Abstract
Apoptotic cell clearance by phagocytes is a fundamental process during development, homeostasis and the resolution of inflammation. However, the demands placed on phagocytic cells such as macrophages by this process, and the limitations these interactions impose on subsequent cellular behaviours are not yet clear. Here, we seek to understand how apoptotic cells affect macrophage function in the context of a genetically tractable Drosophila model in which macrophages encounter excessive amounts of apoptotic cells. Loss of the glial-specific transcription factor Repo prevents glia from contributing to apoptotic cell clearance in the developing embryo. We show that this leads to the challenge of macrophages with large numbers of apoptotic cells in vivo. As a consequence, macrophages become highly vacuolated with cleared apoptotic cells, and their developmental dispersal and migration is perturbed. We also show that the requirement to deal with excess apoptosis caused by a loss of repo function leads to impaired inflammatory responses to injury. However, in contrast to migratory phenotypes, defects in wound responses cannot be rescued by preventing apoptosis from occurring within a repo mutant background. In investigating the underlying cause of these impaired inflammatory responses, we demonstrate that wound-induced calcium waves propagate into surrounding tissues, including neurons and glia of the ventral nerve cord, which exhibit striking calcium waves on wounding, revealing a previously unanticipated contribution of these cells during responses to injury. Taken together, these results demonstrate important insights into macrophage biology and how repo mutants can be used to study macrophage–apoptotic cell interactions in the fly embryo. Furthermore, this work shows how these multipurpose cells can be ‘overtasked’ to the detriment of their other functions, alongside providing new insights into which cells govern macrophage responses to injury in vivo.
Similar content being viewed by others
Introduction
Understanding the interactions between macrophages and apoptotic cells is an important biological question: failures in how immune cells deal with apoptotic cell death can lead to damaging autoimmune conditions1. Apoptotic cell clearance also plays a critical role in the resolution of inflammation and in the reprogramming of immune cells during this process2. Furthermore, interactions between dying cells and macrophages occur at numerous sites of pathology, including at sites of atherosclerosis3 and in the chronically inflamed lungs of patients with chronic obstructive pulmonary disease (COPD)4. As such, macrophage–apoptotic cell interactions have the potential to impact these diseases and many other damaging human conditions1.
Drosophila has proven an excellent organism with which to study innate immunity5, haematopoiesis6 and blood cell function7. Drosophila blood is dominated by macrophage-like cells (plasmatocytes), with these cells making up 95% of the blood cells (hemocytes) in the developing embryo8. These embryonic macrophages disperse over the entire embryo during development, phagocytosing apoptotic cells and secreting matrix as they migrate7. Alongside their functional and morphological similarities to vertebrate macrophages, Drosophila macrophages are specified through the action of related transcription factors to those used in vertebrate haematopoiesis9. Failed dispersal or ablation of these embryonic macrophages leads to developmental abnormalities and failure to hatch to larval stages10,11,12. Drosophila macrophages are also able to respond to injury, mounting inflammatory responses to epithelial wounds13. In embryos and pupae, wounding elicits a rapid calcium wave through the epithelium, a process that requires transient receptor potential (Trp) channel function14,15. The increase in cytoplasmic calcium drives activation of dual oxidase (DUOX) and production of hydrogen peroxide, which is necessary for immune cell recruitment14, resembling events upon tissue damage in higher organisms such as zebrafish16,17.
In addition to macrophages, the other main phagocyte population within Drosophila embryos is the glia of the developing nervous system18. Drosophila embryonic macrophages interact with the developing ventral nerve cord (VNC) and the glial cells that encase it, as they disperse along the ventral side of the embryo19. The VNC contains two populations of glia—the Sim-positive midline glia that help establish the ladder-like structure of neurons early in development and Repo-positive lateral glia20. Repo is a homeodomain transcription factor that specifies glial fate, though defects in repo mutants are not obvious until late in embryonic development, when a decrease in the numbers of glial cells becomes apparent21,22,23. While early glial markers are not lost in repo mutants22,24, Repo is required for the expression of at least some phagocytic receptors used by these cells25. Despite an overlap in the transcription factors used to specify blood cells and glia (e.g., Gcm and Gcm2)26,27, these cells are derived from distinct progenitors and Repo antagonises haematopoiesis to promote a glial fate28. Nonetheless, both macrophages and repo-positive glial cells express a similar repertoire of receptors for apoptotic cells, including Draper and Simu29. The close interactions between these phagocytes means that, should one population fail to clear apoptotic cells, it is likely that the other population would be able to both detect this deficiency and be able to compensate. Loss of apoptotic cell receptors such as Simu leads to a buildup of apoptotic cells within developing embryos30 and this impairs macrophage behaviours, including their developmental dispersal and inflammatory responses31. However, whilst mutants such as simu enable exposure of macrophages to elevated numbers of apoptotic cells in vivo, this approach is not ideal, as it also perturbs receptors that may be involved in immune cell reprogramming32,33.
To investigate interactions between macrophages and apoptotic cells in more detail, we stimulated macrophages with enhanced levels of apoptotic cells using a genetic approach that did not alter the macrophages themselves. By impairing glial differentiation using repo mutants, we increased the number of apoptotic cells macrophages face within the developing embryo. This enhanced apoptotic challenge impaired macrophage dispersal, migration and their inflammatory responses to wounds. In this background, clearance of apoptotic cells by macrophages is not perturbed, and migration can be rescued by preventing apoptosis. Surprisingly, and in contrast to phagocytic receptor mutants, blocking apoptotic cell death in the presence of defective glia failed to rescue wound responses. Further analysis revealed that injury-induced calcium waves propagate beyond the wounded epithelium, and that this process is defective in repo mutants. This suggests that glial cells play an active role in the propagation of even the earliest responses to wounding. Thus, this model provides a unique insight into how macrophage–apoptotic cell interactions dictate macrophage responses to injury and the cell types that contribute to the activation of those responses.
Materials and methods
Fly lines and husbandry
Drosophila melanogaster fruit flies were reared at 25 °C on cornmeal/agar/molasses media (see Supplementary Table 1 for recipe). Depending on the exact alleles and transgenes needed for each experiment, either srp-GAL434 and/or crq-GAL413 were used interchangeably to label Drosophila macrophages in the embryo in combination with UAS-GFP and/or UAS-red stinger35. e22c-GAL4 (VNC and epithelium)36, act5C-GAL4 (ubiquitous)37, da-GAL4 (ubiquitous)38, elav-GAL4 (neuronal)39 and repo-GAL4 (glial cells)40 were used to drive expression in other tissues. UAS-GCaMP6M41 was used to image changes in cytoplasmic calcium concentration. Experiments were conducted on a w1118 background, and the following mutant alleles were used: Df(3L)H9942, repo03702 (see refs. 21,22,23), simu2 (see ref. 30). See Supplementary Table 2 for a full list of genotypes used in this study and sources of the Drosophila lines used. Embryos were collected from apple juice agar plates on which flies had laid overnight at 22 °C. Embryos were washed off plates with distilled water and dechorionated in bleach for 1–2 min. Bleach was thoroughly washed away with distilled water ahead of fixation or mounting of embryos for live imaging. The absence of the fluorescent balancers CTG, CyO dfd, TTG and TM6b dfd43,44 was used to select homozygous mutant embryos after dechorionation.
