Abstract
Nonsense-mediated mRNA decay (NMD) targets premature stop codon (PTC)-containing mRNAs for rapid degradation, and is essential for mammalian embryonic development, brain development and modulation of the stress response. The key event in NMD is the SMG1-mediated phosphorylation of an RNA helicase UPF1 and SMG1 kinase activity is inhibited by SMG8 and SMG9 in an unknown mechanism. Here, we determined the cryo-EM structures of human SMG1 at 3.6 Å resolution and the SMG1–SMG8–SMG9 complex at 3.4 Å resolution, respectively. SMG8 has a C-terminal kinase inhibitory domain (KID), which covers the catalytic pocket and inhibits the kinase activity of SMG1. Structural analyses suggest that GTP hydrolysis of SMG9 would lead to a dramatic conformational change of SMG8–SMG9 and the KID would move away from the inhibitory position to restore SMG1 kinase activity. Thus, our structural and biochemical analyses provide a mechanistic understanding of SMG1–SMG8–SMG9 complex assembly and the regulatory mechanism of SMG1 kinase activity.
Similar content being viewed by others
Introduction
Nonsense-mediated mRNA decay (NMD) plays an important role in rapid degradation of premature stop codon (PTC)-containing mRNAs.1,2,3,4,5 The NMD pathway was initially identified in human cells6 and later recognized to be conserved from yeast to human. Multiple aspects of NMD have substantial clinical significance.7 NMD also functions in posttranscriptional regulation8,9,10,11,12 and is essential for various biological processes including embryonic development and brain development.9 The dysregulations of the NMD pathway have been frequently observed in various human diseases, as exemplified by neurodevelopmental disorders, mental retardation, specific type of cancers, and inherited disorders.13,14,15,16,17
NMD is a fine-tuned process that requires concerted action of several trans-acting factors.9,18,19 The SMG1-mediated phosphorylation of an RNA helicase UPF1 provides a binding platform for NMD factors SMG6 and SMG5-SMG7 for subsequent mRNA degradation.9,20,21,22 SMG1, a 410-KDa protein, belongs to phosphatidylinositol 3-kinase-related kinases (PIKKs) family,23,24 which includes the master regulator of cell growth, mTOR, and three key regulators involved in DNA damage response, ATM, ATR and DNA-PKcs.
In human cells, SMG1 forms a complex with two NMD effectors SMG8 and SMG9, which inhibit the kinase activity of SMG1 in vitro.25,26,27 SMG9 directly binds SMG1 whereas SMG8 binds SMG1 only in the presence of SMG9,26 suggesting the central role of SMG9 in the assembly of SMG1–SMG8–SMG9 complex (designated SMG1C). Crystal structure of C. elegans SMG8–SMG9 core complex reveals that SMG8–SMG9 heterodimer is formed through the contacts between a C-terminal guanine nucleotide-binding (G) domain of SMG9 and an N-terminal G-like domain of SMG8.28 Previous electron microscopic studies revealed low-resolution density map of SMG1 and SMG1C.25,27,29 However, the mechanisms for complex assembly and regulation of SMG1 kinase by SMG8 and SMG9 remain largely unknown.
In this study, we determined the cryo-electron microscopy (EM) structures of human SMG1 and SMG1C complex. The atomic resolution structures reveal distinct structural features of SMG1 compared with other PIKK kinase. SMG8–SMG9 heterodimer stably associates with SMG1 and the C-terminal region of SMG8 inhibits SMG1 kinase activity through covering the catalytic pocket. By comparing with the previously reported structure,28 we proposed a mechanism for activation of SMG1 kinase upon GTP hydrolysis of the SMG9 GTPase. The structural and biochemical analyses together provide structural insights into the assembly of SMG1C complex and the regulation of SMG1 kinase activity. The structures also provide a framework for further studies of the NMD pathway and the design of inhibitors for therapeutic applications.
Results
Structure determination and model building
We purified SMG1 and SMG1C for biochemical and structural analyses (Supplementary information, Fig. S1). The cryo-EM structures were determined using single particle reconstruction. The EM maps of SMG1 and SMG1C were refined to 3.6 Å and 3.4 Å resolution, respectively (Supplementary information, Figs. S2, 3, Table S1 and Movies S1, 2). The structural models were built by homology modeling and manually building into the EM density using the structures of mTOR30 and C. elegans SMG8–SMG9 complex28 as templates.
Overall structure of SMG1
SMG1 is composed of an N-terminal extended α-solenoid consisting of 14 parallel HEAT repeats (N-HEAT) and a C-terminal compact catalytic core domain (CoreSMG1) consisting of middle HEAT repeats (M-HEAT), a FAT domain, a kinase domain (KD) and an FATC domain (Fig. 1 and Supplementary information, Fig. S3). The N-HEAT adopts a C-shaped fold with the N-terminus flanking away from the CoreSMG1. A four-way junction is formed by the N-HEAT, the concave side of the M-HEAT, the TRD2 of the FAT domain, and the C-lobe of KD (Fig. 1d). The M-HEAT consists of 3 parallel HEAT repeats and packs against the C-lobe of KD (Fig. 1e). The HRD of FAT domain packs against and stabilizes the N-lobe of KD. The FAT domain and the M-HEAT together form an open ring, which wraps around and stabilizes KD (Fig. 1f).
