Abstract
Targeting autophagy may serve as a promising strategy for cancer therapy. Ganoderma lucidum polysaccharide (GLP) has been shown to exert promising anti-cancer effects. However, the underlying mechanisms remain elusive. Whether GLP regulates autophagy in cancer has never been reported. In this study, GLP induced the initiation of autophagy in colorectal cancer (CRC) HT-29 and HCT116 cells, as evidenced by enhanced level of LC3-II protein, GFP-LC3 puncta, and increased formation of double membrane vacuoles. However, GLP treatment caused marked increase of p62 expression. Addition of late stage autophagy inhibitor, chloroquine (CQ), further enhanced LC3-II and p62 level, as well as increased autophagosome accumulation, suggesting a blockage of autophagic flux by GLP in CRC cells. We then found GLP blocked autophagosome and lysosome fusion as determined by mRFP-GFP-LC3 colocalization analysis. Mechanistic study revealed that GLP-induced disruption of autophagosome-lysosome fusion is due to reduced lysosome acidification and lysosomal cathepsin activities. Cell viability and flow cytometry assays revealed that GLP-induced autophagosome accumulation is responsible for GLP-induced apoptosis in CRC cells. In line with this, inhibition of autophagy initiation by 3-methyladenine (3-MA), an early stage autophagy inhibitor, attenuated GLP-induced apoptosis. In contrast, suppression of autophagy at late stage by CQ enhanced the anti-cancer effect of GLP. Furthermore, we demonstrated that GLP-induced autophagosome accumulation and apoptosis is mediated via MAPK/ERK activation. Finally, GLP inhibited tumor growth and also inhibited autophagic flux in vivo. These results unveil new molecular mechanism underlying anti-cancer effects of GLP, suggesting that GLP is a potent autophagy inhibitor and might be useful in anticancer therapy.
Similar content being viewed by others
Introduction
Colorectal cancer (CRC) is the third most common malignancy and the third leading cause of cancer-related death in the USA1. Despite the benefits of early screening, surgery, and other therapeutic interventions, the current five-year survival rate for late stage CRC patients is still low2. Therefore, the search for effective and safe agents for CRC prevention and treatment is still urgently needed.
Natural products with highly diverse bioactivities and functions play a vital role in drug discovery for treating diseases. Ganoderma lucidum (G. lucidum) is a medicinal mushroom that has been used in East Asian countries for over 2000 years3,4,5. A substantial number of preclinical and clinical studies have reported that G. lucidum has numerous pharmacological effects, including antioxidant, hypoglycemic, immune-regulatory, anti-diabetic, and anti-cancerous5,6,7,8,9,10. Many studies have demonstrated that GLP is one of the main bioactive components responsible for anti-cancer effects of G. lucidum3,5. Recently, we showed that GLP extracted from the sporoderm-broken spores of G. lucidum significantly inhibited cell proliferation and induced apoptosis in colorectal and prostate cancer cells11,12. However, the molecular mechanisms underlying the anti-cancer effects of GLP remain unclear.
Autophagy is an evolutionarily conserved catabolic process that degrades cytoplasmic materials and provides substrates for energy metabolism during nutrient deprivation and metabolic stress13. Autophagy has been closely related to many human diseases, including obesity, aging, neurodegenerative disorders, and cancer13. The role of autophagy in cancer is complex and differs among various types of cancer14,15. Autophagy inhibits tumor initiation and progression in some cancers, but promotes tumor survival and progression in others14,15. Given these dual effects, therapeutic modulation of autophagy may serve as promising but challenging means for cancer treatment. Autophagy is considered a second type of programmed cell death (PCD)16. Intriguingly, it has been proposed that the interplay between autophagy and apoptosis, the type I PCD, may contribute to the anti-cancer effects of many anti-cancer agents17,18. However, what molecules or signaling pathways mediate the crosstalk between autophagy and apoptosis, whether these two PCDs regulate each other, and how anti-cancer agents affect these processes remain elusive.