Fixation and immunostaining
Dechorionated embryos were fixed and stained as per Roddie et al.31. Antibodies were diluted in PATx (0.1% Triton-X100 (Sigma-Aldrich), 1% BSA (Sigma-Aldrich) in PBS (Oxoid, Thermo Fisher, MA, USA)). Rabbit anti-GFP (ab290 1:1000; Abcam, Cambridge, UK) or mouse anti-GFP (ab1218 1:200; Abcam) were used to detect GFP-labelled macrophages. Rabbit anti-cDCP-1 (9578S 1:1000; Cell Signaling Technologies), mouse anti-Repo (concentrate of clone 8D12 used at 1:1000; Developmental Studies Hybridoma Bank, University of Iowa, USA) or mouse anti-Futch (supernatant of clone 22C10 used at 1:200; Developmental Studies Hybridoma Bank) were also used as primary antibodies. Goat anti-mouse or goat anti-rabbit secondary antibodies conjugated to AlexaFluor568, AlexaFluor488 (A11036 and A11034; Invitrogen, Thermo Fisher) or FITC (115-095-146; Jackson Immunoresearch, Cambridge, UK) were used to detect primary antibodies; these were diluted from stock solutions made according to the recommendations of the supplier (1:400 in PATx). Stained embryos were stored in DABCO mountant (Sigma-Aldrich) and mounted on slides for imaging.
Imaging and analysis
Immunostained embryos were imaged using a ×40 objective lens (CFI Super Plan Fluor ELWD 40x, NA 0.6) on a Nikon A1 confocal. In order to quantify apoptotic cell clearance, embryos containing GFP-labelled macrophages were stained for GFP and cDCP-1. Only those macrophages fully in view on the ventral midline and within a 10-μm deep sub-stack that corresponds to the region between the epidermis and VNC on the ventral midline were analysed. The number of cDCP-1 punctae within macrophage, per embryo (phagocytic index) was calculated from slices in the 10-μm deep sub-stack to provide a measure of apoptotic cell clearance. Sub-stacks were blinded before analysis, and a minimum of four macrophages were counted per embryo. To quantify vacuolation, the number of vacuoles within a single z-slice at which each macrophage exhibited its maximal cross-sectional area was scored. Vacuoles were discriminated via their exclusion of cytoplasmic GFP. Only macrophages on the midline between the epidermis and VNC and fully within view were analysed.
Developmental dispersal was assessed by counting numbers of segments lacking GFP-labelled macrophages on the ventral side of the VNC in stage 13/14 embryos. Embryos that had been fixed and stained for GFP were scored using a Leica MZ205 FA fluorescent dissection microscope with a PLANAPO ×2 objective lens. The same embryos were orientated ventral-side-up, imaged on a Nikon A1 confocal, and numbers of macrophages between epithelium and VNC counted (segments 4–8 were scored).
For live imaging of macrophage morphology, migration speed and wound responses and calcium dynamics upon injury, live embryos were mounted in voltalef oil (VWR) as per Evans et al.19 and imaged on a Perkin Elmer UltraView Spinning Disk system using a ×40 objective lens (UplanSApo ×40 oil, NA 1.3). Embryos that were not ventral-side-up, that rolled during imaging or with large leaks post wounding were excluded from subsequent analyses. Epithelial wounds were made on the ventral side of embryos using a nitrogen-pumped Micropoint ablation laser (Andor, Belfast, UK), as per Evans et al.45. Macrophage inflammatory responses to wounds were imaged and analysed as per Roddie et al.31. Macrophage wound responses (numbers of macrophage at or touching the wound edge at 60 min divided by wound area in μm2) were normalised to control levels. The percentage of macrophages responding (% responders) was calculated by counting the proportion of macrophages present immediately following wounding that reached the wound within 60 min; macrophages already at the wound or absent from the field of view immediately after wounding were not considered in this analysis. Numbers of macrophages in the image taken prior to wounding (pre-wound) were counted as a measure of macrophages available to respond to the injury. The percentage of macrophages leaving the wound (% leavers) is the proportion of macrophages present at the wound site at any point during a 60-min movie that leave the wound site; macrophages retaining contacts with the wound site or edge were not scored as leavers.
For analysis of random migration, 60-min movies of GFP and red stinger double-labelled macrophages were made with macrophage position imaged every 2 min on the ventral midline between the overlying epithelium and VNC. Image stacks were despeckled, and the movements of macrophages within maximum projections tracked as per Roddie et al.31. The manual tracking and Ibidi chemotaxis plugins were used to calculate speed per macrophage, per embryo, in Fiji46.
Calcium dynamics were imaged using GCaMP6M expressed using a range of GAL4 lines. Calcium responses were quantified from average projections of z-stacks collected immediately before and after wounding. Initial wound responses of projections corresponding to epithelial z-slices (from da-GAL4,UAS-GCaMP6M embryos) were quantified by measuring the area of the GCaMP6M response and the corresponding mean gray value (MGV) of this region of interest (ROI) in Fiji immediately after wounding (F1). The same ROI was then used to measure the MGV of GCaMP6M fluorescence in the pre-wound image (F0). Both the area of the initial response and the F1/F0 ratio of MGVs were used as measures of the calcium response to injury (Fig. 7c, d). For analysis of glial responses and the cells within the VNC more broadly (neurons and glia), repo-GAL4 and e22c-GAL4 were used to drive GCaMP6M expression, respectively. Responding glial cells were manually selected using the freehand selection tool in Fiji (Fig. 8h, i), while the outer edge of the vitelline membrane was used to define a ROI to quantify GCaMP6M intensity for projections constructed from deeper volumes of the z-stack (Fig. 8c).
Image processing and statistical analyses
Images were despeckled in Fiji before maximum or average projections were made. All projections or z-stacks to be analysed were blinded ahead of quantification, and Adobe Photoshop was used to assemble figures.
Data are displayed on scattergraphs with lines representing the mean and with standard deviation used as a measure of spread (shown via error bars). Two-sided statistical analyses were performed in GraphPad Prism. Data were not assumed to be normally distributed, and Mann–Whitney and Kruskall–Wallis tests were used. A Pearson test was used to test for correlations between numbers of vacuoles and cell speed (Fig. 4f, g). Details of statistical analyses, post-tests used to correct for multiple comparisons, P values and sample sizes (number of embryos) are reported in figure legends. Experiments were repeated in at least triplicate with embryos taken from laying cages containing greater than 50 adult flies of the same genotype. Minimum sample sizes were estimated based on numbers of embryos sufficient to see a difference of 20% from previous studies. For live-imaging experiments, mutant or control embryos of the correct developmental stage were selected at random from pools of dechorionated embryos of the same genotype and mounted on the same microscope slide for each replicate.
Immunostained embryos are representative examples taken from batches of pooled embryos collected across multiple days. Control and mutant embryos were stained at the same time using master mixes of reagents that was split to stain the different genotypes.
Results
Embryonic macrophages disperse in close contact with the developing central nervous system
Drosophila embryonic macrophages migrate out from the presumptive head region to disperse over the entire embryo during development7. Migration along the developing ventral nerve cord (VNC) is an essential route for macrophage dispersal, with macrophages contacting the overlying epithelium and glial cells on the surface of the nerve cord (Fig. 1a, b). During dispersal, macrophages encounter and clear apoptotic cells (Fig. 1c), while VNC glia also phagocytose dying cells47. The interaction of macrophages and glia suggested to us that impairing glial-mediated apoptotic cell clearance could increase exposure of macrophages to apoptotic cell death in vivo, thus providing a model with which apoptotic cell–macrophage interactions and their effects on macrophage behaviour could be studied in detail.
a, b Single z-slices (a, b) and maximum projections (a’, b’) of immunostained control embryos with GFP-labelled macrophages (green) showing progression of macrophages along both sides of the ventral nerve cord (VNC, Futch staining; magenta) at stage 12 (a) and 14 (b). Arrows indicate the position of VNC; embryos are laterally orientated with anterior to the left, and ventral down. c Ventral views of a stage 15 control embryo immunostained for apoptotic cells (anti-cDCP-1, magenta) and macrophages (anti-GFP, green); panels show single z-slices showing engulfment of apoptotic cells by macrophages (arrows, c) and apoptotic cells within the VNC (arrowheads, c’), and a maximum projection of this region corresponding to a 20-μm deep z-stack (c”); a white line in (c’) indicates the edges of the VNC—macrophages within these lines are sitting within the midline pores that span this structure. d, e Ventral views of stage 15 control and repo mutant embryos immunostained for GFP to show macrophages (d, e) and anti-Repo (d’, e’); macrophages and Repo are green and magenta, respectively, in merged images (d”, e”); arrows in anti-Repo channel indicate non-nuclear staining likely to be cross-reactivity to another epitope. All scale bars represent 10 μm. Genotypes are as follows: w;srp-GAL4,UAS-GFP/ + ;crq-GAL4,UAS-GFP/ + (a, b), w;;crq-GAL4,UAS-GFP (c, d), w;;repo03702,crq-GAL4,UAS-GFP (e).
