Abstract
The hematopoietic stem cell (HSC) niche has been extensively studied in bone marrow, yet a more systematic investigation into the microenvironment regulation of hematopoiesis in fetal liver is necessary. Here we investigate the spatial organization and transcriptional profile of individual cells in both wild type (WT) and Tet2−/− fetal livers, by multiplexed error robust fluorescence in situ hybridization. We find that specific pairs of fetal liver cell types are preferentially positioned next to each other. Ligand-receptor signaling molecule pairs such as Kitl and Kit are enriched in neighboring cell types. The majority of HSCs are in direct contact with endothelial cells (ECs) in both WT and Tet2−/− fetal livers. Loss of Tet2 increases the number of HSCs, and upregulates Wnt and Notch signaling genes in the HSC niche. Two subtypes of ECs, arterial ECs and sinusoidal ECs, and other cell types contribute distinct signaling molecules to the HSC niche. Collectively, this study provides a comprehensive picture and bioinformatic foundation for HSC spatial regulation in fetal liver.
Similar content being viewed by others
Introduction
Hematopoietic stem cells (HSCs) retain the potential to generate all lineages of blood cells throughout the life of the organism. Long-term HSCs arise first in the dorsal aorta of aorta–gonad–mesonephros region of mouse embryos around embryonic day 10.5 (E10.5) through an endothelial-to-hematopoietic transition1. On day 12 of gestation, HSCs migrate to the fetal liver through the blood circulation via the umbilical vein from the placenta. In the fetal liver, HSCs undergo a 38-fold expansion until E162. Finally, HSCs migrate from the liver to the bone marrow where they maintain life-long hematopoiesis.
The microenvironment of HSCs, termed the “HSC niche”, influences fundamental properties of the HSCs, including proliferation, self-renewal, differentiation, and migration3. Various niches are present in different stages of development given the diversity of tissues where HSCs reside, including the yolk sac, aorta–gonad–mesonephros region, placenta, fetal liver, spleen, and bone marrow4,5,6. Distinct niches may have diverse responsibilities to meet the demand of hematopoietic cells during homeostasis and stress7. For example, fetal HSCs are more efficient in regenerating certain cell populations such as CD5+ B cells than adult HSCs8. Another difference is that HSCs are mostly quiescent in bone marrow9, whereas they actively proliferate in fetal liver2. However, in both settings the HSC niche tends to include endothelial cells (ECs), either as part of portal vessels in the fetal liver or arterioles in the bone marrow10,11. How prevalent HSCs are in direct contact with ECs is unknown. The study of HSC niches will further our understanding and treatment of various hematopoietic diseases. Because it is difficult to attain sufficient HSCs for direct clinical use, understanding a niche that supports rapid HSC expansion may offer guidance to amplify HSCs ex vivo.
Several microenvironment-dependent signaling pathways have been shown as essential in regulating the self-renewal and differentiation of HSCs in the bone marrow. These include: CXCL12/CXCR4 signaling, BMP signaling, Mpl/TPO signaling, Tie2/Ang-1 signaling, Wnt signaling, and Notch signaling12. Wnt and Notch are particularly complicated signaling pathways in the niche, because a large number of gene family members are involved in each pathway, and because different levels of pathway activation yield different overall effects on the HSCs and hematopoiesis13,14. How these signaling pathways contribute to fetal liver HSCs needs further investigation.
In addition to signaling pathways in the microenvironment, epigenetic elements also regulate HSC development, heterogeneity and proliferation. In particular, Tet2 encodes a well-known epigenetic modifier that plays important roles in hematopoiesis. TET2 regulates DNA methylation by conversion of 5-methylcytosine (5mC) to 5-hydroxymethylcytosine (5hmC)15,16. Disruption of Tet2 enhances HSC self-renewal, and leads to myeloid transformation17,18. In addition to cell-autonomous functions within the HSC, loss of Tet2 increases bone marrow stromal cell (BMSC) self-renewal and proliferation capacity, and enhances their hematopoietic supportive capacity19. Clinically, loss-of-function mutations of Tet2 are found in clonal hematopoiesis of indeterminate potential20, 26% of myelodysplasia21, 12% of acute myeloid leukemia, and 42% of chronic myelomonocytic leukemia22. Yet the effect of Tet2 in the fetal liver niche context is only partially understood.
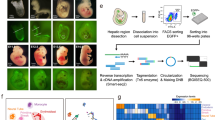
To better understand the function of the HSC niche, a systematic characterization of cell types and active molecular pathways within the HSC microenvironment is required. Single-cell RNA-sequencing (scRNA-seq) allows profiling the heterogeneity of gene expression in individual cells23. However, standard scRNA-seq requires cell dissociation, which destroys the spatial context and cell–cell interactions in the niche. Recent spatial scRNA-seq modalities detect spatial distribution of cell types, but are limited by the spatial resolution (some only partially matching the spatial scale of single cells), detection efficiency, and/or drop-out rate24,25. Single molecule fluorescence in situ hybridization (smFISH) is a powerful tool to map and count RNAs in situ with high spatial resolution and detection efficiency, but traditional smFISH can only measure a few RNA species at a time26. Multiplexed error-robust fluorescence in situ hybridization (MERFISH) is a highly multiplexed FISH method to map hundreds to thousands of RNA species at the single molecule level in single cells27,28,29. MERFISH is based on a combinatorial barcoding and error-robust encoding scheme to label numerous RNA species, and a sequential smFISH strategy to read out the barcodes, each of which is associated with different RNA species27,28,29. Here, we use MERFISH and scRNA-seq to explore hematopoiesis in mouse fetal liver and how it is altered in the absence of TET2. The experimental workflow is illustrated in Supplementary Fig. S1. Using MERFISH, we profile numerous niche factors, signaling factors, and cell type marker transcripts simultaneously at the single cell level in situ. We analyze how cells of different types are spatially organized into microenvironments, and how the cell type and the microenvironment organization jointly affect transcript profiles. Our results show that specific pairs of cell types are enriched as neighbors in the fetal liver microenvironments. Particularly, HSCs locate predominantly in endothelial niches and the majority of HSCs are in direct contact with one or several types of ECs. We further compare the HSC niches in wild type (WT) and Tet2−/− fetal livers and show that loss of Tet2 leads to an increased number of HSCs, as well as increased expression of Wnt and Notch signaling genes within the HSC niche, suggesting a causal link between the molecular and cellular changes and related pathology. In addition, we show that arterial ECs (AECs), sinusoidal ECs (SECs), and other fetal liver cell types offer distinct signaling molecules to the HSC niche.
Results
scRNA-seq and MERFISH identify major cell types in fetal liver
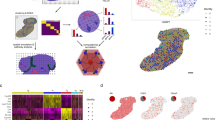
To characterize the gene expression features of different cell types in E14.5 mouse fetal liver, we first identified 11 major cell types in fetal liver with scRNA-seq (Fig. 1a). Cell type assignment was based on the marker genes selected from the Mouse Cell Atlas database30. We performed RNA velocity analysis to derive the developmental relationship among the different cell types (Fig. 1b). RNA velocity was analyzed by calculating the ratio of spliced to unspliced mRNAs in single cells, which predicts the future state of each cell31. According to a two-dimensional representation of the single cell transcriptome and the direction of the arrows in our RNA velocity map, erythroid cells, erythroid progenitors, myeloids, basophils, and neutrophils were connected in a branched development trajectory, while megakaryocytes (MKs) and macrophages were separated from the above mentioned cells; erythroid progenitors differentiated into erythroid cells; and neutrophils were separated into two main subtypes (Neutrophil_Elane and Neutrophil_Ngp) that appear to be differentiated from a common progenitor (Fig. 1b).
a UMAP visualization of cell clustering of 11 major cell types in the WT fetal liver by scRNA-seq (n = 7,635 cells). MK, megakaryocyte; EC, endothelial cell; Pre-B, Pre-B cell. b The observed and the extrapolated future states (arrows) are shown on the pre-defined UMAP. RNA velocity was estimated with spliced and unspliced RNA ratios in single cells. c UMAP plot of MERFISH data of WT fetal liver (n = 40,864 cells). We attained eight major cell types with MERFISH data with an extra small group of “Unknown” cells that we cannot distinguish. d Violin plot of example cell type markers in MERFISH data. x axis shows normalized abundance ranging from 0 to 0.3 for each gene. See Methods for the normalization procedure. e The correlation analysis of total RNA copy numbers from MERFISH imaging vs FPKM values from bulk RNA-seq for each probed RNA species. f The correlation analysis of total RNA copy numbers of two biological replicates of MERFISH for each probed RNA species. g The in situ map of identified cell types in a field of view (left panel) and the detected transcripts of example marker genes plotted in the same field of view (right panel).
We further analyzed the gene ontology (GO) (biological processes) of differentially expressed genes (DEGs) in myeloid cells, MKs, erythroid progenitors, erythroid cells, HSCs, and ECs, in which each cell type was compared to the other cell types (Supplementary Fig. S2). GO analysis of DEGs showed that myeloid cells, as expected, mainly committed themselves to defense response and immune response (Supplementary Fig. S2a); DEGs of MKs showed wound healing, hemostasis, and platelet activation functions (Supplementary Fig. S2b); DEGs of erythroid progenitors were enriched for heme metabolic and biosynthetic processes (Supplementary Fig. S2c); erythroid cells expressed genes related to erythrocyte development and homeostasis (Supplementary Fig. S2d); HSCs expressed genes that regulate the immune system, hematopoietic or lymphoid organ development and hemopoiesis (Supplementary Fig. S2e); and DEGs of ECs were enriched with circulatory system development and vasculature/blood vessel development functions (Supplementary Fig. S2f).