In this study, we sought to examine the effect of GLP on autophagy and to evaluate whether such effect is relevant to the apoptotic effect induced by GLP in CRC, which has never been reported before. We found that GLP served as an autophagy initiation inducer and also a novel autophagic flux inhibitor by interfering with autophagosome-lysosome fusion. In addition, GLP-induced autophagosome accumulation is required for GLP-induced apoptosis in CRC cells. Furthermore, we demonstrated that GLP-induced autophagosome accumulation and apoptosis is mediated by MAPK/ERK activation.
Results
GLP inhibits cell viability and induces autophagy initiation in CRC cells
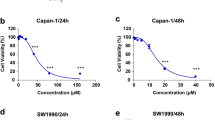
We first examined the effect of GLP on cell viability in HT-29 and HCT116 cells by MTT assay. As shown in Fig. 1a, GLP significantly reduced cell viability in both cells. In order to examine the effect of GLP on autophagy, we evaluated the distribution pattern of GFP-LC3 in CRC cells transiently expressing GFP-LC3, reminiscent of autophagosome formation19. During autophagy, the cytoplasmic form LC3-I is modified to LC3-II, thus, the amount of LC3-II increases with the formation of autophagosomes19. As shown in Fig. 1b, GLP-treated cells exhibited a dramatic increase in the punctuate distribution of GFP-LC3 in CRC cells, whereas autophagy inducer rapamycin (Rap) treated cells displayed less distribution of puncta. Quantitative analysis further confirmed this observation (Fig. 1b). We next confirmed the induction of autophagy initiation by GLP using transmission electron microscopy (TEM) in HT-29 cells. After treating cells with GLP for 24 h, numerous double-membrane autophagic vacuoles were observed in HT-29 cells, but much less in untreated cells (Fig. 1c).
a HT-29 and HCT116 cells were treated with indicated concentrations of GLP for 24, 48, and 72 h. Cell viability was measured by the MTT assay. b HT-29 and HCT116 cells were transfected with GFP-LC3 adenovirus for 24 h, and treated with GLP (5 mg/ml) and Rap (2 μM) for another 24 h. GFP-LC3 puncta was visualized by confocal microscope. The number of GFP-LC3 puncta per cell was quantified and presented as mean ± SE from 100 randomly selected cells (n = 3). Scale bar: 10 µm. c Transmission electron microscopy (TEM) was utilized to observe the formation of autophagosome upon GLP treatment (24 h). Arrows indicate representative double membrane vacuoles. Scale bar: 2 µm. d qRT-PCR determination of the mRNA expression of LC3 and Beclin-1 in HT-29 and HCT116 cells upon treatment with GLP at indicated concentrations for 24 h. e Western blotting analysis of LC3 and Beclin-1 expression in HT-29 and HCT116 cells upon GLP treatment (24 h). Data are presented as the mean ± SE from three independent experiments. *P < 0.05; **P < 0.01; ***P < 0.001 compared with untreated cells. N.S. No significance
Furthermore, GLP induced LC3-II expression at both mRNA and protein levels in a dose-dependent manner in CRC cells (Fig. 1d, e). However, the expression of Beclin-1, a key regulator of autophagy20, was down-regulated by GLP at both the protein and mRNA levels (Fig. 1d, e), suggesting that GLP-induced autophagy is Beclin-1-independnt in CRC cells. Supplementary Fig. 1 shows densitometric analysis of western blots.