Increased efferocytosis by embryonic macrophages in the absence of functional glial cells
Macrophages and glia use the same phagocytic receptors to clear apoptotic cells30,48,49, and the absence of these receptors perturbs efferocytosis, in turn disrupting macrophage function31,45. While this suggests that apoptotic cells modulate macrophage behaviour in flies, these interventions remove genes that also control macrophage fate32. Therefore, in order to expose macrophages to increased amounts of apoptotic cell death without directly impacting their own specification mechanisms or removing important regulators of phagocytosis, we targeted the glia of the VNC. We hypothesised that blocking glial-mediated clearance would lead to macrophages becoming exposed to increased numbers of apoptotic cells due to the inability of glial cells to contribute to this process.
The homeodomain transcription factor Repo is expressed by all glial cells within the developing VNC21,22,23, except the midline glia, which are specified via the action of sim50. Repo is absolutely required for normal specification of these glial cells, and absence of repo leads to the failure to express a variety of phagocytic receptors required for apoptotic cell clearance25. Repo is not expressed by macrophages in the developing embryo (Fig. 1d; Lee and Jones40), and repo03702 mutants lack detectable protein expression in the VNC (Fig. 1e). Consistent with the failed condensation of the VNC in the absence of repo function22, there is a small but statistically significant increase in the length of body segments at stage 15 in repo mutants compared with controls (Supplementary Fig. 1a, b). As per other VNC-specific defects21,22, this phenotype cannot be detected at earlier stages of development (Supplementary Fig. 1c).
To test whether failed glial specification would lead to macrophages encountering increased numbers of apoptotic cells in the developing embryo, we analysed macrophage morphology. Macrophages in repo mutants are highly vacuolated compared with controls (Fig. 2a–c); vacuoles within Drosophila embryonic macrophages typically contain previously engulfed apoptotic cells51. To test whether apoptotic cell clearance by macrophages is increased in repo mutants, control and repo mutant embryos were immunostained for a marker of apoptotic cell death (cleaved DCP-1 (cDCP-1) immunostaining; DCP-1 is cleaved by caspases during apoptosis)52. Macrophages in repo mutants contain far higher numbers of cDCP-1-positive inclusions compared with controls (Fig. 2d–f). Furthermore, macrophages can be labelled using crq-GAL4 or srp-GAL4 in a repo mutant background and efficiently engulf apoptotic cells (Fig. 2), consistent with their normal specification and physiology. There is no difference in the overall numbers of macrophages within embryos in repo mutants compared with controls (Supplementary Fig. 2), suggesting that macrophage proliferation is unaffected by loss of repo function, and that increased levels of apoptotic cell clearance do not lead to macrophage cell death. Taken together, these results suggest that loss of repo function is a suitable tool via which the effects of increased macrophage–apoptotic cell contact can be analysed in vivo.
a, b Ventral views of stage 15 control (a) and repo mutant (b) embryos containing GFP and red stinger-expressing macrophages; lower panels show zooms of macrophages indicated by boxes in upper panels. c Scattergraph of vacuoles per macrophage per embryo from genotypes in (a, b); P < 0.0001 via Mann–Whitney test (n = 24 and 30). d, e Ventral views of stage 15 control (d) and repo mutant (e) embryos containing GFP-labelled macrophages immunostained for GFP (green in merge) and apoptotic cells (anti-cDCP-1, magenta in merge); zooms show macrophages indicated in boxed regions. f Scattergraphs of phagocytic index (cDCP-1 puncta per macrophage per embryo); P < 0.0001 (n = 101 and 69 cells from 6 and 5 embryos, respectively) and P = 0.0043 (n = 6 and 5 embryos, respectively) via Mann–Whitney tests. Scale bars represent 20 μm or 5 μm in zooms; error bars and lines show standard deviation and mean, respectively (c, f); genotypes are w;srp-GAL4,UAS-GFP/srp-GAL4,UAS-red stinger (a, c), w;srp-GAL4,UAS-GFP/srp-GAL4,UAS-red stinger;repo03702 (b, c), w;;crq-GAL4,UAS-GFP (d, f) and w;;repo03702,crq-GAL4,UAS-GFP (e, f).
An increased burden of apoptotic cell clearance is associated with impaired developmental dispersal of macrophages
Apoptotic cells represent the top priority for Drosophila macrophages to respond to within developing embryos53. As a result, increased numbers of apoptotic cells have the potential to disrupt macrophage behaviours, such as their dispersal and recruitment to sites of tissue injury. repo mutants lack gross dispersal defects, with macrophages present along both sides of the VNC at stage 13 (Fig. 3a, b). However, reduced numbers of macrophages are present on the ventral midline in repo mutants (Fig. 3c–e), suggesting an impairment in dispersal, especially since there is no decrease in total numbers of macrophages within repo mutant embryos (Supplementary Fig. 2). Further increasing apoptotic burden in a repo mutant background (via removal of the apoptotic cell clearance receptor Simu) causes large dispersal defects (Fig. 3a, b), comparable to those observed when critical regulators of migration are absent, e.g., SCAR/WAVE51. Combined, these results indicate that challenge of macrophages with excessive amounts of apoptosis can impair developmental dispersal of macrophages.
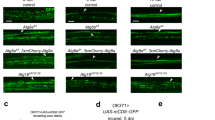
a Lateral views of stage 13/14 control, repo, simu and simu;repo double-mutant embryos containing GFP-labelled macrophages. A complete line of macrophages is present on midline on the ventral side of the VNC in all genotypes, with the exception of simu;repo double mutants (gap indicated via an asterisk); a’ shows zoom of this region. b Scattergraph showing quantification of midline progression defects (numbers of segments lacking macrophages on the ventral side of the VNC); P < 0.0001 (for simu;repo vs each other genotype; no other comparisons are significantly different) via a Kruskall–Wallis test with a Dunn’s multiple comparaison post-test; n = 15 (controls), 17 (repo mutants), 15 (simu mutants) and 11 (simu;repo). c, d Ventral views of stage 13 control and repo mutant embryos containing GFP-expressing macrophages (immunostained via anti-GFP). e Scattergraph showing quantification of numbers of macrophages in five central segments on the ventral midline; P < 0.0001 via Mann–Whitney test, n = 13 (controls), 13 (repo mutants). Scale bars represent 50 μm (a) and 25 μm (c, d); lines and error bars on scattergraphs show mean and standard deviation, respectively; **** indicates P < 0.0001. Genotypes are w1118;;crq-GAL4,UAS-GFP (control), w1118;;P{PZ}repo03702,crq-GAL4,UAS-GFP (repo03702), w1118;simu2;crq-GAL4,UAS-GFP (simu2) and w1118;simu2;P{PZ}repo03702,crq-GAL4,UAS-GFP (simu;repo).