To analyze the distinct fetal liver cell types and their gene expression profiles in real space, we designed a MERFISH probe library that targets 45 cell-type marker genes selected from the scRNA-seq data and 95 niche-associated factors or signaling/marker genes from literature (Supplementary Table S1). We segmented cells with oligo-conjugated wheat germ agglutinin labeling of cell membrane components (Supplementary Fig. S3). The raw images of MERFISH showed distinct RNA foci (Supplementary Fig. S4). The single-cell RNA copy numbers of 132 selected genes (after excluding eight low quality RNA species; see Materials and Methods) allowed us to identify eight major cell types using the fetal liver MERFISH data (Fig. 1c and Supplementary Fig. S5). Note that MERFISH captures a more accurate representation of the proportion of erythroid cells compared to scRNA-seq (Supplementary Fig. S6), which commonly uses red blood cell lysis buffer. The cell types were identified based on the marker gene expression levels (Fig. 1d). Population-level RNA copy numbers measured by MERFISH were highly correlated with FPKM values from bulk RNA sequencing, with a correlation coefficient of 0.79 (Fig. 1e). Biological replicates of MERFISH measurements showed a correlation coefficient of 0.97 (Fig. 1f). These high correlations validated the quality of the MERFISH data. The identified eight major cell types were mapped in situ back onto the tissue sections, revealing their spatial organization (Fig. 1g).
Spatial proximity of pairs of cell types in WT and Tet2 − /− fetal liver
To better understand the spatial organization of cells in fetal liver, we analyzed the enrichment of pairs of cell types in spatial proximity in WT and Tet2−/− fetal livers (Fig. 2). Here, we defined two cells that are less than 20-μm apart as in a neighborhood, and calculated the probability of cell type pairs in neighborhoods. We then calculated the enrichment of cell type pairs in spatial proximity by normalizing this probability to the control probability based on random pairing (see Materials and Methods). The most prominent feature from this spatial proximity analysis was that cells tended to aggregate with cells of the same type in situ (Fig. 2a, b). Due to the highly enriched spatial self-clustering, neighboring pairs of different cell types were mostly depleted when self pairs are calculated together with non-self pairs. To further capture enrichment just between non-self pairs, we removed the self pairs from the analysis (Fig. 2c–f). This allowed us to capture several enriched pairs of different cell types in spatial proximity. In both WT (Fig. 2c, e) and Tet2−/− (Fig. 2d, f) fetal livers, macrophages were paired with erythroid cells; hepatocytes and myeloid cells were paired with erythroid progenitors; hepatocytes were also paired with erythroid cells. In addition, erythroid progenitors were paired with AECs in WT fetal liver (Fig. 2c, e and Supplementary Fig. S7a); macrophages were paired with SECs, and hepatocytes with myeloid cells in Tet2−/− fetal liver (Fig. 2d, f and Supplementary Fig. S7b). In summary, erythroid progenitors lost a neighboring relationship with AECs, myeloids gained a neighboring relationship with hepatocytes, and macrophages gained a neighboring relationship with SECs in Tet2−/− fetal liver (Supplementary Fig. S8). Previous studies have shown that macrophages tended to be surrounded by erythroid cells forming an erythroblastic island and to regulate the differentiation, proliferation, and clearance of erythroid cells32, which is consistent with our observation (Fig. 2g, h and Supplementary Fig. S7c). These results indicate that fetal liver cell types are spatially paired in a non-random fashion, suggesting cell-cell communications.
a, b The enrichment folds of pairs of cell types in spatial proximity with self-pairs in WT (a) and Tet2−/− (b) fetal livers. c, d The enrichment folds of pairs of cell types in spatial proximity without self-pairs in WT (c) and Tet2−/− (d) fetal livers. The color bars are scaled to ±1 (a–d) with red indicating enrichment and blue indicating depletion. Cyan dots represent P values < 0.01 for positive enrichment with false discovery rate correction. e, f The chord diagram plots of significantly enriched pairs in WT (e) and Tet2−/− (f) fetal livers. g, h MERFISH cell type plots of a central macrophage (green) in an erythroid island (red). Cell types were determined using MERFISH data and mapped in situ (left panels). A fourfold magnified image of the region in the white box is shown on the right. AEC, arterial endothelial cell; SEC, sinusoidal endothelial cell; MK, megakaryocyte.
Expression of known niche factors in fetal liver cell types
To explore the expression of various HSC niche factors in different cell types, we calculated the enrichment of gene expression in different cell types over the population-averaged expression profile in both WT (Fig. 3a) and Tet2−/− (Fig. 3b) populations. As expected, cell type markers were enriched in the corresponding cell types. For example, megakaryocyte markers Gnaz, Mrvi1, Vwf, Timp3, and Mmrn1 were enriched in MKs. We found many known HSC niche genes such as Fgf1, Icam1, Cspg4, Il6, Lepr, Angptl2, Eng, Nes, Tgfb2, Nrp1, Efnb2, Il7r, Pecam1, Epcam, Cdh5, Cxcl12, and Ephb4, Wnt genes such as Dkk2, Fzd2, Prickle2, Fzd4, Vangl2, Sfrp1, Fzd1, and Tcf7l1, and Notch genes such as Notch1, Dll1, Jag2, Dll4, Notch4, and Notch3 were enriched in WT AECs or SECs (Fig. 3a); and niche genes such as Fgf2, Cspg4, Angptl2, Il6, Lepr, Ndn, Meis1, Il7r, Pecam1, Eng, Efnb2, Nes, Nrp1, Fgf1, Pdpn, Tgfb2, Epcam, Cdh5, Igf1, Cxcl12, Ephb4, and Egfr, Wnt genes such as Tcf7l1, Fzd3, Prickle2, Fzd2, Fzd4, Sfrp2, Dkk2, Vangl2, Sfrp1, and Fzd1, Notch genes such as Jag2, Jag1, Dll1, Notch4, Dll4, Notch1, and Notch3 were enriched in Tet2−/− AECs or SECs (Fig. 3b). This suggests that ECs are particularly important for supporting the HSC niche. To further analyze the cell–cell communications in the HSC niche, we performed fluorescence-activated cell sorting (FACS) to enrich the HSCs and integrated scRNA-seq datasets of sorted HSCs and of whole fetal liver (Supplementary Fig. S9a). A cellphone analysis33 of ligand/receptor expression using our scRNA-seq data showed that HSCs were potentially closely communicating with ECs (Supplementary Fig. S10a). Several well characterized Wnt and Notch ligand/receptor pairs underlie this potential interaction (Supplementary Fig. S10b).
a, b Gene expression enrichment profiles of probed genes in different cell types in WT (a) and Tet2−/− (b) fetal livers. All genes used in the MERFISH analysis are shown along the x-axes. Log2 enrichment of expression for each gene is shown by a red/blue color scale (on the right) with darker red indicating a stronger enrichment and darker blue a stronger depletion. The niche genes (dark violet), Wnt genes (deep pink), Notch genes (cyan), and other genes (magenta) were ordered based on hierarchical clustering in each category. All the log2 enrichment values of the highlighted niche/Wnt/Notch genes are larger than 1. AEC, arterial endothelial cell; SEC, sinusoidal endothelial cell; MK, megakaryocyte. c, d An example of Kitl positive hepatocytes with nearby Kit positive erythroid progenitors. Cell types were determined using MERFISH data and mapped in situ (c). Individual mRNA molecules from MERFISH were plotted on top of the segmented cell image for Kitl (cyan dots) and Kit (yellow dots) (d). The right panels in c and d are fourfold magnified views of the white boxed regions in the left panels.
Based on the gene expression enrichment analyses, some niche genes were also enriched in non-EC cell types. For instance, Cxcl12, previously reported as crucial for HSC maintenance in bone marrow34,35,36, was enriched in both AECs and hepatocytes. Kitl (Kit ligand), which supports HSC expansion in fetal liver37, was enriched in hepatocytes. Egr1, known to be expressed in HSCs and to coordinate HSC division and migration38, was enriched in hepatocytes (Fig. 3a, b). A full list of gene expression enrichments in different cell types is provided in Supplementary Table S2. One explanation to these observations is that the different cell types offer the corresponding supportive molecules for HSC regulation. However, these findings also raise the possibility that these niche genes may have additional non-HSC functions within the corresponding cell types. Indeed, as we found that hepatocytes were paired with erythroid progenitors in space (Fig. 2c–f), Kitl and Kit were enriched in hepatocytes and erythroid progenitors, respectively (Fig. 3a, b). In situ maps further confirmed that Kitl-positive hepatocytes were usually near kit-positive erythroid progenitor cells (Fig. 3c, d), suggesting ligand–receptor communication between the two cell types.
We also observed that in WT fetal liver Notch1 was enriched in AECs, SECs, and myeloid cells; Notch2 was enriched in myeloid cells; Notch3 was enriched in AECs; Notch4 was enriched in AECs, SECs, and MKs (Fig. 3a). In Tet2−/− fetal liver, Notch1 was enriched in AECs, SECs, and myeloid cells; Notch2 was enriched in hepatocytes and myeloid cells; Notch3 was enriched in AECs and SECs; Notch4 was enriched in SECs and MKs (Fig. 3b). These suggest that different Notch genes may have cell type-specific functions or cell type biases.