GLP inhibits autophagic flux in CRC cells
The assessment of autophagy activity is not only dependent on the measurement of autophagy initiation, but also the degradation of autophagy substrates21. The accumulation of sequestosome-1 or p62, an autophagic cargo protein, inversely correlates with autophagic activity22. Our results showed that p62 expression was significantly increased upon GLP treatment in CRC cells at translational but not transcriptional levels (Fig. 2a, b), suggesting impaired autophagy flux occurred.
a HT-29 and HCT116 cells were treated with GLP at indicated concentrations for 24 h, and the expression of p62 was determined by western blotting analysis. b qRT-PCR analysis of the mRNA expression level of p62 following treatment of HT-29 and HCT116 cells with GLP (5 mg/ml) for 24 h. c–e HT-29 and HCT116 cells were treated with GLP (5 mg/ml), with or without CQ (10 μM) (c), or with or without Rap (2 μM) (d), or with or without 3-MA (5 mM) (e) for 24 h. The protein expressions of LC3 and p62 were determined by western blotting. f, g HT-29 cells were treated with GLP (5 mg/ml), with or without CQ (10 μM) (f), or with or without 3-MA (5 mg/ml) (g) for 24 h. TEM was utilized to observe the formation of autophagosome upon treatments. Arrows indicate representative double membrane vacuoles. Scale bar: 2 µm. Data are presented as Mean ± SE from three independent experiments. N.S. No significance
The effect of GLP on autophagy flux was evaluated by treating cells with late stage autophagy flux inhibitor CQ. As seen in Fig. 2c, both CQ (10 µM) and GLP (5 mg/ml) alone, significantly increased the level of LC3-II and p62 in CRC cells. Moreover, GLP-induced accumulation of LC3-II and p62 was further enhanced by CQ treatment (Fig. 2c), suggesting a synergistic effect on autophagy flux inhibition. In contrast, the autophagy initiation inducer, Rap, upregulated the level of LC3-II but decreased p62 expression in CRC cells, indicating normal autophagy proceeded (Fig. 2d). However, Rap further enhanced GLP-induced LC3-II and p62 expression in both cells, suggesting a synergistic effect with GLP on inducing autophagic autophagosome accumulation (Fig. 2d). We also examined the effect of autophagy early stage inhibitor 3-MA on GLP-induced LC3-II and p62 accumulation in CRC cells. As expected, 3-MA alone slightly inhibited LC3-II, increased p62 expression and further reduced GLP-induced LC3-II expression, but increased p62 expression significantly in HT-29 cells (Fig. 2e). Despite 3-MA significantly reduced GLP-induced LC3-II upregulation and further increased GLP-induced p62 expression in HCT116 cells, it also slightly increased LC3-II expression and reduced p62 expression when treated alone (Fig. 2e). These data suggest that 3-MA may play a complex role in regulating autophagy in these two CRC cells. Supplementary Fig. 2 shows densitometric analysis of western blots.
TEM analysis further confirmed a dramatic increase in the number of autophagic vacuoles upon GLP or CQ treatment in HT-29 cells, and a synergistic effect upon co-treatment with both agents (Fig. 2f). In contrast, 3-MA which inhibited the initiation of autophagy, formed very few autophagic vacuoles than GLP-treated cells (Fig. 2g). Consistent with results from western blotting, 3-MA dramatically reduced GLP-induced vacuole formation in HT-29 cells (Fig. 2g). Taken together, these results suggest that GLP-induced autophagosome accumulation is due to both the initiation of autophagy and inhibition of autophagic flux.
GLP blocks autophagosome-lysosome fusion in CRC cells
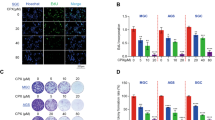
The fusion of autophagosome and lysosome is important for autophagic flux, and inhibition of this process impairs autophagic degradation22. We transfected CRC cells with mRFP-GFP-LC3 adenovirus that serves as a dual-fluorescence pH sensor for autophagic vacuoles and assessed autolysosome formation. As shown in Fig. 3a, treatment with Rap enhanced both green and red dot formation and colocalization gave rise to more red-only puncta, suggesting autolysosome maturation proceeded normally upon Rap treatment in HT-29 and HCT116 cells. Conversely, both the lysosomal acidification inhibitor CQ and GLP increased red and green puncta in CRC cells dramatically, while the yellow punctate fluorescence increased markedly after colocalization, indicating CQ and GLP blocked autophagosome-lysosome fusion in these cells (Fig. 3a, b). Notably, GLP yielded more yellow punctate fluorescence than CQ.
