exhibits a high sensitivity to HBV (strain ayw, D type) infection was named HepG2-2B1 and was used in our experiments. It is well known that hundreds of ISGs are induced in response to IFN stimulation, but the degree of ISG upregulation varies among different cell lines.24 We compiled a list of 285 ISGs that were reported to be significantly induced in several cell lines and constructed lentiviral vectors to express each of them in the HepG2-2B1 cells before HBV challenge.25 The culture medium for each ISG-expressing HepG2-2B1 cell line was changed, the cells were collected 15 days post HBV infection (dpi) and the HBV level was determined by measuring the HBeAg level (Fig. 1a).
Identification of anti-HBV ISGs by cell-based screening. a Scheme of anti-HBV ISG screening. b Dot plots of the HBV level relative to that of the vector control (100%) (as indicated by HBeAg level) in the supernatants collected 15 days post HBV (MOI = 30) infection. Data show the average values from three independent experiments. The supernatant collected from the SAMD4A-expressing cells is indicated. c HepG2-2B1 cells were processed as in a. Levels of HBsAg (left) and HBeAg (right) in the supernatants of the HepG2-2B1 cells measured at different days post infection are shown. Note: the HBV antigens detected in the first few days were exogenously added to promote infection. d dHepaRG cells were infected with lentivirus encoding nothing (vector) or SAMD4A and challenged with HBV. HBsAg (left) and HBeAg (right) were measured on the ninth day after HBV (MOI = 30) infection. dHepaRG, differentiated HepaRG. e HepG2-2B1 cells were processed as in a. HBV RNA and DNA were analyzed by northern blotting (left) and Southern blotting (right) on the 15th day after infection. f HepG2 cells (left) and Huh7 cells (right) were infected with lentivirus encoding nothing (vector) or SAMD4A and transfected with HBV replicon (pTSMP-AD38) plasmids by lipofection. Levels of HBsAg in the supernatants were detected by ELISA on the third day. Data are presented as the means ± SD of triplicate samples, with similar results obtained in three to five independent experiments. ***P < 0.001
Figure 1b shows the relative level of HBeAg in the 15 dpi culture medium of the cells expressing different ISGs. Most ISGs had no or little effect on the propagation of HBV, as the level of HBeAg in the medium of these ISG-expressing cells was comparable with that of the control. Interestingly, several ISGs, including SAMD4A, Indoleamine-pyrrole 2,3-dioxygenase (IDO1), PML, ZAP, ISG20, MYD88, DDX3, and ZBP1, suppressed HBeAg production, suggesting that these ISGs are anti-HBV ISGs. Most of these proteins had been reported to have antiviral functions. ZBP1, also called DAI, is well-established as a cytoplasmic z-DNA sensor that can activate and amplify IFN signaling.26,27,28 PML functions directly on multiple kinds of virus proteins to limit virus replication.29,30,31 Moreover, some of these proteins are reported to directly inhibit HBV replication. The zinc finger protein ZAP recognizes the HBV terminal repeat region, degrades viral RNA and synergizes with multiple ISGs to mediate antivirus activity.32 IDO1, a tryptophan degradation enzyme, inhibits HBV by tryptophan deprivation to efficiently reduce the intracellular DNA level of the virus.33 ISG20 and MYD88 can directly bind to HBV RNA via different target sites and facilitate the decay of viral RNA,34,35 whereas DDX3 can inhibit HBV RNA reverse transcription.36 The inhibitory effect of MKX on HBeAg is most likely due to the toxicity it induces in HepG2 cells (data not shown). Most noticeably, SAMD4A was the protein that exhibited the highest inhibitory activity (more than 95%) on HBV in our screen (Fig. 1b), but its role in suppressing viral propagation has not been previously reported.
We also measured the HBeAg and HBsAg levels at different days after HBV infection and confirmed that overexpression of SAMD4A blocked the production of these two viral antigens in the HepG2-2B1 cells (Fig. 1c) but had no influence on the expression of NTCP (Supplementary Fig. 1A, B). HepaRG is a commonly used cell line for HBV studies, and SAMD4A overexpression also suppressed HBV replication in the differentiated HepaRG cells (Fig. 1d). Thus, we have identified an ISG that can very potently inhibit HBV.
SAMD4A inhibits HBV replication
We then analyzed the effect of SAMD4A on the levels of intracellular HBV protein, RNA and DNA in HBV-infected HepG2-2B1 cells. Western blotting revealed that SAMD4A expression also decreased intracellular HBcAg levels (Supplementary Fig. 1C). Reduction in the three long HBV RNAs (3.5 kb, 2.4 kb, and 2.1 kb) were detected by northern blotting (Fig. 1e left) and the level of HBV RNA reduction was comparable to that of the HBV proteins. The signal emitted by the 0.7 kb viral RNA was too weak to be detected; therefore, we could not determine whether this RNA was influenced by SAMD4A. Southern blotting analysis showed that the amount of viral DNA was significantly decreased in the SAMD4A-expressing cells (Fig. 1e, right). Collectively, our data demonstrated that SAMD4A inhibits HBV propagation. Because of secondary infection by the newly synthesized virus, we were unable to determine the site(s) where SAMD4A exerts its inhibition at this stage.
SAMD4A was reported as an RNA-binding protein that functions to regulate gene expression posttranscriptionally.11,12,37 To characterize the role of SAMD4A in HBV replication, we sought to determine whether SAMD4A inhibits HBV replication when the virus-entry steps are skipped. It is known that HBV cannot infect HepG2 and Huh7 cells due to the low NTCP expression in these cell lines but can be produced if the virus replicon pTSMP-AD38 is transiently transfected into these cell lines. We found that SAMD4A overexpression reduced HBsAg and HBeAg in the HepG2 and Huh7 cells transfected with pTSMP-AD38 (Fig. 1f and Supplementary Fig. 1D). The HepAD38 cell line inductively expresses a tetracycline operator/CMV promoter-controlled HBV genome and is widely used to produce HBV particles. We overexpressed SAMD4A in this cell line and detected profoundly inhibited HBV production (Supplementary Fig. 1E). As virus replication in these cell models bypassed early events, including virus binding, invasion, nucleocapsid translocation, rcDNA release, and conversion into cccDNA, SAMD4A interfered with virus replication at the late stage of the viral life cycle.
