Abstract
Properly responding to DNA damage is vital for eukaryotic cells, including the induction of DNA repair, growth arrest and, as a last resort to prevent neoplastic transformation, cell death. Besides being crucial for ensuring homeostasis, the same pathways and mechanisms are at the basis of chemoradiotherapy in cancer treatment, which involves therapeutic induction of DNA damage by chemical or physical (radiological) measures. Apart from typical DNA damage response mediators, the relevance of cell-intrinsic antiviral signaling pathways in response to DNA breaks has recently emerged. Originally known for combatting viruses via expression of antiviral factors including interferons (IFNs) and establishing of an antiviral state, RIG-I-like receptors (RLRs) were found to be critical for adequate induction of cell death upon the introduction of DNA double-strand breaks. We here show that presence of IRF3 is crucial in this process, most likely through direct activation of pro-apoptotic factors rather than transcriptional induction of canonical downstream components, such as IFNs. Investigating genes reported to be involved in both DNA damage response and antiviral signaling, we demonstrate that IRF1 is an obligatory factor for DNA damage-induced cell death. Interestingly, its regulation does not require activation of RLR signaling, but rather sensing of DNA double-strand breaks by ATM and ATR. Hence, even though independently regulated, both RLR signaling and IRF1 are essential for full-fledged induction/execution of DNA damage-mediated cell death programs. Our results not only support more broadly developing IRF1 as a biomarker predictive for the effectiveness of chemoradiotherapy, but also suggest investigating a combined pharmacological stimulation of RLR and IRF1 signaling as a potential adjuvant regimen in tumor therapy.
Similar content being viewed by others
Introduction
DNA damage is a ubiquitous and existential threat to organisms. Potential causes comprise ionizing radiation (IR), genotoxic chemicals, but also cell-intrinsic mechanisms. Among various possible DNA alterations, the most drastic and impactful are DNA double-strand breaks (DSBs). Complex mechanisms involving detection by ATM, ATR, and downstream processes including the tumor suppressor p53 and checkpoint inhibition, either lead to sufficient repair of the damage or to induction of programmed cell death [1, 2]. The latter mostly comprises apoptosis, but other forms such as necroptosis and pyroptosis have recently been reported as well. Mutations of the central DSB sensors can cause severe diseases such as ataxia telangiectasia, associated with carcinogenesis and serious immunodeficiency [3,4,5]. Originally discovered and best-studied in the context of the antiviral innate immune response, IRF1 has been implicated in the DNA damage response and tumor suppressor functions [6,7,8,9].
Following the IRF1 example, it became apparent that cell-intrinsic antiviral signaling pathways also substantially contribute to DNA damage-induced cell death. Both STING and RIG-I-like receptor (RLR) pathways detect damage-associated molecular patterns (DAMPs), such as endogenous DNA fragments and nuclear RNA, and can trigger cell death [10, 11]. Previously, RIG-I stimulation has been shown to induce death of breast cancer cells, putting forward a potential application in tumor therapy [12]. Typically, the RLRs, RIG-I, and MDA5, are stimulated by non-self RNA in the event of viral infection. Interaction with their adaptor MAVS leads to activation of the transcription factors IRF3, NF-κB p65/RELA and p50/NFKB1. The resulting expression of IFN-stimulated genes (ISGs) and IFNs of type I/III causes the establishment of an antiviral state and, in most cases, effective containment of the invading pathogen. In addition to apoptosis sensitizing effects of NF-κB and IFNs through expression of pro-apoptotic factors, direct cell death mediating effects have recently been reported for MAVS and IRF3 [13, 14]. Chattopadhyay et al. were first to identify and characterize the RLR-induced IRF3-mediated pathway of apoptosis (RIPA) [15]. Stimulation of RLRs with dsRNA or viral infection induces MAVS-dependent ubiquitination of IRF3 and subsequent activation of pro-apoptotic factors independent of IRF3’s transcriptional activity [16]. Furthermore, MAVS was shown to directly interact with procaspase-8, forming so-called MAVS-death-inducing signaling complexes upon viral infection [17].
Here we show that RLR signaling, IRF1, and canonical DNA damage response pathways, comprising ATM/ATR and p53, are involved in efficient triggering of cell death upon DNA damage. We show that these pathways have independent pro-apoptotic capacities, and we present new insights into IRF1’s complex cellular functions.
Methods
Cell culture, cell line generation, and stimulation
Cell lines were grown at 37 °C, 95% humidity, and 5% CO2 in Dulbecco’s modified eagle medium (DMEM high glucose, Life Technologies, Carlsbad, CA, USA), supplemented with final 10% (v/v) fetal calf serum (FCS, Thermo Fisher Scientific, Waltham, MA, USA), 1x non-essential amino acids (Thermo Fisher Scientific), and 100 U/ml penicillin and 100 ng/ml streptomycin (LifeTechnologies). For generation of transgene expressing A549 cell lines by lentiviral transduction, lentiviral particles were produced by transfecting HEK 293T cells with plasmids pCMV-dr8.91, pMD2.G, and the respective retroviral vector (pWPI) using calcium phosphate transfection (CalPhos Mammalian Transfection Kit, Takara Bio Europe, Saint-Germain-en-Laye, France). After 2 days the supernatant was harvested, sterile filtered, and used to transduce target cells two times for 24 h. Transduced cells were selected with antibiotics appropriate for the encoded resistance gene (5 μg/ml blasticidin, MP Biomedicals, Santa Ana, CA, USA; 1 μg/ml puromycin, Sigma Aldrich; 1 mg/ml geneticin (G418), Santa Cruz, Dallas, TX, USA). Knockout (KO) cell lines were generated by clustered regularly interspaced short palindromic repeats (CRISPR)/Cas9 technology. DNA oligonucleotides coding for guideRNAs against the respective genes (sequences shown in Supplementary Table S1) were cloned into the expression vector LentiCRISPRv2 (Feng Zhang, Addgene #52961).
Transduced A549 wild-type cells were selected with puromycin, single-cell clones were isolated, and KO was validated by immunoblotting and functional tests (Fig. S5). We have previously generated, validated and published the following A549-based cell lines: IFNAR1−/− IFNLR1−/− IFNGR−/− (IFNR TKO) [18], IRF1−/− [18], IRF1 OE [18], IRF3−/− [19], MAVS−/− [20], MYD88−/− [20], RELA−/− [18] and RIG-I−/− [21]. A549 RIG-I OE cells and cells stably expressing IRF3-eGFP or histone H2B-mCherry were generated by stable lentiviral transduction as described previously [22]. Cells transduced with non-targeting gRNA (sequence taken from the GeCKO CRISPR v2 library) were used as controls. PH5CH non-neoplastic hepatocytes and HepG2 cells were kindly provided by Dr. Volker Lohmann (Heidelberg University, Heidelberg, Germany). Huh7.5 cells were generously provided by Dr. Charles Rice (Rockefeller University, New York). A549, HepG2 and Huh7.5 were authenticated by SNP typing (Multiplexion, Germany). Cell lines are regularly tested to be free of mycoplasma.
