Abstract
Despite the structural conservation of PTEN with dual-specificity phosphatases, there have been no reports regarding the regulatory mechanisms that underlie this potential dual-phosphatase activity. Here, we report that K27-linked polyubiquitination of PTEN at lysines 66 and 80 switches its phosphoinositide/protein tyrosine phosphatase activity to protein serine/threonine phosphatase activity. Mechanistically, high glucose, TGF-β, CTGF, SHH, and IL-6 induce the expression of a long non-coding RNA, GAEA (Glucose Aroused for EMT Activation), which associates with an RNA-binding E3 ligase, MEX3C, and enhances its enzymatic activity, leading to the K27-linked polyubiquitination of PTEN. The MEX3C-catalyzed PTENK27-polyUb activates its protein serine/threonine phosphatase activity and inhibits its phosphatidylinositol/protein tyrosine phosphatase activity. With this altered enzymatic activity, PTENK27-polyUb dephosphorylates the phosphoserine/threonine residues of TWIST1, SNAI1, and YAP1, leading to accumulation of these master regulators of EMT. Animals with genetic inhibition of PTENK27-polyUb, by a single nucleotide mutation generated using CRISPR/Cas9 (PtenK80R/K80R), exhibit inhibition of EMT markers during mammary gland morphogenesis in pregnancy/lactation and during cutaneous wound healing processes. Our findings illustrate an unexpected paradigm in which the lncRNA-dependent switch in PTEN protein serine/threonine phosphatase activity is important for physiological homeostasis and disease development.
Similar content being viewed by others
Introduction
Phosphatase and tensin homolog (PTEN) is regarded as one of the most important tumor suppressors.1 Although PTEN has high structural conservation with dual-specificity phosphatases (DUSPs),2 research has been predominately focused on its phosphatase activity towards phosphatidylinositol 3,4,5 trisphosphate (PIP3).3 PTEN converts cellular PIP3 into phosphatidylinositol 4,5-bisphosphate (PIP2) and downregulates the protein kinase B (AKT/PKB) pathway.4 In many cancer types, genetic deletion of PTEN leads to elevated cellular PIP3 levels and hyper-activation of AKT.5 However, mice with transgenic expression of activated AKT exhibit a different spectrum of tumor development.6 Therefore, PTEN may possess biological roles other than PIP3 dephosphorylation, such as protein phosphatase activity, which may be involved in regulating homeostasis and human disease.
Recent advances in genomic sequencing demonstrate that long non-coding RNA (lncRNA) molecules are pervasively transcribed and are likely to be important in mediating cellular functions.7 Numerous mega-analyses have indicated that lncRNAs may play critical roles in diverse cellular processes and diseases, including human cancers.8 Genetically, mutations occur frequently in the non-coding regions of the genome, altering the primary sequences of many non-coding RNAs, which is one of the causes of genetic diseases.9 Transcriptionally, dysregulation of lncRNA expression, altered splicing patterns, and microRNA-mediated lncRNA stabilization implicate the importance of lncRNAs in cells.10 Biologically, it has been suggested that lncRNAs help control cell growth, apoptosis, DNA-damage repair, stem cell self-renewal/differentiation,11 and mRNA stability.12
Epithelial–mesenchymal transition (EMT) is hallmarked by a loss of cell–cell contact between epithelial cells and acquisition of invasive properties as mesenchymal phenotypic cells, which is important for tissue repair,13 organ fibrosis,14 cancer metastasis,15 and chemoresistance.16 It has been suggested that transcription factors that regulate EMT, including TWIST1, SNAI1 and YAP, are involved in breast ductal network development.17 However, the underlying mechanisms that regulate TWIST1, SNAI1 and YAP1 during EMT, upon extracellular stimuli, remain largely unknown. Our data shows that high glucose and growth factors including TGF-β, CTGF, SHH, and IL-6 triggered the expression of a lncRNA glucose aroused for EMT activation (GAEA), which bound the RNA-binding E3 ligase MEX3C and facilitated its ligase activity. The activated MEX3C catalyzed K27-linked polyubiquitination (polyUb) of PTEN at lysines 66 and 80 (PTENK27-PolyUb), converting PTEN from a phosphoinositide/tyrosine phosphatase to a serine/threonine phosphatase. PTENK27-polyUb removed phospho-groups from serine/threonine residues in substrates including TWIST1, SNAI1, and YAP1. Upon dephosphorylation, these proteins were stabilized and promoted EMT in a PTEN-dependent manner. Homozygous mice harboring the Pten K80R mutation exhibited impaired EMT during mammary gland morphogenesis in pregnant/lactating mice and during cutaneous wound healing processes. Our results reveal an unappreciated dimension of PTEN function as a protein serine/threonine phosphatase in homeostasis and pathological conditions, with clinical implications.
Results
GAEA enhances the enzymatic activity of MEX3C
Elevated glucose metabolism promotes cancer cell proliferation, cell cycle, and angiogenesis partially through transcriptional regulation.18 To understand the molecular mechanism of glucose-induced EMT, we first determined transcriptional alterations in breast cancer cells that were treated with physiological glucose concentrations (5.0 mM) or high glucose (25 mM). Gene expression profiling indicated that in addition to protein coding genes, transcription of 588 non-coding RNAs were upregulated more than two-fold by high glucose (Fig. 1a, GEO ID: GSE106729). LOC84856 (also named LINC00839 is renamed GAEA) was one of the top ten lncRNA candidates induced by high glucose (Fig. 1a), but not by reintroduction of amino acids or serum (Supplementary Information, Fig. S1a–d). The copy number, transcript size and sgRNA-mediated knockout of GAEA was characterized (Supplementary Information, Fig. S1e–g). A number of transcription factors might be involved in the transcriptional regulation of GAEA upon these stimuli (Supplementary Information, Table S1).
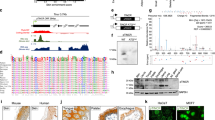
Identification and characterization of GAEA-MEX3C interaction. a Heatmap showing differently expressed coding and non-coding genes in MDA-MB-231 cells with glucose starvation followed by low (5 mM) or high (25 mM) glucose (Glc) stimulation. Red/green indicates increased/decreased fold change. R1, R2 and R3 (biological repeat 1, 2 and 3). b A list of top GAEA-associated proteins identified by RNA pull-down followed by MS analysis in MDA-MB-231 cells. c–g Quantitative measure of poly-Ub chain formation in MEX3C ubiquitin ligase activity assays with indicated recombinant proteins, in the presence indicated ubiquitin (e, f) or MEX3C (g). Error bars, SD. n = 3 independent experiments, Student’s t-test. h RIP-qPCR detection of indicated RNAs retrieved by indicated antibodies in MDA-MB-231 cells. Error bars, SD. n = 3 independent experiments, Student’s t-test. i In vitro RNA–protein binding coupled with dot-blot assays. j RNA EMSA analysis of recombinant MEX3C binding to 1801–1860 fragment of GAEA. RNA EMSA was performed using indicated recombinant protein and biotinylated DNA or RNA probes corresponding to GAEA n.t. 1801–1860 or AR 3’-UTR (32P-labeled, blue, lanes 1-11) or in the presence of cold probes as competitors (green, lanes 12–15). k Immunoblotting (IB) detection using indicated antibodies of MCF10A cells harboring scramble or MEX3C sgRNAs and indicated expression vectors, followed by glucose stimulation (5 or 25 mM). l Quantitative measure of poly-Ub chain formation in MEX3C ubiquitin ligase activity assays with indicated E1–E2–E3 proteins and RNA transcripts. LINK-A was used as a negative lncRNA control and UBE2D3-MDM2 as a negative E2–E3 control. Error bars, SD, n = 3 independent experiments, one-way ANOVA. (n.s. p > 0.05, *p < 0.05, **p < 0.01, and ***p < 0.001)
Mass spectrometry (MS) analysis indicated the proteins that specifically interacted with sense GAEA in cells, including the RNA-binding E3 ligase MEX3C, ubiquitin-conjugating enzyme E2 S (UBE2S), PTEN, and YAP1 (Fig. 1b; Supplementary Information, Table S2). First, we characterized the enzymatic activity of MEX3C in the presence of ubiquitin-activating enzymes (E1, UBE1, also known as UBA1) and ubiquitin-conjugating enzymes (E2, UBE2S). MEX3C catalyzed polyUb chain formation in vitro, which was dependent on Gly76 of ubiquitin (Ub) (Fig. 1c). UBE2D3-MDM2 that catalyzes ubiquitination of p53 served as the positive control.19 Neither UBE2D3-MEX3C nor UBE2S-MDM2 robustly catalyzed the formation of polyUb chains (Fig. 1d). K27R ubiquitin abolished MEX3C-catalyzed polyUb chains, while the ubiquitin mutant with only lysine 27 remaining was as competent as WT Ub in forming polyUb chains (Fig. 1e, f; Supplementary Information, Fig. S1h). Depletion of the RING-type zinc finger domain of MEX3C (ΔZNF) and point mutations, C608S or H625A,20 abolish the E3 ligase activity of MEX3C (Fig. 1g; Supplementary Information, Fig. S1i).
MEX3C directly associated with GAEA in vivo and in vitro (Fig. 1h; Supplementary Information, Fig. S2a, b). Gly343 of the KH2 domain is critical for MEX3C-GAEA binding (Supplementary Information, Fig. S2c). Upon high glucose stimulation, roughly 3000 copies of GAEA may associate with ∼4000 MEX3C protein molecules per MCF10A cell (Supplementary Information, Fig. S2d, e). We then mapped the sequence motif of GAEA, finding nt. 1801–1860 was required for MEX3C binding (Fig. 1i). GAEA U1825G/C1850G (referred as GAEA mut), disrupted the MEX3C-GAEA interaction in vitro (Supplementary Information, Fig. S2f). RNA probe corresponding to GAEA nt. 1801–1860 but not the corresponding DNA probe interacted with WT MEX3C (Fig. 1j, lane 6). The MEX3C G343D mutant or GAEA mut failed to mediate the interaction (lanes 9 and 11). Furthermore, the presence of unlabeled RNA probes corresponding to nt. 1801–1860 of GAEA, but not androgen receptor 3’-UTR (AR 3’-UTR) RNA, competed with biotinylated GAEA RNA probes in binding with MEX3C (Fig. 1j, lane 13). In summary, our data indicate that G343 of MEX3C’s KH2 domain and U1825/C1850 of GAEA are critical for the interaction between GAEA and MEX3C.
The saturation curve experiments indicated that GAEA-MEX3C interaction is U1825/C1850-dependent (Supplementary Information, Fig. S2g). Neither the DNA sequence of GAEA nor the AR 3’-UTR exhibited detectable interaction with MEX3C (Supplementary Information, Fig. S2g). Competition assay indicated that 14.3 nM of the unlabeled GAEA (nt. 1801–1860) achieved 50% inhibition of the interaction between biotin-GAEA and His-tagged MEX3C (Supplementary Information, Fig. S2h), suggesting a high binding affinity between GAEA and MEX3C.
