Abstract
Expression of the adenovirus E1A oncogene sensitizes tumor cells to innate immune rejection by NK cells. This increased NK sensitivity is only partly explained by an E1A-induced increase in target cell surface expression of NKG2D ligands. The post-recognition mechanisms by which E1A sensitizes cells to the apoptotic cell death response to NK injury remains to be defined. E1A sensitizes cells to apoptotic stimuli through two distinct mechanisms—repression of NF-κB-dependent antiapoptotic responses and enhancement of caspase-2 activation and related mitochondrial injury. The current studies examined the roles of each of these post-NKG2D-recognition pathways in the increased sensitivity of E1A-positive target cells to NK killing. Sensitization to NK-induced apoptosis was independent of E1A-mediated repression of cellular NF-κB responses but was dependent on the expression of both caspase-2 and the upstream, caspase-2 activating molecule, PIDD. Target cells lacking caspase-2 or PIDD expression retained E1A-induced increased expression of the NKG2D ligand, RAE-1. NK cell-induced mitochondrial injury of E1A-expressing cells did not require expression of the mitochondrial molecules, Bak or Bax. These results define a PIDD/caspase-2-dependent pathway, through which E1A sensitizes cells to NK-mediated cytolysis independently of and complementarily to E1A-enhanced NKG2D/RAE-1 ligand expression.
Similar content being viewed by others
Introduction
Through its ability to deregulate the cell cycle, E1A can immortalize mammalian cells when expressed after abortive infection or stable transfection, in a manner similar to other DNA tumor virus oncogenes and cellular oncogenes1,2. E1A immortalized cells are not tumorigenic in immunocompetent hosts, since E1A sensitizes cells to the apoptosis-inducing (and therefore tumor rejecting) activities of host immune effector cells, such as natural killer (NK) cells, cytotoxic T lymphocyte (CTL), and activated macrophages3. Data from several experimental animal models support the conclusion that such cellular immune defenses prevent tumor development by E1A-positive tumor cells. Newborn rats are more sensitive to tumor development by E1A-positive sarcoma cells than mature rats. The resistance of mature rats to tumor development by E1A-positive sarcoma cells is the result of increased NK activity in mature rats in comparison to newborn rats4. Studies comparing rejection of E1A-positive tumor cells by immunocompetent mice (T cell positive, NK cell positive), immunocompetent mice depleted of NK cells (T cell positive, NK cell negative), nude mice (T cell negative, NK cell positive), or CD3-ε transgenic mice (T cell negative, NK-negative) demonstrated the ability of the host NK cell response to reject E1A-expressing tumor cells in a T cell-independent manner5. Although the role of NK cells in rejection of E1A-positive tumor cells is well established, the cellular mechanisms through which E1A renders tumor cells sensitive to NK killing and tumor challenge rejection are incompletely defined.
NK killing of E1A-positive tumor target cells requires killer cell receptor interactions with NKG2D ligands in both human (MIC-A/MIC-B, ULBP) and murine tumor cell lines (RAE-1)6. The interactions of NKG2D and its ligands are required for efficient cytolysis of E1A-positive cells, as evidenced by blocking the interaction with blocking antibodies6. Furthermore, overexpression of RAE-1 in E1A-negative mouse tumor cells increases their susceptibility to NK cytolytic activity6. However, E1A-positive cells are still more susceptible to NK killing than E1A-negative/RAE-1-overexpressing cells, suggesting that enhanced RAE-1 expression is not the sole mechanism through which E1A renders cells sensitive to NK cytolytic activity6.
Cytolytic lymphocytes use two main mechanisms to kill target cells—perforin/granzyme and TNF/Fas mediated apoptosis—and E1A expression sensitizes cells to both of these mechanisms7,8,9,10. E1A modulates two cellular pathways to sensitize cells to apoptotic injury, depending on the nature of the injury11,12. E1A sensitizes cells to apoptosis induced through the extrinsic apoptosis pathway (triggered by TNF, Fas ligand, and TRAIL) by repressing transcription of NF-κB-dependent antiapoptotic gene expression13,14,15. We have also demonstrated that E1A sensitizes cells to apoptosis from intrinsic pathway injuries (chemotherapeutic agents and nitric oxide (NO)) through NF-κB-independent cellular mechanisms but is dependent on enhanced PIDD-mediated activation of caspase-211,12,16.
We used rat NK cell killing of E1A-expressing target cells to determine the relative importance of NF-κB-dependent and NF-κB-independent cellular mechanisms of E1A-induced sensitivity to NK cytolytic activity. Rat NK cells are the most well characterized NK cells for systems testing the immunobiology and tumorigenicity of E1A-positive tumor cells. Rat NK can kill E1A-positive targets of mouse, rat, hamster, and human origin; the target cell killing is E1A expression level dependent, and there is a strong correlation between rat NK activity and ontogeny and in vivo rejection of E1A-positive tumor cells4,17,18,19,20. Furthermore, nude rat spleens can be used as a rich source of NK cells for assays such as those reported here, since the spleens contain approximately ten times the NK cytolytic activity of nude mouse spleens against cytolytic-susceptible target cells, such as mouse YAC-1 cells4. In situations where pure NK cell populations are required experimentally, the rat RNK-16 cell line can be used to reproduce nude rat spleen NK cell killing of E1A-positive target cells, and RNK-16 killing of mouse target cells is dependent on killer cell expression of NKG2D19,20,21.
