Abstract
Increasing cell mobility is the basis of tumor invasion and metastasis, and is therefore a therapeutic target for preventing the spread of many types of cancer. Septins are a family of cytoskeletal proteins with GTPase activity, and play a role in many important cellular functions, including cell migration. SEPT9 isoform 1 protein (SEPT9_i1) has been associated with breast tumor development and the enhancement of cell migration; however, the exact mechanism of how SEPT9_i1 might affect breast cancer progression remains to be elucidated. Here, we report that the expression of SEPT9_i1 positively correlated with paxillin, and both were significantly upregulated in invasive breast cancer tissues of patients with lymph node metastases. Lentivirus-mediated shRNA knockdown of SEPT9 in MCF-7 cells diminished tumor cell migration, focal adhesion (FA) maturation and the expression of β-actin, β-tubulin, Cdc42, RhoA, and Rac, whereas overexpression of SEPT9_i1 in SEPT9-knockdown MCF-7 cells promoted cell migration, FA maturation and relevant protein expression. Furthermore, overexpression of SEPT9_i1 in MCF-7 cells markedly increased FAK/Src/paxillin signaling, at least in part through RhoA/ROCK1 upstream activation. Transcriptome profiling suggested that SEPT9_i1 may directly affect “Focal adhesion” and “Regulation of actin cytoskeleton” signaling mechanisms. Finally, overexpression of SEPT9_i1 markedly enhanced lung metastases in vivo 6 weeks after tumor inoculation. These findings suggest that a mechanism of Septin-9-induced aberrant cancer cell migration is through cytoskeletal regulation and FA modulation, and encourages the use of SEPT9 as novel therapeutic target in the prevention of tumor metastasis.
Similar content being viewed by others
Introduction
Distant metastasis of a primary tumor is the major cause of mortality in patients with breast cancer1. Some of the key steps in cancer metastasis are the migration and invasion of tumor cells, and these processes require rearrangement of the cytoskeleton and abnormal cell adhesion. The Rho family of GTPases are master regulators of cytoskeleton, and play key roles in assembly and maintenance of cell–cell contacts and cell migration. Given that Rho GTPases are required for cadherin and integrin function, their activity must be regulated to transition between cell–cell adhesion and cell migration2. Focal adhesions (FAs) are large macromolecular complexes that link the actin cytoskeleton to the extracellular matrix (ECM) to provide traction, and are critical for cell migration3,4. Focal adhesion kinase (FAK), a non-receptor tyrosine kinase, also known as protein tyrosine kinase 2, is a key regulator of growth factor receptor- and integrin-mediated signals, governing fundamental processes in normal and cancer cells through its kinase activity and scaffolding functions5. FAK is comprised of multiple protein binding domains, and tyrosine-phosphorylation of several key residues determine protein recruitment, subcellular localization, and catalytic activity of FAK5. FAK can regulate cytoskeletal dynamics and cell motility by influencing actin polymerization and FA turnover6,7. Studies have shown that increased FAK expression and activity in many cancers is often associated with metastasis and poor clinical outcome, highlighting FAK as a potential determinant of tumor development and cancer progression8,9.
Septins are a highly conserved family of GTP binding proteins that have been recently identified as a fourth major component of the cytoskeleton, interacting with microtubules and microfilaments, and linked to a broad spectrum of cellular functions10. SEPT9, one of the 13 members of septins, located at the Chr. 17q25 locus and is directly implicated in actin dynamics, angiogenesis, bacterial autophagy, cell motility, cell proliferation, cell shape, cytokinesis, microtubule regulation, vesicle targeting, and exocytosis11,12,13,14,15. Alternative splicing results in 18 spliced mRNA variants of SEPT916, and both its expression levels and isoform composition differ among cell types17. SEPT9_i1, one of the largest isoforms, has already been involved in ovarian cancer18, head and neck cancers19, and breast cancer progression14,20.
Although it has been reported that there may be an association between SEPT9_i1 and tumor metastasis14,20, the mechanism(s) by which SEPT9_i1 directly affects tumor progression remain unknown. Here, we demonstrate that SEPT9_i1 promotes breast cancer cell migration, cytoskeletal rearrangements and FA turnover by targeting Rho/ROCK and FAK signaling in vitro and increases the metastatic potential of breast cancer in vivo.
Materials and methods
Cell culture
The human breast cancer cell line MCF-7 was purchased from the Cell Bank of the Chinese Academy of Sciences (Shanghai, China); in addition, HEK293FT cells from Invitrogen/Life Technologies (USA) were used for lentivirus production. MCF-7 cells were incubated at 37 °C and grown in Dulbecco’s Modified Eagle’s Medium (DMEM, Gibco, USA) supplemented with 10% fetal bovine serum (FBS; Biowest, France) and 1% penicillin/streptomycin (HyClone, USA). HEK293FT cells were cultured at 37 °C according to the manufacturer’s instructions. Chemical inhibitors to FAK (10 μM; PF-573228) and ROCK (10 μM; Y-27632) were purchased from Selleck Chemicals (Houston, TX, USA).
Tissue samples
Paraffin-embedded invasive breast cancer tissues and information from 55 female patients that underwent mammectomy were obtained from the Department of Pathology in the Affiliated Hospital of Southwest Medical University from September 2015 to October 2016. None of the patients had received chemotherapy or radiotherapy before surgery. The study complied with the guidelines of the ethics committee of the Affiliated Hospital of Southwest Medical University. Protein expression of SEPT9 and paxillin in slices were detected by immunohistochemistry as detailed below.
Immunohistochemical (IHC) staining and data analysis
IHC staining was performed on paraffin sections (4 µm) which were dried for 30 min at 60 °C, dewaxed with xylene, and rehydrated through graded alcohol series. Antigen retrieval was then conducted in citrate buffer (pH 6.0) at 98 °C for 15 min, and endogenous peroxidase was inactivated with 0.3% H2O2 for 10 min at room temperature. The sections were incubated with rabbit polyclonal anti-SEPT9 (1:150, Abcam, USA) and anti-paxillin antibodies (1:100, Abcam, USA) at 4˚C overnight. After being washed in PBS, the sections were incubated 30 min at room temperature with HRP-conjugated secondary antibody. The immunoperoxidase signal was developed with 3,3’-diaminobenzidine (DAB) solution, and the sections were counterstained with hematoxylin.
Immunohistochemical staining was evaluated at ×400 magnification and scored by two independent pathologists. The staining intensity of tumor cells were categorized as follows: negative (score 0), weak (score 1), moderate (score 2), and strong (score 3). The percentage of positive staining tumor cells was scored as follows: 0 (~5%), 1 (5–25%), 2 (26–50%), 3 (51–75%), and 4 (75–100%). The protein expression of every slice was assessed by using the sum of the staining intensity score and percentage score. Next, this protein expression score was categorized into two groups: 0–3 (low protein expression group) and 4–6 (high protein expression group). The Pearson correlation coefficient was calculated as a measure of correlation between expression of SEPT9 protein and clinicopathological parameters (such as age, tumor size, and the presence of metastases) in breast cancer patients.
