Abstract
The Cav3.2 subtype of T-type calcium channels has been targeted for developing analgesics and anti-epileptics for its role in pain and epilepsy. Here we present the cryo-EM structures of Cav3.2 alone and in complex with four T-type calcium channel selective antagonists with overall resolutions ranging from 2.8 Å to 3.2 Å. The four compounds display two binding poses. ACT-709478 and TTA-A2 both place their cyclopropylphenyl-containing ends in the central cavity to directly obstruct ion flow, meanwhile extending their polar tails into the IV-I fenestration. TTA-P2 and ML218 project their 3,5-dichlorobenzamide groups into the II-III fenestration and place their hydrophobic tails in the cavity to impede ion permeation. The fenestration-penetrating mode immediately affords an explanation for the state-dependent activities of these antagonists. Structure-guided mutational analysis identifies several key residues that determine the T-type preference of these drugs. The structures also suggest the role of an endogenous lipid in stabilizing drug binding in the central cavity.
Similar content being viewed by others
Introduction
Voltage-gated calcium (Cav) channels respond to membrane depolarization, allowing Ca2+ influx and translating electrical signals into intracellular Ca2+-mediated events.1,2,3,4 They play instrumental roles in Ca2+-dependent physiological processes such as neurotransmitter/hormone release, gene expression, and muscle contraction.3,5,6,7,8,9 Precise control of calcium channel activity involves various cellular factors such as receptors, calmodulin, phospholipids, and G proteins.10,11,12,13,14,15,16,17,18 Additionally, these channels can be modulated by a range of ions, toxins, and numerous clinical and investigational drugs.19,20,21,22,23,24
Among the 10 mammalian Cav subtypes (Cav1.1-Cav1.4, Cav2.1-Cav2.3, and Cav3.1-Cav3.3), the Cav3 subfamily, also known as the T-type calcium channels, is characterized by tiny and transient currents, in contrast to the large and long-lasting L-type Cav1 channels.25,26,27,28 Unlike Cav1 and Cav2 members, which require multiple auxiliary subunits for channel activities, the T-type channels function independently.29,30,31 In addition, while Cav1 and Cav2 channels are activated at high voltage, thus also known as high voltage-activated (HVA) channels, Cav3 channels are low voltage-activated (LVA).28,32,33 Their functional distinctions are supported by a large degree of sequence variations between the LVA and HVA channels, offering an opportunity to develop subtype-specific modulators for potential drug discovery.
Cav3.2, encoded by CACNA1H and widely expressed in the central nervous system, regulates neuronal excitability and participates in nociception.34,35 Alterations in its activity are associated with many neurological and neuropsychiatric disorders, including idiopathic generalized epilepsy and pain.36,37,38,39 Studies indicate increased expression and/or activity of Cav3.2 in spinal dorsal horn and in dorsal root ganglion neurons in various inflammatory and neuropathic pain models.36 Silencing or pharmacological inhibition of Cav3.2 channels induces analgesia in rodents.40 Consequently, Cav3.2 is emerging as a promising drug target for the development of next-generation analgesics. Compounds targeting Cav3.2, such as ethosuximide, mibefradil, valproate, zonisamide, pimozide, and certain dihydropyridines (DHPs), demonstrate efficacy in rodent models of acute, inflammatory, and chronic pain.40,41,42
There are ongoing efforts to scrutinize various small molecules with novel chemical skeletons as potential analgesic candidates, such as Z944, TTA-A2, TTA-P2, ML218, and ACT-709478.43,44,45,46 Compared to the approved drugs that usually have limited specificity for T-type channels, these investigational T-type-specific blockers demonstrate stronger affinities for T-type Cav channels. Although these compounds have yet to discriminate between T-type subtypes, the distinctive pharmacokinetic properties of Cav3.2, compared to Cav3.1 and Cav3.3, offer an opportunity to develop bona fide Cav3.2-selective molecules.
Gaining high-resolution insights into the structures of channels complexed with FDA-approved drugs or lead compounds is crucial for guiding de novo drug design or optimization. The structure of Cav3.1 bound to a lead compound Z944 reveals the basic architecture of the T-type channels and the molecular basis for the state-dependent inhibition of Cav3 subfamily by Z944.44 A recent report on the structures of Cav3.3 in complex with mibefradil, pimozide, and otilonium bromide broadens the spectrum of drug recognitions targeting T-type calcium channels.47 In this study, we sought to determine the structures of human Cav3.2 alone and in complex with representative antagonists, with a particular focus on those exhibiting enhanced selectivity for T-type calcium channels.
Results
Structural determination of human Cav3.2
The bottleneck in the structural analysis of human Cav3.2 arises from the low yield of recombinant protein expression. To enhance protein production, various constructs were explored. The I-II loop plays an inhibitory role in Cav3.2 expression and function.48,49 We thereby introduced several internal truncations of different fragments to this region. Eventually, a variant with the deletion of residues 493-772 resulted in an elevated expression level and decent solution behavior (Supplementary information, Fig. S1). This variant was named Cav3.2EM, as it was used for cryogenic electron microscopy (cryo-EM) imaging.
The biophysical properties of Cav3.2EM were verified through whole-cell patch-clamp recordings. Consistent with its improved protein expression, Cav3.2EM exhibited an increased conductance compared with the wild-type (WT) channel. Additionally, both the activation and steady-state inactivation curves of Cav3.2EM demonstrated a slight leftward shift in comparison to WT (Fig. 1a; Supplementary information, Fig. S2 and Table S1).
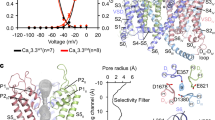
a Electrophysiological characterizations of full-length human Cav3.2WT (black) and truncated Cav3.2EM (red) in HEK293T cells. Voltage-dependent activation and steady-state inactivation curves are presented in the upper and lower panels, respectively. n values indicate the number of independent cells; data are presented as mean ± SEM. Please refer to Materials and Methods and Supplementary information for details. b Cryo-EM reconstruction of human Cav3.2EM. The EM map is color-coded for the four repeats (upper left), and the sugar moieties and lipids are colored bright and pale orange, respectively. The same color scheme is applied throughout the manuscript. The overall structure of human Cav3.2 is shown in a side view (upper right) and top view (bottom). c Structural comparison of the three T-type Cav channels: Cav3.1 (PDB: 6KZO, light green), Cav3.2 (domain colored), and Cav3.3 (PDB: 7WLI, pink). d The ion-conducting path of Cav3.2 is calculated in HOLE70 and illustrated with purple dots (left). The calculated pore radii along the permeation path are depicted as a purple line (middle). Two constriction sites, the SF enclosed by the EEDD motif and the intracellular gate, are shown on the right, in extracellular views. e Ca2+ coordination in the SF. Densities for the SF, prepared in ChimeraX, are contoured at the level of 4 σ, and the potential Ca2+ ion is shown as a green sphere.