We next tested whether SAMD4A-mediated suppression occurs in other subtypes of HBV by transfecting the type A HBV and type C HBV replicons into SAMD4A-overexpressing HepG2 cells and found that both virus subtypes were inhibited by SAMD4A (Supplementary Fig. 1F). To determine whether the suppressive effect of SAMD4A is restricted to HBV, we tested the effect of SAMD4A overexpression on HSV and VSV in HEK293 cells and found that SAMD4A overexpression had no influence on the replication of HSV or VSV (Supplementary Fig. 1G).
We also used the CRISPR-Cas9 method to knock out SAMD4A in HepG2 cells and tested whether genetic deletion of SAMD4A affects HBV replication. In the absence of SAMD4A (Supplementary Fig. 2A), HBV production was doubled (Fig. 2a and Supplementary Fig. 2B). As a previously reported ISG,38 SAMD4A expression is induced by IFN (Supplementary Fig. 2C); therefore, we also examined whether SAMD4A is involved in the IFN-α-mediated inhibition of HBV. HBV was produced by transfecting pTSMP-AD38 plasmids into SAMD4A-KO and wild-type (WT) HepG2 cells. IFN-α pretreatment inhibited HBV production in the WT cells, and the level of inhibition in the KO cells was lower (Fig. 2b and Supplementary Fig. 2D). Hence, the induction of SAMD4A is one of the anti-HBV functions of IFN-α.
Deficiencies of SAMD4A and SAMD4B in the HepG2-2B1 cells enhance the replication of HBV. a HBsAg levels in the supernatants of the WT and SAMD4A-KO HepG2 cells were measured by ELISAs the third day after HBV replicon (pTSMP-AD38) transfection. b WT and SAMD4A-KO HepG2 cells were pretreated with 1000 U IFN-α for 8 h followed by transfection of the HBV replicon (pTSMP-AD38) by lipofection. HBsAg levels in the supernatants were detected by ELISAs on the third day post transfection. The relative HBsAg level was calculated based on the HBsAg level in the supernatant of the control (mock) cells, considered to be 100%. c HBsAg levels in the supernatants of the HepG2-2B1 cells with or without deletion of the SAMD4A, SAMD4B, or both SAMD4A, and SAMD4B genes were measured by ELISAs 15 days post HBV infection (MOI = 30). d The expression of SAMD4A and/or SAMD4B in two independent single clones of SAMD4A and SAMD4B double-KO HepG2-2B1 cells was reconstituted by lentivirus-mediated gene delivery. HBsAg in the supernatants of the HepG2-2B1 cells with or without reconstitution was measured by ELISA at 15 days post HBV infection (MOI = 30). The expression level of SAMD4A or SAMD4B was detected by western blotting. e HepG2 cells were infected with lentivirus carrying nothing (vector) or SAMD4B and transfected with the HBV replicon (pTSMP-AD38) by lipofection. The levels of HBsAg (left) and HBeAg (right) in the supernatants were detected by ELISAs on the third day. f HepAD38 cells were infected with lentivirus carrying nothing (vector) or SAMD4B, and tetracycline was withdrawn to induce HBV propagation. The level of HBsAg in the supernatant was detected by ELISA every day for a 6-day period. Data are presented as the means ± SD of triplicate samples, with similar results obtained in three independent experiments. *P < 0.05, **P < 0.01, and ***P < 0.001
SAMD4B exhibits a similar anti-HBV function as its homolog SAMD4A
SAMD4B is the only homologous protein of SAMD4A in humans, but it is not inducible by IFN. We knocked out SAMD4A and deleted SAMD4B in HepG2-2B1 cells (Supplementary Fig. 2E). Similar to the result shown for the HepG2 cells (Fig. 2a), the SAMD4A defect led to an approximately onefold increase in HBV production in the HepG2-2B1 cells. We observed more HBV production in the double-KO cells, suggesting that SAMD4B is also capable of inhibiting HBV replication (Fig. 2c). Overexpression of either SAMD4A or SAMD4B in double-KO HepG2-2B1 cells rescued the inhibition of HBV replication in these cells. Overexpressing SAMD4A and SAMD4B together had a slight but minimal additive effect on the suppression of HBV replication (Fig. 2d). The suppressive effect of SAMD4B on HBV replication was also observed in HepG2 cells and HepAD38 cells (Fig. 2e, f and Supplementary Fig. 2F). Thus, SAMD4B functions similarly to SAMD4A, and basal SAMD4A and SAMD4B have an inhibitory effect on HBV replication.
The SAM domain is required for SAMD4A and SAMD4B mediation of anti-HBV function
SAMD4A contains one SAM domain and two Smaug similarity regions (SSR1 and SSR2) (Fig. 3a). To determine which domain of SAMD4A is important for limiting HBV replication, we generated domain-deleted SAMD4A mutants and expressed them in HepG2-2B1 cells (Supplementary Fig. 3A). At similar expression levels, the SSR1 and SSR2 deletion mutants behaved similarly to the WT SAMD4A in suppressing HBV replication, whereas the deletion of the SAM domain eliminated the ability of SAMD4A to suppress HBV replication (Fig. 3b and Supplementary Fig. 3B). To confirm the importance of this SAM domain, we generated additional SAMD4A mutants, such as K250Q, A280H, K244A&R247A, in which the SAM domain structure was altered. These SAM domain-defective mutants lost the ability to suppress HBV replication (Fig. 3c and Supplementary Fig. 3C). Although the SAM domain is critical for the suppression of HBV, the expression of the SAM domain alone did not suppress HBV replication.
