Abstract
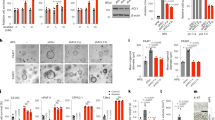
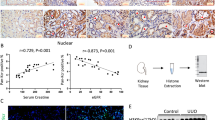
Posttranslational modifications add tremendous complexity to proteomes; however, gaps remain in knowledge regarding the function and regulatory mechanism of newly discovered lysine acylation modifications. Here, we compared a panel of non-histone lysine acylation patterns in metastasis models and clinical samples, and focused on 2-hydroxyisobutyrylation (Khib) due to its significant upregulation in cancer metastases. By the integration of systemic Khib proteome profiling in 20 paired primary esophageal tumor and metastatic tumor tissues with CRISPR/Cas9 functional screening, we identified N-acetyltransferase 10 (NAT10) as a substrate for Khib modification. We further showed that Khib modification at lysine 823 in NAT10 functionally contribute to metastasis. Mechanistically, NAT10 Khib modification enhances its interaction with deubiquitinase USP39, resulting in increased NAT10 protein stability. NAT10 in turn promotes metastasis by increasing NOTCH3 mRNA stability in an N4-acetylcytidine-dependent manner. Furthermore, we discovered a lead compound #7586-3507 that inhibited NAT10 Khib modification and showed efficacy in tumor models in vivo at a low concentration. Together, our findings bridge newly identified lysine acylation modifications with RNA modifications, thus providing novel insights into epigenetic regulation in human cancer. We propose that pharmacological inhibition of NAT10 K823 Khib modification constitutes a potential anti-metastasis strategy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Wang, Y., Zhang, J., Li, B. & He, Q. Y. Advances of proteomics in novel PTM discovery: applications in cancer therapy. Small Methods 3, 1900041 (2019).
Choudhary, C. et al. The growing landscape of lysine acetylation links metabolism and cell signalling. Nat. Rev. Mol. Cell Biol. 15, 536–550 (2014).
Narita, T., Weinert, B. T. & Choudhary, C. Functions and mechanisms of non-histone protein acetylation. Nat. Rev. Mol. Cell Biol. 20, 156–174 (2019).
Deng, G. et al. Loss of heterozygosity and p53 gene mutations in breast cancer. Cancer Res. 54, 499–505 (1994).
Johnson, L. et al. Somatic activation of the K-ras oncogene causes early onset lung cancer in mice. Nature 410, 1111–1116 (2001).
Birkbak, N. J. et al. Telomeric allelic imbalance indicates defective DNA repair and sensitivity to DNA-damaging agents. Cancer Discov. 2, 366–375 (2012).
Zhang, X. et al. Dissecting esophageal squamous-cell carcinoma ecosystem by single-cell transcriptomic analysis. Nat. Commun. 12, 5291 (2021).
Yang, Y. M. et al. Advances in targeted therapy for esophageal cancer. Signal Transduct. Target. Ther. 5, 229 (2020).
Bortolin-Cavaillé, M. L. et al. Probing small ribosomal subunit RNA helix 45 acetylation across eukaryotic evolution. Nucleic Acids Res. 50, 6284–6299 (2022).
Sharma, S. et al. Yeast Kre33 and human NAT10 are conserved 18S rRNA cytosine acetyltransferases that modify tRNAs assisted by the adaptor Tan1/THUMPD1. Nucleic Acids Res. 43, 2242–2258 (2015).
Sas-Chen, A. et al. Dynamic RNA acetylation revealed by quantitative cross-evolutionary mapping. Nature 583, 638–643 (2020).
Arango, D. et al. Acetylation of cytidine in mRNA promotes translation efficiency. Cell 175, 1872–1886.e24 (2018).
Zhang, H. et al. Ketogenesis-generated beta-hydroxybutyrate is an epigenetic regulator of CD8(+) T-cell memory development. Nat. Cell Biol. 22, 18–25 (2020).
Simithy, J. et al. Characterization of histone acylations links chromatin modifications with metabolism. Nat. Commun. 8, 1141 (2017).
