Abstract
The immunological mechanisms underlying the clinical presentation of SARS-CoV-2 infection and those influencing the disease outcome remain to be defined. Myeloid-derived suppressor cells (MDSC) have been described to be highly increased during COVID-19, however, their role remains elusive. We performed an in depth analysis of MDSC in 128 SARS-CoV-2 infected patients. Polymorphonuclear (PMN)-MDSC expanded during COVID-19, in particular in patients who required intensive care treatments, and correlated with IL-1β, IL-6, IL-8, and TNF-α plasma levels. PMN-MDSC inhibited T-cells IFN-γ production upon SARS-CoV-2 peptides stimulation, through TGF-β- and iNOS-mediated mechanisms, possibly contrasting virus elimination. Accordingly, a multivariate regression analysis found a strong association between PMN-MDSC percentage and fatal outcome of the disease. The PMN-MDSC frequency was higher in non-survivors than survivors at the admission time, followed by a decreasing trend. Interestingly, this trend was associated with IL-6 increase in non-survivors but not in survivors. In conclusion, this study indicates PMN-MDSC as a novel factor in the pathogenesis of SARS-CoV2 infection, and open up to new therapeutic options.
Similar content being viewed by others

Introduction
The ongoing COVID-19 pandemic due to the new coronavirus SARS-CoV-2 remains a global health emergency. As of 9 August 2020, there have been 19,462,112 COVID-19 cases with 722,285 deaths reported to the WHO (Ref: WHO Coronavirus Disease situation reports: https://www.who.int/emergencies/diseases/novel-coronavirus-2019/situation-reports/).
The clinical presentations of Covid-19 range from asymptomatic, mild, moderate to severe pneumonia and fulminant disease1.
The immunological mechanisms underlying the heterogeneous clinical expression of SARS-CoV-2 infection and those underlying factors influencing the clinical outcome remain to be defined. Tissue damage has been associated with excessive and uncontrolled immune activation and pro-inflammatory cytokines2,3. A massive infiltration of mononuclear cells has been detected in the lungs, with parallel low levels of hyperactive T cells in the peripheral blood4. Moreover, depletion of lymphocytes and increase of neutrophils in the peripheral blood are changes typically associated with an unfavorable disease course. In particular, it has been reported that the Lymphocyte/Neutrophil ratio is an independent risk factor of mortality for Covid-19 patients5,6,7. Notably, lymphocytes from patients with severe Covid-19 often present an exhausted phenotype8,9, and the macrophage activation syndrome has been described to occur in patients with respiratory failure10. Together, these data indicate that SARS-CoV-2 infection associated with excessive and dysregulated immune activation, and massive migration of cells to the infected tissue, to control viral replication, could contribute to tissue damage.
During evolution, immune system has developed regulatory mechanisms able to control excessive inflammation/activation, such as the induction of inhibitory receptor expression, production of specialized anti-inflammatory cytokines, and expansion of regulatory cells. However, their role during SARS-CoV-2 infection is poorly understood. We recently reported the expansion of myeloid-derived suppressor cells (MDSC) during SARS-CoV-2 infection, correlated with inflammatory milieu11,12. Whether MDSCs play a beneficial anti-inflammatory and/or a detrimental immune suppressive role during COVID-19 has not yet been elucidated.
MDSC, defined in humans as CD11b +CD14−CD33+CD15+ and HLADR-low (polymorphonuclear MDSCs), or CD11b+CD14+CD33+ and HLA-DR-low (monocytic MDSCs), are known to have the remarkable ability to reduce overt inflammation and suppress T-cell responses through several mechanisms, including inducible nitric oxide synthase (iNOS), arginase-1(Arg-1), nicotinamide adenine dinucleotide phosphate oxidase (NOX2), and transforming growth factor beta (TGF-β)13,14,15,16.
In the present work, we provide evidence that early PMN-MDSC expansion inhibits SARS-CoV-2 specific T-cell responses, and might predict fatal outcome.
Materials and methods
Sample size calculation
Sample size was calculated by using a significance level (alpha) of 0.050, and performing a one-sided two-sample equal-variance t-test. Group sample sizes of 96 (no ICU 75%) and 32 (ICU 25%) achieve 95% power to reject the null hypothesis of equal means of MDSC in the two groups, when the population mean difference is more than 15.0 (standard deviation = 22 for both groups). For the follow up analysis, group sample sizes of 59 (survivors 75%) and 19 (non survivors 25%), for a total of 78 patients achieve 90% power to reject the null hypothesis when the population mean difference is more than 17.0.
Study population
Patients with confirmed SARS-CoV-2 infection (n = 128) were treated at the National Institute for Infectious Diseases (INMI) “LazzaroSpallanzani” (Rome, Italy). All patients were symptomatic, ranging from mild to severe (requiring intensive care unit admission, n = 32). Median age was 63 years (IQR 53–75), 88 (68.8%) were males. Healthy individuals (HD, n = 30) were included as controls. Patients were recruited in the study at the time of hospital admission (within 2 weeks from symptoms onset). A group of patients (n = 78) were followed weekly, until 5 weeks after admission. At the time of manuscript preparation, 59 patients recovered and were discharged (survivors), and 19 died (non-survivors).
