Abstract
Diamond–Blackfan anemia (DBA) is a rare, inherited bone marrow failure syndrome, characterized by red blood cell aplasia, developmental abnormalities, and enhanced risk of malignancy. However, the underlying pathogenesis of DBA is yet to be understood. Recently, mutations in the gene encoding ribosomal protein (RP) L18 were identified in DBA patients. RPL18 is a crucial component of the ribosomal large subunit but its role in hematopoiesis remains unknown. To genetically model the ribosomal defect identified in DBA, we generated a rpl18 mutant line in zebrafish, using CRISPR/Cas9 system. Molecular characterization of this mutant line demonstrated that Rpl18 deficiency mirrored the erythroid defects of DBA, namely a lack of mature red blood cells. Rpl18 deficiency caused an increase in p53 activation and JAK2-STAT3 activity. Furthermore, we found inhibitors of JAK2 or STAT3 phosphorylation could rescue anemia in rpl18 mutants. Our research provides a new in vivo model of Rpl18 deficiency and suggests involvement of signal pathway of JAK2-STAT3 in the DBA pathogenesis.
Similar content being viewed by others
Introduction
Diamond–Blackfan anemia (DBA) is an inherited bone marrow failure syndrome characterized by erythroid hypoplasia. It typically presents with macrocytic anemia in infancy, often with normal white blood cell and platelet counts. It is associated with congenital anomalies and a high risk of developing specific cancers, such as acute myeloid leukemia, myelodysplastic syndrome, colon adenocarcinoma, and osteosarcoma1,2. Although corticosteroid therapy and bone marrow transplantation are used to treat the patients3, significant risks and complications are often associated with these therapies. Therefore, novel therapeutic approaches are needed for treating DBA.
Ribosomal protein (RP) haploinsufficiency is the major cause of DBA and at least 19 genes that affect ribosomal biogenesis have been found to be linked to DBA4. Through whole-exome sequencing in patients in DBA, several rare mutations in previously unreported RP genes were found4,5, and one such mutation is a novel nonsynonymous mutation (p.L51S) in rpl18 analysis of pre-rRNA processing in patients’ cells showed an increase in the 36S ribosomal subunit compared with controls, indicating a defect in pre-rRNA processing6. However, no in vivo, organismal level research has elucidated the function of rpl18 in embryonic development.
In this study, an rpl18−/− mutant zebrafish line was generated using the CRISPR/Cas9 system to investigate the role of rpl18 gene in hematopoiesis. Detailed analyses showed that mutation in rpl18 leads to anemia and severe morphological abnormalities in zebrafish. Inhibition of p53 partially rescued erythropoiesis in the mutant. RNA-seq analysis showed that the JAK-STAT signaling pathway was abnormally activated in rpl18 mutant zebrafish. Small molecular inhibitors of STAT3 or JAK2 phosphorylation rescued the anemic phenotype, suggesting that STAT3 and JAK2 could be therapeutic targets for DBA treatment.
Results
CRISPR/Cas9 system generates a targeted mutation of rpl18 −/−
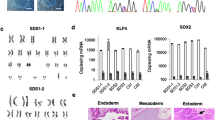
Located on zebrafish chromosome 16, rpl18 encodes a nucleolar protein, which is highly conserved between human and zebrafish with up to 85% amino acid similarity (Fig. S1). To assess whether rpl18 is involved in blood cell development in zebrafish, we used CRISPR/Cas9 system to target exon 3 of the gene and generate rpl18 mutant zebrafish lines (Fig. 1a, b). Injected embryos were raised to adulthood and mutants were screened for germline transmission. Progeny from germline positive carriers were genotyped, and mutations containing a four base pair (bp) deletion were revealed, confirming successful targeting of rpl18 in zebrafish (Fig. 1c). This mutation led to a truncated Rpl18 protein, which contained only 57 amino acids (Fig. 1d). The PCR sequencing results from wild-type (WT) and homozygous larvae are shown in Fig. S2a, b. Compared with WT larvae, homozygous mutant larvae showed a 4 bp deletion (at GCCC, red frame in Fig. S2a). To verify the reliability of the rpl18−/− mutant line, rpl18 transcripts were measured using quantitative real-time PCR (qRT-PCR) (Fig. 1e). At both 24 and 48 hours post fertilization (hpf), rpl18 transcripts were significantly downregulated compared with sibling control embryos. Transcript knockdown was further confirmed by whole-mount in situ hybridization (WISH). Strong expression of rpl18 was observed throughout control embryos at 24 and 48 hpf, however, in our mutants, hardly any rpl18 expression was observed (Fig. 1f).
a, b Illustration describing the target site of rpl18. c, d Genomic sequences and predicted protein sequences based on the DNA sequence in wild-type and rpl18−/− mutants. e qRT-PCR of rpl18 transcript levels of wild-type siblings and rpl18−/− mutants at 24 and 48 hpf. qRT-PCR experiments were performed biological repeats and technical repeats in triplicate (N = 3). ***P < 0.001. f WISH results for rpl18 in siblings and mutants at 24 and 48 hpf, WISH results were verified three times independently (N = 3). All scale bars represent 250 μm.