Generation of MCF-7 SEPT9 knockdown and overexpression cell lines
Inverted and self-complementary hairpin structure single stranded nucleotide targeting SEPT9 mRNA and a scrambled negative control oligonucleotide were designed and synthesized by Sangon Biotechnology Company (Shanghai, China). The sequences were as follows:
SEPT9-shRNA, sense: 5′-CCGGAACACCACACACTGTGAGTTTCTCGAG AACTCACAGTGTGTGGTGTTTTTTTG- 3′
scrambled shRNA, sense: 5′-CCGGAACTCCGAACGTGTCACGTTTCTCGA G AACGTGACACGTTCGGAGTTTTTTG-3′
All above sense and antisense oligos were synthesized, annealed, and ligated into a linearized pLKO.1-TRC cloning vector (Addgene, USA). The lentiviral vector was cotransfected with packaging vectors (pLP1, pLP2, and pLP-VSVG) (Invitrogen, USA) into HEK293FT cells. Supernatants containing lentiviruses were harvested 40 h after transfection, and lentiviruses were used to transduce MCF-7 cells. One day after the transduction, puromycin (InvivoGen, USA) was added to the cell culture medium at the final concentration of 1 μg/ml to generate stable MCF-7 cell lines. MCF-7 cells with stably low expression of SEPT9 were diluted and seeded in 96-well plates at 1 cell/well to isolate monoclonal cell lines that were termed as SEPT9-KD1 and SEPT9-KD2, respectively. Other MCF-7 cells were transfected with lentiviral vectors carrying scrambled shRNA (SEPT9-Control). SEPT9-Overexpression cell lines were generated by transduction of SEPT9-KD1 and SEPT9-KD2 cells with lentiviruses harboring full-length SEPT9_i1-C4Flag rescue gene which was under synonymous mutations in cDNA sequences complementary to the above shRNA sequences avoiding its degradation by the shRNA, and a blasticidin resistance gene. The mutant sequences were as follows: CAC AAC GCA TTG CGA ATT C. After 48 h, the transduced cells were selected by growing cells in DMEM growth medium containing 10 μg/ml blasticidin (InvivoGen, USA), and the expression of the rescue gene was verified by western blot analysis and immunofluorescence staining. The cell lines that rescued SEPT9_i1-C4FLAG in SEPT9-KD1 and SEPT9-KD2 cells were termed as SEPT9-Ov1 and SEPT9-Ov2, respectively.
Wound healing assay
A wound healing (scratch) assay was used to evaluate the ability of SEPT9 to promote migration of MCF-7 cells. Cell lines were seeded into 6-well dishes and grown to 80–90% confluence. A sterile 200 μl pipette tip was used to generate a single wound/scratch across the cell monolayer. The cellular debris from the scraped surface was washed with PBS, and DMEM medium with 2% FBS was added to each well. The images of the cells were taken at 0 h, 24 h, 48 h, and 72 h using phase-contrast microscope. The migration ability of the cells was determined by measuring the width of the monolayer wound for three fields per treatments at 24, 48, and 72 h after scraping, and the migration index was calculated using the following formula, where ‘T’ stands for time (24, 48, or 72 h).
The data shown represent the mean and standard deviation of three independent experiments.
Transwell assays
For migration assays, each cell line was seeded at 5 × 104 in serum-free medium per 8 μm-pore cell culture insert (Corning, USA) that fit into 24-well migration chambers. 500 μl growth medium was added outside of the insert, and plates were incubated at 37 °C for 24 h. The non-migrated cells were scraped from the inner side of the insert, and migrated cells, presenting on the underside of the upper chamber, were fixed with 4% paraformaldehyde (PFA) in PBS and stained with 2% crystal violet-stained. The migrated cells were then photographed under a light microscope at ×100 and were counted in five fields per insert.
Mice and tumor studies
All animal experiments were performed in accordance with Declaration of Helsinki21 in addition to approval granted by the Animal Ethics Committee of Southwest Medical University. Four-week-old female BALB/c nude mice (West China experimental animal center of Sichuan university) were randomized into three groups (n = 6): Control (SEPT9-Control cell line injected), Knockdown (SEPT9-KD1 cell line injected), Overexpression (SEPT9- Ov1 cell line injected). For orthotopic injections, the cells (1 × 106 cells per mouse) were injected into the mammary fat pads of mice. Tumors, lungs, and livers were harvested 6 weeks post-orthotopic injection, fixed in 10% neutral buffered formalin (NBF), embedded in paraffin, sectioned, and stained with hematoxylin and eosin (H&E). For tail vein injections, 5 × 105 cells were injected per mouse, and livers and lungs were harvested at 3 weeks post-injection and examined microscopically by H&E staining. Image J was used to segment and count metastases on liver and lung sections per mouse for both spontaneous and experimental metastasis assays.
Transcriptomic analysis
RNAs from two of the cell lines (SEPT9-KD1 and SEPT9-Ov1) were extracted using Trizol reagent according to manufacturers instructions (Invitrogen, USA); to ensure quality of the RNA samples before sequencing, the NanoDrop 2000 (Thermo Fisher Scientific, USA) and RNA Nano 6000 Assay Kit from the Agilent Bioanalyzer 2100 system (Agilent Technologies, CA, USA) were used to assess the purity, concentration, and integrity of the RNA samples. Transcriptomic analysis was accomplished by Biomarker Technologies (Beijing, China). Differential expression analysis of two conditions/groups was performed using the DESeq R package (1.10.1). The resulting P values were adjusted using the Benjamini and Hochberg’s approach for controlling the false discovery rate (FDR). Genes with an adjusted P-value < 0.05 found by DESeq were assigned as differentially expressed. Differential expression analysis of two samples was performed using the EBSeq R package. The resulting FDRs were adjusted using the posterior probability of being DE (PPDE). The FDR < 0.05 & |log2 (fold change) | ≥ 1 was set as the threshold for significantly differential expression, and the statistical enrichment of differential expression genes in KEGG pathways was implemented by the KOBAS 3.0 software22.
RNA isolation and real-time PCR
1.2 × 106 cells were seeded in 6-well plates, and total RNA was extracted using Trizol reagent (Invitrogen, USA) one day later. First strand cDNA was synthesized using MLV reverse transcriptase (Invitrogen, USA) and random primers. Quantitative PCR amplification was conducted using AceQ qPCR SYBR Green Master Mix (Vazyme, Nanjin, CHN) with primers specific for human ACTB (sense: 5′-CTCCATCCTGGCCTCGCTGT-3′; antisense: 5′-GCTGTCACCTTCACCGTTCC-3′), CDC42 (sense: 5′-ACGACCGCTGAGTTATCCACAAAC-3′; antisense: 5′-ATACTTGACAGCCTTCAGGTCACG-3′)23 and SRC (sense: 5′-CCAGATTGTCAACAACACAGAG-3′; antisense: TCTGACTCCCGTCTGGTGAT-3′). The amplification conditions were: 5 min at 95 °C, 45 cycles of 10 s at 95 °C, 30 s at 53–55 °C, and 40 s at 72 °C, which was performed on a Bio-Rad PCR system (Bio-Rad, CFX96, USA). The relative mRNA expression level was calculated using 2−ΔΔCt method as described previously24, with GAPDH as the internal control.