The study was approved by the Institutional Review Board of the INMI “LazzaroSpallanzani” (approval number: 9/2020) and signed written informed consent was obtained from all patients.
Plasma cytokine levels
Plasma samples were obtained from peripheral blood, after speed centrifugation for 10 min at 2000 rpm and immediately stored at −80 °C. IL1-β, IL-6, IL-8, TNF-α, and TGF-β were measured in plasma samples by using customized automated ELISA assays (ELLA microfluidic analyzer, Protein Simple, Bio-Techne, USA).
Peripheral blood mononuclear cells and PMN-MDSC isolation
PBMCs were isolated from heparin treated whole blood by density gradient centrifugation (Lympholyte-H, cat. CL5020; Cederlane, USA.) PBMCs were suspended in RPMI 1640 (cat. 15040- CV, Corning Incorporated, USA) supplemented with 10% heat-inactivated fetal bovine serum (FBS) (cat. EU S028877, EuroClone, Italy), penicillin/streptomycin solutions,100X (cat. 30-002-CL, Corning Incorporated, USA), 2 mmol/L L-glutamine (cat. M11-004, Corning Incorporated, USA), and 10 mmol/L HEPES buffer (N-2- hydroxyethylpiperazine-N-2-ethane sulfonic acid, cat. 15630106, Gibco, USA), hereafter termed R10.
Magnetic cell isolation technology was used to isolate the PMN-MDSC cells from PBMCs. Positive selections were performed by CD15 direct magnetic labeling (cat. 130-046-601, MiltenyiBiotec, Germany), according to the manufacturer’s procedure. Purity and recovery was >90% and were verified by flow-cytometry (data not shown).
Flow-cytometry
MDSC frequency was evaluated by staining PBMCs with customized Duraclon tubes, (CD11b FITC, HLA-DR ECD, CD14 PC5.5, CD33 PC7, CD45KrO, CD80 APC, DRAQ7, CD56 APC-alexa750, CD19 APC-alexa750, CD3 APC-alexa750, CD15 Pacific Blue, cat. B74697, Beckman Coulter), following manufacturer’s procedures. Data were acquired by CytoFlex flow-cytometer (Beckman Coulter, USA). Data were analyzed by CytExpert or Kaluza software (Beckman Coulter, USA).
Proliferation assay
PBMC were isolated from SARS-CoV-2+ patients. PBMCs and PMN-MDSC-depleted PBMCs (Depleted) were stained with CFDA-SE (Vibrant CFDA SE cell tracer kit, cat. V1288, Invitrogen, USA) following manufacturer’s procedures. PBMCs, PMN-MDSC depleted PBMCs, and PMN-MDSC depleted PBMC plus PMN-MDSC (1/4 ratio) were cultured in 48-well flat plates (500,000 cells/well in R10), stimulated with Staphylococcus enterotoxin B (SEB, 800 ng/ml, cat. S4881, Sigma Aldrich, USA) and maintained at 37 °C in humidified air with 5% CO2. After 4 days, cells were stained with anti-CD3 APC (cat. 555335, BD Biosciences, USA), and acquired by CytoFlex flow cytometer (Beckman Coulter, USA). Data were analyzed by CytExpert software (Beckman Coulter, USA).
SARS-CoV-2 specific T-cell response
SARS-CoV-2 specific T-cell response was performed by stimulating PBMC, PMN-MDSC depleted PBMCs, and Depleted PBMC plus PMN-MDSC (1/4 ratio) with a pool of Spike and Nucleocapside derived peptides (1 μg/ml, Peptide&Elephants GmbH, Germany) for 4 days. Phytohaemagglutinin (PHA, cat. 11082132001, Sigma Aldrich, USA) was used as control for T cell activity. The IFN-γ release was evaluated by using a direct, sandwich chemiluminescence immunoassay (CLIA), and analyzed by LIAISON® XL analyzer, (Diasorin). Results were reported in International Units per mL (IU/mL).
Where indicated, PBMCs, PMN-MDSC depleted PBMCs, and Depleted PBMCs plus MDSC (ratio 1:4), were treated with N(ω)-hydroxy-nor-L-arginine (0.5 mM, cat. 399275, Calbiochem, Germany), neutralizing anti-human TGFβ antibody (10 μg/ml, cat. AB 101 NA, R&D Systems, USA), or L-NG-nitro arginine methyl ester (1 mM, cat.483125, Calbiochem, Germany). The production of IFN-γ was evaluated by ELISA (Quantikine ELISA, cat. PDIF50C, R&D Systems, USA).