Rpl18 deficiency leads to morphological defects and anemia
To explore the role of rpl18 in zebrafish development, we analyzed the temporal and spatial expression of rpl18 at different stages. WISH showed that rpl18 transcripts were detected as early as the one-cell stage in WT zebrafish embryos (Fig. S3a); rpl18 continued to be expressed ubiquitously until 36 hpf (Fig. S3b–e). Then, rpl18 expression became restricted to the eye, head, and pectoral fin bud at 2 days post fertilization (dpf) (Fig. S3f). At 3 dpf, rpl18 expression was detected in the eye, head, liver, and intestine (Fig. S3g).
Prior to 22 hpf, no morphological differences were observed between rpl18 mutant embryos and WT siblings (data not shown); after which, the morphological development of the mutant embryos became gradually abnormal. At 24 hpf, aplasia was seen in the head of Rpl18-deficient embryos (Fig. 2a, b). Small eyes, small heads, and inflation of hindbrain ventricles became apparent between 30 hpf and 2 dpf (Fig. 2c–h). At 3 dpf, the pericardium gradually enlarged and developed edema; additionally, the body length decreased dramatically and the trunk displayed broad apoptosis (Fig. 2i, j). Over time, the severity of the phenotypes increased and the mutation became lethal around 4 dpf.
a–j Phenotypes of wild-type siblings and rpl18−/− mutants (N = 7). At 24 hpf (a, b), aplasia of head region (dashed box) can be seen in mutant. At 30 hpf (c, d), pigmentation was delayed. At 36 hpf (e, f) and 2 dpf (g, h), a smaller head and edema (arrowhead) were present. At 3 dpf (i, j), edema (arrowhead) in heart and shorted tail extension was more apparent. Almost all embryos died at 4 dpf. k, l o-dianisidine staining showed depletion of erythroid cells in rpl18−/− embryos compared with wild-type siblings (arrowheads pointed to the sites of weak o-dianisidine staining, N = 5). m, n rpl18 mRNA (100 pg/embryo) injected to embryos, staining of 3 dpf mutants with o-dianisidine showed a partial recovery (N = 3). All scale bars represent 250 μm.
To investigate the role of rpl18 in the development of red blood cells in this model, anemia was evaluated by o-dianisidine staining. The number of o-dianisidine positive cells was greatly reduced in Rpl18-deficient embryos compared with sibling embryos at 3 dpf (Fig. 2k, l), suggesting a role for rpl18 in the production or maintenance of mature erythroid cells in zebrafish embryos. Moreover, both the morphological defects and lack of erythrocytes were partially rescued in rpl18 mutant embryos upon co-injection of zebrafish rpl18 mRNA (100 pg/embryo) (Figs. 2m, n, and S4) but not p.L51S rpl18 mRNA (data not shown). These data indicate that the p.L51S mutation in RPL18 is likely a pathogenic mutation in patients with DBA.
Maturation but not specification of erythroid cells was affected in Rpl18-deficient embryos
To assess the effect of rpl18 disruption on erythropoiesis in mutant embryos, we performed WISH for several hematopoietic markers throughout different developmental stages. We found the expression level of gata1, one of the earliest markers for specification of erythroid lineage was distinguishable at five-somite stage (Fig. 3a)7, suggesting that the specification of erythroid lineage from hematopoietic cells was not affected. Interestingly, the expression level of gata1 was slightly increased at 24 and 30 hpf (Fig. 3a), qRT-PCR result further confirmed this phenomenon (Fig. 3b). However, the levels of Gata1 protein was reduced in rpl18−/− mutant (Fig. 3c, d), which is consistent with previous study8,9. This finding suggested that there might be some mechanism promoting the mRNA expression of gata1 but impairing the translation of Gata1 protein in Rpl18-deficient embryos. Then we investigated the expression pattern of two globin transcripts hbae1.1 and hbbe1.1. There was a similar expression level of global transcript at 24 hpf, but the expression of hbae1.1 and hbbe1.1 appeared dramatically decreased from 36 to 72 hpf (Fig. 3e, f). Meanwhile, the expression level of lyz and mpx was distinguishable at 30 and 48 hpf (Fig. S5a–e). Next, we analyzed the status of erythrocytes. Evidently, erythroid cells in rpl18 mutant arrested at the basophilic stage at this time point (Fig. 3g), suggesting the maturation of erythrocyte was impaired in rpl18 mutant. Overall, these findings demonstrated that erythropoiesis was specifically affected in rpl18 mutants and the anemia was due to a late-stage terminal maturation of erythroid cells.