Identification of HSCs and characterization of the HSC niches in WT and Tet2 − /− fetal livers by MERFISH
Although the cellphone analysis of scRNA-seq data suggested that ECs communicate with HSCs (Supplementary Fig. S10a), this sequencing-based analysis only provides the potential molecular interactions, but not the direct spatial information. Thus, it cannot directly measure the frequency at which ECs are physically associated with HSCs in space and the molecular and cellular compositions of the HSC niche. To tackle these questions, we set out to identify HSCs in the MERFISH data. To select the specific markers of HSCs, we identified cell types based on known marker genes from our integrated HSC and whole fetal liver scRNA-seq datasets (Supplementary Fig. S9b). We found that Mecom was specifically and highly expressed in the HSC groups (Supplementary Fig. S9c). In addition, Kit was expressed in HSCs (Supplementary Fig. S9c). This agrees with previous studies showing that Mecom was expressed in 60% of mouse long-term HSCs and at high levels in human hematopoietic stem and progenitor cells39,40, and was regarded as critical for long-term HSC function41. Kit was found essential for quiescent HSC maintenance42, and mice lacking Kit showed hematopoietic defects and perinatal death43. Thus we defined HSCs as positive for Kit and Mecom expression, and used this criteria to identify HSCs in MERFISH data (Supplementary Fig. S5).
To characterize the HSC niche in situ, we measured the cell type compositions and gene expression features in HSC niches in their native spatial context by MERFISH. Approximately 53% and 75% of HSCs in WT and Tet2−/− fetal livers, respectively, were in direct contact with ECs, based on MERFISH imaging. However, this may be an underestimate given that our tissue section largely contained a single layer of cells, and an EC might be in contact with an observed HSC but be excluded from the section. In some instances, HSCs were identified in large regions consisting of many AECs, likely forming a central vein (Fig. 4a, b). We defined an HSC niche as a 20-μm-radius area centered at an HSC (Fig. 4c). A comparison between the proportion of ECs within HSC niches vs the proportion of ECs in the entire cell population showed an enrichment of ECs in the HSC niche (Fig. 4d). Specifically, 8% of the cells in the WT HSC niche were ECs compared to 5% overall (21/254 vs 5,876/112,392, P = 0.0422). This enrichment of ECs was even more dramatic for the Tet2−/− HSC niches, where 28% of the cells were ECs compared to 9% of cells being ECs in the entire population (153/550 vs 12,309/137,628, P < 0.001). In addition, erythroid progenitors were increased in the overall population in the Tet2−/− fetal liver in comparison to WT (19% vs 10%, 26,627/137,628 vs 11,460/112,392, P < 0.001).
a An example of an HSC embedded in a vascular wall seen by MERFISH imaging, cell types were determined and mapped in situ (left panel). b Individual mRNA molecule FISH signals were visualized for Mecom (green dots), Kit (yellow dots), Fstl1 (magenta dots), and Pecam1 (cyan dots) in situ. c An example of an HSC niche identified by MERFISH imaging. The niche area is marked by a white circle with a 20-μm radius. The cell boundaries were segmented based on the membrane labeling with wheat germ agglutinin (WGA). The right panels in a and b are fourfold magnified views of the white boxed regions in the left panels. The HSC boundary is shown in red. d The cell type percentages of the HSC niche and of the total cell population were calculated for WT and Tet2−/− fetal livers (WT niche: n = 254 cells; Tet2−/− niche: n = 550 cells; WT overall: n = 112,392 cells; Tet2−/− overall: n = 137,628 cells). The cell type color scheme of d is the same as a and c. AEC, arterial endothelial cell; SEC, sinusoidal endothelial cell; MK, megakaryocyte. e, f The DEGs between the HSC niche and non-niche in WT (e) and Tet2−/− (f) fetal livers. g HSCs were significantly increased in Tet2−/− (n = 3 biological replicates, 0.848 ± 0.183) compared with WT (n = 4 biological replicates, 0.334 ± 0.301) in E14.5 fetal livers (unpaired t-test with Welch’s correction, P = 0.04). h The DEGs between the WT and Tet2−/− HSC niches. In e, f, and h, the horizontal dotted line represents an adjusted P value = 0.05 using a Wilcoxon rank-sum test with false discovery rate correction. The vertical lines represent a fold change = 2. Blue dots are Notch pathway genes, red dots are Wnt pathway genes, purple dots are vasculature genes, and black dots are other genes. All the labeled dots are reproducible significant genes.
We further investigated the DEGs in HSC niches vs outside of the niches. The endothelial arterial marker, Ephrin-B2 (Efnb2) showed increased expression in both WT and Tet2−/− HSC niches in comparison to cells outside of the niches (Fig. 4e, f). Other endothelial markers highly expressed in the Tet2−/− HSC niche included Fstl1, Nrp1, Cdh5, and Clec14a (Fig. 4f). The Notch signal pathway genes Notch3 and Dll4 were significantly upregulated in both WT and Tet2−/− HSC niches (Fig. 4e, f).
Based on the MERFISH results, Tet2−/− mice exhibited an increased number of HSCs when compared with WT (Fig. 4g). This is in agreement with some prior studies showing that LSK and CD150+ HSCs were increased in extramedullary hematopoiesis of adult Tet2−/− mice18, but does not support some other studies reporting that HSC abundance in Tet2−/− fetal livers did not change17. The increased HSC number we observed in Tet2−/− fetal livers may be caused by changes in the HSC niche.
We next compared the DEGs in the HSC niches in Tet2−/− vs WT fetal livers (Fig. 4h). Most notably, several Wnt signaling pathway related genes were significantly upregulated in the Tet2−/− HSC niche. This included canonical Wnt Frizzled (Fzd) receptors Fzd1, Fzd7, and Fzd8. In contrast, Fzd2, Fzd3, Fzd4, and Fzd5, the downstream signaling gene Axin2, and the responsive transcription factors Lef1, Tcf7, Tcf7l1, and Tcf7l2 were not increased. The non-canonical Wnt signaling genes Vangl2 and Prickle2 also showed increases in expression, but not Celsr2. In addition to Wnt signaling agonists, other upregulated genes include Wnt antagonists, such as Nkd2 and Sfrp1, but not Sfrp2. Interestingly, the most significantly upregulated gene in the Tet2−/− HSC niche was a member of the dickkopf family, Dkk3, while there was no enrichment of Dkk2 expression. Whereas DKK2 binds to LRP5/6 and inhibits the interaction of Wnt to the Frizzled receptors44, DKK3 is the most divergent member of the DKK family and does not bind to LRP5/645. Rather, both positive and negative roles have been shown for DKK3 in Wnt signaling45,46.
In addition to Wnt signaling, several other genes important for cell proliferation were also upregulated in the Tet2−/− HSC niche in comparison to the WT HSC niche, including Notch1, Notch3, Cxcl12, Kitl, Fgf2, and Il6 (Fig. 4h). Notch signaling has been shown to increase stem cell numbers in mouse bone marrow47,48. Cxcl12 is critical for HSC maintenance: HSCs were reduced in number and became more quiescent in the absence of CXCL12-abundant reticular cells34,35,36. KIT-ligand (Kitl) induces HSC maintenance in the bone marrow49 and contributes to HSC expansion in the mouse fetal liver37. FGF-2 stimulates HSC proliferation in vivo through Kitl signaling and stromal cell expansion50. Il6 signaling promotes HSPC proliferation in mice51, and has a combinatorial effect with Notch activation on hematopoietic cells52. In summary, our results suggest that loss of Tet2 leads to an increase of HSCs in fetal liver due to the upregulation of Wnt, Notch, and other niche factors that support HSC proliferation.
Distinct signaling compositions of AECs and SECs in HSC niches
Previous studies suggested that both AECs and SECs were essential for HSC regeneration and maintenance5,53. To further identify the specific roles of AECs and SECs, we first analyzed their DEGs (Fig. 5a, b). In both WT and Tet2−/− fetal livers, SECs expressed specific endothelial markers such as Flt4, Stab2, Kdr, Cdh5, Clec14a, Tek, and Lyve1, whereas AECs specifically expressed other endothelial markers such as Fstl1 and Pdgfra. In addition, SECs from both WT and Tet2−/− fetal livers expressed the Notch pathway genes Notch4, Dll1 and Dll4, and the niche factor Lepr, whereas AECs expressed Notch3 and Cxcl12.
a, b The DEGs of AECs vs SECs in WT (a) and Tet2−/− (b) fetal livers by MERFISH. The horizontal dotted line represents an adjusted P value = 0.05 using a Wilcoxon rank-sum test with false discovery rate correction. The vertical lines represent a fold change = 2. Blue dots are Notch pathway genes, red dots are Wnt pathway genes, green dots are niche factors, purple dots are vasculature genes, and black dots are other genes. All the labeled dots are reproducible significant genes. c, d Correlation coefficients of niche genes with cell type markers in WT (c) and Tet2−/− (d) HSC niches. The genes listed in c and d all showed significant correlations (R > 0.1, P < 0.05) with the corresponding cell type markers (dark blue = AEC, yellow = SEC, light blue = hepatocyte, green = macrophage, pink = megakaryocyte). The P values were corrected using false discovery rate.