Quantitative PCR
Total RNA was extracted from PMN-MDSC cells using Direct-zol™ RNA MiniPrep KIT (cat. R1055, Zymo Research Corp., USA) according to the manufacturer’s instructions. For RT-PCR analyses, single-stranded cDNA was obtained by reverse transcription of 1 μg of total RNA using AMV-reverse transcriptase (cat. M5108, Promega, MI, Italia). ArgI, i-NOS and TGF-β mRNA were analyzed by quantitative PCR by using the following primers: Arginase1 F: 5′-CGCCAAGTCCAGAACCATAG-3′ R: 5′-TCCCCATAATCCTTCACATCAC-3′; iNos F: 5′-AGATAAGTGACATAAGTGACCTG-3′ R: 5′-CATTCTGCTGCTTGCTGAG-3′; TGF-β F: 5′-GACATCAACGGGTTCACTAC-3′ prime R: 5′-GTGGAGCTGAAGCAATAGTT-3′. qPCR was performed FastStart Essential DNA Green Master (cat. 06402712001, Roche Life Science, USA) according to the manufacturer’s instructions. The expression levels were normalized to the β2-microlobulin level (β2-microlobulin F: GAGTATGCCTGCCGTGTG, R: AATCCAAATGCGGCATCT) using the equation 2−ΔCt. All qPCR reactions were performed in a Corbett 212 Rotor-gene6000 Real-Time PCR System.
Statistical analysis
GraphPad Prism version 4.00 for Windows (GraphPad Software) and STATA 15.1 were used to perform statistical analyses. The non-parametric Kruskal-Wallis with Dunn’s post test, or the Mann–Whitney test were used to compare continuous variables. Correlations were evaluated with the non-parametric Spearman test. To evaluate the performance of PMN-MDSC frequency as candidate biomarker in discriminating between survivors and non-survivors, a receiver operating characteristic (ROC) curve analysis was performed. Cox regression analysis was used to estimate hazard ratio (HR) and 95%CI of death adjusted for gender and age at admission. A p value < 0.05 was considered statistically significant.
Results
Inflammation promotes PMN-MDSC expansion in COVID-19 patients
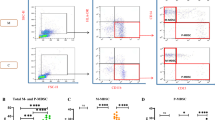
At the admission time, we found a higher frequency of the PMN-MDSC cell population (identified as Lin−, HLA-DR−, CD11b+, CD33+, CD15+, CD14−; Fig. 1A) in 128 hospitalized COVID-19 patients requiring or not requiring intensive care unit (ICU) compared to HD (Fig. 1B). A higher percentage of PMN-MDSC was found in patients requiring ICU compared to patients who did not required ICU care (Fig. 1B). We did not detect monocytic-MDSC in COVID-19 patients or in HD (data not shown). Group sample sizes of 96 (no ICU 75%) and 32 (ICU 25%) achieve 95% power to reject the null hypothesis of equal means of MDSC in the two groups, when the population mean difference is more than 15.0 (standard deviation = 22 for both groups).
A Representative plots of the adopted gating strategy to identify MDSC. Doublets were excluded in the FSC-H/FSC-A dot plot. Live cells were selected as DRAQ7−. In the immunological plot (SideScatter (SSC)/CD45), the CD45+ cells were gated, followed by gating on Lin−(CD3−CD19−CD56−)/HLA-DRlow/− cell. Cells were then selected as CD33+CD11b+, and the expression of CD15+ and CD14− was evaluated. PMN-MDSC frequency was calculated in the immunological gate (CD45+ cells). B Frequency of PMN-MDSC at the admission time in 96 patients who did not require ICU admission, in 32 who did require ICU admissions, and in 30 healthy donors (HD). Results are shown as Box and Whiskers. Kruskal–Walliswith Dunn’s post test was applied. C Correlations between PMN-MDSC frequency and inflammatory cytokines in 78 COVID-19 patients. Non-parametric Spearman correlation was applied, and p < 0.05 was considered significant.
It is well know that the inflammatory cytokines play a central role in inducing the expansion of MDSC17. We quantified, therefore, the IL-1β, IL-6, IL-8, and TNF-α plasma level, and found a significant direct correlation between PMN-MDSC frequency and such inflammatory cytokines (Fig. 1C).
PMN-MDSC inhibit SARS-CoV-2 specific T-cell response
In order to evaluate the suppressive function of PMN-MDSC, we first analyzed the ability of these cells to inhibit the proliferation of CD3+ T cells. We performed PMN-MDSC isolation from PBMC by using CD15 coated microbeads, indeed the CD15+ cells were HLA-DR-, CD11b+`, CD33+ (Supplementary Fig. 1), thus matching the PMN-MDSC phenotype definition. We stimulated either (i) whole PBMC, (ii) PMN-MDSC-depleted (CD15−) PBMC (DEPL), and (iii) PMN-MDSC-depleted PBMCs plus PMN-MDSC (1/4 ratio) (DEPL + MDSC) with SEB, and the proliferation of T cells was tested after four days of culture. We found that PMN-MDSC depletion significantly increase the proliferation rate of CD3+ cells; when PMN-MDSC were added again at 1–4 ratio, the proliferation returned to a similar level as whole PBMC (Fig. 2A, B). To characterize the suppressive ability of PMN-MDSC, the expression of ArgI, TGF-β, and iNOS mRNAs were evaluated in purified PMN-MDSC and DEPL from ten COVID-19 patients. Results showed that PMN-MDSC expressed higher level of Arg1 and iNOS mRNA compared to DEPL (Fig. 2C). Differently, TGF- β was similarly expressed in PMN-MDSC and DEPL. We evaluated the plasmatic level of TGF-β, and found a direct correlation between PMN-MDSC frequency and TGF-β protein in plasma (Fig. 2D), indicating that PMN-MDSC significantly contribute to the production and release of TGF-β.