SAM domain is required for the anti-HBV function of SAMD4A. a Domain structure of SAMD4A. SAM sterile alpha motif, SSR Smaug similarity region. b Different domain-deleted SAMD4A was overexpressed by the lentiviral vector in HepG2-2B1 cells. HBsAg in the supernatants was measured by ELISA every 2 days for 15 days after HBV (MOI = 30) infection. c SAM domain deletion of SAMD4A or point mutation of SAMD4A was overexpressed in HepG2-2B1 cells. The HBsAg level in the supernatant was measured by ELISA on the tenth day post HBV (MOI = 30) infection. DNA or RNA in total lysates of the 10-day post-infection cells were extracted and analyzed by southern blotting and northern blotting, respectively. Lanes 1 and 2 of the northern blot are also presented in Fig. 1e. d SAMD4A with SAM domain deletion or point mutations was overexpressed in HepG2-2B1 cells. The HBV-infected cells were immunostained for HBV core antigen (HBcAg) and counterstained with Hoechst 4 days after the cells were infected. The scale bar is 100 μm. e SAMD4A, its A280H point mutant, or 392–529 aa-deleted mutant was overexpressed in HepG2-2B1 cells. HBsAg levels in the supernatant were measured by ELISAs 15 days post infection (MOI = 30). Data are presented as the means ± SD of triplicate samples, with similar results obtained in three independent experiments. *P < 0.05, ***P < 0.001, and NS P ≥ 0.05
Because of the high degree of homology between SAMD4A and SAMD4B, we also determined the role of the SAM domain in SAMD4B and confirmed the requirement to suppress HBV replication (Supplementary Fig. 3D). We also performed HBcAg immunostaining to evaluate the inhibitory role of SAMD4A and SAMD4B on HBV replication. Overexpression of SAMD4A or SAMD4B dramatically reduced the percentage of HBcAg-positive cells (Fig. 3d and Supplementary Fig. 3E). The reduction in HBcAg immunostaining was not observed in the cells overexpressing the SAM-defective mutants. Thus, the SAM domain plays a critical role in the SAMD4A- and SAMD4B-mediated inhibition of HBV replication, but the SAM domain alone is not sufficient to mediate antiviral activity.
As shown in Fig. 3b, SSR1 and SRR2 were not critical to SAMD4A suppression of HBV. However, the histidine (H) 86 mutation in SSR2 in mouse SAMD4 is known to impair mouse development17 and the mutation in the corresponding histidine (H85) to a proline (P) in human SAMD4A completely blocked its inhibition of HBV replication (Supplementary Fig. 3F), suggesting that the H to P mutation in SAMD4A causes a structural change that makes SAMD4A ineffective. The reported hypophosphorylation of 4EBP1 and S6 observed in SAMD4-expressing H86P mice is not thought to be involved in SAMD4A-mediated inhibition of HBV, as the phosphorylation level of these two proteins is comparable in the WT hepatocytes and in those with SAMD4A ectopic expression and SAMD4A knocked out (Supplementary Fig. 3G).
The C-terminal portion of SAMD4A is required for HBV suppression
To determine whether any portion(s) in addition to SAM is important for HBV suppression by SAMD4A, we generated a series of SAMD4A proteins in which the C-terminal end was partially deleted and we overexpressed these mutants in HepG2-2B1 cells to measure their effect on HBV replication. Although SAMD4A aa 1–533 mediated resistance to HBV replication, SAMD4A aa 1–455 completely lost antiviral function (Supplementary Fig. 4A, B). An alignment of the SAMD4A protein sequences from different species revealed that amino acids 392–529 of the human SAMD4A are conserved. A functional assay showed that the deletion of this C-terminal portion abolished the HBV replication suppressing of SAMD4A (Fig. 3e and Supplementary Fig. 4C). Thus, a conserved region in the C-terminus of SAMD4A is also required for the inhibition of HBV replication.
A conserved SRE stem loop in HBV RNA is targeted by SAMD4A
As SAMD4A affects HBV replication at the late stage of the viral life cycle and a SAM domain-containing protein can interact with RNA by recognizing the SRE, we tested whether SAMD4A binds to HBV RNA. We cotransfected the expression plasmid of 3× Flag-tagged SAMD4A and HBV replicon into HepG2 cells and then immunoprecipitated Flag-SAMD4A with anti-Flag beads. RNA was extracted from the immunoprecipitate and analyzed by real-time PCR for HBV RNA. Convincingly, HBV RNA was not detected in the immunoprecipitate of the IgG control, while a large amount of viral RNA was found in the SAMD4A immunoprecipitate, indicating that SAMD4A can interact with HBV RNA (Fig. 4a). It is important to note that the SAM domain is required for this function, because HBV RNA was negligibly detected in the immunoprecipitate of SAMD4A-ΔSAM. In contrast, the recently generated C-terminal region deletion had no influence on the interaction between SAMD4A and viral RNA (Supplementary Fig. 5A).