Immunofluoresence analysis
Cells were fixed with 4% paraformaldehyde for 15 min at room temperature, permeabilized with 0.2% Triton X-100 (EMD Millipore, GER) in PBS for 15 min, and blocked with 5% BSA for 2 h. Then, cells were incubated with primary antibodies against SEPT9 (Abcam, USA), paxillin (Abcam, USA) and Flag ((Santa Cruz, USA), respectively at 4 °C overnight, followed by incubation for 2 h at room temperature with Alexa Fluor® 488 or 568-conjugated secondary antibodies (Invitrogen/Life technologies, USA). Nuclei were stained with DAPI (Sigma), and F-actin was stained with CF568-Phalloidin (Biotium, USA). Images were acquired using the Laser Scanning Confocal Microscope (Olympus, Japan) at ×600 magnification, and the quantification of FA size and number were quantified with ImageJ (NIH, Bethesda, Maryland, USA) with three fields per well and at least five wells per experment.
Western blotting and COIP analyses
Cells were collected using a cell scraper and suspended in RIPA buffer supplemented with protease inhibitor cocktail and phosphatase inhibitor (Sigma Aldrich, USA). Protein samples were separated by SDS-PAGE and electrically transferred to a PVDF membrane (EMD Millipore, GER). The membrane was then blocked with 5% BSA in TBST (TBS, pH 7.4, 0.2% Tween-20) and incubated at 4 °C overnight with primary antibodies for SEPT9 (Abcam, USA), CDC42 (CST, USA), Rac1/2/3 (CST, USA), t-FAK (CST, USA), p-FAK Y397, Y576/577, Y925 (CST, USA), t-paxillin (Abcam, USA), p-paxillin Y118 (CST, USA), t-Src (CST, USA), p-Src Y419 (CST, USA), p-SrcTyr527 (CST, USA), RhoA (CST, USA), t-ROCK (CST, USA), p-ROCK1 T455 + S456 (CST, USA), β-actin (Santa Cruz, USA), and GAPDH (Santa Cruz, USA). The next day, the membrane was washed 3 times with TBST and incubated with Dylight 680-conjugated secondary antibodies (KPL, USA) in the dark for 2 h.
For COIPs, cells were lysed in cold RIPA lysis buffer supplemented with phosphatase and protease inhibitors (Sigma Aldrich, USA). Lysates were pre-cleared 1 h with binding control protein A/G-agarose beads and incubated overnight with paxillin-Trap agarose beads (Yeasen, USA) at 4 °C. Beads were washed in IP lysis buffer, and bound protein was eluted. All membrane images were acquired with the Li-Cor Odyssey Clx Infrared Imaging System (LI-COR Biotechnology, Lincoln, NE, USA) and densitometry analysis of bands was performed using ImageJ software.
Statistical analysis
Each experiment was performed at least 3 times. Values are shown as means ± SEM. Significance was determined by unpaired Student’s t-test for two-group comparisons and one-way ANOVA for comparisons of more than two groups. Pearson’s correlation analysis was used to analyze the correlation between two indices. P < 0.05 was considered statistically significant.
Results
A significant positive correlation between SEPT9 and nodal metastasis and the expression of paxillin in breast cancer tissues
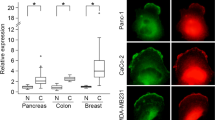
In this study, we first examined the correlation of SEPT9 with clinical characteristics and paxillin in breast cancer samples. Our results showed that SEPT9 and paxillin were overexpressed in 74.5% (41/55) and 54.5% (30/55) of the primary breast cancer clinical samples, respectively (Table 1). The expression levels of SEPT9 and paxillin were significantly increased in the lymph node metastasis group compared to the non-lymph node metastasis group (Fig. 1), and SEPT9 expression was positively correlated with lymph node metastasis (γ = 0.367, P = 0.006). Moreover, there was a positive correlation between SEPT9 and paxillin in the cancer tissues (γ = 0.665, P = 0.000; Table 1).
SEPT9_i1 expression influences the migration of MCF-7 cells in vitro
To examine how SEPT9_i1 functions in the motility of MCF-7 cells in vitro, we inhibited the expression of SEPT9 in MCF-7, which is a low metastatic breast cancer cell line. The two clones stably expressing shSEPT9 (KD1 and KD2) exhibited over 70% protein reduction, while the SEPT9_i1 overexpressed cell lines (Ov1 and Ov2) showed more than tripled compare with scrambled control group (Fig. 2a). Unlike control cells, SEPT9 knockdown MCF-7 cells did not spread out or form protrusions, and SEPT9_i1 overexpression caused markedly altered cell morphology with increased formation of lamellipodia and filopodia, which corroborates previous observations25 (Fig. 2b). Furthermore, SEPT9-Knockdown cells exhibited decreased motility and migration, while SEPT9_i1overexpressed cells exhibited significantly increased motility and migration compared with the scrambled control cells (Fig. 2c, d). These data confirmed that SEPT9_i1 is important for the motility and migration of MCF-7 cells in vitro (Knockdown represents the data from KD1 and Overexpression represents Ov1in these figures, the same below. The data from KD2 and Ov2 showed in Supplementary Fig. 1)
a Western blot for SEPT9_i1 in control, two SEPT9-Knockdown MCF-7 cell lines and two SEPT9-Overexpression cell lines (***p < 0.001). b Phase-contrast microscope images of control, SEPT9-knockdown and SEPT9-Overexpression cells with altered cell morphology (×200). c The motility of cells was measured by a wound healing assay (×40) (*P < 0.05, compared with control group). d Transwell migration assays demonstrating the migration potential of control, SEPT9-Knockdown and SEPT9- Overexpression cells (×100) (*p < 0.05, significant difference versus control; △p < 0.05, significant difference versus knockdown)
Overexpression of SEPT9_i1 promotes metastasis of tumor cells to the lungs
To investigate the effects of SEPT9_i1 expression on breast tumor metastasis in vivo, we injected the engineered MCF-7 cells either into mammary fat pads or the general circulation (via tail vein injection) of BALB/c female nude mice. Palpable mammary tumors were observed within 6 weeks when cells were inoculated into mammary fat pads (Fig. 3a), but no obvious metastases were found in lung or liver sections in mice injected with any of the cell lines (Fig. 3b). However, 3 weeks after tail vein injections, many macro-metastases were evident in lung sections from mice with SEPT9-Overexpression tumors but not mice with SEPT9-Knockdown and scrambled control tumors (Fig. 3c). These data indicate that SEPT9_i1 might promote tumor cell survival and motility in the circulation or metastatic outgrowth at secondary sites, but do not assist escaping from the primary tumor.