a, b HT-29 (a) and HCT116 (b) cells were transfected with mRFP-GFP-LC3 adenovirus and treated with Rap (2 μM), GLP (5 mg/ml), or CQ (10 μM) for 24 h, respectively. Cells were observed to distinguish between autophagosome (yellow puncta) and autolysosome (red puncta) after colocalization analysis using a confocal microscope. Scale bar: 10 µm. c Quantification of the mRNA expression level of LAMP-2 in HT-29 and HCT116 cells upon treatment with GLP at indicated concentrations for 24 h. d Western blotting analysis of LAMP-2 protein expression in CRC cells upon GLP treatment for 24 h. The relative intensity of LAMP-2 was calculated after normalization against β-Actin. Data are presented as the mean ± SE from three independent experiments. *P < 0.05; **P < 0.01; ***P < 0.001 compared with untreated cells
To confirm the above observation, we examined the expression of autophagosome-lysosome fusion marker, lysosome-associated membrane protein 2 (LAMP-2) in CRC cells. Indeed, we found that GLP significantly reduced the expression of LAMP-2 at both mRNA and protein levels in CRC cells (Fig. 3c, d). These results suggest that GLP-induced autophagic flux inhibition in CRC cells is due to impaired autophagosome-lysosome fusion.
GLP inhibits lysosomal acidity and cathepsin activity
Lysosomal enzymes play an important role in autophagic degradation, and the function of lysosomal enzymes are dependent on a narrow range of acidic pH23. We next assessed the effect of GLP on lysosome acidification by staining HT-29 cells with LysoSensor Green DND-189, an acidotropic fluorescent probe that can be trapped in acidic organelles, such as lysosomes. As shown in Fig. 4a, GLP significantly reduced the green fluorescence intensity in HT-29 cells, which is similar to CQ, the positive control. This result indicates that GLP inhibits lysosomal acidity in CRC cells.
a Inhibition of lysosome acidification upon treatment with GLP (5, 10 mg/ml) and CQ (10 μM) for 24 h in HT-29 cells as determined by LysoSensor Green DND-189 (1 mM) staining. Representative fluorescence microscopy images are shown. Scale bar: 20 µm. b Quantification of the mRNA expression levels of lysosomal proteases, CSTB and CTSD, in HT-29 and HCT116 cells upon GLP treatment for 24 h at indicated concentrations. c Western blotting analysis of pro-form and mature-form of CTSB and CTSD proteins in CRC cells upon GLP treatment for 24 h at indicated concentrations. Data are presented as the mean ± SE from three independent experiments. *P < 0.05; **P < 0.01; ***P < 0.001 compared with untreated cells. N.S. No significance
Autophagy is a process by which the cytoplasmic components are degraded by proteases in lysosomes24. We then investigated whether GLP treatment affects the expression of lysosomal cathepsin B and D (CSTB and CSTD). Both cathepsins are produced from precursor forms (pro-cathepsins) into the mature-cathepsins25,26. As shown in Fig. 4b, GLP significantly reduced the expression of CTSB and CTSD at mRNA level. In addition, the expression of the mature-CTSB and mature-CTSD at protein levels was also reduced in CRC cells (Fig. 4c). Taken together, the above results suggest that GLP reduces the acidity of lysosome and inhibits lysosomal cathepsin activities in CRC cells. Supplementary Fig. 3 shows densitometric analysis of western blots.