SRE in HBV RNA interacts with SAMD4A. a Cell lysate prepared from the HepG2 cells cotransfected with plasmids expressing pTSMP-AD38 and SAMD4A-Flag or SAMD4A-Flag with the SAM domain deleted and subjected to immunoprecipitation with anti-Flag antibody or control IgG. HBV RNA in the immunoprecipitate was detected by qPCR using HBV total RNA primers. b Left panel, location of the SRE-like sequences in the HBV genome. Right panel, predicted SRE-like1 and SRE-like2 stem loops. c SAMD4A-expressing plasmid with GFP containing SRE-like1 or SRE-like2 in the 3′-UTR was cotransfected into HEK293T cells. Thirty-six hours after transfection, the GFP fluorescence intensity was detected by flow cytometry. d GFP with SRE-like1 or C1A&G3C SRE-like1 in the 3′-UTR was coexpressed with SAMD4A in HEK293T cells. Cells were observed with a fluorescence microscope 36 h after transfection. The scale bar is 100 μm. e Cell lysates were prepared from the HepG2 cells coexpressing SAMD4A-Flag and pTSMP-AD38 or its SRE-like1 or SRE-like2 mutant, and subjected to immunoprecipitation with anti-Flag antibody. Untagged SAMD4A was used as a control for immunoprecipitation. HBV RNA in the immunoprecipitate of SAMD4A was detected by qPCR using HBV total RNA primers. f SAMD4A-expressing plasmid with pTSMP-AD38 or a SRE-like1 mutant plasmid was cotransfected into HepG2 cells by lipofection. HBsAg levels in the supernatant were detected by ELISA on the third day post transfection. Data are presented as the means ± SD of triplicate samples, with similar results obtained in three independent experiments. **P < 0.01, ***P < 0.001, and NS P ≥ 0.05
We then tried to find the targeting RNA sequence in SAMD4A. The SRE sequence bound by the Drosophila SAMD4A homolog Smaug was defined as a nonspecific stem with a 4- or 5 nt loop containing the CNGG or CNGGN sequence.39 We searched the sequence in the HBV genome and found two candidates located at 1195–1217 nt and 1296–1321 nt. The latter is known as a conserved HBV posttranscriptional regulatory element (HPRE), whereas there is no report on the 1195–1217 nt sequence (Fig. 4b). The 1195–1217 nt sequence is also conserved throughout all 61 HBV variants, for which sequence information is available. We named these two stem-loop SRE-like1 (1195–1217 nt) and SRE-like2 (1296–1321 nt), and examined whether these two stem loops were the target sites of SAMD4A. Each of them was cloned into the 3′-UTR of GFP, as Drosophila Smaug-binding SRE is in the 3′-UTR of mRNA (Supplementary Fig. 5B). Coexpression of the GFP reporter with or without SAMD4A in HEK293T cells revealed that neither stem-loop insertions influenced GFP expression, but SAMD4A coexpression led to decreased GFP expression in the cells with the plasmid containing SRE-like1 but not the plasmid containing SRE-like2 (Fig. 4c and Supplementary Fig. 5C). The guanine at the third position in the loop was reported to bind directly to Smaug and a hydrogen bond may form between cytosine 1 and guanine 4 to maintain the structure of the stem loop.12 We mutated both cytosine 1 and guanine 3 and found that the mutant stem-loop could not be suppressed by SAMD4A, as indicated by the measured GFP fluorescence intensity (Fig. 4d and Supplementary Fig. 5D). We generated an HBV replicon mutant by mutating cytosine 1 and guanine 4 in SRE-like1 and cotransfecting the expression plasmid with 3× Flag-SAMD4A and the mutant HBV replicon into HepG2 cells. We immunoprecipitated SAMD4A and measured the viral RNA level in the precipitate. Viral RNA with a mutated SRE-like1 was not pulled down by SAMD4A (Fig. 4e) and the replication of the mutant virus was not suppressed by SAMD4A overexpression (Fig. 4f). Taken together, our data indicate that SRE-like1 is a functional SRE, and that the binding of SAMD4A to this SRE is required for SAMD4A to suppress HBV replication.
SAMD4A reduces the levels of multiple HBV mRNAs
The HBV genome encodes four different mRNAs (Fig. 4b). To determine whether an SRE controls the expression of viral mRNA, we constructed expression plasmids driven by the CMV promoter to express each of the four HBV mRNAs together with or without SAMD4A in HepG2 cells. SAMD4A coexpression reduced the levels of the 3.5, 2.4, and 2.1 kb but not 0.7 kb viral RNA (Fig. 5a). It appears that SAMD4A selectively reduced the levels of the mRNA containing the SRE, regardless of whether the SRE was in the 3′-UTR of the HBe, HBc, and HBs mRNA (3.5, 2.4, and 2.1 kb, respectively) or in the coding regions in the HBV P protein mRNA (3.5 kb). An SRE in the 3′-UTR of the mRNA reduces gene expression. The aforementioned data suggest that the SRE might also reduce gene expression when it is in the coding region. To determine whether SRE can influence gene expression when it is in the 5′-UTR or coding region, we used the GFP reporter system. We inserted an SRE into the 5′-UTR or coding region of GFP and tested whether overexpression of SAMD4A affected the GFP expression (Supplementary Fig. 5B). As shown in Fig. 5b, c and Supplementary Fig. 5E, F, SAMD4A suppressed the expression of the GFP with the SRE in the coding region but had no effect on the GFP reporter with SRE in its 5′-UTR. Mutated SRE in the coding region also eliminated the SAMD4A-mediated suppression of GFP (Fig. 5c). Thus, viral proteins with SRE in the 3′-UTR or coding region of its mRNA could be suppressed by SAMD4A.
SAMD4A reduces the level of HBV mRNAs. a SAMD4A reduces the levels of the 3.5, 2.4, and 2.1 kb HBV mRNAs but not the 0.7 kb mRNA. SAMD4A and different HBV mRNAs driven by the CMV promoter vector were coexpressed in HepG2 cells. RNA was extracted two days after transfection, and the HBV RNA level was measured by qPCR using HBV total RNA primers. b, c SRE-like1 or its C1A&G3C mutant was inserted in the 5′-UTR (b) or coding sequence (next to the 172nd aa) (c) in GFP. SAMD4A was coexpressed with each of these reporters in HEK293T cells and GFP fluorescence intensity was measured 36 h after transfection. d Chromatin immunoprecipitation (ChIP) in digested chromatin from different HepG2 cells, in which nothing (vector), SAMD4A or SAMD4A A280H was overexpressed, was performed by anti-RNA polymerase antibody or control IgG. Purified DNA was analyzed by qPCR using primers targeting HBV DNA. The results are expressed as the ratio of the experimental group to IgG. e HepG2-2B1 cells were infected with lentivirus encoding nothing (vector), SAMD4A or SAMD4A A280H 5 days post infection with HBV. Actinomycin D (5 μg/mL) was added 24 h after transfection, and the RNA was extracted at the indicated time points. HBV RNA was measured by qPCR using HBV total RNA primers. f A total of 5 μg/mL actinomycin D was added to the WT and SAMD4A-KO HepG2-2B1 cells 5 days post-infection with HBV, and RNA was extracted at the indicated time points. HBV RNA was measured by qPCR using HBV total RNA primers. Data are presented as the means ± SD of triplicate samples, with similar results obtained in three independent experiments. *P < 0.05, **P < 0.01, ***P < 0.001, and NS P > 0.05
To determine whether SAMD4A affects the transcription of HBV, we performed a ChIP assay using an anti-RNA polymerase II antibody. Comparable amounts of HBV DNA were pulled down from the HepG2 cells cotransfected with the HBV replicon and plasmid expressing SAMD4A or SAMD4A A280H, a SAMD4A mutant that lost its function (Fig. 5d). Thus, SAMD4A does not affect HBV transcription.