Structure of SMG1. a Color-coded domain structure of human SMG1, SMG8 and SMG9, and schematic illustration of intermolecular interaction within the SMG1C complex. The same color scheme is used in all structure figures. b, c Ribbon representation of SMG1 structural model (c) and the cryo-EM maps (b) in two different views. The domains are indicated with arrows. d-f Close-up views of the four-way junction (d), the contacts between KD and M-HEAT (e), and the C-terminal core domain (f)
SMG1 is more structurally similar to mTOR than other PIKK family members as represented by ATR and DNA-PKcs30,31,32,33 (Fig. 2 and Supplementary information, Figs. S4, 5). The KDs of the four structures are superimposed well and the critical residues for catalysis are highly conserved (Supplementary information, Fig. S5). SMG1 has structural features distinct from other PIKK kinases (Fig. 2). (1) In other PIKK family members, the N-HEAT domains more or less adopt a twisted super-helical conformation. In contrast, SMG1 has a nearly C-shaped N-HEAT (Fig. 2b–e). (2) Extensive inter-domain contacts (four-way junction) were observed in SMG1 and DNA-PKcs, but not in mTOR or ATR (Fig. 2g–j). This four-way junction may stabilize SMG1 and DNA-PKcs in a monomeric form. In contrast, mTOR and ATR are primarily stabilized upon formation of mTORC1/230,33,34 and ATR-ATRIP31 complexes, respectively. (3) SMG1 has a unique characteristic insertion (residues 2427–3606) that connects the helices kα9 and kα10 of KD and is supposed to be in close proximity to the substrate-binding groove of KD, suggesting its potential regulatory function (Fig. 1b, c). This insertion is invisible according to the EM maps of SMG1 and SMG1C, indicating an intrinsic flexible property. (4) The helix kα9b of mTOR is predicted to block substrate entry.35 In SMG1, kα9b is less ordered and only has a one-turn helix (Fig. 2f), suggesting an open pocket for substrate phosphorylation. (5) An FRB-like domain (FRBL, residues 1960–2085) protrudes out from the N-lobe of KD and adopts a similar conformation to that of mTOR30,33,34,35 and DNA-PKcs.32,36 However, no rapamycin-binding affinity was reported in SMG1. (6) The TRD3 does not adopt a typical TRD conformation and has no direct contact with the KD or the N-HEAT (Fig. 2f). A four-helix insert (residues 1637–1743) protrudes out of the TRD3. This insert was not observed in other PIKK members, suggesting a SMG1-specific function yet to be discovered.
Structural comparison of PIKK family members. a Superimposition of structures of SMG1 (this study), and representative PIKK family members, mTOR in mTORC1 complex (PDB ID: 5WBY),30 DNA-PKcs in DNA-PK holoenzyme (PDB ID: 5Y3R),32 and ATR in ATR-ATRIP complex (PDB ID: 5YZ0).31 The structures are shown in ribbon representations in two different views. b-e The structures of SMG1 (b), mTOR (c), ATR (d) and DNA-PKcs (e) are shown as in a. f Close-up view of the superimposed KDs of SMG1, mTOR and DNA-PKcs. The conformations of helix kα9b, the critical element for regulation of catalysis, are shown in a good view. The kα9b of SMG1 is less ordered than that in mTOR complex, indicating an open catalytic pocket of the SMG1 KD. g-j Close-up views of the four-way junction of SMG1 (g), and equivalent regions in mTOR (h), ATR (i) and DNA-PKcs (j). SMG1 and DNA-PKcs share similar organization of the four-way junction to mediate inter-domain interactions. In contrast, mTOR and ATR have much less interactions in this region. The inter-domain interactions of mTOR and ATR are mediated by dimerization of the kinase proteins (not shown here)
Structure of SMG1C
Within the SMG1C complex, SMG9 occupies the central position and bridges SMG8 and SMG1 (Fig. 3a; Supplementary information, Fig. S3 and Movie S3), consistent with the previous knowledge that SMG8 binds SMG1 only in the presence of SMG9.26 SMG8 consists of an N-terminal G-like domain followed by a stem-like domain composed of a three-helix bundle (Supplementary information, Fig. S6). The G-like domain of SMG8 has no direct contact with SMG1. Helix α9 (residues 343–357) of SMG8 flanks out of the G-like domain and packs against the concave hydrophobic surface formed by repeats HEAT-9 and HEAT-10 of SMG1 (Fig. 3a, b). The equivalent region of SMG9 (residues 463–452) forms an extended coil region with residues F454, L457, and Y460 packing against repeats HEAT-10 and HEAT-11 of SMG1. Moreover, extensive hydrophobic contacts are formed between the M-HEATSMG1 and the edge of SMG9 β-sheet, in agreement with a strong SMG1-SMG9 interaction (Fig. 3c).
Structure of SMG1C. a Ribbon representation of the SMG1C structure in two different views. The intermolecular interfaces are indicated with blue squares. b, c Close-up views of the contacts between SMG1 N-HEAT and SMG8–SMG9 (b) and the contacts between SMG1 M-HEAT and SMG9 (c). The residues involved in the interactions are shown in stick representation and indicated. d Structural comparison of SMG1 and SMG1C. SMG1 is colored in yellow and SMG1C is colored in grey for comparison. The HEAT repeats of N-HEAT are indicated and other regions of SMG1 are omitted for simplicity. The SMG1-binding regions of SMG8 and SMG9 are indicated with arrows
Conformational changes of SMG1 upon SMG1C formation
Structural comparison of SMG1 with the SMG1C complex indicates a similar fold with a root-mean-square deviation (RMSD) of 2.3 Å for 1693 Cα atoms. The major difference exists at the N-HEAT, which adopts a more curved conformation in the SMG1C complex (Fig. 3d). HEAT-11 is the primary turning point between the two structures and the driving force might be derived from SMG8–SMG9, which binds SMG1 and pulls the repeats HEAT-9 towards HEAT-11. As a result, the repeats HEAT-1 to HEAT-8 rotate accordingly and the repeat HEAT-1 moves as far as ~12 Å. The flexible property of the N-HEAT favors the conformational changes of SMG1 and the SMG1C complex assembly.