a WISH showed that expression levels of gata1 in rpl18−/− embryos was slightly increased at 24 and 30 hpf (N = 3). b Relative expression of gata1 mRNA was analyzed by qRT-PCR for rpl18 siblings and mutants (N = 3). *P < 0.05. c Western blot (WB) analysis was performed to determine Gata1 protein level. WB with Gata1 antibody demonstrated its reduction in rpl18−/− mutants at 24 and 30 hpf. d The scatter diagram represented quantification of WB. Compared with siblings, the expression of Gata1 protein in mutants decreased by about 40%. The results of three groups of experiments (each experiment was repeated twice) were statistically analyzed, *P < 0.05. WISH showed that expression levels of hbae1.1 (e) and hbbe1.1 (f) in mutants were unaffected at 24 hpf, but reduced significantly from 36 hpf (N = 3). Scale bars represent 250 μm. g Wright–Giemsa staining of isolated erythroid cells collected from zebrafish embryos at 2.5 dpf. The nucleoli size of erythrocyte in rpl18 mutants was much larger than siblings (N = 4). Scale bars represent 10 μm.
Transcriptome analysis of Rpl18-deficient embryos
Previous studies have shown that RP deficiency leads to the accumulation of nonprocessed pre-rRNA and impaired ribosome biogenesis10. To better understand the mechanism of how rpl18 affects zebrafish development, we used RNA-seq to analyze global transcriptome changes in Rpl18-deficient embryos. There were 663 upregulated genes and 484 downregulated genes in rpl18 mutants compared with the control sample (fold change, 2.0; P value, 0.05) (Fig. 4a). Based on GO annotation, the differentially expressed genes were enriched in the GO terms of cellular process, single organism process, metabolic process, response to stimulus, and immune system process, among others (Fig. 4b). KEGG pathway analysis showed that the p53 signaling pathway, cell cycle, JAK-STAT signaling pathway, and ribosome biogenesis were significantly disrupted in the rpl18 mutants (Fig. 4c). These data confirm that loss of rpl18 does indeed affect ribosome-related gene expression.
a The heat map of the differential gene expression profile for rpl18−/− embryos and siblings assessed by transcriptome sequencing. b The gene ontology (GO) enrichment analysis of biological process indicated that differentially expressed genes occurred in these processes. c Pathway enrichment analysis identified the top 20 of pathway affected in mutants.
Delayed in erythroid maturation is p53 dependent
Ribosome stress has been shown to cause elevated levels of p53 and cell cycle arrest11,12,13. qRT-PCR analysis (Fig. 5a) and WISH data (Fig. 5b) showed that p53 and p53 target genes were increased in Rpl18-deficient embryos. Increased apoptosis was detected via TUNEL staining at 30 and 48 hpf throughout rpl18 mutant embryos (Fig. 5c), suggesting that morphological defects may be partially due to increased apoptosis. To determine whether the defects in rpl18 mutants were p53 dependent, we injected p53 morpholinos into Rpl18-deficient embryos. Knockdown of p53 partially rescued both the morphological defects and anemic phenotype at 3 dpf (Figs. 5d, and S6). Taken together, these findings suggest that p53-dependent pathways are upregulated and involved in the anemic phenotype in rpl18 mutant zebrafish.
a Transcript quantification of p53 and p53 related genes were verified by qRT-PCR. All the p53 related genes were highly expressed in rpl18 mutants at 30 hpf (N = 3), *P < 0.05. **P < 0.01. b WISH showed that p53, p21, and mdm2 expression was elevated in rpl18 mutants at 30 hpf (N = 3). c In situ cell death staining result showed that the signal of apoptosis can be observed thought the body at 30 and 48 hpf (N = 4). d An increase of hemoglobin level appeared in mutants at 3 dpf after p53 knockdown (N = 3). Arrowhead points to the recovery of erythrocytes. All scale bars represent 250 μm.
Increased STAT3 phosphorylation impaired erythroid cell development
Previous studies reported that overexpression of phosphorylated STAT3 has an inhibitory effect on erythroid cell maturation14,15. Our RNA-seq data showed that the JAK-STAT signaling pathway was abnormally activated in Rpl18-deficient embryos (Fig. 4c). More importantly, expression of JAK-STAT pathway-related genes, such as il6, il6st, stat3, irf9, socs3a, and socs3b, were confirmed to be elevated by qRT-PCR (Fig. S7a, b). To investigate whether this pathway and which pathway component affects the maturation of erythroid cells in our DBA model, rpl18 mutant zebrafish embryos were exposed to several small molecular inhibitors targeting different components of the JAK-STAT pathway. We found that three specific inhibitors of STAT3 phosphorylation (Figs. 6a, and S8c–f) and two specific inhibitors of JAK2 phosphorylation (Figs. 6a, and S8k, l) rescued, to various degrees, the anemic but not the morphological phenotype at 3 dpf. No rescue was observed in response to the use of other targeted inhibitors (Figs. S8g–j, and S8m–t). Western blot results further confirmed that inhibition of phosphorylated STAT3 or phosphorylated JAK2 were associated with rescue of the anemic phenotype in our rpl18 mutant DBA model (Fig. 6b, c). To further test the mechanism at a molecular level, constitutively active human STAT3 mRNA was injected into TU embryos. Upon overexpression of constitutively active STAT3 mRNA, o-dianisidine staining revealed reduced hemoglobin at 3 dpf (Fig. 6d), suggesting that overphosphorylation of STAT3 is sufficient to impair erythroid cell development. In addition, injection of dominant-negative human STAT3 mRNA into Rpl18-deficient embryos largely rescued the number of o-dianisidine-positive erythroid cells (Fig. 6e). Taken together, we conclude that deficiency of rpl18 likely induces defect in erythroid cell maturation through JAK2-STAT3 signaling (Fig. 6f).