Rat NK cell injury of E1A-expressing mouse cells was used in these studies to test the hypothesis that E1A-mediated sensitization to NK cytolysis is dependent on the PIDD–caspase-2 apoptotic pathway. Our results showed that NK cell-induced apoptosis of E1A-positive cells is independent of NF-κB cellular responses and that expression of caspase-2 and PIDD is required for NK-cell-induced apoptosis. The resistance of PIDD or caspase-2 knockdown cells to NK cell killing was independent of the level of target cell surface expression of RAE-1, the key NKG2D ligand in NK recognition of E1A-positive mouse cells, since cells with repressed PIDD and caspase-2 expressed a level of cell surface RAE-1 that was equivalent to that of parental E1A-positive cells that have normal levels of PIDD and caspase-2 expression. Collectively, these data indicate that E1A-induced sensitization to NK-mediated apoptosis occurs as the result of two independent but complementary E1A activities—increased recognition of E1A-positive cells by increased cell surface expression of NKG2D ligands, as reported previously, and enhanced activation of the PIDD–Caspase-2 apoptotic signaling pathway6, as defined in this report.
Results
NK cell mediated cytolysis of E1A-expressing cells is dependent on caspase activity
Our laboratory has reported that E1A-expressing cells display both nuclear condensation and DNA fragmentation after incubation with cytolytic lymphocytes, indicating target cell death by apoptosis22. NK cells induce both caspase-dependent and -independent apoptotic pathways in target cells23,24,25. To determine if caspase activity is required for NK cell-induced apoptosis of E1A-positive target cells, we tested RNK-16 cell killing of E1A-positive MCA102 and BMK cells in the absence or presence of the pan-caspase inhibitor, zVAD (100 μM). Both E1A-positive cell types were highly susceptible to killing by RNK-16 cells, when compared with the relevant E1A-negative cells, and, in both cases, E1A-positive target cell killing was repressed in the presence of zVAD (Fig. 1a, b, respectively). Therefore, the cell death response to NK injury was caspase dependent.
a [Cr51]-labeled MCA102 (circle), MCA102 E1A (triangle) cells were incubated with RNK-16 cells at the indicated RNK:target ratios in the absence or presence (inverted triangle) of 100 μM zVAD-fmk. After 6 h, supernatants were collected and % specific Cr51 release was assessed (mean ± SEM; n = 4, ***P < 0.0001, **P ≤ 0.0084, one-way ANOVA). b [Cr51]-labeled BMK (circle) or BMK-E1A (triangle) cells were incubated with RNK-16 cells at the indicated RNK:target ratios in the absence or presence (inverted triangle) of 100 μM zVAD-fmk. After 6 h, supernatants were collected and % specific Cr51 release was assessed (mean ± SEM; n = 4, ***P ≤ 0.0001, one-way ANOVA)
Target cell sensitivity to NK cell-induced apoptosis is not caused by E1A repression of NF-κB-dependent antiapoptotic defenses
To evaluate the role of E1A repression of NF-κB activation in sensitization to NK cell-induced apoptosis, we compared RNK-16 killing of NIH-3T3 cells stably transfected with E1A and known to be highly NK-sensitive (3T3 E1A) with two derivative 3T3 E1A cell lines created to be TNF resistant because of their NF-κB activation responses—3T3 E1ATNFr and 3T3 E1A p6512. Despite the high level, NF-κB-dependent resistance of these derivative cell lines to TNF, they were equally susceptible to killing by RNK-16 cells as parental, E1A-positive cells (3T3 E1A) (Fig. 2). The data in Fig. 2 indicate that the cellular mechanisms through which E1A sensitizes NIH-3T3 cells to NK killing are unrelated to E1A repression of NF-κB-dependent antiapoptotic effects.
[Cr51]-labeled NIH-3T3 (circle), 3T3-E1A (square), 3T3-E1A TNF resistant (TNFr) (triangle) or 3T3-E1A p65 RelA (p65) (diamond) cells were incubated with RNK-16 cells at the indicated RNK:target ratios. After 6 h, supernatants were collected and % specific Cr51 release was assessed (mean ± SEM; n = 3, **P = 0.0015, *P = 0.0126, one-way ANOVA)
NK cells induce mitochondrial injury of E1A-expressing cells
Our studies of other, NF-κB-independent apoptotic stimuli revealed that E1A enhances caspase-dependent, irreversible mitochondrial dysfunction after NO or chemotherapeutic drug injury. Therefore, we tested the NK cell cytotoxicity system, to determine whether E1A-positive cells exhibited a similar loss of mitochondrial function. RNK-16 cells were stained with cell tracker blue and then incubated in contact with 3T3 E1A cells for 4 h, after which mitochondrial membrane potential was assayed, using TMRE staining and flow cytometry (Fig. 3). A non-blue gate was used to assay TMRE staining of E1A-positive cells. The results showed that RNK-16 cells induced a loss of TMRE staining at NK cell:target cell ratios of 5, 10, and 20 to 1. The data presented to this point indicated that the mechanisms through which E1A sensitizes cells to NK killing involve both caspase activation (Fig. 1) and mitochondrial injury (Fig. 3).