SMG8–SMG9 interaction
A well-defined EM density was observed in the GTP-binding pocket of SMG9 G domain, and a GTP molecule and a magnesium cation could be nicely fitted into the EM density (Fig. 4a). The bound GTP in SMG9 is derived from endogenously purified SMG1C because no nucleotide was added during protein purification. No GTP/GDP was observed in SMG8 that lacks a consensus GTP/GDP-binding motif at P loop (Supplementary information, Fig. S4b). The typical GTPase regulatory elements, including the P loop, Switch I and Switch II, are stabilized by the GTP and magnesium cation (Fig. 4b). In SMG9, the magnesium cation is chelated by two oxygen atoms of the β and γ phosphate groups of GTP and residues S218 of P loop and T253 of Switch I. The coordination pattern is similar to that in characteristic GTPases in GTP-bound form.37 The guanine base is sandwiched by hydrophobic residues, M219, F466, L428, and aliphatic side chain of K373 of SMG9, and the interaction is further supported by a hydrogen bond between GTP and residue N372.
Structure of SMG8–SMG9 in SMG1C. a Close-up view of GTP-binding pocket of SMG9 in GTP-bound form. GTP is well covered by the EM map shown in blue mesh. The magnesium cation is shown in red ball. GTP and the residues involved in magnesium cation chelation and guanine base recognition are shown in stick representation and indicated. b Ribbon representation of SMG8–SMG9 in the SMG1C structure. The key GTPase elements, P loop, Switch I and Switch II of SMG9, are indicated. The G domain of SMG9 and G-like domain of SMG8 form a pseudo symmetric heterodimer. c Close-up view of SMG8–SMG9 interface. The residues involved in the SMG8–SMG9 interactions are shown in stick representation
Structure of SMG1C shows that SMG8 and SMG9 form a pseudo symmetric dimer and the interaction is mediated by extensive hydrophobic contacts (Fig. 4b, c). The helices α5 and α8 of SMG8 bind to the Switch I, Switch II (containing α3 and associated regions), and α4 of SMG9. Residues L274, L279 and H297 of SMG9 Switch II bind to residues F219, I221, D224 and I311 of SMG8. Similarly, the α5 and α8 of SMG9 make contacts with the helices α3 and α4 of SMG8 (Fig. 4b, c). Residues W324, F325, M390 and L394 of SMG9 make hydrophobic contacts with residues L58, L162, L163 and W189 of SMG8. Moreover, residues M339, Y515, L518 and L519 of SMG9 make hydrophobic contacts with residues I491 and F495 of the SMG8 three-helix bundle and the interaction is supported by a hydrogen bond formed between Y364 of SMG9 and R540 of SMG8. These contacts support the relative conformation of the SMG8 three-helix bundle to the SMG8–SMG9 complex. The structural observation is consistent with the previous study that double mutations M390R/Y515R of human SMG9 impaired the interaction with SMG8.28
Human SMG9 has undetectable GTPase activity in the context of SMG1C complex because GTP is stably maintained during protein purification (Fig. 4a), suggesting that SMG1C adopts a GTP-bound state. Intriguingly, no GTP or GDP molecule was observed in the C. elegans SMG8–SMG9 complex purified from insect cells and the GDP observed in the crystal structure was added after crystallization,28 suggesting a stable C. elegans SMG8–SMG9 interaction in the absence of GTP or GDP. Thus, the G domain of SMG9 may provide a functional switch to modulate the conformation of SMG8–SMG9 complex relative to SMG1.
Structural difference of SMG8–SMG9 complex in GTP- and GDP-bound forms
Structural comparison of GTP-bound human SMG8–SMG9 complex and GDP-bound C. elegans SMG8–SMG9 complex28 shows that individual G-like domain of SMG8 and G domain of SMG9 adopt similar overall fold (Fig. 5 and Supplementary information, Fig. S6). However, the relative conformation of SMG8 and SMG9 is obviously different in the two structures (Fig. 5a). The Switch I and Switch II of SMG9 are well-ordered in the GTP-bound form but invisible in the C. elegans GDP-bound form. In the GDP-bound form, the intermolecular interactions between the two Switch regions of SMG9 and helices α5 and α8 of SMG8 are not maintained.
