Abstract
The AMP-activated protein kinase (AMPK) is a central regulator of energy homeostasis. Although much has been learned on how low energy status and glucose starvation activate AMPK, how AMPK activity is properly controlled in vivo is still poorly understood. Here we report that UHRF1, an epigenetic regulator highly expressed in proliferating and cancer cells, interacts with AMPK and serves to suppress AMPK activity under both basal and stressed conditions. As a nuclear protein, UHRF1 promotes AMPK nuclear retention and strongly suppresses nuclear AMPK activity toward substrates H2B and EZH2. Importantly, we demonstrate that UHRF1 also robustly inhibits AMPK activity in the cytoplasm compartment, most likely as a consequence of AMPK nucleocytoplasmic shuttling. Mechanistically, we found that UHRF1 has no obvious effect on AMPK activation by upstream kinases LKB1 and CAMKK2 but inhibits AMPK activity by acting as a bridging factor targeting phosphatase PP2A to dephosphorylate AMPK. Hepatic overexpression of UHRF1 showed profound effects on glucose and lipid metabolism in wild-type mice but not in those with the liver-specific knockout of AMPKα1/α2, whereas knockdown of UHRF1 in adipose tissue led to AMPK activation and reduced sizes of adipocytes and lipogenic activity, highlighting the physiological significance of this regulation in glucose and lipid metabolism. Thus, our study identifies UHRF1 as a novel AMPK gate-keeper with critical roles in cellular metabolism.
Similar content being viewed by others
Introduction
The AMP-activated protein kinase (AMPK), a heterotrimeric complex consisting of a catalytic subunit α and two regulatory subunits β and γ, is a central regulator of energy homeostasis and master sensor of glucose and the cellular energy status.1,2,3 Upon activation by low energy status or low glucose, AMPK reprograms cellular metabolism from ATP-utilizing anabolic processes such as gluconeogenesis and lipogenesis to ATP-generating catabolic pathways including fatty acid oxidation and glycolysis by phosphorylating numerous key metabolic proteins including acetyl-CoA carboxylase 1 (ACC1).1,2,3,4 In addition to directly regulating key metabolic proteins in the cytoplasm compartment, AMPK also has multiple critical nuclear functions and rewires cellular metabolism and stress response by targeting key transcriptional and epigenetic regulators, including FOXO3a, EZH2, and TET2, or by directly phosphorylating histone H2B.5,6,7,8 Activation of AMPK requires phosphorylation of Thr172 within the activation loop of AMPKα subunit (p-AMPK). Liver kinase B1 (LKB1) and calcium/calmodulin-dependent protein kinase kinase 2 (CAMKK2) are two primary upstream kinases accounting for the physiological phosphorylation of Thr172 of AMPKα.9,10,11,12 Recent studies have provided evidence that AMPK activation in response to glucose starvation occurs via a non-canonical pathway at the lysosome prior to the decline of cellular energy, suggesting that AMPK is also a surveillance kinase for nutrient stress.13,14,15,16 Several phosphatases are known to suppress AMPK activity through T172 dephosphorylation.17,18,19,20,21 Although nuclear AMPK can be activated by genotoxic damage and other stresses,8,22,23 how AMPK activity is properly controlled under both unstressed and stressed conditions and especially how nuclear AMPK activity is suppressed remain poorly understood.
Ubiquitin-like with plant homeodomain and RING finger domains 1 (UHRF1), also known as NP95 or ICBP90, is a multifunctional protein with a critical role in DNA methylation, DNA damage repair, and cell proliferation.24,25,26,27 Expression of UHRF1 highly correlates with cell proliferation and is markedly elevated in various cancer cells,28,29,30,31,32 suggesting a potential role of UHRF1 in the regulation of cell proliferation and/or cell metabolism. However, despite of a well-established essential role in DNA maintenance methylation, it is unknown if UHRF1 directly regulates cell metabolism.
Here, we demonstrate that UHRF1 functions as a gate-keeper of AMPK activity and controls glucose and lipid metabolism via control of AMPK activity. UHRF1 resides primarily in the nucleus but suppresses AMPK activity both in the cytoplasm and nucleus through dephosphorylating AMPK T-172 by recruiting phosphatase PP2A. UHRF1 is likely a key gate-keeper of nuclear AMPK activity, as we show that UHRF1 suppresses AMPK-catalyzed EZH2 and histone H2B phosphorylation. We also demonstrate with mouse models that UHRF1 controls glucose and lipid metabolism through AMPK. Thus, UHRF1 is not only an epigenetic regulator crucial for DNA maintenance methylation but also has novel functions in regulating cellular metabolism and in coordinating multiple epigenetic pathways through its ability to regulate both cytoplasmic and nuclear AMPK activity.
Results
UHRF1 interacts with and regulates AMPK activity
Because UHRF1 is highly expressed in proliferating cells and many different cancer cells, we hypothesized that UHRF1 may regulate the signaling pathway(s) that controls cell proliferation and/or metabolism. We immunoprecipitated ectopically expressed UHRF1 from HEK293T cells and found that endogenous AMPKα, the catalytic subunit of AMPK (the antibody used recognized both AMPKα1 and AMPKα2 and thus collectively referred to as AMPKα) co-precipitated with UHRF1 (Fig. 1a). We confirmed by immunoprecipitation (IP) and Western blot (WB) assays an interaction between ectopically expressed AMPKα1 and UHRF1 (Fig. 1b) as well as ectopically expressed AMPKα2 and UHRF1 (Fig. 1c). Furthermore, we found that endogenous UHRF1 could be co-immunoprecipitated with AMPKα (Fig. 1d).
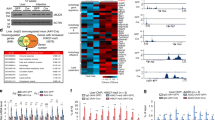
a HEK293T cells were transfected with FLAG-tagged UHRF1, immunoprecipitation was performed using anti-FLAG M2 beads and followed by immunoblotting with antibodies as indicated. F-UHRF1, FLAG-UHRF1. b, c Myc-tagged AMPKα1 (b) or AMPKα2 (c) was co-expressed with FLAG-tagged UHRF1 in HEK293T cells, and immunoprecipitation was performed using anti-FLAG M2 beads and followed by immunoblotting with antibodies as indicated. d AMPKα was immunoprecipitated from HEK293T cellular extracts and detected by immunoblotting using the indicated antibodies. e HEK293T cells were transfected with an increasing amount of HA-UHRF1 and immunoblotting was performed to detect the effect on T172-phosphorylated AMPKα. f HEK293T cells were transfected with Myc-AMPKα1 or Myc-AMPKα2 together with FLAG-tagged UHRF1, and whole-cell extracts were prepared for either direct immunoblotting of UHRF1 or immunoprecipitation of Myc-AMPKα1 or Myc-AMPKα2 using anti-Myc antibody, followed by immunoblotting using anti-T172-phosphorylated AMPKα antibody. g A549, HeLa, and MEF cells were infected with lentiviruses encoding UHRF1 for two days and processed for immunoblotting analyses using antibodies as indicated. h, i HEK293T (h), A549, HeLa, and Bel-7 cells (i) were treated with siRNAs against UHRF1 for three days and processed for immunoblotting analyses using antibodies as indicated. j MEF cells and mouse CGR8 ES cells were treated with control scramble siRNA (siCtrl) or siUhrf1 for three days and processed for immunoblotting analyses using antibodies as indicated. k HCT116 UHRF1flox/- cells were infected with Ad-Cre adenoviruses for three days to knock out the floxed UHRF1 allele and processed for immunoblotting using the indicated antibodies. l HEK293T cells were transfected with plasmids encoding wild-type UHRF1 or mutants with deletion of one of the functional domains for two days and processed for immunoblotting using antibodies as indicated. Note all deletion mutants suppressed AMPK activity like wild-type UHRF1. m HEK293T cells were transfected with plasmid encoding sgUHRF1 + Cas9 or sgDNMT1 + Cas9 and three days after transfection cells were collected for WB analysis or IF staining for 5mC. Scale bar, 50 μm.
Next, we asked whether UHRF1 plays a role in modulating AMPK activity. Ectopic expression of UHRF1 in HEK293T cells markedly diminished the levels of endogenous T172 phosphorylated, the active form AMPK (p-AMPK), in a dose-dependent manner without affecting the levels of AMPKα proteins (Fig. 1e). Consistent with this observation, we found that co-expression of UHRF1 with either AMPKα1 or AMPKα2 reduced their phosphorylated forms without affecting the protein levels (Fig. 1f). Furthermore, ectopic overexpression of UHRF1 was able to markedly reduce the levels of p-AMPK but not the levels of AMPKα in A549 lung cancer cells, HeLa cervical cancer cells and mouse embryonic fibroblast cells (MEFs) (Fig. 1g), indicating a ubiquitous role of UHRF1 in suppressing AMPK activation. Importantly, this inhibitory effect on AMPK is of metabolic consequences, as it also markedly reduced ACC1 phosphorylation (Ser79) and enhanced S6 phosphorylation (S235/S236), the respective biomarkers for AMPK-regulated fatty acid synthesis and protein synthesis1,2,3,4 (Fig. 1g). Conversely, UHRF1 knockdown in various cell lines all resulted in significant AMPK activation, as evident by elevated levels of p-AMPK without change in the levels of AMPKα proteins (Fig. 1h, i). Consistent with the observed activation of AMPK, UHRF1 knockdown resulted in a concomitant increased ACC1 phosphorylation and severely reduced S6 phosphorylation (Fig. 1h, i). Inhibition of AMPK by UHRF1 is not limited to cancer cells, as UHRF1 knockdown in MEFs or mouse ES cells all resulted in AMPK activation (Fig. 1j). In addition, inhibition of AMPK by UHRF1 was also confirmed by Cre recombinase-dependent knockout of UHRF1 in a HCT116 cell line carrying a null UHRF1 allele and floxed UHRF1 allele27 (Fig. 1k). Together these results establish UHRF1 as a universal negative regulator of AMPK activity.
UHRF1 regulates AMPK activity independent of epigenetic regulation and DNA damage
As UHRF1 is critically important for DNA methylation and also plays role in DNA damage repair,24,25,26,27 we next investigated if UHRF1 regulates AMPK activity directly or indirectly through DNA methylation or DNA damage repair. We first tested the ability of a series of UHRF1 deletion mutants to suppress AMPK activity when ectopically expressed in HEK293T cells. Interestingly, deletion of any one of the previously identified UHRF1 functional domains had no significant effect on inhibition of AMPK phosphorylation by UHRF1 (Fig. 1l). As the PHD, SRA, and Ring domains are all required for UHRF1-directed DNA maintenance methylation,24,33,34,35 these results indicate that UHRF1 regulates AMPK independent of its function in DNA methylation. Similarly, as the SRA and Ring domains are also required for UHRF1’s function in DNA repair,27,36 UHRF1 is unlikely to regulate AMPK through DNA damage stress. Consistent with this idea, we found that ectopic expression of ubiquitin E3 ligase activity-deficient UHRF1-H741A or UHRF1-3C/3 A mutants that are defective in DNA methylation function suppressed AMPK as effectively as wild-type UHRF1 (Supplementary information, Fig. S1a–c). In addition, treatment with DNA methylation inhibitor 5-Aza-2’-deoxycytidine (5-Aza-CdR) (Supplementary information, Fig. S1d, e) or knockdown of DNMT1 did not lead to any increase of AMPK activity or ACC1 phosphorylation (Supplementary information, Fig. S1f). Moreover, knockout of UHRF1 but not DNMT1 led to AMPKα activation, yet immunofluorescent staining confirmed that knockout of either UHRF1 or DNMT1 resulted in drastic DNA hypomethylation (Fig. 1m), indicating that UHRF1 regulates AMPK independent of DNA methylation. In further support of this idea, we observed that AMPKα was activated upon knockdown of UHRF1 in DNA methylation-deficient Dnmt1/3a/3b triple-knockout (Dnmt-TKO) mouse ES cells (Supplementary information, Fig. S1g). In addition, UHRF1 appeared to regulate AMPK activity independent of the cell cycle, as knockout of UHRF1 and overexpression of UHRF1 oppositely regulated AMPK activity under the condition with 24 h serum starvation that induced cell cycle arrest (Supplementary information, Fig. S1h, i). Altogether, we conclude that UHRF1 regulates AMPK activity independent of its function in DNA methylation and repair.