Apoptotic cells are responsible for attenuation of macrophage migration in repo mutants
To test the effects of increased apoptotic cell exposure on macrophage migration, we tracked the movements of macrophages on the ventral midline at stage 15 after completion of developmental dispersal, a point at which macrophages exhibit wandering/'random migration'. In repo mutant embryos, macrophages move at significantly slower speeds compared with those in controls (Fig. 4a–e). In order to test whether this attenuation of migration speed is dependent on interactions with apoptotic cells, we removed all developmental apoptosis from a repo mutant background using the Df(3L)H99 deficiency, which deletes the pro-apoptotic genes Hid, reaper and grim42. In these repo mutants that lack apoptosis, there is a significant rescue of macrophage migration speed (Fig. 4c–e), suggesting that it is interactions with apoptotic cells that impair macrophage migration in vivo, rather than defective glial specification per se. There is no correlation between the degree of macrophage vacuolation and migration speed in repo mutants, and only a small correlation in control embryos (Fig. 4f, g). Therefore, engulfment of apoptotic cells is not required to decrease macrophage migration speeds in vivo.
a–d Ventral images of GFP and red stinger-labelled macrophages (green and magenta, respectively) in stage 15 control (a), Df(3L)H99 mutant (b), repo03702 mutant (c) and Df(3L)H99,repo03702 double-mutant embryos (d). Upper panels show initial position of macrophages on the ventral midline; central panels show tracks taken from subsequent 60-min period from the timepoint shown in the upper panel; the lower panel shows overlay of tracks over initial positions of macrophages. e Superplot showing speed per macrophage per embryo (black dots) in μm per min; grey dots represent individual cell speeds to show range of migration speeds in each genotype. Lines and error bars show mean and standard deviation, respectively (embryo averages). Removing apoptosis from a repo mutant background (Df(3L)H99,repo) rescues migration speed (P = 0.0095 via Mann–Whitney test; n = 4, 5, 6 and 6 embryos (left to right)). f, g Scatterplots of macrophage speed and vacuolation for macrophages in control (f) and repo mutant (g) embryos (80 and 103 macrophages analysed from 6 and 10 control and repo mutant embryos, respectively). Linear regression lines shown in magenta. Scale bars represent 20 μm; ns and ** indicate not significant and P < 0.01, respectively. Genotypes are w1118;srp-GAL4,UAS-red stinger/srp-GAL4,UAS-GFP (control), w1118;srp-GAL4,UAS-red stinger/srp-GAL4,UAS-GFP;Df(3L)H99 (Df(3L)H99), w1118;srp-GAL4,UAS-red stinger/srp-GAL4,UAS-GFP;P{PZ}repo03702 (repo03702) and w1118;srp-GAL4,UAS-red stinger/srp-GAL4,UAS-GFP;Df(3L)H99,P{PZ}repo03702 (Df(3L)H99,repo03702).
Functional glia are required for normal macrophage migration to wounds
Given the impaired migration of macrophages in repo mutant embryos, we analysed their inflammatory responses to sterile, laser-induced wounds. Wounding of repo mutant embryos revealed that reduced numbers of macrophages reached wound sites by 60-min post wounding compared with controls (Fig. 5a, b, d). Since there are fewer macrophages present locally at this developmental stage in repo mutants (Fig. 5c), the percentage of cells responding to the injury was also quantified, revealing a significant decrease in the ability of cells to respond to wounds (Fig. 5e; Supplementary Movie 1). Early migration away from wounds does not appear to underlie these defects (Fig. 5f), suggesting that excessive amounts of apoptotic cells turn off wound responses in repo mutants. To further evidence this conclusion, we removed developmental apoptosis, comparing wound responses in Df(3L)H99 mutants and Df(3L)H99,repo double-mutant embryos. Wound responses are defective in Df(3L)H99 mutants32, though we have found this phenotype to exhibit variable penetrance (Fig. 6; Roddie et al.31). Therefore, at best we would expect a rescue of the more severe repo mutant phenotype to Df(3L)H99 levels, though in this instance we did not detect a strong Df(3L)H99 defect in controls (Fig. 6e, f).
a, b Ventral views showing localisation of GFP-labelled macrophages on the ventral midline in control (a, a’) and repo mutant (b, b’) embryos immediately before wounding (a, b) and at 60-min post wounding (a’, b’); white-dotted ellipses indicate wound edges. c Scattergraph showing numbers of macrophages in the field of view ahead of wounding per embryo; P < 0.0001 via Mann–Whitney test (n = 23 and 15 control and repo, respectively). d Scattergraph showing wound responses quantified via density of macrophages at wounds, normalised to control; P < 0.0001 via Mann–Whitney test (n = 23 and 17 control and repo, respectively). e Scattergraph showing percentage of macrophages responding to wounds; P = 0.0003 via Mann–Whitney test (n = 7 and 9 control and repo, respectively). f Scattergraph showing percentage of macrophages that leave the wound (having initially migrated to the wound); P > 0.999 via Mann–Whitney test (n = 7 and 9 control and repo, respectively). Lines and error bars in scattergraphs show mean and standard deviation, respectively; scale bars represent 20 μm; ns, *** and **** indicate not significant (P > 0.05), P < 0.001 and P < 0.0001, respectively. Genotypes are w;;crq-GAL4,UAS-GFP (control) and w;crq-GAL4,UAS-GFP,repo03702 (repo03702).
a–d Ventral views showing localisation of GFP and red stinger-labelled macrophages on the ventral midline in control (a, a’), Df(3L)H99 (b, b’), repo (c, c’) and Df(3L)H99,repo double-mutant embryos (d, d’) immediately before wounding (a–d) and at 60 min post wounding (a’–d’); white-dotted ellipses indicate wound edges. e Scattergraph showing wound responses quantified via density of macrophages at wounds, normalised to control; Mann–Whitney tests used to compare control vs Df(3L)H99 (n = 15 and 7, respectively; P = 0.63) and repo vs Df(3 L)H99,repo (n = 9 and 15, respectively; P > 0.99). f Scattergraph showing percentage of macrophages responding to wounds; Mann–Whitney tests used to compare control vs Df(3L)H99 (n = 9 and 4, respectively P = 0.055) and repo vs Df(3L)H99,repo (n = 6 and 6, respectively; P = 0.17). Lines and error bars in scattergraphs show mean and standard deviation, respectively; scale bars represent 20 μm; ns indicates not significant (P > 0.05). Genotypes are w;srp-GAL4,UAS-GFP/srp-GAL4,UAS-red stinger (control), w;srp-GAL4,UAS-GFP/srp-GAL4,UAS-red stinger;Df(3L)H99 (Df(3L)H99), w;srp-GAL4,UAS-GFP/srp-GAL4,UAS-red stinger;repo03702 (repo03702) and w;srp-GAL4,UAS-GFP/srp-GAL4,UAS-red stinger;Df(3L)H99,repo03702 (Df(3L)H99,repo03702).
In contrast to the rescue of migration speed, removing apoptosis from a repo mutant background failed to rescue either the numbers of cells responding to the injury or the percentage of cells able to respond (Fig. 6). Therefore, the underlying defect in repo mutants that hinders macrophage inflammatory responses does not appear to be contact with apoptotic cells (Fig. 6), in contrast to the effect seen for migration defects (Fig. 4).