We reason that the association between MEX3C and a variety of E2 enzymes may modulate the linkage types of the polyUb chain. To address this hypothesis, we selected four E2 enzymes: UBE2B mediates K48-linked polyUb21; the UBE2N/V1 complex catalyzes K63-linked polyUb22; and UBE2S, which potentially mediates K11-23 and K27-linked polyUb. The MEX3C-deficient cells were rescued with expression of MEX3C WT or G343D (Fig. 1k; Supplementary Information, Fig. S2i). The MEX3C–UBE2B interactions are undetectable (Fig. 1k). The interaction between MEX3C and UBE2N/UBE2V under the low glucose condition, was abolished upon high glucose stimulation; MEX3C G343D mutant showed constituted interactions with UBE2N/UBE2V despite the glucose concentration (Fig. 1k). On the contrary, high glucose induced the interactions between UBE2S and endogenous MEX3C or exogenous MEX3C WT; this interaction was abolished when MEX3C G343D was expressed (Fig. 1k). Expression of GAEA wild type (WT), but not mutant, in GAEA-deficient cells, rescued the interaction between MEX3C and UBE2S upon high glucose stimulation (Supplementary Information, Fig. S2j). These observations suggested that the presence of GAEA may modulate the E2–E3 pair complex formation, leading to higher binding affinity between MEX3C and UBE2S to catalyze the K27-linked polyubiquitination.
In the presence of full-length GAEA, WT MEX3C exhibited more than six-fold enhancement of its ability to catalyze the polyubiquitin chain formation (green bar) (Fig. 1l). In contrast, the GAEA mutant failed to do so (purple bar). The E3 ligase activity of the MEX3C G343D mutant and MDM2 were not affected by the presence of lncRNAs (Fig. 1l). We confirmed these observations using anti-ubiquitin or anti-K27-polyUb specific antibody (lanes 8/14 compared to lanes 7/13 respectively) (Supplementary Information, Fig. S2k). Therefore, our data indicate that GAEA directly associates with MEX3C and enhances its E3 ligase activity in vitro.
PTEN is required for glucose- and growth factors-induced EMT
We confirmed that endogenous PTEN associated with previously reported binding proteins1 (Fig. 2a, green). Upon glucose stimulation, PTEN associated with UBA1, UBE2S, MEX3C, and ubiquitin, which overlaps with GAEA-binding proteins (Fig. 2a; Supplementary Information, Table S3). PTEN was ubiquitinated at K66 and K80 residues (Supplementary Information, Fig. S3a, b). We then synthesized peptides that correspond to the sequences flanking PTEN K66 or K80 with or without branching GG to mimic ubiquitination and generated antibodies referred to as Ub-PTEN (K66) or Ub-PTEN (K80), respectively (Supplementary Information, Fig. S3c, d).
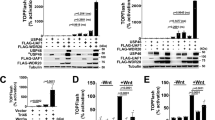
K27-linked polyubiquitination of PTEN promotes EMT. a A list of top PTEN-binding proteins identified by immunoprecipitation and MS analysis in MCF10A PTEN+/+ with or without glucose treatment (5 or 25 mM, 1 h). b Quantitative measure of PTENK27-polyUb using Ub-PTEN (K80) antibody in MCF10A PTEN+/+ cells under the glucose and a panel of indicated growth factor stimulations for 1 h. Statistically significant factors are highlighted as Red. Error bars, SD, n = 3 independent experiments, Student’s t-test. c, d IB detection of PTEN ubiquitination in MCF10A PTEN+/+ and NMuMG cells (c) under the indicated growth factor stimulation for 1 h or in MCF10A PTEN+/+ and MCF10A PTEN-/- cells (d). e Immunofluorescence detection of EMT markers in 3D culture of MCF10A PTEN+/+ and MCF10A PTEN-/- cells, followed by glucose starvation and stimulation (5 or 25 mM, 12 days), or 2-DG (25 mM, 12 days) or in the presences of STF31 (5 µM, 12 days). Scale bars: 50 µm. f, g Immunofluorescence detection using indicated antibodies in MCF10A PTEN+/+ or MCF10A PTEN-/- cells transfected with the indicated expression vectors followed by indicated treatment for 72 h. Scale bars: 50 µm. h Immunofluorescence detection using indicated antibodies in MCF10A PTEN+/+ or MCF10A PTEN-/- cells transducted with lentivirus harboring indicated expression vectors. Scale bars: 50 µm. i Immunofluorescence detection using indicated antibodies in MCF10A PTEN+/+ cells harboring indicated sgRNAs with expression of indicated expression vectors, followed by indicated treatment for 72 h. Scale bars: 50 µm (n.s. p > 0.05, *p < 0.05, **p < 0.01, and ***p < 0.001)
High glucose has been considered a proinflammatory signal.24 Therefore, we screened whether other growth factors/cytokines could induce PTEN polyUb, of which we focused on Ub-PTEN (K80). In addition to high glucose, connective tissue growth factor (CTGF), interleukin 6 (IL-6), sonic hedgehog (SHH) and transforming growth factor beta 1 (TGF-β1), which are central mediators of EMT and cancer metastasis,25,26,27,28 effectively induced polyUb of PTEN as recognized by Ub-PTEN (K80) (Fig. 2b–d).
We hypothesized that these proinflammatory signals might promote EMT. We compared the immortalized human breast basal epithelial MCF10A cells that had either wild type PTEN (PTEN+/+) or that had been genetically engineered to be PTEN deficient (PTEN-/-), under the condition of low glucose (5 mM) or high glucose (25 mM), finding that PTEN is required for glucose-induced EMT (Fig. 2e; Supplementary Information, Fig. S3e, f). The presence of 2DG (2-deoxy-d-glucose) or the glucose transporter inhibitor STF3129 blocked glucose-induced EMT. In MCF10A PTEN-/- cells, high glucose-induced EMT was abolished, suggesting the requisite role of PTEN in promoting glucose-induced EMT (Fig. 2e; Supplementary Information, Fig. S3e, f).
In MCF10A PTEN-/- cells, expression of PTEN WT restored glucose-, TGF-β-, CTGF-, SHH- or IL-6-induced EMT, while expressing the PTEN K66/80R mutant failed to do so (Fig. 2f, g; Supplementary Information, Fig. S3g, h). Transduction of WT GAEA, but not the U1825G/C1850G mutant, induced EMT in MCF10A cells (Fig. 2h; Supplementary Information, Fig. S3i, j). Similarly, expression of MEX3C WT, but not G343D mutant in MEX3C-depleted cells restored EMT upon stimulation (Fig. 2i). These findings indicated the importance of GAEA/MEX3C and PTEN polyUb signaling axis in regulating EMT.
MEX3C catalyzes the K27-linked polyubiquitination of PTEN
MEX3C catalyzed PTEN polyUb at lysine 66 and 80 residues, which was abolished upon expression of PTEN K66/80R, ubiquitin G76A or K27R mutants (Fig. 3a; Supplementary Information, Fig. S4a). Knockdown of GAEA or MEX3C reduced the glucose-induced PTEN polyUb, without affecting global or total ubiquitination of PTEN (Fig. 3b; Supplementary Information, Fig. S4b, c). PTEN K66R, K80R or K66/80R mutants abolished the polyubiquitination at the corresponding sites upon high glucose stimulation (Fig. 3c). Furthermore, expression of MEX3C WT, but not G343D, led to enhanced PTEN polyUb, which is impaired upon GAEA knockdown (Fig. 3d).
MEX3C catalyzes the K27-linked polyubiquitination of PTEN. a In vitro PTEN ubiquitination assay was performed with indicated recombinant proteins, and immunoblotted with indicated antibodies. b, c IB detection using indicated antibodies in cells; which were transfected with indicated siRNAs (b) or expression constructs (c) and were glucose starved followed by glucose stimulation (5 or 25 mM, 1 h). d IB detection using indicated antibodies in cells transducted with lentivirus harboring indicated expression constructs and co-transfected with indicated siRNAs. e Silver staining detection of PTEN ubiquitination in MCF10A PTEN+/+ cells which were transfected with the indicated siRNA and glucose starved followed by glucose stimulation (5 or 25 mM, 1 h). Denatured IP was performed with indicated antibodies and indicated bands were cut to do the MS/MS spectrum. Blue/green box, gel band cut for MS analysis. f Protein score of MS are shown. g A list of MS linkage identification of PTEN and ubiquitin. The linkage type, GG-modified peptides of ubiquitin and the number of recovered peptides are shown. h In vitro ubiquitination assay was performed with indicated recombinant proteins, and the resulting samples were immunoblotted with indicated antibodies. i IB detection using indicated antibodies of cells which were transfected with indicated expression constructs and glucose starved followed by glucose stimulation (5 or 25 mM, 1 h)
To define the subtypes of polyUb chains catalyzed by MEX3C, we immunoprecipitated polyubiquitinated PTEN under denaturing conditions upon glucose stimulation for liquid chromatography–mass spectrometry (LC-MS) (Fig. 3e, f; Supplementary Information, Table S4). PTEN is confirmed to be modified with double Glycine (GG) at K66 and 80 site, and lysine 27 of ubiquitin is the major site to be modified by GG (Fig. 3g). Consistently, MEX3C catalyzed the formation of PTEN polyUb in a K27-dependent manner (referred as PTENK27-polyUb) in vitro and in cells (Fig. 3h, i). Introduction of MEX3C WT, but not the G343D mutant, rescued the MEX3C-dependent PTENK27-PolyUb and EMT (Supplementary Information, Fig. 4d). Depletion of GAEA, but not the glucose-induced lncRNA ZNFX-AS1,30 impaired glucose-induced PTENK27-PolyUb and EMT, which was rescued by expression of WT GAEA, but not the non-MEX3C-binding mutant (Supplementary Information, Fig. S4e, f). Thus, our data demonstrated that the MEX3C-catalyzed PTENK27-polyUb in vivo is required for promoting EMT.
K27-linked polyubiquitination modulates PTEN phosphatase activity. a Graphic illustration of purification of polyubiquitinated PTEN and its subsequent usage in PTEN phosphatase activity or in vitro binding assays with corresponding substrates. b A list of substrates used in PTEN phosphatase activity assays. c Intensity of phospho-substrates and dephosphorylated substrates. d–g Quantitative measure of dephosphorylation of PIP3 (d), p-Tyr (e), p-Ser (f) or p-Thr (g) in PTEN phosphatase activity assays with unmodified or polyubiquitinated PTEN (WT and mutants) at indicated concentrations. Error bars, SD, n = 3 independent experiments. h Summary of ED50 value (mean ± S.E.M, n = 3 independent experiments) of the indicated enzymes based on experiments in d–g. i Activities of PTP1B, PP2A, PTEN or polyubiquitinated PTEN toward PIP3 or phospho-Tyr, phospho-Ser, and phospho-Thr substrates were measured by monitoring the phosphate release
PTENK27-polyUb acts as protein serine/threonine phosphatase
The subcellular localization and half-life of PTEN were not affected by polyubiquitination of PTEN at both K66 and K80 (Supplementary Information, Fig. S5a, b). PTEN, PTENK27-polyUb and PTENK48-polyUb were used to dephosphorylate PIP3 and synthesized generic peptides harboring p-Tyr, p-Ser, or p-Thr,31,32,33 as substrates (Fig. 4a, b and Supplementary Information, Fig. S5c). By incubating PIP3 or p-Tyr-, p-Ser-, p-Thr-containing peptides with BSA, unmodified PTEN, or PTENK27-polyUb, the dephosphorylation of these substrates were determined by MALDI-TOF MS analysis. As expected, PIP3 (MW 981.4 Da) was converted to PIP2 (MW 901.4 Da) in the presence of PTEN; however, PTENK27-polyUb failed to do so (Fig. 4c; Supplementary Information, Fig. S6a, Table S5). The p-Tyr-containing peptide (MW 1118.4 Da) was converted to unphosphorylated form (MW 1038.4 Da) by PTEN but not by PTENK27-polyUb (Fig. 4c; Supplementary Information, Fig. S6b, Table S5). Surprisingly, PTENK27-polyUb, but not PTEN, dephosphorylated both p-Ser- and p-Thr-containing peptides (MW 1244.5 Da to 1164.5 Da; and 909.5 Da to 829.5 Da respectively) (Fig. 4c; Supplementary Information, Fig. S6c, d, Table S5).