Following the functional validation of Cav3.2EM, we performed standard cryo-EM single particle analysis and obtained a three-dimensional (3D) EM reconstruction at an overall resolution of 3.0 Å. This structure is henceforth referred to as Cav3.2Apo. Two glycan chains were observed, attaching to Asn345 and Asn1466 in the extracellular loops in repeats I and III (ECLI and ECLIII) via N-link (Fig. 1b; Supplementary information, Figs. S3, S4 and Table S2). Previous studies have also suggested the importance of these N-glycosylation sites in regulating the functional expression of Cav3.2.50,51,52
The overall structure of Cav3.2Apo displays an inactivated state nearly identical to that observed for Cav3.1 and Cav3.3,44,47 with root-mean-square deviations (RMSDs) of 0.58 Å over 847 Cα atoms and 1.16 Å over 948 Cα atoms, respectively (Fig. 1c). All four voltage-sensing domains (VSDs) adopt the depolarized or “up” conformation, and the intracellular gate is tightly twisted. The closed intracellular gate comprises three layers of hydrophobic residues. The first assembly site beneath the central cavity includes conserved residues Leu415, Leu1010, Phe1550, and Val1855, a common feature observed in the structures of all three members of Cav3 channels (Fig. 1d). A spherical density, likely corresponding to a Ca2+ ion, is surrounded by four residues, Glu378, Glu974, Asp1504, and Asp1808 (the EEDD motif) in the selectivity filter (SF) (Fig. 1e).
Structural determination of Cav3.2 with different antagonists
Next, we set out to determine the structures of Cav3.2 in complex with representative T-type channel-selective antagonists. To validate the action of these compounds on both WT Cav3.2 channel and Cav3.2EM variant, we conducted whole-cell patch-clamp recordings in HEK293T cells (Supplementary information, Fig. S5 and Tables S3, S4). The characterizations indicated that Cav3.2EM exhibited comparable potency across all tested compounds compared to the WT channel. Each antagonist was individually incubated with purified Cav3.2 protein at a final concentration at least 10-fold higher than its IC50 value before cryo-sample preparation. Following similar protocols for cryo-EM data acquisition and analysis, we successfully resolved the structures of Cav3.2 in complex with four compounds, ACT-709478, TTA-A2, TTA-P2, and ML218, with overall resolutions ranging from 2.8 Å to 3.2 Å. For simplicity, we will refer to these structures as Cav3.2-ACT/TA/TP/ML, with “ACT” representing ACT-709478, “TA” for TTA-A2, “TP” for TTA-P2, and “ML” for ML218 (Supplementary information, Fig. S6 and Table S2).
All four complex structures resemble the apo form, with the compounds each accommodated within the pore domain (PD) (Supplementary information, Fig. S6q). The four compounds can be further categorized into two groups: ACT-709478 and TTA-A2 insert through the IV-I fenestration, while TTA-P2 and ML218 dock on the II-III fenestration. Apart from the fenestration binding site, the other end of these elongated compounds is nestled within the central cavity. The fenestration-accommodating binding poses immediately suggest the molecular basis for their state-dependent pore-blocking mechanism. In the following text, we will illustrate their binding details, which will facilitate future drug design and optimization.
Coordination of ACT-709478 and TTA-A2
Both ACT-709478 and TTA-A2 were well resolved. The distinct structural features within the densities, like the trifluoromethyl group in ACT-709478 and the methyl group in TTA-A2, enabled reliable model building of these small-molecule compounds (Supplementary information, Fig. S6).
The head groups, characterized by the cyclopropylphenyl group in TTA-A2 or the trifluoromethyl cyclopropylphenyl moiety in ACT-709478, reside in the center of the cavity, which we named site C in Nav channels53,54 (Fig. 2a, b). These groups are coordinated similarly by several hydrophobic residues from the S6 tetrahelical bundles, including Phe408 and Asn412 on S6I, Phe1007 and Leu1010 on S6II, Val1546, Leu1547, and Phe1550 on S6III, and Gln1848, Leu1851, and Val1852 on S6IV (Fig. 2c, d).
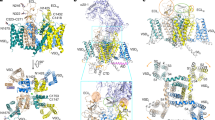
a Chemical structures of ACT-709478 and TTA-A2. The two compounds share a common core structure, featuring a cyclopropylphenyl head (highlighted in red), an aromatic tail (highlighted in green), and an amide linker. b Structural basis for pore block by ACT-709478 or TTA-A2. Both molecules traverse the central cavity, with one end inserting into the IV-I fenestration. The two structures, named Cav3.2-ACT and Cav3.2-TA, are superimposed relative to the PD. ACT-709478 and TTA-A2 are shown as brown and black sticks, respectively, and only the PD of Cav3.2-ACT is shown. c Detailed coordination of ACT-709478 and TTA-A2. The aromatic tail groups of ACT-709478 and TTA-A2, which vary in details, can both be accommodated in the IV-I fenestration. Potential H-bonds are highlighted with red dashed lines. d Schematic representation of residues constituting the binding site for ACT-709478 (upper) and TTA-A2 (lower). Residues within a 4 Å cutoff distance to the ligand are shown, with the binding pocket and potential H-bonds indicated by gray dashed contour and red dashed lines, respectively.
Both tail groups, despite their distinct chemical structures, wedge into the IV-I fenestration, indicating the fenestration’s versatile adaptability to diverse molecules. Per our recently proposed nomenclature system for the druggable sites on Nav channels,53 this site will be described as site F4. The tail groups in the IV-I fenestration are surrounded by hydrophobic residues, Leu377, Ile403, Ser407, and Phe408 from repeat I, as well as Phe1756, Phe1802, Ser1805, and Thr1806 from repeat IV. In the case of ACT-709478, the nitrogen within the cyanopyridine ring is further stabilized through a hydrogen bond (H-bond) with the hydroxyl group from Ser1805. Furthermore, both internal amide linkages are H-bonded with Asn412 and Gln1848, contributing to the stability of these unique binding poses (Fig. 2c, d).
An α-to-π transition of S6II in the presence of TTA-A2
ACT-709478 and TTA-A2 share a similar pharmacophore featuring a cyclopropylphenyl head, an aromatic tail, and an amide linkage. However, there are local structural variations in the presence of these two compounds. While the conformation of Cav3.2-ACT is nearly identical to that of the apo channel, structural rearrangements occur in the presence of TTA-A2, as exemplified by an α-to-π transition in the middle of the S6II segment (Fig. 3a).
a An α-to-π transition observed in the S6II segment upon TTA-A2 binding. Left: Red arrows indicate the structural differences between Cav3.2-ACT (gray) and Cav3.2-TA (domain colored). An enlarged view that highlights the change of S6II is shown in the inset. Right: Rotation of the bottom half of the S6II helix in the presence of TTA-A2, but not ACT-709478. b Closure of the I-II fenestration in Cav3.2-TA. Corresponding surface views of the four sides of the PD are presented for Cav3.2-ACT (upper) and Cav3.2-TA (lower).