3T3-E1A cells were incubated with cell tracker blue-stained RNK-16 cells at the indicated ratios of RNK:E1A cells for 4 h and then stained with TMRE. Histograms show non-blue gated cells (3T3-E1A cells) and are representative of four individual experiments. Percentage of non-blue cells are shown above the markers
E1A-induced cellular sensitization to NK killing does not require Bak or Bax
The data in Fig. 3 indicated that NK killing of E1A-positive cells is associated with a marked loss of mitochondrial membrane potential. Such injury-induced mitochondrial dysfunction in cells destined for apoptotic death can be mediated by the mitochondrial effects of proapoptotic, Bcl-2 family members, including Bid, Bak, and Bax. Stimulus induced cleavage of Bid to its active form, tBid, triggers homodimerization of Bak and Bax, both of which alter mitochondrial membranes to cause loss of mitochondrial membrane potential and leakage of cytochrome c into the cytosol26. Granzyme B, produced by NK cells during cytolytic injury, can cause target cell mitochondrial depolarization and apoptosis both through this Bid/Bak/Bax pathway and through mechanisms that are independent of this pathway27,28,29.
To determine whether E1A-induced target cell sensitization to NK cell-induced mitochondrial injury involves the Bid/Bak/Bax pathway, we obtained Bak, Bax, and Bak/Bax single and double knockout BMK cells that were transformed with E1A and dominant-negative mutant p5330. To confirm the role of Bak and Bax in the intrinsic apoptotic pathway in these cells, we treated wild type, Bak deficient (−/−), Bax deficient (−/−) or Bak/Bax double-deficient cells with ceramide. Ceramide triggers the intrinsic apoptotic pathway and results in mitochondrial injury that is mediated through Bak and Bax31,32,33,34. Wild type, Bak−/− and Bax−/− BMK cells were sensitive to ceramide-induced apoptosis, whereas Bak/Bax double-deficient cells were resistant (Fig. 4a). These results are similar to those reported with TNF-α and cycloheximide treatment30. As shown in Fig. 4b, cells expressing E1A and sufficient in Bak or Bax, deficient in either Bak or Bax or deficient in both Bak and Bax were equivalently sensitive to RNK-16 induced cytolysis. In the absence of both Bak and Bax, RNK-mediated apoptosis still required caspase activity (Fig. 4c). These data show that E1A enhancement of the intrinsic (mitochondrial injury) apoptosis pathway activated by NK cells is independent of Bid/Bak/Bax mechanisms.
a [H3]-thymidine-labeled BMK-E1A, BMK-E1A-Bak−/− (BMK Bak), BMK-E1A-Bax−/− (BMK Bax), or BMK-E1A-Bak/Bax−/−/−/− (BMK DKO) cells were incubated with ceramide (100 µM) overnight. Supernatants were collected and % specific thymidine release was assessed (mean ± SEM; n = 5, ***P = 0.001, one-way ANOVA). b [Cr51]-labeled BMK (circle), BMK-E1A (triangle), BMK-E1A-Bak−/− (square), BMK-E1A-Bax−/− (inverted triangle) or BMK-E1A-Bak/Bax−/−/−/− (double knockout = DKO, diamond) cells were incubated with RNK-16 cells at the indicated RNK:target ratios. After 6 h, supernatants were collected and % specific 51Cr release was assessed (mean ± SEM; n = 4, ***P ≤ 0.0003, one-way ANOVA). c [Cr51]-labeled BMK (circle) or BMK-E1A-Bak/Bax−/−/−/− (double knockout = DKO, diamonds) cells were incubated with RNK-16 cells at the indicated RNK:target ratios in the absence (filled diamond) or presence (open diamond) of 100 μM zVAD-fmk. After 6 h, supernatants were collected and % specific 51Cr release was assessed (mean ± SEM; n = 4, ***P ≤ 0.0001, **P = 0.0016, one-way ANOVA)
NK-mediated cytolysis of E1A-expressing cells requires Caspase-2 and PIDD expression
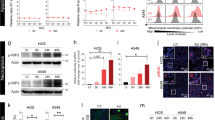
We have reported that E1A sensitization to intrinsic apoptosis, induced by both NO and etoposide, requires expression of both caspase-2 and its main activating platform member PIDD11,12. Both injuries induce caspase-dependent apoptosis and mitochondrial injury similar to what we observed with RNK-mediated injury of E1A-expressing cells in this study. We used E1A-positive, PIDD (E1A iPIDD), and caspase-2 (E1A iC2) shRNA knockdown cells to determine whether the PIDD–caspase-2 pathway is required for NK-mediated cytolysis of E1A-expressing target cells (Fig. 5a, b)11,12. Both types of knockdown cells were significantly less sensitive to lysis by RNK-16 NK cells (Fig. 5c) and nude rat splenic NK cells (Fig. 5d) than parental control 3T3 E1A cells, indicating that PIDD and caspase-2 are required for E1A sensitization to NK killing.