a rpl18 sibling and mutant embryos were treated with BP (T3708, STAT3 inhibitor) or AZ (T6309, JAK2 inhibitor) (N = 5). At 3 dpf, o-dianisidine staining of mutant embryos showed obvious recovery with BP (2.5 μM) treatment, and mutants treated with AZ (0.1 μM) also be seen partially restored (arrowhead). b The total amounts of Stat3 (t-Stat3) and the phosphorylation status of endogenous Stat3 protein (p-Stat3) was evaluated in rpl18 siblings and mutants. The protein levels of t-Stat3 and p-Stat3 had a significant increase in mutants, compared with wild-type (WT) embryos at 30 hpf. Both the t-Stat3 and p-Stat3 were dramatically decreased in Rpl18-deficient embryos treated with BP or AZ separately, as shown by western blotting. c The scatter diagram represented quantification of western blots. The results of three groups of experiments (twice technical repeated for each group) were statistically analyzed, *P < 0.05. d The o-dianisidine staining displayed that most of erythroid cells disappeared (arrow) in wild-type embryos after injection the STAT3CA (constitutively active STAT3 with the double mutation A661C N663C) mRNA, compared with sibling embryos at 3 dpf (N = 3). e Erythrocytes were partially rescued in rpl18 mutants that had been injected with STAT3DN (dominant-negative STAT3) mRNA at 3 dpf (N = 3). Black arrowhead points to the recovery of erythrocytes. f Model of the defective erythroid maturation in Rpl18 deficiency through increased phosphorylated Stat3 expression. All scale bars represent 250 μm.
Discussion
DBA has been studied for several decades and, although mutations in many RPs have been reported to cause this disease, there are still some unidentified pathogenic variants in DBA patients. In this work, we successfully mutagenized a newly identified locus, rpl18, in zebrafish using CRISPR/Cas9 gene-editing technology. Rpl18 deficiency caused gross morphological defects and an anemic phenotype, which is consistent with previous reports in zebrafish. Not surprisingly, p53 and its downstream genes were elevated in rpl18 mutants. Inhibition of p53 activity using p53 morpholinos only partially rescued both the morphological defects and anemic phenotype, suggesting that other factors or pathways are likely involved in pathogenesis of rpl18-induced DBA.
Morphological and erythroid defects can be ameliorated in Rps19-deficient zebrafish embryos treated with L-leucine or exogenous nucleosides16,17, but no recognizable improvements were observed in rpl18 mutants in response to these treatments (data not shown). Danilova et al. reported that the innate immune system, including interferon and inflammatory cytokines, was activated in Rps19-deficient zebrafish and RPS19-deficient human primary cells18. The expression of related genes, such as tlr9, was also increased in Rpl18-deficient embryos (Fig. S9). We observed that the JAK-STAT pathway was significantly activated in our DBA model. Upon analyzing the RNA-seq data, this same phenomenon occurred in the rps19 mutant zebrafish DBA model19. Either inhibition of STAT3 or JAK2 phosphorylation partially rescued the anemic phenotype but not the morphological defects in rpl18 mutant zebrafish, suggesting that the role of the JAK2-STAT3 pathway is specific to the maturation of erythroid cells. Like in rpl18 mutants, restoration of hemoglobin levels was observed upon inhibition of STAT3 phosphorylation in rps19 morphant zebrafishes (Fig. S10), suggesting that overphosphorylation of STAT3 may be a phenomenon common to different genetic models of DBA.
Previous studies have shown that phosphorylated STAT3 inhibits GATA1 through a direct protein–protein interaction, and GATA1 can reverse STAT3-mediated γ-globin gene silencing in human erythroid cells; in our rpl18 mutant DBA model, a slight increase in gata1 mRNA expression but reduced levels of Gata1 protein was observed at 24 and 30 hpf. Therefore, a positive feedback mechanism between phosphorylated STAT3 and GATA1 may exist, where overphosphorylation of STAT3 promotes gata1 mRNA expression but meanwhile inhibits the translation of GATA1 protein. Further experiments are required to test these hypotheses.