Stimulation was performed with doxorubicin (DOX, Hölzel Diagnostika, Cologne, Germany), etoposide (ETO, Cell Signaling Technology, Danvers, MA, USA), or cells were transfected with in vitro transcribed and chromatographically purified 200 bp 5’ppp-dsRNA [23], poly(C) (Sigma-Aldrich), and poly(I:C) (Sigma-Aldrich) using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA) following the manufacturer’s protocol. Cells were γ-irradiated with doses of 0–30 Gy using a Gammacell 40 Exactor (Best Theratronics, Ottawa, Canada).
Real-time imaging of cell death
A549 cells stably expressing histone H2B mCherry [19] were seeded at density of 2 × 103 cells per 96-well. The next day, cells were stimulated with 1–2 µM DOX (10 h), 25 µM ETO (10 h), 0.1 ng/ml dsRNA (8 h), or γ-IR. DMSO (Carl Roth, Karlsruhe, Germany), poly(C) [23], and mock irradiation were used as appropriate controls. Post treatment, fresh medium was supplemented with 1:10 000 IncuCyte® Cytotox Green Reagent (Sartorius, Göttingen, Germany) to determine dead cells. Total cell number and dead cells were monitored every 2 h using a 10× magnification in an IncuCyte® S3 Live-Cell Analysis System (Sartorius). For IFN pre-stimulation, 200 IU/ml IFN-β (IFN-β1, Bioferon, Laupheim, Germany) or IFN-γ (R&D Systems, Minneapolis, MN, USA) were added at the time of seeding. For inhibitor administration, 40 µM Z-VAD-FMK (Z-VAD, R&D Systems) and 10 µM Necrostatin-7 (Nec-7, Sigma Aldrich), or 25 µM TPCA-1 (Sigma Aldrich) were added 2 h prior treatment. IncuCyte® Software (2019B Rev2, Sartorius) was used to mask cells in phase contrast images. Calculations were performed applying the following settings: red fluorescence: segmentation top-hat, radius 100 µM, threshold (GCU) 0.4, edge split sensitivity −35, area 60–1000 µm2, integrated intensity ≥60; green fluorescence: segmentation top-hat, radius 100 µM, threshold (GCU) 10, edge split sensitivity −40, area 100–700 µm2, eccentricity ≤0.8, mean intensity 7–1000, and integrated intensity ≥2500. Percentage of dead cells was calculated relative to total cell count. Data represent the results of at least three biologically independent experiments. For curve charts, results were normalized to the control cell line of each replicate. Bars represent non-normalized means 36 h post treatment.
Immunofluorescence microscopy and determination of cellular IRF3 distribution
Fluorescence microscopy was performed to visualize phosphorylated histone H2A.X. After 4 h treatment with 2 µM DOX or DMSO, or 1 h post γ-IR with 20 Gy or 0 Gy, cells were permeabilized with −20 °C methanol and fixed with 4% paraformaldehyde. To block non-specific background, cells were incubated with 1 % (w/v) bovine serum albumin (BSA) and 10 % (v/v) FCS for 30 min. Primary antibodies specific for phospho-H2A.X (Cell Signaling Technology, 9718, 1:1000) were applied at 4 °C over-night. Slides were incubated with Alexa Fluor® 488 anti-rabbit (ThermoFisher Scientific, A11008, 1:1000) and DAPI (ThermoFisher Scientific, D1306, 1:5000) for 1 h. For determination of cellular IRF3 distribution, A549 cells stably expressing IRF3-eGFP and histone H2B-mCherry were stimulated either with DOX or poly(I:C) for 12 h. Fluorescence was visualized using a Primovert microscope (Carl Zeiss, Jena, Germany).
Immunoblotting
Stimulated cells were lysed in Laemmli sample buffer, and digested with Benzonase® Nuclease (Merck Millipore, Burlington, MA, USA). For inhibitor administration, 20 µM KU-55933 (Sigma-Aldrich), 25 µM Rabusertib (Hölzel Diagnostika), 25 µM TPCA-1 (Sigma Aldrich), or 10 µM VE-822 (Hölzel Diagnostika) were added 2 h prior treatment. For stimulation with IFNs, 200 IU/ml IFN-α (PBL Assay Science, Piscataway, NJ, USA), IFN-β, or IFN-γ were applied over-night. Lysed samples were further denatured at 95 °C for 5 min and cleared from detritus. Resulting protein extracts were subjected to 10 % (w/v) SDS-polyacrylamide gel electrophoresis and transferred to PVDF membranes (Bio-Rad, Hercules, CA, USA, 0.2 µm pore size). Upon incubation with 5% (w/v) BSA for 2 h to block non-specific background, membranes were probed using antibodies specific for β-actin (Sigma-Aldrich, A5441, 1:5000), calnexin (Enzo Biochem, Farmingdale, NY, USA, ADI-SPA-865-F, 1:1000), CASP3 (Cell Signaling Technology, 9662 S, 1:1000), CASP9 (Cell Signaling Technology, 9508, 1:1000), IRF1 (Cell Signaling Technology, 8478 S, 1:1000), phospho-IRF3 (pS396, ThermoFisher Scientific, MA5-14947, 1:1000), JAK1 (Cell Signaling Technology, 3332 S, 1:1000), MDA5 (Enzo Biochem, ALX-210-935, 1:1000), NFKB1 (p50) (Abcam, Cambridge, UK, ab32360, 1:1000), p53 (Santa Cruz Biotechnology, Dallas, TX, USA, sc-126, 1:1000), or STAT1 (BD Biosciences, Franklin Lakes, NJ, USA, 610115, 1:1000) at 4 °C over-night. For detection, anti-rabbit horseradish peroxidase (HRP) (Sigma-Aldrich, A6154-5X1ML, 1:20 000) or anti-mouse HRP (Sigma-Aldrich, A4416-5X1ML, 1:10,000) were applied for 1 h, membranes were covered with Amersham ECL Prime Western Blotting Detection Reagent (ThermoFisher Scientific) for 1 min, and luminescence was detected using a sensitive CCD camera system (ECL ChemoCam Imager 3.2, INTAS Science Imaging Instruments, Göttingen, Germany). Full, uncropped/unedited images of the main figure western blots are shown in Fig. S9.
Densitometric analysis of the protein bands was performed using ImageJ (1.52e). Data shown represent the results of at least three biologically independent experiments.
Quantitative PCR with reverse transcription
Upon stimulation, cells were lysed and total RNA was isolated with the Monarch RNA isolation kit (New England Biolabs, Ipswich, MA, USA), following the manufacturer’s protocol. After extraction, complementary DNA (cDNA) was generated using the High Capacity cDNA Reverse Transcription kit (ThermoFisher Scientific). Determination of messenger RNA (mRNA) expression was performed using iTaq Universal SYBR® Green Supermix (Bio-Rad) on a CFX96 real-time-system (Bio-Rad). Sequences of specific exon-spanning PCR primers are shown in Supplementary Table S2. GAPDH mRNA was used as a housekeeping gene control and relative expression determined by 2ΔCt (thus, not normalizing to reference condition).
Cell viability
A549 cells were seeded at a density of 6 × 103 cells per 96-well. Upon treatment with 2 µM DOX or DMSO for 24 h, cell viability was determined using the CellTiter-Glo® luminescent cell viability assay (Promega, Madison, WI, USA) following the manufacturer’s protocol. Luciferase activity was measured using a Mithras LB 943 multimode reader (Berthold Technologies, Bad Wildbad, Germany).