Characterization of the interaction between UHRF1 and AMPK and AMPK repression by UHRF1
We next aimed to define the interaction between UHRF1 and AMPK as well as the UHRF1 functional domain(s) required for the inhibition of AMPK activation. Consistent with the functional results in Fig. 1l, none of the previously characterized UHRF1 functional domains are essential for UHRF1 to interact with AMPK33 (Supplementary information, Fig. S2a, b). Both the N-terminal 1–300aa and 300–C fragments of UHRF1 were sufficient to inhibit AMPK activity (Supplementary information, Fig. S2c) and interact with both AMPKα1 and AMPKα2 (Supplementary information, Fig. S2d, e). The interaction with AMPK is likely required for UHRF1-mediated AMPK inhibition, as the UHRF1 N-terminal 1–133 aa fragment defective in AMPK interaction (Supplementary information, Fig. S2f) is inactive in inhibiting AMPK activity (Supplementary information, Fig. S2g). In vitro, GST pulldown assays demonstrated that both UHRF1-1–300 and UHRF1-300–C regions can directly interact with AMPKα1 (Supplementary information, Fig. S2h). Furthermore, in vitro GST pulldown demonstrated that the UHRF1-133–300 fragment is capable of binding AMPKα1 (Supplementary information, Fig. S2i). Thus, at least two regions within UHRF1 can interact with and repress AMPK activity, with 133–300 aa as one of the interaction domains.
UHRF1 suppresses basal as well as glucose starvation and other stress-induced AMPK activation
Having established that UHRF1 suppresses AMPK activity in various cells, we next investigated how UHRF1 influences AMPK function in response to glucose starvation and low energy stress in HEK293T or MEFs. The representative results in Figs. 2a, b show that knockdown of UHRF1 elevated the levels of phosphorylated AMPK under both glucose-rich (2.9-fold in HEK293T and 3.6-fold in MEFs) and glucose starvation conditions (from 4.1-fold to 8.1-fold in HEK293T and 3.7-fold to 9.2-fold in MEFs). Notably, a similar level of AMPK activation was observed when UHRF1 knockdown was compared to glucose starvation alone (2.9-fold vs 4.1-fold in HEK293T and 3.6-fold vs 3.7-fold in MEFs), highlighting a critical role of UHRF1 in suppressing AMPK activity under conditions without glucose starvation. Similar results were observed when experiments were carried out using wild-type and Uhrf1 null mouse ES cells (Fig. 2c). Furthermore, UHRF1 knockdown elevated AMPK activity under stress conditions with deprivation of all nutrients (glucose, glutamine, and serum) (Fig. 2d) or AICAR (an AMP analog that potentially activates AMPK) treatment37 (Fig. 2e). In contrast, we found that ectopic overexpression of UHRF1 significantly suppressed AMPK activation induced by either glucose starvation (Fig. 2f–g) or ER stress induced by Thapsigargin (Tg) treatment, which promotes Ca2+ release from the ER by inhibiting the ER Ca2+-ATPase pump38 (Fig. 2h), and the inhibition on AMPK activation correlated with inhibition of ACC1 phosphorylation (Fig. 2f–h). These gain- and loss-of-function experiments thus demonstrated the role of UHRF1 in suppressing both basal and stress-induced AMPK activity.
a, b HEK293T or MEF cells were transfected with shRNAs as indicated and cultured in the glucose-free medium for 4 h before being harvested for immunoblotting analysis. GS, glucose starvation. SE, short exposure; LE, long exposure. The relative levels of p-AMPK were shown through quantification of WB results using Image J, with the level of p-AMPK in the control setting as 1. This applied to all p-AMPK quantification in this figure. c Uhrf1−/− and Uhrf1+/+ mouse ES cells were deprived of glucose for 4 h before cells were harvested for immunoblotting analysis. d HEK293T were transfected with siRNA as indicated. Cells were cultured with medium free of glucose, glutamine, and serum for 4 h before being harvested for immunoblotting analyses using the indicated antibodies. EBSS, MEM with Earle’s Balanced Salt Solution without glucose, glutamine and serum. e HEK293T were transfected with siRNAs as indicated followed by treatment with 2 mM AICAR for 4 h. Cells were then harvested for immunoblotting analysis. f HEK293T cells transfected with control plasmid or plasmid expressing HA-UHRF1 were cultured in the glucose-free medium for 0, 1, 2, or 3 h before being harvested for immunoblotting with antibodies as indicated. The relative levels of p-ACC1 were shown as quantification of WB results using Image J, normalized to the level of p-ACC1 in the control. g NIH3T3 cells transfected with control plasmid or plasmid expressing FLAG-UHRF1 were cultured in the glucose-free medium for 0, 1, 3, or 5 h before being harvested for immunoblotting using antibodies as indicated. h HEK293T cells were transfected with plasmids as indicated. Cells were treated with Thapisgargin (Tg) for various times as indicated before being harvested for immunoblotting. i FLAG-UHRF1 was co-expressed with Myc-AMPKα1 or Myc-AMPKα2 in HEK293T cells as indicated. Cells were then treated with or without glucose starvation for 4 h before harvested for IP-WB analysis using antibodies as indicated. M-AMPKα1, Myc-AMPKα1; M-AMPKα2, Myc-AMPKα2. j HEK293T cells were transfected with Myc-AMPKα1 with or without FLAG-UHRF1 and harvested for IP-WB analysis. M-AMPKα1, Myc-AMPKα1. Note that Myc-AMPKα1 coimmunoprecipitated with FLAG-UHRF1 was devoid of T172 phosphorylation. k UHRF1 regulates ACC1 and S6 phosphorylation through AMPK. WB analysis showing that knockdown of UHRF1 elevated ACC1 phosphorylation and suppressed S6 phosphorylation and that ectopic overexpression of UHRF1 suppressed ACC1 phosphorylation and enhanced S6 phosphorylation in wild-type but not AMPKα1/α2-DKO MEFs. l Nile red (red) and DAPI (blue) staining showing knockdown of UHRF1 by siRNA markedly reduced the levels of lipid droplets in wild-type but not in AMPKα1/α2-DKO MEF cells. Scale bar, 10 μm. Three independent experimental data were quantified using Image J. m Oil red (red) and DAPI (blue) staining showing knockdown of UHRF1 by siRNA markedly reduced the levels of lipid droplets in HEK293T and A549 cells. Scale bar, 50 μm for HEK293T and 20 μm for A549.
Having determined that UHRF1 also suppresses stress-induced AMPK activation, we wondered whether the interaction between UHRF1 and AMPK could be regulated by stress. We carried out co-IP experiments with cells transfected with FLAG-UHRF1 and or Myc-AMPKα1 or Myc-AMPKα2 and treated with or without 4 h glucose starvation. The representative results in Fig. 2i revealed that the interaction between UHRF1 and AMPK is reduced under glucose starvation, suggesting that this interaction is dynamic and influenced by cellular glucose status. Indeed, further analysis revealed that the UHRF1-bound AMPKα proteins were essentially dephosphorylated (Fig. 2j), in agreement with the role of UHRF1 in suppressing AMPK activation.
UHRF1 regulates cellular lipid metabolism and glycolysis through AMPK
Having demonstrated that UHRF1 regulates AMPK activity and ACC1 phosphorylation, we next examined whether UHRF1 regulates lipid metabolism and glycolysis, as these processes are controlled by AMPK and ACC1.37,38,39 First, we examined if UHRF1 regulates ACC1 phosphorylation via AMPK. To this end, we found that UHRF1-knockdown-induced ACC1 phosphorylation and UHRF1-overexpression-suppressed ACC1 phosphorylation were all abolished in AMPKα1/α2 double knockout (DKO) MEF cells (Fig. 2k). Similarly, UHRF1 had no regulatory effect on S6 phosphorylation in AMPKα1/α2-DKO MEF cells (Fig. 2k). These results suggest that UHRF1 most likely regulates cell metabolism including lipid metabolism through its effect on AMPK. To test this directly, we performed Nile red staining on control wild-type MEFs and AMPKα1/α2-DKO MEF cells with or without treatment with siUHRF1. The image and quantitative results in Fig. 2l showed that UHRF1 knockdown by siRNA dramatically reduced the levels of lipid droplets in wild-type but not in AMPKα1/α2-DKO MEF cells. In addition, oil red staining assay revealed that knockdown of UHRF1 in HEK293T and A549 cells all resulted in substantial reduction of lipid droplet contents (Fig. 2m), suggesting a broad role for UHRF1 in regulating lipid metabolism via regulation of AMPK.
As AMPK is known to regulate the expression of glycolytic genes,39,40 we performed a quantitative RT-PCR analysis of glycolytic-related genes in both wild-type MEFs and AMPKα1/α2-DKO MEF cells with or without UHRF1 knockdown. This analysis revealed that UHRF1 knockdown significantly reduced the expression of glycolytic genes in wild-type but not AMPKα1/α2-DKO MEF cells (Supplementary information, Fig. S3a), whereas ectopic UHRF1 expression had an opposite effect primarily in wild-type but not AMPKα1/α2-DKO MEF cells (Supplementary information, Fig. S3b). Consistent with the observed alteration in glycolytic gene expression, UHRF1 overexpression elevated the levels of lactate accumulation in media of wild-type MEF culture but not in AMPKα1/α2-DKO MEF culture (Supplementary information, Fig. S3c). Previous studies indicated that AMPK regulates glycolytic gene expression through HIF1α,39 possibly indirectly through the inhibition of mTORC1.41 Consistently, we observed that knockdown of UHRF1 led to AMPK activation and a concomitant marked reduction of HIF1α protein levels in HEK293T and A549 cells (Supplementary information, Fig. S3d, e). Importantly, UHRF1 regulates HIF1α protein level in an AMPK-dependent manner, as neither UHRF1 knockdown nor UHRF1 ectopic expression significantly altered HIF1α protein levels in AMPKα1/α2-DKO MEF cells (Supplementary information, Fig. S3f, g).
