Abstract
Human coronaviruses (HCoVs), including severe acute respiratory syndrome coronavirus (SARS-CoV) and 2019 novel coronavirus (2019-nCoV, also known as SARS-CoV-2), lead global epidemics with high morbidity and mortality. However, there are currently no effective drugs targeting 2019-nCoV/SARS-CoV-2. Drug repurposing, representing as an effective drug discovery strategy from existing drugs, could shorten the time and reduce the cost compared to de novo drug discovery. In this study, we present an integrative, antiviral drug repurposing methodology implementing a systems pharmacology-based network medicine platform, quantifying the interplay between the HCoV–host interactome and drug targets in the human protein–protein interaction network. Phylogenetic analyses of 15 HCoV whole genomes reveal that 2019-nCoV/SARS-CoV-2 shares the highest nucleotide sequence identity with SARS-CoV (79.7%). Specifically, the envelope and nucleocapsid proteins of 2019-nCoV/SARS-CoV-2 are two evolutionarily conserved regions, having the sequence identities of 96% and 89.6%, respectively, compared to SARS-CoV. Using network proximity analyses of drug targets and HCoV–host interactions in the human interactome, we prioritize 16 potential anti-HCoV repurposable drugs (e.g., melatonin, mercaptopurine, and sirolimus) that are further validated by enrichment analyses of drug-gene signatures and HCoV-induced transcriptomics data in human cell lines. We further identify three potential drug combinations (e.g., sirolimus plus dactinomycin, mercaptopurine plus melatonin, and toremifene plus emodin) captured by the “Complementary Exposure” pattern: the targets of the drugs both hit the HCoV–host subnetwork, but target separate neighborhoods in the human interactome network. In summary, this study offers powerful network-based methodologies for rapid identification of candidate repurposable drugs and potential drug combinations targeting 2019-nCoV/SARS-CoV-2.
Similar content being viewed by others
Introduction
Coronaviruses (CoVs) typically affect the respiratory tract of mammals, including humans, and lead to mild to severe respiratory tract infections1. In the past two decades, two highly pathogenic human CoVs (HCoVs), including severe acute respiratory syndrome coronavirus (SARS-CoV) and Middle East respiratory syndrome coronavirus (MERS-CoV), emerging from animal reservoirs, have led to global epidemics with high morbidity and mortality2. For example, 8098 individuals were infected and 774 died in the SARS-CoV pandemic, which cost the global economy with an estimated $30 to $100 billion3,4. According to the World Health Organization (WHO), as of November 2019, MERS-CoV has had a total of 2494 diagnosed cases causing 858 deaths, the majority in Saudi Arabia2. In December 2019, the third pathogenic HCoV, named 2019 novel coronavirus (2019-nCoV/SARS-CoV-2), as the cause of coronavirus disease 2019 (abbreviated as COVID-19)5, was found in Wuhan, China. As of 24 February 2020, there have been over 79,000 cases with over 2600 deaths for the 2019-nCoV/SARS-CoV-2 outbreak worldwide; furthermore, human-to-human transmission has occurred among close contacts6. However, there are currently no effective medications against 2019-nCoV/SARS-CoV-2. Several national and international research groups are working on the development of vaccines to prevent and treat the 2019-nCoV/SARS-CoV-2, but effective vaccines are not available yet. There is an urgent need for the development of effective prevention and treatment strategies for 2019-nCoV/SARS-CoV-2 outbreak.
Although investment in biomedical and pharmaceutical research and development has increased significantly over the past two decades, the annual number of new treatments approved by the U.S. Food and Drug Administration (FDA) has remained relatively constant and limited7. A recent study estimated that pharmaceutical companies spent $2.6 billion in 2015, up from $802 million in 2003, in the development of an FDA-approved new chemical entity drug8. Drug repurposing, represented as an effective drug discovery strategy from existing drugs, could significantly shorten the time and reduce the cost compared to de novo drug discovery and randomized clinical trials9,10,11. However, experimental approaches for drug repurposing is costly and time-consuming12. Computational approaches offer novel testable hypotheses for systematic drug repositioning9,10,11,13,14. However, traditional structure-based methods are limited when three-dimensional (3D) structures of proteins are unavailable, which, unfortunately, is the case for the majority of human and viral targets. In addition, targeting single virus proteins often has high risk of drug resistance by the rapid evolution of virus genomes1.
Viruses (including HCoV) require host cellular factors for successful replication during infection1. Systematic identification of virus–host protein–protein interactions (PPIs) offers an effective way toward elucidating the mechanisms of viral infection15,16. Subsequently, targeting cellular antiviral targets, such as virus–host interactome, may offer a novel strategy for the development of effective treatments for viral infections1, including SARS-CoV17, MERS-CoV17, Ebola virus18, and Zika virus14,19,20,21. We recently presented an integrated antiviral drug discovery pipeline that incorporated gene-trap insertional mutagenesis, known functional drug–gene network, and bioinformatics analyses14. This methodology allows to identify several candidate repurposable drugs for Ebola virus11,14. Our work over the last decade has demonstrated how network strategies can, for example, be used to identify effective repurposable drugs13,22,23,24,25,26,27 and drug combinations28 for multiple human diseases. For example, network-based drug–disease proximity sheds light on the relationship between drugs (e.g., drug targets) and disease modules (molecular determinants in disease pathobiology modules within the PPIs), and can serve as a useful tool for efficient screening of potentially new indications for approved drugs, as well as drug combinations, as demonstrated in our recent studies13,23,27,28.
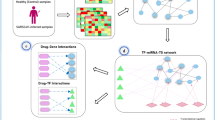
In this study, we present an integrative antiviral drug repurposing methodology, which combines a systems pharmacology-based network medicine platform that quantifies the interplay between the virus–host interactome and drug targets in the human PPI network. The basis for these experiments rests on the notions that (i) the proteins that functionally associate with viral infection (including HCoV) are localized in the corresponding subnetwork within the comprehensive human PPI network and (ii) proteins that serve as drug targets for a specific disease may also be suitable drug targets for potential antiviral infection owing to common PPIs and functional pathways elucidated by the human interactome (Fig. 1). We follow this analysis with bioinformatics validation of drug-induced gene signatures and HCoV-induced transcriptomics in human cell lines to inspect the postulated mechanism-of-action in a specific HCoV for which we propose repurposing (Fig. 1).
Our network-based methodology combines a systems pharmacology-based network medicine platform that quantifies the interplay between the virus–host interactome and drug targets in the human PPI network. a Human coronavirus (HCoV)-associated host proteins were collected from literatures and pooled to generate a pan-HCoV protein subnetwork. b Network proximity between drug targets and HCoV-associated proteins was calculated to screen for candidate repurposable drugs for HCoVs under the human protein interactome model. c, d Gene set enrichment analysis was utilized to validate the network-based prediction. e Top candidates were further prioritized for drug combinations using network-based method captured by the “Complementary Exposure” pattern: the targets of the drugs both hit the HCoV–host subnetwork, but target separate neighborhoods in the human interactome network. f Overall hypothesis of the network-based methodology: (i) the proteins that functionally associate with HCoVs are localized in the corresponding subnetwork within the comprehensive human interactome network; and (ii) proteins that serve as drug targets for a specific disease may also be suitable drug targets for potential antiviral infection owing to common protein–protein interactions elucidated by the human interactome.