A close structural comparison of Cav3.2-ACT/TA reveals that the conformational deviation is caused by the minor difference in the head group. The only divergence of the head group of the two molecules pertains to the presence of an additional trifluoromethyl group in ACT-709478. This protrusion would clash with Phe1007 in a π-helical configuration but align well with the α-helix. Yet, Phe1007 provides a favorable environment for accommodating the smaller cyclopropylphenyl group in TTA-A2, explaining the π form of S6II in Cav3.2-TA (Fig. 3a). As a result of the minor rotation of Phe1007, the I-II fenestration, present in the apo channel and Cav3.2-ACT, diminishes in Cav3.2-TA, and the gating residue on S6II shifts from Leu1010 to Val1011 in Cav3.2-TA, with the intracellular gate remaining closed (Figs. 1d, 3b; Supplementary information, Fig. S7).
TTA-P2 and ML218 bind through the II-III fenestration
TTA-P2 and ML218 also share a similar chemical structure, characterized by a 3,5-dichlorobenzamide head and an aliphatic tail (Fig. 4a). They also display a similar binding paradigm, with the head adhering to the II-III fenestration (Site F2), and the tail projecting into the cavity (site C) (Fig. 4b). Despite an ~30° deviation of the binding poses for the two head groups, the accommodation site within the fenestration is similar. The environment is primarily hydrophobic, enclosed by residues from S5II, P1II, S6II, and S6III segments, including Leu922 on S5II, Leu971 on P1II, Asn1003 and Phe1007 on S6II, Lys1503 in the P-loop, and Leu1539, Ser1543, Leu1547, and Phe1550 on S6III (Fig. 4c, d).
a Chemical structures of ML218 and TTA-P2. b The two molecules exhibit similar binding poses. Structures of Cav3.2-ML (with ML218) and Cav3.2-TP (with TTA-P2) are superimposed relative to the PD. Only the PD structure of Cav3.2-ML is shown as semi-transparent cartoon. c The binding poses of ML218 and TTA-P2 deviate with an ~30° rotation of the 3,5-dichlorobenzamide head within the II-III fenestration. Detailed coordination of ML218 and TTA-P2 in the II-III fenestration are presented in the middle and right panels, respectively. d Schematic representation of residues constituting the binding site for ML218 or TTA-P2 within a 4-Å cutoff distance. The binding pocket and potential H-bonds are indicated by a gray dashed contour and red dashed lines, respectively. e Coordination of the aliphatic tail of the two compounds in the central cavity of Cav3.2. f Both structures adopt a π configuration in the middle of the S6II segment.
The aliphatic tail of both TTA-P2 and ML218 reclines across the central cavity, directly obstructing ion permeation. The cavity site is formed by Leu415 on S1I, Phe1007 on S6II, and Ser1543, Leu1547, and Phe1550 on S6III (Fig. 4d, e) Similar to the conformation observed in the presence of TTA-A2, S6II adopts a π-helix conformation upon binding to TTA-P2 or ML218. On one end, the 3,5-dichlorobenzamide head group would encounter spatial repulsion with Leu1006 in an α-helix conformation; on the other end, the aliphatic tail could be further stabilized by Phe1007 through a π-H interaction, thus favoring the π-helix form (Fig. 4f).
Stabilization of antagonist binding by an endogenous lipid
Examination of the 3D EM maps for all the structures, including that of the apo channel, identifies a well-resolved density in the cavity, which can be best fitted with a phosphatidylethanolamine (PE) molecule (Fig. 5a, b; Supplementary information, Fig. S8). The two hydrophobic tails penetrate the III-IV fenestration, and the polar head inserts into the central cavity, directly contributing to drug coordination. Similar paradigm has been observed in drug-bound structures of Cav1.1, Cav3.1, and Cav3.3.22,44,47
a A conserved endogenous lipid stabilizes antagonist association within the central cavity. The EM densities for the lipid and antagonist are contoured at the similar level of 4.5–5 σ. The antagonists show more diverse docking poses in the absence of the endogenous lipid. Predicted binding poses in the absence of the lipid are represented as semi-transparent sticks, while the structure-determined binding pose is shown as solid sticks. b Identical binding pose of the lipid in all blocker-bound Cav3.2 structures. Only the pore-forming helices of Cav3.2-TA are shown in an extracellular view of the superimposed structures of the antagonist-bound pore cavity. c Less favored binding of antagonists to Cav3.2 in the absence of the lipid. Predicted binding poses in the absence of the endogenous lipid exhibit larger variability, as indicated by the increased RMSD values. The average binding free energy (ΔGbinding) was calculated using Prime-MM/GBSA.
To investigate the role of the lipid in drug binding, we performed molecular docking simulations with or without the lipid. The docking poses in the presence of the lipid align well with the experimental structures, with RMSD values of the predicted binding poses < 3.0 Å compared to the corresponding experimental structures. In contrast, in the absence of the lipid, the antagonists display more diverse docking poses, resulting in a broader range of RMSD and ΔGbinding values (Fig. 5a, c). Among these, flipped poses are even more favored for ACT-709478 and TTA-A2 in the absence of the lipid (Fig. 5a). Our computational analyses support the role of the lipid in stabilizing the accommodation of the antagonists in the cavity, yet the physiological relevance and the identity and specificity of the endogenous lipids that may facilitate drug binding awaits further characterizations.
Structural basis for ligand selectivity on T-type Cav channels
The four structures presented here, along with our previously reported Cav3.1-Z944 complex structure,44 offer important insights into T type-specific inhibition by these selective antagonists. Sequence alignment reveals that several residues involved in ligand binding, including Leu377, Gln973, Phe1007, Leu1010, Val1011, Lys1503, Leu1539, Leu1540, Ser1543, Val1546, Leu1547, Phe1556, Ser1805, Gln1848, Leu1851, Val1852, and Val1855, most of which are positioned on the P-loops and the S6 tetrahelical bundle, vary from those in the HVA Cav channels (Fig. 6a).
a Structure-guided sequence analysis to identify the determinants for antagonist selectivity. Sequence comparison of human Cav channels for the antagonist-binding residues is shown. The dashes represent residues in other subtypes that are identical to the corresponding ones in Cav3.2. b Functional validation of residues critical to TTA-A2 selectivity. Several residues in Cav3.2 were mutated to corresponding ones in the HVA channels. The responses of these mutants to TTA-A2 were examined through whole-cell patch-clamp recordings. c A magnified view of the coordination of TTA-P2 and ML218 in the superimposed structures of Cav3.2-TP and Cav3.2-ML. d F1007L confers Cav3.2 resistance to ML218, but not TTA-P2. Experimental details are provided in Materials and Methods and Supplementary information, Figs. S9–S11, and Tables S3 and S4. e Potential molecular basis for the distinct responses to TTA-P2 and ML218 by Cav3.2-F1007L. The additional fluorine atom in the piperidine ring of TTA-P2 may interfere with the essential π-H interaction between Phe1007 and the piperidine ring, potentially resulting in lower potency on Cav3.2, which is largely unaffected by the Leu substitution.