Haploinsufficiency of RPs has been implicated in several bone marrow failure disorders including DBA20,21. The first animal model of DBA was generated in mouse. But unlike DBA patients, heterozygous deletion of rps19 did not have a hematopoietic phenotype, and the homozygous mouse was embryonic lethal22. The similar phenomenon was observed in rps19 mutant zebrafish and other known DBA-related knockout fish23,24. But in our rpl18-induced DBA model, we observed a noteworthy phenomenon. Generally, we should get two-thirds of heterozygous rpl18+/− offspring from in cross of two heterozygous parents, but we could only get one-fifth rpl18+/− fish (Table S1), which suggests that some heterozygous fishes could have died before we screened. In addition, the identified heterozygous fish had a shorter life span (5 months), which means heterozygous zebrafish with the same genotype (rpl18+/−) has different phenotype. This phenomenon has been noted for over five generations. In patients, why individuals with the same mutation can present with different symptoms or different severity of disease is still unknown25. Here we evaluated the erythroid production in the heterozygous mutants. Quantitative analysis of RBC mass showed that the different heterozygous adults had different and less hemoglobin concentration than WT. Moreover, Wright–Giemsa staining analysis indicated that heterozygous mutants have more immature erythrocytes compared with the WT (Fig. S11b). These results demonstrated that in zebrafish, the same mutation can also result in different severity of the disease, which mimics the symptoms of the DBA in patients.
To the best of our knowledge, this is the first demonstration that the anemic defect in a zebrafish DBA model can be rescued by inhibition of STAT3 or JAK2 phosphorylation. This finding enhances our understanding of the pathogenesis of the disease. It is worth noting that one drug targeting STAT3 that is in clinical development, ochromycinone (STA-21)26, can attenuate the anemic phenotype in our zebrafish DBA model (Fig. S12). Based on our findings, therapies targeting JAK2-STAT3 signaling should be further explored as potential treatments for DBA.
Materials and methods
Zebrafish maintenance
Zebrafish was raised and maintained under standard laboratory conditions at 28.5 °C. WT TU zebrafish were used in this study. All zebrafish experiments were approved by Institutional Animal Care and Use Committee of Peking University. All zebrafish embryos or adults used in this study were chosen randomly. Embryos or adults were genotyped after analysis.
Cas9 mRNA and gRNA synthesis
For making zCas9 mRNA, the template was linearized by XbaI (pT3TS-zCas9) digestion27 and purified using TIANquik mini purification kit (TIANGEN). Capped zCas9 mRNA was synthesized using T3 mMESSAGE mMACHINE kit (Ambion) and purified using an RNeasy FFPE kit (Qiagen). All sgRNAs (single-guide RNAs) templates in this study were prepared according to the cloning-independent sgRNA generation method28. All gRNAs were transcribed in vitro using the T7 RNA Polymerase (TaKaRa), and purified using RNeasy® FFPE Kit (Qiagen). The size and quality of resulting mRNA and sgRNAs was confirmed by electrophoresis through a 1.5% agarose gel.
Generation of the rpl18 −/− homozygous mutants
A solution (1–2 nl) containing zCas9 mRNA (200 ng/μl) and sgRNA (30 ng/μl) was co-injected into one-cell-stage zebrafish embryos. These embryos were incubated at 28.5 °C for observation of phenotypes and PCR amplification. Injected embryos were raised as F0 to adulthood. F0 adult zebrafish was outcrossed to a WT fish (Tuebingen) and offspring embryos were collected. The region flanking of the target site of offspring embryos were PCR amplified using forward primer: 5′-ATGCGTCAGTTTTGGCCAGT-3′ and reverse primer: 5′-TGTGGACGGAGAGCTTTTCA-3′. The purified PCR product was made by TA cloning using pEASY-T1 Simple Cloning Kit (TransGen) and then sequencing. Offspring of germline transmitted F0 zebrafishes were raised as F1 to adulthood. To identify germline transmitted F1 fishes, the tail of sexually mature F1 was cut off to obtain the genome, which was then genotyped and sequencing using forward primer: 5′-ACACCTCTGTGCTTTTTGTGC-3′ and reverse primer: 5′-ACATACCCTGTCTCCCTTGC-3′. Pairs of rpl18+/− F1 heterozygotes incrossed and generated the rpl18−/− homozygous mutants.
Constructs
The zebrafish rpl18 gene were amplified and subcloned into vector PCS2+ for mRNA synthesis following NotI (NEB) digestion. Human STAT3DN (a dominant-negative STAT3), STAT3CA (constitutively active STAT3 with the double mutation A661C N663C) constructs have been described previously29. The sequences of p53 MO was descried by Langherinrich et al.30.
qRT-PCR
Total RNA was isolated from whole embryos (30–35 embryos per group) at specific stages, using TRIzol® Reagent (Life Technologies), and following the manufacture’s protocol. The cDNA was reverse transcribed from 1 μg total RNA using PrimerScriptTM RT reagent kit (TaKaRa) for next real-time quantitative PCR analysis. All quantitative PCR was performed biological repeat in triplicates (each biological repeat contains three technical repeats) using TB Green® Premix Ex Taq™ (TaKaRa) and the CFX96 real-time PCR detection system (Bio-Rad). The used PCR primer sequences are listed in Table S2.