Neurons and glia respond to injury through changes in their calcium dynamics
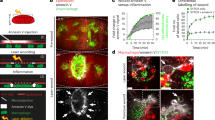
Since removing the apoptotic cell burden from macrophages in repo mutants failed to improve their inflammatory responses to wounds (Fig. 6), we investigated whether upstream signalling mechanisms that form part of the normal response to wounds remain intact in this mutant background. Laser ablation of embryos triggers the rapid spread of a calcium wave through the epithelium, away from the site of damage (Fig. 7a; Supplementary Movie 2; Razzell et al.14). This wave of calcium activates hydrogen peroxide production via DUOX, which is required for migration of macrophages to wound sites14,45,54. Imaging calcium dynamics after wounding using the cytoplasmic calcium sensor GCaMP6M41, we were unable to identify a difference immediately post wounding in the epithelial calcium responses of repo mutant embryos compared with controls (Fig. 7a–d; Supplementary Movie 2). However, in performing these experiments we noticed that the calcium response was not limited to the epithelium, with this signal visible deeper within the embryo, including within the neurons and glia of the VNC (Fig. 7e–g)—a structure located immediately underneath the epithelium on the ventral side of the embryo (Fig. 1a, b). These changes in intracellular calcium were not limited to the damaged tissue and extended away from the necrotic core of the wound (Fig. 7e–g; Supplementary Movie 3), with changes in calcium levels particularly striking within the axons of the CNS (Fig. 7g). Strikingly, quantification of calcium responses in sub-epithelial regions upon wounding showed a reduced calcium response in repo mutants compared with controls (Fig. 8a–c), suggesting that glial cells are responsive to injury and contribute to damage-induced signalling.
a, b Calcium levels imaged via ubiquitous expression of GCaMP6M (da-GAL4,UAS-GCaMP6M) on wounding of the ventral surface of control (a) and repo mutant (b) stage 15 embryos; images show pre-wound calcium levels, immediately after wounding (0 min) and 5 min after wounding. Dotted lines show edges of embryo and wound edges in pre-wound and 5 min images, respectively. c Scattergraph showing area of GCaMP6M response immediately after wounding; Mann–Whitney test used to compare control vs repo (n = 35 and 47, respectively; P = 0.66). d Scattergraph showing ratio of GCaMP6M intensity in wound area before and immediately after wounding; Mann–Whitney test used to compare control vs repo (n = 23 and 26, respectively; P = 0.22). e–g maximum projections of superficial/epithelial regions (0–5 μm, e), superficial half of the VNC (10–20 μm from surface, f) and deeper/dorsal half of the VNC (20–30 μm from the surface, g) of the ventral side of a wounded control embryo containing ubiquitous expression of GCaMP6M; panels show pre-wound calcium levels, immediately after wounding (0 min) and 5 min after wounding. Dotted lines show edges of epithelial wound at 5 min, solid circles show physical damage at deeper regions of the embryo; embryo has not been orientated in order to show larger region of the response to wounding. Lines and error bars in scattergraphs show mean and standard deviation, respectively; scale bars represent 20 μm; ns indicates not significant (P > 0.05). Genotypes are w;;da-GAL4,UAS-GCaMP6M (control) and w;;repo03702,da-GAL4,UAS-GCaMP6M (repo).
a, b Calcium levels within the VNC imaged via expression of UAS-GCaMP6M via e22c-GAL4 (projections assembled from slice range 30–10-μm deep from surface of the embryo) on wounding of the ventral surface of control (a) and repo mutant (b) stage 15 embryos; images show pre-wound calcium levels and immediately after wounding (0 min). c Scattergraph showing GCaMP6M response within VNC immediately after wounding (F1/F0) of control and repo mutants labelled using e22c-GAL4,UAS-GCaMP6M; Mann–Whitney test used to compare control vs repo (n = 37 and 30, respectively; P = 0.0051). d Calcium levels in neuronal cells (labelled using elav-GAL4 to drive UAS-GCaMP6M expression) on wounding of the ventral surface of a stage 15 embryo; images show pre-wound calcium levels, immediately after wounding (0 min) and 5-min after wounding. e–g Calcium levels in glial cells (labelled using repo-GAL4 to drive UAS-GCaMP6M expression) on wounding of the ventral surface of control (e, g) and repo (f) mutant stage 15 embryos; images show pre-wound calcium levels, immediately after wounding (0 min) and 5-min after wounding. g shows embryo wounded more laterally and subsequent spread of calcium signal along glial cells to more lateral positions; dotted line shows equivalent position in the 0 min and 2 min post-wound timepoint images. h, i Scattergraphs showing GCaMP6M responses in glial cells immediately after wounding (h, F1/F0) and the initial area (μm2) of the GCaMP6M response in control and repo mutant embryos labelled via repo-GAL4,UAS-GCaMP6M (i); Mann–Whitney test used to compare control vs repo (n = 16 and 12, respectively; P = 0.42 (h) and P = 0.0003 (i)). Asterisks show position of wounds in pre-wound images; scale bars represent 20 μm; lines and error bars show mean and standard deviation in all scattergraphs; **, *** and ns denote P < 0.01, P < 0.001 and not significant (P > 0.05), respectively. Genotypes are w;e22c-GAL4,UAS-GCaMP6M (a, c), w;e22c-GAL4,UAS-GCaMP6M;repo03702 (b, c), elav-GAL4/w;UAS-GCaMP6M/+ (d), w;repo-GAL4,UAS-GCaMP6M (e, g, h–i) and w;repo-GAL4,UAS-GCaMP6M;repo03702 (f, h, i).
repo is required for normal calcium responses to injury in the developing embryo
Using the tissue-specific drivers elav-GAL4 (neurons) and repo-GAL4 (glia; n.b. repo-GAL4 has been shown to label residual glial cells in a repo null mutant background—this reporter does not require repo function for expression of GAL4)27 to express GCaMP6M confirmed that both neurons and glial cells within the VNC respond to injury by transiently increasing their cytosolic calcium levels (Fig. 8d, e; Supplementary Movies 4 and 5). Changes in cytoplasmic calcium are transmitted beyond the confines of the wound (Fig. 8d, g) and remain elevated at wound sites several hours post injury (Supplementary Fig. 3). These responses suggest that these cells communicate damage signals away from sites of tissue damage, potentially contributing to inflammatory recruitment of macrophages and regenerative processes. In addition, the transmission of calcium signals distal to the site of physical injury suggests that these responses are not simply limited to those cells damaged during the wounding process.
In repo mutants, while there is less glial proliferation, residual glial cells are present, but lack late glial markers21,22 and normal patterns of phagocytic receptor expression25. In contrast to the wild-type situation, wounding of repo mutants with glial-specific expression of GCaMP6M showed a reduction in the calcium response on injury (Fig. 8e, f, h, i). Taken together, these data indicate that, in contrast to our present understanding, tissues other than the epithelium undergo alterations in calcium signalling upon injury. Furthermore, the reduced calcium responses observed in repo mutants on wounding may therefore contribute to the reduced inflammatory responses undertaken by macrophages.
Discussion
Clearance of apoptotic cells is associated with reprogramming of phagocytes such as macrophages towards anti-inflammatory states and is part of the process of resolution of inflammation2. Here, we show that by impairing glial specification using repo mutants, macrophages can be challenged with increased levels of apoptotic cell death in vivo. Challenging ‘wild-type’ macrophages in this way causes them to become engorged with phagocytosed apoptotic cells. The excessive amounts of apoptosis macrophages face in repo mutants appear associated with impaired macrophage developmental dispersal, slowed migration speeds and attenuated inflammatory responses. However, while macrophage migration speeds can be rescued by removing apoptosis from a repo mutant background, wound responses are not improved, suggesting interactions with apoptotic cells do not represent the primary cause of this phenotype. Instead, in contrast to our current understanding, tissues in addition to the wounded epithelium are wound-responsive, and glia and the neurons they support are likely to contribute to immune cell recruitment and repair mechanisms in Drosophila.
Macrophages in repo mutants become highly vacuolated with those vacuoles containing apoptotic cells. At early stages, much apoptotic cell death around the developing VNC corresponds to dying epidermal cells at segmental boundaries55, but by stage 15 most apoptosis is confined to the VNC56. How macrophages access apoptotic cells that repo mutant glia have failed to clear remains to be determined precisely. However, the blood–brain barrier (BBB), which is comprised of glial cells, is yet to form at the stages we have analysed57. Normally macrophages are excluded from the VNC, but can enter this structure in repo mutants at later stages25, suggesting the BBB fails to form in an appropriate manner. Interestingly, active migration of dying neurons has been observed in the zebrafish brain to aid their clearance by macrophages58, and apoptotic cells are expelled towards the periphery of the VNC during development47. Therefore, a similar process may occur in flies facilitating contact between macrophages and apoptotic cells in repo mutants. In the absence of macrophages, glial cells become engorged with apoptotic cells, highlighting the interplay and competition for apoptotic cells between these phagocytes47.