Results
Phylogenetic analyses of 2019-nCoV/SARS-CoV-2
To date, seven pathogenic HCoVs (Fig. 2a, b) have been found:1,29 (i) 2019-nCoV/SARS-CoV-2, SARS-CoV, MERS-CoV, HCoV-OC43, and HCoV-HKU1 are β genera, and (ii) HCoV-NL63 and HCoV-229E are α genera. We performed the phylogenetic analyses using the whole-genome sequence data from 15 HCoVs to inspect the evolutionary relationship of 2019-nCoV/SARS-CoV-2 with other HCoVs. We found that the whole genomes of 2019-nCoV/SARS-CoV-2 had ~99.99% nucleotide sequence identity across three diagnosed patients (Supplementary Table S1). The 2019-nCoV/SARS-CoV-2 shares the highest nucleotide sequence identity (79.7%) with SARS-CoV among the six other known pathogenic HCoVs, revealing conserved evolutionary relationship between 2019-nCoV/SARS-CoV-2 and SARS-CoV (Fig. 2a).
a Phylogenetic tree of coronavirus (CoV). Phylogenetic algorithm analyzed evolutionary conservation among whole genomes of 15 coronaviruses. Red color highlights the recent emergent coronavirus, 2019-nCoV/SARS-CoV-2. Numbers on the branches indicate bootstrap support values. The scale shows the evolutionary distance computed using the p-distance method. b Schematic plot for HCoV genomes. The genus and host information of viruses was labeled on the left by different colors. Empty dark gray boxes represent accessory open reading frames (ORFs). c–e The 3D structures of SARS-CoV nsp12 (PDB ID: 6NUR) (c), spike (PDB ID: 6ACK) (d), and nucleocapsid (PDB ID: 2CJR) (e) shown were based on homology modeling. Genome information and phylogenetic analysis results are provided in Supplementary Tables S1 and S2.
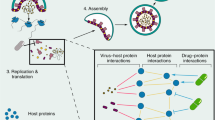
HCoVs have five major protein regions for virus structure assembly and viral replications29, including replicase complex (ORF1ab), spike (S), envelope (E), membrane (M), and nucleocapsid (N) proteins (Fig. 2b). The ORF1ab gene encodes the non-structural proteins (nsp) of viral RNA synthesis complex through proteolytic processing30. The nsp12 is a viral RNA-dependent RNA polymerase, together with co-factors nsp7 and nsp8 possessing high polymerase activity. From the protein 3D structure view of SARS-CoV nsp12, it contains a larger N-terminal extension (which binds to nsp7 and nsp8) and polymerase domain (Fig. 2c). The spike is a transmembrane glycoprotein that plays a pivotal role in mediating viral infection through binding the host receptor31,32. Figure 2d shows the 3D structure of the spike protein bound with the host receptor angiotensin converting enznyme2 (ACE2) in SARS-CoV (PDB ID: 6ACK). A recent study showed that 2019-nCoV/SARS-CoV-2 is able to utilize ACE2 as an entry receptor in ACE2-expressing cells33, suggesting potential drug targets for therapeutic development. Furthermore, cryo-EM structure of the spike and biophysical assays reveal that the 2019-nCoV/SARS-CoV-2 spike binds ACE2 with higher affinity than SARS-CoV34. In addition, the nucleocapsid is also an important subunit for packaging the viral genome through protein oligomerization35, and the single nucleocapsid structure is shown in Fig. 2e.
Protein sequence alignment analyses indicated that the 2019-nCoV/SARS-CoV-2 was most evolutionarily conserved with SARS-CoV (Supplementary Table S2). Specifically, the envelope and nucleocapsid proteins of 2019-nCoV/SARS-CoV-2 are two evolutionarily conserved regions, with sequence identities of 96% and 89.6%, respectively, compared to SARS-CoV (Supplementary Table S2). However, the spike protein exhibited the lowest sequence conservation (sequence identity of 77%) between 2019-nCoV/SARS-CoV-2 and SARS-CoV. Meanwhile, the spike protein of 2019-nCoV/SARS-CoV-2 only has 31.9% sequence identity compared to MERS-CoV.
HCoV–host interactome network
To depict the HCoV–host interactome network, we assembled the CoV-associated host proteins from four known HCoVs (SARS-CoV, MERS-CoV, HCoV-229E, and HCoV-NL63), one mouse MHV, and one avian IBV (N protein) (Supplementary Table S3). In total, we obtained 119 host proteins associated with CoVs with various experimental evidence. Specifically, these host proteins are either the direct targets of HCoV proteins or are involved in crucial pathways of HCoV infection. The HCoV–host interactome network is shown in Fig. 3a. We identified several hub proteins including JUN, XPO1, NPM1, and HNRNPA1, with the highest number of connections within the 119 proteins. KEGG pathway enrichment analysis revealed multiple significant biological pathways (adjusted P value < 0.05), including measles, RNA transport, NF-kappa B signaling, Epstein-Barr virus infection, and influenza (Fig. 3b). Gene ontology (GO) biological process enrichment analysis further confirmed multiple viral infection-related processes (adjusted P value < 0.001), including viral life cycle, modulation by virus of host morphology or physiology, viral process, positive regulation of viral life cycle, transport of virus, and virion attachment to host cell (Fig. 3c). We then mapped the known drug–target network (see Materials and methods) into the HCoV–host interactome to search for druggable, cellular targets. We found that 47 human proteins (39%, blue nodes in Fig. 3a) can be targeted by at least one approved drug or experimental drug under clinical trials. For example, GSK3B, DPP4, SMAD3, PARP1, and IKBKB are the most targetable proteins. The high druggability of HCoV–host interactome motivates us to develop a drug repurposing strategy by specifically targeting cellular proteins associated with HCoVs for potential treatment of 2019-nCoV/SARS-CoV-2.
a A subnetwork highlighting the HCoV–host interactome. Nodes represent three types of HCoV-associated host proteins: targetgable (proteins can be targeted by approved drugs or drugs under clinical trials), non-targetable (proteins do not have any known ligands), neighbors (protein–protein interaction partners). Edge colors indicate five types of experimental evidence of the protein–protein interactions (see Materials and methods). 3D three-dimensional structure. b, c KEGG human pathway (b) and gene ontology enrichment analyses (c) for the HCoV-associated proteins.
Network-based drug repurposing for HCoVs
The basis for the proposed network-based drug repurposing methodologies rests on the notions that the proteins that associate with and functionally govern viral infection are localized in the corresponding subnetwork (Fig. 1a) within the comprehensive human interactome network. For a drug with multiple targets to be effective against an HCoV, its target proteins should be within or in the immediate vicinity of the corresponding subnetwork in the human protein–protein interactome (Fig. 1), as we demonstrated in multiple diseases13,22,23,28 using this network-based strategy. We used a state-of-the-art network proximity measure to quantify the relationship between HCoV-specific subnetwork (Fig. 3a) and drug targets in the human interactome. We constructed a drug–target network by assembling target information for more than 2000 FDA-approved or experimental drugs (see Materials and methods). To improve the quality and completeness of the human protein interactome network, we integrated PPIs with five types of experimental data: (1) binary PPIs from 3D protein structures; (2) binary PPIs from unbiased high-throughput yeast-two-hybrid assays; (3) experimentally identified kinase-substrate interactions; (4) signaling networks derived from experimental data; and (5) literature-derived PPIs with various experimental evidence (see Materials and methods). We used a Z-score (Z) measure and permutation test to reduce the study bias in network proximity analyses (including hub nodes in the human interactome network by literature-derived PPI data bias) as described in our recent studies13,28.