A Representative flow-cytometry histogram plots of the proliferation rate of CD3+ cells upon stimulation with SEB. B Percentage of CD3+ proliferating cells in PBMC, PMN-MDSC depleted PBMC (DEPL), and DEPL + PMN-MDSC (1/4 ratio) from 5 patients. Results are shown as mean ± SEM. Paired T test was performed. p < 0.05 was considered significant. C mRNA of TGF-β, ArgI, and iNOS in purified PMN-MDSC from 10 COVID-19 patients. Results are shown as Box and Whiskers. D Correlation between PMN-MDSC and plasmatic TGF-β in 22 COVID-19 patients. Non-parametric Spearman correlation was applied, and p < 0.05 was considered significant.
We then evaluated the impact of PMN-MDSC on SARS-CoV-2 specific T cell response. The stimulation of PBMC with SARS-CoV-2 peptides induced only a very low IFN-γ release from T cells. In contrast, PMN-MDSC depletion was able to significantly improve the antigen-specific T-cell response, defined by the increase of IFN-γ production upon SARS-CoV-2 peptide stimulation. This specific T-cell response was switched off when PMN-MDSC were added again (1:4 ratio) as shown by the complete inhibition of the IFN-γ release (Fig. 3A). Similar results were obtained by using PHA that trigger the CD3 complex (Fig. 3A). To define the possible mechanisms used by PMN-MDSC to inhibit SARS-CoV-2 specific T-cell response, we stimulated PBMC with SARS-CoV-2 peptides in the presence of neutralizing anti-TGF-β antibody, an inhibitor of ArgI (N(ω)-hydroxy-nor-L-arginine, Nor-NOHA), and an inhibitor of iNOS (L-NG-nitro arginine methyl ester, L-NAME). Figure 3 panel B shows that anti-TGF-β and L-NAME treatments were able to increase the capacity of T cells to produce IFN-γ in 100% (four out of four), and 75% (three out of four) patients, respectively, indicating that TGF-β and iNOS were involved in the PMN-MDSC mediated suppressive activities.
A IFN-γ production from PBMC, PMN-MDSC depleted PBMC (DEPL), and DEPL + PMN-MDSC (1/4 ratio) from 11 patients upon stimulation with SARS-CoV-2 peptides or with PHA. Results are shown as Box and Whiskers. Kruskal–Walliswith Dunn’s post test was applied. B IFN-γ production from PBMC from 4 patients upon stimulation with SARS-CoV-2 peptides (CTR) and neutralizing anti-human TGF-β, Nor-NOHA, or L-NAME.
High PMN-MDSC frequency is associated with SARS-CoV-2 fatal infection
T-cell response is central in the adaptive immune-mediated elimination of pathogens. Since we showed the detrimental impact of PMN-MDSC on T-cell response to SARS-CoV-2, we wondered whether PMN-MDSC frequency could be predictive of disease outcome. We retrospectively grouped patients into different groups, namely, those who recovered (survivors, n = 59) and those who succumbed to the disease (non-survivors, n = 19), and analyzed the relative PMN-MDSC frequency at the time of admission. We found a significantly higher frequency of PMN-MDSC in the non-survivor compared with the survivor group (Fig. 4A), suggesting that PMN-MDSC percentage could be predictive of disease outcome. To validate this hypothesis, we performed a ROC analysis and found that PMN-MDSC frequency at the admission could distinguish between survivors and non-survivors (Fig. 4B; area under the curve 0.82, p < 0.0001), identifying a cut off value of 54.91% (77.8% and 78% of sensibility and specificity, respectively). This data were further confirmed by estimating the HR of death for MDSC in a Cox regression analysis adjusting for age and gender: the increase of 1% in PMN-MDSC frequency was independently associated with an augmentation of 3% of risk of fatal outcome (Table 1).
A PMN-MDSC frequency at the time of admission grouped in survivors (n = 59) and non-survivors (n = 19). Results are shown as Box and Whiskers. Mann–Whitney test was applied, and a p value < 0.05 was considered significant. B Receiver operating characteristic (ROC) curve for the PMN-MDSC frequency as marker of COVID-19 fatal outcome.
PMN-MDSC frequency kinetic
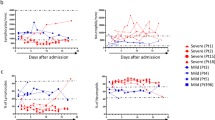
To evaluate whether PMN-MDSC frequency persisted high during the time of infection, a subgroup of patients was monitored weekly until discharged/death. While at the admission survivors and non-survivor did differ in PMN-MDSC frequency, this was not observed during the longitudinal follow-up: both groups, survivors and non-survivors, showed a decreasing trend of PMN-MDSC percentage after admission (Fig. 5A).