The vacuolated macrophages in repo mutants exhibit impaired dispersal and reduced migration speeds, phenotypes consistent with other Drosophila mutants that perturb apoptotic cell clearance, including SCAR/WAVE51 and simu31. Similarly, removal of the apoptotic clearance burden via ablation of the apoptotic machinery improves migration speeds in these mutant backgrounds. This suggests that apoptotic cells are responsible for these phenotypes and that the ‘migratory substrate' (i.e., the VNC) is sufficiently developed to support normal motility. Structural defects in the VNC are also unlikely to account for perturbed developmental dispersal of macrophages, since we and others have found that VNC defects are not apparent until after completion of macrophage dissemination21,22. Excessive production of find-me cues—chemoattractants released from apoptotic cells to alert phagocytes to their presence59—represents one potential explanation, with such cues distracting or overwhelming macrophages. Consistent with this, apoptotic cells have been inferred to be the most highly prioritised migratory cue for macrophages in the fly embryo53, and the degree of vacuolation does not correlate with decreased migration speeds in repo or simu mutants31, revealing that apoptotic cells need not be engulfed in order to affect migration speeds. Currently, this hypothesis remains difficult to investigate further, since the nature and identity of find-me cues have yet to be discovered in this organism. Alternatively, reprogramming of macrophages to different transcriptional and/or activation states may underlie these phenotypic changes60,61. While little evidence for such reprogramming exists in Drosophila, infection can drive Drosophila macrophage metabolism towards aerobic glycolysis62, resembling pro-inflammatory activation of vertebrate myeloid cells. Another recent paper used single-cell RNA-sequencing to identify subsets of larval blood cells with molecular signatures reminiscent of immune cell activation63. Therefore, emerging data suggest that Drosophila macrophages are more similar to their vertebrate counterparts than previously anticipated and reprogramming via apoptotic cell clearance remains a viable explanation for the changes in their behaviour in repo mutants.
Phagocytes such as Drosophila macrophages may have to decide whether to engulf or move: for instance both dendritic cells and Dictyostelium amoebae pause during macropinocytosis64,65, a process related to phagocytosis. Similarly, lysosomal storage disorders that lead to vacuolation are associated with perturbed immune cell migration in patient-derived cells66,67,68 and experimental models69. Sequestration of regulators of both phagocytosis and motility, such as the lysosomal Trp channel Trpml (the fly homologue of TRPML1/MCOLN1)70, by excessive phagocytic cup formation in the face of elevated apoptotic cell burdens, could antagonise the ability of phagocytes to migrate. repo mutants represent a model to further understand how apoptotic cells regulate changes in macrophage behaviour in vivo, such as their migration and phagocytic capacities, especially since initial specification of macrophages does not appear compromised in this background and repo plays no direct functional role in these cells.
Macrophage inflammatory responses were not improved by blocking apoptosis in repo mutants. This was surprising, since preventing apoptosis in simu mutants, another background in which macrophages face large numbers of apoptotic cells, did improve macrophage responses to injury31. Thus it is unlikely that apoptotic cell–macrophage interactions represent the primary cause of wound recruitment defects in repo mutants. Faster and more sensitive imaging technologies alongside the use of alternative drivers to express genetically encoded calcium reporters enabled capture of larger volumes of the embryo during wounding in comparison to previous studies14. This revealed that those tissues surrounding the damaged epithelium, such as the neurons and glia of the VNC, were also responsive to injury. Loss of repo function causes defects in glial specification, proliferation21,22,23 and also leads to a perturbed calcium response in the VNC and glia. Potentially, repo drives a transcriptional programme enabling glia to respond to injury. Failed glial specification may have a ‘knock-on' effect and hinder the damage responses of neurons, which are supported by glial cells. Alternatively, the decreased number or mispositioning of glial cells in repo mutants21 may decrease the amplitude or spread of wound signals activated by injury, such as via defective cell–cell contacts between glia. Impairing the ability of surrounding tissues to respond to injury may therefore perturb the generation of wound cues required for normal macrophage migration to sites of tissue damage and could potentially impact evolutionarily conserved CNS repair processes71, especially since calcium waves in the developing Drosophila wing disc are required for regeneration following mechanical injury72. While these results mean that repo mutants are less suitable to study apoptotic cell–macrophage interactions after injury, this work has changed our understanding of how wound responses are regulated in Drosophila.
The use of calcium as a regulator of wound responses is conserved across evolution with calcium waves visible upon transection of zebrafish larval fins17, and immediately after wounding Xenopus embryos73 and the C. elegans epidermis74. Release of calcium from internal stores during wave propagation is also a commonality in these models. The mechanism of activation in the Drosophila CNS remains to be established, but in zebrafish tailfin wounds the release of ATP from damaged cells may activate P2Y receptors, leading to subsequent release of calcium from internal stores via PKC signalling75. Calcium waves can also be observed within the developing zebrafish brain upon wounding, with these dependent upon glutamate-mediated activation of NMDA receptors. NMDA receptors in turn regulate ATP-dependent recruitment of microglial cells to sites of injury76. Traumatic brain injury has long been associated with rapid decreases in extracellular calcium (e.g., Young et al.77; for review see Weber78) and laser wounding has been proposed as a useful technique to model this79. Indeed, buffering calcium dynamics within damaged neurons in flies can be neuroprotective80, suggesting this system could uncover novel therapeutic strategies for traumatic brain injury.
In summary, we have shown that increasing the apoptotic cell burden placed on macrophages impairs their normal behaviour, dampening their migratory abilities and impairing their dispersal; repo mutants will thus provide a useful model to understand the numerous functions of macrophages relating to contact with apoptotic cells that do not depend upon intact glial function. Furthermore, we have uncovered that wound-induced calcium waves spread beyond the epithelium into neighbouring tissues, potentially enabling them to contribute to recruitment of macrophages and subsequent repair processes. How these waves spread and how epithelial and non-epithelial tissues integrate these responses to coordinate inflammation and repair remain key questions for future work. Since specification of macrophages is not perturbed in repo mutants, this model will prove useful to examine macrophage–apoptotic cell interactions in vivo in more detail and shed light on cellular interactions that are fundamental for normal development and homeostasis.
Data availability
Supplementary information is available at Cell Death and Disease’s website. Fly lines and raw data are available on request from I.R.E.
References
Morioka, S., Maueröder, C. & Ravichandran, K. S. Living on the edge: efferocytosis at the interface of homeostasis and pathology. Immunity 50, 1149–1162 (2019).
Serhan, C. N. & Savill, J. Resolution of inflammation: the beginning programs the end. Nat. Immunol. 6, 1191–1197 (2005).
Bäck, M., Yurdagul, A., Tabas, I., Öörni, K. & Kovanen, P. T. Inflammation and its resolution in atherosclerosis: mediators and therapeutic opportunities. Nat. Rev. Cardiol. https://doi.org/10.1038/s41569-019-0169-2 (2019).
McCubbrey, A. L. & Curtis, J. L. Efferocytosis and lung disease. Chest 143, 1750–1757 (2013).
Buchon, N., Silverman, N. & Cherry, S. Immunity in Drosophila melanogaster—from microbial recognition to whole-organism physiology. Nat. Rev. Immunol. 14, 796–810 (2014).
Banerjee, U., Girard, J. R., Goins, L. M. & Spratford, C. M. Drosophila as a genetic model for hematopoiesis. Genetics 211, 367–417 (2019).
Ratheesh, A., Belyaeva, V. & Siekhaus, D. E. Drosophila immune cell migration and adhesion during embryonic development and larval immune responses. Curr. Opin. Cell Biol. 36, 71–79 (2015).