Nuclear UHRF1 robustly suppresses AMPK activity in both nucleus and cytoplasm
UHRF1 is mainly a nuclear protein24,32 and AMPK is primarily localized and activated in lysosome and cytoplasm,42 raising the question in which subcellular compartment UHRF1 suppresses AMPK activity. We thus knocked down or overexpressed UHRF1 in different cells and performed subcellular fractionation to determine AMPK activities in cytoplasmic and nuclear compartments. Representative results (Fig. 3a, b and Supplementary information, Fig. S4a) showed that UHRF1 was indeed detected nearly exclusively in the nuclear fraction and that the majority of AMPK is found in the cytoplasmic fraction. However, knockdown of UHRF1 in these cells resulted in increased levels of T172-phosphorylated, active AMPK in both nuclear and cytoplasmic fractions and under both non-stress and glucose starvation condition (Fig. 3a, b and Supplementary information, Fig. S4a). In agreement with activation of nuclear AMPK upon UHRF1 knockdown, WB analysis showed elevated levels of Thr311-phosphorylated EZH2 (p-EZH2T311) and Ser36-phosphorylated histone H2B (p-H2BS36), two previously reported AMPK nuclear substrates,6,8 upon knockdown of UHRF1 (Fig. 3c and Supplementary information, Fig. S4b). A role of UHRF1 in suppression of nuclear AMPK activity became more evident under the condition of UHRF1 overexpression, as ectopic UHRF1 expression led to a significant increase of nuclear AMPKα and a concomitant decreased cytoplasmic AMPKα pool in MEF cells, yet the level of active nuclear AMPKα increased only marginally under both non-stress or glucose starvation condition (Fig. 3d). Similarly, ectopic expression of UHRF1 led to a variable increased level of nuclear AMPKα and a concomitant decreased level of cytoplasmic AMPKα in HEK293T and NIH3T3 cells (Fig. 3e, f), yet the levels of active nuclear AMPKα (p-AMPK) were either unaltered (in HEK293T cells) or even reduced (in NIH3T3 cells). Ectopic overexpression of UHRF1 also markedly inhibited AICAR-induced AMPK activation, AMPK-catalyzed EZH2 T311 phosphorylation, and H2B S36 phosphorylation (Fig. 3g), further demonstrating the role of UHRF1 in suppressing nuclear AMPK activity. Importantly, we found the regulation of UHRF1 on EZH2 and H2B phosphorylation is dependent on AMPK, as this effect is abolished in AMPKα-DKO MEFs (Fig. 3h). Together these results suggest that nuclear UHRF1 proteins contribute to AMPK nuclear retention and robust suppression of nuclear AMPK kinase activity.
a, b Immunoblotting analysis showing knockdown of UHRF1 activated both cytoplasmic and nuclear AMPK in HEK293T (a) and MEFs (b). HEK293T cells were transfected with siCtrl or siUHRF1, and cultured in glucose-containing or glucose-free medium for 4 h, followed by subcellular fractionation and immunoblotting analysis. MEF cells were transfected with siRNA against Uhrf1, followed by fractionation and immunoblotting using the indicated antibodies. The relative levels of p-AMPK were shown through quantification of WB results using Image J. For whole-cell extracts, the level of p-AMPK in the control cells without glucose starvation was set as 1, and the results were shown as relative p-AMPK vs AMPK. For cytosol and nuclear fractions, the level in the control cytosol was designated as 1, and the results were shown as relative p-AMPK vs AMPK. c UHRF1 controls AMPK nuclear substrates EZH2 and H2B phosphorylation. HEK293T cells were transfected with siCtrl or siUHRF1, and were then cultured in glucose-free medium for 4 h, followed by immunoblotting analysis. d Immunoblotting analysis showing ectopic UHRF1 expression suppressed glucose starvation-induced activation of cytoplasmic and nuclear AMPK. MEF cells were infected with control or UHRF1 encoding lentiviruses, and cultured in glucose-containing or glucose-free medium for 4 h, followed by subcellular fractionation and immunoblotting analysis. The quantification was completed as in a. SE, short exposure; LE, long exposure for all in this figure. e, f Immunoblotting analysis showing ectopic UHRF1 expression suppressed glucose starvation-induced activation of cytoplasmic and nuclear AMPK in HEK293T (e) and NIH3T3 cells (f). HEK293T and NIH3T3 cells were transfected with control or UHRF1 expressing plasmid for two days, and cultured in glucose-free medium for 4 h before being harvested for subcellular fractionation and immunoblotting analysis. g Immunoblotting analysis showing ectopic UHRF1 expression suppressed AICAR-induced AMPK activation and phosphorylation of H2BS36 and EZH2. MEF cells were infected by lentiviruses expressing control vector or UHRF1, 48 h after viral infection cells were treated with or without 2 mM AICAR for 4 h, followed by preparation of whole-cell extracts for immunoblotting analyses. h Immunoblotting analysis showing UHRF1 regulates EZH2 and H2BS36 phosphorylation in AMPK-dependent manner. Control and AMPKα1/α2-DKO MEF cells were treated with control or siUhrf1 for two days before harvested for immunoblotting analysis. i, j Live cell imaging showing UHRF1 increased AMPK nuclear occupancy. Representative live cell images of Hela cells expressing either mCherry-AMPKα1 alone or both GFP-UHRF1 and mCherry-AMPKα1 (i) and quantitative results of the relative ratio of nuclear vs cytoplasmic AMPK red fluorescence intensity in HeLa cells expressing mCherry-AMPKα1 alone or both mCherry-AMPKα1 and GFP-UHRF1 (j). Scale bar, 5 μm. Values were measured from ten different representative cells (n = 10). The results represent the means ± SD. two-tailed, paired t-test. k, l Photobleaching (FRAP) experiments showing UHRF1 enhanced AMPK nuclear retention. Representative images show the difference of mCherry-AMPKα1 fluorescence recovery in HeLa cells expressing mCherry-AMPKα1 plus GFP or mCherry-AMPKα1 plus GFP-UHRF1 after photobleaching of nuclear mCherry-AMPKα1 (k) and the corresponding data were quantified (l). Scale bar, 5 μm. The quantitative results represent the means ± SD of three independent experiments, two-tailed, paired t-test.
The ability of UHRF1 in suppressing cytoplasmic AMPK activity is likely explained by shuttling of AMPKα between cytoplasm and nucleus43 and by increased retention of AMPKα in the nucleus, presumably as a result of AMPKα-UHRF1 interaction. In support of this idea, live-cell imaging experiments showed that ectopically expressed UHRF1 led to increased levels of nuclear AMPKα (Fig. 3i, j). Furthermore, photobleaching experiments showed that UHRF1 markedly promoted the recovery of nuclear AMPKα1 (Fig. 3k, l). We observed more mCherry-AMPKα1 molecules moving into the nucleus when co-expressed with UHRF1 and mCherry-AMPKα1 molecules shuffle between cytoplasm and nucleus as dot foci (Supplementary information, videos S1–4). On the contrary, knockdown of UHRF1 slowed down the fluorescent recovery of nuclear GFP-AMPKα1 (Supplementary information, Fig. S4c). These data suggest a working model that UHRF1 mainly regulates AMPK activity in the nuclear compartment, but is able to control global AMPK activity as a consequence of AMPK dynamic shuttling between cytoplasm and nucleus.
UHRF1 does not appear to suppress AMPK activation through upstream kinases
To elucidate the mechanism by which UHRF1 suppresses AMPK activity, we first asked if UHRF1 suppresses AMPKα activation by inhibiting LKB110,44 and/or CAMKK2,11,45 two major upstream kinases that activate AMPK by phosphorylating T172 in the activation loop of AMPKα. A549 lung cancer cell line is deficient of LKB1 activity.46 The repression of AMPK activity in A549 cells by UHRF1 (Fig. 1g–i) argued for a LKB1-independent mechanism. Consistent with this idea, knockdown of LKB1 in HEK293T and HCT116 cells had no obvious effects on UHRF1 knockdown-induced AMPK activation (Supplementary information, Fig. S5a, b). Similarly, treatment with STO-609, a CAMKK2-specific inhibitor,47 also had no effect on UHRF1 knockdown-induced AMPK activation in these cells (Supplementary information, Fig. S5c, d). However, UHRF1 knockdown-induced AMPK activation was severely impaired if we inactivated both LKB1 and CAMKK2 with siLKB1 and STO-609 (Supplementary information, Fig. S5e, f), indicating that in these cells either LKB1 or CAMKK2 can activate AMPK. On the other hand, STO-609 treatment alone was sufficient to block UHRF1 knockdown-induced AMPK activation in A549 cells (Supplementary information, Fig. S5g), consistent with the fact that A549 cells are deficient of LKB1.46 Taken together, these results provide evidence that UHRF1 is unlikely to suppress AMPK activity via inhibition of LKB1 or CAMKK2, although these kinases are required for AMPK activation upon UHRF1 knockdown.
The ATM kinase was shown to mediate AMPK activation by phosphorylating AMPK at T172.48 UHRF1 knockdown could cause DNA damage and led to activation of ATM kinase, raising the possibility that UHRF1 knockdown may activate AMPK via ATM. We found that knockdown of UHRF1 led to elevated levels of phosphorylated H2AX (γ-H2AX), a surrogate marker of DNA damage, in A549 (Supplementary information, Fig. S5g) but not in HCT116 cells (Supplementary information, Fig. S5h). Nevertheless, treatment with ATM inhibitor Ru-5593349 did not affect AMPK activation upon UHRF1 knockdown in both A549 and HCT116 (Supplementary information, Fig. S5g, h), indicating that UHRF1 suppresses AMPK activation independent of ATM.
UHRF1 suppresses AMPK activation by promoting PP2A-dependent dephosphorylation
We next tested whether UHRF1 suppresses AMPK activity by enhancing AMPKα dephosphorylation. MEF cells without or with ectopic UHRF1 overexpression were first cultured in glucose-free medium for 4 h to induce robust AMPK activation. The cells were then switched to the medium with 25 mM glucose for various times. Subsequent WB analysis indicated that ectopic UHRF1 overexpression led to a much more rapid AMPKα dephosphorylation than control cells (Fig. 4a, b). Re-emergence of AMPK activation at 1.5 h was likely due to depletion of glucose at this time point. This result suggests that UHRF1 is able to promote AMPKα dephosphorylation.
a MEF cells were infected with lentiviruses expressing control vector or UHRF1 and 48 h after viral infection cells were cultured in glucose-free medium for 4 h to induce AMPK activation. Cells were then switched to the medium with 25 mM glucose for various times, followed by immunoblotting analysis. The relative levels of p-AMPK were determined using the level of p-AMPK in 0 h samples for both control and UHRF1-expressing cells as 1. b Quantitation of p-AMPK levels upon glucose replenish in (a). The results represent the means ± SD of three independent experiments, two-tailed, paired t-test. c HEK293T cells were transfected with indicated plasmids and after 48 h cells were harvested for IP-WB analysis using antibodies as indicated. M-AMPKα1 as Myc-AMPKα1 and M-AMPKα2 as Myc-AMPKα2. d, e HEK293T cells were transfected with FLAG-WIP1 (d) or FALG-PPM1E (e) together with FLAG-UHRF1 and Myc-AMPKα1 plasmids, and after 48 h cells were harvested for IP using Myc antibody and WB using antibodies as indicated. f HEK293T whole-cell extracts were immunoprecipitated with anti-PPP2CA antibody and followed by immunoblotting analysis using antibodies as indicated. g MEF cells were infected by lentiviruses expressing control vector or UHRF1. 48 h after viral infection, cells were harvested for IP-WB analysis using antibodies as indicated. h, i HEK293T cells were transfected with plasmids in the presence or absence of siUHRF1 as indicated. 48 h post-transfection the cells were harvested for IP-WB analysis using antibodies as indicated. Note that knockdown UHRF1 markedly diminished the interaction between PPP2CA and AMPKα1 (h) and the interaction between PPP2CA and AMPKα2 (i). j HEK293T cells were treated with siRNAs as indicated and were harvested for WB analysis using antibodies as indicated. k HEK293T cells were transfected with control vector or UHRF1 and siRNAs as indicated. Cells were harvested 48 h post-transfection for WB analysis using antibodies as indicated. l HEK293T cells were transfected with FLAG-PPP2CA and/or siUHRF1 as indicated. Cells were harvested 48 h post-transfection for WB analysis using antibodies as indicated. m HEK293T cells were transfected with indicated siRNAs and were harvested 48 h post-transfection for WB analysis using antibodies as indicated.