Fang, Y. et al. Histone crotonylation promotes mesoendodermal commitment of human embryonic stem cells. Cell Stem Cell 28, 748–763.e7 (2021).
Huang, H. et al. Landscape of the regulatory elements for lysine 2-hydroxyisobutyrylation pathway. Cell Res. 28, 111–125 (2018).
Dominissini, D. & Rechavi, G. N(4)-acetylation of cytidine in mRNA by NAT10 regulates stability and translation. Cell 175, 1725–1727 (2018).
Xu, W. S., Parmigiani, R. B. & Marks, P. A. Histone deacetylase inhibitors: molecular mechanisms of action. Oncogene 26, 5541–5552 (2007).
Tang, X. et al. SIRT7 antagonizes TGF-β signaling and inhibits breast cancer metastasis. Nat. Commun. 8, 318–318 (2017).
MacPherson, L. et al. HBO1 is required for the maintenance of leukaemia stem cells. Nature 577, 266–270 (2020).
Li, X. et al. Deubiquitinase USP39 and E3 ligase TRIM26 balance the level of ZEB1 ubiquitination and thereby determine the progression of hepatocellular carcinoma. Cell Death Differ. 28, 2315–2332 (2021).
Wu, J. et al. USP39 regulates DNA damage response and chemo-radiation resistance by deubiquitinating and stabilizing CHK2. Cancer Lett. 449, 114–124 (2019).
Liu, X. et al. NAT10 regulates p53 activation through acetylating p53 at K120 and ubiquitinating Mdm2. EMBO Rep. 17, 349–366 (2016).
Arango, D. et al. Direct epitranscriptomic regulation of mammalian translation initiation through N4-acetylcytidine. Mol. Cell 82, 2797–2814.e11 (2022).
van Nes, J. et al. A NOTCH3 transcriptional module induces cell motility in neuroblastoma. Clin. Cancer Res. 19, 3485–3494 (2013).
Allfrey, V. G., Faulkner, R. & Mirsky, A. E. Acetylation and methylation of histones and their possible role in the regulation of RNA synthesis. Proc. Natl. Acad. Sci. USA 51, 786–794 (1964).
Zhang, H. et al. GSK-3beta-regulated N-acetyltransferase 10 is involved in colorectal cancer invasion. Clin. Cancer Res. 20, 4717–4729 (2014).
Liu, H. Y. et al. Acetylation of MORC2 by NAT10 regulates cell-cycle checkpoint control and resistance to DNA-damaging chemotherapy and radiotherapy in breast cancer. Nucleic Acids Res. 48, 3638–3656 (2020).
Tsai, K. et al. Acetylation of cytidine residues boosts HIV-1 gene expression by increasing viral RNA stability. Cell Host Microbe 28, 306–312.e6 (2020).
Majumder, S. et al. Targeting Notch in oncology: the path forward. Nat. Rev. Drug Discov. 20, 125–144 (2021).
Deng, L. et al. The role of ubiquitination in tumorigenesis and targeted drug discovery. Signal Transduct. Target. Ther. 5, 11 (2020).
Harrigan, J. A., Jacq, X., Martin, N. M. & Jackson, S. P. Deubiquitylating enzymes and drug discovery: emerging opportunities. Nat. Rev. Drug Discov. 17, 57–78 (2018).
Fioravanti, R. et al. Targeting histone acetylation/deacetylation in parasites: an update (2017–2020). Curr. Opin. Chem. Biol. 57, 65–74 (2020).
Xu, W. W. et al. Cancer cell-secreted IGF2 instigates fibroblasts and bone marrow-derived vascular progenitor cells to promote cancer progression. Nat. Commun. 8, 14399 (2017).
Shimada, Y. et al. Characterization of 21 newly established esophageal cancer cell lines. Cancer 69, 277–284 (1992).
Liao, L. et al. Anti-HIV drug elvitegravir suppresses cancer metastasis via increased proteasomal degradation of m6A methyltransferase METTL3. Cancer Res. 82, 2444–2457 (2022).