A PMN-MDSC frequency in survivors (red line) and non-survivor (black line) patients was monitored weekly. Results are expressed as median and IQR. Mann–Whitney or Friedman test were applied to compare the PMN-MDSC frequency between the two groups at different time points or to compare different time points, respectively. A p value < 0.05 was considered significant. B Kinetic of the indicated cytokines in survivors (red line) and non-survivor (black line) patients. Results are expressed as median and IQR. Mann–Whitney or Friedman test were applied to compare the cytokine levels between the two groups at different time points or to compare different time points, respectively. A p value < 0.05 was considered significant.
Since a correlation between PMN-MDSC and inflammatory cytokines was observed, we analyzed IL-1b, IL-6, IL-8, and TNF-a at the same time points, and found that, accordingly with PMN-MDSC frequency, the plasma IL-8 level was higher in non-survivors than in survivors at the admission time, and decreased to level comparable to survivors at later time points (Fig. 5B). Interestingly, plasma IL-6 levels in the group of non-survivors increased after two weeks from admission, resulting significantly higher than in survivors (Fig. 5B). We did not observe any significant difference for TNF-α and IL-1β.
Discussion
We identified the PMN-MDSC frequency as an early marker of COVID-19 fatal outcome involved in the inhibition of SARS-CoV-2 specific T-cell response. The increased proportion of circulating MDSCs, mainly described in cancer, is attracting great interest in infectious diseases18,19,20,21,22 Although MDSC typically arise in in peripheral tissues to dampen inflammation and inflammation-induced tissue damage, a detrimental role of this cell population in cancer and infectious diseases has been widely reported21,23,24. We recently found an expansion of PMN-MDSC in the peripheral blood of COVID-19 patients compared to HD, in particular in patients experiencing severe disease11,12. We observed an expansion of PMN-MDSC in non-ICU and ICU requiring patients, with the latter group presenting with a higher PMN-MDSC frequency. Following the recommendation for MDSC nomenclature and characterization25, the cells phenotypically here identified as PMN-MDSC were confirmed to act as suppressive cells, since they were able to effectively inhibit T-cell proliferation.
It has been proposed that inflammation promotes MDSC expansion and up-regulation of their suppressive capability17. In COVID-19 patients, a hyper-inflammation has been described2,26 that possibly induced PMN-MDSC differentiation and expansion. Indeed, we observed a correlation between PMN-MDSC frequency and the plasma level of IL-1β, IL-6, IL-8, and TNF-α. Interestingly, patients who will ultimately succumbed to SARS-CoV-2 infection showed a higher frequency of PMN-MDSC at the time of admission as compared to patients who ultimately survived, suggesting that PMN-MDSC frequency can be used as a prognostic marker of disease outcome. The non-survivors PMN-MDSC frequency matched with a high IL-8 level at the admission time, which decreased at later time points, in parallel to the reducing trend of PMN-MDSC. Among the pro-inflammatory mediators, IL-8 has been linked to the recruitment of MDSCs27,28, suggesting that IL-8 could have a major role in PMN-MDSC maintenance. Of note, in non-survivors a significant increase of IL-6 was found after 2 weeks from admission, suggesting that, the decrease of PMN-MDSC frequency could account for IL-6 rise. High IL-6 levels has been recently associated with fatal COVID-19 infection29, suggesting that immune suppression, possibly mediated by expanded MDSC, could be highly beneficial in reducing inflammation.
PMN-MDSC from COVID-19 patients expressed typical mRNA associated with MDSC suppressive functions such as ArgI, TGF-β, and iNOS, providing evidence that these cells exhibited a highly suppressive potential. Moreover, PMN-MDSC frequency correlated with the plasma level of TGF-β, indicating that PMN-MDSC contribute to TGF-β release, which, in turn, may act as a potent enhancer of the MDSC inhibition function30. L-Arginine, nitric oxide, and TGFβ pathways have been reported to play a major role in the regulation of innate and adaptive immune response31,32,33. In particular, these molecules influence human T-cell proliferation, differentiation, and survival34. The importance of the specific T cells has been demonstrated for virus clearance, for limiting tissue damage, and dampening overactive innate immune responses35,36,37. Two recently published studies demonstrated a SARS-CoV-2 specific T-cell response in convalescent/recovered patients38,39. However, timing, composition, magnitude, persistence, and protective role of T-cell response during COVID-19 are still to be fully elucidated. Our data showed a low SARS-CoV-2 specific T-cell response during COVID-19, defined by IFN-γ production, which was increased by depleting MDSC, suggesting that the low T cell responses are due to in vivo suppressive activity, not associated with the absence of antigen-specific T cells. We also found that TGF-β and iNOS produced by PMN-MDSC mediated their suppressive activity. PMN-MDSC from COVID-19 patients also inhibit PHA-induced IFN-γ production, indicating a powerful suppressive activity on T cells.
The suppressive potential of expanded MDSC can inhibit also NK-cell function16. The observation of reduction in the granzime A content of NK cells in patients with COVID-1940 could therefore also be associated with the MDSC-induced suppression of NK activity. We have also reported a negative correlation between PMN-MDSC frequency and perforin expressing NK cells11, consolidating that PMN-MDSC are indeed involved in inhibiting the cytotoxic potential of NK cells.