The analysis above included AECs and SECs inside and outside of HSC niches. To explore the co-expression of various niche factors among cells within HSC niches, we calculated the correlation coefficients of pairs of genes co-expressed in the same HSC niche cells in the WT (Fig. 5c) and Tet2−/− (Fig. 5d) fetal livers using the MERFISH data. Note the scRNA-seq procedure removes a large population of ECs in fetal liver (Supplementary Fig. S6), and has lower cell number throughput than MERFISH in general (Fig. 1a with 136 ECs out of 7,635 cells in total and Fig. 1c with 2,546 ECs out of 40,864 cells in total). Besides, to analyze the gene co-expression in niche requires the cellular spatial distribution information, which is not preserved by scRNA-seq, but by MERFISH. We found that in WT HSC niches, niche genes Nes, Nrp1, Cxcl12, Lepr, Bmp5, Tet2, Cspg4, Angpt1, and Epcam, Wnt and Notch signaling genes Dll4, Notch3, Sfrp1, Jag1, Fzd1, Tcf7l2, and Ep300 were correlated with AEC, but not SEC, marker genes. In contrast, niche genes Tgfb2, Il7r, Pdpn, Meis1, Mpl, Ephb4, Eng, and Tet1, and Wnt and Notch signaling genes Prickle2, Dll1, Rbpj, and Notch4 were correlated with SEC, but not AEC, marker genes (Fig. 5c). In Tet2−/− HSC niches, niche genes Igf1, Meis1, Angptl2, Cspg4, Fgf2, Ndn, Egfr, Gfap, Tet2, and Angpt1, and Wnt and Notch signaling genes Sfrp1, Notch3, Fzd1, Sfrp2, Fzd2, Vangl2, Notch2, Tcf7l1, Fzd7, Tcf7l2, Fzd4, Fbxw7, Fzd8, and Rbpj were correlated with AEC marker genes; niche gene Pdpn, and Notch signaling genes Dll4 and Jag2 were correlated with SEC marker genes (Fig. 5d). Note that the expression of Tet2 mRNA in the Tet2−/− mice is expected based on the knockout design that removes only exons 8 through 1154. These results indicate that AEC and SEC may contribute different niche molecules to HSC niche in fetal liver.
We also observed several niche or signaling genes correlated with markers of other cell types in the HSC niche. For example, in WT HSC niches, Egfr, Notch2, and Fzd2 were co-expressed with hepatocyte marker genes; Fgf1, Cdh5, Efnb2, Bmp2, Bmp7, and Tcf7 were co-expressed with macrophage marker genes; and Il6, Nkd2, Pecam1, Lef1, and Ndn were co-expressed with megakaryocyte marker genes (Fig. 5c). In Tet2−/− HSC niches, Kitl, Egr1, Dkk3, Bmp7, Tcf7, Icam1, and Fzd3 were co-expressed with hepatocyte marker genes; Jag1, Lef1, Dkk2, Prickle2, Dll1, Notch4, Maml1, Tgfb2, and Eng were co-expressed with megakaryocyte marker genes (Fig. 5d). A full list of correlations between niche/signaling genes and cell type marker genes is provided in Supplementary Table S3. These results suggest that cell types other than ECs may also provide additional supporting molecules to the HSC niche.
Discussion
In summary, using MERFISH and scRNA-seq we showed that fetal livers display intricate cellular and molecular organizations, indicating potential cell–cell communications and collaborations. HSCs locate predominantly in endothelial niches and the majority of HSCs directly contact one or more ECs (AECs and/or SECs). AECs and SECs possess different molecules and may establish distinct functions for HSCs. Other cell types may also contribute to HSC regulation, as well as regulating each other (e.g., hepatocytes may regulate erythroid progenitor development). Loss of Tet2 led to an increased number of HSCs and an increased expression of Wnt and Notch signaling genes as well as other niche factors within the HSC niches, which potentially drives the HSC expansion.
Single cell in situ transcriptomics provides a valuable tool to answer many long-standing questions in biology that require an appreciation of tissue architecture. Previous studies using high throughput RNA smFISH-like techniques, including STARmap55, seqFISH+56, or MERFISH27,28,29, often dealt with tissues with well-separated cell bodies (e.g., brain tissue) or cell lines grown in vitro at densities that minimized cell–cell contact. Under these conditions, it is possible to perform cell segmentation by Nissl staining or RNA density. In contrast, the fetal liver contains many closely packed cells. Thus, to effectively detect cell boundary and segment single cells in fetal liver tissue sections is challenging. In our study, we used WGA, which marks the plasma membrane of eukaryotic cells by binding to the N-acetylglucosamine of glycoproteins and glycolipids57. Moreover, we attached WGA to an oligonucleotide to facilitate the visualization of WGA binding pattern in a manner compatible with MERFISH tissue clearing, and successfully segmented single cells in fetal liver tissue sections (Supplementary Fig. S3). We expect this procedure to be widely applicable to many tissue types, and to enable MERFISH single cell analyses of tissues with tightly packed cells.
Consistent with prior work10, we found that fetal liver HSCs highly colocalize with ECs. Extending Khan et al.’s observation that Efnb2-positive portal vessels were abutting Nestin+ cells, which were enriched for fetal liver HSC niche and expansion factors10, we discovered that Efnb2 is more highly expressed in the fetal liver HSC niche than outside of the niche. Not elucidated in previous studies, we revealed that the majority of HSCs are directly touching at least one EC. Furthermore, we demonstrated that both AECs and SECs can be present in the HSC niches, with differences in transcripts enriched in these two cell types. Dll4, in particular, is enriched in the fetal liver HSC niche compared to non-niche cells (Fig. 4e, f). This is consistent with a prior report that vascular DLL4 acted as a key Notch ligand that controlled early hematopoietic differentiation58 and that Notch signaling promoted stem cell self-renewal and inhibited differentiation47. Interestingly Dll4 is associated with AECs in WT niche (Fig. 5c), but with SECs in Tet2−/− niche (Fig. 5d). This switch may underlie the changed HSC behavior upon Tet2 knockout. Fzd1, which is associated with AECs but not with SECs (Fig. 5c, d), is a specific receptor of Wnt3a which is known to be particularly important for regulating HSC fate59,60. This suggests a specific importance of AECs to provide Fzd1 in the niche. Experimental validation of our findings will require the development of AEC- and SEC-specific transgenic Cre lines to specifically knock out the niche factor genes from the appropriate cell population to observe their influence on HSC functions. We also illustrated the expression of other genes known to influence HSCs as being correlated with specific cell types. Taken together, our MERFISH findings suggest that many different types of cells contribute various signaling components to the HSC niche.
Given the importance of the epigenetic modifier Tet2 to HSC development, we evaluated the HSC niche of Tet2−/− fetal liver. Kunimoto et al. showed that disruption of Tet2 enhanced the self-renewal and long-term repopulating capacity of fetal liver HSCs in vivo17. We found increased numbers of HSCs by MERFISH in the Tet2−/− fetal liver, which can be explained by both HSC cell-autonomous and non-cell-autonomous mechanisms. TET2 is an epigenetic regulator responsible for the hydroxylation of 5mCs to 5-hmCs, leading to DNA demethylation15,61,62. Loss of TET2 leads to both direct and indirect changes in gene expression63,64. Use of conditional knockouts and competitive bone marrow transplantation assays has shown that Tet2 deletion augments the hematopoietic stem/progenitor cell population in a cell-autonomous manner18,54,65,66. In addition, loss of Tet2 from bone marrow mesenchymal stromal cells also augments BMSC self-renewal and proliferation, which enhances their hematopoietic supportive capacity19. Similarly, loss of Tet2 leads to an increase of the EC population in fetal liver (Fig. 4d), which may promote HSC expansion in a non-cell-autonomous manner. Therefore, Tet2 deficiency may affect HSC functions in cell-autonomous and non-cell-autonomous manners.
Khan et al. suggested that WT HSCs did not expand by increasing Kitl, Angptl2, or Igf2 in Nestin+ HSC-niche cells, because these factors were expressed at a similar level at E12, E13, and E14.5 in fetal liver10. However, in the Tet2−/− setting, we found that HSC expansion factors such as Cxcl12, Kitl, Fgf2, and Il6 are increased when compared with WT fetal liver. In addition, we observed several Wnt and Notch signaling genes were strongly enriched in the Tet2−/− HSC niche compared to non-niche cells and compared to the WT HSC niche. The complex and nuanced native microenvironment that we elucidated with MERFISH is difficult to recapitulate by in vitro coculture systems. We look forward to the generation of advanced in vitro systems where pure HSC and niche cells can be precisely isolated and cultured in order to help functionally test some of the niche interactions learned by MERFISH. Deletion of each of the identified niche factor genes in the Tet2 null setting would be necessary to validate their influence on fetal liver HSC functions. In addition, we report that Kitl-positive hepatocytes are usually adjacent to Kit-positive erythroid progenitor cells, which suggests ligand-receptor communications between the two cell types. This observation of a potential hematopoietic signaling function of hepatocytes is consistent with previous studies that Kitl-expressing fetal liver hepatoblasts support the expansion of Kit-expressing HSCs and potential survival or expansion of hematopoietic progenitors37. Functional data also show that hepatoblast-depleted mouse embryos display a decrease in Kitl gene expression and a reduction of the Kit+ HSCs and hematopoietic progenitors67,68. Previous reports also show that hepatocyte growth factors can directly control erythroid progenitor proliferation and differentiation69. Using MERFISH to further refine the gene expression map of the HSC microenvironment under different physiological or pathological conditions will improve our understanding of HSC expansion and renewal and of ways to manipulate them.