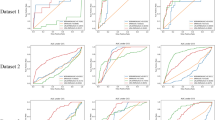
In total, we computationally identified 135 drugs that were associated (Z < −1.5 and P < 0.05, permutation test) with the HCoV–host interactome (Fig. 4a, Supplementary Tables S4 and 5). To validate bias of the pooled cellular proteins from six CoVs, we further calculated the network proximities of all the drugs for four CoVs with a large number of know host proteins, including SARS-CoV, MERS-CoV, IBV, and MHV, separately. We found that the Z-scores showed consistency among the pooled 119 HCoV-associated proteins and other four individual CoVs (Fig. 4b). The Pearson correlation coefficients of the proximities of all the drugs for the pooled HCoV are 0.926 vs. SARS-CoV (P < 0.001, t distribution), 0.503 vs. MERS-CoV (P < 0.001), 0.694 vs. IBV (P < 0.001), and 0.829 vs. MHV (P < 0.001). These network proximity analyses offer putative repurposable candidates for potential prevention and treatment of HCoVs.
a A subnetwork highlighting network-predicted drug-HCoV associations connecting 135 drugs and HCoVs. From the 2938 drugs evaluated, 135 ones achieved significant proximities between drug targets and the HCoV-associated proteins in the human interactome network. Drugs are colored by their first-level of the Anatomical Therapeutic Chemical (ATC) classification system code. b A heatmap highlighting network proximity values for SARS-CoV, MERS-CoV, IBV, and MHV, respectively. Color key denotes network proximity (Z-score) between drug targets and the HCoV-associated proteins in the human interactome network. P value was computed by permutation test.
Discovery of repurposable drugs for HCoV
To further validate the 135 repurposable drugs against HCoVs, we first performed gene set enrichment analysis (GSEA) using transcriptome data of MERS-CoV and SARS-CoV infected host cells (see Methods). These transcriptome data were used as gene signatures for HCoVs. Additionally, we downloaded the gene expression data of drug-treated human cell lines from the Connectivity Map (CMAP) database36 to obtain drug–gene signatures. We calculated a GSEA score (see Methods) for each drug and used this score as an indication of bioinformatics validation of the 135 drugs. Specifically, an enrichment score (ES) was calculated for each HCoV data set, and ES > 0 and P < 0.05 (permutation test) was used as cut-off for a significant association of gene signatures between a drug and a specific HCoV data set. The GSEA score, ranging from 0 to 3, is the number of data sets that met these criteria for a specific drug. Mesalazine (an approved drug for inflammatory bowel disease), sirolimus (an approved immunosuppressive drug), and equilin (an approved agonist of the estrogen receptor for menopausal symptoms) achieved the highest GSEA scores of 3, followed by paroxetine and melatonin with GSEA scores of 2. We next selected 16 high-confidence repurposable drugs (Fig. 5a and Table 1) against HCoVs using subject matter expertise based on a combination of factors: (i) strength of the network-predicted associations (a smaller network proximity score in Supplementary Table S4); (ii) validation by GSEA analyses; (iii) literature-reported antiviral evidence, and (iv) fewer clinically reported side effects. Specifically, we showcased several selected repurposable drugs with literature-reported antiviral evidence as below.
a Network-predicted evidence and gene set enrichment analysis (GSEA) scores for 16 potential repurposable drugs for HCoVs. The overall connectivity of the top drug candidates to the HCoV-associated proteins was examined. Most of these drugs indirectly target HCoV-associated proteins via the human protein–protein interaction networks. All the drug–target-HCoV-associated protein connections were examined, and those proteins with at least five connections are shown. The box heights for the proteins indicate the number of connections. GSEA scores for eight drugs were not available (NA) due to the lack of transcriptome profiles for the drugs. b–e Inferred mechanism-of-action networks for four selected drugs: b toremifene (first-generation nonsteroidal-selective estrogen receptor modulator), c irbesartan (an angiotensin receptor blocker), d mercaptopurine (an antimetabolite antineoplastic agent with immunosuppressant properties), and e melatonin (a biogenic amine for treating circadian rhythm sleep disorders).
Selective estrogen receptor modulators
An overexpression of estrogen receptor has been shown to play a crucial role in inhibiting viral replication37. Selective estrogen receptor modulators (SERMs) have been reported to play a broader role in inhibiting viral replication through the non-classical pathways associated with estrogen receptor37. SERMs interfere at the post viral entry step and affect the triggering of fusion, as the SERMs’ antiviral activity still can be observed in the absence of detectable estrogen receptor expression18. Toremifene (Z = –3.23, Fig. 5a), the first generation of nonsteroidal SERM, exhibits potential effects in blocking various viral infections, including MERS-CoV, SARS-CoV, and Ebola virus in established cell lines17,38. Compared to the classical ESR1-related antiviral pathway, toremifene prevents fusion between the viral and endosomal membrane by interacting with and destabilizing the virus membrane glycoprotein, and eventually inhibiting viral replication39. As shown in Fig. 5b, toremifene potentially affects several key host proteins associated with HCoV, such as RPL19, HNRNPA1, NPM1, EIF3I, EIF3F, and EIF3E40,41. Equilin (Z = –2.52 and GSEA score = 3), an estrogenic steroid produced by horses, also has been proven to have moderate activity in inhibiting the entry of Zaire Ebola virus glycoprotein and human immunodeficiency virus (ZEBOV-GP/HIV)18. Altogether, network-predicted SERMs (such as toremifene and equilin) offer candidate repurposable drugs for 2019-nCoV/SARS-CoV-2.
Angiotensin receptor blockers
Angiotensin receptor blockers (ARBs) have been reported to associate with viral infection, including HCoVs42,43,44. Irbesartan (Z = –5.98), a typical ARB, was approved by the FDA for treatment of hypertension and diabetic nephropathy. Here, network proximity analysis shows a significant association between irbesartan’s targets and HCoV-associated host proteins in the human interactome. As shown in Fig. 5c, irbesartan targets SLC10A1, encoding the sodium/bile acid cotransporter (NTCP) protein that has been identified as a functional preS1-specific receptor for the hepatitis B virus (HBV) and the hepatitis delta virus (HDV). Irbesartan can inhibit NTCP, thus inhibiting viral entry45,46. SLC10A1 interacts with C11orf74, a potential transcriptional repressor that interacts with nsp-10 of SARS-CoV47. There are several other ARBs (such as eletriptan, frovatriptan, and zolmitriptan) in which their targets are potentially associated with HCoV-associated host proteins in the human interactome.
Immunosuppressant or antineoplastic agents
Previous studies have confirmed the mammalian target of rapamycin complex 1 (mTORC1) as the key factor in regulating various viruses’ replications, including Andes orthohantavirus and coronavirus48,49. Sirolimus (Z = –2.35 and GSEA score = 3), an inhibitor of mammalian target of rapamycin (mTOR), was reported to effectively block viral protein expression and virion release effectively50. Indeed, the latest study revealed the clinical application: sirolimus reduced MERS-CoV infection by over 60%51. Moreover, sirolimus usage in managing patients with severe H1N1 pneumonia and acute respiratory failure can improve those patients’ prognosis significantly50. Mercaptopurine (Z = –2.44 and GSEA score = 1), an antineoplastic agent with immunosuppressant property, has been used to treat cancer since the 1950s and expanded its application to several auto-immune diseases, including rheumatoid arthritis, systemic lupus erythematosus, and Crohn’s disease52. Mercaptopurine has been reported as a selective inhibitor of both SARS-CoV and MERS-CoV by targeting papain-like protease which plays key roles in viral maturation and antagonism to interferon stimulation53,54. Mechanistically, mercaptopurine potentially target several host proteins in HCoVs, such as JUN, PABPC1, NPM1, and NCL40,55 (Fig. 5d).