RNA-seq analysis
Two independent batches of samples were collected for RNA-seq. The first group between mutants and siblings, total RNA was isolated from whole embryos (70–80 embryos per group) at 28 hpf. The Second group, total RNA was extracted at 30 hpf. TRIzol® Reagent (Life Technologies) was used.
RNA concentration and purity were measured by nanodrop2000 (Thermo). Total RNA concentrations ≥ 200 μg/μl and the purity RIN ≥ 7, 28S/18S ≥ 1.0. The mRNA was enriched by Oligo (dT) beads, then the enriched mRNA was fragmented into short fragments (200–700 nt) and reverse synthesized into first-strand cDNA using random primers. The second-strand cDNA was performed end reparation, poly-A addition and Illumina sequencing adapters ligation following purification with QiaQuick PCR extraction kit. After the PCR amplification, these products were sequenced by Illumina HiSeqTM.
For bioinformatics analysis, raw reads containing adapter, low quality reads (Q value ≤ 20) and reads containing unknown nucleotides (more than 10%) were removed. Then the RNA removed reads mapped to the zebrafish (Danio Rerio) reference genome by TopHat2 (version 2.0.3.12)31. Differentially expressed genes were analyzed by edgeR package (fold change > 2, FDR < 0.05) in a comparison as significant DEGs31. DEGs were subjected to enrichment analysis of gene ontology and pathway. For GO analysis, all DEGs were mapped to GO terms in the Gene Ontology database, and gene numbers were calculated for every term32. For pathway enrichment, KEGG was the major public pathway-related database. The calculated P value was execute FDR correction (FDR ≤ 0.05 as a threshold)33.
WISH
Embryos of different periods were fixed in 4% paraformaldehyde (PFA) solution overnight at 4 °C. After fixation and gradient dehydration, embryos can store at −20 °C for long time. Linearized plasmids were employed as templates to synthesize digoxigenin-labeled antisense probes using the DIG RNA labeling mix (Roche) and T7 RNA Polymerase (TaKaRa). After synthesis, these probes were purified by the RNeasy® FFPE Kit (Qiagen). Finally, WISH was performed as described using probes including rpl18, gata1, hbae1.1, hbbe1.1, lyz, mpx, p53, p21, and mdm2. The stained embryos were photographed using a microscope (ZEISS, Imager.A1).
Genotype identification
For genotyping, the genome was extracted from the embryos when imaging was finished. Generally, an embryo was placed in a centrifuge tube with immersing in 10 μl of 50 mM NaOH and was incubated at 95 °C for 20 min twice. After vortexing, 1 μl of 1 M Tris-HCl (pH = 8.0) was added to the previous solution to mix well. Finally, the prepared DNA was amplified for sequencing to identify genotypes.
Protein extraction and western bolt
Thirty to thirty-five whole embryos at 30 hpf were collected and lysed in M-PER™ mammalian protein extraction reagent (Thermo) that had been mixed with protease and phosphatase inhibitors. Protein extraction experiment needs to be done on ice. Protein concentration was measured with Pierce® BCA protein assay reagent (Thermo). Ten to fifteen milligrams of total protein was used for SDS-PAGE (Tris glycine) electrophoresis, and the next procedure of western blot need to refer to protocols of different antibodies. The following antibodies for zebrafish were used in this project: GAPDH (GeneTex), GATA1 (GeneTex), STAT3 (Abcam), anti-phospho-STAT3 (Tyr708) (MBL), anti-mouse IgG HRP-linked antibody (CST), and anti-rabbit IgG HRP-linked antibody (CST). Imaging results were quantified using ImageJ software.
o-Dianisidine staining
Embryos were fixed at 4 °C using 4% PFA followed by four washes (5 min each) in 1X PBST. Then embryos were dyed using o-dianisidine staining solution (40% anhydrous ethanol, 0.65% hydrogen peroxide, 10 mM sodium acetate, and 0.6 mg/ml o-dianisidine [Sigma]) in the dark for 50 min. After incubation, embryos were washed four times (5 min each) in 1X PBST and then added into the bleach solution (1% potassium hydroxide, 3% hydrogen peroxide) until the pigmentation was removed. Embryos were then washed in 1X PBST and imaged by microscope (ZEISS, Imager.A1).
TUNEL staining
Thirty and forty-eight hours post fertilization embryos were fixed with 4% PFA overnight at 4 °C followed by five washes (5 min each) in 1X PBST. Then embryos were treated with acetone for 5 min and digested with proteinase K (Roche) (10 μg/ml) at room temperature for 15–20 min. After each of these treatments, 1X PBST washing was required for next step. TUNEL staining was performed using the in situ Cell Death Detection Kit and TMR Red (Roche). The reaction was stopped after 1.5 h dark treatment and immediately imaged. The pictures were captured by a fluorescence microscope (ZEISS, Discovery.V2.0).