a 1 × 106 cells per group were injected into the mammary fat pads of nude mice. Shown are representative tumors that formed 6 weeks post-injection. b Representative H&E-stained sections of lungs and livers of mice with control, knockdown and overexpression group tumors 6 weeks post-orthotopic injection. c Representative H&E-stained sections of lungs and livers of mice with control, knockdown and overexpression group tumors 3 weeks after tail vein injection
RNA-seq highlights SEPT9-induced transcriptional changes in FA and actin cytoskeleton regulation
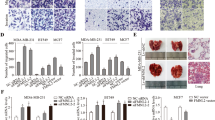
RNA-seq of two cell lines (SEPT9-KD1 and SEPT9-Ov1) was performed. Using FC (Fold change) > 2 and FDR < 0.05 as the statistical cutoffs, we identified 331 differentially expressed genes (DEGs) between SEPT9-KD1 and SEPT9-Ov1 cell lines. Among these genes, 79 genes were upregulated with SEPT9_i1 overexpression, and 252 genes were down-regulated (Fig. 4a). The cellular processes affected by SEPT9 overexpression were evaluated by mapping the DEGs to reference canonical pathways in the KEGG (Kyoto Encyclopedia of Genes and Genomes) database. Eleven DEGs were categorized in the “FA” pathway and five DEGs were in the “Regulation of actin cytoskeleton” pathway (Fig. 4b). Changes in these two pathways are commonly observed during cell migration and changes in cell motility4,6, so these categorical annotations provided evidence of a role for SEPT9 in established signaling mechanisms. The heatmap for these and some cancer related DEGs was shown in Fig. 4c. β-actin, CDC42 and SRC mRNA were proved to conform to the changes of DEGs by real-time RT-PCR (Fig. 4d).
a Volcano plots for all the genes comparing SEPT9-KD1 and SEPT9- Ov1. The red and green dots indicated up- and down-regulated DEGs with FDR < 0.05, while black dots indicated those without statistical significance. b KEGG pathway terms enriched by DEGs between SEPT9-KD1 and SEPT9- Ov1 cell lines, here we listed reference canonical pathways (all p < 0.05) belonging to “Cellular Process” and “Environmental Information Processing”. c Heatmap plot of some related differentially expressed genes across all samples. Columns represent genes and rows represent samples. d β-actin, CDC42 and SRC mRNA expression were analyzed by real-time RT-PCR (***p < 0.001, *p < 0.05)
SEPT9_i1 influences the expressions of GTPases and interacts with stress fibers, microtubules
Confocal microscopy and 3D image reconstruction were used to more precisely examine the effects of SEPT9_i1 expression on the cytoskeletal network of MCF-7 cells. The actin meshwork was thin and devoid of organized stress fibers in the central region of SEPT9-Knockdown cells rather than in control and SEPT9-Overexpression cells (Fig. 5a). Similarly, a network of microtubules also displayed thin and disorder in SEPT9-Knockdown cells (Fig. 5b). In addition, septin-9 appeared to be partially co-localized with stress fibers and microtubules in SEPT9-Ov and scrambled control cells obviously (Fig. 5a, b). Western blotting showed that gene and protein expression of β-actin are down-regulated in SEPT9 knockdown cells as well as β-tubulin protein (Fig. 5c, d). Additionally, GTPases Cdc42, Rac and RhoA, which are critical for actin network organization, were all decreased in SEPT9-KD cells compared to control, and increased in SEPT9-Ov cells (Fig. 5e). Taken together, the possible interaction of septin-9 with F-actin and microtubules, combined with the altered expression of β-actin and β-tubulin, Cdc42, Rac, RhoA in SEPT9 engineered cells lends insight into mechanisms by which SEPT9_i1 may contribute to organization of cytoskeleton in breast cancer cells.
a Confocal images of septin-9 (green) and actin stress fibers (phalloidin; red) in control, knockdown and overexpression group cells. 3D reconstruction reveals that septin-9 filaments colocalize with F-actin stress fibers. b Confocal images of septin-9 (green) and microtubules (red) in control, knockdown and overexpression group cells. It reveals that septin-9 filaments colocalize with microtubules. c, d β-actin and β-tubulin protein expression were analyzed by western blot (*p < 0.05). e Cdc42, Rac and RhoA protein expression were analyzed by western blot
SEPT9_i1 interacts with paxillin and promotes maturation of FAs
Analysis of paxillin IF in MCF7 cells revealed that the size and number of FAs in SEPT9 knockdown cells were reduced compared to the SEPT9-Ov line, which resulted in large, membrane-associated FAs (Fig. 6a, b). We also observed that part of SEPT9_i1 fibers closer to the cell edge directly connected to FAs same as the stress fibers (Fig. 6a). COIP assay discovered that SEPT9_i1 interacted with paxillin (Fig. 6c), and western blotting assays revealed that paxillin protein were decreased in SEPT9-KD cells, and SEPT9-Ov increased expression compared with scrambled control cells (Fig. 6d). Combined with observations of SEPT9_i1-induced changes in actin and microtubules cytoskeleton, these data indicate that SEPT9_i1-mediated FA maturation may be related to the organization of the cytoskeleton, and expression and localization of paxillin. (Supplementary Fig. 2)
a IF of FA proteins paxillin in control, SEPT9-KD and SEPT9-Ov cells. IF of SEPT9_i1 (red) and paxillin (green) in SEPT9-Ov cells. b Quantification of FAs and average FA size per nucleus (*p < 0.05, vs control group; △p < 0.05, vs Knockdown group). c Interaction between SEPT9_i1 and paxillin by COIP. d Western blot confirming the inhibition in the expression of paxillin in knockdown group and overexpression of SEPT9_i1 promoting the expression of paxillin (*p < 0.05, ***p < 0.001)
Overexpression of SEPT9 in MCF-7 cells activates RhoA/ROCK1 and FAK/Src/paxillin signaling, and promotes cell migration
FAK is a non-receptor tyrosine kinase linked to FA dynamics26 and the FAK/Src/paxillin signaling cascade is known to be involved in several cellular processes, including migration and adhesion27. In addition, RhoA/ROCK1 signaling plays a key role in stabilizing actin filaments and regulation of the actin cytoskeleton28. Given our observations of an association between SEPT9 and actin, we investigated proteins in the FAK/Src/paxillin and RhoA/ROCK1 signaling pathways, as well as their phosphrorylation. We found that SEPT9 knockdown abolished activation of the FAK-Src-paxillin pathway, where p-FAK (Y397, Y576/577 and Y925), p-Src (Y416), p-paxillin (Y118) (Fig. 7a), and p-ROCK1 (T455 + S456) (Fig. 7b) were all inhibited compared to control cells (Supplementary Fig. 3). Compared to the total proteins, phosphorylation of FAK and ROCK1 were significantly higher in SEPT9_i1 overexpression cells than in control or SEPT9-Knockdown cells (Fig. 7c). These results suggest that SEPT9_i1 can mediate activation of the FAK/Src/paxillin and RhoA/ROCK1 pathways, which could affect cell migration.