Caspase activity
A549 cells were seeded at density of 6 × 103 cells per 96-well. In all, 48 h post treatment with 0–2 µM DOX for 10 h, caspase-3/7 activity was determined using the Apo-ONE® homogeneous caspase-3/7 assay (Promega) following the manufacturer’s instructions. Resulting fluorescence was measured using the Mithras LB 943 multimode reader (Berthold Technologies).
Statistical analysis and reproducibility
Sample size was chosen in accordance with the general standards in the field; no statistical method was used to predetermine the sample size. If not stated otherwise, all experiments were done in three biologically independent repetitions (n = 3), with multiple technical replicates (≥3) per experiment; experiments were not randomized, researchers were not blinded. Individual experiments were excluded only in case of technical failure. Shown are mean and standard deviation (error bars) of the replicate means. For all datasets that underwent statistical comparison, normality of the data could be assumed and variance was similar between the tested samples. Hence, a paired, two-tailed Student’s t-test was employed; *p ≤ 0.05, **p ≤ 0.01, ***p ≤ 0.001, ****p ≤ 0.0001.
Results
Apoptosis induction via DNA damage response pathway in A549 cells
To investigate the molecular links between DNA damage-induced cell death and innate immune signaling, we used A549 human lung carcinoma cells, as they are immunocompetent with intact virus sensing and IFN pathways [18]. Furthermore, we previously generated numerous functional knockouts (KOs) of components of the antiviral system (see methods) and could successfully generate KOs of further components of the antiviral and DNA damage pathways. Cells were treated with DNA DSB inducers, specifically γ-IR or the topoisomerase II inhibitors doxorubicin (DOX) and etoposide (ETO), and the resulting cell death was monitored on single-cell level by real-time imaging.
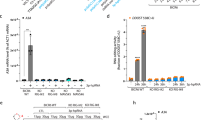
Treatment of A549 cells with DOX resulted in pronounced cell death (Fig. 1A) and a corresponding reduction of bulk cell viability (Fig. 1B), accompanied by the detection of the DNA damage marker phospho-histone H2A.X by immunofluorescence (Fig. 1C). As in DMSO control conditions no cell death was observed (Fig. 1A), for the clarity of presentation we omitted this control in the following figures but provide the control data in the supplements. In order to characterize the type of cell death predominant upon DOX-induced DNA damage, we first evaluated activation of caspase-3 and -7 being pivotal markers of apoptosis. DOX treatment activated caspase-3 and -7 in a dose-dependent manner (Fig. 1D). Conversely, we treated cells with the pan-caspase inhibitor Z-VAD in combination with Nec-7 to prevent spill-over to necroptosis [24], or depleted caspase-3 or -9. Both approaches resulted in a significant reduction of cell death upon DOX treatment (Fig. 1E, F, H). These findings confirmed prior reports that cell death driven by DOX is mainly due to apoptosis [25]. Next, we investigated typical components of the DNA damage response upstream of caspase activation. In line with p53’s (TP53) essential role in inducing apoptosis, depletion of p53 showed a significant reduction of cell death (Fig. 1G, H). Interestingly, TP53−/− had the opposite effects at late time points, elevating cell death for time points >54 h (Fig. 1G). Amongst others, p53 induces apoptosis via activation of PUMA and NOXA. Accordingly, we found PUMA and NOXA transcript levels to be increased in DOX-treated cells (Fig. 1I), supporting a canonical DNA damage response through p53 in DOX-treated A549 cells.
A Percentage of dead A549 cells relative to total cells counted over time post DOX or DMSO treatment. B Cell viability of A549 cells post DOX treatment for 24 h. C Immunofluorescence of phosphorylated histone H2A.X (S139) (cyan) and DAPI-stained nuclei (magenta) in A549 cells post DOX treatment for 4 h. See Fig. S1A for wider image. D Caspase-3/7 activity of A549 cells 24 h post DOX treatment for 10 h. E–H Percentage of dead A549 cells with caspase (and necroptosis) inhibition or functional KO of the indicated genes relative to total cells counted over time (E–G) or 36 h (H) post DOX treatment. See Fig. S1B–D for DMSO controls. I A549 cells were treated with 1 µM DOX or DMSO for 24 h. PUMA and NOXA mRNA transcripts were determined by qRT-PCR. A, B, D–I Data shown represent the results of at least three biologically independent experiments.
Relevance of innate antiviral immunity pathways in DNA damage-induced cell death
In order to investigate the contribution of antiviral signaling cascades to the induction of DSB-induced cell death, we compared the impact of the major antiviral pathways using KOs of their respective signaling adapters. We observed DOX-induced cell death to be significantly reduced only by MAVS depletion (RLR signaling), but not so in the absence of STING (cGAS signaling), TRIF (TLR3 signaling), or MYD88 (general TLR signaling) (Fig. 2A–C). Despite RLR signaling appeared to play a major role, neither canonical IRF3 phosphorylation nor its nuclear translocation could be detected (Fig. 2D, E). Consistently, there was also no characteristic RLR-mediated induction of ISGs, such as IFIT1 (Fig. 2F).
A–C Percentage of dead A549 cells with functional KO of the indicated genes relative to total cells counted over time (A, B) or 36 h (C) post DOX treatment. See Fig. S2 for DMSO controls. D A549 were stimulated with 1 µM DOX or 1 ng/ml dsRNA for 8 h. Phosphorylated IRF3 (S396) was determined by western blot. Two replicate lanes per condition: A549 wild-type cells (left lanes) and A549 cells with a KO of an irrelevant gene (IRF1, right lanes). Uncropped image of western blot shown in Fig. S9A. E A549 cells were stimulated with 1 µM DOX or 2 µg/ml poly(I:C) for 12 h. Cellular distribution of IRF3 eGFP (cyan) and histone H2B (magenta) was visualized by immunofluorescence microscopy. F A549 cells were stimulated with 1 µM DOX or 10 ng/ml dsRNA for 24 h. IFIT1 mRNA transcripts were determined by qRT-PCR. A–C, F Data shown represent the results of at least three biologically independent experiments.
Given the observed relevance of MAVS in DOX-induced cell death, we further analysed the effect of specific RLR depletion. Both RIG-I−/− and MDA5−/− reduced cell death upon DOX treatment, however, RIG-I exhibited a considerably stronger effect (Fig. 3A, C). Reciprocally, RIG-I overexpression (OE) markedly increased cell death upon DOX treatment (but not in untreated conditions, compare Fig. S3B), underlining the decisive role of RLR signaling in this process (Fig. 3B, C). In order to determine the factors responsible for mediating cell death downstream of MAVS, we further examined the influence of transcription factors IRF3 and NF-κB p65/RELA. We observed that depletion of either factor significantly reduced DOX-induced cell death (Fig. 3D, F), what would be in line with a putative role for type I/III IFNs, whose transcription is co-dependent on both. However, using IFN-“blind” A549 IFNAR1−/− IFNLR1−/− IFNGR−/− (IFNR TKO) cells, we demonstrated that this effect was independent of a response mediated by secreted IFNs (Fig. 3E, F), which was further confirmed using STAT1−/− cells (Fig. S3E). This was in accordance with the lack of ISG expression observed previously (Fig. 2F). Thus, IRF3, and possibly NF-κB, appear to have death sensitizing effects distinct from their classical transcriptional activity in the antiviral program.