To quantitatively measure the phosphatase activity of PTEN, PTENK27-polyUb, and PTENK48-polyUb, we performed PTEN ELISA assay to measure conversion of PIP3 to PIP2 (Fig. 4d) or in vitro dephosphorylation assays to measure the release of the organic phosphate from p-Tyr-, p-Ser-, p-Thr-containing peptides (Fig. 4e–g). We included PTP1B and PP2A as positive controls for tyrosine phosphatase34 and serine/threonine phosphatase,35 respectively. Conversion of PIP3 to PIP2 was robustly catalyzed by either PTEN (red line) or PTENK48-polyUb (purple line), but not by PTENK27-polyUb (green line) (Fig. 4d). K66R or K80R single mutants (orange and black lines respectively) dephosphorylated PIP3 as equally efficient as unmodified PTEN (Fig. 4d). Similarly, for p-Tyr containing peptides, PTEN (red line), PTENK48-polyUb (purple line), K66R mutant, and K80R mutant all showed similar activity in catalyzing this reaction. In contrast, PTENK27-polyUb (green line) failed to dephosphorylate p-Tyr-containing substrate (Fig. 4e). Further, PTENK27-polyUb (green line) but not unmodified PTEN (red line), exhibited adequate phosphatase activity toward p-Ser- and p-Thr-containing substrates (Fig. 4f, g). We calculated the median effective dose (ED50) value (nM) of each enzyme that was summarized in Fig. 4h. PTEN dephosphorylated PIP3 and p-Tyr with ED50 value of 0.72 and 11.4 nM, respectively, but had undetectable activity toward p-Ser- and p-Thr-containing substrates. In contrast, PTENK27-polyUb showed undetectable activity toward PIP3 and p-Tyr but moderate phosphatase activity toward p-Ser and p-Thr substrates (ED50 values of 17.4 and 24.7 nM, respectively) (Fig. 4h). PTEN (K66R)K27-polyUb, and PTEN (K80R)K27-polyUb functioned similarly to unmodified PTEN (Fig. 4h).
We further determined the specific phosphatase activity for each enzyme as measured in pmol of substrate/minute/µg of enzyme. 1 µg of PTEN catalyzed roughly 7.5 pmol of PIP3 or 3 pmol of p-Tyr per minute. In contrast, PTENK27-polyUb catalyzed about 0.2 pmol of PIP3, 0 pmol of p-Tyr, and 2.5–3.0 pmol of p-Ser- and p-Thr-containing peptides per minute (Fig. 4i). We further validated that the USP2, a general deubiquitination enzyme,36 was able to deubiquitinate PTENK27-polyUb, thus restoring its PIP3 and p-Tyr phosphatase activities (Supplementary Information, Fig. S6e–g). Taken together, our data indicate that K27-linked polyubiquitination of PTEN at K66 and K80 converted PTEN from a phosphoinositide/protein tyrosine phosphatase to a protein serine/threonine phosphatase.
The depth of PTEN catalytic cleft determines substrate preference
For dual-specificity phosphatases, the conserved catalytic core contains the CXXXGXXR motif, where a specific distance between C (Cys) and R (Arg) is critical for substrate binding.37 Because the polar head groups of phosphoinositol and p-Tyr are larger than that of p-Ser or p-Thr, the distance between C124 and R130 of PTEN could be essential for substrate preference.38 The 3-dimensional structure of PTEN revealed that the P loop (catalytic core) contains HC124KAGKGR130. The ~8 Å catalytic pocket of PTEN can fit p-Tyr, p-Ser, and p-Thr side chains, which is between the 9 Å deep cleft of PTP1B and the approximately 6 Å shallow cleft of PP2A.39 Therefore, we hypothesize that this might be the reason that PTEN and PTENK27-polyUb exhibit different binding affinities to PIP3, p-Tyr, p-Ser, or p-Thr substrates.
To address this hypothesis, we generated biotinylated-PIP3 and biotinylated-BSA-conjugated p-Tyr, p-Ser, and p-Thr amino acids for in vitro enzyme–substrate overlay assay (Fig. 5a). Purified His-tagged PTEN, PTENK27-polyUb, PTEN (K66R) K27-polyUb, PTEN (K80R) K27-polyUb, and PTENK48-polyUb were applied on the dot-blots to probe their binding with different substrates. It is clear that unmodified PTEN, PTEN (K66R) K27-PolyUb, PTEN (K80R) K27-PolyUb, and PTENK48-polyUb bound to PIP3 and BSA-p-Tyr; however, PTENK27-polyUb directly bound to BSA-p-Ser and BSA-p-Thr residues (Fig. 5a). Next, using an alpha assay, we observed that PTEN exhibited robust association with PIP3 and BSA-p-Tyr, moderate binding with BSA-p-Ser and -Thr, but undetectable binding with BSA alone (Supplementary Information, Fig. S7a–e).
K27-linked polyubiquitination alters PTEN substrate binding. a Enzyme–substrate overlay assay showing the binding capability of unmodified and polyubiquitinated PTEN to indicated substrates. Biotinylated PIP3 or indicated BSA-conjugated phospho-amino acids were spotted on nylon membrane as substrate strip. b Schematic diagram of a proposed model showing how polyubiquitination alters the catalytic pocket conformation of PTEN, which can be mimicked by insertion (increasing pocket depth) or deletion (decreasing pocket depth) mutations. c–f Quantitative measure of dephosphorylation of PIP3 (c), p-Tyr (d), p-Ser (e) or p-Thr (f) in PTEN phosphatase activity assays with unmodified or polyubiquitinated PTEN (WT or indicated mutants). Error bars, SD, n = 3 independent experiments (n.s. p > 0.05, *p < 0.05, and ***p < 0.001, Student’s t-test). g–j Competition binding assay to determine Kd for interaction between unmodified or polyubiquitinated PTEN (WT or indicated mutants) and biotin-PIP3 (g), biotin-BSA-p-Tyr (h), biotin-BSA-p-Ser (i), biotin-BSA-p-Thr (j) in the presence of unlabeled substrates respectively. Error bars, SD, n = 3 independent experiments. k Summary of Kd value (mean ± S.E.M, n = 3 independent experiments) of the interaction between indicated enzymes and substrates
To examine whether the distance between C124 and R130 determines PTEN’s substrate preference, we inserted one glycine between every two adjacent amino acids between C124 and R130 to generate insertion mutants Ins125G, Ins126G, Ins127G, and InsG129G (due to the position of Gly129, Ins128G and Ins129G have the same sequence) (Fig. 5b). Moreover, we generated deletion mutants of PTEN (ΔG127 and ΔG129), due to concerns that deletion of other amino acids between C124 and R130 may disrupt protein folding or the catalytic pocket of PTEN (Fig. 5b).
Bacterially-expressed PTEN WT, insertion or deletion mutants in vitro ubiquitinated (Supplementary Information, Fig. S7f), were subjected to PTEN phosphatase activity assay (substrates listed in Fig. 4b). Insertion of an additional glycine either before or after G129 of PTEN (Ins129G) abolished all of the proteins’ phosphatase activity, which is consistent with the notion that the natural variant G129E of PTEN has no enzymatic activity40 (Fig. 5c–f). Consistent with our previous findings, PTEN dephosphorylated PIP3 and p-Tyr; PTENK27-polyUb efficiently dephosphorylated p-Ser and p-Thr. However, both unmodified and ubiquitinated deletion mutants (ΔG127 and ΔG129) failed to exhibit noticeable phosphatase activity towards PIP3 and p-Tyr, but had potent phosphatase activity toward p-Ser and p-Thr. In contrast, all insertion mutants showed efficient phosphatase activity toward PIP3 and p-Tyr- but not p-Ser- or p-Thr-containing substrates regardless of the ubiquitination status (Fig. 5c–f). In summary, our data suggest that PTEN with one Gly deletion (shallow cleft) functions as a protein serine/threonine phosphatase, whereas PTEN with one Gly insertion (deep cleft) acts as a phosphoinositide/protein tyrosine phosphatase (Fig. 5b). Interestingly, none of the phosphatase activity of these PTEN mutants was regulated by K27-linked polyubiquitination, suggesting that K27-linked polyubiquitination on K66 and K80 of PTEN phosphatase domain fine-tuned the conformation of the catalytic loop to accommodate p-Ser/p-Thr side chains, converting PTENK27-polyUb to a protein serine/threonine phosphatase.
To further support this point, we determined the Kd values for the binding affinity of these PTEN mutants to biotinylated-PIP3 or biotinylated-BSA-conjugated p-Tyr, p-Ser or p-Thr residues (Fig. 5g–j). As summarized in Fig. 5k, unmodified PTEN exhibited high binding affinity to PIP3 and p-Tyr (Kd = 4.0 and 123.9 nM, respectively) and moderate binding with p-Ser and p-Thr (Kd = 720 and 559 nM, respectively). In contrast, PTENK27-polyUb exhibited 5–10 fold reduction in binding to PIP3/p-Tyr and about 8–14-fold enhancement in binding to p-Ser/p-Thr (Fig. 5k). Similarly to PTENK27-polyUb (red line), deletion mutants (ΔG127 and ΔG129, green and violet line, respectively) exhibited reduced binding to PIP3 and p-Tyr but enhanced binding to p-Ser and p-Thr residues (Fig. 5g–j). In contrast, insertion mutants of PTEN, Ins125G (orange line), Ins126G (black line), and Ins127G (brown line), exhibited enhanced binding to PIP3 and p-Tyr but impaired binding to p-Ser and p-Thr residues compared to unmodified PTEN (blue line) (Fig. 5g–j). Therefore, our data indicate that K27-linked ubiquitination of PTEN at K66 and K80 modulate substrate binding affinity of PTEN, hence switching PTEN from a phosphoinositide/tyrosine phosphatase to a serine/threonine phosphatase. The dimerization of PTEN and the PDZ domain-mediated binding of PTEN to the lipid membrane could also be modulated by the K27-linked polyUb. Technical difficulties in resolving the precise structure of PTENK27-polyUb prevents complete understanding of the mechanism behind ubiquitination-dependent alteration of PTEN’s enzymatic activity at this stage.