Materials and methods
Animal preparation and tissue sectioning
E14.5 pregnant female C57BL/6 WT and Tet2−/− mice were used for fetal liver preparation. All mice were maintained under 12 h light/12 h darkness cycles at 22 °C. All animal experiments were approved by the Institutional Animal Care and Use Committee of Yale University. Pregnant females were euthanized by isoflurane inhalation and cervical dislocation. E14.5 fetal livers were dissected from embryos in cold dulbecco’s phosphate-buffered saline (DPBS) and embedded in Tissue-Tek Cryomold (VWR, 25608-916) with optimal cutting temperature compound (Tissue-Tek O.C.T.; VWR, 25608-930). Frozen tissue blocks were stored at −80 °C before cryosection. E14.5 fetal liver frozen blocks were equilibrated to −15 °C for 1 h and then cryosectioned into 10-μm-thick slices using a Leica CM1950 machine at −15 °C. Tissue sections were post-fixed immediately with 4% PFA (Electron Microscopy Sciences, 15700) in DPBS while on salinized and poly-L-lysine (Millipore, 2913997) treated coverslips (Bioptechs, 40 mm Coverslip #1.5Thick) for 20 min at room temperature. After washing out the PFA with DPBS, tissue sections were stored in 100% ethanol at −20 °C. Data were collected from four WT fetal livers at E14.5 and two Tet2 knockout fetal livers at E14.5.
MERFISH probe design and construction
Primary probes of MERFISH were designed using the 16-bit Hamming-distance-4 code with 140 barcodes as previously described27,28,29. The 140-gene library contained 45 cell type marker genes from our 10× Genomics single cell sequencing data of E14.5 fetal liver and 95 HSC niche-associated genes from the literature. We chose 98 of the library gene isoforms that were long enough to construct 48 target regions using a non-overlap design and 42 of the library gene isoforms with an overlap design as previously published70. Each MERFISH oligo had a 30-nt targeting region, three 20-nt readout regions, and two 20-nt priming regions. The template oligonucleotide sequences for MERFISH were listed in Supplementary Table S4. The gene FPKM values of MERFISH library genes ranged from 0.16 to 794 based on matched bulk RNA-sequencing.
Primary probe synthesis
The primary probes were constructed from a complex oligonucleotide pool that was ordered from CustomArray, Genscript27,28,29. Primary probes were synthesized via limited cycle PCR to create the templates for in vitro transcription (The forward PCR primer (with T7 promoter) sequence is GCCGTACGGATAATACGACTCACTATAGGG GCGTCGTTATGGTGCAACGT, and the reverse PCR primer sequence is TTGTCGCACGTTCGGTGTCG). These templates were converted into RNA through in vitro transcription, then converted back to single stranded DNA using reverse transcription (The reverse transcription primer sequence is TTGTCGCACGTTCGGTGTCG). RNA templates were removed from the DNA oligos via alkaline hydrolysis. Finally, the DNA oligos were purified through a Zymo column and concentrated via vacuum drying.
Silanization of coverslips
We coated the coverslips with a silane layer containing an allyl moiety to stabilize a polyacrylamide (PA) film introduced in the later “Tissue embedding and clearing” step29. Briefly, the coverslips were cleaned in a 1:1 mixture of 37% (vol/vol) HCl and methanol for 30 min at room temperature. Then, coverslips were rinsed in deionized water three times and in 70% ethanol once. Coverslips were dried in a 70 °C oven before being immersed in 0.1% (vol/vol) triethylamine (Millipore, TX1200-11) and 0.2% (vol/vol) allytrichlorosilane (Sigma, 107778-5 G) in chloroform for 30 min at room temperature. Finally, the coverslips were washed once with chloroform and once with 100% ethanol, then dried in a 70 °C oven for 1 h and stored in a drying basin.
WGA conjugation
WGA (VECTOR, B-1025) was labeled with a copper-free click crosslinking agent using NHS-ester chemistry similar to previous reports70,71. DBCO-PEG5-NHS Ester (Kerafast, FCC310) was diluted to a concentration of 10 mM in anhydrous dimethyl sulfoxide (DMSO). Overall, 2.7 μL of the solution was then combined with 100 μL of 2 mg/mL WGA in DPBS. After 1 h at room temperature the reaction was terminated via an Amicon Ultra-0.5 Centrifugal Filter Unit (Millipore, UFC501024) at 14,000× g for 15 min, and then 1,000× g for 2 min to collect the DBCO labeled WGA. On average, the procedure produced ~2 DBCOs per WGA. Oligonucleotide probes, ordered from IDT and diluted to 100 μM in DPBS, contained the desired sequence (named OS_61r, /5Acryd/CGGTACGCACTTCCGTCGACGCAATAGCTC/3AzideN/) as well as a 5′-acrydite, to allow cross-linking to the PA gel, and a 3′-azide, to allow cross-linking to the DBCO-labeled WGA. A total of, 20 μL of the oligonucleotide was added to 100 μL of the DBCO-labeled WGA at a final concentration of ~2 mg/mL. This reaction was incubated at 4 °C for at least 12 h. The residual oligonucleotides were washed away from samples during primary probe staining.
Primary probe staining
The primary probe staining procedure was similar to those in previous reports27,28,29. Tissue sections were stained with oligo-conjugated WGA in 1× hank’s balanced salt solution (HBSS) buffer for 20 min at 37 °C. Samples were post-fixed in 4% PFA for 10 min. Tissue sections were permeabilized using 0.5% Triton in DPBS for 20 min at room temperature before primary probe staining. Tissue samples were incubated for 5 min in pre-hybridization buffer composed of 50% (vol/vol) formamide and 2 mM Ribonucleoside vanadyl complexes (Sigma-Aldrich, R3380) in 2× saline sodium citrate (2× SSC) (Invitrogen, AM9765). Tissue samples were then stained with primary probes in primary hybridization buffer, containing 24-28 μM primary probes, 50% (vol/vol) formamide, 0.1% yeast tRNA (Invitrogen, 1885325), 1% (vol/vol) murine RNase inhibitor (NEB, M0314S), 10% (wt/vol) dextran sulfate (Millipore, S4030), and 2 μM anchor probe (a 15-nt sequence of alternating dT and thymidine-locked nucleic acid with a 5′ -acrydite modification) in 2× SSC, in a humidity chamber at 37 °C for 24 h. After staining, samples were washed for 15 min with 2× SSCT (2× SSC, 0.1% (vol/vol) Tween 20) twice at 60 °C and then once for 15 min at room temperature.
Tissue embedding and clearing
To anchor RNAs and the oligos conjugated with WGA, the primary probe stained samples were embedded in 8% PA gels29. Briefly, a 1:30,000 dilution of 0.1-μm-diameter carboxylate-modified yellow-green fluorescent beads (Invitrogen, F8800) in 2× SSC containing 1% (vol/vol) VRC (NEB, S1402) was first incubated with the stained samples for 10 min at room temperature. Then, the samples were washed for 2 min with a de-gassed PA gel solution, containing 8% (vol/vol) 19:1 acrylamide/bis-acrylamide (Bio-Rad, 1610144), 60 mM Tris·HCl pH 8, 0.3 M NaCl, 0.03% (wt/vol) ammonium persulfate (Sigma, A3678-25G), and 0.15% (vol/vol) TEMED (AMERICANBIO.COM, AB02020-00050). To cast a thin PA gel film on the tissue samples, 50 μL of this PA gel solution was added to the surface of a glass plate that had been pretreated for 10 min with 1 mL GelSlick solution (Lonza, 50640). The sample coverslip was washed with PA gel solution and the excess solution on the coverslip was gently removed by dipping the edge of the coverslip on a Kimwipe tissue. Then, the sample coverslip was inverted and placed onto this 50 μL droplet to form a thin layer of PA gel between the coverslip and the glass plate. The PA gel was cast for 1.5 h at room temperature. The coverslip was then gently separated from the glass plate and incubated in digestion buffer for 12 h in a humid 37 °C incubator. The digestion buffer contained 2× SSC, 2% (vol/vol) SDS, and 1% (vol/vol) proteinase K (ThermoFisher, AM2548). The tissue was then washed with 2× SSC supplemented with 1% Murine RNase inhibitor on a shaker, three times each for 15 min.
MERFISH imaging
After primary probe hybridization and tissue clearing, the samples were sequentially hybridized with different secondary probes, imaged and washed as previously reported27,28,29. We used Alexa Fluro 750/647-conjugated 20-nt secondary probes with sequences complementary to the readout regions of the primary probes (The secondary probe sequences are the same as previously reported)28,29 and an ATTO 565-conjugated 30-nt probe (OS_61, /5ATTO565N/AGAGCTATTGCGTCGACGGAAGTGCGTACCG) with the complementary sequence of the WGA-conjugated oligonucleotide (OS_61r). To label the cell membrane, we hybridized OS_61 (6 nM probes) in secondary hybridization buffer (2× SSC, 20% (vol/vol) ethylene carbonate (Sigma-Aldrich, E26258), 0.05% murine RNase inhibitor) to the samples at room temperature on a shaker for 15 min. Then we used a Bioptech’s FCS2 flow chamber and a home-built fluidics system to perform automatic buffer exchange during the multiple rounds of secondary hybridization. For each round of secondary hybridization, the samples were first incubated with the secondary hybridization buffer with 3 nM appropriate secondary probes for 10 min. We then sequentially washed the samples with 2 mL secondary wash buffer (20% (vol/vol) ethylene carbonate in 2× SSC) for 5 min and 2 mL imaging buffer with an oxygen scavenging system (50 mM Tris-HCl pH 8.0, 10% (wt/vol) glucose (Sigma, G8270), 2 mM Trolox (Sigma-Aldrich, 238813), 0.5 mg/mL glucose oxidase (Sigma-Aldrich, G2133), 40 μg/mL catalase (Sigma-Aldrich, C30), 0.05% (vol/vol) murine RNase inhibitor in 2× SSC) for 5 min. The imaging buffer in the input tube of the fluidics system was covered with a layer of mineral oil (Sigma, 330779) to prevent continuous oxidation. Next, the samples were imaged at multiple fields of view. For each field of view, we took z-stack images with 750, 647, 560, and 488 nm laser illuminations. The range of the z-stacks was 9 μm overall, with 16 steps of 0.6 μm intervals. After each round of imaging, we flowed secondary probe removal buffer (65% formamide in 2× SSC) for 10 min to wash out secondary probes.