Wood, W. & Jacinto, A. Drosophila melanogaster embryonic haemocytes: masters of multitasking. Nat. Rev. Mol. Cell Biol. 8, 542–551 (2007).
Evans, C. J., Hartenstein, V. & Banerjee, U. Thicker than blood. Dev. Cell 5, 673–690 (2003).
Sears, H. C., Kennedy, C. J. & Garrity, P. A. Macrophage-mediated corpse engulfment is required for normal Drosophila CNS morphogenesis. Development 130, 3557–3565 (2003).
Olofsson, B. & Page, D. T. Condensation of the central nervous system in embryonic Drosophila is inhibited by blocking hemocyte migration or neural activity. Dev. Biol. 279, 233–243 (2005).
Defaye, A. et al. Genetic ablation of Drosophila phagocytes reveals their contribution to both development and resistance to bacterial infection. J. Innate Immun. 1, 322–334 (2009).
Stramer, B. et al. Live imaging of wound inflammation in Drosophila embryos reveals key roles for small GTPases during in vivo cell migration. J. Cell Biol. 168, 567–573 (2005).
Razzell, W., Evans, I. R., Martin, P. & Wood, W. Calcium flashes orchestrate the wound inflammatory response through duox activation and hydrogen peroxide release. Curr. Biol. 23, 424–429 (2013).
Antunes, M., Pereira, T., Cordeiro, J. V., Almeida, L. & Jacinto, A. Coordinated waves of actomyosin flow and apical cell constriction immediately after wounding. J. Cell Biol. 202, 365–379 (2013).
Niethammer, P., Grabher, C., Look, A. T. & Mitchison, T. J. A tissue-scale gradient of hydrogen peroxide mediates rapid wound detection in zebrafish. Nature 459, 996–999 (2009).
Yoo, S. K., Starnes, T. W., Deng, Q. & Huttenlocher, A. Lyn is a redox sensor that mediates leukocyte wound attraction in vivo. Nature 480, 109–112 (2011).
Kurant, E. Keeping the CNS clear: glial phagocytic functions in Drosophila. Glia 59, 1304–1311 (2011).
Evans, I. R., Hu, N., Skaer, H. & Wood, W. Interdependence of macrophage migration and ventral nerve cord development in Drosophila embryos. Development 137, 1625–1633 (2010).
Crews, S. T. Drosophila embryonic CNS development: neurogenesis, gliogenesis, cell fate, and differentiation. Genetics 213, 1111–1144 (2019).
Campbell, G. et al. RK2, a glial-specific homeodomain protein required for embryonic nerve cord condensation and viability in Drosophila. Development 120, 2957–2966 (1994).
Halter, D. A. et al. The homeobox gene repo is required for the differentiation and maintenance of glia function in the embryonic nervous system of Drosophila melanogaster. Development 121, 317–332 (1995).
Xiong, W. C., Okano, H., Patel, N. H., Blendy, J. A. & Montell, C. Repo encodes a glial-specific homeo domain protein required in the Drosophila nervous system. Genes Dev. https://doi.org/10.1101/gad.8.8.981 (1994).
Yuasa, Y. et al. Drosophila homeodomain protein REPO controls glial differentiation by cooperating with ETS and BTB transcription factors. Development 130, 2419–2428 (2003).
Shklyar, B. et al. Developmental regulation of glial cell phagocytic function during Drosophila embryogenesis. Dev. Biol. 393, 255–269 (2014).
Bernardoni, R., Vivancos, V. & Giangrande, A. Glide/Gcm is expressed and required in the scavenger cell lineage. Dev. Biol. 191, 118–130 (1997).
Alfonso, T. B. & Jones, B. W. gcm2 promotes glial cell differentiation and is required with glial cells missing for macrophage development in Drosophila. Dev. Biol. 248, 369–383 (2002).
Trébuchet, G. et al. The repo homeodomain transcription factor suppresses hematopoiesis in Drosophila and preserves the glial fate. J. Neurosci. https://doi.org/10.1523/JNEUROSCI.1059-18.2018 (2019).
Melcarne, C., Lemaitre, B. & Kurant, E. Phagocytosis in Drosophila: from molecules and cellular machinery to physiology. Insect Biochem. Mol. Biol. 109, 1–12 (2019).
Kurant, E., Axelrod, S., Leaman, D. & Gaul, U. Six-microns-under acts upstream of draper in the glial phagocytosis of apoptotic neurons. Cell 133, 498–509 (2008).
Roddie, H. G., Armitage, E. L., Coates, J. A., Johnston, S. A. & Evans, I. R. Simu-dependent clearance of dying cells regulates macrophage function and inflammation resolution. PLoS Biol. https://doi.org/10.1371/journal.pbio.2006741 (2019).
Weavers, H., Evans, I. R., Martin, P. & Wood, W. Corpse engulfment generates a molecular memory that primes the macrophage inflammatory response. Cell 165, 1658–1671 (2016).
A-Gonzalez, N. et al. Phagocytosis imprints heterogeneity in tissue-resident macrophages. J. Exp. Med. 214, 1281–1296 (2017).
Brückner, K. et al. The PDGF/VEGF receptor controls blood cell survival in Drosophila. Dev. Cell 7, 73–84 (2004).
Stramer, B. et al. Clasp-mediated microtubule bundling regulates persistent motility and contact repulsion in Drosophila macrophages in vivo. J. Cell Biol. 189, 681–689 (2010).
Yoffe, K. B., Manoukian, A. S., Wilder, E. L., Brand, A. H. & Perrimon, N. Evidence for engrailed-independent wingless autoregulation in Drosophila. Dev. Biol. https://doi.org/10.1006/dbio.1995.1243 (1995).
Ito, K., Awano, W., Suzuki, K., Hiromi, Y. & Yamamoto, D. The Drosophila mushroom body is a quadruple structure of clonal units each of which contains a virtually identical set of neurones and glial cells. Development 124, 761–771 (1997).
Wodarz, A., Hinz, U., Engelbert, M. & Knust, E. Expression of crumbs confers apical character on plasma membrane domains of ectodermal epithelia of Drosophila. Cell 82, 67–76 (1995).
Luo, L., Joyce Liao, Y., Jan, L. Y. & Jan, Y. N. Distinct morphogenetic functions of similar small GTPases: Drosophila Drac1 is involved in axonal outgrowth and myoblast fusion. Genes Dev. https://doi.org/10.1101/gad.8.15.1787 (1994).
Lee, B. P. & Jones, B. W. Transcriptional regulation of the Drosophila glial gene repo. Mech. Dev. 122, 849–862 (2005).
Chen, T.-W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
White, K. et al. Genetic control of programmed cell death in Drosophila. Science 264, 677–683 (1994).
Halfon, M. S. et al. New fluorescent protein reporters for use with thedrosophila gal4 expression system and for vital detection of balancer chromosomes. Genesis 34, 135–138 (2002).
Le, T. et al. A new family of Drosophila balancer chromosomes with a w-dfd-GMR yellow fluorescent protein marker. Genetics 174, 2255–2257 (2006).
Evans, I. R., Rodrigues, F. S. L. M., Armitage, E. L. & Wood, W. Draper/CED-1 mediates an ancient damage response to control inflammatory blood cell migration in vivo. Curr. Biol. 25, 1606–1612 (2015).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Sonnenfeld, M. J. & Jacobs, J. R. Macrophages and glia participate in the removal of apoptotic neurons from the Drosophila embryonic nervous system. J. Comp. Neurol. 359, 644–652 (1995).
Freeman, M. R., Delrow, J., Kim, J., Johnson, E. & Doe, C. Q. Unwrapping glial biology: Gcm target genes regulating glial development, diversification, and function. Neuron 38, 567–580 (2003).
Manaka, J. et al. Draper-mediated and phosphatidylserine-independent phagocytosis of apoptotic cells by Drosophila hemocytes/macrophages. J. Biol. Chem. 279, 48466–48476 (2004).