Several phosphatases including PP1, PP2A, PP2C, PPM1D (WIP1), and PPM1E have been shown to dephosphorylate AMPKα in vitro or in vivo.17,18,19,20,21 We confirmed that ectopic expression of the catalytic subunits of these phosphatases in HEK293T cells could lead to substantial dephosphorylation of endogenous AMPKα (Supplementary information, Fig. S6a). Among these phosphatases, PPP2CA (catalytic subunit of PP2A), PPM1D, and PPMIE were mainly localized in the nucleus (Supplementary information, Fig. S6b) and also interacted with UHRF1 in co-IP assay (Supplementary information, Fig. S6c, d). Importantly, UHRF1 appeared to bridge the interaction between AMPK and PPP2CA, as we found that UHRF1 overexpression dramatically enhanced the interaction of AMPKα1 and AMPKα2 with PPP2CA (Fig. 4c) but not PPM1D and PPM1E (Fig. 4d, e and Supplementary information, Fig. S6e, f). We also detected an interaction between endogenous PPP2CA and UHRF1 by immunoprecipitation of PPP2CA (Fig. 4f), and observed that ectopic UHRF1 expression enhanced the interaction of endogenous AMPKα and PPP2CA (Fig. 4g). Furthermore, we found that knockdown of UHRF1 markedly decreased the interaction of both AMPKα1 and AMPKα2 with PPP2CA (Fig. 4h, i), indicating that UHRF1 is required for bridging the interaction between AMPKα and PPP2CA. Moreover, both UHRF1-1–300 and 300–C fragments could promote the interaction between AMPKα1 and PPP2CA (Supplementary information, Fig. S6g, h), consistent with the observation that both the regions are capable of inhibiting AMPK activity. These results suggest that UHRF1 bridges the interaction selectively between AMPK and PPP2CA which does not appear to involve other phosphatases and thus may exploit a PP2A-dependent dephosphorylation mechanism to regulate AMPK activity.
To test if UHRF1 regulates AMPK activity through PP2A, we ectopically expressed UHRF1 in HEK293T cells that were first treated with siRNAs to knock down either PPP2CA or PPM1E. Although knockdown of either PPP2CA or PPM1E resulted in AMPK activation, we found that knockdown of PPP2CA consistently rendered a more robust activation (Fig. 4j). Importantly, PPP2CA but not PPM1E knockdown abrogated UHRF1 knockdown-induced AMPK activation (Fig. 4j) as well as UHRF1-overexpression-induced AMPK inactivation (Fig. 4k). Furthermore, UHRF1 knockdown abolished PPP2CA overexpression-induced AMPK dephosphorylation (Fig. 4l) as well as PPP2CA knockdown-induced AMPK phosphorylation (Fig. 4m), indicating that PP2A suppresses AMPK activation in a UHRF1-dependent manner. Subcellular fractionation experiments revealed that knockdown of PPP2CA led to activation of both cytoplasmic and nuclear fraction of AMPK (Supplementary information, Fig. S6i). Taken together, these results suggest that UHRF1 suppresses AMPK activity primarily by acting as a bridging factor between phosphatase PP2A and AMPK.
Characterization of the PP2A holoenzyme that dephosphorylates AMPK
To further characterize the interaction between UHRF1 and AMPK and PP2A, we performed gel filtration chromatography of 293 T nuclear extracts with or without ectopic expression of UHRF1 (Fig. 5a). Using HDAC3 as a molecular weight marker,50 which is known to exist as more than 1 million dalton corepressor complexes, we found that both endogenous and ectopic expressed UHRF1 distributed in a broad range of fractions, suggesting the existence of various UHRF1 complexes. Interestingly, AMPKα was found to show extensive co-fractionation with UHRF1, and furthermore, ectopic overexpression of UHRF1 led to a substantially increased level of co-fractionated AMPKα among fractions 3–17 (Fig. 5a), consistent with increased nuclear AMPKα retention upon UHRF1 overexpression. Despite of the markedly increased AMPKα upon UHRF1 overexpression, the levels of p-AMPK in these fractions only increased marginally, consistent with nuclear AMPK being mainly dephosphorylated. Co-fractionation of PPP2CA with UHRF1 was also observed, and ectopic overexpression of UHRF1 also led to an increased level of co-fractionated PPP2CA, although less pronounced compared to AMPKα. Although co-fractionation in gel filtration does not necessarily mean the existence of protein-protein interaction, we confirmed by subsequent IP-WB analysis of the peak co-fractionated fractions the existence of UHRF1-AMPK and UHRF1-PPP2CA interactions (Fig. 5b). Increased co-IP for both AMPKα and PPP2CA was observed in UHRF1 overexpression samples, consistent with our notion that UHRF1 interacts with both AMPKα and PPP2CA.
a Gel filtration analysis of nuclear extracts from control and UHRF1 overexpressed HEK293T cells with Superdex 200. The collected odd fractions were used for WB analysis. HDAC3 serves as a molecular weight marker for gel filtration. Note UHRF1 overexpression was associated with a substantial increase of AMPKα in various fractions, as a result of AMPKα nuclear accumulation. b The indicated fractions from gel filtration in a were used for IP-WB analysis using antibodies as indicated. c Identification of nuclear AMPKα2-associated proteins by mass spectrometry. HEK293T cells were transfected with plasmids encoding Myc-AMPKα2 and nuclear extracts were prepared 48 h post-transfection. Myc-AMPKα2 and associated proteins were purified by anti-Myc antibody beads and subjected to protein identification by mass spectrometry. Also shown are the numbers of unique peptides. d IP-WB analysis of UHRF1-associated PP2A subunits. FLAG-UHRF1 was expressed in HEK293T cells and the resulting nuclear extracts were used for IP-WB analysis using antibodies as indicated. e WB analyses showing the effect of knocking down PPP2R5D on UHRF1-regulated AMPK activation. f Immunofluorescent staining of HA-LAMP2-AMPKα1 in HeLa cells. Scale bar, 10 μm. g WB analysis of whole-cell extracts derived from HA-LAMP2-AMPKα1-expressing HEK293T cells. The relative ratio (6.5) of phosphorylated HA-LAMP2-AMPKα1 vs phosphorylated AMPKα was determined by Image J after adjusting the level of HA-LAMP2-AMPKα1 and AMPKα proteins. h Knockdown of UHRF1 activated endogenous AMPKα but not lysosome-anchored HA-LAMP2-AMPKα1. HEK293T cells were transfected with HA-LAMP2-AMPKα1 and treated with control siRNAs or siUHRF1 as indicated. 48 h after siRNA treatment, the cells were collected and subjected to WB analysis using either anti-AMPKα antibody (left panel) or anti-p-AMPKα antibody (middle panel). MEF cells were infected by lentiviruses expressing control vector or UHRF1. The relative activation was determined by Image J. i Ectopic overexpression of UHRF1 suppressed endogenous AMPKα but not lysosome-anchored HA-LAMP2-AMPKα1. The experiments were performed as in h.
As PP2A is a collection of various holoenzymes, we aimed to identify the specific PP2A holoenzyme(s) that interacts with AMPK. To this end, we isolated nuclear Myc-AMPKα2 complexes by immune-affinity purification and identified the associated proteins by IC-MS analysis. This analysis revealed the presence of UHRF1, PPP2CA, and the PP2A scaffold subunit PPP2R1A, in addition to the expected AMPK subunits such as AMPKβ2 and AMPKγ1 (Fig. 5c). Although we failed to identify a PP2A regulatory subunit in our IC-MS analysis of AMPKα2 associated proteins, a previous study reported that the regulatory subunit PPP2R5D is required for AMPK dephosphoryation by PP2A.51 Consistently, we detected PPP2R1A and PPP2R5D along with PPP2CA in immunoprecipitates of nuclear FLAG-UHRF1 (Fig. 5d). Importantly, knockdown of PPP2R5D essentially abrogated AMPK activation induced by knockdown of UHRF1 (Fig. 5e, left panel) and AMPK suppression induced by overexpression of UHRF1 (Fig. 5e, right panel). Together these data demonstrate that the PPP2CA/PPP2R1A/PPP2R5D holoenzyme is likely the major PP2A that catalyzes UHRF1-induced AMPK dephosphorylation, although the involvement of the other types of PP2A complexes could not be excluded. We also found that the interaction of AMPKα–UHRF1–PPP2CA was compromised under glucose starvation (Supplementary information, Fig. S6j, k), suggesting that UHRF1 mediated AMPK suppression could be partially relieved under nutrient-poor conditions to allow optimal AMPK activation.
Suppression of cytoplasm AMPK by UHRF1 is dependent on AMPK cytoplasm-nucleus shuttling
To test if AMPK cytoplasm-nucleus shuttling is required for suppression of AMPK activity in the cytoplasm compartment by UHRF1, we constructed a LAMP2-AMPKα1 fusion protein. LAMP2 is a lysosomal membrane protein and a previous study had successfully targeted AMPKβ to lysosomes by fusing LAMP2 to AMPKβ.42 We found the HA-LAMP2-AMPKα1 protein exhibited a typical lysosome staining pattern when ectopically expressed in HeLa cells (Fig. 5f). WB analysis using anti-AMPKα antibody detected HA-LAMP2-AMPKα1 as two distinct bands, in addition to the endogenous AMPKα (Fig. 5g, left panel). Interestingly, WB analysis using anti-p-AMPKα revealed that, in contrast to the endogenous AMPKα, the top full-length HA-LAMP2-AMPKα1 was extensively phosphorylated at T-172 (Fig. 5g, middle panel). Quantitative analysis indicated an approximately 6.5-fold more T-172 phosphorylation on HA-LAMP2-AMPKα1 than endogenous AMPKα (Fig. 5g, right panel). This observation indicates the lysosomal localized HA-LAMP2-AMPKα1 is highly activated, consistent with the previous result.42 The lower band HA-LAMP2-AMPKα1 is much less phosphorylated, and the reason for this difference is currently unknown.
Having established a lysosomal targeting of AMPKα1 by LAMP2, we next tested whether knockdown and overexpression of UHRF1 affected HA-LAMP2-AMPKα1 activity. As shown in Fig. 5h, we found that knockdown of UHRF1 activated endogenous AMPKα1 but not HA-LAMP2-AMPKα1. Conversely, ectopic overexpression of UHRF1 suppressed endogenous AMPKα but not HA-LAMP2-AMPKα1 under conditions of glucose starvation (Fig. 5i). Thus, UHRF1 is not able to suppress the AMPK activity if the AMPK proteins were immobilized to the lysosome, which prevents AMPK cytoplasm-nucleus shuttling.
UHRF1 regulates glucose and lipid metabolism in mice
Having demonstrated that UHRF1 has a profound effect on AMPK activity under both basal and stress conditions, we next investigated whether UHRF1 regulates AMPK activity and metabolism in physiological conditions. We first examined UHRF1 expression levels in mouse major metabolic organs, including liver, muscle, and adipose tissues. As shown in Supplementary information, Fig. S7a, the levels of UHRF1 mRNA were relatively lower in liver and muscle, and higher in adipose tissues. Based on this expression pattern, we explored the effect on AMPK activity and metabolism upon UHRF1 overexpression in the liver and knockdown in adipose tissues.