We then examined whether SAMD4A affects HBV mRNA stability. SAMD4A- and SAMD4A A280H-overexpressing HepG2-2B1 cells were infected with HBV followed by the addition of actinomycin D to block transcription; these cells were collected at different time points after actinomycin D treatment. The primer pairs used to target the common sequence of the four different viral RNAs were used in real-time PCR to quantitate total HBV mRNA. Overexpression of SAMD4A but not SAMD4A A280H shortened the half-life of the HBV mRNA (Fig. 5e). Facilitating the effect of SAMD4A in reducing the stability of the HBV mRNA, the viral mRNA degradation in the SAMD4A-KO cells was slower than that in the WT cells (Fig. 5f). As SAMD4A did not affect the transcription of the HBV mRNA, mRNA instability may be the cause of the reduction in HBV mRNA induced by SAMD4A. As the SAMD4A-mediated reduction in HBV mRNA is comparable to that of proteins (Fig. 1c, e), the promotion of HBV RNA degradation may be the major mechanism of SAMD4A-mediated HBV suppression.
4E-T and CCR4–NOT complexes interact with SAMD4A but contribute little to SAMD4A-mediated suppression of HBV
As an interaction between Drosophila Smaug and CUP protein was detected, it was suggested that this interaction links Smaug with eIF4E, which competitively inhibits the protein translation initiated by the eIF4E–eIF4G complex.11,37,40 Smaug may also destabilize mRNA, as it interacts with CCR4, a core component of the CCR4–NOT deadenylation complex.41,42 To verify the involvement of these complexes in the SAMD4A-mediated suppression of HBV, we first tested whether SAMD4A interacts with the human homolog of eIF4E and CUP (named 4E-T or EIF4ENIF1 in humans), as well as CCR4 and NOT (named CNOT6 and CNOT8, respectively, in humans), by performing a coexpression assay of HEK293T cells. Consistent with previous research, both eIF4E and 4E-T were coimmunoprecipitated with SAMD4A in human cells, as were CNOT6 and CNOT8 (Fig. 6a). Although all the interactions were independent of the SAM domain (Supplementary Fig. 6A), deletion of the C-terminal portion abolished the interaction between SAMD4A and eIF4E or 4E-T, but not CNOT6 or CNOT8 (Supplementary Fig. 6B).
4E-T-eIF4E and CCR4–NOT complexes partially contribute to SAMD4A-mediated HBV suppression. a HEK293T cells were transfected with the indicated plasmids. Cell lysates were immunoprecipitated with anti-Flag antibody and blotted with anti-HA antibody. The total cell lysates were analyzed by western blotting with anti-HA and anti-flag antibodies. b WT HepG2-2B1 or two independent single clones of 4E-T-KO (left) CNOT6-KO (middle) and 4E-T/CNOT6 double-KO (right) HepG2-2B1 cells were infected with lentivirus carrying nothing (vector) or SAMD4A and challenged with HBV (MOI = 30). HBsAg levels in the supernatants were measured by ELISA on the tenth day post infection with HBV. Relative levels were calculated by using the HBsAg level of the empty virus-infected cells (vector), considered to be 100%. Data are presented as the means ± SD of triplicate samples, with similar results obtained in three independent experiments. *P < 0.05 and **P < 0.01
We then analyzed the function of the eIF4E–eIF4G complex and CCR4–NOT complex in SAMD4A-mediated antiviral events by using shRNA to downregulate the expression of 4E-T and CNOT6 (Supplementary Fig. 6C). Knocking down either 4E-T or CNOT6 negligibly reduced the antiviral ability of SAMD4A (Supplementary Fig. 6D). We also generated 4E-T-KO and CNOT6-KO HepG2-2B1 cells, and measured the anti-HBV capacity of SAMD4A in these cells. Consistent with the knockdown results, deletion of either 4E-T or CNOT6 only partially suppressed the SAMD4A-mediated inhibition of HBV (Fig. 6b). We also generated double KOs of 4E-T and CNOT6, and found no additive effect of the deletion of these two genes (Fig. 6b). These data suggest that the functions of 4E-T and CNOT6 complexes are coupled in SAMD4A-mediated HBV suppression. Because of the limited effect caused by deleting the 4E-T and CNOT6 genes, other unknown mechanism(s) may play major roles in SAMD4A-mediated suppression of HBV.
SAMD4 family members suppress HBV replication in vivo
As mouse hepatocytes cannot support productive HBV infection even when human NTCP is engineered to be expressed in them, there is no ideal mouse model for studying HBV infection. However, the effect of SAMD4A on HBV replication can be evaluated in mice by hydrodynamically injecting pTSMP-AD38 and SAMD4A- or SAMD4B-expressing plasmids into mice, which enables HBV production in mouse liver in the presence or absence of SAMD4A or SAMD4B overexpression.43,44 Serum and liver tissues were harvested and detected for viral antigens 3 days postinjection. The expression of SAMD4A or SAMD4B significantly reduced the levels of HBsAg and HBeAg secreted into the serum and the intrahepatic viral RNA level (Fig. 7a and Supplementary Fig. 7A-C). In addition immunohistochemical assays of the liver tissue revealed higher numbers of anti-HBcAg- and anti-HBsAg-stained cells in the control group than in the SAMD4A-overexpressing group (Fig. 7b). Moreover, mouse SAMD4, the homolog SAMD4A in mice, exhibited suppression of HBV replication similar to that in the cells in vitro (Supplementary Fig. 7D).