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Tan, X. P. et al. Lomerizine 2HCl inhibits cell proliferation and induces protective autophagy in colorectal cancer via the PI3K/Akt/mTOR signaling pathway. MedComm 2, 453–466 (2021).
Zheng, C. C. et al. Targeting PFKL with penfluridol inhibits glycolysis and suppresses esophageal cancer tumorigenesis in an AMPK/FOXO3a/BIM-dependent manner. Acta Pharm. Sin. B 12, 1271–1287 (2022).
Hu, H. F. et al. Anti-allergic drug azelastine suppresses colon tumorigenesis by directly targeting ARF1 to inhibit IQGAP1-ERK-Drp1-mediated mitochondrial fission. Theranostics 11, 1828–1844 (2021).
Xu, W. W. et al. IGF2 induces CD133 expression in esophageal cancer cells to promote cancer stemness. Cancer Lett. 425, 88–100 (2018).
Xu, W. W. et al. Direct targeting of CREB1 with imperatorin inhibits TGFβ2-ERK signaling to suppress esophageal cancer metastasis. Adv. Sci. 7, 2000925 (2020).
Xu, W. W. et al. Genome-wide identification of key regulatory lncRNAs in esophageal cancer metastasis. Signal Transduct. Target. Ther. 6, 88 (2021).
Hu, H. F. et al. Identification of miR-515-3p and its targets, vimentin and MMP3, as a key regulatory mechanism in esophageal cancer metastasis: functional and clinical significance. Signal Transduct. Target. Ther. 5, 271 (2020).
Acknowledgements
This work was supported by the National Key R&D Program of China (2021YFC2501000, 2021YFC2501900), the National Natural Science Foundation of China (82273141, 82073196, 31961160727, 81973339, 82071372), Key Laboratory of Guangdong Higher Education Institutes (2021KSYS009), the Outstanding Scholar Program of Bioland Laboratory (2018GZR110102002), and the Science and Technology Program of Guangzhou (202007030012). We thank Profs. Didier Trono and Vladislav Verkhusha for the plasmids obtained from Addgene.
Author information
Authors and Affiliations
Contributions
B.L. conceived and designed the study; L.L., Y.H., S.J.L., X.M.Y., Z.C.L., Y.Y.L., J.Y., G.G.Z., C.M.D., X.W., Y.D.Z., T.Y.X. and C.C.Z. acquired, analyzed and interpreted the data; L.L., Y.H., S.J.L. and X.M.Y. performed statistical analysis and drafted the manuscript; H.Y., C.C., A.L., Z.G.L. and J.B.L. provided technical and/or material support and critically revised the manuscript for important intellectual content; B.L. supervised the study. All authors edited and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
All animal experiments were approved by the Ethics Committee for Animal Experiments of Guangzhou Medical University (G2022-084), and mice were cared under standard conditions according to institutional guidelines. The human ESCC specimens were collected in accordance with the Declaration of Helsinki and was approved by the Ethics Committee of Shanghai Chest Hospital, Shanghai Jiao Tong University (No. KS(Y)22278). Informed consent was obtained from each participant.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liao, L., He, Y., Li, SJ. et al. Lysine 2-hydroxyisobutyrylation of NAT10 promotes cancer metastasis in an ac4C-dependent manner. Cell Res 33, 355–371 (2023). https://doi.org/10.1038/s41422-023-00793-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41422-023-00793-4
This article is cited by
-
Targeting the NAT10/NPM1 axis abrogates PD-L1 expression and improves the response to immune checkpoint blockade therapy
Molecular Medicine (2024)
-
Dissecting the oncogenic properties of essential RNA-modifying enzymes: a focus on NAT10
Oncogene (2024)
-
Bridging tissue repair and epithelial carcinogenesis: epigenetic memory and field cancerization
Cell Death & Differentiation (2024)
-
Alternative splicing and related RNA binding proteins in human health and disease
Signal Transduction and Targeted Therapy (2024)
-
N4-acetylcytidine modifies primary microRNAs for processing in cancer cells
Cellular and Molecular Life Sciences (2024)