To overexpress UHRF1 in mouse liver, we first injected male C57BL/6 J mice with adenoviruses expressing UHRF1 (Ad-UHRF1) or control GFP (Ad-GFP) via tail vein. Subsequent WB analysis of mouse liver tissues confirmed overexpression of UHRF1 in Ad-UHRF1 compared to Ad-GFP injected mice (Fig. 6a). Importantly, UHRF1 overexpression in liver resulted in a marked reduction of phosphorylated AMPK and insignificant change of total AMPK (Fig. 6a). Compared to control Ad-GFP-injected mice, Ad-UHRF1-injected mice exhibited higher blood glucose levels at both fed and fasted states (Fig. 6b), as well as insulin resistance as revealed by glucose and insulin tolerance tests (Fig. 6c, d). Moreover, Ad-UHRF1-injected mice had elevated liver triacylglycerol (TG) contents (Fig. 6e) and this was confirmed by haematoxylin and eosin (H&E) and Oil Red O staining (Fig. 6f), suggesting that forced overexpression of UHRF1 led to increased hepatic lipid deposition and hepatosteatosis. Consistent with inhibition of AMPK activity in Ad-UHRF1-injected mouse liver tissues (Fig. 6a), elevated levels of expression were detected for key genes involved in gluconeogenesis (G6pc and Pepck) and triglyceride synthesis (Fasn) and transcription factor Srebp1c that controls lipogenic gene expression in Ad-UHRF1-injected mouse liver tissues (Fig. 6g). We further observed that overexpression of UHRF1 in primary hepatocytes by Ad-UHRF1 infection repressed AMPK activity and phosphorylation of ACC1 (Fig. 6h) and elevated TG contents (Supplementary information, Fig. S7b) and expression of gluconeogenic and lipogenic genes (Supplementary information, Fig. S7c), recapitulating the metabolic results observed in Ad-UHRF1-infected mouse livers.
a–g Male C57BL/6 J mice were injected with adenoviruses expressing UHRF1 (Ad-UHRF1) or GFP (Ad-GFP) via tail vein to overexpress UHRF1 in liver (n = 6). Mice were sacrificed at day 12 after Ad-UHRF1 or Ad-GFP injection, followed by WB analysis of UHRF1 and p-AMPK levels in liver tissues (a). The fed and fasted (8 h) blood glucose levels were measured at day 4 (b), and glucose tolerance tests (c) were performed at day 6 and insulin tolerance tests (d) at day 9. Liver samples were also used for TG content analysis (e), H&EH&E and Oil Red O staining (scale bar, 50 μm) (f), and mRNA expression analysis of genes related to glucose and lipid metabolism (g). h Primary hepatocytes were isolated from male C57BL/6 J mice and infected with Ad-GFP or Ad-UHRF1 and followed by WB analysis three days after viral infection. i Top diagram illustrating the generation of liver-specific UHRF1 overexpression transgenic mouse model. Lower panel showing WB analysis of liver tissues derived from control WT and UHRF1 overexpression (UHRF1-OE) mice using antibodies as indicated. WT and UHRF1-OE mice were euthanized after a 12-h overnight fasting, and liver tissues were dissected. j The blood glucose levels of random (fed) and fasted (8 h) WT and UHRF1-OE mice (n = 7). k, l Glucose tolerance tests (k) and insulin tolerance tests (l) of WT and UHRF1-OE mice (n = 7). m Liver TG levels of WT and UHRF1-OE mice (n = 7). n Representative H&E and Oil Red O staining showing TG accumulation in the liver of UHRF1-OE but not WT mice, scale bar, 50 μm. o mRNA levels of genes related to glucose and lipid metabolism in liver tissues of WT and UHRF1-OE mice (n = 7).
To test further the roles of UHRF1 in control of hepatic lipid and glucose metabolism, we generated a conditional liver-specific UHRF1 overexpression transgenic mouse model (Fig. 6i top panel). Consistent with Ad-UHRF1-mediated UHRF1 overexpression, we observed in this transgenic mouse model that liver-specific UHRF1 overexpression resulted in AMPK inactivation (Fig. 6i), increased serum glucose levels (Fig. 6j), moderate insulin resistance as revealed by glucose and insulin tolerance tests (Fig. 6k–i), elevated TG contents (Fig. 6m) and lipid deposition in liver (Fig. 6n), and increased expression of gluconeogenic and lipogenic genes (Fig. 6o). Taken together, both adenoviral and transgenic-based liver-specific UHRF1 overexpression in mice established further a role of UHRF1 in control of AMPK activity and revealed for the first time its profound effect on physiological glucose and lipid metabolism.
As UHRF1 has a relatively higher expression level in adipose tissue, we explored if UHRF1 regulates AMPK in adipose by injection of specific siUhrf1 unilaterally on one side of inguinal fat pad of high fat diet (HFD)-fed mice, and injection of scramble control siRNA on the other side of fat pad as control. Representative results (Fig. 7a) showed that knockdown of UHRF1 dramatically increased the levels of p-AMPK and p-ACC1 in adipose tissues. In addition, the sizes of adipocytes were decreased in siUhrf1-injected inguinal fat pad (Fig. 7b), accompanied with increased expression of lipolytic genes and reduced expression of lipogenic genes Fasn and Elovl6 without changes in inflammatory responses (Fig. 7c). Taken together, these data show in vivo that UHRF1 functions as a critical suppressor of AMPK and affects energy metabolism in both liver and adipose tissues.
a–c C57BL/6 J mice with 4 weeks of HFD were injected with siNC or siUhrf1 into the inguinal fat pads for 2 days (n = 5). Inguinal adipose tissues were isolated and processed for WB analysis of UHRF1, p-AMPK and p-ACC1 levels (a), H&E staining (b, left panel, scale bar, 50 μm) and quantitative measurement of adipocyte sizes (b, right panel), and qRT-PCR analysis of mRNA levels of genes related to lipolysis, lipogenesis, and macrophage (c). All results are shown as means ± SEM. *P < 0.05, **P < 0.01, n.s., not significant. d–h, AMPKαDKO mice were injected with Ad-UHRF1 or Ad-GFP via tail vein to overexpress UHRF1 in the liver (n = 3). The fed and fasted (8 h) blood glucose levels were measured at day 4 (d). Glucose tolerance tests (e) and insulin tolerance tests (f) were performed at day 6 and day 9, respectively. Mice were sacrificed at day 12 after Ad-UHRF1 or Ad-GFP injection and liver samples were used for TG content analysis (g) and mRNA expression analysis of genes related to glucose and lipid metabolism (h). i Working model illustrating how nuclear UHRF1 controls AMPK activity. UHRF1 resides primarily in the nucleus and promotes AMPKα dephosphorylation by PP2A. AMPKα cycles between nucleus and cytoplasm via protein trafficking and its nuclear retention regulated by UHRF1 leads to inactivation of cytoplasmic AMPK.
UHRF1 regulates glucose and lipid metabolism in mice through AMPK
To investigate whether UHRF1 regulates hepatic metabolic homeostasis through inhibition of AMPK, we made use of liver-specific AMPKα1/α2 knockout (AMPKα-LKO) mouse model.52 AMPKα-LKO mice were injected with Ad-UHRF1 or Ad-GFP as above. Subsequent analyses showed that, unlike the case of control mice (Fig. 6b–g), Ad-UHRF1 mediated UHRF1 overexpression did not significantly affect the levels of fed and fasted blood glucose (Fig. 7d), insulin sensitivity (Fig. 7e, f), liver TG contents (Fig. 7g) and expression of genes involved in gluconeogenesis and lipid synthesis (Fig. 7h), suggesting that the metabolic effects of UHRF1 in the liver are dependent on AMPK.
Discussion
UHRF1 is best known for its key role in targeting DNMT1 to DNA replication forks and consequently DNA maintenance methylation. However, UHRF1 was actually initially identified as a 95-kDa protein that was highly expressed in the S phase of normal mouse thymocytes and constitutively overexpressed in T cell lymphoma cells.32 Since then, a large body of literature has accumulatively documented a widespread UHRF1 overexpression in highly proliferating cells and cancer cells and demonstrated an essential role of UHRF1 in cell proliferation.28,30,34,53 Although DNA methylation is required for somatic cell proliferation and thus may explain the UHRF1 essentiality in cell proliferation and oncogenic function,54 its elevated expression in proliferating and cancer cells strongly suggests a role beyond DNA methylation. We thus examined the potential links between UHRF1 and key cellular signaling pathways that govern cell proliferation and metabolism. This effort allowed us to observe a selective interaction between UHRF1 and AMPKα, among several key signaling kinases we have tested (Fig. 1a). The interaction between UHRF1 and AMPK is confirmed by various co-IP experiments and in vitro pulldown and multiple regions of UHRF1 display interaction with AMPK (Fig. 1b–d and Supplementary information, Fig. S2). Importantly, we found that UHRF1 broadly and negatively regulates AMPK activity in various cells we have tested, including multiple cancer cell lines, MEFs, mouse ESCs, and mouse primary hepatocytes (Figs. 1e–k, 6h) and in mouse liver and adipose tissues (Figs. 6, 7). In addition, we found that UHRF1 suppresses AMPK activity under both basal and stress-induced activation conditions (Fig. 2) and acts independent of its function in DNA methylation (Supplementary information, Fig. S1), highlighting a gate-keeper role of UHRF1 in control of AMPK activity. Furthermore, we found that UHRF1 negatively regulates AMPK activity both in the cytoplasm and nucleus (Fig. 3) and does so not by blocking AMPK activation via upstream kinases LKB1, CAMKK2, and ATM (Supplementary information, Fig. S5). Instead, UHRF1 inactivates AMPK through dephosphorylation by bridging the interaction between AMPK and PP2A phosphatase (Figs. 4–5 and Supplementary information, Fig. S6). Thus, we identify for the first time UHRF1 as a novel gate-keeper of AMPK activity (Fig. 7i).
The finding of UHRF1 as an AMPK gate-keeper also helps reveal a previously unexpected role for UHRF1 in governing lipid and glucose metabolism and protein synthesis in cell and animal levels (Figs. 2, 6, 7; Supplementary information, Fig S3). Using both wild-type and AMPKα1/α2-DKO MEF cells, we show by both gain-of-function and loss-of-function experiments that UHRF1 profoundly suppresses ACC1 phosphorylation but promotes S6 phosphorylation and this effect is dependent on AMPK (Fig. 2k), implying a role of UHRF1 in promoting fatty acid synthesis and protein synthesis via suppressing AMPK. Consistently, knockdown of UHRF1 markedly reduced lipid droplet level in wild-type but not in AMPKα1/α2-DKO MEF cells (Fig. 2l) and in HEK293T and A549 cells (Fig. 2m). Gene expression analysis revealed that UHRF1 regulates fatty acid metabolism-related and glycolysis-related gene expression in an AMPK-dependent manner (Supplementary information, Fig. S3a, b). Importantly, we show by two independent approaches, adenovirus-driven, and transgenic gene-driven liver-specific UHRF1 expression, that enhanced UHRF1 expression robustly suppressed endogenous AMPK activity in livers, leading to elevation of both fed and fast glucose levels and accumulation of lipid droplets, and impaired glucose tolerance and insulin sensitivity (Fig. 6). As the same adenovirus-driven liver-specific UHRF1 expression had no significant effect on fed and fast glucose levels, glucose tolerance and insulin sensitivity, fatty acid and glucose metabolism-related gene expression in liver-specific AMPKα1/α2-DKO mice (Fig. 7), it is clear that UHRF1 regulates glucose and lipid metabolism to a large extent through its inhibitory effect on AMPK. The physiological role of UHRF1 in metabolism is further demonstrated by the knockdown of UHRF1 in adipose tissues, showing that knockdown of UHRF1 resulted in reduced sizes of adipocytes, reduced TG content, and reduced expression of lipogenic genes (Fig. 7a–c). Altogether these data provide evidence that UHRF1 plays an important role in regulating glucose and lipid metabolism under physiological conditions via its gate-keeper function on AMPK.