a–c Cell lysates from Control, two cell lines with SEPT9 knockdown and two cell lines with SEPT9_i1 overexpressed were immunoblotted for total FAK, Src, paxillin, RhoA, ROCK1, their phosphorylated protein levels and phosphorylation ratio. d Effects of PF-573228 and Y-27632 on the expression of FAK-Src-paxillin and RhoA-ROCK1 signaling related proteins (*p < 0.05, ***p < 0.001)
To shed light on the upstream and downstream relationship between FAK/Src/paxillin and RhoA/ROCK1 signaling in SEPT9-mediated MCF-7 cell migration, a selective FAK inhibitor, PF-573228, and a ROCK inhibitor, Y-27632, were used in SEPT9 overexpression MCF-7 cells. The transwell and wound healing assays revealed that the migration and motility of the cells were both inhibited after treatment with the inhibitors, and both the number and size of FAs in these cells were reduced (Supplementary Fig. 4). Additionally, the FAK/Src/paxillin signaling pathway was inactivated by Y-27632, while the activity of RhoA-ROCK1 signaling was not affected after treatment with the FAK inhibitor PF-573228 (Fig. 7d). These data indicated that RhoA-ROCK1 might play a role upstream of FAK-Src-paxillin signaling.
Discussion
SEPT9_i1, a product of SEPT9 transcript variant 1 (SEPT9_v1) was shown to be frequently increased in breast tumors, and has already been involved in these tumors progression29. In this study, we used shRNA lentivirus vector to knockdown the expression of SEPT9 isoforms in MCF-7 cells significantly including SEPT9_i1, and the reduced cell motility, cell migration and the formation of invadopodia were detected. These effects were recovered when rescued the expression of foreign SEPT9-i1 specially, and these results agree fairly well with the findings in human prostate cancer cells30 and immortalized murine embryonic fibroblasts in mice homozygous deletion of SEPT931. Importantly, although SEPT9_i1 overexpression cells were unable to metastasize successfully from an orthotopic site, they were able to colonize the lung when injected directly into the circulation, which indicated that SEPT9_i1 can promote the survival and migration in the circulation and outgrowth in the lung.
Cell migration requires reorganization of actin and microtubules, which plays a critical role in cancer metastasis32. SEPT9 has been shown to be involved in the reconstruction process by interacting with actin filaments directly and indirectly33,34. A recent study demonstrated that septin-9 binds to F-actin in a highly polymorphic fashion and protects actin filaments from depolymerization by cofilin and myosin, hence, SEPT9 could maintain the integrity of growing and contracting actin filaments34. However, actin filaments were on average slightly thinner in SEPT9del/del cells compared to SEPT9cond/cond cells31. This study showed that septin-9 partially co-localized with stress fibers; when SEPT9 was knocked down, actin fibers were reduced and positioned to the cortical edge of the cell, while SEPT9_i1 overexpression substantially increased stress fibers. Based on these observations, we hypothesized that if the protective effect of SEPT9_i1 fibers on F-actin was reduced by knockdown of SEPT9, then the depolymerization of F-actin would be enhanced. However, when the protective effect was recovered with overexpression of SEPT9, the depolymerization of F-actin was reduced and stability of F-actin was enhanced. In mammalian cells, septin depletion affects MT organization, dynamics, and post-translational modifications35,36. SEPT9_i1 has already been proposed to participate in the resistance to MT-disrupting agents37, and cell resistance to paclitaxel38. It also is the only septin in which specific repeated motifs might allow MT binding and bundling39. We observed the coalignment between SEPT9_i1 and microtubule, and also showed microtubule organization was altered after SEPT9 knockdown, and recovered when SEPT9_i1 overexpressed. Interestingly, the expression of β-actin and β-tubulin was decreased in SEPT9-knockdown cells; whether this is a direct or indirect association is unclear and requires further mechanistic studies.
Cell migration also consists of well-defined steps that include extension of the leading edge and the formation of immature FAs; FA maturation and cell body translocation; and the FA disassembly and rear retraction40. During this cycle, FA dynamics and cytoskeletal organization conjoin to drive this coordinated process4. The actin cytoskeleton assembles into branched networks or bundles to generate mechanical force for critical cellular processes such as establishment of polarity, adhesion, and migration41. FAs undergo a maturation process, which is known to be induced by biochemical or physical stimuli, to vary FA size, quantity, distribution, dynamics, and composition40. This process is regulated by tension forces from actin stress fibers40,42,43. Microtubules are also involved in the organization and turnover of FAs44,45. Microtubules play an important role in the delivery of proteins to subcellular locations including the FAs and they are critical in paxillin assembly in cell periphery and lamellipodia46. Recent research indicates that in the leading edge of motile renal epithelia, a novel network of septin filaments promotes cell motility by reinforcing the organization of the lamellar stress fiber network and the stability of nascent FAs33. Our work adds to a growing body of work linking SEPT9, F-actin, microtubules and FAs to tumor cell motility, migration, and metastasis, especially the link between SEPT9_i1 and paxillin was first suggested by IF and COIP analysis. Whether the interaction between SEPT9_i1 and paxillin can influence the stability of paxillin protein, and then caused the difference expression of paxillin protein similar to Hif-1α47, We will make further research. Furthermore, we had revealed that SEPT9 knockdown might influence the cargo transport through the microtubules in previous research (Supplementary Fig. 2), and in this research, SEPT9 knockdown decreased the expression of β-tubulin, microtubules organization, as well as the number and size of FAs. Hence, SEPT9_i1 might influence on the turnover of FAs partly through altered the microtubule delivery function of paxillin. Not only that, we observed that in contrast to SEPT9-knockdown cells, the FAs in SEPT9_i1 overexpression cells were larger and closer to the edge of the cells, which implies SEPT9_i1 might promote cell migration through facilitating actin and microtubule cytoskeleton reorganization, the stabilization of cytoskeleton, and advancing FA maturation.