Conformational switch of SMG8–SMG9 and regulation of SMG1 kinase activity. a Ribbon representations of human SMG8–SMG9GTP in SMG1C and C. elegans SMG8–SMG9GDP structures with two SMG9 superimposed. Two different views are shown and key structural elements are indicated. Human (H) and C. elegans (C) SMG8 are colored in lemon and magenta, respectively. Human and C. elegans SMG9 are colored in light blue and dark green, respectively. b C. elegans SMG8–SMG9GDP and human SMG1C structures are shown with two SMG9 superimposed. Left panel: the EM map of the SMG8 three-helix bundle and the extended region (KID) shown in isosurface representation. KID is right above the catalytic pocket of the SMG1 KD. Right panel: the left EM map fitted with the three-helix bundle of C. elegans SMG8–SMG9GDP structure with SMG9GDP and human SMG9GTP superimposed. Human and C. elegans SMG8 are labeled the same as in a
Structural comparison shows that if SMG9 switches from GTP-bound to GDP-bound form in SMG1C, the helix α8 of SMG8 would move as far as ~10 Å and the G-like domain of SMG8 would rotate by ~10° relative to SMG9 from both top and front views (Fig. 5a). The orientation of the three-helix bundle and the associated unassigned region of SMG8 would also rotate accordingly (Fig. 5b; Supplementary information, Fig. S6 and Movie S3). As a result, this unassigned region (referred to as kinase inhibitory domain, KID) would move away from the top of the catalytic pocket of the SMG1 KD.
Inhibition of SMG1 kinase activity by the SMG8 KID
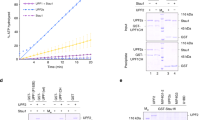
KID is in close proximity to the catalytic pocket of the SMG1 KD and partially covers the substrate entry groove (Fig. 5b, left panel). Such a conformational arrangement may provide a steric hindrance to prohibit efficient access of protein substrate, suggesting an inhibitory role of KID in regulating kinase activity of SMG1. The EM map reveals that KID is very likely derived from the C-terminus of SMG8 because it directly extends out from the three-helix bundle (Fig. 5b and Supplementary information, Fig. S6). To test this hypothesis, we performed an in vitro kinase assay using purified SMG1 with full-length human UPF1 as a substrate. The kinase activity of SMG1 largely decreased in the presence of increasing amounts of wild-type SMG8–SMG9 (Fig. 6a, lanes 1–5). In contrast, SMG8–SMG9 with the deletion of SMG8 C-terminus (residues 643–991), refers to as SMG8ΔKID-SMG9, still binds SMG1 (Supplementary information, Fig. S7b, lanes 6–8), but has no obvious inhibitory effect (Fig. 6a, lanes 6–7). There is no obvious change in the interaction between UPF1 and SMG1 in the presence of SMG8–SMG9 or SMG8ΔKID–SMG9 (Supplementary information, Fig. S7b, lanes 10–12), suggesting that UPF1 and SMG8–SMG9 may be recruited to distinct sites of SMG1. Therefore, we speculate that the access of UPF1 phosphorylation site (S1096) to SMG1 catalytic pocket is blocked by the KID of SMG8 due to steric hindrance.
In vitro kinase assays. a, b The effect of various SMG1 and SMG8–SMG9 deletions (a) and mutants (b) on UPF1 phosphorylation. In vitro kinase assays were performed using various purified full length and deletions of SMG1 and SMG8–SMG9 with purified human full-length UPF1 as a substrate. The reactions were subjected to SDS-PAGE followed by western blotting using antibody against phospho-S1096 of UPF1 (top panels). The reactions (~4-fold amounts of that in western blotting) were subjected to SDS-PAGE and the input proteins were visualized by silver staining. SMG1/SMG1ΔC (2.5 nM) and the full-length UPF1 (38.9 nM) were used in all the reactions. The relative activities were quantified by densitometry calculations of western blotting results and shown below each blot. In a, increasing amounts (2.5 nM, 4.9 nM and 14.8 nM) of SMG8–SMG9 (wild type) were used in the reactions in lanes 3–5 and 10–12, respectively. Increasing amounts (2.5 nM and 4.9 nM) of SMG8ΔKID–SMG9 were used in the reactions in lanes 6–7 and 13–14, respectively. In b, increasing amounts (0.8 nM, 2.5 nM and 4.9 nM) of individually purified SMG8 and SMG9 mutants were used in the reactions, respectively
In line with the previous study,25 the deletion of SMG1 C-insertion (residues 2432–3600) largely increases the kinase activity of SMG1 (Fig. 6a, lanes 1–2, 8–9), indicating that the C-insertion inhibits SMG1 kinase activity. Interestingly, similar to SMG1 full-length protein, the kinase activity of SMG1ΔC was inhibited by SMG8–SMG9 but not SMG8ΔKID–SMG9 (Fig. 6a, lanes 10–14). Thus, SMG1 C-insertion and SMG8 KID serve as two independent inhibitory elements to regulate SMG1 kinase activity.
To test the effect of SMG9 nucleotide-binding status on the regulation of SMG1 kinase activity, we performed in vitro kinase assay using four SMG9 GTP binding defective mutants (S218A, S218E, S218N, D269A). Surprisingly, these SMG9 mutants largely decreased their interactions with SMG8 (Supplementary information, Fig. S8). The addition of individually purified wild-type SMG8 and SMG9 mutants showed no obvious inhibition on SMG1 kinase activity (Fig. 6b). C. elegans SMG8 and SMG9 form stable heterodimer in the absence of GTP or GDP.28 Thus, these mutations may impair SMG8–SMG9 interaction. Human SMG8–SMG9 heterodimer might be well maintained if GTP hydrolysis occurs. Nevertheless, whether and how GTP hydrolysis of SMG9 occurs in physical condition, and if yes, how it affects SMG8 association in the SMG1C complex await further investigation.