GLP induces autophagosome formation by activating MAPK/ERK signaling
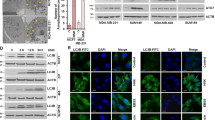
Autophagy is also regulated by many signaling pathways, including mTOR, AMPK, and MAPK27. We next examined the role of these signaling pathways in GLP-induced autophagic deregulation. As shown in Fig. 5a, GLP (5 mg/ml) transiently induced the phosphorylation of mTOR and MAPK/ERK, but reduced AMPKα phosphorylation within 2 h in CRC cells. In addition, GLP dose dependently induced mTOR and MAPK/ERK phosphorylation, but reduced AMPKα phosphorylation in CRC cells when treated for 1 h (Fig. 5b). We then extended time of treatment, and found GLP-induced MAPK/ERK phosphorylation remained high at 24 h (Fig. 5c). The result of p-AMPKα is confusing which did not show a clear trend upon prolonged treatment. Interestingly, the phosphorylation of mTOR was not further increased by GLP at 4 and 8 h in both cells, but started to decrease at 12 h in CRC cells and maintained to be inhibited at 24 h (Fig. 5c). Furthermore, at 24 h, GLP inhibited the phosphorylation of mTOR and AMPKα, but increased the phosphorylation of MAPK/ERK in a dose-dependent manner in both cells (Fig. 5d).
a HT-29 and HCT116 cells were treated with or without GLP (5 mg/ml) for 15, 30, 60, and 120 min. b HT-29 and HCT116 cells were treated with 5 mg/ml of GLP for 1 h at indicated concentrations. c HT-29 and HCT116 cells were treated with GLP (5 mg/ml) for 4, 8, 12, and 24 h. d HT-29 and HCT116 cells were treated with 5 mg/ml of GLP for 24 h at indicated concentrations. e HT-29 and HCT116 cells were pre-treated with PD98059 (20 µM) for 2 h, then co-treated with GLP (5 mg/ml) for additional 24 h. (For a–d, the expressions of both phosphorylated and total mTOR, AMPKα, and MAPK/ERK were determined by western blotting. The relative intensities of LC3-II and p-MAPK/ERK were calculated after normalization against β-Actin and t-MAPK/ERK, respectively. Data are presented as the Mean ± SE from three independent experiments. *P < 0.05; **P < 0.01; ***P < 0.001 compared with untreated cells
Previous studies have demonstrated that MAPK/ERK is an important regulator of autophagy28. In order to examine whether activated MAPK/ERK mediates GLP-induced autophagy in CRC cells, we pre-treated both cells with 20 µM of PD98059, a MAPK/ERK inhibitor, for 2 h and then treated with GLP for additional 24 h. Western blotting analysis revealed that the expression of GLP-induced LC3-II was significantly downregulated by PD98059 in CRC cells (Fig. 5e), suggesting that MAPK/ERK activation is responsible for GLP-induced autophagy initiation. Supplementary Fig. 4 shows densitometric analysis of western blots.
Accumulation of autophagosome confers cytotoxicity in CRC cells
We then examined whether autophagosome accumulation and MAPK/ERK activation play roles in GLP-induced cytotoxicity and apoptosis in CRC cells. As shown in Fig. 6a, b, addition of 3-MA or PD98059 significantly increased GLP-reduced cell viability in CRC cells. Furthermore, results from western blotting and flow cytometry showed that GLP alone significantly induced poly (ADP-ribose) (PARP) cleavage and apoptosis in both cells (Fig. 6c–e). However, both 3-MA and PD98059 significantly rescued GLP-induced PARP cleavage and apoptosis in CRC cells (Fig. 6c–e).