Anti-inflammatory agents
Inflammatory pathways play essential roles in viral infections56,57. As a biogenic amine, melatonin (N-acetyl-5-methoxytryptamine) (Z = –1.72 and GSEA score = 2) plays a key role in various biological processes, and offers a potential strategy in the management of viral infections58,59. Viral infections are often associated with immune-inflammatory injury, in which the level of oxidative stress increases significantly and leaves negative effects on the function of multiple organs60. The antioxidant effect of melatonin makes it a putative candidate drug to relieve patients’ clinical symptoms in antiviral treatment, even though melatonin cannot eradicate or even curb the viral replication or transcription61,62. In addition, the application of melatonin may prolong patients’ survival time, which may provide a chance for patients’ immune systems to recover and eventually eradicate the virus. As shown in Fig. 5e, melatonin indirectly targets several HCoV cellular targets, including ACE2, BCL2L1, JUN, and IKBKB. Eplerenone (Z = –1.59), an aldosterone receptor antagonist, is reported to have a similar anti-inflammatory effect as melatonin. By inhibiting mast-cell-derived proteinases and suppressing fibrosis, eplerenone can improve survival of mice infected with encephalomyocarditis virus63.
In summary, our network proximity analyses offer multiple candidate repurposable drugs that target diverse cellular pathways for potential prevention and treatment of 2019-nCoV/SARS-CoV-2. However, further preclinical experiments64 and clinical trials are required to verify the clinical benefits of these network-predicted candidates before clinical use.
Network-based identification of potential drug combinations for 2019-nCoV/SARS-CoV-2
Drug combinations, offering increased therapeutic efficacy and reduced toxicity, play an important role in treating various viral infections65. However, our ability to identify and validate effective combinations is limited by a combinatorial explosion, driven by both the large number of drug pairs and dosage combinations. In our recent study, we proposed a novel network-based methodology to identify clinically efficacious drug combinations28. Relying on approved drug combinations for hypertension and cancer, we found that a drug combination was therapeutically effective only if it was captured by the “Complementary Exposure” pattern: the targets of the drugs both hit the disease module, but target separate neighborhoods (Fig. 6a). Here we sought to identify drug combinations that may provide a synergistic effect in potentially treating 2019-nCoV/SARS-CoV-2 with well-defined mechanism-of-action by network analysis. For the 16 potential repurposable drugs (Fig. 5a, Table 1), we showcased three network-predicted candidate drug combinations for 2019-nCoV/SARS-CoV-2. All predicted possible combinations can be found in Supplementary Table S6.
a The possible exposure mode of the HCoV-associated protein module to the pairwise drug combinations. An effective drug combination will be captured by the “Complementary Exposure” pattern: the targets of the drugs both hit the HCoV–host subnetwork, but target separate neighborhoods in the human interactome network. ZCA and ZCB denote the network proximity (Z-score) between targets (Drugs A and B) and a specific HCoV. SAB denotes separation score (see Materials and methods) of targets between Drug A and Drug B. b–d Inferred mechanism-of-action networks for three selected pairwise drug combinations: b sirolimus (a potent immunosuppressant with both antifungal and antineoplastic properties) plus dactinomycin (an RNA synthesis inhibitor for treatment of various tumors), c toremifene (first-generation nonsteroidal-selective estrogen receptor modulator) plus emodin (an experimental drug for the treatment of polycystic kidney), and d melatonin (a biogenic amine for treating circadian rhythm sleep disorders) plus mercaptopurine (an antimetabolite antineoplastic agent with immunosuppressant properties).
Sirolimus plus Dactinomycin
Sirolimus, an inhibitor of mTOR with both antifungal and antineoplastic properties, has demonstrated to improve outcomes in patients with severe H1N1 pneumonia and acute respiratory failure50. The mTOR signaling plays an essential role for MERS-CoV infection66. Dactinomycin, also known actinomycin D, is an approved RNA synthesis inhibitor for treatment of various cancer types. An early study showed that dactinomycin (1 μg/ml) inhibited the growth of feline enteric CoV67. As shown in Fig. 6b, our network analysis shows that sirolimus and dactinomycin synergistically target HCoV-associated host protein subnetwork by “Complementary Exposure” pattern, offering potential combination regimens for treatment of HCoV. Specifically, sirolimus and dactinomycin may inhibit both mTOR signaling and RNA synthesis pathway (including DNA topoisomerase 2-alpha (TOP2A) and DNA topoisomerase 2-beta (TOP2B)) in HCoV-infected cells (Fig. 6b).
Toremifene plus Emodin
Toremifene is among the approved first-generation nonsteroidal SERMs for the treatment of metastatic breast cancer68. SERMs (including toremifene) inhibited Ebola virus infection18 by interacting with and destabilizing the Ebola virus glycoprotein39. In vitro assays have demonstrated that toremifene inhibited growth of MERS-CoV17,69 and SARA-CoV38 (Table 1). Emodin, an anthraquinone derivative extracted from the roots of rheum tanguticum, has been reported to have various anti-virus effects. Specifically, emdoin inhibited SARS-CoV-associated 3a protein70, and blocked an interaction between the SARS-CoV spike protein and ACE2 (ref. 71). Altogether, network analyses and published experimental data suggested that combining toremifene and emdoin offered a potential therapeutic approach for 2019-nCoV/SARS-CoV-2 (Fig. 6c).
Mercaptopurine plus Melatonin
As shown in Fig. 5a, targets of both mercaptopurine and melatonin showed strong network proximity with HCoV-associated host proteins in the human interactome network. Recent in vitro and in vivo studies identified mercaptopurine as a selective inhibitor of both SARS-CoV and MERS-CoV by targeting papain-like protease53,54. Melatonin was reported in potential antiviral infection via its anti-inflammatory and antioxidant effects58,59,60,61,62. Melatonin indirectly regulates ACE2 expression, a key entry receptor involved in viral infection of HCoVs, including 2019-nCoV/SARS-CoV-2 (ref. 33). Specifically, melatonin was reported to inhibit calmodulin and calmodulin interacts with ACE2 by inhibiting shedding of its ectodomain, a key infectious process of SARS-CoV72,73. JUN, also known as c-Jun, is a key host protein involving in HCoV infectious bronchitis virus74. As shown in Fig. 6d, mercaptopurine and melatonin may synergistically block c-Jun signaling by targeting multiple cellular targets. In summary, combination of mercaptopurine and melatonin may offer a potential combination therapy for 2019-nCoV/SARS-CoV-2 by synergistically targeting papain-like protease, ACE2, c-Jun signaling, and anti-inflammatory pathways (Fig. 6d). However, further experimental observations on ACE2 pathways by melatonin in 2019-nCoV/SARS-CoV-2 are highly warranted.
Discussion
In this study, we presented a network-based methodology for systematic identification of putative repurposable drugs and drug combinations for potential treatment of 2019-nCoV/SARS-CoV-2. Integration of drug–target networks, HCoV–host interactions, HCoV-induced transcriptome in human cell lines, and human protein–protein interactome network are essential for such identification. Based on comprehensive evaluation, we prioritized 16 candidate repurposable drugs (Fig. 5) and 3 potential drug combinations (Fig. 6) for targeting 2019-nCoV/SARS-CoV-2. However, although the majority of predictions have been validated by various literature data (Table 1), all network-predicted repurposable drugs and drug combinations must be validated in various 2019-nCoV/SARS-CoV-2 experimental assays64 and randomized clinical trials before being used in patients.