A–F Percentage of dead A549 cells with functional KO or OE of the indicated genes relative to total cells counted over time (A, B, D, E) or 36 h (C, F) post DOX treatment. See Fig. S3A–D for DMSO controls. Data shown represent the results of at least three biologically independent experiments.
Taken together, we demonstrated that RLR signaling strongly contributes to the induction of cell death after DNA damage and that this function is independent of IFN secretion and the induction of canonical ISGs.
Role of IRF1 in DNA damage-induced cell death
Another transcription factor of the IRF family important for antiviral defenses [6, 18], IRF1, has previously also been implicated with the DNA damage response [26]. We hypothesized that upon genotoxic insult, IRF1 might be a downstream target of the RLR/IRF3 pathway, as reported for virus infection [18], and thereby link RLR activity to the DNA damage response. Indeed, upon DOX treatment, we observed IRF1 upregulation at the mRNA (Fig. 4A) and protein level (Figs. 4B and S4A). Of note, IRF1 induction occurred independently of the presence of p53 (Figs. 4B and S4A). In order to determine the relevance of IRF1 to cell death, we next tested IRF1−/− cells in DOX treatment. Strikingly, IRF1 depletion almost completely abolished DOX-induced cell death (Fig. 4D, G). Conversely, increasing IRF1 abundance, either by OE through stable transduction or by pre-stimulation of cells with IFN-β or IFN-γ, markedly increased cell death upon DOX treatment (Fig. 4D, E, G), and the percentage of dead cells correlated with IRF1 levels in western blot (Figs. 4C and S4B, C). Notably, neither IFN stimulation alone, nor DOX treatment in IFN-primed but IRF1-depleted cells did induce cell death (Fig. S4E–G). Surprisingly, the same phenotype was observed in RIG-I−/− conditions (Fig. S4H), in which IRF1 was present, suggesting an independent requirement on both, RLR signaling and IRF1 induction, for proper triggering and/or execution of cell death. Similar observations were also made after ETO treatment (Fig. S5A, C, D), ruling out DOX-specific effects.
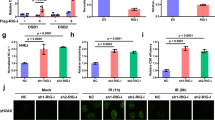
A A549 cells were treated with 1 µM DOX or DMSO for the indicated durations. IRF1 mRNA transcripts were determined by qRT-PCR. B A549 cells or A549 TP53−/− were treated with 1 µM DOX or DMSO for 10 h. Levels of IRF1 were determined by western blot. Uncropped image of western blot shown in Fig. S9B. Quantification of replicate western blots in Fig. S4A. C A549 cells were mock treated or stimulated with IFN-β or IFN-γ over-night. Levels of IRF1 were determined by western blot. Uncropped image of western blot shown in Fig. S9C. Data with additional DOX-treatment in Fig. S4B. D–G Percentage of dead A549 cells with functional KO or OE of IRF1, or post IFN pre-stimulation relative to total cells counted over time (D–F) or 36 h (G) post DOX or γ-IR (20 Gy) treatment. DMSO controls shown in Fig. S4D, E, G. H Percentage of dead cells upon IFN-γ treatment in absence or presence of γ-IR (20 Gy) at 36 h. I A549 cells were γ-irradiated. After 10 h IRF1 protein levels were determined by western blot. Uncropped image of western blot shown in Fig. S9D. A, D–H Data shown represent the results of at least three biologically independent experiments.
The fundamental importance of IRF1 was additionally demonstrated in response to γ-IR. Although irradiation did induce DNA damage in A549 cells (Fig. S5F, G), we could neither observe induction of IRF1 expression nor any cell death upon administration of up to 30 Gy (Fig. 4F, G, I). Strikingly, induction of cell death upon γ-IR was restored under conditions of elevated IRF1 levels, such as stable OE or IFN-γ pre-stimulation (Fig. 4F–H). In line with this, cells in which γ-IR naturally leads to an upregulation of IRF1 expression, such as PH5CH cells, did exhibit a dose-dependent induction of cell death (Fig. S5H, I).
Thus, we showed that besides clear involvement of p53 and RLR signaling, IRF1 is essential for proper triggering of cell death upon DNA damage. IFNs, in particular IFN-γ, sensitize cells for DNA damage-induced cell death through upregulation of IRF1.
Regulation of IRF1 expression upon DNA damage
Above we have shown that RLR/IRF3 signaling as well as expression of IRF1 are important for DNA damage-induced cell death. Furthermore, across all conditions of DNA damage-induced cell death that we tested for IRF1 expression, we found the transcription factor to be upregulated. We now aimed to confirm whether IRF1 is in fact induced as a downstream target of RLR signaling. We first investigated the induction of IRF1 expression after RIG-I stimulation using dsRNA as a canonical, highly specific agonist [23]. Indeed, we observed a fully RLR-dependent (RIG-I, MAVS, IRF3) increase of IRF1 levels, with a partial contribution of p65/RELA and IFN signaling (IFNR TKO) (Fig. 5A), in line with a recent report of our lab [18]. dsRNA-stimulation furthermore also led to the induction of cell death, which was fully abolished upon depletion of the RLR signaling components RIG-I, MAVS, or IRF3 (Fig. 5B, D). Depletion of p65/RELA and the IFN receptors (IFNR TKO) had minor pro-survival effects, suggesting a potential role for transcription-independent RIPA with a possible but limited role for IFN signaling and ISG induction (Fig. 5C, D). Interestingly and in clear contrast to the situation upon DNA damage, dsRNA-induced cell death was independent of IRF1 and, for unknown reasons, rather increased in IRF1−/− conditions (Fig. 5C). Nonetheless, experimentally elevating IRF1 levels markedly increased the percentage of dead cells also in this setting (Fig. S6E–G).
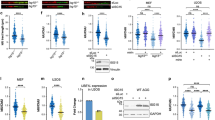
A A549 cells with functional KO of the indicated genes were mock stimulated with poly(C) or stimulated with 2 ng/ml 5’ppp-dsRNA for 6 h. IRF1 mRNA transcripts were determined by qRT-PCR; fold-induction over mock stimulation is shown. Non-normalized data shown in Fig. S6A + B. B–D Percentage of dead A549 cells with functional KO of the indicated genes relative to total cells counted over time (B, C) or 36 h (D) post dsRNA stimulation. Mock controls (poly[C] stimulation) shown in Fig. S6C + D. E–G A549 cells with functional KO of the indicated genes or administration of the indicated inhibitors were treated with 2 µM DOX or DMSO for 6 h. Levels of IRF1 were determined by western blot. Uncropped images of western blots shown in Fig. S9E–G. Quantification of replicate (n ≥ 3) western blots shown in Fig. S6H + I. A–D Data shown represent the results of at least three biologically independent experiments.