Nambu, J. R., Franks, R. G., Hu, S. & Crews, S. T. The single-minded gene of Drosophila is required for the expression of genes important for the development of CNS midline cells. Cell 63, 63–75 (1990).
Evans, I. R., Ghai, P. A., Urbančič, V., Tan, K. L. & Wood, W. SCAR/WAVE-mediated processing of engulfed apoptotic corpses is essential for effective macrophage migration in Drosophila. Cell Death Differ. 20, 709–720 (2013).
Song, Z., McCall, K. & Steller, H. DCP-1, a Drosophila cell death protease essential for development. Science 275, 536–540 (1997).
Moreira, S., Stramer, B., Evans, I., Wood, W. & Martin, P. Prioritization of competing damage and developmental signals by migrating macrophages in the Drosophila embryo. Curr. Biol. 20, 464–470 (2010).
Hunter, M. V., Willoughby, P. M., Bruce, A. E. E. & Fernandez-Gonzalez, R. Oxidative stress orchestrates cell polarity to promote embryonic wound healing. Dev. Cell 47, 377–387.e4 (2018).
Pazdera, T. M., Janardhan, P. & Minden, J. S. Patterned epidermal cell death in wild-type and segment polarity mutant Drosophila embryos. Development 125, 3427–3436 (1998).
Rogulja-Ortmann, A., Lüer, K., Seibert, J., Rickert, C. & Technau, G. M. Programmed cell death in the embryonic central nervous system of Drosophila melanogaster. Development 134, 105–116 (2007).
Schwabe, T., Bainton, R. J., Fetter, R. D., Heberlein, U. & Gaul, U. GPCR signaling is required for blood-brain barrier formation in Drosophila. Cell 123, 133–144 (2005).
Van Ham, T. J., Kokel, D. & Peterson, R. T. Apoptotic cells are cleared by directional migration and elmo1-dependent macrophage engulfment. Curr. Biol. https://doi.org/10.1016/j.cub.2012.03.027 (2012).
Medina, C. B. & Ravichandran, K. S. Do not let death do us part: ‘find-me’ signals in communication between dying cells and the phagocytes. Cell Death Differ. 23, 979–989 (2016).
Fadok, V. A. et al. Macrophages that have ingested apoptotic cells in vitro inhibit proinflammatory cytokine production through autocrine/paracrine mechanisms involving TGF-beta, PGE2, and PAF. J. Clin. Invest. 101, 890–898 (1998).
Murray, P. J. Macrophage polarization. Annu. Rev. Physiol. 79, 541–566 (2017).
Krejčová, G. et al. Drosophila macrophages switch to aerobic glycolysis to mount effective antibacterial defense. eLife 8, e50414 (2019).
Cattenoz, P. B. et al. Temporal specificity and heterogeneity of Drosophila immune cells. EMBO J. 39, e104486 (2020).
Chabaud, M. et al. Cell migration and antigen capture are antagonistic processes coupled by myosin II in dendritic cells. Nat. Commun. 6, 7526 (2015).
Veltman, D. M. Drink or drive: competition between macropinocytosis and cell migration. Biochem. Soc. Trans. 43, 129–132 (2015).
Bettman, N., Avivi, I., Rosenbaum, H., Bisharat, L. & Katz, T. Impaired migration capacity in monocytes derived from patients with Gaucher disease. Blood Cells, Mol. Dis. https://doi.org/10.1016/j.bcmd.2014.12.003 (2015).
Morell, G. P., Niaudet, P., Jean, G. & Descamps-Latscha, B. Altered oxidative metabolism, motility, and adherence in phagocytic cells from cystinotic children. Pediatr. Res. https://doi.org/10.1203/00006450-198512000-00023 (1985).
Aker, M., Zimran, A., Abrahamov, A., Horowitz, M. & Matzner, Y. Abnormal neutrophil chemotaxis in Gaucher disease. Br. J. Haematol. https://doi.org/10.1111/j.1365-2141.1993.tb08270.x (1993).
Berg, R. D. et al. Lysosomal disorders drive susceptibility to tuberculosis by compromising macrophage migration. Cell 165, 139–152 (2016).
Edwards-Jorquera, S. S., Bosveld, F., Bellaïche, Y. A., Lennon-Duménil, A.-M. & Glavic, Á. Trpml controls actomyosin contractility and couples migration to phagocytosis in fly macrophages. J. Cell Biol. 219, e201905228 (2020).
Kato, K., Losada-Perez, M. & Hidalgo, A. Gene network underlying the glial regenerative response to central nervous system injury. Dev. Dyn. 247, 85–93 (2018).
Restrepo, S. & Basler, K. Drosophila wing imaginal discs respond to mechanical injury via slow InsP3R-mediated intercellular calcium waves. Nat. Commun. 7, 12450 (2016).
Clark, A. G. et al. Integration of single and multicellular wound responses. Curr. Biol. 19, 1389–1395 (2009).
Xu, S. & Chisholm, A. D. A Gαq-Ca2+ signaling pathway promotes actin-mediated epidermal wound closure in C. elegans. Curr. Biol. 21, 1960–1967 (2011).
de Oliveira, S. et al. ATP modulates acute inflammation in vivo through dual oxidase 1-derived H2O2 production and NF-κB activation. J. Immunol. 192, 5710–5719 (2014).
Sieger, D., Moritz, C., Ziegenhals, T., Prykhozhij, S. & Peri, F. Long-range Ca2+ waves transmit brain-damage signals to microglia. Dev. Cell 22, 1138–1148 (2012).
Young, W., Yen, V. & Blight, A. Extracellular calcium ionic activity in experimental spinal cord contusion. Brain Res. https://doi.org/10.1016/0006-8993(82)90677-1 (1982).
Weber, J. Calcium homeostasis following traumatic neuronal injury. Curr. Neurovasc. Res. https://doi.org/10.2174/1567202043480134 (2005).
Shannon, E. K. et al. Multiple mechanisms drive calcium signal dynamics around laser-induced epithelial wounds. Biophys. J. 113, 1623–1635 (2017).
Avery, M. A. et al. WldS prevents axon degeneration through increased mitochondrial flux and enhanced mitochondrial Ca2+ buffering. Curr. Biol. 22, 596–600 (2012).
Acknowledgements
We thank Will Wood (University of Edinburgh) for advice and support, Frederico Rodrigues (University of Bristol) for performing preliminary experiments and Karen Plant (University of Sheffield) for technical support. Imaging work was performed in the Wolfson Light Microscopy Facility, using the Perkin Elmer spinning disk (MRC grant G0700091 and Wellcome grant 077544/Z/05/Z) and Nikon A1 confocal/TIRF (Wellcome grant WT093134AIA) microscopes. This work would not be possible without the Bloomington Drosophila Stock Centre (NIH P40OD018537) and Flybase (NIH and MRC grants U41 HG000739 and MR/N030117/1, respectively). We thank the Drosophila community for sharing Drosophila reagents (see Supplementary Table 2). We are grateful to Darren Robinson (Wolfson LMF) and the Fly Facility staff (University of Sheffield) for their assistance and Phil Elks, Simon Johnston, Steve Renshaw and Martin Zeidler (University of Sheffield) for critical reading and feedback on the paper. This work was supported by a Sir Henry Dale Fellowship awarded to I.R.E. by Wellcome and The Royal Society (102503/Z/13/Z) and a University of Sheffield Medical School PhD position awarded to I.R.E. and H.G.R.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by B. Joseph
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Armitage, E.L., Roddie, H.G. & Evans, I.R. Overexposure to apoptosis via disrupted glial specification perturbs Drosophila macrophage function and reveals roles of the CNS during injury. Cell Death Dis 11, 627 (2020). https://doi.org/10.1038/s41419-020-02875-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-020-02875-2