We acknowledge several limitations in the current study. Although 2019-nCoV/SARS-CoV-2 shared high nucleotide sequence identity with other HCoVs (Fig. 2), our predictions are not 2019-nCoV/SARS-CoV-2 specific by lack of the known host proteins on 2019-nCoV/SARS-CoV-2. We used a low binding affinity value of 10 μM as a threshold to define a physical drug–target interaction. However, a stronger binding affinity threshold (e.g., 1 μM) may be a more suitable cut-off in drug discovery, although it will generate a smaller drug–target network. Although sizeable efforts were made for assembling large scale, experimentally reported drug–target networks from publicly available databases, the network data may be incomplete and some drug–target interactions may be functional associations, instead of physical bindings. For example, Silvestrol, a natural product from the flavagline, was found to have antiviral activity against Ebola75 and Coronaviruses76. After adding its target, an RNA helicase enzyme EIF4A76, silvestrol was predicted to be significantly associated with HCoVs (Z = –1.24, P = 0.041) by network proximity analysis. To increase coverage of drug–target networks, we may use computational approaches to systematically predict the drug-target interactions further25,26. In addition, the collected virus–host interactions are far from completeness and the quality can be influenced by multiple factors, including different experimental assays and human cell line models. We may computationally predict a new virus–host interactome for 2019-nCoV/SARS-CoV-2 using sequence-based and structure-based approaches77. Drug targets representing nodes within cellular networks are often intrinsically coupled with both therapeutic and adverse profiles78, as drugs can inhibit or activate protein functions (including antagonists vs. agonists). The current systems pharmacology model cannot separate therapeutic (antiviral) effects from those predictions due to lack of detailed pharmacological effects of drug targets and unknown functional consequences of virus–host interactions. Comprehensive identification of the virus–host interactome for 2019-nCoV/SARS-CoV-2, with specific biological effects using functional genomics assays79,80, will significantly improve the accuracy of the proposed network-based methodologies further.
Owing to a lack of the complete drug-target information (such as the molecular “promiscuity” of drugs), the dose–response and dose–toxicity effects for both repurposable drugs and drug combinations cannot be identified in the current network models. For example, Mesalazine, an approved drug for inflammatory bowel disease, is a top network-predicted repurposable drug associated with HCoVs (Fig. 5a). Yet, several clinical studies showed the potential pulmonary toxicities (including pneumonia) associated with mesalazine usage81,82. Integration of lung-specific gene expression23 of 2019-nCoV/SARS-CoV-2 host proteins and physiologically based pharmacokinetic modeling83 may reduce side effects of repurposable drugs or drug combinations. Preclinical studies are warranted to evaluate in vivo efficiency and side effects before clinical trials. Furthermore, we only limited to predict pairwise drug combinations based on our previous network-based framework28. However, we expect that our methodology remain to be a useful network-based tool for prediction of combining multiple drugs toward exploring network relationships of multiple drugs’ targets with the HCoV–host subnetwork in the human interactome. Finally, we aimed to systematically identify repurposable drugs by specifically targeting nCoV host proteins only. Thus, our current network models cannot predict repurposable drugs from the existing anti-virus drugs that target virus proteins only. Thus, combination of the existing anti-virus drugs (such as remdesivir64) with the network-predicted repurposable drugs (Fig. 5) or drug combinations (Fig. 6) may improve coverage of current network-based methodologies by utilizing multi-layer network framework16.
In conclusion, this study offers a powerful, integrative network-based systems pharmacology methodology for rapid identification of repurposable drugs and drug combinations for the potential treatment of 2019-nCoV/SARS-CoV-2. Our approach can minimize the translational gap between preclinical testing results and clinical outcomes, which is a significant problem in the rapid development of efficient treatment strategies for the emerging 2019-nCoV/SARS-CoV-2 outbreak. From a translational perspective, if broadly applied, the network tools developed here could help develop effective treatment strategies for other emerging viral infections and other human complex diseases as well.
Methods and materials
Genome information and phylogenetic analysis
In total, we collected DNA sequences and protein sequences for 15 HCoVs, including three most recent 2019-nCoV/SARS-CoV-2 genomes, from the NCBI GenBank database (28 January 2020, Supplementary Table S1). Whole-genome alignment and protein sequence identity calculation were performed by Multiple Sequence Alignment in EMBL-EBI database (https://www.ebi.ac.uk/) with default parameters. The neighbor joining (NJ) tree was computed from the pairwise phylogenetic distance matrix using MEGA X84 with 1000 bootstrap replicates. The protein alignment and phylogenetic tree of HCoVs were constructed by MEGA X84.
Building the virus–host interactome
We collected HCoV–host protein interactions from various literatures based on our sizeable efforts. The HCoV-associated host proteins of several HCoVs, including SARS-CoV, MERS-CoV, IBV, MHV, HCoV-229E, and HCoV-NL63 were pooled. These proteins were either the direct targets of HCoV proteins or were involved in critical pathways of HCoV infection identified by multiple experimental sources, including high-throughput yeast-two-hybrid (Y2H) systems, viral protein pull-down assay, in vitro co-immunoprecipitation and RNA knock down experiment. In total, the virus–host interaction network included 6 HCoVs with 119 host proteins (Supplementary Table S3).
Functional enrichment analysis
Next, we performed Kyoto Encyclopedia of Genes and Genomes (KEGG) and Gene Ontology (GO) enrichment analyses to evaluate the biological relevance and functional pathways of the HCoV-associated proteins. All functional analyses were performed using Enrichr85.
Building the drug–target network
Here, we collected drug–target interaction information from the DrugBank database (v4.3)86, Therapeutic Target Database (TTD)87, PharmGKB database, ChEMBL (v20)88, BindingDB89, and IUPHAR/BPS Guide to PHARMACOLOGY90. The chemical structure of each drug with SMILES format was extracted from DrugBank86. Here, drug–target interactions meeting the following three criteria were used: (i) binding affinities, including Ki, Kd, IC50, or EC50 each ≤10 μM; (ii) the target was marked as “reviewed” in the UniProt database91; and (iii) the human target was represented by a unique UniProt accession number. The details for building the experimentally validated drug–target network are provided in our recent studies13,23,28.
Building the human protein–protein interactome
To build a comprehensive list of human PPIs, we assembled data from a total of 18 bioinformatics and systems biology databases with five types of experimental evidence: (i) binary PPIs tested by high-throughput yeast-two-hybrid (Y2H) systems; (ii) binary, physical PPIs from protein 3D structures; (iii) kinase-substrate interactions by literature-derived low-throughput or high-throughput experiments; (iv) signaling network by literature-derived low-throughput experiments; and (v) literature-curated PPIs identified by affinity purification followed by mass spectrometry (AP-MS), Y2H, or by literature-derived low-throughput experiments. All inferred data, including evolutionary analysis, gene expression data, and metabolic associations, were excluded. The genes were mapped to their Entrez ID based on the NCBI database92 as well as their official gene symbols based on GeneCards (https://www.genecards.org/). In total, the resulting human protein–protein interactome used in this study includes 351,444 unique PPIs (edges or links) connecting 17,706 proteins (nodes), representing a 50% increase in the number of the PPIs we have used previously. Detailed descriptions for building the human protein–protein interactome are provided in our previous studies13,23,28,93.