Imaging system
For imaging, we used a home-built microscope with a Nikon Ti2-U body, a Nikon CFI Plan Apo Lambda 60× Oil (NA1.40) objective lens, and an active auto-focusing system as described before72. A 750-nm laser (2RU-VFL-P-500-750-B1R, MPB Communications) was used to excite and image Alexa Fluor 750 on secondary probes. A 647-nm laser (2RU-VFL-P-1000-647-B1R, MPB Communications) was used to excite and image Alexa Fluor 647 on secondary probes. A 560-nm laser (2RU-VFL-P-1000-560-B1R, MPB Communications) was used to excite and image ATTO 565 on cell membrane probe OS_61. A 488-nm laser (2RU-VFL-P-500-488-B1R, MPB Communications) was used to excite and image the yellow-green fiducial beads for drift correction. The four laser lines were directed to the samples using a multi-band dichroic mirror (488/561/647/752rpc-UF2, Chroma) on the excitation path. On the emission path, we had a multi-band emission filter (488/561/647–656/752 m, Chroma) and a Hamamatsu Orca Flash 4.0 V3 camera. The pixel size of our system was measured to be 107.9 nm per pixel. To automatically scan and image multiple fields of view, we used a computer-controlled motorized x-y sample stage (SCAN IM 112 × 74, Marzhauser).
scRNA-seq
For sorted HSC scRNA-seq, fetal liver tissues were isolated from E14.5 C57BL/6 pregnant mice and in cold 1× PBS dissociated into single cell suspension by pipetting up and down through 15 mL pipette. Red blood cells were removed using 1× RBC lysis buffer (diluted into 1× in deionized water from 10× RBC lysis buffer, Santa Cruz Biotechnology, sc-296258) at room temperature for 5 min. Cells were incubated for 5 min at room temperature and mixed constantly by inverting the tube. After that, cells were centrifuged for 5 min and then the supernatant was carefully removed without disturbing the pellet cells. Cells were resuspended in cold 1× PBS with 0.5% BSA, passed through a 40 μm strainer (Falcon, 352235). Fetal liver cells were used for FACS to isolate HSCs as described below. For whole fetal liver scRNA-seq, fetal liver tissues were isolated from E14.5 C57BL/6 pregnant mice and digested by dissociation enzyme (Accutase, A1110501) for 10 min at room temperature. Red blood cells were removed using 1× RBC lysis buffer (Santa Cruz Biotechnology, sc-296258) at 4 °C. Then, cells were passed through a 40 μm strainer (Falcon, 352235) and re-suspended in cold DPBS with 0.04% (wt/vol) BSA. We measured cell viability with a TC20TM Automated Cell Counter (Bio-Rad), and then took 8,000 cells for 10× library preparation using Chromium Single Cell 3′ Reagent Kits (v2). The libraries were run on an Illumina HiSeqX10 as 150 bp paired-end reads and occupied one full lane.
FACS sorting of fetal liver HSC
HSCs were sorted under the same condition as previously described73. Before sorting HSCs, we added Biotin Lineage depletion cocktail (anti-Mac1, anti-CD3ε, anti-B220, anti-Gr-1, anti-CD11b, and anti-Ter119) and streptavidin Particles Plus-DM (BD IMag™, mouse hematopoietic progenitor (stem) cell enrichment set). Lineage positive cells were depleted by transferring cells to a magnetic stand. Enriched lineage negative cells were further stained with antibodies (LSK CD150+CD34−CD48−Flt3−) listed in Supplementary Table S5. HSCs were sorted with FACS AriaII (BD Biosciences) from the Yale Cell Sorter Core Facility.
Processing of scRNA-seq data
The FASTQ format results were analyzed using Cell Ranger (https://support.10xgenomics.com/single-cell-gene-expression/software/pipelines/latest/using/tutorial_ov) for sample demultiplexing, barcode processing, and gene counting. The reference genome mm10/GRCm38 build by 10× genomics was used to align inserted cDNA. According to cell barcode and unique molecular identifiers, 9,448 single fetal liver cells and 8,613 single fetal liver HSCs were identified. Further analysis was performed using the Seurat R package74, including quality filtering, the identification of highly variable genes, dimensionality reduction, standard unsupervised clustering, and the discovery of DEGs. We removed low-quality cells that had fewer than 500 detected genes and genes expressed in less than three cells, as well as cells with more than 10% of the transcripts coming from mitochondrial genes. Next, we normalized the data by the total expression, multiplied the data by the scale factor 10,000, added 1 and log-transformed the result. To remove batch effects, we used the Seurat alignment method CCA (canonical correlation analysis) for data integration. We used 2,500 anchor features for the canonical correlation analysis. The subspaces were aligned on the basis of the first 30 canonical correlation vectors to generate a combined aligned dataset to use for further integrative analysis. To find out HSC markers at the transcriptome level, we further reduced dimensionality of the entire integrated dataset using Uniform Manifold Approximation and Projection (UMAP), whereby the major cell type identification was based on the Mouse Cell Atlas30. We used the cell type terms and the corresponding marker genes reported in the associated web database: http://bis.zju.edu.cn/MCA/gallery.html?tissue=Fetal-Liver. To analyze the GO of hematopoietic cells, we calculated the DEGs using Seurat, which was used as an input for GO analysis with Clusterprofiler75. For the RNA velocity analysis, we first generated the spliced and unspliced RNA loom file using velocyto, and then performed the velocity analysis using velocity.R in Seurat, and the cells were visualized using UMAP. For the cellphone analysis, we first converted the mouse genes to human homologous genes using biomaRt76, and then analyzed the interaction between different cell types using the integrated dataset via CellphoneDB33.
Bulk RNA sequencing
Total RNA was extracted from E14.5 fetal liver using RNeasy Mini Kit (QIAGEN, 74104). mRNA was isolated using the Dynabeads mRNA Purification Kit (THERMOFISHER, 61006). The cDNA libraries were produced with a NEBNext® Ultra™ II RNA Library Prep Kit for Illumina® (NEB, E7770S) and NEBNext® Multiplex Oligos for Illumina® (NEB, E7335S) according to the manufacturer’s description. Finally, the cDNA libraries were sequenced on an Illumina HiSeqX10 platform with 150 bp paired-end reads.
MERFISH data processing
All MERFISH analyses in this study were performed with MATLAB (MATLAB, 9.6.0.1150989 (R2019a) Update 4). To efficiently analyze MERFISH images, we implemented a pixel-based MERFISH analysis pipeline similar to that introduced in a previous report27,28,29. We corrected the sample drift between different rounds of MERFISH imaging based on the fiducial beads and decoded the RNA signals according to each drift-corrected RNA image. We matched the decoded RNA signals with our MERFISH codebook (Supplementary Table S1). The molecular counts detected by MERFISH for each RNA species were compared with the FPKMs from bulk RNA-seq by Pearson correlation analysis. MERFISH data were used for further analysis only when the correlation coefficient was greater than 0.7. When correlating MERFISH copy numbers and bulk RNA-seq FPKMs, we found the expression of a small set RNA species (eight RNA species, including Angpt2, Ddx4, Dkk1, Gata3, Hoxb5, Selp, Wnt2, Wnt4) were not consistent with bulk RNA-seq data. These outlier genes were removed before the next steps of analyses. The displayed correlation analysis of FPKM vs total RNA copy numbers for WT (Fig. 1e) and the correlation of total RNA copy numbers from two MERFISH experiments (Fig. 1f) were from one of the biological replicates.
We segmented the tissue section into single cells based on the WGA labeling pattern using a watershed algorithm, and assigned RNA counts into individual cells based on cell boundaries to generate the gene by cell expression matrix as previously described77. When analyzing single-cell RNA data, we removed “cells” larger than 20,000 pixels or less than 2,500 pixels in area as these were usually empty regions or non-cell particles in the tissue section. We removed partial cells overlapping the edges of each field of view. We removed cells with < 10 detected RNA molecules to ensure high quality in the cell type identification analyses. To cluster cells based on the RNA copy number profiles, we applied the Louvain–Jaccard clustering algorithm78, and visualized the cell clusters with UMAP. We identified the cell types of the clusters based on the RNA expression patterns in the clusters. The displayed MERFISH single cell cluster result (Fig. 1c) was from one of the WT biological replicates.