a, b HT-29 and HCT116 cells were treated with GLP (5 mg/ml) with or without 3-MA (5 mM) or with or without PD98059 (20 μM) for 24 h. The cell viability was then determined by MTT assay. c, d HT-29 and HCT116 cells were treated with GLP (5 mg/ml) with or without 3-MA (5 mM) (c), or with or without PD98059 (d) for 24 h. The expression of PARP was determined by western blotting. The relative intensities of cleaved-PARP in each cell line were calculated after normalization against β-Actin. e HT-29 and HCT116 cells were treated with GLP (5 mg/ml) with or without 3-MA (5 mM), or with or without PD98059 (20 μM) for 24 h. Apoptosis was analyzed by flow cytometry. Percentage of total apoptotic cells with both early and late apoptosis was calculated and presented. Data are presented as the mean ± SE from three independent experiments. *P < 0.05; **P < 0.01; ***P < 0.001. N.S. No significance
Furthermore, we analyzed the effect of increasing the accumulation of autophagosomes on cell viability and apoptosis upon GLP treatment in CRC cells. As shown in Fig. 7a, b, both the late stage autophagy inhibitor CQ and early stage autophagy inducer Rap, which increase autophagosome accumulation, significantly enhanced GLP-induced cell death in both cells. In addition, GLP-induced PARP cleavage and apoptosis were further increased by CQ or Rap (Fig. 7c–e). Taken together, these results demonstrate that GLP-induced autophagic flux disruption, and thus autophagosome accumulation contributes to GLP-induced cytotoxicity and apoptosis in CRC cells, which maybe mediated via MAPK/ERK activation in CRC cells.
a, b HT-29 and HCT116 cells were treated with GLP (5 mg/ml) with or without CQ (10 μM), or Rap (2 μM) for 24 h. The cell viability was then determined by MTT assay. c, d HT-29 and HCT116 cells were treated with GLP (5 mg/ml) with or without CQ (10 μM) (c), or with or without Rap (2 μM) (d) for 24 h. The expression of PARP was determined by western blotting. The relative intensities of cleaved-PARP in each cell line were calculated after normalization against β-Actin. e HT-29 and HCT116 cells were treated with GLP (5 mg/ml), with or without CQ (10 μM) or with or without Rap (2 μM) for 24 h. Apoptosis was analyzed by flow cytometry. Percentage of total apoptotic cells with both early and late apoptosis was calculated and presented. Data are presented as the mean ± SE from three independent experiments. *P < 0.05; **P < 0.01; ***P < 0.001. N.S. No significance
GLP inhibits tumor growth and inhibits autophagy in vivo
To determine the antitumor activity of GLP in vivo, male BALB/C nude mice were injected with HT-29 cells subcutaneously and then treated with GLP (150 mg/kg and 300 mg/kg). As shown in Fig. 8a–d, GLP treatment significantly decreased tumor volume and weight compared with control group, while the body weight of mice was not affected by GLP treatments. Next, we examined the regulation of autophagy by GLP in tumor samples. As shown in Fig. 8e, consistent with in vitro experiments, GLP increased the levels of LC3-II, p62, p-MAPK/ERK, and decreased the levels of Beclin-1 and p-AMPKα in xenograft tumors. Moreover, the levels of mature-CTSB and mature-CTSD were decreased in tumor sample from high-dose treatment (Fig. 8e). However, the expression of p-mTOR was increased in GLP-treated tumor samples (Fig. 8e). We also examined the apoptosis index by terminal deoxynucleotidyl transferase dUTP nick end labeling (TUNEL) staining. Unfortunately, GLP did not significantly increase apoptosis in xenograft tumors (Supplementary Fig. 5), though western blotting analysis showed that PARP cleavage might be higher in GLP-treated tumor samples than controls (Fig. 8e). These results indicate that GLP inhibits tumor growth and autophagy flux in vivo, but the role of GLP-induced autophagy inhibition in the tumor growth in xenografts is unknown in vivo.