The ROC analysis confirmed the detrimental role of PMN-MDSC during COVID-19, as their high frequency at early stage of the disease correlated with fatal disease outcome. A logistic regression analysis showed that patients with an early increased PMN-MDSC frequency had a high risk of mortality after adjustment for other cofounders. Altogether, our data indicate that a high PMN-MDSC frequency, even if reduces inflammation, can inhibit both adaptive and innate anti-viral immune response, thus preventing virus elimination and ultimately patients’ recovery.
In conclusion, despite the low number of patients analyzed in this report, this explorative study indicates new biologically and clinically relevant factors in the pathogenesis of SARS-CoV2 infection, namely the MDSC and TGF-β-mediated suppression of virus specific T cell function. Our data also highlights the rational for a possible use of therapeutic approaches focused on reducing MDSC number/function, thereby increasing anti-virus directed T-cell responses as a viable therapeutic approach for patients with cancer41,42.
References
Borges do Nascimento, I. J. et al. Novel Coronavirus Infection (COVID-19) in Humans: A Scoping Review and Meta-Analysis. J. Clin. Med. https://doi.org/10.3390/jcm9040941 (2020).
Huang, C. et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 395, 497–506 (2020).
Guan, W. J. et al. Clinical characteristics of coronavirus disease 2019 in China. N. Engl. J. Med. 382, 1708–1720 (2020).
Xu, Z. et al. Pathological findings of COVID-19 associated with acute respiratory distress syndrome. Lancet Respir. Med. 8, 420–422 (2020).
Chen, L. et al. Risk factors for death in 1859 subjects with COVID-19. Leukemia 34, 2173–2183 (2020).
Yan, X. et al. Neutrophil to lymphocyte ratio as prognostic and predictive factor in patients with coronavirus disease 2019: a retrospective cross-sectional study. J. Med. Virol. 92, 2573–2581 (2020).
Liu, J. et al. Neutrophil-to-lymphocyte ratio predicts critical illness patients with 2019 coronavirus disease in the early stage. J. Transl. Med. 18, 206–020-02374-0 (2020).
Zheng, M. et al. Functional exhaustion of antiviral lymphocytes in COVID-19 patients. Cell. Mol. Immunol. 17, 533–535 (2020).
Diao, B. et al. Reduction and functional exhaustion of T cells in patients with coronavirus disease 2019 (COVID-19). Front. Immunol. 11, 827 (2020).
McGonagle, D., Sharif, K., O’Regan, A. & Bridgewood, C. The role of cytokines including interleukin-6 in COVID-19 induced pneumonia and macrophage activation syndrome-like disease. Autoimmun. Rev. 19, 102537 (2020).
Bordoni, V. et al. An inflammatory profile correlates with decreased frequency of cytotoxic cells in COVID-19. Clin. Infect. Dis. https://doi.org/10.1093/cid/ciaa577 (2020).
Agrati, C. et al. Expansion of myeloid-derived suppressor cells in patients with severe coronavirus disease (COVID-19). Cell Death Differ. 27, 3196–3207 (2020).
Solito, S. et al. A human promyelocytic-like population is responsible for the immune suppression mediated by myeloid-derived suppressor cells. Blood 118, 2254–2265 (2011).
Rodriguez, P. C. et al. Arginase I in myeloid suppressor cells is induced by COX-2 in lung carcinoma. J. Exp. Med. 202, 931–939 (2005).
Lee, B. R. et al. Elevated endoplasmic reticulum stress reinforced immunosuppression in the tumor microenvironment via myeloid-derived suppressor cells. Oncotarget 5, 12331–12345 (2014).
Li, H., Han, Y., Guo, Q., Zhang, M. & Cao, X. Cancer-expanded myeloid-derived suppressor cells induce anergy of NK cells through membrane-bound TGF-beta 1. J. Immunol. 182, 240–249 (2009).
Gabrilovich, D. I. & Nagaraj, S. Myeloid-derived suppressor cells as regulators of the immune system. Nat. Rev. Immunol. 9, 162–174 (2009).
Telatin, V. et al. In chronic hepatitis C infection, myeloid-derived suppressor cell accumulation and T cell dysfunctions revert partially and late after successful direct-acting antiviral treatment. Front. Cell. Infect. Microbiol. 9, 190 (2019).
El Daker, S. et al. Granulocytic myeloid derived suppressor cells expansion during active pulmonary tuberculosis is associated with high nitric oxide plasma level. PLoS One 10, e0123772 (2015).
du Plessis, N. et al. Increased frequency of myeloid-derived suppressor cells during active tuberculosis and after recent mycobacterium tuberculosis infection suppresses T-cell function. Am. J. Respir. Crit. Care Med. 188, 724–732 (2013).
Agrati, C. et al. Myeloid derived suppressor cells expansion persists after early ART and may affect CD4 T cell recovery. Front. Immunol. 10, 1886 (2019).