To analyze the HSC niches in situ, we determined HSCs according to the expression of Mecom and Kit. First, we calculated the 99.95 percentile of cells based on the RNA copy numbers of Mecom and set this copy number as the Mecom threshold, where cells with a greater Mecom RNA copy number than this threshold were considered to be potential HSCs. Then, we identified HSCs from potential HSCs whose copy numbers of Kit were equal to or greater than 1. After the HSC identification, we defined a niche by selecting cells surrounding an HSC within a radius of 20 μm. For all the DEGs and gene co-expression analyses, we first normalized the expression of each gene in each cell: the copy number of each RNA species in the cell was divided by the copy number of all labeled RNA species in the cell, and was then multiplied by 1,000. The normalized copy numbers are proportional to the fractions of total RNA counts that belong to each RNA species within the cells.
To calculate the gene expression enrichment for each cell type, we first calculated the average gene expression profile of each cell type, and then divided the profile by the average gene expression profile of the entire cell population. The result was then log2 transformed. For better display, we further clustered and reordered the niche genes, Wnt genes, Notch genes, and other genes using hierarchical clustering. The displayed log2 gene expression enrichment matrices for WT and Tet2−/− (Fig. 3a, b) were derived from all biological replicates.
In the gene-gene correlation analysis, we calculated the correlation coefficients between the expression levels of pairs of genes among individual cells in the HSC niche. The displayed correlation coefficients for WT and Tet2−/− niche (Fig. 5c, d) were derived from the niche cells from all biological replicates. When a niche gene was correlated with multiple marker genes of the same cell type, we displayed the highest correlation coefficient. The P values for testing the multiple hypotheses were adjusted using false discovery rate. All significant correlations, defined as correlations with correlation coefficients R > 0.1 and adjusted P < 0.05, were listed in the Supplementary Table S3.
The enrichment of pairs of cell types in spatial proximity was calculated by first measuring the probability of finding each pair of cell types within 20 μm distance, and then normalizing this probability to the control probability derived from randomly pairing cell types given their abundance in the whole population. The P values of the enrichment were calculated by chi-square test and corrected using false discovery rate. The enrichment folds were log2 transformed for display. The displayed enrichment of pairs of cell types in spatial proximity for WT and Tet2−/− (Fig. 2a–f) were from all the biological replicates in each case. To determine reproducible enrichment observations (cyan dots in Fig. 2a–d), an enrichment needs to be reproducibly significant (P < 0.01) in three individual biological replicates (out of four total biological replicates for WT and three total biological replicates for Tet2−/−).
All “Adjusted P value” shown in this paper were corrected for multiple hypothesis testing using false discovery rate. To analyze single cell data for reproducibility, we divided the data into two replicate groups (each group contains multiple independent biological replicates) for WT and Tet2−/−, and performed the DEGs analysis for each replicate group. Only genes that were significant in the total data (with a fold change ≥ 2 one way or the other, and P < 0.05) and in both replicate groups (with a fold change trend consistent with the total data and P < 0.1) were reported as significant DEGs. Any genes that were not significant in either of the replicate groups were disregarded as unreproducible findings. The displayed DEG volcano plots show results from all biological replicates.
Data availability
Raw imaging data are available from S.W. on request. Analyzed imaging data are available at: https://campuspress.yale.edu/wanglab/HSCMERFISH. Bulk RNA sequencing data from this work are available at: GSE172128. Single cell RNA sequencing data from this paper are available at: GSE172127.
Code availability
Data were collected using open source python codes from https://github.com/ZhuangLab/storm-control. All MATLAB programs and R scripts for data analysis are available at https://campuspress.yale.edu/wanglab/HSCMERFISH.
References
Boisset, J. C. et al. In vivo imaging of haematopoietic cells emerging from the mouse aortic endothelium. Nature 464, 116–120 (2010).
Ema, H. & Nakauchi, H. Expansion of hematopoietic stem cells in the developing liver of a mouse embryo. Blood 95, 2284–2288 (2000).
Pinho, S. & Frenette, P. S. Haematopoietic stem cell activity and interactions with the niche. Nat. Rev. Mol. Cell Biol. 20, 303–320 (2019).
Gao, X., Xu, C., Asada, N. & Frenette, P. S. The hematopoietic stem cell niche: from embryo to adult. Development https://doi.org/10.1242/dev.139691 (2018).
Shestopalov, I. A. & Zon, L. I. Stem cells: the right neighbour. Nature 481, 453–455 (2012).
Orkin, S. H. & Zon, L. I. Hematopoiesis: an evolving paradigm for stem cell biology. Cell 132, 631–644 (2008).
Wei, Q. & Frenette, P. S. Niches for hematopoietic stem cells and their progeny. Immunity 48, 632–648 (2018).
Hardy, R. R. & Hayakawa, K. Perspectives on fetal derived CD5+ B1 B cells. Eur. J. Immunol. 45, 2978–2984 (2015).
Wilson, A. et al. Hematopoietic stem cells reversibly switch from dormancy to self-renewal during homeostasis and repair. Cell 135, 1118–1129 (2008).
Khan, J. A. et al. Fetal liver hematopoietic stem cell niches associate with portal vessels. Science 351, 176–180 (2016).
Kunisaki, Y. et al. Arteriolar niches maintain haematopoietic stem cell quiescence. Nature 502, 637–643 (2013).
Chotinantakul, K. & Leeanansaksiri, W. Hematopoietic stem cell development, niches, and signaling pathways. Bone Marrow Res. 2012, 270425 (2012).
Luis, T. C. et al. Canonical wnt signaling regulates hematopoiesis in a dosage-dependent fashion. Cell Stem Cell 9, 345–356 (2011).
Lampreia, F. P., Carmelo, J. G. & Anjos-Afonso, F. Notch signaling in the regulation of hematopoietic stem cell. Curr. Stem Cell Rep. 3, 202–209 (2017).
Ito, S. et al. Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification. Nature 466, 1129–1133 (2010).
Gjini, E. et al. A zebrafish model of myelodysplastic syndrome produced through tet2 genomic editing. Mol. Cell Biol. 35, 789–804 (2015).
Kunimoto, H. et al. Tet2 disruption leads to enhanced self-renewal and altered differentiation of fetal liver hematopoietic stem cells. Sci. Rep. 2, 273 (2012).
Moran-Crusio, K. et al. Tet2 loss leads to increased hematopoietic stem cell self-renewal and myeloid transformation. Cancer Cell 20, 11–24 (2011).
Li, R. et al. TET2 loss dysregulates the behavior of bone marrow mesenchymal stromal cells and accelerates Tet2(-/-)-driven myeloid malignancy progression. Stem Cell Rep. 10, 166–179 (2018).
Steensma, D. P. Clinical consequences of clonal hematopoiesis of indeterminate potential. Blood Adv. 2, 3404–3410 (2018).
Langemeijer, S. M. et al. Acquired mutations in TET2 are common in myelodysplastic syndromes. Nat. Genet. 41, 838–842 (2009).
Abdel-Wahab, O. et al. Genetic characterization of TET1, TET2, and TET3 alterations in myeloid malignancies. Blood 114, 144–147 (2009).
Tang, F. et al. mRNA-Seq whole-transcriptome analysis of a single cell. Nat. Methods 6, 377–382 (2009).
Rodriques, S. G. et al. Slide-seq: a scalable technology for measuring genome-wide expression at high spatial resolution. Science 363, 1463–1467 (2019).
Lee, J. H. et al. Fluorescent in situ sequencing (FISSEQ) of RNA for gene expression profiling in intact cells and tissues. Nat. Protoc. 10, 442–458 (2015).
Femino, A. M., Fay, F. S., Fogarty, K. & Singer, R. H. Visualization of single RNA transcripts in situ. Science 280, 585–590 (1998).
Chen, K. H., Boettiger, A. N., Moffitt, J. R., Wang, S. & Zhuang, X. RNA imaging. Spatially resolved, highly multiplexed RNA profiling in single cells. Science 348, aaa6090 (2015).
Moffitt, J. R. et al. High-throughput single-cell gene-expression profiling with multiplexed error-robust fluorescence in situ hybridization. Proc. Natl. Acad. Sci. 113, 11046–11051 (2016).
Moffitt, J. R. et al. High-performance multiplexed fluorescence in situ hybridization in culture and tissue with matrix imprinting and clearing. Proc. Natl. Acad. Sci. 113, 14456–14461 (2016).
Han, X. et al. Mapping the mouse cell atlas by microwell-seq. Cell 172, 1091–1107 (2018).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Klei, T. R., Meinderts, S. M., van den Berg, T. K. & van Bruggen, R. From the cradle to the grave: the role of macrophages in erythropoiesis and erythrophagocytosis. Front. Immunol. 8, 73 (2017).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. CellPhoneDB: inferring cell-cell communication from combined expression of multi-subunit ligand-receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
Greenbaum, A. et al. CXCL12 in early mesenchymal progenitors is required for haematopoietic stem-cell maintenance. Nature 495, 227–230 (2013).
Ding, L. & Morrison, S. J. Haematopoietic stem cells and early lymphoid progenitors occupy distinct bone marrow niches. Nature 495, 231–235 (2013).
Omatsu, Y. et al. The essential functions of adipo-osteogenic progenitors as the hematopoietic stem and progenitor cell niche. Immunity 33, 387–399 (2010).
Chou, S. & Lodish, H. F. Fetal liver hepatic progenitors are supportive stromal cells for hematopoietic stem cells. Proc. Natl. Acad. Sci. USA 107, 7799–7804 (2010).