SAMD4A inhibits HBV production in the mice models. a, b SAMD4A- or SAMD4B-expressing plasmids (30 μg/mouse) and pTSMP-AD38 (15 μg/mouse) were coadministered in 8-week-old C57BL/6 J male mice by tail vein hydrodynamic injection. HBsAg level in serum was measured by ELISA on the third day (a); representative immunohistochemistry pictures of HBcAg and HBsAg in the liver tissue (left), and statistical graphs (right) are shown in b. The scale bar is 100 μm. c, d pTSMP-AD38 plasmid (12 μg/mouse) was injected into 8-week-old Samd4f/f or Albumin-Cre Samd4f/f male mice. HBsAg level in serum was measured by ELISA on the third day postinjection (c). Representative pictures of HBcAg and HBsAg in hepatocytes of the Samd4f/f or Albumin-Cre Samd4f/f mice (left), and the statistical graphs (right) are shown in d. e Eight-week-old HBV transgenic male mice were injected with 0.2 mL 0.9% NaCl solution (control) or 1 × 1012 AAV8 encoding nothing (vector) or SAMD4A in 0.2 mL of 0.9% NaCl solution. HBsAg level in serum was detected by ELISA before injection (mock) and every week after injection. Data from the mock samples and those obtained 4 weeks after injection are shown. f Database (GSE83148) containing gene expression data of 122 HBV-infected liver tissues and 6 healthy samples were analyzed with GEO2R. The levels of SAMD4A (left) and SAMD4B (right) in healthy people, HBV-low patients (HBV DNA ≤ 106), and HBV-high patients (HBV DNA > 106) are shown. Data are presented as the means ± SD of multiple samples (n ≥ 6). *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001, and NS P > 0.05
As we found that the expression of SAMD4 in mouse liver can be induced by injection of the HBV replicon (Supplementary Fig. 7E), analysis of HBV replication in SAMD4-deficient mice can be used to address the anti-HBV role of SAMD4 in vivo. As Samd4 mutant mice died postnatally,17 we conditionally knocked out Samd4 in the liver by crossing Albumin (Alb)-Cre mice with floxed Samd4 allele-carrying mice. The mice with Samd4 gene deletion in the liver grew normally and we did not find a difference in the size and morphology of the livers in the WT and mutant mice (Supplementary Fig. 7F, G). After pTSMP-AD38 hydrodynamic injection, the serum of the Samd4f/f Alb-Cre mice contained a significantly higher level of HBsAg and HBeAg than that of the Samd4f/f mice, with the intrahepatic HBV RNA level also higher (Fig. 7c and Supplementary Fig. 7H, I). In addition, the number of HBcAg- and HBeAg-positive cells was dramatically increased in the SAMD4-deficient mice compared with the number in the WT mice (Fig. 7d).
In the coinjection experiments described above, HBV replication and SAMD4A expression was initiated simultaneously. We also needed to determine whether SAMD4A inhibits HBV replication after the virus had already propagated in vivo. As an HBV-producing transgenic mouse strain was available, we used AAV to express SAMD4A in the transgenic mice and analyzed viral antigens in their serum. The expression of SAMD4A decreased viral HBsAg levels in the transgenic mice (Fig. 7e). Collectively, the mouse experiments demonstrated that HBV replication can be suppressed by SAMD4A and SAMD4B in vivo.
SAMD4A and SAMD4B levels were negatively correlated with the HBV titers in samples from chronic hepatitis B patients
Zhou et al.45 analyzed the gene expression profiles of liver tissues from 122 patients with chronic hepatitis B and 6 healthy people. The patient samples were divided into HBV-high (viral DNA > 106 copies/mL) and HBV-low (viral DNA < 106 copies/mL) groups in this study. We compared SAMD4A and SAMD4B expression levels in the healthy people, HBV-low patients, and HBV-high patients, and found a negative correlation between the HBV titers and SAMD4A and SAMD4B levels in the patient samples (Fig. 7f). Interestingly, the SAMD4A level in the healthy individuals was comparable to that in the HBV-infected patients, but the SAMD4B level was significantly elevated in the hepatitis B patients. This finding suggests that, although SAMD4B is not an ISG, it might be a gene that is regulated during host defense against HBV.
Discussion
IFN-α is widely used to treat hepatitis B, but its anti-HBV mechanism is not fully understood. In this study, we screened and identified SAMD4A as the strongest antiviral ISG from nearly three hundred ISGs identified (Fig. 1a, b). SAMD4A and its homolog SAMD4B inhibit HBV by promoting HBV RNA degradation. SAMD4A and SAMD4B suppresses HBV replication in vitro and in mouse models, and their expression levels were negatively correlated with the HBV titers in hepatitis B patient samples.
SAMD4A suppresses all subtypes of HBV but not HSV or VSV. A deep literature search revealed that SAMD4A was identified in a screen of anti-Togaviridae virus factors, indicating that the viral suppression function of SAMD4A is not limited to HBV.25,46 It is highly likely that SAMD4A exhibits an inhibitory effect on all viruses with mRNAs that contains an SRE loop. The SRE sequence is conserved in HBV, suggesting that it may be essential for the viral life cycle. Several stem loops, such as the ε region and HPRE, were previously reported to be contained in HBV RNA, where it controls encapsidation and viral RNA nuclear export.47 Other anti-HBV factors, such as ISG20,34 RIG-I,48 DDX3,36 and MYD88,35 inhibit virus replication by targeting ε or HPRE. However, the antiviral ability of these factors may interfere with each other because all of them target ε or HPRE. The SRE loop in HBV RNA was revealed in this study and is the unique site targeted by SAMD4A. We showed that SAMD4A can recognize and degrade RNA containing the SRE motif in the 3′-UTR or coding sequence but not in the 5′-UTR, which is an important piece of information for the future identification of other SAMD4A-regulated RNAs.