PTEN dephosphorylates master regulators of EMT in vivo
In MDA-MB-231 (with PTEN expression) and MDA-MB-468 (PTEN null) cells, high glucose induced the elevation of PIP3 levels, which was abolished when PTEN was genetically deleted, or MEX3C/GAEA/PTEN were knocked down by siRNA (Supplementary Information, Fig. S8a–c). The enzymatic activity of cellular PI3K in converting PIP2 to PIP3 was not significantly affected (Supplementary Information, Fig. S8d). Expression of PTEN WT, but not K66/80R mutant to MCF10A PTEN-/-, or MDA-MB-468 cells, rescued the high glucose-triggered elevated PIP3 levels (Supplementary Information, Fig. S8e–h). PTEN has been reported to dephosphorylate p-IRS1 upon IGF1 stimulation.41 Expression of PTEN WT, but not K66/80R mutant, rescued IGF1-induced IRS1 dephosphorylation under the condition of low or high glucose, without affecting the enzymatic activities of SHP2 and PP2A (Supplementary Information, Fig. S9a–f). These data suggest that PTENK27-polyUb exhibited reduced phosphoinositide/protein tyrosine phosphatase activity.
We explored the proteins candidates that potentially gain p-Ser and p-Thr upon PTEN-/- or expression of PTEN K66/80 R mutant (Supplementary Information, Fig. S10a, left panel), and identified the proteins that associated with PTENK27-polyUb but not with unmodified PTEN upon high glucose (Supplementary Information, Fig. S10a, right panel), finding that TWIST1, SNAI1 and YAP potentially being dephosphorylated by PTENK27-polyUb (Supplementary Information, Fig. S10b and Table S6).
To comprehensively demonstrate the protein phosphatase activity of PTEN with or without K27-polyUb, we determined phospho-proteomics profiling in MCF10A PTEN+/+, PTEN-/- cells expressing PTEN WT, Y138L42 or PTEN K66/80R (Supplementary Information, Fig. S10c). Hierarchical clustering analysis and principal component analysis (PCA) suggested that the three biological repeats were faithfully clustered together (Supplementary Information, Fig. S10d, e). We identified three categories of phospho-peptides representing PTEN protein phosphatase-independent substrates (potentially PIP3-dependent); and PTEN protein phosphatase-dependent substrates, which could be further defined as PTENK27-polyUb-independent and PTENK27-polyUb-dependent substrates (Fig. 6a). TWIST1 (Ser123) and SNAI1 (Ser96) exhibited phosphorylation in PTEN-/- cells expressing PTEN K66/80R mutant (Fig. 6a, top section), suggesting that TWIST1 and SNAI1 are endogenous substrates of PTENK27-polyUb as a protein serine/threonine phosphatase.
Characterization of Ser/Thr-phosphatase activity of polyubiquitinated PTEN in vivo. a Heat map and hierarchical clustering based on the intensities (log) of phospho-peptides of PTEN+/+, PTEN-/- MCF10A cells expressing blank vector, PTEN WT, Y138L or K66/80R mutants. Columns and rows are reordered by hierarchical clustering using the genotype. 3 independent biological repeats for each experimental condition were shown. b In vitro dephosphorylation assay were performed with 32P-labled substrates in the presence of unmodified or polyubiquitinated PTEN, and analyzed by IB or autoradiography. c Stability of TWIST1 and SNAI1 proteins after CHX treatment (100 µg/mL) for indicated time in glucose-starved MCF10A PTEN+/+ or MCF10A PTEN-/-cells followed by glucose re-stimulation (5 mM or 25 mM). d, e IB detection of indicated proteins in MCF10A PTEN+/+ or MCF10A PTEN-/-cells transfected with indicated vectors followed by glucose starvation with or without MG132 pre-treatment (5 µM, 6 h) and re-stimulation of glucose (5, 25 mM, 1 h). f IB detection using indicated antibodies in MCF10A PTEN+/+ and MCF10A PTEN-/- cells transfected with indicated expression vectors and siRNAs subjected to glucose starvation and followed by glucose re-stimulation (5 or 25 mM)
We validated the protein Ser/Thr phosphatase activity of PTENK27-polyUb toward p-TWIST1, p-SNAI1, phosphorylated by GSK3β43,44 and p-YAP1, phosphorylated by c-Abl45 or AKT,46 respectively (Fig. 6b). The Ser/Thr-phosphorylated TWIST1, SNAI1, and YAP1 were robustly dephosphorylated by PTENK27-polyUb but not unmodified PTEN; however, Tyr-phosphorylated YAP was dephosphorylated by PTP1B but not by PTENK27-polyUb (Fig. 6b).
It is interesting to find that 10 out of 16 PTENK27-polyUb-dependent substrates contain the consensus sequence (S/TxxxS/T) that is potentially subjected to phosphorylation-dependent degradation (phosphodegron) (Fig. 6a, right column). We further found that high glucose triggered the dephosphorylation and stabilization of TWIST1 and SNAI1 proteins in a PTEN-dependent manner, with the mRNA levels of TWIST, SNAI1 or YAP1 unaffected (Fig. 6c; Supplementary Information, Fig. S11a, b). Rescue experiments indicated that glucose-induced protein accumulation and dephosphorylation of TWIST1 and SNAI1 was abolished by PTEN-/-, which was rescued by expression of PTEN WT, but not PTEN K66/80R mutant (Fig. 6d, e).
TWIST1 and SNAI1 are key transcription factors regulating EMT.47 Upon high glucose, knockdown of TWIST1 or SNAI1 led to partially reduced vimentin-positive cells in PTEN-proficient cells or PTEN WT-expressing cells (Fig. 6f; Supplementary Information, Fig. S11c).
Glucose triggers YAP dephosphorylation and nuclear accumulation to activate YAP target genes.48 We observed diminished YAP phosphorylation at Ser127 and Ser397 upon high glucose stimulation, which was PTENK27-polyUb-dependent (Supplementary Information, Fig. S11d, e). Knockdown of MEX3C, or PTEN led to impairment of the dephosphorylation of YAP1, but not the phosphorylated LATS1 or the protein levels of PP2A-A, -B, and -C (Supplementary Information, Fig. S11f). Knockdown of GAEA, MEX3C, or PTEN also abolished activation of glucose-induced, YAP1-dependent target gene expression (Supplementary Information, Fig. S11g). These results indicate that polyubiquitination-dependent PTEN dual phosphatase activity switch correlates with the phosphorylation status of its cellular substrates.
Ubiquitination of PTEN is required for tissue remodeling and repair
We surveyed the status of GAEA and PTENK27-polyUb in normal human tissues, finding that GAEA and PTENK27-polyUb are detectable in the epidermal layer of skin, mammary gland ducts, intestinal villi, and kidney tubules, with low manifestation in cerebrum, lung, soft tissue and thyroid (Fig. 7a; Supplementary Information, Fig. S12a). We generated a genetically edited mouse model that has replaced lysine 80 with arginine in the FVB genetic background using CRISPR (clustered regularly-interspaced short palindromic repeats)/Cas (CRISPR associated) system-mediated genomic editing49 (Fig. 7b). The single-nucleotide mutant (AAA→AGA) was confirmed by Sanger sequencing and SNP (single nucleotide polymorphism) genotyping (Supplementary Information, Fig. S12b, c). Homozygous Pten K80R exhibited minimal effect on animal homeostasis (Supplementary Information, Fig. S12d). Male or female PtenK80R/K80R exhibits similar tumor susceptibility compared to PtenWT/WT or PtenWT/K80R mice (Supplementary Information, Fig. S12e). PTENK27-polyUb may play a functional role in type II EMT, which is involved in tissue remodeling and repair.50
K27-linked polyubiquitination of PTEN is required for mammary gland morphogenesis. a Immunohistochemistry (IHC) staining of Ub-PTEN (K80) using normal human tissues. Scale bar: 200 µm. b Schematic of the generation of PtenK80R knockin allele using CRISP-CAS9 method. c Representative whole mounts of inguinal mammary glands of 3 weeks or 9 weeks PtenWT/WT and PtenK80R/K80R virgin mice. Scale bar: 1 mm (left panel). Quantification of ductal extension as measured for the longest duct (right panel) (n = 5 animal per group). d Representative images of the second branch of mammary glands of 9-week old PtenWT/WT and PtenK80R/K80R virgin mice. Scale bar: 500 µm. Quantification of the number of secondary branch points in indicated mice (n = 5 animal per group). e Representative images of H&E staining of alveoli in the PtenWT/WT and PtenK80R/K80R luminal epithelial cells during different stages of pregnancy (P15.5 and L0.5) (left panel) and quantification of the percent alveoli per fat pad (right panel) (n = 5 animal per group). Scale bar: 100 µm. f, g Representative images of immunolabeling for Ub-PTEN (K80)+ (f) or Vimentin+ (g) cells in the PtenWT/WT and PtenK80R/K80R luminal epithelial cells during different stages of pregnancy (P15.5 and L0.5) (left panel) and quantification of the percent Ub-PTEN (K80)+ cells (f) or Vimentin+ cells (g) per visual field (×400) (right panel) (n = 5 animal per group). White arrows: Ub-PTEN (K80)+ cells (f) or Vimentin+ cells (g); scale bar: 50 µm. h Representative images of SNAI1/SLUG IHC staining in the PtenWT/WT and PtenK80R/K80R luminal epithelial cells during different stages of pregnancy (P15.5 and L0.5) (left panel) and quantification of the percent SNAI1/SLUG+ cells per visual field (×200) (right panel) (n = 5 animal per group). Black arrowheads-SNAI1/SLUG+ cells; scale bar: 100 µm. All error bars, SD, Student’s t-test (n.s. p > 0.05, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001)
In 9-week old virgin mice, PtenK80R/K80R mammary glands exhibited no changes in ductal length (Fig. 7c), but impairments in morphogenesis compared to wild-type littermates (Fig. 7d). On pregnancy day 15.5 and lactation day 0.5, mammary glands of PtenK80R/K80R mice exhibited reduced ductal morphogenesis compared to those of PtenWT/WT littermates (Fig. 7e). However, proliferation and apoptosis rates of PtenK80R/K80R mammary glands were comparable to those of wild-type littermates (Supplementary Information, Fig. S12f, g). PTENK27-polyUb was detected during pregnancy and lactation (Fig. 7f). The expression of vimentin and SNAI1/Slug in the mammary gland epithelium (E-cadherin-positive cells) was reduced in PtenK80R/K80R mice during pregnancy and lactation (Fig. 7g, h). These data suggest that PTENK27-polyUb may facilitate biological EMT in vivo.