a Expression of Casp-2, E1A and actin in 3T3, 3T3-E1A and E1A-iC2 cell lines (originally published in12. b Expression of PIDD, E1A and actin in 3T3, 3T3-E1A and E1A-iPIDD cell lines11. c [Cr51]-labeled 3T3 (circle), 3T3-E1A (square), E1A-iC2 (inverted triangle), and E1A-iPIDD (triangle) cells were incubated with nude rat splenic NK cells at the indicated spleen cell:target ratios. % specific 51Cr release was assessed (mean ± SEM; n = 4, ***P ≤ 0.0002, **P = 0.0012, one-way ANOVA). d [Cr51]-labeled 3T3 (circle), 3T3-E1A (square), E1A-iC2 (inverted triangle), and E1A-iPIDD (triangle) cells were incubated with RNK-16 cells at the indicated RNK:target ratios. % specific 51Cr release was assessed (mean ± SEM; n = 4, ***P ≤ 0.0008, one-way ANOVA). d RAE-1 expression on 3T3 (filled histogram) compared to 3T3-E1A (open histogram), E1A-iC2 (open histogram), and E1A-iPIDD (open histogram) cells. Data shown are representative of four individual experiments
Caspase-2 and PIDD-deficient cells retain increased RAE-1 expression
We have reported that E1A expression induces increased cell surface expression of the NKG2D ligands, RAE-1 (mouse cells) and MIC-A/B and ULPBP (human cells) and that this increased expression of NKG2D ligands on mouse or human cells results in an increased susceptibility to NK cell killing6. Rat NK cell killing of mouse tumor cells is dependent on the interaction of NKG2D and NKG2D ligands21,35. To exclude the possibility that selection of the clonally derived, E1A-positive caspase-2 (E1A iC2) and PIDD (E1A iPIDD) knockdown cells might have co-selected for cells with reduced NKG2D ligand expression, RAE-1 cell surface expression was examined by flow cytometry. Figure 5e shows that both E1A iC2 and E1A iPIDD cells retain the same level of cell surface expression of RAE-1 as the E1A-positive parental cells from which the clones were derived. E1A expression level in the E1A iC2 and E1A iPIDD cells was also comparable to that of the parental cells11,12. In addition to RAE-1, Mult-1 and H60 are other mouse NKG2D ligands. Expression of H60 was not enhanced in E1A-expressing cells (data not shown) and Mult-1 expression was also not altered by expression of E1A6, indicating that, in this model system, RAE-1 expression is the sole NKG2D ligand. In summary, these data indicated that E1A expression mediates increased susceptibility of target cells to NK killing through two independent and complementary pathways. The first is increased target cell recognition through increased cell surface NKG2D ligand (e.g., RAE-1) expression. The second is enhancement of the target cell proapoptotic response to NK cell injury, through the PIDD–caspase-2 pathway.
Discussion
Expression of E1A sensitizes cells to the cytolytic machinery of NK cells. Previously, it has been shown that E1A expression results in increased expression of NKG2D ligands on the cell surface allowing for increased recognition by and increased sensitivity to lysis by NK cells6. However, increased expression of NKG2D ligand without E1A expression did not fully replicate the sensitizing effect of E1A6. The data presented here define PIDD–caspase-2 as the key, post-recognition cellular pathway through which E1A enhances target cell susceptibility to NK-cell-mediated cytolytic activity, as a complementary mechanism to E1A-induced increased expression of NKG2D ligands6. These results and our previously reported data also suggest that PIDD-dependent caspase-2 activation enhancement is a common cellular mechanism through which E1A sensitizes cells to a variety of stimuli of the intrinsic apoptosis pathway, including macrophage-NO and proapoptotic chemotherapeutic drugs11,12,16.
NK cells can kill target cells using multiple effector mechanisms, including production of TNF-superfamily members and transfer of cytolytic granules composed of perforin and granzyme at the NK cell/target cell synapse36. E1A utilizes two general mechanisms to sensitize cells to apoptotic cell death, depending on whether the proapoptotic stimulus triggers the extrinsic or the intrinsic apoptotic pathway. Sensitization to the extrinsic apoptotic pathway, triggered by members of the TNF superfamily, involves E1A repression of NF-κB-dependent transcription of antiapoptotic molecules13,14,37,38. The current data show that NK cells can also induce caspase dependent apoptotic cell death of E1A-expressing cells (Fig. 1) through mechanisms that are independent of E1A repression of NK-κB-dependent, antiapoptotic defenses (Fig. 2) but that mediate mitochondrial injury (Fig. 3). We have reported that induction of the intrinsic apoptotic pathway in E1A-expressing cells by NO and chemotherapeutic drugs requires PIDD-dependent activation of caspase-2 (refs. 11,12,16). The current studies extend that mechanistic pattern to NK cell killing of E1A-expressing target cells (Fig. 5c, d).
The multiple roles of caspase-2 in apoptosis remain to be completely defined. Caspase-2 can function as both an initiator and an effector caspase39. In its role as an initiator caspase, caspase-2 requires the formation of high molecular weight protein complexes, using the initiation platform called the PIDDosome40. The requirement, in our studies, for both PIDD and caspase-2 expression for efficient E1A sensitization of target cells to NK cell killing indicates that caspase-2 is functioning as an initiator caspase, during NK triggering of target cell apoptosis (Fig. 5c, d). These data and those published previously strongly suggest that E1A expression enhances caspase-2 activation through the PIDDosome in response to a variety of intrinsic apoptotic injuries11,16, including those mediated by NK cells.
Our observations provide insights into other possible cellular mechanisms through which E1A might enhance NK cell-induced, PIDD and caspase-2 dependent apoptosis of target cells. The short-term (6 h) NK assays used for these experiments reflect killing induced by cytolytic granules (composed of perforin and granzymes). Granzymes can cleave several cellular target molecules, including Bid, caspase-3, Hsp70, and Hsp90. Therefore, the question arises about whether E1A affects one or more of those granzyme-targeted cell molecule interactions in a way that enhances NK cell triggered PIDD–caspase-2 activation.