To identify the residues that underlie the subtype-specific sensitivity to these inhibitors, we started with in silico molecular docking and binding free energy calculation for Cav3.2 mutants each with a single locus substituted with the corresponding residue from the HVA channels (Supplementary information, Fig. S9a). The computational analysis suggests that many mutations would have distinct impact on different inhibitors. For example, L377M, Q1848A/S, and L1851V/I/M resulted in decreased binding energy for ACT-709478 and TTA-A2, but not TTA-P2 or ML218. On the other hand, F1007L reduces the affinity to TTA-A2 and ML218, but has little effect on ACT-709478 and TTA-P2 (Supplementary information, Fig. S9).
Based on these clues, we generated a number of corresponding Cav3.2 mutants and characterized their responses to the drugs using whole-cell patch-clamp recordings in HEK293T cells. Several mutations indeed attenuate the potency of the drugs, as exemplified by L377M, F1007L, Q1848A, L1851M and L1851I for TTA-A2, and F1007L for ML218. Consistent with the computational results, F1007L shows little effect on the potency of TTA-P2, which shares a similar binding pose to ML218 (Fig. 6b–d; Supplementary information, Figs. S9–S11 and Tables S3, S4).
The difference in F1007L’s responses to ML218 and TTA-P2 may be attributed to the additional fluorine atom in the piperidine ring of TTA-P2, which interferes with the π-H interaction between Phe1007 and the piperidine ring, leading to an ~30° deviation of the binding poses for the 3,5-dichlorobenzamide heads. Therefore, the potency of TTA-P2 on Cav3.2, regardless of the F1007L mutation, is lower than that of ML218 (Fig. 6c–e; Supplementary information, Fig. S9, Table S4). As there is no fluorine atom in the piperidine group of Z944, an analog of ML218 and TTA-P2, we also performed similar analysis on the complex structure of Z944-bound Cav3.1. Its overall binding pose in Cav3.1 is closer to that of ML218 in Cav3.2 than TTA-P2. Replacing the allelic Phe956 with Leu in Cav3.1 significantly reduces the potency of Z944 on Cav3.1 in a similar manner to that of ML218 on Cav3.2,44 thereby confirming the distinct sensitivity of Phe1007 in Cav3.2, or Phe956 in Cav3.1, in ligand recognition (Supplementary information, Figs. S9–S11 and Table S4).
It is noted that K1503F or K1503G, which only slightly affects the binding energy, does not show a significant impact on the potency of ML218 on Cav3.2 (Fig. 6d). A re-examination of the Z944 response by the corresponding Cav3.1 mutants, K1462F or K1462G, reveals some technical issues in the previous electrophysiological characterizations, which are summarized in the legend of Supplementary information, Table S4. Using a corrected protocol, we show that K1462F or K1462G does not reduce Cav3.1’s sensitivity to Z944 either (Supplementary information, Table S4).44
Taken together, our structural, computational, and functional analyses reveal the complexity underlying the determinants for the subtype specificity by selective inhibitors, and suggest that targeting these critical residues could be a viable strategy for designing selective T-type Cav channel blockers.
Discussion
Given its pivotal role in epilepsy and pain, Cav3.2 has emerged as a promising target for the development of anti-epileptics and analgesics.39,55 Dozens of pathological mutations, including those associated with autism, amyotrophic lateral sclerosis, hyperaldosteronism familial 4, and epilepsy, are directly linked to Cav3.2 (Supplementary information, Table S5).38,56,57,58,59,60 The high-resolution structures of Cav3.2 enable precise mapping of nineteen mutations located in the resolved regions. These mutations exhibit widespread distribution across various VSDs, ECLs, and the pore (Supplementary information, Fig. S12), establishing a structural foundation for understanding disease mechanisms.
The structures reported here and previously44,47 together reveal a common paradigm for T-type channel-selective pore blockers (Fig. 7a). These elongated molecules directly obstruct ion flow by occupying site C; meanwhile, the other end targets either site F2 (TTA-P2, ML218, Z944, pimozide, mibefradil, and otilonium bromide) or site F4 (ACT-709478 and TTA-A2) to stabilize the inactivated state. It is noted that the DHP compounds selectively target the III-IV fenestration (site F3) of Cav1 channels, which are also known as the DHP receptors,21 while sites F2, F4, and C appear to be the promising regions in the T-type channels for developing selective blockers. Further chemical modifications of critical residues hold the promise to alter drug sensitivity and selectivity, providing opportunities for precise modulation of Cav subtypes.
a Small-molecule T-type Cav blockers generally exhibit two binding poses within the cavity. Shown here are two perpendicular views of the superimposed PD of T-type Cav channels with inhibitors bound to the central cavity. TTA-A2 and ACT-709478 interact with the IV-I fenestration, while all others are associated with the II-III fenestration. b A common α-to-π transition of the S6II segment when accommodating different antagonists. Among the various resolved structures of T-type Cav channels in complex with antagonists, ACT-709478 is the only one that binds to the α form.
Except Cav3.2-ACT, which remains unchanged from the apo channel, a conserved α-to-π transition in the middle of the S6II segment of all T-type channels is found in the presence of all other antagonists (Fig. 7b). Such secondary structural transition is also frequently observed in Nav channels bound with pore blockers or even gating modifier toxins which bind to VSDs.61 The π-form S6 is usually associated with sealed fenestration(s), shrunk cavity volume, and contracted intracellular gate, physical features that all antagonize channel conductance. This observation suggests that the π-form structures might represent potentially more favored templates for Cav- or Nav-targeting drug design.
Last but not least, small molecule interactions within the scaffold of the spacious cavity have been manifested by the drug-drug interaction between the antiviral sofosbuvir/MNI-1 and the anti-arhythmic amiodarone in the PD of Cav1 channels.15,62 Endogenous lipids also represent an important class of small molecules. It is therefore not surprising that one or even more phospholipid molecules might interfere with antagonist binding by modifying the contour and chemical composition of the binding pocket. The role of lipids and other endogenous molecules, such as hormones and metabolites, in drug binding should also be investigated when targeting voltage-gated ion channels for drug discovery.