Discussion
The structural and biochemical analyses indicate that SMG8–SMG9 binds to SMG1 so that the KID (C-terminal part of SMG8) provides steric hindrance to block substrate entry and inhibits SMG1 kinase activity (Fig. 7). The SMG1 kinase activity might be regulated upon GTP hydrolysis of SMG9 triggered by a GTPase-activating protein (GAP) yet to be discovered. The kinase activity of SMG1 could be promptly and precisely regulated by GTP hydrolysis and GDP-GTP exchange. Such a GTPase-mediated regulatory mechanism is distinct from that of other PIKK kinases and may be consistent with its functional roles in the regulation of NMD processes. Previous study shows that SMG1 C-insertion inhibits SMG1 kinase activity.25 We demonstrate that SMG1 C-insertion and SMG8 KID independently inhibit SMG1 activity, suggesting that two upstream signals independently and/or collectively control the activation level of SMG1 kinase.
A working model for the regulation of SMG1 kinase activity. Right panel: schematic representation of an inactive conformation of SMG1C as shown in this work. The KID domain is right above the catalytic pocket of the SMG1 KD and provides a steric hindrance for substrate entry. SMG9 is in GTP-bound form and supports the position of KID relative to KD. Left panel: schematic representation of an active complex in which SMG9 is in GDP-bound form. KID in this conformation is away from the catalytic pocket. The conformational switches might be regulated by decreased SMG8–SMG9 interaction, GAP (GTPase-activating protein) and GEF (Guanine nucleotide exchange factor) of SMG9 yet to be discovered
Interestingly, both mTOR and SMG1 are regulated by GTPases and SMG1 is structurally more similar to mTOR than other PIKK kinases. The GTP-bound small GTPase RHEB (Ras homolog enriched in brain) allosterically activates mTORC1 by realigning active-site residues of mTOR for catalysis.30 In contrast, our structure suggests that the GTP-bound SMG9 allosterically inhibits SMG1 kinase activity by fixing the KID domain of SMG8 in the conformation that blocks substrate access.
A SURF complex consisting of SMG1, UPF1, eRF1, and eRF3, has been proposed to connect NMD and translation termination, two tightly coupled processes.10 The GTPase activity of the translational GTPase eRF3 is essential for efficient translation termination. Whether, and if yes how, the two GTPases, eRF3 and SMG9, act together to regulate translation termination and NMD activation needs further investigation.
SMG1 mediates phosphorylation of an RNA helicase UPF1 that serves as a key player in the NMD pathway. Previous studies show that the C-terminal insertion domain of SMG1 interacts with SMG8 and SMG9 and promotes high-affinity UPF1 binding to SMG1C.25 However, we did not observe the C-terminal insertion domain according to the EM map. The structure of SMG1 in complex with UPF1 and other binding partners may provide the underlying mechanisms for substrate recruitment and phosphorylation.
Materials and methods
Protein expression and purification
The ORFs of human SMG1, SMG8 and SMG9 were individually subcloned into a modified pMLink vector, which contains N-terminal FLAG tag and Protein A tag. Preparation of SMG1C complex was performed as previously reported.25 Three plasmids were co-transfected into 293 F suspension cells. After being cultured at 37 °C for 3 days, cells were harvested and lysed in buffer containing 50 mM HEPES (pH 8.0), 450 mM NaCl, 10% glycerol and 3 mM DTT supplemented with 1 mM PMSF, 1 μg/mL Aprotinin, 1 μg/mL Pepstatin, 1 μg/mL Leupeptin at 4 °C. The supernatant was incubated with IgG beads for 2 h, and unbound proteins were extensively washed away with buffer containing 50 mM HEPES (pH 8.0), 150 mM NaCl, 0.05% CHAPS and 3 mM DTT. The fusion proteins were digested using TEV protease overnight to remove tags and the complex was eluted with a buffer containing 20 mM HEPES (pH 8.0), 200 mM NaCl, 0.05% CHAPS and 3 mM DTT. The eluted proteins were further purified using an anion exchange column (Source Q) in HEPES (pH 8.0) buffer with increasing concentrations of NaCl from 0 to 1000 mM. The SMG8–SMG9 complex in excess was eluted when NaCl concentration reached 150–250 mM, whereas the SMG1C complex was eluted when NaCl concentration reached 250–350 mM. The peak fractions of SMG1C were collected, concentrated for EM sample preparation and stored at −80 °C for kinase assays. SMG1, SMG8–SMG9, SMG8ΔKID–SMG9 and full-length UPF1 were similarly prepared. SMG8ΔKID represents SMG8 containing residues 1–642.
To prepare SMG1C complex sample for EM study, the peak fractions of Source Q was further treated by gradient fixation (Grafix).38 The gradient was generated from a light solution containing 10% (v/v) glycerol, 20 mM HEPES (pH 8.0), 300 mM NaCl, 3 mM DTT, and a heavy solution containing 30% (v/v) glycerol, 20 mM HEPES (pH 8.0), 300 mM NaCl, 3 mM DTT and 0.01% (v/v) glutaraldehyde. Centrifugation was performed at 38,000 rpm in a Beckman SW41Ti swinging bucket rotor for 18 h at 4 °C. The peak fractions were collected and quenched with 100 mM Tris-HCl (pH 7.4). The cross-linked SMG1C complex was concentrated to 3.0 mg/mL, further dialyzed at 4 °C for 6 h and then applied to cryo-EM grids.