Previous studies have shown that granzymes can induce cleavage of Bid, which results in Bid mediated activation of Bax and Bak leading to mitochondrial injury26,41,42,43. Our data show that E1A-expressing cells remain sensitive to NK cell lysis in the absence of Bak and/or Bax (Fig. 4b), indicating that granzyme cleavage of Bid and subsequent Bak/Bax mediated mitochondrial injury are not required for E1A sensitization to NK cell-induced apoptosis. Our data on the lack of a role of Bcl-2 family members in modulating the susceptibility of E1A-expressing cells to NK killing are in agreement with those of others and ours that show that both NK and CTL can induce cell death of E1A-positive cells during expression of the adenoviral gene E1B 19K, which is a viral ortholog of Bcl-2 (refs. 8,27,44,45), which inhibits the proapoptotic effects of Bak and Bax46.
Cytolytic lymphocyte granzyme B can cleave capase-3, which can, in turn, cleave and activate caspase-2, thereby potentially bypassing the requirement for PIDD/PIDDosome activation of caspase-2 in the apoptotic response47,48,49. It is possible, therefore, that there might be both PIDD-dependent, initiator caspase-2 activation and PIDD-independent, caspase-3 enhanced downstream caspase-2 activation in E1A-positive target cells injured by NK cells. However, the marked reduction in NK susceptibility of E1A-positive, PIDD knockdown cells (E1A iPIDD; Fig. 5c, d)—almost to the level of cytolytic-resistant, E1A-negative cells—suggests that most of the E1A enhancing effect is PIDD-dependent and therefore at the level of initiator caspase-2 activity and not a downstream, post-caspase-3 effector caspase activity50.
Heat-shock proteins (Hsp70 and Hsp90) are PIDD chaperones that stabilize PIDD. Hsp90 binds full-length PIDD and is required for PIDD processing to both PIDD-C and PIDD-CC, the caspase activating form of PIDD51. Hsp70 and Hsp90 interact with PIDD, resulting in auto-proteolysis of PIDD from full-length to the active PIDD-CC form52. The presence of Hsp90 bound to PIDD-CC also acts as a negative regulator of caspase-2, by blocking the interaction of caspase-2 with PIDD-CC53. Therefore, the balance of Hsp interactions with PIDD and caspase-2 might determine the course of the cellular apoptotic response to a given stimulus. It is possible that NK cell granzyme activity and target cell E1A effects could affect this balance. Granzyme B can cleave Hsp70 and Hsp90 in target cells54,55,56. E1A induces increased expression of Hsp90 (ref. 57) and Hsp70 (ref. 58) and, therefore, might increase Hsp-mediated stabilization and processing of PIDD. Clarification of these intermolecular interactions among heat-shock proteins, PIDD, caspase-2, and E1A, in the context of NK cell injury of target cells, will require further study. Considering the variety of cellular molecular targets of E1A and granzymes that might impinge on the PIDD/caspase-2 activation process, it is not unlikely that there could be other mechanisms that might be involved in this E1A enhancing effect on the cellular apoptotic response.
E1A increases NKG2D ligand expression that causes increased target cell recognition by NK cells6. However, increased NKG2D ligand expression, in the absence of E1A, does not induce comparable target cell susceptibility to NK cell-induced killing and does not prevent tumor formation in mice6. These results suggest that there is a post-recognition mechanism through which E1A sensitizes cells to NK cell injury and that is independent of and complementary to effects on NKG2D ligand expression. In the current study, the two cell lines created to express reduced levels of either caspase-2 (E1A iC2) or PIDD (E1A iPIDD) retained the E1A-related increase in cell surface RAE-1 expression (Fig. 5e) but were significantly less susceptible to NK killing than the parental, E1A-positive cells from which they were derived (Fig. 5c, d). In summary, the data presented in this report support the conclusion that the E1A enhancing effect on the PIDD–caspase-2 activation response is the major post-recognition mechanism through which E1A enhances the target cell death response to NK cell injury. Considering the similar findings in our previous studies of NO and chemotherapy-induced apoptosis of E1A-expressing cells, we propose that E1A-mediated enhancement of PIDD-dependent activation of caspase-2 activity is the common pathway through which E1A sensitizes cells to stimuli that trigger the intrinsic apoptosis pathway.
Methods and materials
Cell lines and cell line characterization
The C57/B6-dervied, methylcholanthrene-induced sarcoma cell line, MCA102, was provided by Dr. N. Restifo (National Institutes of Health, Bethesda, MD) and maintained in Dulbecco's modified Eagle's medium (DMEM) with antibiotics and 5% fetal bovine serum (FBS)59. MCA102 E1A cells have been described37. E1A transformed baby mouse kidney cells (BMK) from wild type (wt; Bak+/+, Bax+/+), Bak−/−, Bax−/− and Bak−/− Bax−/− mice were obtained from Eileen White and maintained in DMEM with antibiotics and 10% FBS30. An NIH-3T3 cell line expressing Ad5 E1A 12S protein (MT12–1; herein referred to as 3T3 E1A) and its derivatives were maintained in DMEM supplemented with antibiotics and 5% calf serum12,60. The TNF-resistant, E1A-expressing NIH-3T3 cell lines, TR1 (herein referred to as 3T3 E1ATNFr) and A1 (herein referred to as 3T3 E1A p65), have been characterized12. These cell lines were derived from MT12–1/3T3 E1A cells, either through selection for survival in TNF (3T3 E1ATNFr) or through stable overexpression of the key NF-κB subunit, p65/RelA (3T3 E1A p65). 3T3-E1A+shRNA Caspase-2 cells (E1A-iC2) and 3T3-E1A+shRNA PIDD cells (E1A-iPIDD) have been characterized11,12. RNK-16 cells were provided by M. Nakamura (University of California at San Francisco) and were maintained in RPMI supplemented with 10% fetal bovine serum and β-mercaptoethanol61.