Materials and methods
Transient expression of human Cav3.2 in HEK293F cells
The codon-optimized cDNA of human CACNA1H for Cav3.2 isoform 1 (2353 residues, Uniprot Q95180-1) was synthesized by BGI Geneland Scientific Co., Ltd. To generate Cav3.2EM, residues 493-772 were deleted using a standard two-step PCR method. Both the full-length Cav3.2 and the truncated Cav3.2EM were subcloned into the pCAG vector with Twin-Strep-tag and Flag-tag in tandem at the amino terminus. All the plasmids for transient expression were confirmed through DNA sequencing. HEK293F suspension cells (Thermo Fisher Scientific, R79007) were cultured in FreeStyle 293 medium (Thermo Fisher Scientific) at 37 °C, supplied with 5% CO2 under 60% humidity. Transfection of the cells with plasmids was carried out when the cell density reached 1.5–2.0 × 106 cells/mL. For each 1-L cell culture, a mixture of 1.5 mg expression plasmids for Cav3.2 and 3 mg 40-kDa linear polyethylenimines (Polysciences) in 50 mL fresh medium was incubated for 15–30 min, and then added to the cell culture to achieve transient expression of the human Cav3.2 complex.
Preparation of Cav3.2 alone and drug-bound complexes
For one batch of protein purification, 15 L of transfected HEK293F cells were harvested ~48 h after transfection by centrifugation at 3600 × g for 10 min and resuspended in the lysis buffer containing 25 mM HEPES (pH 7.4), 150 mM NaCl, 2 mM CaCl2 and protease inhibitor cocktail (Selleckchem). After sonication on ice, the suspension was supplemented with n-dodecyl-β-D-maltopyranoside (Anatrace) to a final concentration of 1% (w/v), and cholesteryl hemisuccinate Tris salt (Anatrace) to 0.1% (w/v). After incubation at 4 °C for 2 h, the mixture was centrifuged at 16,000 × g for 45 min, and the supernatant was applied to anti-Flag M2 affinity gel (Sigma) for affinity purification. The resin was rinsed five times with the wash buffer (buffer W) that contains 25 mM HEPES (pH 7.4), 150 mM NaCl, 2 mM CaCl2, 0.06% GDN, and protease inhibitor cocktail. The target proteins were eluted with buffer W supplemented with 0.2 mg/mL Flag peptide (synthesized by GenScript). The eluent was concentrated using a 100-kDa cut-off Amicon filter unit (Millipore) and further purified through size-exclusion chromatography (Superose 6 10/300 GL, GE Healthcare) that was pre-equilibrated in the buffer containing 25 mM HEPES (pH 7.4), 150 mM NaCl, 2 mM CaCl2 and 0.02% GDN. Among the tested constructs of Cav3.2, Cav3.2EM afforded decent protein yield that was suitable for cryo-EM analysis. The peak fractions were pooled and concentrated to a final concentration of ~12 mg/mL.
To prepare drug-bound complexes, different drug (ACT-709478, TTA-A2, TTA-P2, or ML218) was added to concentrated Cav3.2 proteins at the final concentration of 1 mM, and the mixtures were incubated at 4 °C for 45 min before preparation of cryo-EM grids.
Cryo-EM sample preparation and data acquisition
The QuantiFoil grids (R1.2/1.3 300 mesh, Quantifoil) or UltrAuFoil grids (R1.2/1.3 300 mesh, Quantifoil) underwent glow-discharge using easiGlow (PELCO) at 15 mA for 15 s at 0.37 mBar. Prior to sample application, the Vitrobot Mark IV chamber was pre-cooled to 10 °C with 100% humidity. Subsequently, 3 μL of concentrated Cav3.2-Apo/ACT/TA/TP/ML sample was applied to the freshly treated grid surface, followed by blotting with filter paper for 4 s. The grid was then swiftly plunged into liquid ethane and stored in liquid nitrogen.
For automated data collection, the grids were loaded onto a 300 kV Titan Krios G4 (Thermo Fisher) equipped with a Falcon IV detector (Thermo Fisher). Micrographs were captured using EPU (Thermo Fisher) in Falcon IV EC mode at a nominal magnification of 75,000 ×, resulting in a calibrated pixel size of 1.036 Å. Each movie stack in EER format was exposed for 6 s, accumulating a total electron dose of ~40 e−/Å2. Subsequently, the movie stacks underwent alignment, summation, and dose-weighting using cryoSPARC live.63
Cryo-EM data processing
A total of 2,954/4,999/5,775/4,614/5,241 cryo-EM micrographs were collected for Cav3.2-Apo/ACT/TA/TP/ML, respectively. During cryoSPARC live preprocessing, patched CTF estimation was implemented. For the Cav3.2Apo dataset, 504 particles from 25 micrographs were manually picked to generate 2D templates through 2D classification. Subsequently, 1,812,275 particles were picked using the selected 2D templates through the template picker, and 2D classification was performed with bin-4 particles. 2D Class averages with different views were chosen as new references for the template picker. Two batches of picked particles underwent independent 2D classifications, and the well-classified particles were selected and merged. The Cav3.1 structure (EMD-0791) was low-pass filtered and, together with a decoy junk density, used for heterogeneous refinement with bin-4 particles. The best class was selected and re-extracted to bin-2 for multiple rounds of continuous heterogeneous refinement. Subsequently, 565,821 particles were re-extracted into bin1 and subjected to another three rounds of heterogeneous refinement, resulting in a 3.4 Å NU-refinement result. This map was applied as a new heterogeneous refinement reference, considering higher frequency information. Through additional rounds of heterogeneous refinement and a final clean-up round of 2D classification, 103,429 particles were selected, leading to a reconstruction at 3.0 Å. A similar processing workflow was applied to other ligand-bound Cav3.2 datasets, utilizing the apo-structure as the initial model.
Model building and refinement
The Cav3.2Apo initial model was automatically generated with ModelAngelo64 and subsequently subjected to manual examination and adjustments in COOT.65 Refinement against the corresponding map was then executed using the Real-space Refinement option in PHENIX.66 Additional structure optimization was conducted with ISOLDE,67 followed by a conclusive round of Real-space Refinement in PHENIX. The apo structure served as the new initial model for ligand-bound structures, employing the same model-building process. Detailed validation results for the model refinement are presented in Supplementary information, Table S2.