The ORF of human UPF1-S1096 peptide corresponding to residues 1090–1103 (QIDVALSQDSTYQG) of UPF1 was subcloned into a modified pGEX-6P-1 vector. The plasmid was transformed into E. coli BL21 (DE3) cells and the GST fusion protein was overexpressed as previously reported.23 The fusion protein (GST-UPF1-S1096) was purified by glutathione resin and eluted by an elution buffer containing 20 mM HEPES (pH 8.0), 150 mM NaCl, 3 mM DTT, 10 mM GSH. It was further purified by gel filtration chromatography using Superdex 75 (10/300 GL) in a buffer containing 20 mM HEPES (pH 8.0), 150 mM NaCl and 3 mM DTT. The peak fractions were collected and stored at −80 °C for in vitro kinase assay.
Pull-down assay
Supernatant of 4×proA-tagged SMG1 were incubated with IgG for 2 h at 4 °C and immobilized to the indicated resin before pull-down assay. In the IgG pull-down assay, the purified full-length UPF1 (0.28 µM), SMG8–SMG9 (wild type) (0.3 µM), SMG8ΔKID–SMG9 (0.39 µM) and SMG8–SMG9 (S218E) (0.3 µM) were added and incubated with IgG-4×proA-tagged SMG1 (0.033 µM) or IgG resin in 200 µL binding buffer containing 50 mM HEPES (pH 7.4), 150 mM NaCl, 3 mM DTT, 0.01% CHAPS and 10% (v/v) glycerol for 2 h at 4 °C, respectively. After being washed for 3 times with binding buffer, the bound proteins were subjected to SDS-PAGE and stained with Coomassie blue.
In vitro kinase assays
In vitro kinase assays were performed in a reaction buffer containing 20 mM HEPES (pH 7.5), 50 mM NaCl, 10% (w/v) glycerol, 1 mM DTT, 0.1% BSA (w/v), 10 mM MnCl2.27,39 The purified full-length UPF1 and GST-UPF1-S1096 serves as substrates,27 and the purified SMG1 or SMG1C complex was used as a kinase. Reactions were initiated by the addition of 5 mM ATP, and incubated for 15 min at 30 °C. The reactions were terminated by the addition of SDS-PAGE sample loading buffer and analyzed by western blotting with antibodies against phospho-p53-S15 (ABclonal, Cat# AP0083), which has been used to target phospho-UPF1-S1096.40,41
Co-immunoprecipitation
The plasmids containing the ORFs of SMG8 and SMG9 were co-transfected to 293 F cells. After being cultured at 37 °C for 3 days, the cells were harvested and lysed in lysis buffer containing 50 mM HEPES (pH 7.4), 150 mM NaCl, 0.1% CHAPS, 3 mM DTT, 10% (v/v) glycerol and protease inhibitor cocktail. Supernatants were incubated with IgG resin at 4 °C for 2 h. After being washed for three times with lysis buffer, the immobilized proteins were subjected to SDS-PAGE and visualized by Coomassie blue staining.
EM grid preparation
Negative staining was used to evaluate the protein quality. In brief, 5 µL of freshly purified SMG1 or SMG1C complex (~50 nM) was applied onto copper grids supported by a thin layer of glow-discharged carbon film (Zhongjingkeyi Technology Co., Ltd). After adsorption for 1 min, uranyl acetate (2% w/v) was used for negative staining at room temperature. The negative stained grid was examined using FEI Talos L120C operated at 120 kV.
Cryo-grid preparation was performed using FEI Vitrobot mark IV operated at 9 °C and 100% humidity. Aliquots of 4 µL of freshly purified SMG1 or SMG1C complex (~1 mg/mL) was applied to glow-discharged holey carbon grids (Quantifoil Au, R1.2/1.3, 300 mesh). The grids were blotted using filter paper and then flash-plunged into liquid ethane pre-cooled in liquid nitrogen.
EM data acquisition
For SMG1, cryo-EM data collection was performed at Center of Cryo-Electron Microscopy (CCEM), Zhejiang University. All the cryo-images were recorded on FEI Titan Krios by Gatan K2 Summit camera operated in counting mode using SerialEM.42 The magnification is 29,000, corresponding to a final pixel size of 1.01 Å. For SMG1C, data collection was performed at the Institute of Biophysics of the Chinese Academy of Sciences. All the cryo-images were recorded on FEI Titan Krios by Gatan K2 Summit camera operated in super-resolution mode using SerialEM.42 The magnification is 22,500, corresponding to a final pixel size of 1.04 Å by applying a binning factor of 2. For each image stack, a total dose of about 50 electrons were equally fractioned into 32 frames. Defocus values was set in a range of −1.8 to −2.5 μm for data collection.
Image processing
MotionCor2s43 was used to pre-process the movie stack through all frames. The contrast transfer function parameters were estimated by Gctf from non-dose-weighted micrographs and other procedures of data processing were performed with RELION 3.0 using the dose-weighted micrographs.44,45 The data processing was shown in Supplementary information, Fig. S2. For SMG1, a reported 3.63 Å resolution (corrected gold-standard FSC 0.143 criteria) map was generated after B-factor sharpening. For SMG1C complex, a 3.43 Å (corrected gold-standard FSC 0.143 criteria) map was generated as the final reconstruction. We also performed further refinement with different local masks but did not yield better density maps. Chimera46 was used to visualize and evaluate the 3D maps, and the local resolution map was calculated using ResMap.47
Model building
The atomic model of SMG1C complex was built with COOT48 using the cryo-EM structure of mTOR (PDB ID: 5WBY)30 and the crystal structure of C. elegans SMG8–SMG9 complex (PDB ID: 5NKK)28 as references. The GTP molecule was manually built into the corresponding density. The model of SMG1 in the SMG1C complex was used as initial model for building model of SMG1 monomer. Structures were refined with phenix.real_space_refine.49
Data availability
The cryo-EM maps have been deposited in the EM Databank under accession numbers: EMD-0836 (SMG1) and EMDB-0837 (SMG1–SMG8–SMG9). The corresponding models have been deposited in the Protein Data Bank as PDB IDs: 6L53 (SMG1) and 6L54 (SMG1–SMG8–SMG9).