As reported in Drosophila,21 the SAM domain is required for SAMD4A-mediated antiviral function. Moreover, we showed that the C-terminal domain in SAMD4A is also important for SAMD4A-mediated suppression of HBV (Fig. 3e). Its requirement for the binding of 4E-T may contribute to the suppression of HBV replication. Since the contribution of the 4E-T and CCR4–NOT complexes in SAMD4A-mediated HBV suppression is limited, the major mechanism(s) by which SAMD4A suppresses gene expression awaits further investigation.
The in vivo suppression of HBV by SAMD4A was demonstrated by overexpressing human SAMD4A, human SAMD4B or mouse SAMD4 and hepatocyte-specific gene deletion of Samd4 in mice. Overexpression of SAMD4A in the HBV-producing transgenic mice also inhibited HBV propagation. The negative correlation of SAMD4A and SAMD4B levels and HBV titers in human hepatitis patients supports the physiological relevance of SAMD4A/B-mediated suppression of HBV (Fig. 7f). Therefore, future SAMD4A/B agonist screening may provide valuable information for therapeutic inventions of hepatitis B.
References
Perz, J. F., Armstrong, G. L., Farrington, L. A., Hutin, Y. J. F. & Bell, B. P. The contributions of hepatitis B virus and hepatitis C virus infections to cirrhosis and primary liver cancer worldwide. J. Hepatol. 45, 529–538 (2006).
Yan, H. et al. Sodium taurocholate cotransporting polypeptide is a functional receptor for human hepatitis B and D virus. Elife 1, e00049 (2012).
Li, H. et al. HBV life cycle is restricted in mouse hepatocytes expressing human NTCP. Cell Mol. Immunol. 11, 175–183 (2014).
Tuttleman, J. S., Pourcel, C. & Summers, J. Formation of the pool of covalently closed circular viral-DNA in hepadnavirus-infected cells. Cell 47, 451–460 (1986).
Tong, S. P. & Revill, P. Overview of hepatitis B viral replication and genetic variability. J. Hepatol. 64, S4–S16 (2016).
Newbold, J. E. et al. The covalently closed duplex form of the hepadnavirus genome exists in-situ as a heterogeneous population of viral minichromosomes. J. Virol. 69, 3350–3357 (1995).
Ghany, M. & Liang, T. J. Drug targets and molecular mechanisms of drug resistance in chronic hepatitis B. Gastroenterology 132, 1574–1585 (2007).
Perrillo, R. Benefits and risks of interferon therapy for hepatitis B. Hepatology 49, S103–S111 (2009).
Wieland, S. F., Asabe, S., Engle, R. E., Purcell, R. H. & Chisari, F. V. Limited hepatitis B virus replication space in the chronically hepatitis C virus-infected liver. J. Virol. 88, 5184–5188 (2014).
Tan, G. Y., Song, H. X., Xu, F. C. & Cheng, G. H. When hepatitis B virus meets interferons. Front. Microbiol. 9, 1611 (2018).
Smibert, C. A., Wilson, J. E., Kerr, K. & Macdonald, P. M. Smaug protein represses translation of unlocalized nanos mRNA in the Drosophila embryo. Gene Dev. 10, 2600–2609 (1996).
Aviv, T., Lin, Z., Ben-Ari, G., Smibert, C. A. & Sicheri, F. Sequence-specific recognition of RNA hairpins by the SAM domain of Vts1p. Nat. Struct. Mol. Biol. 13, 168–176 (2006).
Baez, M. V. & Boccaccio, G. L. Mammalian smaug is a translational repressor that forms cytoplasmic foci similar to stress granules. J. Biol. Chem. 280, 43131–43140 (2005).
Baez, M. V. et al. Smaug1 mRNA-silencing foci respond to NMDA and modulate synapse formation. J. Cell Biol. 195, 1141–1157 (2011).
Fernandez-Alvarez, A. J., Pascual, M. L., Boccaccio, G. L. & Thomas, M. G. Smaug variants in neural and non-neuronal cells. Commun. Integr. Biol. 9, e1139252 (2016).
de Haro, M. et al. Smaug/SAMD4A restores translational activity of CUGBP1 and suppresses CUG-induced myopathy. PLoS Genet. 9, e1003445 (2013).
Chen, Z. et al. Mutation of mouse Samd4 causes leanness, myopathy, uncoupled mitochondrial respiration, and dysregulated mTORC1 signaling. Proc. Natl Acad. Sci. USA 111, 7367–7372 (2014).
Niu, N. N. et al. RNA-binding protein SAMD4 regulates skeleton development through translational inhibition of Mig6 expression. Cell Discov. 3, 16050 (2017).
Zhang, Y. L. et al. Robust in vitro assay for analyzing the neutralization activity of serum specimens against hepatitis B virus. Emerg. Microbes Infect. 8, 724–733 (2019).
Gripon, P. et al. Infection of a human hepatoma cell line by hepatitis B virus. Proc. Natl Acad. Sci. USA 99, 15655–15660 (2002).
Wu, Y. et al. Sleeping beauty transposon-based system for rapid generation of HBV-replicating stable cell lines. J. Virol. Methods 234, 96–100 (2016).
Zhang, T. Y. et al. Prolonged suppression of HBV in mice by a novel antibody that targets a unique epitope on hepatitis B surface antigen. Gut 65, 658–671 (2016).
Zhang, D. W. et al. RIP3, an energy metabolism regulator that switches TNF-induced cell death from apoptosis to necrosis. Science 325, 332–336 (2009).
Ivashkiv, L. B. & Donlin, L. T. Regulation of type I interferon responses. Nat. Rev. Immunol. 14, 36–49 (2014).
Schoggins, J. W. et al. A diverse range of gene products are effectors of the type I interferon antiviral response. Nature 472, 481–U545 (2011).
Takaoka, A. et al. DAI (DLM-1/ZBP1) is a cytosolic DNA sensor and an activator of innate immune response. Nature 448, 501–U514 (2007).