Our finding that UHRF1 governs AMPK activity also uncovers a potential role of UHRF1 in coordinating cellular energy metabolism and cell proliferation. As a master regulator of cellular metabolism, activation of AMPK suppresses ATP-utilizing anabolic processes and thus also suppresses cell proliferation.1,2,55 Reduced AMPK activity has been observed in various human cancers.56,57 The tumor-suppressive functions of AMPK have been linked to the ability of activated AMPK to inactivate the mTOR signaling pathways including protein synthesis,58 to inhibit the glycolytic pathway,39 and to arrest cell cycle by inducing p53,59 etc. Inhibition of AMPK by UHRF1 is thus likely to promote cell proliferation and impair the tumor-suppressive functions of AMPK. This could explain at least in part the intimate relation between elevated UHRF1 expression and cell proliferation as well as widespread overexpression of UHRF1 in human cancers.28,30,34,53
Our finding that nuclear UHRF1 suppresses AMPK activation also provides a reasonable explanation for differential AMPK activity in the cytoplasm and nucleus observed in the previous studies.42 Strong evidence indicates that AMPK is primarily activated in the cytoplasm and/or lysosomes and that nuclear AMPK is poorly activated by glucose starvation.42 As UHRF1 is highly enriched in the nucleus, it is ideal to inactivate nuclear AMPK by bridging an interaction between PP2A and AMPK. UHRF1, AMPKα and PPP2CA exhibited extensive co-fractionation and co-IP in gel filtration experiments (Fig. 5a, b), further supporting the interaction among these proteins. The PP2A phosphatases are composed of various scaffold and regulatory subunits. Both the catalytic subunit PPP2CA and scaffold subunit PPP2R1A were detected in purified AMPKα2 complexes (Fig. 5c), whereas PPP2CA, PPP2R1A and regulatory subunit PPP2R5D were found to co-immunoprecipitated with nuclear UHRF1 (Fig. 5d). Furthermore, knockdown of PPP2R5D impaired UHRF1’s ability to regulate AMPK (Fig. 5e) supporting a key role for the PPP2R1A/PPP2R5D/PPP2CA PP2A complex in dephosphorylation of nuclear AMPK. However, we do not exclude the possibility that UHRF1 may also promote AMPK dephosphorylation through additional phosphatase(s). Alternatively, UHRF1 may also control AMPK activity by promoting PP2A activity. Future studies are needed to better define the molecular mechanisms by which UHRF1 negatively control AMPK activity.
Because nuclear UHRF1 is capable of suppressing cytoplasmic AMPK (Fig. 3a, b, d–f), we postulate this is likely due to the cytoplasm-nucleus shuttling of AMPK.43 In support of this idea, our live-cell imaging experiments showed that AMPK is indeed rapidly shuttled between cytoplasm and nucleus (Fig. 3k–l). Furthermore, we show that the lysosomal anchored LAMP2-AMPKα2 is not regulated by UHRF1 (Fig. 5f–i), demonstrating that cytoplasm-nuclear shuttling is most likely required for suppression of cytoplasm AMPK by UHRF1. We also found that UHRF1 promotes AMPK nuclear retention (Figs. 3d, f, i, j, 5a, b), presumably due to the protein-protein interaction between UHRF1 and AMPK. The increased AMPK nuclear retention upon increased UHRF1 expression further supports AMPK is shuttling between cytoplasm and nucleus. Thus, the inactivation of AMPK by nuclear UHRF1 would result in a lower AMPK activity in the nucleus and a higher AMPK activity in cytoplasm.42
Our finding that UHRF1 has a profound role in controlling nuclear AMPK activity also unravels a potential role of UHRF1 in coordinating the intricate balance of multiple epigenetic pathways including H3K27 methylation, H2B phosphorylation, DNA methylation and demethylation. Activated AMPK has been shown to activate stress-related gene expression by phosphorylating histone H2B at serine 36,8 to control the activity of exonuclease Exo1 to prevent aberrant fork resection during replication stress,22 to suppress H3K27 methylation and oncogenic function by phosphorylating methyltransferase EZH2 at T311,6 to stabilize TET2 and promote the conversion of methylcytosine in DNA to 5-hydroxymethylcytosine (5hmC) by phosphorylating TET2 at serine 99,7 and to activate CARM1-dependent autophagy transcriptional program by phosphorylating FOXO3,5 and thus can profoundly impact nuclear epigenetic landscapes and function. Consistent with the aforementioned reports, we found that UHRF1 suppresses both EZH2 T311 and H2B S36 phosphorylation in an AMPK-dependent manner (Fig. 3c, g, h, and Supplementary information, Fig. S4b). As nuclear AMPK activity is constantly suppressed by UHRF1, we surmise that sustained AMPK activation in the nucleus could lead to aberrant inactivation of EZH2-mediated polycomb silencing and aberrant activation of stress response genes, which are likely detrimental to proper regulation of epigenome landscape and thus need to be tightly controlled. Our finding that UHRF1 serves as a gate-keeper for nuclear AMPK also raises an intriguing question whether UHRF1 plays a central role in coordinating DNA methylation with EZH2-mediated H3K27 methylation, H2BS36 phosphorylation-regulated gene expression, TET2-mediated DNA demethylation, and FOXO3a-regulated autophagy transcription, as these epigenetic and transcriptional events are all regulated by AMPK. As a key epigenetic regulator that is highly expressed in proliferating and cancer cells, UHRF1 is considered a therapeutic target for cancer. Our finding that UHRF1 functions as a gate-keeper of AMPK and controls cell metabolism provides another incentive for targeting this novel molecule.
Materials and methods
Plasmids
Expression plasmids for various proteins were constructed in the pCDNA3.1 vector for transient transfection, in pLVX-IRES for lentivirus packaging (stabe expression). The lentiviral-based vector PLKO.1 was used for the expression of shRNA in different cells. The nucleotide sequence of human UHRF1 shRNA is as follows: 5′-GCCTTTGATTCGTTCCTTCTT-3′ (#1) and 5′-GCGCTGGCTCTCAACTGCTTT-3′ (#2).
Reagents
Rabbit antibodies pAMPKα-T172 (#2535), AMPKα (#2532), pACC1Ser79 (#3661), LKB1 (#3047), EZH2 (#5246), pEZH2 (#27888), AMPKα2 (#2757), pS6 (#4858), H2B120ub (#5546), HIF1α (#36169) antibodies were purchased from Cell Signaling Technology. Rabbit antibodies AMPKα1 (CY5326), ACC1 (CY5575), mTOR (CY3456), GSk3β (AY0503), Akt1 (Ab3327), γH2AX (Ab3322), and DNMT1 (CY6649) were purchased from Abways Technology. Rabbit antibodies PPM1E (ab137023) and PI3K (ab32089) were purchased from Abcam. Rabbit antibodies PPP2CA (13482-1-AP) and RPS6 (14823-1-AP) were purchased from Proteintech. Rabbit pH2BS36 antibody was purchased from ECMBiosciences. The chemicals STO-609, AICAR, metformin, and thapsigargin were purchased from Selleck. Nile Red Staining kit (ab228553) was purchased from Abcam.
Cell culture, plasmid transfection, and lentivirus infection
HEK293T, HCT116, BEL7402, and MEFs were maintained in DMEM medium (Gibco) and A549 were maintained in 1640 medium (Gibco) supplemented with 10% fetal bovine serum (FBS), 1% penicillin/streptomycin (Invitrogen) at 37 °C in a humidified incubator containing 5% CO2. All cell lines used in the study were regularly tested for mycoplasma contamination. For glucose starvation, cells were washed with PBS, then incubated with glucose-free DMEM supplemented with 10% dialysed FBS. Transient transfections were performed using lipofectamine 2000 according to the manufacturer’s instructions. Cells were harvested for subsequent WB, IP-WB, or RT-PCR analysis 48 h after transfection. Lentiviruses for cell infection were packaged in HEK293T cells using Lipofectamine 2000 transfection according to the manufacturer’s protocols. 48 h post-transfection, the medium was collected and added to the cells for infection. The cells were incubated for another 24 h and then replaced with fresh medium.
IP
Cells were washed with PBS and ice-cold lysis buffer (50 mM Tris-HCl, pH 7.4, 150 mM NaCl, 1% NP-40, 1 mM EDTA, 2 mM PMSF, protease inhibitor) was added. Cells were lysed for 30 min at 4 °C with occasional vortexing. The lysates were centrifugated for 10 min at 12,000 rpm. The supernatants (whole cell extracts) were incubated with various antibodies plus protein A-agarose beads as indicated (1:50 for immunoprecipitating AMPKα and 1:100 for PPP2CA) or anti-Flag M2, anti-Myc, or anti-HA beads (Sigma-Aldrich) for 5 h at 4 °C. The beads were washed five times in washing buffer (20 mM Tris-HCl, pH 8.0, 100 mM NaCl, 1 mM EDTA, 0.5% NP-40, and 1 mM PMSF). Bound proteins were boiled with SDS-PAGE sample buffer, and separated by SDS-PAGE followed by WB analysis.
WB
Total proteins were extracted by lysing cells in lysis buffer containing protease inhibitors as above. Protein samples were separated by SDS-PAGE and transferred onto polyvinylidene fluoride (PVDF) membranes. After blocking with 7% non-fat milk in PBST (PBS with Tween-20), membranes were incubated with the primary antibody and then bound to the secondary antibodies. After incubation with the corresponding secondary antibodies, PVDF membranes were analyzed using an Odyssey infrared imaging system (LI-COR Biosciences).
GST pull-down assay
The cDNA that encodes UHRF1 amino acids 1–300 (UHRF1-1–300), 300–793 (UHRF1-300–C), and 133–300 (UHRF1-133–300) and full length of human UHRF1 was respectively cloned into a pGEX-4T1 vector and transfected into E. coli BL21. Purification of GST-fusion proteins was carried out according to the manufacturer’s instructions. FLAG-AMPKα1 protein was purified from HEK293T cells transfected with FLAG-AMPKα1 plasmid using anti-FLAG M2 beads according to the manufacturer’s instruction. GST pull-down assays were carried out using 2 μg immobilized GST fusion proteins plus ~25 ng FLAG-AMPKα1 in binding buffer (25 mM Tris-HCl, pH 8.0, 150 mM NaCl, 1% NP-40, 2 mM EDTA, 1× protease inhibitor cocktail, and 1 mM DTT) at 4 °C for 4 h on a rotator. The beads were washed three times with binding buffer, and the bound FLAG-AMPKα1 was detected by WB analysis.
Nile red staining
Cells were washed using PBS, fixed using 4% formaldehyde solution, and stained for 30 min with Nile red solution according to the manufacturer’s protocols to visualize lipid droplets. Cell nuclei were counterstained 4,6-diamidino-2-phenylindole (DAPI) and images acquired by laser confocal microscopy (Carl Zeiss).