Isolation of NK cells and cytotoxicity assays
Use of nude rats for isolation of spleen NK cells was approved by the Edward Hines, Jr VA Hospital IACUC committee. Rat splenic NK cells were isolated from 2-month-old nude rats as previously described with modifications62. Spleen cells were treated with ACK lysis buffer (Lonza) to lyse red blood cells. Resulting cells were resuspended in 2 ml RPMI+10% FBS and put on a nylon wool column for 1 h at 37 °C. Columns were washed with 10 ml of warm medium to elute the B-cell-depleted cell population. Aliquots of the resulting cell suspensions were stained with anti-CD161 (Clone 10/78 AbCam), >80% of the B-cell-depleted population was CD161+. 51Cr-release cytotoxicity assays were performed as described63.
RNK-16 mediated effects on target cell mitochondrial membrane permeability
RNK-16 cells were stained with 80 μM Cell Tracker Blue (CTB) (Molecular Probes) and were incubated at the indicated ratios with 3T3-E1A cells in 24-well plates for 6 h at 37 °C. After 6 h, all cells were stained with 100 nM TMRE (Molecular Probes). All cells were collected with trypsin-EDTA and processed for flow cytometry on a BD LSR. A non-blue gate was used to separate 3T3-E1A cells from CTB-RNK-16 cells for analysis of mitochondrial membrane potential of the 3T3-E1A cells. CTB-stained RNK-16 cells retained equivalent cytolytic activity as non-stained RNK-16 cells.
Extracellular RAE-1 staining
Cells were harvested in 2 mM EDTA-PBS and stained with either PE-conjugated monoclonal anti-mouse panspecific RAE-1 antibody or Rat IgG2A isotype control (R&D Systems), and flow cytometry was performed on an Accuri C6.
Statistical analysis
Graph Pad Prism 6 was used for statistical analysis. One-way ANOVA was first performed followed by post hoc analysis by the Holm–Sidak method to determine significance of differences between two specific data sets. For all analyses, P < 0.05 was considered a significant difference.
References
Felsani, A., Mileo, A. M. & Paggi, M. G. Retinoblastoma family proteins as key targets of the small DNA virus oncoproteins. Oncogene 25, 5277–5285 (2006).
Ralston, R. & Bishop, M. The protein products of the myc and myb oncogenes and adenovirus E1a are structurally related. Nature 306, 803–806 (1983).
Cook, J. L. & Routes, J. M. Role of the innate immune response in determining the tumorigenicity of neoplastic cells. 106, 99–107; discussion 107–108, 143–160 (2001).
Cook, J. L. & Lewis, A. M. Immunological surveillance against DNA-virus-transformed cells: correlations between natural killer cell cytolytic competence and tumor susceptibility of athymic rodents. J. Virol. 61, 2155–2161 (1987).
Routes, J. M., Ryan, S., Li, H., Steinke, J. & Cook, J. L. Dissimilar immunogenicities of human papillomavirus E7 and adenovirus E1A proteins influence primary tumor development. Virology 277, 48–57 (2000).
Routes, J. M. et al. Adenovirus serotype 5 E1A sensitizes tumor cells to NKG2D-dependent NK cell lysis and tumor rejection. J. Exp. Med. 202, 1477–1482 (2005).
Cook, J. L., Potter, T. A., Bellgrau, D. & Routes, B. A. E1A oncogene expression in target cells induces cytolytic susceptibility at a post-recognition stage in the interaction with killer lymphocytes. Oncogene 13, 833–842 (1996).
Klefstrom, J. et al. c-Myc and E1A induced cellular sensitivity to activated NK cells involves cytotoxic granules as death effectors. Oncogene 18, 2181–2188 (1999).
Berke, G. The binding and lysis of target cells by cytotoxic lymphocytes: molecular and cellular aspects. Annu. Rev. Immunol. 12, 735–773 (1994).
Arase, H., Arase, N. & Saito, T. Fas-mediated cytotoxicity by freshly isolated natural killer cells. J. Exp. Med. 181, 1235–1238 (1995).
Radke, J. R., Siddiqui, Z. K., Figueroa, I. & Cook, J. L. E1A enhances cellular sensitivity to DNA-damage-induced apoptosis through PIDD-dependent caspase-2 activation. Cell Death Discov. 2, 16076 (2016).
Radke, J. R., Siddiqui, Z. K., Miura, T. A., Routes, J. M. & Cook, J. L. E1A oncogene enhancement of caspase-2-mediated mitochondrial injury sensitizes cells to macrophage nitric oxide-induced apoptosis. J. Immunol. 180, 8272–8279 (2008).
Shao, R. et al. E1A sensitizes cells to tumor necrosis factor-induced apoptosis through inhibition of IkappaB kinases and nuclear factor kappaB activities. J. Biol. Chem. 274, 21495–21498 (1999).
Cook, J. L., Walker, T. A., Worthen, G. S. & Radke, J. R. Role of the E1A Rb-binding domain in repression of the NF-kappa B-dependent defense against tumor necrosis factor-alpha. Proc. Natl Acad. Sci. USA 99, 9966–9971 (2002).
Janaswami, P. M., Kalvakolanu, D. V., Zhang, Y. & Sen, G. C. Transcriptional repression of interleukin-6 gene by adenoviral E1A proteins. J. Biol. Chem. 267, 24886–24891 (1992).