We then utilized the cutaneous wound healing model51 to investigate the role of PTENK27-polyUb-dependent EMT in wound margins, finding that wound closure was significantly reduced in PtenK80R/K80R mice (Fig. 8a–c). PTENK27-polyUb in the neo-epidermis of 3-day wound tissues was significantly reduced in 6-day wound tissues and absent in PtenK80R/K80R skin (Fig. 8d, e). PtenK80R/K80R wounds exhibited reduced expression of vimentin, SNAI1/Slug and YAP1 in the neo-epidermis (Fig. 8f–k). The status of TWIST1 in neo-epidermis is undetectable under our experimental setting (Supplementary Information, Fig. S13a). The proliferation and apoptosis of skin epidermis were minimal affected in PtenK80R/K80R wound (Supplementary Information, Fig. S13b, c). Taken together, we demonstrated that PTENK27-polyUb is dynamically regulated upon physical or pathological conditions, with important roles in regulating EMT during tissue remodeling and repair.
K27-linked polyubiquitination of PTEN mediates EMT in vivo. a, b Representative macroscopic views of PtenWT/WT and PtenK80R/K80R mice skin wounds at days 1–6 after wounding (a). Scale bar: 1 mm. The morphometric analysis of the wound closure areas (b) (error bars, SEM. n = 5 animals per group, Student’s t-test). c Representative histological sections of post-wounded skin area of PtenWT/WT and PtenK80R/K80R mice at day 3; scale bar 200 μm, wm-wound margin; dotted line-basement membrane; dashed lines-delineated epidermal basal cells; ne- new epidermal cells. d, e Representative images of immunolabeling for Ub-PTEN (K80) in post-wounded skin area of PtenWT/WT and PtenK80R/K80R mice at day 3 and day 6 (d) and quantification of the percent ub-PTEN (K80)+ cells per visual field (×200) (e) (n = 5 animals per group). White arrows-ub-PTEN (K80)+ cells; scale bar: 100 µm. f, g Representative images of immunolabeling for E-cadherin and Vimentin in post-wounded skin area of PtenWT/WT and PtenK80R/K80R mice at day 6 (f) and quantification of the percent Vimentin+ cells per visual field (×200) (g) (n = 5 animals per group,). White arrows-Vimentin+ cells; scale bar: 100 µm. h, i Representative images of IHC staining of SNAI1 in post-wounded skin area of PtenWT/WT and PtenK80R/K80R mice at day 6 (h) and quantification of the percent SNAI1+ cells per field (i) (n = 5 animals per group). Black arrows-SNAI1+ cells; scale bar: 100 µm. j, k Representative images of immunolabeling for E-cadherin and YAP1 in post-wounded skin area of PtenWT/WT and PtenK80R/K80R mice at day 6 (j) and quantification of the percent YAP1+ cells per visual field (×200) (k) (n = 5 animals per group). White arrowheads-YAP1+ cells; scale bar: 100 µm. For (e, g, i, k), error bars, SD, Student’s t-test. (n.s. p > 0.05, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001)
Discussion
EMT has been demonstrated to be required for embryonic development, tissue remodeling/repair, and cancer metastasis.50 Many factors can induce the transformation of epithelial cells to mesenchymal cells.25 Our findings indicate that in addition to high glucose, CTGF, IL-6, SHH, and TGF-β1 effectively induce PTENK27-polyUb in human and mouse mammary gland epithelial cells. MEX3C is critical for catalyzing K27-linked polyubiquitination of PTEN at K66 and K80. PTENK27-polyUb potentially undergoes a conformational change, which might transform PTEN from a phosphoinositide/protein tyrosine phosphatase to a protein serine/threonine phosphatase. We identified potential p-Ser/p-Thr-containing protein substrates of PTENK27-polyUb, including TWIST1, SNAI1, and YAP1. Consequently, the PTENK27-polyUb-dependent dephosphorylation of TWIST1, SNAI1, and YAP1 resulted in accumulation of these proteins and promotion of EMT (Supplementary Information, Fig. S14).
Although PTEN has been considered one of the most important tumor suppressors, PTEN also plays important roles during homeostasis and pathological conditions,52 such as lactation,53 wound healing processes54 and organ fibrosis.55 To overcome the role of PTEN in inhibiting cell migration and proliferation, cells develop a variety of mechanisms. In addition to genetic depletion, non-sense mutations, and protein degradation,56 our data suggests that PTENK27-polyUb may exhibit reduced phosphoinositide phosphatase activity and adequate protein serine/threonine phosphatase activity. Upon stimulation by glucose or growth factors, the level of unmodified PTEN is reduced with concurrent upregulation of PTENK27-polyUb. Global phospho-proteomics profiling suggested that PTENK27-polyUb dephosphorylates a distinct cohort of protein substrates with phosphoserine/threonine residues. It is interesting that the majority of PTENK27-polyUb-dephosphorylated protein substrates harbor a consensus sequence that is potentially subjected to phosphorylation-dependent degradation (phosphodegron).57 PTENK27-polyUb-dependent dephosphorylation events may antagonize phosphorylation-dependent degradation, leading to protein stabilization. As a consequence, the potentially stabilized TWIST1,58 MYC,59 SNAI1,60 CASC5,61 PRLR,62 FGD1,63 CCNE1,64 and GLI365 may promote cell proliferation, EMT, invasion, and stemness, which are critical for homeostasis and pathological conditions. The substrates and corresponding cellular effects of PTENK27-polyUb could be diverse in a cellular/tissue context-specific manner. It is also possible that other post-translational modifications modulate the phosphatase activities of PTEN, which requires further examination.
Genetic depletion of Pten leads to hyper-activation of the PIP3/AKT pathway; on the contrary, PtenK80R/K80R results in unaffected phosphoinositide phosphatase activity and degradation of TWIST1/SNAI1/YAP1 proteins. Therefore, PtenK80R/K80R mice are non-tumor-prone. Although PTEN depletion generally leads to enhanced growth and invasion in cancer cells,66 PTEN loss leads to unaffected or reduced cell invasion in normal mammary epithelial cells.67 This notion is consistent with our observation that PTEN depletion leads to activation of the AKT pathway but degradation of TWIST1/SNAI1/YAP1. Several instances of cellular and genetic evidence indicated that AKT1, but not AKT2 or AKT3, serves as a negative regulator of EMT and breast cancer metastasis.68 AKT1 has been shown to phosphorylate TWIST1 at Ser42 to promote EMT69 or at Ser123 to facilitate β-TrCP-mediated TWIST1 degradation and inhibition of EMT.70 One explanation is that AKT1-dependent TWIST1 phosphorylation at Ser123 could be removed by PTENK27-polyUb under high glucose conditions, leading to activation of the AKT pathway and stabilization of the TWIST1 protein simultaneously.
K27-linked polyubiquitination might trigger a potential conformational change in the phosphatase domain of PTEN, which is a topic of further investigation. This effect depends on polyubiquitination at both K66 and K80, which is supported by our results indicating that mutation at either K66 or K80 failed to switch on this protein Ser/Thr phosphatase activity. We speculate that the two large polyubiquitin chains located on the phosphatase domain fine-tune the structure of the catalytic cleft, which alters substrate recognition by PTENK27-polyUb.
The two KH domains of MEX3C serve as potential RNA-binding domains. We demonstrated that upon GAEA association, MEX3C catalyzes K27-linked polyUb chain formation effectively. It is possible that the presence of GAEA manipulates the E2 that associates with MEX3C, leading to K27-linked polyUb chain formation. Consistent with this notion, MEX3C or GAEA mutants that failed to mediate this interaction abolished glucose-induced, PTENK27-polyUb-mediated EMT. It has been suggested that the UCAY RNA motif may be involved in the interactions with the KH domain. Hence, it is highly likely that other human or mouse lncRNAs containing similar RNA motifs may associate with and modulate the enzymatic activity of MEX3C.
Materials and methods
In vivo murine models
All animal-based research were approved by the institutional animal care and use committee (IACUC) of the University of Texas M.D. Anderson Cancer Center (MDACC). CRISPR/Cas9-mediated gene editing mice were generated by homologous recombination and implanted in FVB (Charles River Laboratories) blastocysts as previously described.39 Briefly, isogenic genomic fragments flanking PTEN K80 site containing the K80R mutation were cloned as EGE-LC-028 (K80R) expression vector. Two sgRNAs flanking targeting loci were cloned, in vitro transcribed and validated by Sanger sequencing and electrophoresis (Biocytogen). The targeting vector and sgRNAs were injected to blastocysts and the high-percentage male chimeric mice were crossed with female FVB mice to obtain transmitted founders. Genotyping of PtenK80R/K80R mice was determined by TaqMan® SNP genotyping technology using purified genomic DNA from mouse tails (Applied StemCell & Zymo Research). All mice were on a FVB genetic background. For mammary gland development experiments, 3-week- and 9-week-old female PtenWT/WT or PtenK80R/K80R littermates were used; for pregnancy and lactation experiment, 6-week-old PtenWT/WT and PtenK80R/K80R male mice were crossed to female mice with the same age and genotype. We checked the plugs the second day, and the pregnancy female mice were euthanized at P15.5 and L0.5 (P: pregnancy; L: lactation). The animal experiment was set up to use 5–12 mice per group to detect a 2-fold difference with power of 80% and at the significance level of 0.05 by a two-sided test for significant studies. The sample sized was chosen based on our previous experience and indicated for each experiment in the figure legend. Investigators were not blinded for group allocation, but were blinded for the assessment of the phenotypic outcome assessed by histological analyses. Animals were housed randomly and no animals were excluded from the analysis.
Whole-mount analysis and histological analysis
For whole-mount staining, the fourth 3-week- and 9-week-old female PtenWT/WT or PtenK80R/K80R mice mammary glands were excised and spread on microscope slides. The tissues were fixed in Carnoy’s fixative, washed in 70% ethanol, hydrated by passing through ethanol with decreasing concentrations, and stained in carmine alum stain overnight at room temperature. After staining, the slides were dehydrated through increasing ethanol concentrations, cleared in xylene, and mounted under a coverslip. For histological analysis, lactation mice mammary gland, wound healing mice skin and mouse major organ specimens were fixed overnight in neutral buffered formalin, dehydrated and embedded in paraffin. Tissue blocks were sectioned into 5 μm and stained with hematoxylin and eosin. For the analysis of the mammary gland development experiments and wound healing, five ×100 or ×200 visual fields were randomly selected for each carmine alum staining and H&E staining by a grid intersection analysis using Adobe Photoshop. Representative images were acquired with a Leica microscope.
Cutaneous wound healing and analysis
Full-thickness wounds were generated in the mouse dorsal skin under sterile conditions as previously described.51 Briefly, 6-week-old male or female mice were subjected to two excisional wounds with a sterile disposable biopsy punch (5 mm in diameter; ThermoFisher Scientific), after shaving and cleaning procedure. Each wound site was digitally photographed at the indicated time intervals, and wound areas were determined on photographs using Adobe PhotoShop (version 7.0; Adobe Systems). Wounds and their surrounding areas, including the scab and epithelial margins, were cut at the indicated time intervals for H & E, immunohistochemistry and immunofluorescence staining.