Lactate measurements
Extracellular lactate levels were measured using Lactate production assay kit (Sigma) according to the manufacturer’s protocols.
Immunofluorescence
For immunofluorescence staining, cells on coverslips were washed with 1×PBS before fixation in 4% fresh paraformaldehyde at room temperature for 30 min. The coverslips were then incubated with 1% Triton X-100 on ice for 15 min, blocked with 5% bovine serum albumin in 37 °C incubator for 60 min, and incubated with primary antibody for 2 h. The coverslips were washed three times with PBST and then incubated with Texas green-conjugated secondary antibody against mouse or rabbit IgG for 1 h. DNA was stained with DAPI. Cells were mounted and visualized under a confocal microscope (Carl Zeiss). Fluorescent images were quantified with ZEN software.
Fluorescence recovery after photobleaching (FRAP)
GFP-UHRF1 and mCherry-AMPKα1 were transfected into HeLa cells. FRAP experiments were performed in a Live Cell imaging chamber (ZEISS) with ZEISS LSM880 laser scanning confocal microscopy. The cytoplasm or nuclear red fluorescence was bleached with a 405 nm laser. GFP fluorescence signal was collected over time. The fluorescence intensity of the bleached area was obtained by ZEN software black edition (ZEISS).
qRT–PCR
For qRT-PCR analysis, total RNAs were extracted from the indicated cell lines and reverse transcribed using kits purchased from Transgene. The quantity of mRNA was calculated using the ΔΔCt method and β-Actin was used as controls. The following mouse primers were used in this study.
β-Actin, forward (fwd) 5′-TAGCCATCCAGGCTGTGCTG-3′, reverse (rev) 5′-CAGGATCTTCATGAGGTAGTC-3′;
UHRF1, fwd 5′-CCACACCGTGAACTCTCTGTC-3′, rev 5′-AGGCGCACATCATAATCGAAG-3′;
ALDOA, fwd 5′-AGTCCACCGGAAGCATTGC-3′, rev 5′-CAGCCCCTGGGTAGTTGTC-3′;
HK1, fwd 5′-AACGGCCTCCGTCAAGATG-3′, rev 5′-GCCGAGATCCAGTGCAATG-3′;
HK2, fwd 5′-CTAAGGGGTTCAAGTCCAGTGG-3′, rev 5′-AGACCAATCTCGCAGTTCTGA-3′;
LDHA, fwd 5′-TTCAGCGCGGTTCCGTTAC-3′, rev 5′-CCGGCAACATTCACACCAC-3′;
PDK1, fwd 5′-AGGATCAGAAACCGGCACAAT-3′, rev 5′-GTGCTGGTTGAGTAGCATTCTAA-3′;
PFKP, fwd 5′-CGCCTATCCGAAGTACCTGGA-3′, rev 5′-CCCCGTGTAGATTCCCATGC-3′;
PGK1, fwd 5′-ATGTCGCTTTCCAACAAGCTG-3′, rev 5′-GCTCCATTGTCCAAGCAGAAT-3′.
Subcellular fractionation
The cells were rinsed with ice-cold PBS and then added buffer A (10 mM HEPES, pH 7.9, 1.5 mM MgCl2, 10 mM KCl, 0.5 mM DTT, 0.5% NP40 and protease inhibitor cocktail). After 20 min incubation on ice, the cells were centrifuged at 13,000 rpm for 10 min. The resultant supernatant was then used as a cytosolic fraction. The pellet was then homogenized with buffer B (20 mM HEPES, pH 7.9, 0.4 M NaCl, 1 mM EDTA, 1 mM DTT, and 1 mM PMSF). After 10 min of vigorous vortexing, the homogenates were centrifuged for 20 min at 13,000 rpm and the resultant supernatants were then used as nuclear fractions.
Mass spectrometry analysis
The immunoprecipitated material was digested overnight with trypsin, and peptides preparation and mass spectrometry analysis were performed as previously described.60
Gel filtration chromatography
Three-milligram nuclear proteins in 500 μL buffer B (10 mM Tris-HCl, pH 7.6, 150 mM NaCl, 2 mM MgCl2) were loaded onto a 24 mL superdex 200 10/300 GL column. The eluted fractions (0.5 ml each) were collected and analyzed by WB or IP-WB using the indicated antibodies.
Mice
Male C57BL/6 J mice aged 8–12 weeks were purchased from Shanghai Laboratory Animal Company (SLAC, Shanghai, China). Hepatocyte-specific UHRF1 transgenic mice were generated by the company Cyagen Biosciences. In brief, UHRF1 was cloned into the Cre/loxP conditional ROSA26 targeting vector and used for generating transgenic mouse lines, and liver-specific overexpression of UHRF1 was achieved by crossing with mice expressing Cre-recombinase under the control of the albumin promoter. WT and liver-specific AMPKα1/α2 knockout mice (AMPKα-LKO) were generated as previously reported.52 All mice were housed at 21 °C ± 1 °C with humidity of 55% ± 10% and a 12-h light/12-h dark cycle, with free access to chow diet and water. All studies with mice were reviewed and approved by the Institutional Animal Use and Care Committee of East China Normal University.
Adenoviruses and siRNA delivery
Adenoviruses expressing murine UHRF1 or GFP (Ad-UHRF1, Ad-GFP) were constructed by Hanbio Biotechnology Co. Ltd., (Shanghai, China) with a full-length UHRF1 or GFP coding sequence. Overexpression of hepatic UHRF1 or GFP was achieved by means of tail vein injection of Ad-UHRF1 or Ad-GFP (2 × 109 plaque-forming units) in male C57BL/6 J mice or WT and AMPKα-LKO mice. The random (fed) and fasted blood glucose levels were measured at day 4, and the insulin tolerance tests and glucose tolerance tests were conducted at day 6 and day 9 after adenoviruses injection, respectively. At day 12, mice were euthanized, and tissues were dissected for further analysis. To silence UHRF1 in inguinal white adipose tissue (iWAT), siRNA targeting UHRF1 and a negative control siRNA were synthesized with the following sequences: siRNA (NC) 5′-UUCUCCGAACGUGUCACGUTT-3′, siUHRF1 5′-GGAAGUGUUUCACGUGGAATT-3′ (designed and synthesized by Genepharma, Shanghai, China). Male C57BL/6 J mice were randomly assigned to either an HFD (60% fat, 20% protein, and 20% carbohydrates; Research Diets) for 4 weeks. In vivo transfer of siRNA into the inguinal fat pads of HFD-fed mice was achieved by a commercialized kit (Entranster-in vivo; Engreen Biosystem Co. Ltd., Beijing, China) as previously reported.61
Primary hepatocyte isolation and adenovirus treatment
Primary hepatocytes were isolated from 8 weeks old male C57BL/6 J mice by collagenase perfusion as previously described.62 Hepatocytes were seeded at a final density of 0.5 × 106 cells/well in 6-well plates and cultured in DMEM supplemented with 10% FBS and 1% penicillin-streptomycin. Cells were washed with PBS at 6 h, and the medium was replaced with fresh 10% FBS high-glucose DMEM complete medium. After 24 h, primary hepatocytes were infected with Ad-UHRF1 or Ad-GFP for 36 h, and then cultured with serum-free DMEM medium for 16 h and collected for further analysis.
Glucose and insulin tolerance tests
For glucose tolerance tests (GTT), mice were fasted for 16 h, followed by intraperitoneal (i.p.) glucose injection (1.5 g/kg body weight). For insulin tolerance tests (ITT), mice received an i.p. injection of insulin (1 U/kg body weight). Blood glucose levels were measured using an automated glucose monitor by tail punch at 0, 15, 30, 60, and 90 min after glucose or insulin injections.
Hepatic triglyceride assay
Liver samples with steel beads in tissue lysis buffer (140 mM NaCl, 50 mM Tris-HCl, pH 8.0, and 1% Triton-X100) were homogenized using a TissueLyser (Jingxin) on ice. Triglyceride concentration in the lysates was measured using Triglyceride Quantification Kit (BioVision) according to the manufacturer’s protocols.
Histological analysis
For Oil Red O staining, 10 μm frozen sections were prepared from snap-frozen liver tissues, and fixed in 10% buffered formalin. The sections were then stained with 0.5% Oil Red O (Sigma) to evaluate hepatic lipid content. For H&E staining, liver and inguinal fat tissues were fixed overnight in 4% formalin, and embedded in paraffin. Paraffin-sections (5μm thick) were deparaffinized, rehydrated, and stained with H&E. Images were acquired by a digital camera (Olympus). The size of adipocytes in iWAT was quantified using Image J.
Statistical analysis
Two-tailed Students t-test was used as specified in the figure legends. ns donates no significant difference; * denotes P < 0.05; ** denotes P < 0.01; and *** denotes P < 0.001. Values are expressed as means ± SEM or means ± SD.
References
Herzig, S. & Shaw, R. J. AMPK: guardian of metabolism and mitochondrial homeostasis. Nat. Rev. Mol. Cell Biol. 19, 121–135 (2018).
Lin, S. C. & Hardie, D. G. AMPK: sensing glucose as well as cellular energy status. Cell Metab. 27, 299–313 (2018).
Hardie, D. G., Ross, F. A. & Hawley, S. A. AMPK: a nutrient and energy sensor that maintains energy homeostasis. Nat. Rev. Mol. Cell Biol. 13, 251–262 (2012).
Carling, D., Zammit, V. A. & Hardie, D. G. A common bicyclic protein kinase cascade inactivates the regulatory enzymes of fatty acid and cholesterol biosynthesis. FEBS Lett. 223, 217–222 (1987).
Shin, H. J. et al. AMPK-SKP2-CARM1 signalling cascade in transcriptional regulation of autophagy. Nature 534, 553–557 (2016).
Wan, L. et al. Phosphorylation of EZH2 by AMPK suppresses PRC2 methyltransferase activity and oncogenic function. Mol. Cell 69, 279–291 (2018).
Wu, D. et al. Glucose-regulated phosphorylation of TET2 by AMPK reveals a pathway linking diabetes to cancer. Nature 559, 637–641 (2018).
Bungard, D. et al. Signaling kinase AMPK activates stress-promoted transcription via histone H2B phosphorylation. Science 329, 1201–1205 (2010).
Hawley, S. A. et al. Complexes between the LKB1 tumor suppressor, STRAD alpha/beta and MO25 alpha/beta are upstream kinases in the AMP-activated protein kinase cascade. J. Biol. 2, 28 (2003).
Woods, A. et al. LKB1 is the upstream kinase in the AMP-activated protein kinase cascade. Curr. Biol. 13, 2004–2008 (2003).
Woods, A. et al. Ca2+/calmodulin-dependent protein kinase kinase-beta acts upstream of AMP-activated protein kinase in mammalian cells. Cell Metab. 2, 21–33 (2005).
Hawley, S. A. et al. Calmodulin-dependent protein kinase kinase-beta is an alternative upstream kinase for AMP-activated protein kinase. Cell Metab. 2, 9–19 (2005).
Zhang, Y. L. et al. AMP as a low-energy charge signal autonomously initiates assembly of AXIN-AMPK-LKB1 complex for AMPK activation. Cell Metab. 18, 546–555 (2013).
Zhang, C. S. et al. The lysosomal v-ATPase-Ragulator complex is a common activator for AMPK and mTORC1, acting as a switch between catabolism and anabolism. Cell Metab. 20, 526–540 (2014).