FAK is ubiquitously expressed in many different cells and tissues, and functions as both a protein tyrosine kinase and scaffold, mediating and regulating specific signals initiated at the sites of integrin clusters of FAs48. Several types of signaling events can initiate FAK activation, which leads to a number of cellular processes, including attachment, migration, invasion, proliferation, and survival49. The well-documented example of FAK function involves clustering of the integrins and the subsequent co-clustering of proteins such as talin and paxillin with the cytoplasmic tail of integrins50,51. FAK contains multiple tyrosine and serine phosphorylation sites that can induce conformational changes52, including Y397, Y576/Y577, and Y925. The FAK–Src complex phosphorylates and/or recruits several downstream FA proteins, including paxillin, p130Cas, etc., initiating specific cellular signaling pathways and responses53. Deregulated FAK/Src signaling occurs in a variety of tumors8,50,54. FAK/Src-dependent phosphorylation of paxillin Y31/118 exposes a vinculin binding site55 and recruits other adhesion molecules. Increased adhesion forces then transmit stronger mechanical signals to promote focal adhesion complex turnover in the process of cell migration7,56,57,58. In this study, our novel findings suggest a relationship between SEPT9 and activation of the FAK/Src/paxillin pathway. FAK-Y397, -Y576/577, -Y925, and Src-Y416 were all found to be phosphorylated in SEPT9_i1 overexpression cells, which implied the activation of FAK-Src-paxillin signaling. However, the results from transcriptomic analysis showed that ITGB4 is one of DEGs between SEPT9-KD1 and SEPT9-Ov1 cell lines, which suggested that the activation of FAK-Src-paxillin signaling might partly result from the increased expression of ITGB4 as the previous researches reported59,60. Notably, previous studies have demonstrated that the maturation of FAs is force dependent55,61, and the phosphorylation of paxillin plays a central role in this process58. In line with this, we found that there was a reduction in the phosphopaxillin-to-paxillin ratio when SEPT9_i1 was overexpressed (Fig. 7c), which is consistent with evidence that the relative level of phosphopaxillin is negatively regulated by force61,62.
The ability of Rho GTPase family members to regulate cytoskeletal dynamics, cell adhesion, and cell migration points to a central role in cancer cell invasion and metastasis. Of this family, RhoA, Rac, and Cdc42 typically control the actin and microtubule cytoskeleton organization, cell motility and cell adhersion32. Constitutively activated (GTPase deficient) mutants of Rho and Rac were found to induce the assembly of contractile actin and myosin filaments (stress fibers) and actin-rich surface protrusions (lamellipodia). Cdc42 has also been shown to promote the formation of actin-rich, finger-like membrane extensions (filopodia). These specialized actin structures are believed to provide the driving force for cell migration25. In this research, the protein expressions of RhoA, Rac and Cdc42 were all reduced in SEPT9-Knockdown cells and restored when SEPT9_i1 was overexpressed, and so was CDC42 mRNA (Fig. 4). Corresponding changes in actin and microtubules cytoskeleton, microtubule dynamics and cell morphology were also seen. These GTPase proteins have been known to reorganize cytoskeletons and to regulate cell migration via activation of effector proteins such as ROCK63,64,65,66. As a major signaling node for the interfaces of cell–cell adhesion, RhoA signaling is triggered when mechanical forces are applied to a variety of cell adhesion molecules67, and it has been a main focal point in studies of cellular responses to mechanical forces68. Activated RhoA can stimulate actin polymerization69 and also stabilize F-actin by activating ROCK70. It is well established that the Rho/ROCK signaling pathway is associated with cancer invasion and metastasis, such as breast cancer71,72, ovarian cancer63, colon cancer73, and gastric cancer65.
Previous studies have demonstrated that there is crossover and interaction between RhoA/ROCK1 and FAK/Src/paxillin signaling in promoting cell migration, and they may share upstream or downstream effector molecules and signaling processes74,75,76. Here, we also demonstrated that RhoA/ROCK1 signaling may play an important role in activation of FAK/Src/paxillin signaling, which was similar to the molecular mechanism of the hypoxia-induced breast cancer cell migration76. Earlier reports had identified that SA-RhoGEF (septin-associated RhoGEF) and Rhotekin can binds with SEPT9 and altered endogenous septin filament structures. They are novel regulators organizing mammalian septin structures and provide a link between septins and Rho signaling77,78. These results remind us SEPT9_i1-induced activation of RhoA/ROCK1 might be through interaction with these Rho regulators.
In summary, our results provide in vitro and in vivo evidence for the contribution of SEPT9_i1 to the motility of breast cancer cells. These studies showed that SEPT9 overexpression increased cell migration and motility through reorganization of cytoskeleton. Activation of the RhoA/ROCK1 and FAK/Src/paxillin pathways promoted FA maturation, which was consistent with KEGG pathway analysis. These results provide a basis for further investigation into targeting SEPT9-related signaling in order to effectively decrease the metastatic potential of tumor cells in breast and other carcinomas.
References
Wood, S. L., Westbrook, J. A. & Brown, J. E. Omic-profiling in breast cancer metastasis to bone: implications for mechanisms, biomarkers and treatment. Cancer Treat. Rev. 40, 139–152 (2014).
Collins, C. & Nelson, W. J. Running with neighbors: coordinating cell migration and cell-cell adhesion. Curr. Opin. Cell Biol. 36, 62–70 (2015).
Ridley, A. J. Life at the leading edge. Cell 145, 1012–1022 (2011).
Webb, D. J., Parsons, J. T. & Horwitz, A. F. Adhesion assembly, disassembly and turnover in migrating cells–over and over and over again. Nat. Cell Biol. 4, E97–E100 (2002).
Lee, B. Y., Timpson, P., Horvath, L. G. & Daly, R. J. FAK signaling in human cancer as a target for therapeutics. Pharm. Ther. 146, 132–149 (2015).
Fife, C. M., McCarroll, J. A. & Kavallaris, M. Movers and shakers: cell cytoskeleton in cancer metastasis. Br. J. Pharm. 171, 5507–5523 (2014).
Webb, D. J. et al. FAK-Src signalling through paxillin, ERK and MLCK regulates adhesion disassembly. Nat. Cell Biol. 6, 154–161 (2004).
Weiner, T. M., Liu, E. T., Craven, R. J. & Cance, W. G. Expression of focal adhesion kinase gene and invasive cancer. Lancet 342, 1024–1025 (1993).
Hess, A. R. et al. Focal adhesion kinase promotes the aggressive melanoma phenotype. Cancer Res 65, 9851–9860 (2005).
Mostowy, S. & Cossart, P. Septins: the fourth component of the cytoskeleton. Nat. Rev. Mol. Cell Biol. 13, 183–194 (2012).
Nagata, K. et al. Filament formation of MSF-A, a mammalian septin, in human mammary epithelial cells depends on interactions with microtubules. J. Biol. Chem. 278, 18538–18543 (2003).
Gonzalez, M. E., Makarova, O., Peterson, E. A., Privette, L. M. & Petty, E. M. Up-regulation of SEPT9_v1 stabilizes c-Jun-N-terminal kinase and contributes to its pro-proliferative activity in mammary epithelial cells. Cell Signal 21, 477–487 (2009).
Estey, M. P., Di Ciano-Oliveira, C., Froese, C. D., Bejide, M. T. & Trimble, W. S. Distinct roles of septins in cytokinesis: SEPT9 mediates midbody abscission. J. Cell Biol. 191, 741–749 (2010).
Connolly, D. et al. Septin 9 isoform expression, localization and epigenetic changes during human and mouse breast cancer progression. Breast Cancer Res 13, R76 (2011).