Phosphoproteomics and data analysis
Samples were prepared as previously described71 with minor modification. Briefly, MCF10A PTEN+/+ or PTEN-/- cells stably expressing blank vector, PTEN wild type, Y138L or K66/80R mutant were subjected to glucose starvation for 16 h followed by glucose stimulation (25 mM, 1 h). The cell lysates were freshly prepared using the ProteoPrep Zwitterionic Cell Lysis Kit, Mass Spec Grade (Protea) with Anti-RNase, Protease/Phosphatase Inhibitor Cocktail, and N-ethylmaleimide (NEM) (to preserve the native ubiquitination) supplemented in the lysis buffer. Immunoprecipitation using anti-phospho-Tyrosine, anti-phosphor-Serine/Threonine antibodies as a pool were performed using Dynabeads® Co-Immunoprecipitation Kit (Thermo Fisher Scientific) according to manufacturer’s instruction. Antibodies used in this study are listed in Supplementary Information, Table S8. The eluted protein complexes were denatured, reduced, alkylated and digested with immobilized trypsin (Promega). For TiO2 enrichment of phosphopeptides, packed tips were sequentially washed with glycolic acid buffer 2, 99/1 H2O/ACN and eluted with four sequential washes of 5% NH4OH in 1% ACN. Phosphopeptide pellets were resuspended in 20 μL of 0.1% TFA, and 4 μL was loaded into an LC–MS/MS system at MD Anderson Cancer Center Proteomics Facility.
Raw MS/MS data were processed using MaxQuant (version 1.5.8.3)72 with the Andromeda search engine73 against the human SwissProt database (20258 sequences, 02/28/2018) plus the standard MaxQuant contaminant database. Enzyme specificity was set to trypsin and up to two missed cleavages were allowed. Carbamidomethyl (C) was set as a fixed modification and oxidation (M), acetyl (protein N-term) and phospho (STY) as variable modifications. A false discovery rate cutoff of 1% was used at the PSM and protein levels. Reverse and contaminant matches were removed from the analysis. For phosphopeptide identification, an Andromeda minimum score and minimum delta score threshold of 40 and 17 were used, respectively, and only confidently localized phosphosites (localization probability ≥ 0.75) were used for downstream analysis. For quantification, only phosphopeptides consistently identified in all three replicates in at least one condition were analyzed. Heatmap was generated by metaX.74
Immunohistochemistry and immunofluorescence
For immunohistochemistry, formalin-fixed paraffin-embedded mammary glands and skins were deparaffinized. After blocking in 5% BSA/0.05% Tween-20, primary antibodies were applied. Antibodies used in this study are listed in Supplementary Information, Table S8. For all staining, counterstaining with hematoxylin was performed and DAB positivity was analyzed in five visual fields at ×200 magnification.
For immunofluorescence, cells or frozen tissues were fixed with 4% PFA at 4 °C for 10 min at room temperature. Both types of samples were blocked in 10% normal goat serum (NGS), PBS with 0.25% Triton X-100 in for 1 h at room temperature. For staining of 3D culture in Matrigel, 20 µg/mL goat anti-mouse F(ab´)2 fragment was included in primary block buffer to block the immunoreactive mouse IgG species in the Matrigel. Antibodies used in this study are listed in Supplementary Information, Table S8. Slides were mounted with ProLong Gold anti-fade reagent (Life Technologies) and imaged with confocal microscope (Zeiss). Five random fields were selected for each mouse and the number of E-cadherin-positive cells were scored for the presence or absence of vimentin, Ub-PTEN (K80), TWIST, or YAP1 co-staining. Quantification was performed using PerkinElmer InForm Analysis Software. Pseudocoloring and non-linear adjustment (γ changes) were applied in order for the printed images to accurately represent the quality of the high-resolution screens.
Cell culture and treatments
Human cancer cell lines MDA-MB-231, MDA-MB-468, A549, human embryonic kidney cell line HEK293T and Mus musculus mammary gland epithelial cell line NMuMG were purchased from American Type Culture Collection (ATCC) and Characterized Cell Line Core Facility (MD Anderson Cancer Center) and were maintained in Dulbecco’s modified eagle medium (DMEM) supplemented with 10% fetal bovine serum (FBS) at 37 °C in 5% CO2 (v/v). MCF10A and MCF10A PTEN-/- (Sigma-Aldrich) cells were maintained in DMEM/F12 medium supplemented with 5% horse serum, 200 ng/mL epidermal growth factor, 500 ng/mL hydrocortisone, 100 ng/mL cholera toxin and 10 µg/mL insulin at 37 °C in 5% CO2 (v/v). For 3D culture, the cells were mixed with 2% growth factor reduced (GFR) Matrigel (BD Biosciences) and then seeded in the eight-well chamber slide (Thermo Fisher Scientific) pre-coated with GFR Matrigel for culture at 37 °C in 5% CO2 (v/v) until 15 days. The cells have been authenticated and tested for mycoplasma contamination by vendors. For glucose treatment, cells were seeded in proper dishes. Twenty-four hours later, cells were washed and refreshed with glucose-free DMEM with 10% dialyzed FBS. Sixteen hours later, cells were treated with D-glucose (5 or 25 mM; Sigma-Aldrich) for indicated time in various experiments. For growth factor treatment, cells were serum starved for 24 h followed by treatment with Angiotensin II (100 ng/mL), BMP2 (100 ng/mL), CCL18 (100 ng/mL), CoCl2 (250 µM), Cripto-1 (100 ng/mL), hCTGF (10 ng/mL), mCTGF (10 ng/mL), EGF (10 ng/mL), FGF2 (10 ng/mL), HGF (20 ng/mL), IGF-1 (50 ng/mL), IL-1 (10 ng/mL), hIL-6 (10 ng/mL), mIL-6 (10 ng/mL), IL-8 (10 ng/mL), MMP2 (100 ng/mL), OSM (50 ng/mL), PDGF (10 ng/mL), PGE2 (10 ng/mL), SDII-1, hSHH (10 ng/mL), mSHH (10 ng/mL), hTGF-β (10 ng/mL), mTGF-β (10 ng/mL), TNF-α (10 ng/mL), Wnt3α (50 ng/mL) or CCL21 (100 nM) (Peprotech) for 1 h or 72 h as indicated.
Plasmids construction, transfection and lentiviral transduction
The full-length of MEX3C and PTEN mammalian expression vectors were obtained from OriGene. Ubiquitin and its corresponding mutants were constructed into pCMV-HA mammalian expression vector (Clontech). Bacterial expression vectors for His6-tagged MEX3C (wild-type and mutants) were constructed by subcloning the corresponding gene sequences into pET-28a vector. His6-tagged PTEN (wild-type and mutants) were constructed into the pET-DEST42 vectors using the Gateway system (Life Technologies). The point or domain deletion mutants were generated from the wild-type sequence using QuikChange™ Lightning Site-Directed Mutagenesis Kit (Agilent Technologies). Plasmids transfections were performed using Lipofectamine3000 (Life Technologies), or electroporation using 4D-Nucleofector™ System (Lonza) according to manufacturer’s instruction. Lentiviruses were produced in HEK293T cells with ViraPower™ Lentiviral Expression System (Thermo Scientific).
siRNA, recombinant proteins, antibodies and oligonucleotides
Non-targeting control siRNAs (D-001320), and ON-TARGETplus SMARTpool siRNA targeting MEX3C (L-006989), and PTEN (L-003023) from GE Healthcare Dharmacon were used in this study. The following recombinant proteins were used in this study. BostonBiochem: GST-UBA1, GST-UBE2D3, UBE2S, MDM2 and Ubiquitin (wild-type, His6-Ubiquitin-AA, G76A, K6R, K11R, K27R, K29R, K33R, K48R, K63R, K0, K6 only, K6 only, K11 only, K27 only, K29 only, K33 only, K48 only, K63 only, L73P); Lifesensors: USP2 and NEDD4; Novus Biologicals: GST-FMRP, GST-PTEN, GST-MEX3C, GST-Twist1, GST-SNAI1 and GST-YAP1; Cayman Chemical: PP2A; R&D Systems: PTP1B; OriGene: FLAG-PTEN. Millipore: His6-GSK3β, His6-AKT, and His6-ABL; Recombinant His6-MEX3C and His6-PTEN proteins were expressed in E. coli strain BL21-CodonPlus (DE3)-RIPL (Agilent Technologies) and purified using HisPur™ Cobalt Spin Columns (Life Technologies). The antibodies and oligonucleotides used in this study are listed in Supplementary Information, Tables S8, S9 respectively.
RNA isolation, lncRNAs microarray, qRT-PCR, immunoprecipitation and immunoblotting
Total RNA was isolated from cells using RNeasy Mini Kit (QIAGEN) following the manufacturer’s protocol. Human lncRNA microarray 3.0 analyses at ArrayStar Inc. were performed as previously described.75 Primer sequences for RT-QPCR are listed in Supplementary Information Table S9. Cells were homogenized in 1 × RIPA buffer (EMD Millipore) supplemented with Protease/Phosphatase Inhibitor Cocktail (Pierce, Thermo Scientific), Panobinostat (Selleck chemicals) and Methylstat (Sigma-Aldrich). Lysates were cleared by centrifugation at 13,000 rpm for 15 min at 4 °C. Supernatants were analyzed for immunoprecipitation with the indicated antibodies and the immunoprecipitated proteins were either subjected to immunoblotting or protein identification by mass spectrometry.
Denaturing immunoprecipitation, silver staining and mass spectrometry
For the denaturing immunoprecipitation, cell pellets were lysed using TSD buffer (50 mM Tris–HCl, pH 7.5, 1% SDS, 5 mM DTT), boil for 10 mins. The cleared lysates were used for immunoprecipitation as described above. The elution were loaded on NuPAGE 4–12% Bis–Tris gel (GenScript). Then the gel was stained using Pierce Silver Stain for Mass Spectrometry kit (Thermo Fisher Scientific) according to the manufacturer’s instructions. After silver staining, specific bands were excised, destained and digested with immobilized trypsin (Promega) for MS analysis at MD Anderson Cancer Center Proteomics Facility.
PIP3 mass ELISA and PI3K activity assay
Total cellular PIP3 was quantified using PI(3,4,5)P3 mass ELISA kits (Echelon Biosciences) following the manufacturer’s protocol. PI3K activity in cells with indicated treatments was measured using PI3-Kinase Activity ELISA: Pico (Echelon Biosciences).
E3LITE MEX3C ubiquitin ligase assay and in vitro ubiquitination assay
E3LITE Customizable Ubiquitin Ligase assay (LifeSensors) were performed according to the manufacture’s instruciton. In vitro ubiquitination assays were performed using an ubiquitin protein conjugation kit (BostonBiochem).