Radke, J. R., Figueroa, I., Routes, J. M. & Cook, J. L. PIDD-dependent activation of caspase-2-mediated mitochondrial injury in E1A-induced cellular sensitivity to macrophage nitric oxide-induced apoptosis. Cell Death Discov. 4, 35 (2018).
Cook, J. L., Ikle, D. N. & Routes, B. A. Natural killer cell ontogeny in the athymic rat. Relationship between functional maturation and acquired resistance to E1A oncogene-expressing sarcoma cells. J. Immunol. 155, 5512–5518 (1995).
Cook, J. L., Wilson, B. A., Wolf, L. A. & Walker, T. A. E1A oncogene expression level in sarcoma cells: an independent determinant of cytolytic susceptibility and tumor rejection. Oncogene 8, 625–635 (1993).
Routes, J. M., Ryan, J. C., Ryan, S. & Nakamura, M. MHC class I molecules on adenovirus E1A-expressing tumor cells inhibit NK cell killing but not NK cell-mediated tumor rejection. Int. Immunol. 13, 1301–1307 (2001).
Miura, T. A. et al. Expression of an E1A/E7 chimeric protein sensitizes tumor cells to killing by activated macrophages but not NK cells. J. Virol. 78, 4646–4654 (2004).
Lacotte, S. et al. Alloimmune activation promotes anti-cancer cytotoxicity after rat liver transplantation. PLoS ONE 9, e91515 (2014).
Cook, J. L., Routes, B. A., Walker, T. A., Colvin, K. L. & Routes, J. M. E1A oncogene induction of cellular susceptibility to killing by cytolytic lymphocytes through target cell sensitization to apoptotic injury. Exp. Cell Res. 251, 414–423 (1999).
Pardo, J. et al. The biology of cytotoxic cell granule exocytosis pathway: granzymes have evolved to induce cell death and inflammation. Microbes Infect. 11, 452–459 (2009).
Voskoboinik, I., Whisstock, J. C. & Trapani, J. A. Perforin and granzymes: function, dysfunction and human pathology. Nat. Rev. Immunol. 15, 388–400 (2015).
Waterhouse, N. J. et al. Cytotoxic T lymphocyte-induced killing in the absence of granzymes A and B is unique and distinct from both apoptosis and perforin-dependent lysis. J. Cell Biol. 173, 133–144 (2006).
Wei, M. C. et al. Proapoptotic BAX and BAK: a requisite gateway to mitochondrial dysfunction and death. Science 292, 727–730 (2001).
Thomas, D. A., Scorrano, L., Putcha, G. V., Korsmeyer, S. J. & Ley, T. J. Granzyme B can cause mitochondrial depolarization and cell death in the absence of BID, BAX, and BAK. Proc. Natl Acad. Sci. USA 98, 14985–14990 (2001).
Wang, G. Q. et al. Resistance to granzyme B-mediated cytochrome c release in Bak-deficient cells. J. Exp. Med. 194, 1325–1337 (2001).
Heibein, J. A. et al. Granzyme B-mediated cytochrome c release is regulated by the Bcl-2 family members bid and Bax. J. Exp. Med. 192, 1391–1402 (2000).
Degenhardt, K., Sundararajan, R., Lindsten, T., Thompson, C. & White, E. Bax and Bak independently promote cytochrome C release from mitochondria. J. Biol. Chem. 277, 14127–14134 (2002).
Kim, H. J., Mun, J. Y., Chun, Y. J., Choi, K. H. & Kim, M. Y. Bax-dependent apoptosis induced by ceramide in HL-60 cells. FEBS Lett. 505, 264–268 (2001).
Dejean, L. M. et al. Oligomeric Bax is a component of the putative cytochrome c release channel MAC, mitochondrial apoptosis-induced channel. Mol. Biol. Cell 16, 2424–2432 (2005).
Chipuk, J. E. et al. Sphingolipid metabolism cooperates with BAK and BAX to promote the mitochondrial pathway of apoptosis. Cell 148, 988–1000 (2012).
Zamorano, S. et al. A BAX/BAK and cyclophilin D-independent intrinsic apoptosis pathway. PLoS ONE 7, e37782 (2012).
Wai, L.-E., Garcia, J. A., Martinez, O. M. & Krams, S. M. Distinct roles for the NK cell-activating receptors in mediating interactions with dendritic cells and tumor cells. J. Immunol. 186, 222–229 (2011).
Paul, S. & Lal, G. The molecular mechanism of natural killer cells function and its importance in cancer immunotherapy. Front. Immunol. 8, 1124 (2017).
Routes, J. M. et al. Adenovirus E1A oncogene expression in tumor cells enhances killing by TNF-related apoptosis-inducing ligand (TRAIL). J. Immunol. 165, 4522–4527 (2000).
Hu, B. et al. Enhanced TRAIL sensitivity by E1A expression in human cancer and normal cell lines: inhibition by adenovirus E1B19K and E3 proteins. Biochem. Biophys. Res. Commun. 325, 1153–1162 (2004).
Bouchier-Hayes, L. & Green, D. R. Caspase-2: the orphan caspase. Cell Death Differ. 19, 51–57 (2012).
Tinel, A. & Tschopp, J. The PIDDosome, a protein complex implicated in activation of caspase-2 in response to genotoxic stress. Science 304, 843–846 (2004).
Waterhouse, N. J. et al. A central role for Bid in granzyme B-induced apoptosis. J. Biol. Chem. 280, 4476–4482 (2005).
Alimonti, J. B., Shi, L., Baijal, P. K. & Greenberg, A. H. Granzyme B induces BID-mediated cytochrome c release and mitochondrial permeability transition. J. Biol. Chem. 276, 6974–6982 (2001).