Vollbrecht, T. et al. Chronic progressive HIV-1 infection is associated with elevated levels of myeloid-derived suppressor cells. AIDS 26, F31–F37 (2012).
Wang, Y., Ding, Y., Guo, N. & Wang, S. MDSCs: Key Criminals of Tumor Pre-metastatic Niche Formation. Front. Immunol. 10, 172 (2019).
Venet, F., Demaret, J., Gossez, M. & Monneret, G. Myeloid cells in sepsis-acquired immunodeficiency. Ann. N. Y. Acad. Sci. 1–15 (2020).
Bronte, V. et al. Recommendations for myeloid-derived suppressor cell nomenclature and characterization standards. Nat. Commun. 7, 12150 (2016).
Qin, C. et al. Dysregulation of immune response in patients with COVID-19 in Wuhan, China. Clin. Infect. Dis. 71, 762–768 (2020).
Alfaro, C. et al. Tumor-produced interleukin-8 attracts human myeloid-derived suppressor cells and elicits extrusion of neutrophil extracellular traps (NETs). Clin. Cancer Res. 22, 3924–3936 (2016).
Tobin, R. P. et al. IL-6 and IL-8 Are linked with myeloid-derived suppressor cell accumulation and correlate with poor clinical outcomes in melanoma patients. Front. Oncol. 9, 1223 (2019).
Ruan, Q., Yang, K., Wang, W., Jiang, L. & Song, J. Clinical predictors of mortality due to COVID-19 based on an analysis of data of 150 patients from Wuhan, China. Intensive Care Med. 46, 846–848 (2020).
Lee, C. R., Lee, W., Cho, S. K. & Park, S. G. Characterization of multiple cytokine combinations and TGF-beta on differentiation and functions of myeloid-derived suppressor cells. Int. J. Mol. Sci. https://doi.org/10.3390/ijms19030869 (2018).
Kim, S. H., Roszik, J., Grimm, E. A. & Ekmekcioglu, S. Impact of l-arginine metabolism on immune response and anticancer immunotherapy. Front. Oncol. 8, 67 (2018).
Pickup, M., Novitskiy, S. & Moses, H. L. The roles of TGFbeta in the tumour microenvironment. Nat. Rev. Cancer 13, 788–799 (2013).
Blesson, S. et al. Analysis of the mechanisms of human cytotoxic T lymphocyte response inhibition by NO. Int. Immunol. 14, 1169–1178 (2002).
Geiger, R. et al. L-Arginine modulates T cell metabolism and enhances survival and anti-tumor activity. Cell 167, 829–842. e13 (2016).
Palm, N. W. & Medzhitov, R. Not so fast: adaptive suppression of innate immunity. Nat. Med. 13, 1142–1144 (2007).
Kim, K. D. et al. Adaptive immune cells temper initial innate responses. Nat. Med. 13, 1248–1252 (2007).
Zhao, J., Zhao, J. & Perlman, S. T cell responses are required for protection from clinical disease and for virus clearance in severe acute respiratory syndrome coronavirus-infected mice. J. Virol. 84, 9318–9325 (2010).
Grifoni, A. et al. Targets of T Cell Responses to SARS-CoV-2 Coronavirus in Humans with COVID-19 Disease and Unexposed Individuals. Cell 181, 1489–1501.e15 (2020).
Ni, L. et al. Detection of SARS-CoV-2-Specific Humoral and Cellular Immunity in COVID-19 Convalescent Individuals. Immunity 52, 971–977.e3 (2020).
Mazzoni, A. et al. Impaired immune cell cytotoxicity in severe COVID-19 is IL-6 dependent. J. Clin. Invest 130, 4694–4703 (2020).
Tobin, R. P. et al. Targeting myeloid-derived suppressor cells using all-trans retinoic acid in melanoma patients treated with Ipilimumab. Int. Immunopharmacol. 63, 282–291 (2018).
Eriksson, E., Wenthe, J., Irenaeus, S., Loskog, A. & Ullenhag, G. Gemcitabine reduces MDSCs, tregs and TGFbeta-1 while restoring the teff/treg ratio in patients with pancreatic cancer. J. Transl. Med. 14, 282 (2016).