References
Holbrook, J. A., Neu-Yilik, G., Hentze, M. W. & Kulozik, A. E. Nonsense-mediated decay approaches the clinic. Nat. Genet. 36, 801–808 (2004).
Chang, Y. F., Imam, J. S. & Wilkinson, M. E. The nonsense-mediated decay RNA surveillance pathway. Annu. Rev. Biochem. 76, 51–74 (2007).
Popp, M. W. & Maquat, L. E. Organizing principles of mammalian nonsense-mediated mRNA decay. Annu Rev. Genet. 47, 139–165 (2013).
Maquat, L. E. Nonsense-mediated mRNA decay: splicing, translation and mRNP dynamics. Nat. Rev. Mol. Cell Biol. 5, 89–99 (2004).
Popp, M. W. & Maquat, L. E. Leveraging rules of nonsense-mediated mRNA decay for genome engineering and personalized medicine. Cell 165, 1319–1322 (2016).
Maquat, L. E., Kinniburgh, A. J., Rachmilewitz, E. A. & Ross, J. Unstable beta-globin mRNA in mRNA-deficient beta o thalassemia. Cell 27, 543–553 (1981).
Behm-Ansmant, I. et al. mRNA quality control: an ancient machinery recognizes and degrades mRNAs with nonsense codons. FEBS Lett. 581, 2845–2853 (2007).
Lykke-Andersen, S. & Jensen, T. H. Nonsense-mediated mRNA decay: an intricate machinery that shapes transcriptomes. Nat. Rev. Mol. Cell Biol. 16, 665–677 (2015).
Karousis, E. D., Nasif, S. & Muhlemann, O. Nonsense-mediated mRNA decay: novel mechanistic insights and biological impact. Wiley Interdiscip. Rev. RNA 7, 661–682 (2016).
Karousis, E. D. & Muhlemann, O. Nonsense-mediated mRNA decay begins where translation ends. Cold Spring Harb. Perspect. Biol. 11, a032862 (2019). pii.
Nickless, A., Bailis, J. M. & You, Z. S. Control of gene expression through the nonsense-mediated RNA decay pathway. Cell Biosci. 7, 26 (2017).
Guan, Q. et al. Impact of nonsense-mediated mRNA decay on the global expression profile of budding yeast. PLoS Genet. 2, e203 (2006).
Liu, C. et al. The UPF1 RNA surveillance gene is commonly mutated in pancreatic adenosquamous carcinoma. Nat. Med. 20, 596–598 (2014).
Tarpey, P. S. et al. Mutations in UPF3B, a member of the nonsense-mediated mRNA decay complex, cause syndromic and nonsyndromic mental retardation. Nat. Genet. 39, 1127–1133 (2007).
Nguyen, L. S. et al. Transcriptome profiling of UPF3B/NMD-deficient lymphoblastoid cells from patients with various forms of intellectual disability. Mol. Psychiatry 17, 1103–1115 (2012).
Nguyen, L. S. et al. Contribution of copy number variants involving nonsense-mediated mRNA decay pathway genes to neuro-developmental disorders. Hum. Mol. Genet. 22, 1816–1825 (2013).
Mort, M., Ivanov, D., Cooper, D. N. & Chuzhanova, N. A. A meta-analysis of nonsense mutations causing human genetic disease. Hum. Mutat. 29, 1037–1047 (2008).
Hug, N., Longman, D. & Caceres, J. F. Mechanism and regulation of the nonsense-mediated decay pathway. Nucleic Acids Res. 44, 1483–1495 (2016).
Lykke-Andersen, J., Shu, M. D. & Steitz, J. A. Human Upf proteins target an mRNA for nonsense-mediated decay when bound downstream of a termination codon. Cell 103, 1121–1131 (2000).
Yamashita, A. Role of SMG-1-mediated Upf1 phosphorylation in mammalian nonsense-mediated mRNA decay. Genes Cells 18, 161–175 (2013).
Jonas, S., Weichenrieder, O. & Izaurralde, E. An unusual arrangement of two 14-3-3-like domains in the SMG5-SMG7 heterodimer is required for efficient nonsense-mediated mRNA decay. Genes Dev. 27, 211–225 (2013).
Eberle, A. B., Lykke-Andersen, S., Muhlemann, O. & Jensen, T. H. SMG6 promotes endonucleolytic cleavage of nonsense mRNA in human cells. Nat. Struct. Mol. Biol. 16, 49–55 (2009).
Yamashita, A., Ohnishi, T., Kashima, I., Taya, Y. & Ohno, S. Human SMG-1, a novel phosphatidylinositol 3-kinase-related protein kinase, associates with components of the mRNA surveillance complex and is involved in the regulation of nonsense-mediated mRNA decay. Genes Dev. 15, 2215–2228 (2001).