Zhang, C. S. et al. Fructose-1,6-bisphosphate and aldolase mediate glucose sensing by AMPK. Nature 548, 112–116 (2017).
Li, M. et al. Transient receptor potential V channels are essential for glucose sensing by aldolase and AMPK. Cell Metab. 30, 508–524 (2019).
Marley, A. E. et al. Biochemical characterization and deletion analysis of recombinant human protein phosphatase 2C alpha. Biochem. J. 320, 801–806 (1996).
Gimeno-Alcaniz, J. V. & Sanz, P. Glucose and type 2A protein phosphatase regulate the interaction between catalytic and regulatory subunits of AMP-activated protein kinase. J. Mol. Biol. 333, 201–209 (2003).
Voss, M. et al. Ppm1E is an in cellulo AMP-activated protein kinase phosphatase. Cell Signal. 23, 114–124 (2011).
Salminen, A., Kaarniranta, K. & Kauppinen, A. Age-related changes in AMPK activation: Role for AMPK phosphatases and inhibitory phosphorylation by upstream signaling pathways. Ageing Res. Rev. 28, 15–26 (2016).
Chandrashekarappa, D. G., McCartney, R. R. & Schmidt, M. C. Ligand binding to the AMP-activated protein kinase active site mediates protection of the activation loop from dephosphorylation. J. Biol. Chem. 288, 89–98 (2013).
Li, S. et al. Ca(2+)-stimulated AMPK-dependent phosphorylation of Exo1 protects stressed replication forks from aberrant resection. Mol. Cell 74, 1123–1137 (2019).
Vara-Ciruelos, D. et al. Genotoxic damage activates the AMPK-alpha1 isoform in the nucleus via Ca(2+)/CaMKK2 signaling to enhance tumor cell survival. Mol. Cancer Res. 16, 345–357 (2018).
Sharif, J. et al. The SRA protein Np95 mediates epigenetic inheritance by recruiting Dnmt1 to methylated DNA. Nature 450, 908–912 (2007).
Bostick, M. et al. UHRF1 plays a role in maintaining DNA methylation in mammalian cells. Science 317, 1760–1764 (2007).
Liang, C. C. et al. UHRF1 is a sensor for DNA interstrand crosslinks and recruits FANCD2 to initiate the Fanconi anemia pathway. Cell Rep. 10, 1947–1956 (2015).
Tian, Y. et al. UHRF1 contributes to DNA damage repair as a lesion recognition factor and nuclease scaffold. Cell Rep. 10, 1957–1966 (2015).
Jenkins, Y. et al. Critical role of the ubiquitin ligase activity of UHRF1, a nuclear RING finger protein, in tumor cell growth. Mol. Biol. Cell 16, 5621–5629 (2005).
Bronner, C. et al. The UHRF family: oncogenes that are drugable targets for cancer therapy in the near future? Pharmacol. Ther. 115, 419–434 (2007).
Li, J. et al. Activated MEK/ERK pathway drives widespread and coordinated overexpression of UHRF1 and DNMT1 in cancer cells. Sci. Rep. 9, 907 (2019).
Jia, Y. et al. Negative regulation of DNMT3A de novo DNA methylation by frequently overexpressed UHRF family proteins as a mechanism for widespread DNA hypomethylation in cancer. Cell Discov. 2, 16007 (2016).
Fujimori, A. et al. Cloning and mapping of Np95 gene which encodes a novel nuclear protein associated with cell proliferation. Mamm. Genome 9, 1032–1035 (1998).
Liu, X. et al. UHRF1 targets DNMT1 for DNA methylation through cooperative binding of hemi-methylated DNA and methylated H3K9. Nat. Commun. 4, 1563 (2013).
Kong, X. et al. Defining UHRF1 domains that support maintenance of human colon cancer DNA methylation and oncogenic properties. Cancer Cell 35, 633–648 (2019).
Rothbart, S. B. et al. Multivalent histone engagement by the linked tandem Tudor and PHD domains of UHRF1 is required for the epigenetic inheritance of DNA methylation. Genes Dev. 27, 1288–1298 (2013).
Zhang, H. et al. A cell cycle-dependent BRCA1-UHRF1 cascade regulates DNA double-strand break repair pathway choice. Nat. Commun. 7, 10201 (2016).
Sullivan, J. E. et al. Inhibition of lipolysis and lipogenesis in isolated rat adipocytes with AICAR, a cell-permeable activator of AMP-activated protein kinase. FEBS Lett. 353, 33–36 (1994).
Tamas, P. et al. Regulation of the energy sensor AMP-activated protein kinase by antigen receptor and Ca2+ in T lymphocytes. J. Exp. Med. 203, 1665–1670 (2006).
Faubert, B. et al. AMPK is a negative regulator of the Warburg effect and suppresses tumor growth in vivo. Cell Metab. 17, 113–124 (2013).
Kishton, R. J. et al. AMPK is essential to balance glycolysis and mitochondrial metabolism to control T-ALL cell stress and survival. Cell Metab. 23, 649–662 (2016).
Shackelford, D. B. et al. mTOR and HIF-1alpha-mediated tumor metabolism in an LKB1 mouse model of Peutz-Jeghers syndrome. Proc. Natl. Acad. Sci. USA 106, 11137–11142 (2009).
Zong, Y. et al. Hierarchical activation of compartmentalized pools of AMPK depends on severity of nutrient or energy stress. Cell Res. 29, 460–473 (2019).
Kodiha, M., Rassi, J. G., Brown, C. M. & Stochaj, U. Localization of AMP kinase is regulated by stress, cell density, and signaling through the MEK-ERK1/2 pathway. Am. J. Physiol. Cell Physiol. 293, C1427–C1436 (2007).
Shaw, R. J. et al. The tumor suppressor LKB1 kinase directly activates AMP-activated kinase and regulates apoptosis in response to energy stress. Proc. Natl. Acad. Sci. USA 101, 3329–3335 (2004).
Hurley, R. L. et al. The Ca2+/calmodulin-dependent protein kinase kinases are AMP-activated protein kinase kinases. J. Biol. Chem. 280, 29060–29066 (2005).
Sanchez-Cespedes, M. et al. Inactivation of LKB1/STK11 is a common event in adenocarcinomas of the lung. Cancer Res. 62, 3659–3662 (2002).
Tokumitsu, H. et al. STO-609, a specific inhibitor of the Ca(2+)/calmodulin-dependent protein kinase kinase. J. Biol. Chem. 277, 15813–15818 (2002).
Fu, X., Wan, S., Lyu, Y. L., Liu, L. F. & Qi, H. Etoposide induces ATM-dependent mitochondrial biogenesis through AMPK activation. PLoS One 3, e2009 (2008).
Hickson, I. et al. Identification and characterization of a novel and specific inhibitor of the ataxia-telangiectasia mutated kinase ATM. Cancer Res. 64, 9152–9159 (2004).
Li, J. et al. Both corepressor proteins SMRT and N-CoR exist in large protein complexes containing HDAC3. EMBO J. 19, 4342–4350 (2000).
Zhu, X. N. et al. PP2A-AMPKalpha-HSF1 axis regulates the metal-inducible expression of HSPs and ROS clearance. Cell. Signal. 26, 825–832 (2014).
Han, Y. et al. Post-translational regulation of lipogenesis via AMPK-dependent phosphorylation of insulin-induced gene. Nat. Commun. 10, 623 (2019).
Ashraf, W. et al. The epigenetic integrator UHRF1: on the road to become a universal biomarker for cancer. Oncotarget 8, 51946–51962 (2017).
Chen, T. et al. Complete inactivation of DNMT1 leads to mitotic catastrophe in human cancer cells. Nat. Genet. 39, 391–396 (2007).
Gonzalez, A., Hall, M. N., Lin, S. C. & Hardie, D. G. AMPK and TOR: The Yin and Yang of Cellular Nutrient Sensing and Growth Control. Cell Metab. 31, 472–492 (2020).
Hadad, S. M. et al. Histological evaluation of AMPK signalling in primary breast cancer. BMC Cancer 9, 307 (2009).
Tong, W. H. et al. The glycolytic shift in fumarate-hydratase-deficient kidney cancer lowers AMPK levels, increases anabolic propensities and lowers cellular iron levels. Cancer Cell 20, 315–327 (2011).
Gwinn, D. M. et al. AMPK phosphorylation of raptor mediates a metabolic checkpoint. Mol. Cell 30, 214–226 (2008).
Jones, R. G. et al. AMP-activated protein kinase induces a p53-dependent metabolic checkpoint. Mol. Cell 18, 283–293 (2005).
Tan, G. H. et al. PRRT2 deficiency induces paroxysmal kinesigenic dyskinesia by regulating synaptic transmission in cerebellum. Cell Res. 28, 90–110 (2018).
Zhou, J. N. et al. MicroRNA-125b attenuates epithelial-mesenchymal transitions and targets stem-like liver cancer cells through small mothers against decapentaplegic 2 and 4. Hepatology 62, 801–815 (2015).
Lu, Y. et al. Periostin promotes liver steatosis and hypertriglyceridemia through downregulation of PPARalpha. J. Clin. Invest. 124, 3501–3513 (2014).
Acknowledgements
We thank all other members of the J.W. laboratory for suggestions and technical assistance. We thank Q.Y. Lei (Fudan University, Shanghai, China) for kindly providing the AMPKα1/α2 double-knockout MEFs. We thank X.H. Feng (Zhejiang University, Hangzhou, China) for kindly providing FLAG-PPP2CA, FLAG-PP2Cα, FLAG-PP2Cβ and FLAG-PPM1E expression constructs. We also thank Wen Chen (Sun Yat-sen University, Guangzhou, China) for kindly providing antibody and shRNA plasmid for PPP2R5D. This study is supported by grants from the Ministry of Science and Technology of China (2017YFA054201) and the National Natural Science Foundation of China (31730048 and 31961133009 to J.W.), the Shanghai Science and Technology Committee (20JC1411500 to J.W.) and the National Natural Science Foundation of China (91957120 to S.H.L.).
Author information
Authors and Affiliations
Contributions
The study was designed by X.X., G.D., X.M. and J.W., all biochemical and most cellular experiments were performed by X.X., G.D., Y.D., X.H., Y.H. and Z.C.; X.H., X.X., W.W., and W.L. did live cell imaging experiments. C.L., X.C. and J.L. carried out experiments with liver-specific UHRF1 overexpression and liver-specific AMPKα1/α2 knockout mice. X.X., G.D., C.L., X.M. and J.W. wrote the manuscript and all members edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Xu, X., Ding, G., Liu, C. et al. Nuclear UHRF1 is a gate-keeper of cellular AMPK activity and function. Cell Res 32, 54–71 (2022). https://doi.org/10.1038/s41422-021-00565-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41422-021-00565-y
This article is cited by
-
AMPK activation attenuates central sensitization in a recurrent nitroglycerin-induced chronic migraine mouse model by promoting microglial M2-type polarization
The Journal of Headache and Pain (2024)
-
AKT1 regulates UHRF1 protein stability and promotes the resistance to abiraterone in prostate cancer
Oncogenesis (2023)
-
ASCL1 activates neuronal stem cell-like lineage programming through remodeling of the chromatin landscape in prostate cancer
Nature Communications (2022)
-
UHRF1: a novel metabolic guardian restricting AMPK activity
Cell Research (2022)
-
UHRF1 overexpression promotes osteosarcoma metastasis through altered exosome production and AMPK/SEMA3E suppression
Oncogenesis (2022)