Purification of ubiquitinated PTEN and PTEN phosphatase activity assay
Recombinant His6-PTEN (WT or mutants) was incubated with ubiquitin, E1 and indicated E3-E2 pair to form K27-Poly- or K48-Poly-ubiquitinated PTEN, respectively. The polyubiquitinated PTEN was purified by His6-TUBE1 (LifeSensors) according to vendor. The phosphoinositide phosphatase activity of PTEN was detected using PTEN Activity ELISA kit (Echelon Biosciences). Phospho-Tyr, phospho-Ser/Thr phosphatase activity of unmodified or polyubiquitinated PTEN was assessed by incubating the indicated enzymes (200–0.006 nM) with indicated generic phospho-peptide substrates in in vitro phosphatase buffer (25 mM MOPS at pH 7.0 containing 50 mM NaCl, 1 mM DTT and 0.05% Tween® 20 and 50 mM Tris–HCl at pH 7.0 containing 0.1 mM CaCl2, 1 mM NiCl2, 125 µg/mL BSA and 0.05% Tween® 20, respectively) for 30 min at 37 °C. The phosphate released during enzymatic phosphatase assays was measured by Malachite Green Phosphate Detection Kit (Cell Signaling Technologies). A standard curve derived from the provided phosphate standard was used to convert A630nm readings obtained in an assay into nanomoles of phosphate. For the mass spectrometry analysis, 1 µg of PI(3,4,5)P3 diC8 (Echelon Biosciences) or synthesized phospho-Tyr, -Ser or -Thr-containing peptides were incubated with 500 ng wild-type or polyubiquitinated PTEN in the reaction buffer described above. Resulting peptides were purified by ZipTip and analyzed by MALDI-TOF mass spectrometry.
Recombinant GST-tagged TWIST1, SNAI1 or YAP proteins were incubated with 50 µL of in vitro kinase assay buffer II (SignalChem) containing 10 µCi [γ-32P]ATP and His6-tagged protein kinase (GSK3β for TWIST1 and SNAI1; AKT for YAP1 serine phosphorylation; c-Abl for YAP1 tyrosine phosphorylation) for 1 h at 30 °C. 32P-TWIST1, 32P-SNAI1, 32P-Ser-YAP1 or 32P-Tyr-YAP1 were purified by removing the kinases using HisPur™ Ni-NTA Magnetic Beads (ThermoFisher Scientific) and incubated with PTEN or PTENK27-polyUb as described above. The reactions were separated by SDS–PAGE and detected by autoradiography or immunoblotting with indicated antibodies.
Determination of Kd value using Alpha assay
Alpha binding assay using Streptavidin Donor beads and anti-His6 AlphaLISA® Acceptor Beads (PerkinElmer) was used to determine Kd for the PTEN: PIP3, or PTEN: phospho-amino acid interactions (PerkinElmer) according to manufacturer’s instruction. Saturation curve and competition curve were generated to determine Kd of the interaction between biotin-PIP3, biotin-BSA-pTyr, -pSer, -pThr or biotin-BSA as indicated and His6-PTEN in Alpha format, in which unlabeled PIP3 (10 µM to 0.01 nM) and BSA-pTyr, -pSer, or -pThr (400 µM to 0.04 nM) were used as competitor, respectively. The saturation binding curve and competitive inhibition curve were calculated based on Alpha signal readings by fitting to a nonlinear regression “saturation binding” model and a “log (inhibitor) vs. response-Variable slope (four parameters)” model, respectively. The Kd values were calculated using GraphPad Prism 6 software.
Measurement of RNA copy number and protein molecule number
The RNA copy number per cell were performed as previously described.76 Briefly, total RNA of MCF10A cells with or without high glucose (25 mM) stimulation were extracted and detected by qRT-PCR. Standard curve was generated by a serial dilution of in vitro transcribed GAEA RNA, using 717166.3 as molecular weight, and total RNA per cell was estimated to be 20 pg. The absolute molecule number of MEX3C and CD147 were determined using recombinant protein with a serial-dilution as standard curves and cell lysates from 10,000 cells using MEX3C ELISA kit (MyBioSource) and CD147 ELISA kit (R&D systems).
GAEA transcription factor prediction
Through searching the GAEA gene ID (LOC84856 or LINC00839) using the (https://genome.ucsc.edu/), we chose the 500 bp of GAEA promotor gene sequence to do the transcription prediction using two online tools.
Data analysis and statistics
The experiment was set up to use 3–5 samples/repeats per experiment/group/condition to detect a 2-fold difference with power of 80% and at the significance level of 0.05 by a two-sided test for significant studies. For immunoblotting, immunostaining, immunohistochemical staining, 3D culture, the representative images are shown. Each of these experiments was independently repeated for 3–5 times. Analyses of relative gene expression were determined using the 2−ΔΔCt method with GAPDH or B2M as the internal reference genes. Results are reported as mean ± standard error of the mean (S.E.M.) or standard deviation (SD) of at least three independent experiments. Each exact n value is indicated in the corresponding figure legend. Statistical analysis was performed using GraphPad Prism 7 software. Comparisons were performed using unpaired Student’s t-test or one-way ANOVA (*p < 0.05, **p < 0.01, ***p < 0.001 and ****p < 0.0001) as indicated in individual figures. A p-value < 0.05 was considered statistically significant. The investigators were not blinded to allocation during experiments and outcome assessment.
Accessions
The raw lncRNAs microarray data has been deposit to GEO (ID: GSE106729).
References
Song, M. S., Salmena, L. & Pandolfi, P. P. The functions and regulation of the PTEN tumour suppressor. Nat. Rev. Mol. Cell Biol. 13, 283–296 (2012).
Denu, J. M. & Dixon, J. E. A catalytic mechanism for the dual-specific phosphatases. Proc. Natl Acad. Sci. USA 92, 5910–5914 (1995).
Lee, J. O. et al. Crystal structure of the PTEN tumor suppressor: implications for its phosphoinositide phosphatase activity and membrane association. Cell 99, 323–334 (1999).
Maehama, T. & Dixon, J. E. The tumor suppressor, PTEN/MMAC1, dephosphorylates the lipid second messenger, phosphatidylinositol 3,4,5-trisphosphate. J. Biol. Chem. 273, 13375–13378 (1998).
Tamura, M. et al. Inhibition of cell migration, spreading, and focal adhesions by tumor suppressor PTEN. Science 280, 1614–1617 (1998).
Blanco-Aparicio, C., Renner, O., Leal, J. F. & Carnero, A. PTEN, more than the AKT pathway. Carcinogenesis 28, 1379–1386 (2007).
Robinson, V. L. Rethinking the central dogma: noncoding RNAs are biologically relevant. Urol. Oncol. 27, 304–306 (2009).
Guttman, M. et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458, 223–227 (2009). nature07672 [pii].
Hrdlickova, B., de Almeida, R. C., Borek, Z. & Withoff, S. Genetic variation in the non-coding genome: Involvement of micro-RNAs and long non-coding RNAs in disease. Biochim. Biophys. Acta 1842, 1910–1922 (2014).
St Laurent, G., Wahlestedt, C. & Kapranov, P. The Landscape of long noncoding RNA classification. Trends Genet. 31, 1910–1922 (2014).
Wilusz, J. E., Sunwoo, H. & Spector, D. L. Long noncoding RNAs: functional surprises from the RNA world. Genes Dev. 23, 1494–1504 (2009). 23/13/1494 [pii].
Johnsson, P., Lipovich, L., Grander, D. & Morris, K. V. Evolutionary conservation of long non-coding RNAs; sequence, structure, function. Biochim. Biophys. Acta 1840, 1063–1071, (2014).
Stone, R. C. et al. Epithelial-mesenchymal transition in tissue repair and fibrosis. Cell Tissue Res. 365, 495–506, (2016).
Kalluri, R. & Neilson, E. G. Epithelial-mesenchymal transition and its implications for fibrosis. J. Clin. Invest. 112, 1776–1784, (2003).
Kalluri, R. & Weinberg, R. A. The basics of epithelial-mesenchymal transition. J. Clin. Invest. 119, 1420–1428, (2009).
Fischer, K. R. et al. Epithelial-to-mesenchymal transition is not required for lung metastasis but contributes to chemoresistance. Nature 527, 472–476, (2015).
Okubo, T. et al. Down-regulation of promoter 1.3 activity of the human aromatase gene in breast tissue by zinc-finger protein, snail (SnaH). Cancer Res. 61, 1338–1346 (2001).
Masur, K. et al. Diabetogenic glucose and insulin concentrations modulate transcriptome and protein levels involved in tumour cell migration, adhesion and proliferation. Br. J. Cancer 104, 345–352, (2011).
Saville, M. K. et al. Regulation of p53 by the ubiquitin-conjugating enzymes UbcH5B/C in vivo. J. Biol. Chem. 279, 42169–42181, (2004).
Katoh, S., Tsunoda, Y., Murata, K., Minami, E. & Katoh, E. Active site residues and amino acid specificity of the ubiquitin carrier protein-binding RING-H2 finger domain. J. Biol. Chem. 280, 41015–41024, (2005).
Polge, C. et al. UBE2B is implicated in myofibrillar protein loss in catabolic C2C12 myotubes. J. Cachexia Sarcopenia Muscle 7, 377–387, (2016).
Zhang, L., Xu, M., Scotti, E., Chen, Z. J. & Tontonoz, P. Both K63 and K48 ubiquitin linkages signal lysosomal degradation of the LDL receptor. J. Lipid Res. 54, 1410–1420, (2013).
Wu, T. et al. UBE2S drives elongation of K11-linked ubiquitin chains by the anaphase-promoting complex. Proc. Natl Acad. Sci. USA 107, 1355–1360, (2010).
Dandona, P., Chaudhuri, A., Ghanim, H. & Mohanty, P. Proinflammatory effects of glucose and anti-inflammatory effect of insulin: relevance to cardiovascular disease. Am. J. Cardiol. 99, 15B–26B, (2007).
Lamouille, S., Xu, J. & Derynck, R. Molecular mechanisms of epithelial-mesenchymal transition. Nat. Rev. Mol. Cell Biol. 15, 178–196, (2014).
Sonnylal, S. et al. Connective tissue growth factor causes EMT-like cell fate changes in vivo and in vitro. J. Cell. Sci. 126, 2164–2175, (2013).
Yoo, Y. A. et al. Sonic hedgehog pathway promotes metastasis and lymphangiogenesis via activation of Akt, EMT, and MMP-9 pathway in gastric cancer. Cancer Res. 71, 7061–7070, (2011).
Sullivan, N. J. et al. Interleukin-6 induces an epithelial-mesenchymal transition phenotype in human breast cancer cells. Oncogene 28, 2940–2947 (2009).
Chan, D. A. et al. Targeting GLUT1 and the Warburg effect in renal cell carcinoma by chemical synthetic lethality. Sci. Transl. Med. 3, 94ra70, (2011).
Biswas, S. et al. Endothelin-1 regulation is entangled in a complex web of epigenetic mechanisms in diabetes. Physiol. Res. 67, S115–S125 (2018).
Zhang, Z. Y. et al. Substrate specificity of the protein tyrosine phosphatases. Proc. Natl Acad. Sci. USA 90, 4446–4450 (1993).
Donella Deana, A. et al. An investigation of the substrate specificity of protein phosphatase 2C using synthetic peptide substrates; comparison with protein phosphatase 2A. Biochim. Biophys. Acta 1051, 199–202 (1990).
Pinna, L. A. & Donella-Deana, A. Phosphorylated synthetic peptides as tools for studying protein phosphatases. Biochim. Biophys. Acta 1222, 415–431 (1994).