Whole-cell electrophysiology
HEK293T cells were cultured in Dulbecco’s Modified Eagle Medium (BI) supplemented with 4.5 mg/mL glucose and 10% (v/v) fetal bovine serum (BI). For patch-clamp recordings, the cells were plated onto glass coverslips and transiently co-transfected with the Cav3.2 variant plasmids and eGFP in the presence of lipofectamine 2000 (Invitrogen). Cells with green fluorescence were selected for patch-clamp recording 18–36 h after transfection. Experiments were performed at room temperature. No authentication was performed for the commercially available cell line. Mycoplasma contamination was not tested.
Whole-cell Cav3.2 Ca2+ currents were recorded in HEK293T cells using an EPC10-USB amplifier with Patchmaster software v2*90.2 (HEKA Elektronik), filtered at 3 kHz (low-pass Bessel filter) and sampled at 50 kHz. The borosilicate pipettes (Sutter Instrument) used in all experiments had a resistance of 2–4 MΩ and series resistance was compensated by 70%–80%. The electrodes were filled with the internal solution composed of 125 mM CsCl, 10 mM EGTA, 2 mM CaCl2, 1 mM MgCl2, 5 mM Na2ATP, 10 mM HEPES, pH adjusted to 7.4 with CsOH. The bath solutions contained 5 mM CaCl2, 166 mM TEA-Cl, 10 mM HEPES, pH adjusted to 7.4 with TEA-OH. For the long-term drug delivery tests, the internal and external solutions were altered. The borosilicate pipettes were filled with a solution consisting of 130 mM Cs-methanesulfonate, 10 mM TEA-Cl, 10 mM EGTA, 5 mM MgCl2, 5 mM Na-ATP, 10 mM HEPES (pH 7.4 with CsOH) with a resistance of 2–4 MΩ, whereas the bath solutions contained 110 mM CsCl, 40 mM TEA-Cl, 5 mM BaCl2, 1 mM MgCl2, 10 mM D-glucose, and 10 mM HEPES (pH 7.4 with TEA-OH). Mutants with small currents were recorded with 10 mM BaCl2 in the external solution instead of 5 mM. Data were analyzed using Fitmaster 2.90.5 (HEKA Elektronik), Origin (OriginLab), and GraphPad Prism (GraphPad Software).
The voltage dependence of ion current (I-V) was analyzed using a protocol consisting of steps from a holding potential of –100 mV for 200 ms to voltages ranging from –90 mV to +50 mV for 150 ms in 5 mV increments. The linear component of leaky currents and capacitive transients were subtracted using the P/4 procedure. For activation property analysis, the equation G = I/(V – Vr) was utilized, where Vr, the reversal potential, represents the voltage at which the current is zero. For the activation curves, conductance (G) was normalized and plotted against the voltage from –90 mV to 0 mV or 20 mV. For voltage-dependent steady-state inactivation, cells were clamped at a holding potential of –100 mV and applied to step pre-pulses from –110 mV to –10 mV for 1000 ms with an increment of 5 mV. Subsequently, the Ca2+ currents were recorded at the test pulse of –30 mV for 100 ms. The peak currents under the test pulses were normalized and plotted against the prepulse voltage. Activation and inactivation curves were fitted with a Boltzmann function to determine V1/2 and slope values. The time course of inactivation data from the peak current at –30 mV was fitted with a single exponential equation: y = A1 exp(−x/τinac) + y0, where A1 was the relative fraction of current inactivation, τinac was the time constant, x was the time, and y0 was the amplitude of the steady-state component.
To investigate the inhibition of Cav3.2 variants by different drugs, cells were held at –100 mV and stepped to –30 mV for 150 ms. Drugs were dissolved in dimethyl sulfoxide (final concentration < 0.1%, Sigma) to make a stock solution. Solutions with the indicated concentrations were freshly prepared and perfused to the recording cell for up to several minutes to achieve the maximal blockade using a multichannel perfusion system (VM8, ALA). Prior to drug perfusion, cells were recorded for 5 min to establish a stable peak current. The concentration-response curve was fitted using the equation Y = Bottom + (Top – Bottom)/(1 + 10(logIC50 – X) × Hill Slope), where IC50 represented the concentration of the drugs that blocked 50% of the current and X denoted the log of drug concentration, and Hill Slope indicated the slope factor.
Data are presented as mean ± standard error of the mean (SEM), with n representing the number of experimental cells from which recordings were obtained. Statistical significance was assessed using an unpaired t-test with Welch’s correction, one-way ANOVA analysis, and extra sum-of-squares F test.
Molecular docking and binding free energy calculation
The molecular docking analysis employed the Schrödinger Suite 2018-1 (Schrödinger, Inc.). Initial 3D configurations of small molecules were generated and optimized using the LigPrep program68 with the OPLS3 force field.69 The protein structure was prepared using default settings within Protein Preparation Wizard, utilizing the coordinates from the Cav3.2–drug complexes. Subsequently, molecular docking simulations were executed with the extra-precision docking method (Glide XP) in the Glide program. The top-scored binding pose aligns well with the binding pose resolved in the cryo-EM structure and was selected for further analysis. Subtype-specific residues within the drug-binding site were mutated based on sequence alignment to those in other subtypes. The Prime-molecular mechanics/generalized Born surface area (MM/GBSA) method was employed to calculate the relative binding free energy changes (ΔΔG) for each mutant compared to the WT, while keeping the ligand and other residues fixed.
Data availability
The data that support this study are available from the corresponding authors upon reasonable request. The cryo-EM maps have been deposited in the Electron Microscopy Data Bank (EMDB) under accession codes EMD-43991 (Cav3.2Apo), EMD-43995 (Cav3.2-ACT), EMD-43992 (Cav3.2-TA), EMD-43993 (Cav3.2-TP), and EMD-43994 (Cav3.2-ML). The coordinates have been deposited in the Protein Data Bank (PDB) under accession codes 9AYG (Cav3.2Apo), 9AYL (Cav3.2-ACT), 9AYH (Cav3.2-TA), 9AYJ (Cav3.2-TP), and 9AYK (Cav3.2-ML).
References
Catterall, W. A. Voltage-gated calcium channels. Cold Spring Harb. Perspect. Biol. 3, a003947 (2011).
Dolphin, A. C. A short history of voltage-gated calcium channels. Br. J. Pharmacol. 147(1), S56–S62 (2006).
McDonald, T. F., Pelzer, S., Trautwein, W. & Pelzer, D. J. Regulation and modulation of calcium channels in cardiac, skeletal, and smooth muscle cells. Physiol. Rev. 74, 365–507 (1994).
Clapham, D. E. Calcium signaling. Cell 131, 1047–1058 (2007).
Takahashi, M., Seagar, M. J., Jones, J. F., Reber, B. F. & Catterall, W. A. Subunit structure of dihydropyridine-sensitive calcium channels from skeletal muscle. Proc. Natl. Acad. Sci. USA 84, 5478–5482 (1987).