a Image of the tumor samples from each group at necropsy: control; 150 mg/kg and 300 mg/kg. b Average final tumor weight of each group. c Growth curve of tumor volume in nude mice. d Effect of GLP treatment on body weight of nude mice. e Expression of LC3, p62, Beclin-1, p-mTOR, p-AMPKα, p-MAPK/ERK, Mature-CTSB, and Mature-CTSD in xenograft tumor at protein levels was determined by western blotting. β-Actin used as internal control. Data are presented as the mean ± SE from 7 xenograft tumor samples of each group. *P < 0.05; **P < 0.01; ***P < 0.001 compared with control group
Discussion
The role of autophagy in cancer is complex and context dependent29. Overwhelming preclinical and clinical evidence suggests that suppression of autophagy effectively inhibit cancer development and often sensitizing cancer cells to chemotherapeutic agents30,31,32. CQ and its analog, hydroxychloroquine (HCQ), are the only autophagy inhibitor being used for treating cancer in clinical trials33. However, Phase I/II clinical trials with CQ and HCQ showed mixed results with varying levels of efficacy, highlighting the needs to develop alternative autophagy inhibitors33,34,35. Numerous studies have indicated that GLP inhibits tumorigenesis through inducing apoptosis, inhibiting invasion, and regulating immune response3,5,8. However, there has been no report regarding the effect of GLP on autophagy in cancer. As summarized in Supplementary Fig. 6, our study demonstrates that GLP induced autophagy initiation and suppressed autophagic degradation through inhibiting lysosomal acidification and downregulating lysosomal enzymes. The accumulation of autophagosomes that resulted from autophagic flux disruption contributes to GLP-induced apoptosis through MAPK/ERK activation in CRC cells.
Although defined as type II PCD, controversy exists over whether autophagy alone can drive cell death and the molecular mechanisms underlying autophagic cell death are poorly understood16,36. Autophagy and apoptosis could cross-regulate each other and often occur in a sequence in which autophagy precedes apoptosis17. In line with our results, bodies of evidence suggest that disruption of the late stage of autophagy leads to excessive accumulation of autophagosomes, which has the potential to turn autophagy into a destruction process and induce apoptosis37,38,39,40,41. This notion is further supported by results obtained after adding Rap to CRC cells in our study. The autophagy inducer Rap induced normal process of autophagy and showed no effect on apoptosis in CRC cells. However, apoptosis was enhanced when cells were co-treated with GLP and Rap, suggesting that autophagosome accumulation plays a key role in GLP-induced cell death. In addition, blocking of autophagosome formation by autophagy in early stage inhibitor 3-MA dramatically rescued GLP-induced apoptosis. In contrast, the late stage autophagy inhibitor CQ which increases autophagosome formation, enhanced GLP-induced cytotoxicity and apoptosis. These results suggest that GLP-induced autophagosome accumulation is responsible for its pro-apoptotic effect in CRC cells. Similarly, a substantial number of studies have shown that other chemotherapy drugs induced the accumulation of autophagosomes due to autophagic flux disruption also confers cytotoxicity21,42,43,44,45,46.
Using both genetic and chemical approaches, Button et al. found defects in autophagosome-lysosome fusion or lysosome alone do not induce sufficient cellular toxicity. However, it was the increased autophagosome synthesis and accumulation that directly induce cell death in three different cancer cell models45. Together with our results, autophagosome accumulation through inhibiting autophagy may be a valuable strategy for cancer therapy by anti-cancer agent, such as GLP. However, in neurodegenerative or other diseases, lowering autophagosome accumulation may have a therapeutic value instead47.
Autophagy and apoptosis are under the control of multiple molecules and signaling pathways16. The classical regulation of autophagy is governed by the mTOR pathway, upon inhibition by AMPK27. However, apart from classical role of mTOR, numerous studies have reported that autophagy induction could also be mTOR-independent process48,49,50,51. Furthermore, some studies found that inhibition of mTOR would rather impair the autophagic flux, thus leads to enhanced autophagosome accumulation rather than autophagy induction52,53. These observations may support our findings that a later inhibition (after 8 h) of mTOR might be important in GLP-induced autophagy flux disruption. In our study, we also found that the regulation of autophagy by GLP may be AMPK-independent, as supported by studies from others54,55. Interestingly, Vucicevic et al. reported that Compound C, a specific AMPK inhibitor, actually inhibited autophagy through an AMPK-independent inhibition of mTOR in cancer cells56.