Network proximity measure
We posit that the human PPIs provide an unbiased, rational roadmap for repurposing drugs for potential treatment of HCoVs in which they were not originally approved. Given C, the set of host genes associated with a specific HCoV, and T, the set of drug targets, we computed the network proximity of C with the target set T of each drug using the “closest” method:
where d(c, t) is the shortest distance between gene c and t in the human protein interactome. The network proximity was converted to Z-score based on permutation tests:
where \(\overline {d_r}\) and σr were the mean and standard deviation of the permutation test repeated 1000 times, each time with two randomly selected gene lists with similar degree distributions to those of C and T. The corresponding P value was calculated based on the permutation test results. Z-score < −1.5 and P < 0.05 were considered significantly proximal drug–HCoV associations. All networks were visualized using Gephi 0.9.2 (https://gephi.org/).
Network-based rational prediction of drug combinations
For this network-based approach for drug combinations to be effective, we need to establish if the topological relationship between two drug–target modules reflects biological and pharmacological relationships, while also quantifying their network-based relationship between drug targets and HCoV-associated host proteins (drug–drug–HCoV combinations). To identify potential drug combinations, we combined the top lists of drugs. Then, “separation” measure SAB was calculated for each pair of drugs A and B using the following method:
where \(\left\langle {d_ \cdot } \right\rangle\) was calculated based on the “closest” method. Our key methodology is that a drug combination is therapeutically effective only if it follows a specific relationship to the disease module, as captured by Complementary Exposure patterns in targets’ modules of both drugs without overlapping toxic mechanisms28.
Gene set enrichment analysis
We performed the gene set enrichment analysis as an additional prioritization method. We first collected three differential gene expression data sets of hosts infected by HCoVs from the NCBI Gene Expression Omnibus (GEO). Among them, two transcriptome data sets were SARS-CoV-infected samples from patient’s peripheral blood94 (GSE1739) and Calu-3 cells95 (GSE33267), respectively. One transcriptome data set was MERS-CoV-infected Calu-3 cells96 (GSE122876). Adjusted P value less than 0.01 was defined as differentially expressed genes. These data sets were used as HCoV–host signatures to evaluate the treatment effects of drugs. Differential gene expression in cells treated with various drugs were retrieved from the Connectivity Map (CMAP) database36, and were used as gene profiles for the drugs. For each drug that was in both the CMAP data set and our drug–target network, we calculated an enrichment score (ES) for each HCoV signature data set based on previously described methods97 as follows:
ESup and ESdown were calculated separately for the up- and down-regulated genes from the HCoV signature data set using the same method. We first computed aup/down and bup/down as
where j = 1, 2, …, s were the genes of HCoV signature data set sorted in ascending order by their rank in the gene profiles of the drug being evaluated. The rank of gene j is denoted by V(j), where 1 ≤ V(j) ≤ r, with r being the number of genes (12,849) from the drug profile. Then, ESup/down was set to aup/down if aup/down > bup/down, and was set to −bup/down if bup/down > aup/down. Permutation tests repeated 100 times using randomly generated gene lists with the same number of up- and down-regulated genes as the HCoV signature data set were performed to measure the significance of the ES scores. Drugs were considered to have potential treatment effect if ES > 0 and P < 0.05, and the number of such HCoV signature data sets were used as the final GSEA score that ranges from 0 to 3.
Data availability
All predicted repurposable drugs and network-predicted drug combinations can be freely accessed at https://github.com/ChengF-Lab/2019-nCoV.
Code availability
All codes can be freely accessed at https://github.com/ChengF-Lab/2019-nCoV.
References
Zumla, A., Chan, J. F., Azhar, E. I., Hui, D. S. & Yuen, K. Y. Coronaviruses—drug discovery and therapeutic options. Nat. Rev. Drug Discov. 15, 327–347 (2016).
Paules, C. I., Marston, H. D. & Fauci, A. S. Coronavirus infections—more than just the common cold. JAMA 323, 707–708 (2020).
de Wit, E., van Doremalen, N., Falzarano, D. & Munster, V. J. SARS and MERS: recent insights into emerging coronaviruses. Nat. Rev. Microbiol. 14, 523–534 (2016).
de Wilde, A. H., Snijder, E. J., Kikkert, M. & van Hemert, M. J. Host factors in coronavirus replication. Curr. Top. Microbiol. Immunol. 419, 1–42 (2018).
Chen, N. et al. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: a descriptive study. Lancet 395, 507–513 (2020).
Li, Q. et al. Early transmission dynamics in Wuhan, China, of novel coronavirus-infected pneumonia. N. Engl. J. Med. https://doi.org/10.1056/NEJMoa2001316 (2020) (in press).
Greene, J. A. & Loscalzo, J. Putting the patient back together—social medicine, network medicine, and the limits of reductionism. N. Engl. J. Med. 377, 2493–2499 (2017).
Avorn, J. The $2.6 billion pill-methodologic and policy considerations. N. Engl. J. Med. 372, 1877–1879 (2015).
Cheng, F. In silico oncology drug repositioning and polypharmacology. Methods Mol. Biol. 1878, 243–261 (2019).
Cheng, F., Hong, H., Yang, S. & Wei, Y. Individualized network-based drug repositioning infrastructure for precision oncology in the panomics era. Brief Bioinformatics 18, 682–697 (2017).
Cheng, F., Murray, J. L. & Rubin, D. H. Drug repurposing: new treatments for Zika virus infection? Trends Mol. Med. 22, 919–921 (2016).
Santos, R. et al. A comprehensive map of molecular drug targets. Nat. Rev. Drug Discov. 16, 19–34 (2017).
Cheng, F. et al. Network-based approach to prediction and population-based validation of in silico drug repurposing. Nat. Commun. 9, 2691 (2018).
Cheng, F. et al. Systems biology-based investigation of cellular antiviral drug targets identified by gene-trap insertional mutagenesis. PLoS Comput. Biol. 12, e1005074 (2016).
Yang, S., Fu, C., Lian, X., Dong, X. & Zhang, Z. Understanding human-virus protein-protein interactions using a human protein complex-based analysis framework. mSystems 4, e00303 (2019).
Liu, C. et al. Computational network biology: data, models, and applications. Phys. Rep. https://doi.org/10.1016/j.physrep.2019.12.004 (2020) (in press).
Dyall, J. et al. Repurposing of clinically developed drugs for treatment of Middle East respiratory syndrome coronavirus infection. Antimicrob. Agents Chemother. 58, 4885–4893 (2014).
Johansen, L. M. et al. FDA-approved selective estrogen receptor modulators inhibit Ebola virus infection. Sci. Transl. Med. 5, 190ra179 (2013).
He, S. et al. Repurposing of the antihistamine chlorcyclizine and related compounds for treatment of hepatitis C virus infection. Sci. Transl. Med. 7, 282ra249 (2015).
Barrows, N. J. et al. A screen of FDA-approved drugs for inhibitors of Zika virus infection. Cell Host. Microbe 20, 259–270 (2016).
Xu, M. et al. Identification of small-molecule inhibitors of Zika virus infection and induced neural cell death via a drug repurposing screen. Nat. Med. 22, 1101–1107 (2016).
Cheng, F. et al. Prediction of drug-target interactions and drug repositioning via network-based inference. PLoS Comput. Biol. 8, e1002503 (2012).
Cheng, F. et al. A genome-wide positioning systems network algorithm for in silico drug repurposing. Nat. Commun. 10, 3476 (2019).
Zeng, X. et al. deepDR: a network-based deep learning approach to in silico drug repositioning. Bioinformatics 35, 5191–5198 (2019).
Zeng, X. et al. Target identification among known drugs by deep learning from heterogeneous networks. Chem. Sci. 11, 1775–1797 (2020).