Van Goor, F., Zivadinovic, D., Martinez-Fuentes, A. J. & Stojilkovic, S. S. Dependence of pituitary hormone secretion on the pattern of spontaneous voltage-gated calcium influx. Cell type-specific action potential secretion coupling. J. Biol. Chem. 276, 33840–33846 (2001).
Olivera, B. M., Miljanich, G. P., Ramachandran, J. & Adams, M. E. Calcium channel diversity and neurotransmitter release: the omega-conotoxins and omega-agatoxins. Annu. Rev. Biochem. 63, 823–867 (1994).
Flavell, S. W. & Greenberg, M. E. Signaling mechanisms linking neuronal activity to gene expression and plasticity of the nervous system. Annu. Rev. Neurosci. 31, 563–590 (2008).
Li, B., Tadross, M. R. & Tsien, R. W. Sequential ionic and conformational signaling by calcium channels drives neuronal gene expression. Science 351, 863–867 (2016).
Ben-Johny, M. & Yue, D. T. Calmodulin regulation (calmodulation) of voltage-gated calcium channels. J. Gen. Physiol. 143, 679–692 (2014).
Halling, D. B., Aracena-Parks, P. & Hamilton, S. L. Regulation of voltage-gated Ca2+ channels by calmodulin. Sci. STKE 2005, re15 (2005).
Van Petegem, F., Chatelain, F. C. & Minor, D. L. Jr Insights into voltage-gated calcium channel regulation from the structure of the CaV1.2 IQ domain-Ca2+/calmodulin complex. Nat. Struct. Mol. Biol. 12, 1108–1115 (2005).
Suh, B. C., Leal, K. & Hille, B. Modulation of high-voltage activated Ca(2+) channels by membrane phosphatidylinositol 4,5-bisphosphate. Neuron 67, 224–238 (2010).
Yao, X. et al. Structures of the R-type human Ca(v)2.3 channel reveal conformational crosstalk of the intracellular segments. Nat. Commun. 13, 7358 (2022).
Gao, S. et al. Structural basis for human Ca(v)1.2 inhibition by multiple drugs and the neurotoxin calciseptine. Cell 186, 5363–5374.e16 (2023).
Dolphin, A. C. G protein modulation of voltage-gated calcium channels. Pharmacol. Rev. 55, 607–627 (2003).
Dolphin, A. C. Mechanisms of modulation of voltage-dependent calcium channels by G proteins. J. Physiol. 506(1), 3–11 (1998).
Wolfe, J. T., Wang, H., Howard, J., Garrison, J. C. & Barrett, P. Q. T-type calcium channel regulation by specific G-protein betagamma subunits. Nature 424, 209–213 (2003).
Catterall, W. A. et al. Voltage-gated ion channels and gating modifier toxins. Toxicon 49, 124–141 (2007).
Catterall, W. A., Lenaeus, M. J. & Gamal El-Din, T. M. Structure and pharmacology of voltage-gated sodium and calcium channels. Annu. Rev. Pharmacol. Toxicol. 60, 133–154 (2020).
Zhao, Y. et al. Molecular basis for ligand modulation of a mammalian voltage-gated Ca(2+) channel. Cell 177, 1495–1506.e12 (2019).
Gao, S. & Yan, N. Structural basis of the modulation of the voltage-gated calcium ion channel Ca(v) 1.1 by dihydropyridine compounds*. Angew Chem. Int. Ed. Engl. 60, 3131–3137 (2021).
Gao, S., Yao, X. & Yan, N. Structure of human Ca(v)2.2 channel blocked by the painkiller ziconotide. Nature 596, 143–147 (2021).
Neumaier, F., Dibue-Adjei, M., Hescheler, J. & Schneider, T. Voltage-gated calcium channels: Determinants of channel function and modulation by inorganic cations. Prog. Neurobiol. 129, 1–36 (2015).
Nowycky, M. C., Fox, A. P. & Tsien, R. W. Three types of neuronal calcium channel with different calcium agonist sensitivity. Nature 316, 440–443 (1985).
Ertel, E. A. et al. Nomenclature of voltage-gated calcium channels. Neuron 25, 533–535 (2000).
Catterall, W. A., Perez-Reyes, E., Snutch, T. P. & Striessnig, J. International union of pharmacology. XLVIII. Nomenclature and structure-function relationships of voltage-gated calcium channels. Pharmacol. Rev. 57, 411–425 (2005).
Doering, C. J. & Zamponi, G. W. Molecular pharmacology of high voltage-activated calcium channels. J. Bioenerg. Biomembr. 35, 491–505 (2003).
Perez-Reyes, E. Molecular characterization of T-type calcium channels. Cell Calcium 40, 89–96 (2006).
Arikkath, J. & Campbell, K. P. Auxiliary subunits: essential components of the voltage-gated calcium channel complex. Curr. Opin. Neurobiol. 13, 298–307 (2003).
Randall, A. D. & Tsien, R. W. Contrasting biophysical and pharmacological properties of T-type and R-type calcium channels. Neuropharmacology 36, 879–893 (1997).
Perez-Reyes, E. Molecular physiology of low-voltage-activated t-type calcium channels. Physiol. Rev. 83, 117–161 (2003).
Talley, E. M. et al. Differential distribution of three members of a gene family encoding low voltage-activated (T-type) calcium channels. J. Neurosci. 19, 1895–1911 (1999).
Francois, A. et al. The low-threshold calcium channel Cav3.2 determines low-threshold mechanoreceptor function. Cell Rep. 10, 370–382 (2015).
Bourinet, E. et al. Silencing of the Cav3.2 T-type calcium channel gene in sensory neurons demonstrates its major role in nociception. EMBO J. 24, 315–324 (2005).
Cai, S., Gomez, K., Moutal, A. & Khanna, R. Targeting T-type/CaV3.2 channels for chronic pain. Transl. Res. 234, 20–30 (2021).
Choi, S. et al. Attenuated pain responses in mice lacking Ca(V)3.2 T-type channels. Genes Brain Behav. 6, 425–431 (2007).
Heron, S. E. et al. Genetic variation of CACNA1H in idiopathic generalized epilepsy. Ann. Neurol. 55, 595–596 (2004).
Khosravani, H. et al. Gating effects of mutations in the Cav3.2 T-type calcium channel associated with childhood absence epilepsy. J. Biol. Chem. 279, 9681–9684 (2004).
Snutch, T. P. & Zamponi, G. W. Recent advances in the development of T-type calcium channel blockers for pain intervention. Br. J. Pharmacol. 175, 2375–2383 (2018).
Matar, N. et al. Zonisamide block of cloned human T-type voltage-gated calcium channels. Epilepsy Res. 83, 224–234 (2009).