These findings confirmed that, despite not being essential for cell death induction, IRF1 is induced downstream of RLR signaling, at least when stimulated by a strong RIG-I specific agonist. We next investigated whether this would also be the case in the context of DNA damage. Unexpectedly, upon treatment of cells with DOX, induction of IRF1 expression was neither affected by depletion of RLR nor of IFN signaling components, including JAK1 (Figs. 5E, F and S6H). This suggested IRF1 expression is induced independently of and coincidentally with antiviral RLR signaling upon DNA damage. We therefore hypothesized sensing of DNA damage might directly induce IRF1. To test this, we treated cells with specific inhibitors of the prototypical DSB sensors ATM and ATR, as well as potential downstream pathways. We found IRF1 induction upon DOX-treatment to be completely blocked by the ATM inhibitor KU-55933 [27] and the ATR inhibitor VE-822 [28], suggesting important roles of these sensors in activation of IRF1 (Figs. 5G and S6I).
As IRF1 expression has previously been shown to be NF-κB sensitive [29], we employed the common pan-NF-κB and JAK1 inhibitor TPCA-1 [30, 31]. Remarkably, TPCA-1 treatment strongly diminished IRF1 expression upon DOX treatment, and even virtually completely depleted basal expression levels (Figs. 5G and S6I). This effect could further be confirmed upon RLR-stimulation with dsRNA (Fig. S7A) and even upon IFN-γ treatment, which is a strong and well-studied canonical inducer of IRF1 (Fig. S7B). We could rule out a cell line (A549) specific effect by testing three other human cell lines, PH5CH, HeLa, and Huh7.5 (Fig. S7C). To our knowledge, this striking effect of TPCA-1 on IRF1 expression has not been reported before. Again, corroborating IRF1’s crucial role in DNA damage-induced cell death, supressing IRF1 induction by TPCA-1 also reduced cell death in DOX-treated A549, PH5CH, HeLa, and Huh7.5 cells (Fig. S7D).
Finally, we aimed to identify which signaling pathway and NF-κB subunit would be responsible for IRF1 expression upon triggering the DNA damage response. As reported in literature, ATR may signal through CHK1 to activate p50/NFKB1, a potential target of TPCA-1 [32, 33]. We therefore inhibited CHK1 by Rabusertib [34] prior to DOX-treatment. However, our experiments did not reveal any effect of CHK1 inhibition or p50/NFKB1 depletion on IRF1 levels (Figs. 5G and S6I, J). We hence conclude that a so far elusive pathway downstream of the ATM/ATR system induces IRF1.
Taken together, we demonstrated that IRF1 expression upon DOX-treatment is induced by the DSB sensors ATM/ATR rather than RLR signaling. This induction is independent of CHK1 signaling. Additionally, we identified a previously unappreciated IRF1-depleting effect of the NF-κB inhibitor TPCA-1.
Discussion
Cells, particularly of multicellular organisms, have elaborate systems in place ensuring the integrity of their genome, as DNA damage poses severe risks of accumulating tumorigenic mutations or alterations. In response to excessive DNA damage beyond the potential of being properly repaired, cells trigger the execution of cell death programs, most commonly apoptosis [35]. This is also exploited for common cancer chemoradiotherapies, in which excessive DNA damage is radiologically (e.g., γ-IR) or pharmacologically (e.g., DOX or ETO) introduced, leading to the induction of cell death programs particularly in dividing tissues such as tumors. Elucidating the underlying mechanisms of how DNA damage molecularly leads to cell death is crucial to a better understanding of the circumstances leading to cancer and the pathways relevant for chemoradiotherapy. While classical DNA damage checkpoint control via p53 has been investigated thoroughly [1], much less is known about the relevance and contribution of non-canonical pathways. For example, a ground-breaking study surprisingly found the antiviral type I IFN pathway essential for certain chemotherapies’ efficacy [36]. Cytostatic and pro-apoptotic effects of IFNs have long been noticed [37,38,39]; however, it remained unresolved what triggered the production of IFNs in the studied context in the first place. Recent data also revealed cell-intrinsic triggering of cell death upon activation of antiviral signaling adapters, such as MAVS and STING. Interestingly, this was not only the case for viral infections, but also in response to DNA damage [10, 11, 40].
In the present study, we confirm this interrelationship between DNA damage response and antiviral signaling pathways, and we demonstrate a strong dependence of DOX- and ETO-triggered cell death on the presence of intact RLR/MAVS signaling. In clear contrast to recently published data, other branches of the cell-intrinsic antiviral defense, such as the TLR or the cGAS/STING system [10, 41, 42], did not affect DOX-induced cell death in our experimental setup; MyD88 KO even appeared to increase cell death (Fig. 2B). Instead, the cytosolic RNA sensors RIG-I and, to a lesser extent, MDA5 were triggered and essential for the induction of cell death. This is in line with a study by Ranoa et al. suggesting small nuclear RNAs U1 and U2 translocate into the cytoplasm in irradiated cells and trigger RIG-I activation [11]. In our experimental system, an intact RIG-I/MDA5-MAVS-IRF3 axis was essential for the full extent of cell death observed upon DNA damage; however, we could not observe canonical transcriptional activities of IRF3, such as the induction of IFN genes or ISGs. While the relevance of both IRF3 and p65/RELA suggested the involvement of IFNB expression, KO of the receptors for all three types of IFNs (IFNR TKO) did not impact cell death. A plausible mechanism for this IFN-independent triggering of apoptosis is RIPA, involving LUBAC-dependent ubiquitylation of IRF3 and subsequent activation of pro-apoptotic BH3-only proteins [16]. The clear contribution of p65/RELA in our experiments might be through its transcriptional activation of further pro-apoptotic proteins [43]. To our knowledge, cooperative effects between RIPA and NF-κB have not been described before and may be an interesting subject for future investigations.
Efficient sensing of nuclear DSBs and triggering an appropriate response is critical for cell survival upon DNA damage, or for initiating cell death and preventing potentially cancerous transformation. As expected, we observed an clear role for p53, highlighting its central function in checkpoint control, coordinating DNA damage repair and triggering apoptosis as a last resort [44]. Interestingly, depletion of p53 strongly reduced the number of apoptotic cells at early time points, but increased cell death at later times. Thus, absence of p53 led to a lack of induction of apoptosis in response to DOX-mediated DSBs at first, but likely massive accumulation of unrepaired DNA damage eventually led to increased, putatively necrotic cell death [45]. As a factor potentially linking the DNA damage response and antiviral signaling, we investigated the role of the multifunctional transcription factor IRF1, as it is known to be involved in both the DNA damage response [8, 26] and IFN signaling [6, 18, 46]. Indeed, we found that IRF1 was considerably upregulated upon DOX and ETO treatment as well as γ-IR in different cell lines. Interestingly, only in A549 cells, described to be relatively radioresistant as a common characteristic for non-small cellular lung cancers [47], IRF1 was not appreciably induced upon irradiation. We also observed reduced histone H2A.X phosphorylation after γ-IR compared to DOX treatment, but potential underlying mechanisms are only partially understood and may comprise several processes [48, 49]. Nonetheless, we could further corroborate this clear correlation between IRF1 induction and triggering/execution of a cell death program on a functional level. Experimentally increasing IRF1 levels by stable OE or by pre-treatment of cells with IFN-γ, known as a strong inducer of IRF1 [46], radioresistance of A549 cells could be overcome. A similar effect has previously been demonstrated in T cells [26]. In our experiments, increased IRF1 expression also led to a sensitization towards DOX-treatment. Vice versa, IRF1 KO almost completely rescued cell survival upon DOX-, ETO-, and γ-IR-induced DNA damage. These observations clearly establish a fundamentally important role of IRF1 in DNA damage-induced cell death. This is in accordance with literature suggesting IRF1 as a biomarker for radioresistance in tumor cells [50]. For example, extremely radioresistant osteosarcomas were shown to exhibit significantly reduced IRF1 expression levels [51]. Our data further support establishing IRF1 as a predictive biomarker in chemoradiotherapy in tumor patients.