Zeng, X. et al. Network-based prediction of drug-target interactions using an arbitrary-order proximity embedded deep forest. Bioinformatics https://doi.org/10.1093/bioinformatics/btaa010 (2020) (in press).
Fang, J. et al. Network-based translation of GWAS findings to pathobiology and drug repurposing for Alzheimer’s disease. MedRxiv. https://doi.org/10.1101/2020.01.15.20017160 (2020).
Cheng, F., Kovacs, I. A. & Barabasi, A. L. Network-based prediction of drug combinations. Nat. Commun. 10, 1197 (2019).
Forni, D., Cagliani, R., Clerici, M. & Sironi, M. Molecular evolution of human coronavirus genomes. Trends Microbiol. 25, 35–48 (2017).
Kirchdoerfer, R. N. & Ward, A. B. Structure of the SARS-CoV nsp12 polymerase bound to nsp7 and nsp8 co-factors. Nat. Commun. 10, 2342 (2019).
Li, F., Li, W., Farzan, M. & Harrison, S. C. Structure of SARS coronavirus spike receptor-binding domain complexed with receptor. Science 309, 1864–1868 (2005).
Lu, R. et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet 395, 565–574 (2020).
Zhou, P. et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature https://doi.org/10.1038/s41586-020-2012-7 (2020).
Wrapp, D. et al. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science https://doi.org/10.1126/science.abb2507 (2020).
Chang, C. K., Chen, C. M., Chiang, M. H., Hsu, Y. L. & Huang, T. H. Transient oligomerization of the SARS-CoV N protein-implication for virus ribonucleoprotein packaging. PLoS ONE 8, e65045 (2013).
Lamb, J. et al. The Connectivity Map: using gene-expression signatures to connect small molecules, genes, and disease. Science 313, 1929–1935 (2006).
Lasso, G. et al. A structure-informed atlas of human-virus interactions. Cell 178, 1526–1541.e1516 (2019).
de Wilde, A. H. et al. Screening of an FDA-approved compound library identifies four small-molecule inhibitors of Middle East respiratory syndrome coronavirus replication in cell culture. Antimicrob. Agents Chemother. 58, 4875–4884 (2014).
Zhao, Y. et al. Toremifene interacts with and destabilizes the Ebola virus glycoprotein. Nature 535, 169–172 (2016).
Emmott, E. et al. The cellular interactome of the coronavirus infectious bronchitis virus nucleocapsid protein and functional implications for virus biology. J. Virol. 87, 9486–9500 (2013).
V’Kovski, P. et al. Determination of host proteins composing the microenvironment of coronavirus replicase complexes by proximity-labeling. Elife 8, e42037 (2019).
Moskowitz, D. W. & Johnson, F. E. The central role of angiotensin I-converting enzyme in vertebrate pathophysiology. Curr. Top. Med. Chem. 4, 1433–1454 (2004).
Seko, Y. Effect of the angiotensin II receptor blocker olmesartan on the development of murine acute myocarditis caused by coxsackievirus B3. Clin. Sci. 110, 379–386 (2006).
Erlandson, K. M. et al. The impact of statin and angiotensin-converting enzyme inhibitor/angiotensin receptor blocker therapy on cognitive function in adults with human immunodeficiency virus infection. Clin. Infect. Dis. 65, 2042–2049 (2017).
Wang, X. J. et al. Irbesartan, an FDA approved drug for hypertension and diabetic nephropathy, is a potent inhibitor for hepatitis B virus entry by disturbing Na(+)-dependent taurocholate cotransporting polypeptide activity. Antivir. Res. 120, 140–146 (2015).
Ko, C. et al. The FDA-approved drug irbesartan inhibits HBV-infection in HepG2 cells stably expressing sodium taurocholate co-transporting polypeptide. Antivir. Ther. 20, 835–842 (2015).
Hong, M. et al. Identification of a novel transcriptional repressor (HEPIS) that interacts with nsp-10 of SARS coronavirus. Viral Immunol. 21, 153–162 (2008).
McNulty, S., Flint, M., Nichol, S. T. & Spiropoulou, C. F. Host mTORC1 signaling regulates andes virus replication. J. Virol. 87, 912–922 (2013).
Stohr, S. et al. Host cell mTORC1 is required for HCV RNA replication. Gut 65, 2017–2028 (2016).
Wang, C. H. et al. Adjuvant treatment with a mammalian target of rapamycin inhibitor, sirolimus, and steroids improves outcomes in patients with severe H1N1 pneumonia and acute respiratory failure. Crit. Care Med. 42, 313–321 (2014).
Dyall, J. et al. Middle East respiratory syndrome and severe acute respiratory syndrome: current therapeutic options and potential targets for novel therapies. Drugs 77, 1935–1966 (2017).
Karran, P. & Attard, N. Thiopurines in current medical practice: molecular mechanisms and contributions to therapy-related cancer. Nat. Rev. Cancer 8, 24–36 (2008).
Chen, X., Chou, C. Y. & Chang, G. G. Thiopurine analogue inhibitors of severe acute respiratory syndrome-coronavirus papain-like protease, a deubiquitinating and deISGylating enzyme. Antivir. Chem. Chemother. 19, 151–156 (2009).
Cheng, K. W. et al. Thiopurine analogs and mycophenolic acid synergistically inhibit the papain-like protease of Middle East respiratory syndrome coronavirus. Antivir. Res. 115, 9–16 (2015).
Chen, H., Wurm, T., Britton, P., Brooks, G. & Hiscox, J. A. Interaction of the coronavirus nucleoprotein with nucleolar antigens and the host cell. J. Virol. 76, 5233–5250 (2002).
Rainsford, K. D. Influenza (“Bird Flu”), inflammation and anti-inflammatory/analgesic drugs. Inflammopharmacology 14, 2–9 (2006).
Garcia, C. C., Guabiraba, R., Soriani, F. M. & Teixeira, M. M. The development of anti-inflammatory drugs for infectious diseases. Discov. Med. 10, 479–488 (2010).
Silvestri, M. & Rossi, G. A. Melatonin: its possible role in the management of viral infections-a brief review. Ital. J. Pediatr. 39, 61 (2013).
Srinivasan, V., Mohamed, M. & Kato, H. Melatonin in bacterial and viral infections with focus on sepsis: a review. Recent Pat. Endocr. Metab. Immune Drug Discov. 6, 30–39 (2012).
Tan, D. X., Korkmaz, A., Reiter, R. J. & Manchester, L. C. Ebola virus disease: potential use of melatonin as a treatment. J. Pineal Res. 57, 381–384 (2014).
Tan, D. X., Manchester, L. C., Terron, M. P., Flores, L. J. & Reiter, R. J. One molecule, many derivatives: a never-ending interaction of melatonin with reactive oxygen and nitrogen species? J. Pineal Res. 42, 28–42 (2007).
Galano, A., Tan, D. X. & Reiter, R. J. On the free radical scavenging activities of melatonin’s metabolites, AFMK and AMK. J. Pineal Res. 54, 245–257 (2013).
Xiao, J., Shimada, M., Liu, W., Hu, D. & Matsumori, A. Anti-inflammatory effects of eplerenone on viral myocarditis. Eur. J. Heart Fail. 11, 349–353 (2009).
Wang, M. et al. Remdesivir and chloroquine effectively inhibit the recently emerged novel coronavirus (2019-nCoV) in vitro. Cell Res. https://doi.org/10.1038/s41422-020-0282-0 (2020).
Tan, X. et al. Systematic identification of synergistic drug pairs targeting HIV. Nat. Biotechnol. 30, 1125–1130 (2012).