Denning, G., Jamieson, L., Maquat, L. E., Thompson, E. A. & Fields, A. P. Cloning of a novel phosphatidylinositol kinase-related kinase - Characterization of the human SMG-1 RNA surveillance protein. J. Biol. Chem. 276, 22709–22714 (2001).
Deniaud, A. et al. A network of SMG-8, SMG-9 and SMG-1 C-terminal insertion domain regulates UPF1 substrate recruitment and phosphorylation. Nucleic Acids Res. 43, 7600–7611 (2015).
Yamashita, A. et al. SMG-8 and SMG-9, two novel subunits of the SMG-1 complex, regulate remodeling of the mRNA surveillance complex during nonsense-mediated mRNA decay. Genes Dev. 23, 1091–1105 (2009).
Arias-Palomo, E. et al. The nonsense-mediated mRNA decay SMG-1 kinase is regulated by large-scale conformational changes controlled by SMG-8. Genes Dev. 25, 153–164 (2011).
Li, L., Lingaraju, M., Basquin, C., Basquin, J. & Conti, E. Structure of a SMG8-SMG9 complex identifies a G-domain heterodimer in the NMD effector proteins. RNA 23, 1028–1034 (2017).
Melero, R. et al. Structures of SMG1-UPFs complexes: SMG1 contributes to regulate UPF2-dependent activation of UPF1 in NMD. Structure 22, 1105–1119 (2014).
Yang, H. et al. Mechanisms of mTORC1 activation by RHEB and inhibition by PRAS40. Nature 552, 368–373 (2017).
Rao, Q. et al. Cryo-EM structure of human ATR-ATRIP complex. Cell Res. 28, 143–156 (2018).
Yin, X., Liu, M., Tian, Y., Wang, J. & Xu, Y. Cryo-EM structure of human DNA-PK holoenzyme. Cell Res. 27, 1341–1350 (2017).
Chen, X. et al. Cryo-EM structure of human mTOR complex 2. Cell Res. 28, 518–528 (2018).
Yang, H. et al. 4.4 Å Resolution cryo-EM structure of human mTOR Complex 1. Protein Cell 7, 878–887 (2016).
Yang, H. et al. mTOR kinase structure, mechanism and regulation. Nature 497, 217–223 (2013).
Sibanda, B. L., Chirgadze, D. Y., Ascher, D. B. & Blundell, T. L. DNA-PKcs structure suggests an allosteric mechanism modulating DNA double-strand break repair. Science 355, 520–524 (2017).
Mishra, A. K. & Lambright, D. G. Invited review: Small GTPases and their GAPs. Biopolymers 105, 431–448 (2016).
Kastner, B. et al. GraFix: sample preparation for single-particle electron cryomicroscopy. Nat. Methods 5, 53–55 (2008).
Clerici, M. et al. Structural and functional analysis of the three MIF4G domains of nonsense-mediated decay factor UPF2. Nucleic Acids Res. 42, 2673–2686 (2014).
Kim, S. T., Lim, D. S., Canman, C. E. & Kastan, M. B. Substrate specificities and identification of putative substrates of ATM kinase family members. J. Biol. Chem. 274, 37538–37543 (1999).
Page, M. F., Carr, B., Anders, K. R., Grimson, A. & Anderson, P. SMG-2 is a phosphorylated protein required for mRNA surveillance in Caenorhabditis elegans and related to Upf1p of yeast. Mol. Cell Biol. 19, 5943–5951 (1999).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. Elife 7, e42166 (2018).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Pettersen, E. F. et al. UCSF Chimera-a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, (213–221 (2010).
Acknowledgements
We thank Center of Cryo-Electron Microscopy, Zhejiang University School of Medicine, Center for Biological Imaging of Institute of Biophysics of Chinese Academy of Sciences, and National Center for Protein Science Shanghai for the supports on cryo-EM data collection and data analyses. We thank the Biomedical Core Facility, Fudan University for the supports on Mass Spectrometry analyses. This work was supported by the National key R&D program of China (2016YFA0500700), the National Natural Science Foundation of China (31830107, 31821002, 31425008), the National Ten-Thousand Talent Program (Y.X.), the National Program for support of Top-Notch Young Professionals (Y.X.), and the Strategic Priority Research Program of the Chinese Academy of Sciences (XDB08000000).
Author information
Authors and Affiliations
Contributions
L.Z., L.L., Y.Q., and Y.X. designed the experiments. L.Z. purified the proteins and performed biochemical analyses. Y.Q. prepared the cryo-EM sample, collected the data and determined the structure. Y.Q. and L.L. built the structural model. L.Z., L.L., Y.Q., and Y. X. analyzed the data and wrote the manuscript. Y.X. supervised the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Zhu, L., Li, L., Qi, Y. et al. Cryo-EM structure of SMG1–SMG8–SMG9 complex. Cell Res 29, 1027–1034 (2019). https://doi.org/10.1038/s41422-019-0255-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41422-019-0255-3
This article is cited by
-
Messenger RNA Surveillance: Current Understanding, Regulatory Mechanisms, and Future Implications
Molecular Biotechnology (2024)
-
Identification of a novel compound heterozygous SMG9 variants in a Chinese family with heart and brain malformation syndrome using whole exome sequencing
BMC Medical Genomics (2022)
-
Residue-wise local quality estimation for protein models from cryo-EM maps
Nature Methods (2022)
-
The activation mechanisms of master kinases in the DNA damage response
Genome Instability & Disease (2021)