Acknowledgements
We gratefully acknowledge the Collaborators Members of INMI COVID-19 study group: Maria Alessandra Abbonizio, Amina Abdeddaim, Chiara Agrati, Fabrizio Albarello, Gioia Amadei, Alessandra Amendola, Mario Antonini, Tommaso Ascoli Bartoli, Francesco Baldini, Raffaella Barbaro, Barbara Bartolini, Rita Bellagamba, Martina Benigni, Nazario Bevilacqua, Gianlugi Biava, Michele Bibas, Licia Bordi, Veronica Bordoni, Evangelo Boumis, Marta Branca, Donatella Busso, Marta Camici, Paolo Campioni, Maria Rosaria Capobianchi, Alessandro Capone, Cinzia Caporale, Emanuela Caraffa, Ilaria Caravella, Fabrizio Carletti, Concetta Castilletti, Adriana Cataldo, Stefano Cerilli, Carlotta Cerva, Roberta Chiappini, Pierangelo Chinello, Carmine Ciaralli, Stefania Cicalini, Francesca Colavita, Angela Corpolongo, Massimo Cristofaro, Salvatore Curiale, Alessandra D’Abramo, Cristina Dantimi, Alessia De Angelis, Giada De Angelis, Maria Grazia De Palo, Federico De Zottis, Virginia Di Bari, Rachele Di Lorenzo, Federica Di Stefano, Gianpiero D’Offizi, Davide Donno, Francesca Faraglia, Federica Ferraro, Lorena Fiorentini, Andrea Frustaci, Matteo Fusetti, Vincenzo Galati, Roberta Gagliardini, Paola Gallì, Gabriele Garotto, Saba Gebremeskel Tekle, Maria Letizia Giancola, Filippo Giansante, Emanuela Giombini, Guido Granata, Maria Cristina Greci, Elisabetta Grilli, Susanna Grisetti, Gina Gualano, Fabio Iacomi, Giuseppina Iannicelli, Giuseppe Ippolito, Eleonora Lalle, Simone Lanini, Daniele Lapa, Luciana Lepore, Raffaella Libertone, Raffaella Lionetti, Giuseppina Liuzzi, Laura Loiacono, Andrea Lucia, Franco Lufrani, Manuela Macchione, Gaetano Maffongelli, Alessandra Marani, Luisa Marchioni, Andrea Mariano, Maria Cristina Marini, Micaela Maritti, Alessandra Mastrobattista, Giulia Matusali, Valentina Mazzotta, Paola Mencarini, Silvia Meschi, Francesco Messina, Annalisa Mondi, Marzia Montalbano, Chiara Montaldo, Silvia Mosti, Silvia Murachelli, Maria Musso, Emanuele Nicastri, Pasquale Noto, Roberto Noto, Alessandra Oliva, Sandrine Ottou, Claudia Palazzolo, Emanuele Pallini, Fabrizio Palmieri, Carlo Pareo, Virgilio Passeri, Federico Pelliccioni, Antonella Petrecchia, Ada Petrone, Nicola Petrosillo, Elisa Pianura, Carmela Pinnetti, Maria Pisciotta, Silvia Pittalis, Agostina Pontarelli, Costanza Proietti, Vincenzo Puro, Paolo Migliorisi Ramazzini, Alessia Rianda, Gabriele Rinonapoli, Silvia Rosati, Martina Rueca, Alessandra Sacchi, Alessandro Sampaolesi, Francesco Sanasi, Carmen Santagata, Alessandra Scarabello, Silvana Scarcia, Vincenzo Schininà, Paola Scognamiglio, Laura Scorzolini, Giulia Stazi, Fabrizio Taglietti, Chiara Taibi, Roberto Tonnarini, Simone Topino, Francesco Vaia, Francesco Vairo, Maria Beatrice Valli, Alessandra Vergori, Laura Vincenzi, Ubaldo Visco-Comandini, Pietro Vittozzi, Mauro Zaccarelli. INMI authors are supported by the Italian Ministry of Health (Ricerca Corrente, COVID-2020-12371735 and COVID-2020-12371817). Profs Ippolito and Zumla are co-principal investigator of the Pan-African Network on Emerging and Re-emerging Infections (PANDORA-ID-NET), funded by the European & Developing Countries Clinical Trials Partnership, supported under Horizon 2020. Sir Zumla is in receipt of a National Institutes of Health Research senior investigator award. Prof Maeurer is a member of the innate immunity advisory group of the Bill & Melinda Gates Foundation, and is funded by the Champalimaud Foundation.
Author information
Authors and Affiliations
Contributions
Conceptualization: A.S. and C.A.; Experimental design: A.S., C.A., and M.R.C.; Flow cytometry analysis: G.G., V.B., E.C., and R.C.; Functional experiments: A.S., G.G., M.T., and P.C.; Cytokine analysis: S.N. and E.T.; Statistical analysis: P.L. and A.S.; Patient management: L.M., N.P., A.A., G.D., F.P., and E.N.; Writing the paper: A.S., G.G., and C.A.; Revised the paper: M.M., A.Z., G.I., and F.L.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by H.-U. Simon
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sacchi, A., Grassi, G., Bordoni, V. et al. Early expansion of myeloid-derived suppressor cells inhibits SARS-CoV-2 specific T-cell response and may predict fatal COVID-19 outcome. Cell Death Dis 11, 921 (2020). https://doi.org/10.1038/s41419-020-03125-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-020-03125-1
This article is cited by
-
Longitudinal single cell atlas identifies complex temporal relationship between type I interferon response and COVID-19 severity
Nature Communications (2024)
-
Impairment of neutrophil functions and homeostasis in COVID-19 patients: association with disease severity
Critical Care (2022)
-
Increased circulating myeloid-derived suppressor cells in vivax malaria and severe falciparum malaria
Malaria Journal (2022)
-
Immunophenotype and function of circulating myeloid derived suppressor cells in COVID-19 patients
Scientific Reports (2022)
-
Clinical perspectives on the age-related increase of immunosuppressive activity
Journal of Molecular Medicine (2022)