Our finding strongly suggested IRF1 to be the functional link between the DNA damage response and the antiviral system, with RLR signaling (either directly or via the IFN/JAK/STAT cascade) leading to transcriptional activation of IRF1. However, KO experiments clearly refuted this hypothesis. Neither KO of essential factors of the RLR pathway nor of IFN signaling components abolished IRF1 induction upon DNA damage, suggesting that RLR signaling may activate IRF1 post-translationally. Generally, IRF1 is thought to be only regulated on a transcriptional level [46]. However, one study reports the requirement for “licensing” of IRF1 to become fully active, which required TLR signaling and MYD88 [52]. In preliminary experiments, we did not find any evidence for post-translational modifications in our setting, but this may warrant deeper investigations in the future. Alternatively, IRF1 might enhance the transcriptional response of IRF3, as reported before [53]. While we cannot rule out this possibility, the virtually complete inhibition of cell death in IRF1−/− despite abundant presence of IRF3 makes this unlikely. In another study, we have also not found any indication of a dampening of IRF3 responses in A549 IRF1−/− cells [18], and we see no effect of IRF1 KO on IRF3 phosphorylation (Fig. 2D, last lane). Notably, despite IRF1 being critically important for cell death induction in our system, IRF1 (over-)expression alone did not suffice to elicit cell death programs. We therefore suspect RLR signaling and IRF1 activity to cooperate further downstream, putatively via the transcriptional activation of complementary cell death promoting factors.
It is interesting to note that cell death is also elicited upon RLR stimulation by dsRNA (the canonical way to trigger antiviral signaling). Also in this case, IRF1 is induced, but strictly dependent on RIG-I and to a lesser extent dependent on IFN signaling. Surprisingly, however, depletion of IRF1 did not affect the cell death rate upon dsRNA stimulation, pointing towards transcription-independent mechanisms such as RIPA [15]. Still, KO of NF-κB (RELA) or the IFN receptors (IFNR TKO) affect cell death, suggesting some transcriptional regulation, which, however, was independent of IRF1. This may suggest that full-fledged RLR signaling upon dsRNA encounter induces a sufficiently broad transcriptional response, which (in contrast to the situation upon DNA damage) itself is capable of triggering apoptosis or other cell death pathways. Strikingly, even in dsRNA stimulation, ectopic OE of IRF1 or pre-treatment of cells with IFN-γ led to a notable increase in the number of dying cells, putatively by the same cooperative pro-apoptotic/cell death promoting effects observed in the case of DNA damage. This observation of a general sensitization for cell death by IRF1 is in line with data showing that IRF1 OE enhances apoptosis in breast or gastric cancer treatment [54,55,56]. It is further plausible to speculate that reported pro-apoptotic effects of type I IFN [57, 58] would also be mediated by upregulation of IRF1 through homodimeric STAT1 transcription factor complexes (GAF) inadvertently formed early upon IFNAR engagement [59]. This could mechanistically explain how IFN-α improved chemotherapy response and overall survival in a murine tumor model [36]. Thus, evidence further accumulates suggesting IRF1-inducing agents to be more broadly considered as adjuvants in tumor therapy.
Two central questions remain: firstly, which cell death promoting factors are specifically induced by IRF1 upon DNA damage that so potently sensitize cells to committing suicide upon (slight) RLR triggering. To this end, we are currently investigating IRF1-dependent candidate genes induced upon DOX-treatment at a transcriptomic level. Secondly, how is IRF1 induced upon DNA damage in the first place if not through classical STAT1:STAT1 activity. In our study, we found its transcriptional regulation to be fully independent of RLR signaling and p53 but completely reliant on DNA DSB sensing via ATM and ATR. Still, the downstream pathway leading to IRF1 expression remains elusive. While p65/RELA or p50/NFKB1 depletion did not affect IRF1 induction, it was almost completely abolished by TPCA-1, a commonly known inhibitor of NF-κB. Interestingly, TPCA-1 considerably reduced baseline IRF1 expression independent of the cell line used, and could even abolish the strong induction upon IFN-γ treatment. Thus, in addition to its inhibitory effects on NF-κB, JAK1, and STAT3 [30, 31, 60], TPCA-1 appears to specifically and very efficiently inhibit the activity of an essential transcription factor for IRF1.
In conclusion, our study highlights the relevance of the antiviral RLR system for the proper and timely induction of cell death upon DNA damage. We provide evidence for independent but cooperative involvement of p53, IRF1, and IRF3 activity upon detection of DNA DSBs by the ATM/ATR machinery. We show that elevating expression levels of IRF1 lead to the sensitization towards cell death across different genotoxic insults, such as chemotherapeutics, γ-IR or cytosolic dsRNA (i.e. virus infection). These data corroborate a fundamental role for IRF1 and considerable involvement of RLR signaling in DNA damage-mediated cell death and suggest future exploration of IRF1 inducers, such as IFN-γ, together with low-dose RIG-I agonists for their potential as highly efficacious adjuvants in chemoradiotherapy. Additionally, our findings support IRF1 as a biomarker predictive for chemo- and radio-sensitivity of tumors.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The raw data acquired for this study are available from the corresponding author on reasonable request.
References
Sullivan KD, Gallant-Behm CL, Henry RE, Fraikin J-L, Espinosa JM. The p53 circuit board. Bioch Biophys Acta (BBA) - Rev Cancer. 2012;1825:229–44.
Shiloh Y, Ziv Y. The ATM protein kinase: regulating the cellular response to genotoxic stress, and more. Nat Rev Mol Cell Biol. 2013;14:197–210.
Shao L, Fujii H, Colmegna I, Oishi H, Goronzy JJ, Weyand CM. Deficiency of the DNA repair enzyme ATM in rheumatoid arthritis. J Exp Med. 2009;206:1435–49.
Deng X, Ljunggren-Rose A, Maas K, Sriram S. Defective ATM-p53–mediated apoptotic pathway in multiple sclerosis. Ann Neurol. 2005;58:577–84.
Hecht F, Hecht BK. Cancer in Ataxia-telangiectasia patients. Cancer Genet Cytogenet. 1990;46:9–19.
Fujita T, Sakakibara J, Sudo Y, Miyamoto M, Kimura Y, Taniguchi T. Evidence for a nuclear factor(s), IRF-1, mediating induction and silencing properties to human IFN-beta gene regulatory elements. EMBO J. 1988;7:3397–405.
Yamane D, Feng H, Rivera-Serrano EE, Selitsky SR, Hirai-Yuki A, Das A, et al. Basal expression of interferon regulatory factor 1 drives intrinsic hepatocyte resistance to multiple RNA viruses. Nat Microbiol. 2019;4:1096–104.
Harada H, Kitagawa M, Tanaka N, Yamamoto H, Harada K, Ishihara M, et al. Anti-oncogenic and oncogenic potentials of interferon regulatory factors-1 and -2. Science. 1993;259:971.