Wright–Giemsa staining
Zebrafish embryos at 60 hpf were anesthetized by tricaine, then were immersed in mixture solution (40% FBS [HyClone] and a final concentration of 5 mM EDTA solution in 1X PBS). Red blood cells were collected from the heart using a microinjection needle. The cell solution was centrifuged at 1000 rpm for 5 min and the supernatant was discarded. Then the enriched cells were smeared evenly on slide and air-dried rapidly. After fixing in the methanol for 5 min, the slide was soaked in rapid Wright–Giemsa staining solution (BBI Life Sciences) for 10 min. Finally, slides were rinsed with deionized water. Images were captured with 100× lens after air dry. Assessment of erythrocyte status was based on previously published article34.
Hemoglobin quantification
Peripheral blood was collected from anaesthetized WT and rpl18+/− heterozygous zebrafish (4 months old, both male and female) by cutting the tail35. To prevent clotting, pipette tips and scissors were treated with 1% heparin solution in advance. Hemoglobin was measured with the Hemoglobin Assay Kit (Sigma-Aldrich, MAK115) according to the manufacturer’s protocol.
Inhibitor treatments
Most of inhibitors used were purchased from TargetMol, including T3708 (BP-1-102), T6309 (AZ960), T2814 (cryptotanshinone), T4234 (HJC0152 hydrochloride), T4216 (STAT5 inhibitor), T1038 (fludarabine), T1849 (momelotinib), T6122 (CEP-33779), T3620 (solcitinib), T3998 (itacitinib), and T4657 (WHI-P97). Only ochromycinone (STA-21) were purchased from Selleck. All inhibitors were dissolved in DMSO to a storage concentration of 10 mM and stored at −80 °C. It is important to avoid repeated freezing and thawing for inhibitor activity. Embryos from the same parent were divided into control and treatment groups at 6 hpf. Every treatment group was raised in 1X Holt buffer with all kinds of inhibitors and control groups was raised in in 1X Holt buffer. Embryos were observed and collected for the following experiments at 3 dpf. Each inhibitor treatment has at least three biological replications and three technical replications.
Statistics and reproducibility
Experiments were biologically repeated at least three times independently, and the number of times an experiment was repeated (N) had stated in the legend. All statistical data were calculated and graphed using GraphPad Prism 8 Software. Then, the mean and standard deviation were calculated. For two groups’ comparison, P values were calculated using two-tailed unpaired Student’s t test and less than 0.05 was considered as significant.
Data availability
All data generated or analyzed during this study are included in this article and its supplementary information files. The raw RNA-seq data was uploaded to NCBI SRA database. The SRA accession number: PRJNA597556.
References
Lipton, J. M. & Ellis, S. R. Diamond Blackfan anemia 2008-2009: broadening the scope of ribosome biogenesis disorders. Curr. Opin. Pediatr. 22, 12–19 (2010).
Vlachos, A., Rosenberg, P. S., Atsidaftos, E., Alter, B. P. & Lipton, J. M. Incidence of neoplasia in Diamond Blackfan anemia: a report from the Diamond Blackfan Anemia Registry. Blood 119, 3815–3819 (2012).
Aspesi, A., Borsotti, C. & Follenzi, A. Emerging therapeutic approaches for Diamond Blackfan anemia. Curr. Gene Ther. 18, 327–335 (2018).
Ulirsch, J. C. et al. The genetic landscape of Diamond-Blackfan anemia. Am. J. Hum. Genet. 103, 930–947 (2018).
Ikeda, F. et al. Exome sequencing identified RPS15A as a novel causative gene for Diamond-Blackfan anemia. Haematologica 102, e93–e96 (2017).
Mirabello, L. et al. Novel and known ribosomal causes of Diamond-Blackfan anaemia identified through comprehensive genomic characterisation. J. Med. Genet. 54, 417–425 (2017).
Detrich, H. W. 3rd et al. Intraembryonic hematopoietic cell migration during vertebrate development. Proc. Natl Acad. Sci. USA 92, 10713–10717 (1995).
Ludwig, L. S. et al. Altered translation of GATA1 in Diamond-Blackfan anemia. Nat. Med. 20, 748–753 (2014).
Khajuria, R. K. et al. Ribosome levels selectively regulate translation and lineage commitment in human hematopoiesis. Cell 173, 90–103 e119 (2018).
Aspesi, A. & Ellis, S. R. Rare ribosomopathies: insights into mechanisms of cancer. Nat. Rev. Cancer 19, 228–238 (2019).
Golomb, L., Volarevic, S. & Oren, M. p53 and ribosome biogenesis stress: the essentials. FEBS Lett. 588, 2571–2579 (2014).
Boultwood, J., Pellagatti, A. & Wainscoat, J. S. Haploinsufficiency of ribosomal proteins and p53 activation in anemia: Diamond-Blackfan anemia and the 5q-syndrome. Adv. Biol. Regul. 52, 196–203 (2012).
Ear, J. et al. A Zebrafish Model of 5q-Syndrome using CRISPR/Cas9 targeting RPS14 reveals a p53-independent and p53-dependent mechanism of erythroid failure. J. Genet. Genom. 43, 307–318 (2016).