Hall, P. A. & Russell, S. E. Mammalian septins: dynamic heteromers with roles in cellular morphogenesis and compartmentalization. J. Pathol. 226, 287–299 (2012).
McIlhatton, M. A. et al. Genomic organization, complex splicing pattern and expression of a human septin gene on chromosome 17q25.3. Oncogene 20, 5930–5939 (2001).
Stanbery, L. & Petty, E. M. Steps solidifying a role for SEPT9 in breast cancer suggest that greater strides are needed. Breast Cancer Res 14, 101 (2012).
Scott, M., McCluggage, W. G., Hillan, K. J., Hall, P. A. & Russell, S. E. Altered patterns of transcription of the septin gene, SEPT9, in ovarian tumorigenesis. Int J. Cancer 118, 1325–1329 (2006).
Stanbery, L. et al. High SEPT9_v1 expression is associated with poor clinical outcomes in head and neck squamous cell carcinoma. Transl. Oncol. 3, 239–245 (2010).
Gonzalez, M. E. et al. High SEPT9_v1 expression in human breast cancer cells is associated with oncogenic phenotypes. Cancer Res 67, 8554–8564 (2007).
Hellmann, F., Verdi, M., Schlemper, B. R. Jr. & Caponi, S. 50th anniversary of the Declaration of Helsinki: the double standard was introduced. Arch. Med Res 45, 600–601 (2014).
Mao, X., Cai, T., Olyarchuk, J. G. & Wei, L. Automated genome annotation and pathway identification using the KEGG Orthology (KO) as a controlled vocabulary. Bioinformatics 21, 3787–3793 (2005).
Hamel, M. et al. Identification of differentially expressed markers in human follicular cells associated with competent oocytes. Hum. Reprod. 23, 1118–1127 (2008).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25, 402–408 (2001).
Etienne-Manneville, S. & Hall, A. Rho GTPases in cell biology. Nature 420, 629–635 (2002).
Parsons, J. T., Martin, K. H., Slack, J. K., Taylor, J. M. & Weed, S. A. Focal adhesion kinase: a regulator of focal adhesion dynamics and cell movement. Oncogene 19, 5606–5613 (2000).
Wu, G. S. et al. Ganoderiol A-enriched extract suppresses migration and adhesion of MDA-MB-231 cells by inhibiting FAK-SRC-paxillin cascade pathway. PLoS One 8, e76620 (2013).
Narumiya, S., Tanji, M. & Ishizaki, T. Rho signaling, ROCK and mDia1, in transformation, metastasis and invasion. Cancer Metastasis Rev. 28, 65–76 (2009).
Connolly, D. et al. Septin 9 amplification and isoform-specific expression in peritumoral and tumor breast tissue. Biol. Chem. 395, 157–167 (2014).
Amir, S., Golan, M. & Mabjeesh, N. J. Targeted knockdown of SEPT9_v1 inhibits tumor growth and angiogenesis of human prostate cancer cells concomitant with disruption of hypoxia-inducible factor-1 pathway. Mol. Cancer Res 8, 643–652 (2010).
Fuchtbauer, A. et al. Septin9 is involved in septin filament formation and cellular stability. Biol. Chem. 392, 769–777 (2011).
Ridley, A. J. Rho GTPase signalling in cell migration. Curr. Opin. Cell Biol. 36, 103–112 (2015).
Dolat, L. et al. Septins promote stress fiber-mediated maturation of focal adhesions and renal epithelial motility. J. Cell Biol. 207, 225–235 (2014).
Smith, C. et al. Septin 9 exhibits polymorphic binding to F-actin and inhibits myosin and cofilin activity. J. Mol. Biol. 427, 3273–3284 (2015).
Spiliotis, E. T. Regulation of microtubule organization and functions by septin GTPases. Cytoskeleton (Hoboken) 67, 339–345 (2010).
Bowen, J. R., Hwang, D., Bai, X., Roy, D. & Spiliotis, E. T. Septin GTPases spatially guide microtubule organization and plus end dynamics in polarizing epithelia. J. Cell Biol. 194, 187–197 (2011).
Amir, S. & Mabjeesh, N. J. SEPT9_V1 protein expression is associated with human cancer cell resistance to microtubule disrupting agents. Cancer Biol. Ther. 6, 1926–1931 (2007).
Targa, B. et al. Septin filament coalignment with microtubules depends on SEPT9_i1 and tubulin polyglutamylation, and is an early feature of acquired cell resistance to paclitaxel. Cell Death Dis. 10, 54 (2019).
Bai, X. et al. Novel septin 9 repeat motifs altered in neuralgic amyotrophy bind and bundle microtubules. J. Cell Biol. 203, 895–905 (2013).
Kuo, J. C. Focal adhesions function as a mechanosensor. Prog. Mol. Biol. Transl. Sci. 126, 55–73 (2014).
Smith, M. A., Hoffman, L. M. & Beckerle, M. C. LIM proteins in actin cytoskeleton mechanoresponse. Trends Cell Biol. 24, 575–583 (2014).
Gardel, M. L., Schneider, I. C., Aratyn-Schaus, Y. & Waterman, C. M. Mechanical integration of actin and adhesion dynamics in cell migration. Annu Rev. Cell Dev. Biol. 26, 315–333 (2010).
Burridge, K. & Wittchen, E. S. The tension mounts: Stress fibers as force-generating mechanotransducers. J. Cell Biol. 200, 9–19 (2013).
Etienne-Manneville, S. Microtubules in cell migration. Annu. Rev. Cell Developmental Biol. 29, 471–499 (2013).
Stehbens, S. & Wittmann, T. Targeting and transport: how microtubules control focal adhesion dynamics. J. Cell Biol. 198, 481–489 (2012).
Hu, Y. L. et al. Roles of microfilaments and microtubules in paxillin dynamics. Biochem Biophys. Res Commun. 348, 1463–1471 (2006).
Golan, M. & Mabjeesh, N. J. SEPT9_i1 is required for the association between HIF-1alpha and importin-alpha to promote efficient nuclear translocation. Cell Cycle 12, 2297–2308 (2013).
Frame, M. C., Patel, H., Serrels, B., Lietha, D. & Eck, M. J. The FERM domain: organizing the structure and function of FAK. Nat. Rev. Mol. Cell Biol. 11, 802–814 (2010).
van Nimwegen, M. J. & van de Water, B. Focal adhesion kinase: a potential target in cancer therapy. Biochemical Pharmacol. 73, 597–609 (2007).
Mitra, S. K. & Schlaepfer, D. D. Integrin-regulated FAK-Src signaling in normal and cancer cells. Curr. Opin. Cell Biol. 18, 516–523 (2006).
Lawson, C. et al. FAK promotes recruitment of talin to nascent adhesions to control cell motility. J. Cell Biol. 196, 223–232 (2012).