Perez-Reyes, E., Van Deusen, A. L. & Vitko, I. Molecular pharmacology of human Cav3.2 T-type Ca2+ channels: block by antihypertensives, antiarrhythmics, and their analogs. J. Pharmacol. Exp. Ther. 328, 621–627 (2009).
Francois, A. et al. State-dependent properties of a new T-type calcium channel blocker enhance Ca(V)3.2 selectivity and support analgesic effects. Pain 154, 283–293 (2013).
Zhao, Y. et al. Cryo-EM structures of apo and antagonist-bound human Ca(v)3.1. Nature 576, 492–497 (2019).
Xiang, Z. et al. The discovery and characterization of ML218: A novel, centrally active T-type calcium channel inhibitor with robust effects in STN neurons and in a rodent model of Parkinson’s disease. ACS Chem. Neurosci. 2, 730–742 (2011).
Nam, G. T-type calcium channel blockers: a patent review (2012-2018). Expert Opin. Ther. Pat. 28, 883–901 (2018).
He, L. et al. Structure, gating, and pharmacology of human Ca(V)3.3 channel. Nat. Commun. 13, 2084 (2022).
Monteil, A. et al. Inhibition of Cav3.2 T-type calcium channels by its intracellular I-II loop. J. Biol. Chem. 290, 16168–16176 (2015).
Shcheglovitov, A. et al. Alternative splicing within the I-II loop controls surface expression of T-type Ca(v)3.1 calcium channels. FEBS Lett. 582, 3765–3770 (2008).
Weiss, N., Black, S. A., Bladen, C., Chen, L. & Zamponi, G. W. Surface expression and function of Cav3.2 T-type calcium channels are controlled by asparagine-linked glycosylation. Pflugers Arch. 465, 1159–1170 (2013).
Ficelova, V. et al. Functional identification of potential non-canonical N-glycosylation sites within Ca(v)3.2 T-type calcium channels. Mol. Brain 13, 149 (2020).
Orestes, P. et al. Reversal of neuropathic pain in diabetes by targeting glycosylation of Ca(V)3.2 T-type calcium channels. Diabetes 62, 3828–3838 (2013).
Wu, Q. et al. Structural mapping of Na(v)1.7 antagonists. Nat. Commun. 14, 3224 (2023).
Li, Z., Wu, Q. & Yan, N. A structural atlas of druggable sites on Na(v) channels. Channels 18, 2287832 (2024).
Becker, A. J. et al. Transcriptional upregulation of Cav3.2 mediates epileptogenesis in the pilocarpine model of epilepsy. J. Neurosci. 28, 13341–13353 (2008).
Vitko, I. et al. The I-II loop controls plasma membrane expression and gating of Ca(v)3.2 T-type Ca2+ channels: a paradigm for childhood absence epilepsy mutations. J. Neurosci. 27, 322–330 (2007).
Heron, S. E. et al. Extended spectrum of idiopathic generalized epilepsies associated with CACNA1H functional variants. Ann. Neurol. 62, 560–568 (2007).
Splawski, I. et al. CACNA1H mutations in autism spectrum disorders. J. Biol. Chem. 281, 22085–22091 (2006).
Rzhepetskyy, Y., Lazniewska, J., Blesneac, I., Pamphlett, R. & Weiss, N. CACNA1H missense mutations associated with amyotrophic lateral sclerosis alter Cav3.2 T-type calcium channel activity and reticular thalamic neuron firing. Channels 10, 466–477 (2016).
Stringer, R. N. et al. A rare CACNA1H variant associated with amyotrophic lateral sclerosis causes complete loss of Ca(v)3.2 T-type channel activity. Mol. Brain 13, 33 (2020).
Huang, G. et al. High-resolution structures of human Na(v)1.7 reveal gating modulation through α-π helical transition of S6(IV). Cell Rep. 39, 110735 (2022).
Yao, X. et al. Structural basis for the severe adverse interaction of sofosbuvir and amiodarone on L-type Ca(v) channels. Cell 185, 4801–4810.e13 (2022).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Jamali, K. et al. Automated model building and protein identification in cryo-EM maps. Nature https://doi.org/10.1038/s41586-024-07215-4 (2024).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D Struct. Biol. 75, 861–877 (2019).
Croll, T. I. ISOLDE: a physically realistic environment for model building into low-resolution electron-density maps. Acta Crystallogr. D Struct. Biol. 74, 519–530 (2018).
Sastry, G. M., Adzhigirey, M., Day, T., Annabhimoju, R. & Sherman, W. Protein and ligand preparation: parameters, protocols, and influence on virtual screening enrichments. J. Comput. Aided Mol. Des. 27, 221–234 (2013).
Harder, E. et al. OPLS3: a force field providing broad coverage of drug-like small molecules and proteins. J. Chem. Theory Comput. 12, 281–296 (2016).
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. A. & Sansom, M. S. HOLE: a program for the analysis of the pore dimensions of ion channel structural models. J. Mol. Graph 14, 354–360 (1996).
Acknowledgements
We thank the cryo-EM facility at Princeton Imaging and Analysis Center. This work is supported by the National Institutes of Health (23307-G0001-10013397-101), the National Natural Science Foundation of China (32330052), Beijing Frontier Research Center for Biological Structure, and Start-up funds from Tsinghua University. N.Y. was supported by the Shirley M. Tilghman endowed professorship from Princeton University in 2017–2022. X.F. has been supported by the HFSP long-term fellowship (LT000754/2020-L) from the International Human Frontier Science Program Organization (HFSPO) from 2020 to 2023.
Author information
Authors and Affiliations
Contributions
N.Y., J.H. and X.F. conceived the project. J.H., X.F., X.J., J.C., C.L., Q.G., T.L., A.D., P.L. and N.Y. designed experiments; J.H. and X.F. performed all experiments related to cryo-EM studies, including protein expression and purification; J.H. performed all computational simulations, including molecular docking and binding free energy calculations; X.J., J.C., C.L., Q.G. and T.L. carried out experiments related to electrophysiology; J.H., X.F., X.J. J.C., C.L., Q.G., T.L., A.D., P.L. and N.Y. analyzed the data; J.H., X.F., X.J., and N.Y. wrote the manuscript with input from all authors. All authors approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Huang, J., Fan, X., Jin, X. et al. Structural basis for human Cav3.2 inhibition by selective antagonists. Cell Res 34, 440–450 (2024). https://doi.org/10.1038/s41422-024-00959-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41422-024-00959-8
This article is cited by
-
A tale of two calcium channels: structural pharmacology of Cav2.1 and Cav3.2
Cell Research (2024)
-
Author Correction: Cryo-EM structures of apo and antagonist-bound human Cav3.1
Nature (2024)
-
Structural biology and molecular pharmacology of voltage-gated ion channels
Nature Reviews Molecular Cell Biology (2024)