Min, I. M. et al. The transcription factor EGR1 controls both the proliferation and localization of hematopoietic stem cells. Cell Stem Cell 2, 380–391 (2008).
Christodoulou, C. et al. Live-animal imaging of native haematopoietic stem and progenitor cells. Nature 578, 278–283 (2020).
Zeng, Y. et al. Tracing the first hematopoietic stem cell generation in human embryo by single-cell RNA sequencing. Cell Res. 29, 881–894 (2019).
Zhang, Y. et al. PR-domain-containing Mds1-Evi1 is critical for long-term hematopoietic stem cell function. Blood 118, 3853–3861 (2011).
Thoren, L. A. et al. Kit regulates maintenance of quiescent hematopoietic stem cells. J. Immunol. 180, 2045–2053 (2008).
Waskow, C., Paul, S., Haller, C., Gassmann, M. & Rodewald, H. R. Viable c-Kit(W/W) mutants reveal pivotal role for c-kit in the maintenance of lymphopoiesis. Immunity 17, 277–288 (2002).
Mao, B. et al. Kremen proteins are Dickkopf receptors that regulate Wnt/beta-catenin signalling. Nature 417, 664–667 (2002).
Leonard, J. L. et al. The Dkk3 gene encodes a vital intracellular regulator of cell proliferation. PLoS ONE 12, e0181724 (2017).
Ferrari, N. et al. Dickkopf-3 links HSF1 and YAP/TAZ signalling to control aggressive behaviours in cancer-associated fibroblasts. Nat. Commun. 10, 130 (2019).
Stier, S., Cheng, T., Dombkowski, D., Carlesso, N. & Scadden, D. T. Notch1 activation increases hematopoietic stem cell self-renewal in vivo and favors lymphoid over myeloid lineage outcome. Blood 99, 2369–2378 (2002).
Karanu, F. N. et al. The notch ligand jagged-1 represents a novel growth factor of human hematopoietic stem cells. J. Exp. Med. 192, 1365–1372 (2000).
Ding, L., Saunders, T. L., Enikolopov, G. & Morrison, S. J. Endothelial and perivascular cells maintain haematopoietic stem cells. Nature 481, 457–462 (2012).
Itkin, T. et al. FGF-2 expands murine hematopoietic stem and progenitor cells via proliferation of stromal cells, c-Kit activation, and CXCL12 down-regulation. Blood 120, 1843–1855 (2012).
Zhao, J. L. et al. Conversion of danger signals into cytokine signals by hematopoietic stem and progenitor cells for regulation of stress-induced hematopoiesis. Cell Stem Cell 14, 445–459 (2014).
Varnum-Finney, B., Brashem-Stein, C. & Bernstein, I. D. Combined effects of Notch signaling and cytokines induce a multiple log increase in precursors with lymphoid and myeloid reconstituting ability. Blood 101, 1784–1789 (2003).
Xu, C. et al. Stem cell factor is selectively secreted by arterial endothelial cells in bone marrow. Nat. Commun. 9, 2449 (2018).
Ko, M. et al. Ten-eleven-translocation 2 (TET2) negatively regulates homeostasis and differentiation of hematopoietic stem cells in mice. Proc. Natl. Acad. Sci. USA 108, 14566–14571 (2011).
Wang, X. et al. Three-dimensional intact-tissue sequencing of single-cell transcriptional states. Science 361, eaat5691 (2018).
Eng, C.-H. L. et al. Transcriptome-scale super-resolved imaging in tissues by RNA seqFISH+. Nature 568, 235–239 (2019).
Chazotte, B. Labeling membrane glycoproteins or glycolipids with fluorescent wheat germ agglutinin. Cold Spring Harb. Protoc. 2011, pdb prot5623 (2011).
Tikhonova, A. N. et al. The bone marrow microenvironment at single-cell resolution. Nature 569, 222–228 (2019).
Voloshanenko, O., Gmach, P., Winter, J., Kranz, D. & Boutros, M. Mapping of Wnt-Frizzled interactions by multiplex CRISPR targeting of receptor gene families. FASEB J. 31, 4832–4844 (2017).
Luis, T. C. & Staal, F. WNT proteins: environmental factors regulating HSC fate in the niche. Ann. N. Y. Acad. Sci. 1176, 70–76 (2009).
Tahiliani, M. et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 324, 930–935 (2009).
Zhang, H. et al. TET1 is a DNA-binding protein that modulates DNA methylation and gene transcription via hydroxylation of 5-methylcytosine. Cell Res. 20, 1390–1393 (2010).
Solary, E., Bernard, O. A., Tefferi, A., Fuks, F. & Vainchenker, W. The ten-eleven translocation-2 (TET2) gene in hematopoiesis and hematopoietic diseases. Leukemia 28, 485–496 (2014).
Hon, GaryC. et al. 5mC oxidation by Tet2 modulates enhancer activity and timing of transcriptome reprogramming during differentiation. Mol. Cell 56, 286–297 (2014).
Li, Z. et al. Deletion of Tet2 in mice leads to dysregulated hematopoietic stem cells and subsequent development of myeloid malignancies. Blood 118, 4509–4518 (2011).
Quivoron, C. et al. TET2 inactivation results in pleiotropic hematopoietic abnormalities in mouse and is a recurrent event during human lymphomagenesis. Cancer Cell 20, 25–38 (2011).
Schmelzer, E. Hepatic progenitors of the fetal liver: interactions with hematopoietic stem cells. Differentiation 106, 9–14 (2019).
Sugiyama, D., Kulkeaw, K., Mizuochi, C., Horio, Y. & Okayama, S. Hepatoblasts comprise a niche for fetal liver erythropoiesis through cytokine production. Biochem. Biophys. Res. Commun. 410, 301–306 (2011).
Galimi, F. et al. Hepatocyte growth factor induces proliferation and differentiation of multipotent and erythroid hemopoietic progenitors. J. Cell Biol. 127, 1743–1754 (1994).
Wang, G., Moffitt, J. R. & Zhuang, X. Multiplexed imaging of high-density libraries of RNAs with MERFISH and expansion microscopy. Sci. Rep. 8, 4847 (2018).
Gong, H. et al. Simple method to prepare oligonucleotide-conjugated antibodies and its application in multiplex protein detection in single cells. Bioconjug. Chem. 27, 217–225 (2016).
Wang, S., Moffitt, J. R., Dempsey, G. T., Xie, X. S. & Zhuang, X. Characterization and development of photoactivatable fluorescent proteins for single-molecule-based superresolution imaging. Proc. Natl. Acad. Sci. USA 111, 8452–8457 (2014).
Yang, J. et al. Single cell transcriptomics reveals unanticipated features of early hematopoietic precursors. Nucleic Acids Res. 45, 1281–1296 (2017).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 e1821 (2019).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Durinck, S., Spellman, P. T., Birney, E. & Huber, W. Mapping identifiers for the integration of genomic datasets with the R/Bioconductor package biomaRt. Nat. Protoc. 4, 1184–1191 (2009).
Liu, M. et al. Multiplexed imaging of nucleome architectures in single cells of mammalian tissue. Nat. Commun. 11, 2907 (2020).
Shekhar, K. et al. Comprehensive classification of retinal bipolar neurons by single-cell transcriptomics. Cell 166, 1308–1323 (2016).
Acknowledgements
We thank Drs. Diane Krause for helpful discussions and Jun Lu for the Tet2−/− mice. We also thank Dr. Jianming Zeng (University of Macau), and all the members of his bioinformatics team, for generously sharing their experience and codes in scRNA-seq analysis. This work is in part supported by Yale Cooperative Center of Excellence in Hematology (YCCEH) Pilot Project Grant (awarded by the NIDDK under U54DK106857), the National Nature Science Foundation of China (81770173), Guangdong Natural Science Foundation (2018B030308004), and the National Key R&D Program of China (2018YFA0507800). X.P. is supported by Pearl River Talents Program Local Innovative and Research Teams (2017BT01S131). S.G.K. is supported by NHLBI Grant 1R01HL131793. M.L. is supported by the China Scholarship Council (CSC) grant.
Author information
Authors and Affiliations
Contributions
S.W., S.G.K., X.P., and S.M.W. conceived the study. Y.L. and M.L. performed the MERFISH experiments and Y.L. and J.Y. performed the scRNA-seq experiments. Y.L., M.L., and S.W. analyzed data. Y.L., X.P., S.G.K., and S.W. wrote the manuscript with input from M.L., J.Y., and S.M.W.
Corresponding authors
Ethics declarations
Conflict of interest
S.W. is one of the inventors on a patent applied for by Harvard University related to MERFISH. The remaining authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lu, Y., Liu, M., Yang, J. et al. Spatial transcriptome profiling by MERFISH reveals fetal liver hematopoietic stem cell niche architecture. Cell Discov 7, 47 (2021). https://doi.org/10.1038/s41421-021-00266-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41421-021-00266-1
This article is cited by
-
scGIST: gene panel design for spatial transcriptomics with prioritized gene sets
Genome Biology (2024)
-
High resolution spatial profiling of kidney injury and repair using RNA hybridization-based in situ sequencing
Nature Communications (2024)
-
PROST: quantitative identification of spatially variable genes and domain detection in spatial transcriptomics
Nature Communications (2024)
-
Spatially resolved analysis of pancreatic cancer identifies therapy-associated remodeling of the tumor microenvironment
Nature Genetics (2024)
-
Fetal liver development and implications for liver disease pathogenesis
Nature Reviews Gastroenterology & Hepatology (2023)