Foley, H. A. et al. Stat3 beta inhibits gamma-globin gene expression in erythroid cells. J. Biol. Chem. 277, 16211–16219 (2002).
Yao, X. et al. Role of STAT3 and GATA-1 interactions in gamma-globin gene expression. Exp. Hematol. 37, 889–900 (2009).
Zhang, Y. et al. Defects of protein production in erythroid cells revealed in a zebrafish Diamond-Blackfan anemia model for mutation in RPS19. Cell Death Dis. 5, e1352 (2014).
Danilova, N. et al. The role of the DNA damage response in zebrafish and cellular models of Diamond Blackfan anemia. Dis. Model. Mech. 7, 895–905 (2014).
Danilova, N. et al. Innate immune system activation in zebrafish and cellular models of Diamond Blackfan Anemia. Sci. Rep. 8, 5165 (2018).
Jia, Q. et al. Transcriptome analysis of the zebrafish model of Diamond-Blackfan anemia from RPS19 deficiency via p53-dependent and -independent pathways. PLoS ONE 8, e71782 (2013).
Dutt, S. et al. Haploinsufficiency for ribosomal protein genes causes selective activation of p53 in human erythroid progenitor cells. Blood 117, 2567–2576 (2011).
Ito, E., Konno, Y., Toki, T. & Terui, K. Molecular pathogenesis in Diamond-Blackfan anemia. Int. J. Hematol. 92, 413–418 (2010).
Matsson, H. et al. Targeted disruption of the ribosomal protein S19 gene is lethal prior to implantation. Mol. Cell. Biol. 24, 4032–4037 (2004).
Danilova, N., Sakamoto, K. M. & Lin, S. Ribosomal protein S19 deficiency in zebrafish leads to developmental abnormalities and defective erythropoiesis through activation of p53 protein family. Blood 112, 5228–5237 (2008).
Danilova, N., Sakamoto, K. M. & Lin, S. Ribosomal protein L11 mutation in zebrafish leads to haematopoietic and metabolic defects. Br. J. Haematol. 152, 217–228 (2011).
Vlachos, A. & Muir, E. How I treat Diamond-Blackfan anemia. Blood 116, 3715–3723 (2010).
Song, H., Wang, R., Wang, S. & Lin, J. A low-molecular-weight compound discovered through virtual database screening inhibits Stat3 function in breast cancer cells. Proc. Natl Acad. Sci. USA 102, 4700–4705 (2005).
Jao, L. E., Wente, S. R. & Chen, W. Efficient multiplex biallelic zebrafish genome editing using a CRISPR nuclease system. Proc. Natl Acad. Sci. USA 110, 13904–13909 (2013).
Bassett, A. R., Tibbit, C., Ponting, C. P. & Liu, J. L. Highly efficient targeted mutagenesis of Drosophila with the CRISPR/Cas9 system. Cell Rep. 4, 220–228 (2013).
Su, F. et al. Protein tyrosine phosphatase Meg2 dephosphorylates signal transducer and activator of transcription 3 and suppresses tumor growth in breast cancer. Breast Cancer Res. 14, R38 (2012).
Langheinrich, U., Hennen, E., Stott, G. & Vacun, G. Zebrafish as a model organism for the identification and characterization of drugs and genes affecting p53 signaling. Curr. Biol. 12, 2023–2028 (2002).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578 (2012).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Kanehisa, M. et al. KEGG for linking genomes to life and the environment. Nucleic Acids Res. 36, D480–D484 (2008).
Qian, F. et al. Distinct functions for different scl isoforms in zebrafish primitive and definitive hematopoiesis. PLoS Biol. 5, e132 (2007).
Zang, L., Shimada, Y., Nishimura, Y., Tanaka, T. & Nishimura, N. Repeated blood collection for blood tests in adult zebrafish. J. Vis. Exp. 102, e53272 (2015).
Acknowledgements
We thank Liuwen Huang for zebrafish husbandry. We thank Michelle Goody, PhD, from Liwen Bianji, Edanz Editing China (www.liwenbianji.cn/ac), for editing the English text of a draft of this manuscript. This work was supported by the National Natural Science Foundation of China (31801209) and Shenzhen Science and Technology Innovation Commission (JCYJ20180302153633991).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by H.-U. Simon
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, C., Lu, M., Lin, S. et al. The nuclear gene rpl18 regulates erythroid maturation via JAK2-STAT3 signaling in zebrafish model of Diamond–Blackfan anemia. Cell Death Dis 11, 135 (2020). https://doi.org/10.1038/s41419-020-2331-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-020-2331-5
This article is cited by
-
SpG and SpRY variants expand the CRISPR toolbox for genome editing in zebrafish
Nature Communications (2022)
-
Molecular Subgroup Classification in Alzheimer’s Disease by Transcriptomic Profiles
Journal of Molecular Neuroscience (2022)