Kindrachuk, J. et al. Antiviral potential of ERK/MAPK and PI3K/AKT/mTOR signaling modulation for Middle East respiratory syndrome coronavirus infection as identified by temporal kinome analysis. Antimicrob. Agents Chemother. 59, 1088–1099 (2015).
Lewis, E. L., Harbour, D. A., Beringer, J. E. & Grinsted, J. Differential in vitro inhibition of feline enteric coronavirus and feline infectious peritonitis virus by actinomycin D. J. Gen. Virol. 73, 3285–3288 (1992).
Zhou, W. B., Ding, Q., Chen, L., Liu, X. A. & Wang, S. Toremifene is an effective and safe alternative to tamoxifen in adjuvant endocrine therapy for breast cancer: results of four randomized trials. Breast Cancer Res. Treat. 128, 625–631 (2011).
Cong, Y. et al. MERS-CoV pathogenesis and antiviral efficacy of licensed drugs in human monocyte-derived antigen-presenting cells. PLoS ONE 13, e0194868 (2018).
Schwarz, S., Wang, K., Yu, W., Sun, B. & Schwarz, W. Emodin inhibits current through SARS-associated coronavirus 3a protein. Antivir. Res. 90, 64–69 (2011).
Ho, T. Y., Wu, S. L., Chen, J. C., Li, C. C. & Hsiang, C. Y. Emodin blocks the SARS coronavirus spike protein and angiotensin-converting enzyme 2 interaction. Antivir. Res. 74, 92–101 (2007).
Lambert, D. W., Clarke, N. E., Hooper, N. M. & Turner, A. J. Calmodulin interacts with angiotensin-converting enzyme-2 (ACE2) and inhibits shedding of its ectodomain. FEBS Lett. 582, 385–390 (2008).
Dai, J., Inscho, E. W., Yuan, L. & Hill, S. M. Modulation of intracellular calcium and calmodulin by melatonin in MCF-7 human breast cancer cells. J. Pineal Res. 32, 112–119 (2002).
Fung, T. S. & Liu, D. X. Activation of the c-Jun NH2-terminal kinase pathway by coronavirus infectious bronchitis virus promotes apoptosis independently of c-Jun. Cell Death Dis. 8, 3215 (2017).
Biedenkopf, N. et al. The natural compound silvestrol is a potent inhibitor of Ebola virus replication. Antivir. Res. 137, 76–81 (2017).
Muller, C. et al. Broad-spectrum antiviral activity of the eIF4A inhibitor silvestrol against corona- and picornaviruses. Antivir. Res. 150, 123–129 (2018).
Halder, A. K., Dutta, P., Kundu, M., Basu, S. & Nasipuri, M. Review of computational methods for virus-host protein interaction prediction: a case study on novel Ebola-human interactions. Brief. Funct. Genomics 17, 381–391 (2018).
Bedi, O., Dhawan, V., Sharma, P. L. & Kumar, P. Pleiotropic effects of statins: new therapeutic targets in drug design. Naunyn Schmiedebergs Arch. Pharmacol. 389, 695–712 (2016).
Li, Q. et al. Integrative functional genomics of hepatitis C virus infection identifies host dependencies in complete viral replication cycle. PLoS Pathog. 10, e1004163 (2014).
Gebre, M., Nomburg, J. L. & Gewurz, B. E. CRISPR-Cas9 genetic analysis of virus-host interactions. Viruses 10, 55 (2018).
Kim, J. H. et al. Acute eosinophilic pneumonia related to a mesalazine suppository. Asia Pac. Allergy 3, 136–139 (2013).
Gupta, A. & Gulati, S. Mesalamine induced eosinophilic pneumonia. Respir. Med. Case Rep. 21, 116–117 (2017).
Chiang, C. W. et al. Translational high-dimensional drug interaction discovery and validation using health record databases and pharmacokinetics models. Clin. Pharmacol. Ther. 103, 287–295 (2018).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549 (2018).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Law, V. et al. DrugBank 4.0: shedding new light on drug metabolism. Nucleic Acids Res. 42, D1091–D1097 (2014).
Yang, H. et al. Therapeutic target database update 2016: enriched resource for bench to clinical drug target and targeted pathway information. Nucleic Acids Res. 44, D1069–D1074 (2016).
Gaulton, A. et al. ChEMBL: a large-scale bioactivity database for drug discovery. Nucleic Acids Res. 40, D1100–D1107 (2012).
Liu, T. Q., Lin, Y. M., Wen, X., Jorissen, R. N. & Gilson, M. K. BindingDB: a web-accessible database of experimentally determined protein-ligand binding affinities. Nucleic Acids Res. 35, D198–D201 (2007).
Pawson, A. J. et al. The IUPHAR/BPS Guide to PHARMACOLOGY: an expert-driven knowledgebase of drug targets and their ligands. Nucleic Acids Res. 42, D1098–D1106 (2014).
Apweiler, R. et al. UniProt: the Universal Protein knowledgebase. Nucleic Acids Res. 32, D115–D119 (2004).
Coordinators, N. R. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 44, D7–D19 (2016).
Smith, I. N., Thacker, S., Seyfi, M., Cheng, F. & Eng, C. Conformational dynamics and allosteric regulation landscapes of germline PTEN mutations associated with autism compared to those associated with cancer. Am. J. Hum. Genet. 104, 861–878 (2019).
Reghunathan, R. et al. Expression profile of immune response genes in patients with severe acute respiratory syndrome. BMC Immunol. 6, 2 (2005).
Josset, L. et al. Cell host response to infection with novel human coronavirus EMC predicts potential antivirals and important differences with SARS coronavirus. mBio 4, e00165–00113 (2013).
Yuan, S. et al. SREBP-dependent lipidomic reprogramming as a broad-spectrum antiviral target. Nat. Commun. 10, 120 (2019).
Sirota, M. et al. Discovery and preclinical validation of drug indications using compendia of public gene expression data. Sci. Transl. Med. 3, 96ra77 (2011).
Acknowledgements
This work was supported by the National Heart, Lung, and Blood Institute of the National Institutes of Health (NIH) under Award Number K99 HL138272 and R00 HL138272 to F.C. The content of this publication does not necessarily reflect the views of the Cleveland Clinic.
Author information
Authors and Affiliations
Contributions
F.C. conceived the study. Y.Z. and Y.H. developed the network methodology and performed all computational experiments. J.S., Y.Z., Y.H., and W.M. performed data analysis. F.C., Y.Z., and Y.H. wrote and critically revised the manuscript with contributions from other co-authors.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhou, Y., Hou, Y., Shen, J. et al. Network-based drug repurposing for novel coronavirus 2019-nCoV/SARS-CoV-2. Cell Discov 6, 14 (2020). https://doi.org/10.1038/s41421-020-0153-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41421-020-0153-3
This article is cited by
-
A network-based drug prioritization and combination analysis for the MEK5/ERK5 pathway in breast cancer
BioData Mining (2024)
-
The cGAS-STING pathway in COPD: targeting its role and therapeutic potential
Respiratory Research (2024)
-
Generic model to unravel the deeper insights of viral infections: an empirical application of evolutionary graph coloring in computational network biology
BMC Bioinformatics (2024)
-
Exploring potential therapeutic combinations for castration-sensitive prostate cancer using supercomputers: a proof of concept study
Scientific Reports (2024)
-
Molecular docking as a tool for the discovery of novel insight about the role of acid sphingomyelinase inhibitors in SARS- CoV-2 infectivity
BMC Public Health (2024)