Kleinschmidt, E. G. & Schlaepfer, D. D. Focal adhesion kinase signaling in unexpected places. Curr. Opin. Cell Biol. 45, 24–30 (2017).
Hauck, C. R. et al. Inhibition of focal adhesion kinase expression or activity disrupts epidermal growth factor-stimulated signaling promoting the migration of invasive human carcinoma cells. Cancer Res 61, 7079–7090 (2001).
Ji, H. F. et al. Overexpression of focal adhesion kinase correlates with increased lymph node metastasis and poor prognosis in non-small-cell lung cancer. J. Cancer Res Clin. Oncol. 139, 429–435 (2013).
Pasapera, A. M., Schneider, I. C., Rericha, E., Schlaepfer, D. D. & Waterman, C. M. Myosin II activity regulates vinculin recruitment to focal adhesions through FAK-mediated paxillin phosphorylation. J. Cell Biol. 188, 877–890 (2010).
Galbraith, C. G., Yamada, K. M. & Sheetz, M. P. The relationship between force and focal complex development. J. Cell Biol. 159, 695–705 (2002).
Wu, J. C. et al. Focal adhesion kinase-dependent focal adhesion recruitment of SH2 domains directs SRC into focal adhesions to regulate cell adhesion and migration. Sci. Rep. 5, 18476 (2015).
Zaidel-Bar, R., Milo, R., Kam, Z. & Geiger, B. A paxillin tyrosine phosphorylation switch regulates the assembly and form of cell-matrix adhesions. J. Cell Sci. 120, 137–148 (2007).
Gan, L. et al. Extracellular matrix protein 1 promotes cell metastasis and glucose metabolism by inducing integrin beta4/FAK/SOX2/HIF-1alpha signaling pathway in gastric cancer. Oncogene 37, 744–755 (2018).
Pak, J. H. et al. Clonorchis sinensis excretory-secretory products promote the migration and invasion of cholangiocarcinoma cells by activating the integrin beta4-FAK/Src signaling pathway. Mol. Biochem Parasitol. 214, 1–9 (2017).
Zaidel-Bar, R. Early molecular events in the assembly of matrix adhesions at the leading edge of migrating cells. J. Cell Sci. 116, 4605–4613 (2003).
Zaidel-Bar, R., Kam, Z. & Geiger, B. Polarized downregulation of the paxillin-p130CAS-Rac1 pathway induced by shear flow. J. Cell Sci. 118, 3997–4007 (2005).
Ghasemi, A., Hashemy, S. I., Aghaei, M. & Panjehpour, M. RhoA/ROCK pathway mediates leptin-induced uPA expression to promote cell invasion in ovarian cancer cells. Cell Signal 32, 104–114 (2017).
Wei, L., Surma, M., Shi, S., Lambert-Cheatham, N. & Shi, J. Novel insights into the roles of Rho kinase in cancer. Archivum Immunol. Therapiae Exp 64, 259–278 (2016).
Genda, T. et al. Cell motility mediated by rho and Rho-associated protein kinase plays a critical role in intrahepatic metastasis of human hepatocellular carcinoma. Hepatology 30, 1027–1036 (1999).
Itoh, K. et al. An essential part for Rho-associated kinase in the transcellular invasion of tumor cells. Nat. Med 5, 221–225 (1999).
Lessey, E. C., Guilluy, C. & Burridge, K. From mechanical force to RhoA activation. Biochemistry 51, 7420–7432 (2012).
Chen, C. S. Mechanotransduction - a field pulling together? J. Cell Sci. 121, 3285–3292 (2008).
Watanabe, N. et al. p140mDia, a mammalian homolog of Drosophila diaphanous, is a target protein for Rho small GTPase and is a ligand for profilin. EMBO J. 16, 3044–3056 (1997).
Maekawa, M. et al. Signaling from Rho to the actin cytoskeleton through protein kinases ROCK and LIM-kinase. Science 285, 895–898 (1999).
Chan, C. H. et al. Deciphering the transcriptional complex critical for RhoA gene expression and cancer metastasis. Nat. Cell Biol. 12, 457–467 (2010).
Liu, S., Goldstein, R. H., Scepansky, E. M. & Rosenblatt, M. Inhibition of rho-associated kinase signaling prevents breast cancer metastasis to human bone. Cancer Res 69, 8742–8751 (2009).
Leve, F. et al. Lysophosphatidic acid induces a migratory phenotype through a crosstalk between RhoA-Rock and Src-FAK signalling in colon cancer cells. Eur. J. Pharm. 671, 7–17 (2011).
Lim, Y. et al. PyK2 and FAK connections to p190Rho guanine nucleotide exchange factor regulate RhoA activity, focal adhesion formation, and cell motility. J. Cell Biol. 180, 187–203 (2008).
Iwanicki, M. P. et al. FAK, PDZ-RhoGEF and ROCKII cooperate to regulate adhesion movement and trailing-edge retraction in fibroblasts. J. Cell Sci. 121, 895–905 (2008).
Gilkes, D. M. et al. Hypoxia-inducible factors mediate coordinated RhoA-ROCK1 expression and signaling in breast cancer cells. Proc. Natl Acad. Sci. USA 111, E384–E393 (2014).
Ito, H. et al. Possible role of Rho/Rhotekin signaling in mammalian septin organization. Oncogene 24, 7064–7072 (2005).
Nagata, K. & Inagaki, M. Cytoskeletal modification of Rho guanine nucleotide exchange factor activity: identification of a Rho guanine nucleotide exchange factor as a binding partner for Sept9b, a mammalian septin. Oncogene 24, 65–76 (2005).
Acknowledgements
This work was supported by the Natural Science Foundation of Education Office of Sichuan province, China (grant numbers 12ZB056).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by A. Oberst
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zeng, Y., Cao, Y., Liu, L. et al. SEPT9_i1 regulates human breast cancer cell motility through cytoskeletal and RhoA/FAK signaling pathway regulation. Cell Death Dis 10, 720 (2019). https://doi.org/10.1038/s41419-019-1947-9
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-019-1947-9
This article is cited by
-
METTL16 regulates the mRNA stability of FBXO5 via m6A modification to facilitate the malignant behavior of breast cancer
Cancer & Metabolism (2024)
-
Tuftelin1 drives experimental pulmonary fibrosis progression by facilitating stress fiber assembly
Respiratory Research (2023)
-
FRMD3 inhibits the growth and metastasis of breast cancer through the ubiquitination-mediated degradation of vimentin and subsequent impairment of focal adhesion
Cell Death & Disease (2023)
-
Septin11 promotes hepatocellular carcinoma cell motility by activating RhoA to regulate cytoskeleton and cell adhesion
Cell Death & Disease (2023)
-
SEPT9_v2, frequently silenced by promoter hypermethylation, exerts anti-tumor functions through inactivation of Wnt/β-catenin signaling pathway via miR92b-3p/FZD10 in nasopharyngeal carcinoma cells
Clinical Epigenetics (2020)