Hou, Q. et al. Granzyme H induces apoptosis of target tumor cells characterized by DNA fragmentation and Bid-dependent mitochondrial damage. Mol. Immunol. 45, 1044–1055 (2008).
Routes, J. M. & Cook, J. L. E1A gene expression induces susceptibility to killing by NK cells following immortalization but not adenovirus infection of human cells. Virology 210, 421–428 (1995).
Cook, J. L., Routes, B. A., Leu, C. Y., Walker, T. A. & Colvin, K. L. E1A oncogene-induced cellular sensitization to immune-mediated apoptosis is independent of p53 and resistant to blockade by E1B 19 kDa protein. Exp. Cell Res. 252, 199–210 (1999).
Cheng, E. H. et al. BCL-2, BCL-X(L) sequester BH3 domain-only molecules preventing BAX- and BAK-mediated mitochondrial apoptosis. Mol. Cell 8, 705–711 (2001).
Metkar, S. S. et al. Granzyme B activates procaspase-3 which signals a mitochondrial amplification loop for maximal apoptosis. J. Cell Biol. 160, 875–885 (2003).
Li, H. et al. Activation of caspase-2 in apoptosis. J. Biol. Chem. 272, 21010–21017 (1997).
Paroni, G., Henderson, C., Schneider, C. & Brancolini, C. Caspase-2-induced apoptosis is dependent on caspase-9, but its processing during UV- or tumor necrosis factor-dependent cell death requires caspase-3. J. Biol. Chem. 276, 21907–21915 (2001).
Adrain, C., Murphy, B. M. & Martin, S. J. Molecular ordering of the caspase activation cascade initiated by the cytotoxic T lymphocyte/natural killer (CTL/NK) protease granzyme B. J. Biol. Chem. 280, 4663–4673 (2005).
Bock, F. J., Peintner, L., Tanzer, M., Manzl, C. & Villunger, A. P53-induced protein with a death domain (PIDD): master of puppets? Oncogene 31, 4733–4739 (2012).
Tinel, A. et al. Regulation of PIDD auto-proteolysis and activity by the molecular chaperone Hsp90. Cell Death Differ. 18, 506–515 (2011).
Bouchier-Hayes, L. et al. Characterization of cytoplasmic caspase-2 activation by induced proximity. Mol. Cell 35, 830–840 (2009).
Bredemeyer, A. J., Carrigan, P. E., Fehniger, T. A., Smith, D. F. & Ley, T. J. Hop cleavage and function in granzyme B-induced apoptosis. J. Biol. Chem. 281, 37130–37141 (2006).
Hostetter, D. R., Loeb, C. R. K., Chu, F. & Craik, C. S. Hip is a pro-survival substrate of granzyme B. J. Biol. Chem. 282, 27865–27874 (2007).
Loeb, C. R. K., Harris, J. L. & Craik, C. S. Granzyme B proteolyzes receptors important to proliferation and survival, tipping the balance toward apoptosis. J. Biol. Chem. 281, 28326–28335 (2006).
Simon, M. C. et al. Selective induction of human heat shock gene transcription by the adenovirus E1A gene products, including the 12S E1A product. Mol. Cell. Biol. 7, 2884–2890 (1987).
Simon, M. C., Fisch, T. M., Benecke, B. J., Nevins, J. R. & Heintz, N. Definition of multiple, functionally distinct TATA elements, one of which is a target in the hsp70 promoter for E1A regulation. Cell 52, 723–729 (1988).
Mulé, J. J., Yang, J. C., Afreniere, R. L., Shu, S. Y. & Rosenberg, S. A. Identification of cellular mechanisms operational in vivo during the regression of established pulmonary metastases by the systemic administration of high-dose recombinant interleukin 2. J. Immunol. 139, 285–294 (1987).
Cook, J. L. et al. Role of tumor necrosis factor-alpha in E1A oncogene-induced susceptibility of neoplastic cells to lysis by natural killer cells and activated macrophages. J. Immunol. 142, 4527–4534 (1989).
Reynolds, C. W., Bere, E. W. J. & Ward, J. M. Natural killer activity in the rat. III. Characterization of transplantable large granular lymphocyte (LGL) leukemias in the F344 rat. J. Immunol. 132, 534–540 (1984).
Cook, J. L., Hibbs, J. B. J. & Lewis, A. M. J. DNA virus-transformed hamster cell-host effector cell interactions: level of resistance to cytolysis correlated with tumorigenicity. Int. J. Cancer 30, 795–803 (1982).
Krantz, C. K., Routes, B. A., Quinlan, M. P. & Cook, J. L. E1A second exon requirements for induction of target cell susceptibility to lysis by natural killer cells: implications for the mechanism of action. Virology 217, 23–32 (1996).
Acknowledgements
We thank Eileen White for the E1A-transformed Bak−/−, Bax−/− and Bak/Bax−/− MEF cells. We thank Zeba K. Siddiqui and Kristin R. Morris for technical support and Dennis Stevens for critical reading of the manuscript. The views expressed here are those of the authors and do not reflect the opinion or policy of the VA or the Government of the United States.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by A. Rufini
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Radke, J.R., Routes, J.M. & Cook, J.L. E1A oncogene induced sensitization to NK cell induced apoptosis requires PIDD and Caspase-2. Cell Death Discov. 5, 110 (2019). https://doi.org/10.1038/s41420-019-0189-z
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41420-019-0189-z