Doherty GM, Boucher L, Sorenson K, Lowney J. Interferon regulatory factor expression in human breast cancer. Ann Surg. 2001;233:623–9.
Härtlova A, Erttmann SF, Raffi FAM, Schmalz AM, Resch U, Anugula S, et al. DNA damage primes the type I interferon system via the cytosolic DNA sensor STING to promote anti-microbial innate immunity. Immunity 2015;42:332–43.
Ranoa DRE, Parekh AD, Pitroda SP, Huang X, Darga T, Wong AC, et al. Cancer therapies activate RIG-I-like receptor pathway through endogenous non-coding RNAs. Oncotarget 2016;7:26496–515.
Elion DL, Jacobson ME, Hicks DJ, Rahman B, Sanchez V, Gonzales-Ericsson PI, et al. Therapeutically active RIG-I agonist induces immunogenic tumor cell killing in breast cancers. Cancer Res. 2018;78:6183.
Besch R, Poeck H, Hohenauer T, Senft D, Häcker G, Berking C, et al. Proapoptotic signaling induced by RIG-I and MDA-5 results in type I interferon–independent apoptosis in human melanoma cells. J Clin Investig. 2009;119:2399–411.
Goubau D, Romieu-Mourez R, Solis M, Hernandez E, Mesplède T, Lin R, et al. Transcriptional re-programming of primary macrophages reveals distinct apoptotic and anti-tumoral functions of IRF-3 and IRF-7. Eur J Immunol. 2009;39:527–40.
Chattopadhyay S, Marques JT, Yamashita M, Peters KL, Smith K, Desai A, et al.Viral apoptosis is induced by IRF-3-mediated activation of Bax. EMBO J. 2010;29:1762–73.
Chattopadhyay S, Kuzmanovic T, Zhang Y, Wetzel Jaime L, Sen Ganes C. Ubiquitination of the transcription factor IRF-3 activates RIPA, the apoptotic pathway that protects mice from viral pathogenesis. Immunity 2016;44:1151–61.
El Maadidi S, Faletti L, Berg B, Wenzl C, Wieland K, Chen ZJ, et al. A novel mitochondrial MAVS/Caspase-8 platform links RNA virus–induced innate antiviral signaling to Bax/Bak-independent apoptosis. J Immunol. 2014;192:1171.
Wüst S, Schad P, Burkart S, Binder M. Comparative analysis of six IRF family members in alveolar epithelial cell-intrinsic antiviral responses. Cells 2021;10:2600.
Urban C, Welsch H, Heine K, Wust S, Haas DA, Dachert C, et al. Persistent innate immune stimulation results in IRF3-mediated but caspase-independent cytostasis. Viruses. 2020;12:635.
Krischuns T, Gunl F, Henschel L, Binder M, Willemsen J, Schloer S, et al. Phosphorylation of TRIM28 enhances the expression of IFN-beta and proinflammatory cytokines during HPAIV infection of human lung epithelial cells. Front Immunol. 2018;9:2229.
Willemsen J, Wicht O, Wolanski JC, Baur N, Bastian S, Haas DA, et al. Phosphorylation-dependent feedback inhibition of RIG-I by DAPK1 identified by kinome-wide siRNA screening. Mol Cell. 2017;65:403–15 e8.
Binder M, Kochs G, Bartenschlager R, Lohmann V. Hepatitis C virus escape from the interferon regulatory factor 3 pathway by a passive and active evasion strategy. Hepatology. 2007;46:1365–1374.
Binder M, Eberle F, Seitz S, Mücke N, Hüber CM, Kiani N, et al. Molecular mechanism of signal perception and integration by the innate immune sensor retinoic acid-inducible gene-I (RIG-I). J Biol Chem. 2011;286:27278–87.
Vandenabeele P, Vanden Berghe T, Festjens N. Caspase inhibitors promote alternative cell death pathways. Sci STKE. 2006;2006:pe44.
Wang S, Konorev EA, Kotamraju S, Joseph J, Kalivendi S, Kalyanaraman B. Doxorubicin induces apoptosis in normal and tumor cells via distinctly different mechanisms. intermediacy of H(2)O(2)- and p53-dependent pathways. J Biol Chem. 2004;279:25535–43.
Tamura T, Ishihara M, Lamphier MS, Tanaka N, Oishi I, Aizawa S, et al. An IRF-1-dependent pathway of DNA damage-induced apoptosis in mitogen-activated T lymphocytes. Nature 1995;376:596–9.
Hickson I, Zhao Y, Richardson CJ, Green SJ, Martin NMB, Orr AI, et al. Identification and characterization of a novel and specific inhibitor of the ataxia-telangiectasia mutated kinase ATM. Cancer Res. 2004;64:9152.
Fokas E, Prevo R, Pollard JR, Reaper PM, Charlton PA, Cornelissen B, et al. Targeting ATR in vivo using the novel inhibitor VE-822 results in selective sensitization of pancreatic tumors to radiation. Cell Death Dis. 2012;3:e441–e.
Tong A-J, Liu X, Thomas BJ, Lissner MM, Baker MR, Senagolage MD, et al. A stringent systems approach uncovers gene-specific mechanisms regulating inflammation. Cell 2016;165:165–79.
Cataldi M, Shah NR, Felt SA, Grdzelishvili VZ. Breaking resistance of pancreatic cancer cells to an attenuated vesicular stomatitis virus through a novel activity of IKK inhibitor TPCA-1. Virology 2015;485:340–54.
Podolin PL, Callahan JF, Bolognese BJ, Li YH, Carlson K, Davis TG, et al. Attenuation of murine collagen-induced arthritis by a novel, potent, selective small molecule inhibitor of IκB kinase 2, TPCA-1 (2-[(Aminocarbonyl)amino]-5-(4-fluorophenyl)-3-thiophenecarboxamide), occurs via reduction of proinflammatory cytokines and antigen-induced T cell proliferation. J Pharmacol Exp Ther. 2005;312:373.
Bartek J, Lukas J. Chk1 and Chk2 kinases in checkpoint control and cancer. Cancer Cell. 2003;3:421–9.
Schmitt Adam M, Crawley Clayton D, Kang S, Raleigh David R, Yu X, Wahlstrom Joshua S, et al. p50 (NF-κB1) Is an effector protein in the cytotoxic response to DNA methylation damage. Mol Cell. 2011;44:785–96.
King C, Diaz H, Barnard D, Barda D, Clawson D, Blosser W, et al. Characterization and preclinical development of LY2603618: a selective and potent Chk1 inhibitor. Investig. New Drugs. 2014;32:213–26.
Norbury CJ, Zhivotovsky B. DNA damage-induced apoptosis. Oncogene 2004;23:2797–808.
Sistigu A, Yamazaki T, Vacchelli E, Chaba K, Enot DP, Adam J, et al. Cancer cell–autonomous contribution of type I interferon signaling to the efficacy of chemotherapy. Nat Med. 2014;20:1301.
Balkwill F, Watling D, Taylor-Papadimitriou J. Inhibition by lymphoblastoid interferon of growth of cells derived from the human breast. Int J Cancer. 1978;22:258–65.
Widau RC, Parekh AD, Ranck MC, Golden DW, Kumar KA, Sood RF, et al. RIG-I–like receptor LGP2 protects tumor cells from ionizing radiation. Proc Natl Acad Sci USA. 2014;111:E484.