Brown-Shimer, S. et al. Molecular cloning and chromosome mapping of the human gene encoding protein phosphotyrosyl phosphatase 1B. Proc. Natl Acad. Sci. USA 87, 5148–5152 (1990).
Stone, S. R. et al. The nucleotide sequence of the cDNA encoding the human lung protein phosphatase 2A alpha catalytic subunit. Nucleic Acids Res. 16, 11365 (1988).
Stevenson, L. F. et al. The deubiquitinating enzyme USP2a regulates the p53 pathway by targeting Mdm2. EMBO J. 26, 976–986, (2007).
Camps, M., Nichols, A. & Arkinstall, S. Dual specificity phosphatases: a gene family for control of MAP kinase function. FASEB J. 14, 6–16 (2000).
Leslie, N. R. & Downes, C. P. PTEN function: how normal cells control it and tumour cells lose it. Biochem. J. 382, 1–11 (2004).
Xu, Y. et al. Structure of the protein phosphatase 2A holoenzyme. Cell 127, 1239–1251 (2006).
Myers, M. P. et al. The lipid phosphatase activity of PTEN is critical for its tumor supressor function. Proc. Natl Acad. Sci. USA 95, 13513–13518 (1998).
Shi, Y. et al. PTEN is a protein tyrosine phosphatase for IRS1. Nat. Struct. Mol. Biol. 21, 522–527, (2014).
Davidson, L. et al. Suppression of cellular proliferation and invasion by the concerted lipid and protein phosphatase activities of PTEN. Oncogene 29, 687–697, (2010).
Lander, R. et al. Interactions between Twist and other core epithelial-mesenchymal transition factors are controlled by GSK3-mediated phosphorylation. Nat. Commun. 4, 1542, (2013).
Zhou, B. P. et al. Dual regulation of snail by GSK-3beta-mediated phosphorylation in control of epithelial-mesenchymal transition. Nat. Cell Biol. 6, 931–940, (2004).
Levy, D., Adamovich, Y., Reuven, N. & Shaul, Y. Yap1 phosphorylation by c-Abl is a critical step in selective activation of proapoptotic genes in response to DNA damage. Mol. Cell 29, 350–361, (2008).
Basu, S., Totty, N. F., Irwin, M. S. & Sudol, M. & Downward, J. Akt phosphorylates the Yes-associated protein, YAP, to induce interaction with 14-3-3 and attenuation of p73-mediated apoptosis. Mol. Cell 11, 11–23 (2003).
Yang, J. & Weinberg, R. A. Epithelial-mesenchymal transition: at the crossroads of development and tumor metastasis. Dev. Cell 14, 818–829 (2008).
Wang, W. et al. AMPK modulates Hippo pathway activity to regulate energy homeostasis. Nat. Cell Biol. 17, 490–499, (2015).
Yang, H. et al. One-step generation of mice carrying reporter and conditional alleles by CRISPR/Cas-mediated genome engineering. Cell 154, 1370–1379 (2013).
Kalluri, R. EMT: when epithelial cells decide to become mesenchymal-like cells. J. Clin. Invest. 119, 1417–1419, (2009).
Ganguli-Indra, G. Protocol for cutaneous wound healing assay in a murine model. Methods Mol. Biol. 1210, 151–159, (2014).
Nguyen, K. T. et al. Essential role of Pten in body size determination and pancreatic beta-cell homeostasis in vivo. Mol. Cell Biol. 26, 4511–4518, (2006).
Wang, Z. et al. Pten regulates development and lactation in the mammary glands of dairy cows. PLoS ONE 9, e102118, (2014).
Zhao, M. PTEN: a promising pharmacological target to enhance epithelial wound healing. Br. J. Pharmacol. 152, 1141–1144, (2007).
Miyoshi, K. et al. Epithelial Pten controls acute lung injury and fibrosis by regulating alveolar epithelial cell integrity. Am. J. Respir. Crit. Care Med. 187, 262–275, (2013).
Salmena, L., Carracedo, A. & Pandolfi, P. P. Tenets of PTEN tumor suppression. Cell 133, 403–414 (2008).
Holt, L. J. Regulatory modules: Coupling protein stability to phopshoregulation during cell division. FEBS Lett. 586, 2773–2777 (2012).
Yang, J. et al. Twist, a master regulator of morphogenesis, plays an essential role in tumor metastasis. Cell 117, 927–939 (2004).
Dang, C. V. MYC on the path to cancer. Cell 149, 22–35 (2012).
Wang, Y., Shi, J., Chai, K., Ying, X. & Zhou, B. P. The Role of Snail in EMT and Tumorigenesis. Curr. Cancer Drug Targets 13, 963–972 (2013).
Urata, Y. N., Takeshita, F., Tanaka, H., Ochiya, T. & Takimoto, M. Targeted knockdown of the kinetochore protein D40/Knl-1 inhibits human cancer in a p53 status-independent manner. Sci. Rep. 5, 13676 (2015).
Clevenger, C. V. Role of prolactin/prolactin receptor signaling in human breast cancer. Breast Dis. 18, 75–86 (2003).
Ayala, I. et al. Faciogenital dysplasia protein Fgd1 regulates invadopodia biogenesis and extracellular matrix degradation and is up-regulated in prostate and breast cancer. Cancer Res. 69, 747–752, (2009).
Keyomarsi, K., Tucker, S. L. & Bedrosian, I. Cyclin E is a more powerful predictor of breast cancer outcome than proliferation. Nat. Med. 9, 152, (2003).
Matise, M. P. & Joyner, A. L. Gli genes in development and cancer. Oncogene 18, 7852–7859, (1999).
Tamura, M., Gu, J., Takino, T. & Yamada, K. M. Tumor suppressor PTEN inhibition of cell invasion, migration, and growth: differential involvement of focal adhesion kinase and p130Cas. Cancer Res. 59, 442–449 (1999).
Ghosh, S., Varela, L., Sood, A., Park, B. H. & Lotan, T. L. mTOR signaling feedback modulates mammary epithelial differentiation and restrains invasion downstream of PTEN loss. Cancer Res. 73, 5218–5231, (2013).
Hutchinson, J. N., Jin, J., Cardiff, R. D., Woodgett, J. R. & Muller, W. J. Activation of Akt-1 (PKB-alpha) can accelerate ErbB-2-mediated mammary tumorigenesis but suppresses tumor invasion. Cancer Res. 64, 3171–3178 (2004).
Xue, G. et al. Akt/PKB-mediated phosphorylation of Twist1 promotes tumor metastasis via mediating cross-talk between PI3K/Akt and TGF-beta signaling axes. Cancer Discov. 2, 248–259, (2012).
Li, C. W. et al. AKT1 inhibits epithelial-to-mesenchymal transition in breast cancer through phosphorylation-dependent Twist1 degradation. Cancer Res. 76, 1451–1462, (2016).
Wilkes, E. H., Terfve, C., Gribben, J. G., Saez-Rodriguez, J. & Cutillas, P. R. Empirical inference of circuitry and plasticity in a kinase signaling network. Proc. Natl Acad. Sci. USA 112, 7719–7724, (2015).
Tyanova, S., Temu, T. & Cox, J. The MaxQuant computational platform for mass spectrometry-based shotgun proteomics. Nat. Protoc. 11, 2301–2319 (2016).
Cox, J. et al. Andromeda: a peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 10, 1794–1805, (2011).
Wen, B., Mei, Z., Zeng, C. & Liu, S. metaX: a flexible and comprehensive software for processing metabolomics data. BMC Bioinform. 18, 183, (2017).
Xing, Z. et al. lncRNA directs cooperative epigenetic regulation downstream of chemokine signals. Cell 159, 1110–1125 (2014).
Fey, A. et al. Establishment of a real-time PCR-based approach for accurate quantification of bacterial RNA targets in water, using Salmonella as a model organism. Appl. Environ. Microbiol. 70, 3618–3623 (2004).
Acknowledgements
We thank Mr. D. Aten for assistance with figure presentation and Peter K. Park and Sergey D. Egranov for manuscript preparation. This research work is partially supported by National Cancer Institute (NCI) CPTAC award [U24 CA210954], Cancer Prevention and Research Institutes of Texas [CPRIT RR160027], McNair Medical Institute at The Robert and Janice McNair Foundation to B.Z. This work was supported in part by Cancer Prevention Research Institute of Texas (CPRIT) grant number RP130397 and NIH grant number 1S10OD012304-01 to D.H.H. This work was supported by National Institutes of Health Pathway to Independence Award (R00CA166527), National Cancer Institute R01 award (1 R01 CA218036-01), Cancer Prevention Research Institute of Texas First-time Faculty Recruitment Award (R1218) grants, Department of Defense Breakthrough award (BC151465), Andrew Sabin Family Foundation Fellows award, and AACR-Bayer Innovation and Discovery Grants to L.Q.Y. and National Institutes of Health Pathway to Independence Award (R00DK094981), National Cancer Institute R01 award (1R01CA218025-01, 1R01CA231011-01), Department of Defense Breakthrough award BC180196, and Cancer Prevention Research Institute of Texas Individual Investigator Research Award (150094 and 180259) to C.R.L.
Author information
Authors and Affiliations
Contributions
L.Q.Y. and C.R.L. conceived of the project and designed the experiments. Q.S.H. and S.Y.W. executed the primary studies with assistance of Y.Y.Z., Q.S.H., C.L.L. and Y.J.L. performed the experiments with genetic animal models. D.H. executed mass spectrometry analysis. B.W. and B.Z., with assistance of J.Y. performed the phospho-proteomics data analysis. Y.Q.Y. and L.H. ensured the statistical analysis. Clinical specimens were ascertained and processed by J.R.M. and L.Q.Y. The histological staining and corresponding analysis were performed by K.L., C.L.L. and Q.S.H. H.H. and T.K.N. assisted with genetic mouse model breeding. P.K.P. and S.D.E. assisted with manuscript drafting. M.C.H. contributed to experimental design and data interpretation. L.Q.Y. and C.R.L. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Hu, Q., Li, C., Wang, S. et al. LncRNAs-directed PTEN enzymatic switch governs epithelial–mesenchymal transition. Cell Res 29, 286–304 (2019). https://doi.org/10.1038/s41422-018-0134-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41422-018-0134-3
This article is cited by
-
TC2N inhibits distant metastasis and stemness of breast cancer via blocking fatty acid synthesis
Journal of Translational Medicine (2024)
-
GPR162 activates STING dependent DNA damage pathway as a novel tumor suppressor and radiation sensitizer
Signal Transduction and Targeted Therapy (2023)
-
The cancer-testis lncRNA lnc-CTHCC promotes hepatocellular carcinogenesis by binding hnRNP K and activating YAP1 transcription
Nature Cancer (2022)
-
The deadly cross-talk between Hippo pathway and epithelial–mesenchymal transition (EMT) in cancer
Molecular Biology Reports (2022)
-
UBE2S promotes cell chemoresistance through PTEN-AKT signaling in hepatocellular carcinoma
Cell Death Discovery (2021)