Kuriakose, T. et al. ZBP1/DAI is an innate sensor of influenza virus triggering the NLRP3 inflammasome and programmed cell death pathways. Sci. Immunol. 1, aag2045 (2016).
Chen, Q. Y. et al. DNA-dependent activator of interferon-regulatory factors inhibits hepatitis B virus replication. World J. Gastroenterol. 18, 2850–2858 (2012).
Wang, W. H., Studach, L. L. & Andrisani, O. M. Proteins ZNF198 and SUZ12 are down-regulated in hepatitis B virus (HBV) X protein-mediated hepatocyte transformation and in HBV replication. Hepatology 53, 1137–1147 (2011).
Tavalai, N., Papior, P., Rechter, S. & Stamminger, T. Nuclear domain 10 components promyelocytic leukemia protein and hDaxx independently contribute to an intrinsic antiviral defense against human cytomegalovirus infection. J. Virol. 82, 126–137 (2008).
Regad, T. & Chelbi-Alix, M. K. Role and fate of PML nuclear bodies in response to interferon and viral infections. Oncogene 20, 7274–7286 (2001).
Mao, R. C. et al. Inhibition of hepatitis B virus replication by the host zinc finger antiviral protein. PLoS Pathog. 9, e1003494 (2013).
Mao, R. C. et al. Indoleamine 2,3-dioxygenase mediates the antiviral effect of gamma interferon against hepatitis B virus in human hepatocyte-derived cells. J. Virol. 85, 1048–1057 (2011).
Liu, Y. J. et al. Interferon-inducible ribonuclease ISG20 inhibits hepatitis B virus replication through directly binding to the epsilon stem-loop structure of viral RNA. PLoS Pathog. 13, e1006296 (2017).
Li, J. H. et al. Inhibition of hepatitis B virus replication by MyD88 involves accelerated degradation of pregenomic RNA and nuclear retention of Pre-S/S RNAs. J. Virol. 84, 6387–6399 (2010).
Wang, H. F., Kim, S. & Ryu, W. S. DDX3 DEAD-Box RNA helicase inhibits hepatitis B virus reverse transcription by incorporation into nucleocapsids. J. Virol. 83, 5815–5824 (2009).
Nelson, M. R., Leidal, A. M. & Smibert, C. A. Drosophila Cup is an eIF4E-binding protein that functions in Smaug-mediated translational repression. EMBO J. 23, 150–159 (2004).
Brodsky, L. I. et al. A novel unsupervised method to identify genes important in the anti-viral response: application to interferon/ribavirin in hepatitis C patients. PLoS ONE 2, e584 (2007).
Aviv, T. et al. The RNA-binding SAM domain of Smaug defines a new family of post-transcriptional regulators. Nat. Struct. Biol. 10, 614–621 (2003).
Nishimura, T. et al. The eIF4E-binding protein 4E-T is a component of the mRNA decay machinery that bridges the 5′ and 3′ termini of target mRNAs. Cell Rep. 11, 1425–1436 (2015).
Semotok, J. L. et al. Smaug recruits the CCR4/POP2/NOT deadenylase complex to trigger maternal transcript localization in the early drosophila embryo. Curr. Biol. 15, 284–294 (2005).
Zaessinger, S., Busseau, I. & Simonelig, M. Oskar allows nanos mRNA translation in Drosophila embryos by preventing its deadenylation by Smaug/CCR4. Development 133, 4573–4583 (2006).
Huang, L. R., Wu, H. L., Chen, P. J. & Chen, D. S. An immunocompetent mouse model for the tolerance of human chronic hepatitis B virus infection. Proc. Natl Acad. Sci. USA 103, 17862–17867 (2006).
Yang, P. L., Althage, A., Chung, J. & Chisari, F. V. Hydrodynamic injection of viral DNA: a mouse model of acute hepatitis B virus infection. Proc. Natl Acad. Sci. USA 99, 13825–13830 (2002).
Zhou, W. et al. Predictive model for inflammation grades of chronic hepatitis B: large-scale analysis of clinical parameters and gene expressions. Liver Int. 37, 1632–1641 (2017).
Hubel, P. et al. A protein-interaction network of interferon-stimulated genes extends the innate immune system landscape. Nat. Immunol. 20, 493–49 (2019).
Pollack, J. R. & Ganem, D. An RNA stem-loop structure directs hepatitis-B virus genomic RNA encapsidation. J. Virol. 67, 3254–3263 (1993).
Sato, S. et al. The RNA sensor RIG-I dually functions as an innate sensor and direct antiviral factor for hepatitis B virus. Immunity 42, 123–132 (2015).
Acknowledgements
This work was supported by the National Scientific and Technological Major Project (2017ZX10202203-003), the National Natural Science Foundation of China (81788101, 31420103910, and 81630042), the 111 Project (B12001), and the National Science Foundation of China for Fostering Talents in Basic Research (J1310027).
Author information
Authors and Affiliations
Contributions
Y.W., X.F., Q.Y., N.X., and J.H. conceived and designed the experiments. Y.W., X.F., Y.S., R.L., Y.L., J.W., and X.L. performed the experiments. Y.W., X.F., Y.S., G.F., N.X., and J.H. analyzed the data. Y.W., X.F., and J.H. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Wang, Y., Fan, X., Song, Y. et al. SAMD4 family members suppress human hepatitis B virus by directly binding to the Smaug recognition region of viral RNA. Cell Mol Immunol 18, 1032–1044 (2021). https://doi.org/10.1038/s41423-020-0431-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41423-020-0431-x
Keywords
This article is cited by
-
Interferon-γ-induced GBP1 is an inhibitor of human papillomavirus 18
BMC Women's Health (2024)
-
RNA binding protein SAMD4: current knowledge and future perspectives
Cell & Bioscience (2023)
-
Restoration of HBV-specific CD8+ T-cell responses by sequential low-dose IL-2 treatment in non-responder patients after IFN-α therapy
Signal Transduction and Targeted Therapy (2021)