Chiappinelli Katherine B, Strissel Pamela L, Desrichard A, Li H, Henke C, Akman B, et al. Inhibiting DNA methylation causes an interferon response in cancer via dsRNA including endogenous retroviruses. Cell 2015;162:974–86.
Guo Q, Chen X, Chen J, Zheng G, Xie C, Wu H, et al. STING promotes senescence, apoptosis, and extracellular matrix degradation in osteoarthritis via the NF-κB signaling pathway. Cell Death Dis. 2021;12:13.
Harberts E, Fishelevich R, Liu J, Atamas SP, Gaspari AA. MyD88 mediates the decision to die by apoptosis or necroptosis after UV irradiation. Innate Immun. 2013;20:529–39.
Huang TT, Wuerzberger-Davis SM, Wu Z-H, Miyamoto S. Sequential Modification of NEMO/IKKγ by SUMO-1 and ubiquitin mediates NF-κB activation by genotoxic stress. Cell 2003;115:565–76.
Wang P, Qiu W, Dudgeon C, Liu H, Huang C, Zambetti GP, et al. PUMA is directly activated by NF-κB and contributes to TNF-α-induced apoptosis. Cell Death Differ. 2009;16:1192–202.
Gatz SA, Wiesmüller L. p53 in recombination and repair. Cell Death Differ. 2006;13:1003–16.
Boege Y, Malehmir M, Healy ME, Bettermann K, Lorentzen A, Vucur M, et al. A dual role of caspase-8 in triggering and sensing proliferation-associated DNA damage, a key determinant of liver cancer development. Cancer Cell. 2017;32:342–59.e10.
Feng H, Zhang Y-B, Gui J-F, Lemon SM, Yamane D. Interferon regulatory factor 1 (IRF1) and anti-pathogen innate immune responses. PLoS Pathog. 2021;17:e1009220.
Kim W, Youn H, Seong KM, Yang HJ, Yun YJ, Kwon T, et al. PIM1-activated PRAS40 regulates radioresistance in non-small cell lung cancer cells through interplay with FOXO3a, 14-3-3 and protein phosphatases. Radiat Res. 2011;176:539–52.
Yang HJ, Kim N, Seong KM, Youn H, Youn B. Investigation of radiation-induced transcriptome profile of radioresistant non-small cell lung cancer A549 cells using RNA-seq. PLoS ONE. 2013;8:e59319.
Pawlik A, Alibert O, Baulande S, Vaigot P, Roux DT-L. Transcriptome characterization uncovers the molecular response of hematopoietic cells to ionizing radiation. Radiat Res. 2010;175:66–82.
Zhang Q, Bing Z, Tian J, Wang X, Liu R, Li Y, et al. Integrating radiosensitive genes improves prediction of radiosensitivity or radioresistance in patients with oesophageal cancer. Oncol Lett. 2019;17:5377–88.
Jones KB, Salah Z, Del Mare S, Galasso M, Gaudio E, Nuovo GJ, et al. miRNA signatures associate with pathogenesis and progression of osteosarcoma. Cancer Res. 2012;72:1865–77.
Negishi H, Fujita Y, Yanai H, Sakaguchi S, Ouyang X, Shinohara M, et al. Evidence for licensing of IFN-γ-induced IFN regulatory factor 1 transcription factor by MyD88 in Toll-like receptor-dependent gene induction program. Proc Natl Acad Sci USA. 2006;103:15136.
Wang J, Li H, Xue B, Deng R, Huang X, Xu Y, et al. IRF1 promotes the innate immune response to viral infection by enhancing the activation of IRF3. J Virol. 2020;94:e01231–20.
Bouker KB, Skaar TC, Riggins RB, Harburger DS, Fernandez DR, Zwart A, et al. Interferon regulatory factor-1 (IRF-1) exhibits tumor suppressor activities in breast cancer associated with caspase activation and induction of apoptosis. Carcinogenesis 2005;26:1527–35.
Kim PKM, Armstrong M, Liu Y, Yan P, Bucher B, Zuckerbraun BS, et al. IRF-1 expression induces apoptosis and inhibits tumor growth in mouse mammary cancer cells in vitro and in vivo. Oncogene 2004;23:1125–35.
Tan L, Yuan J, Zhu W, Tao K, Wang G, Gao J. Interferon regulatory factor-1 suppresses DNA damage response and reverses chemotherapy resistance by downregulating the expression of RAD51 in gastric cancer. Am J Cancer Res. 2020;10:1255–70.
Thyrell L, Erickson S, Zhivotovsky B, Pokrovskaja K, Sangfelt O, Castro J, et al. Mechanisms of Interferon-alpha induced apoptosis in malignant cells. Oncogene 2002;21:1251–62.
Choi EA, Lei H, Maron DJ, Wilson JM, Barsoum J, Fraker DL, et al. Stat1-dependent induction of tumor necrosis factor-related apoptosis-inducing ligand and the cell-surface death signaling pathway by interferon β in human cancer cells. Cancer Res. 2003;63:5299.
Kok F, Rosenblatt M, Teusel M, Nizharadze T, Gonçalves Magalhães V, Dächert C, et al. Disentangling molecular mechanisms regulating sensitization of interferon alpha signal transduction. Mol Syst Biol. 2020;16:e8955–e.
Nan J, Du Y, Chen X, Bai Q, Wang Y, Zhang X, et al. TPCA-1 is a direct dual inhibitor of STAT3 and NF-κB and regresses mutant EGFR-associated human non–small cell lung cancers. Mol Cancer Ther. 2014;13:617.
Acknowledgements
We want to thank Joschka Willemsen and Leanne Strauß for having started our lab’s research line on RLR-related cell death and providing valuable preliminary data, Hendrik Welsch for help with the IncuCyte instrument and fruitful discussions, Maike Drechsler for superb technical assistance, Volker Lohmann and Charlie Rice for provision of cell lines, and Ralf Bartenschlager for providing an excellent research environment and supervising the doctoral thesis of D.Y.Z.. We are grateful to Eva Schnober, Hartmut Hengel, and the rest of the team for managing the fantastic Integrated Research Training Group “Immunovirology” as part of TRR179, supporting D.Y.Z. with a doctoral fellowship.
Funding
This research was funded by the German Research Foundation (Deutsche Forschungsgemeinschaft, DFG), project 272983813 CRC-TR 179 (TP11) and project BI1693/1-2. Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
This study was conceived and designed by M.B. and D.Y.Z., critical intellectual input was provided by V.G.M., experiments were performed by D.Y.Z. with assistance and contributions by S.W., S.S.B., and V.G.M., data was evaluated by D.Y.Z. and M.B., the manuscript was written by D.Y.Z. and M.B. and edited and approved of by all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
This research did not involve human or animal material; ethical approval was not required.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Professor Rami Aqeilan
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zander, D.Y., Burkart, S.S., Wüst, S. et al. Cooperative effects of RIG-I-like receptor signaling and IRF1 on DNA damage-induced cell death. Cell Death Dis 13, 364 (2022). https://doi.org/10.1038/s41419-022-04797-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-022-04797-7
This article is cited by
-
Exploiting RIG-I-like receptor pathway for cancer immunotherapy
Journal of Hematology & Oncology (2023)