Beclin-1 is essential for the formation and extension of pre-autophagosomal structure20. However, numerous studies suggest that there are Beclin-1-independent mechanisms of autophagy and autophagy-independent roles for Beclin-1 in cancer57,58,59,60,61,62,63. Our study showed that the accumulation of autophagosomes induced by GLP is not accompanied by the upregulation of Beclin-1, but rather Beclin-1 is inhibited by GLP in CRC cells. Although suggested as a tumor suppressor gene, role of Belin-1 in cancer is very complicated and controversial64. Recent studies reported that deletion of Beclin-1 expression by RNAi promoted apoptosis in CRC cells65,66. Clinical studies have reported that high expression of Beclin-1 in human advanced colorectal cancer tumors is negativity correlated with the survival rate of patients67. These data suggest that inhibition of Beclin-1 by GLP may serve as autophagy-independent mechanism underlying the anti-cancer effects of GLP in CRC cells. However, more studies are needed to definitively determine this hypothesis by using RNAi method that completely deletes Beclin-1 in CRC cells in future.
MAPK has been shown to play an important role in autophagy regulation28. However, the precise role of MAPK/ERK in autophagy initiation or maturation has not been defined and yields controversial results28. Accumulating evidence indicates that impaired autophagy through disruption of lysosomal function can be triggered by drug-induced activation of MAPK/ERK and thus increases cell death68,69. Sustained activation of MAPK/ERK has been shown to inhibit the maturation step of the autophagy process68. Activation of MAPK/ERK signaling pathway has also been reported to impair autophagy by promoting degradation of Fork head Box O1 (FOXO1) in cancer cells70. FOXO1 has been shown to be a protein involved in the dynamic control of autophagy, which might be essential for maintenance of autophagic flux in cancer cells and neurons71,72,73,74. It will be interesting to examine whether effect of GLP on MPAK/ERK activation is through targeting FOXO1 in CRC cells in future. Our study suggests that GLP-induced initiation of autophagy and apoptosis is MAPK/ERK-dependent. However, whether MAPK/ERK plays a role in GLP-induced autophagic flux disruption also need to be further examined in future studies.
One discrepancy in our study is that although autophagy flux was inhibited both in vitro and in vivo, GLP failed to increase apoptosis index in CRC tumor samples. One explanation could be that cancer progression in vivo is largely influenced by the tumor microenvironment75. Previously, we reported that GLP induced significant increase of necrosis, another form of cell death, in HCT116 xenograft tumors11. Both hypoxia and immunity within the tumor play a role that determine how autophagy, apoptosis, and necrosis coordinate with each other75. It will be interesting in future to dissect the specific role of GLP-induced autophagy inhibition in cell death in vivo.
In summary, our study demonstrates that GLP triggers apoptosis in CRC by suppressing autophagic flux and inducing autophagosome accumulation. We provide insights into the molecular mechanisms by which GLP elicits the anti-cancer effects against CRC. Our findings provide a basis for potential use of GLP as an adjuvant or direct autophagy inhibitor for CRC treatment. Our results also suggest that increasing the accumulation of autophagosomes by anti-cancer agents may have therapeutic value for CRC.
Materials and methods
Materials
Cell culture and reagents
Human colon cancer cell lines HT-29 and HCT116 were obtained from American Type Culture Collection (ATCC, VA, USA). Cells were maintained in a humidified atmosphere of 5% CO2 at 37 ℃ in Dulbecco’s modified Eagle’s medium (DMEM, Gibco, Grand Island, NY, USA) supplemented with 10% fetal bovine serum (Gemini, Grand Island, NY, USA) and 1% penicillin-streptomycin (Gibco, USA). [3-(4, 5-dimethylthia-zol-2-yl)-2, 5-diphenyltetrazolium bromide] (MTT) and 3-MA were purchased from Sigma-Aldrich (Saint Louis, Missouri, USA). Rap was purchased from Yuanye Biotech (Shanghai, China). CQ was obtained from MedChemExpress Technology (New Jersey, UAS). PD98059 was purchased from Calbiochem (Darmstadt, Germany).
Preparation of GLP
The powder of sporodum-broken spores of G. lucidum was obtained from Shouxiangu Institute of Rare Medicine Plant (Wuyi, Zhejiang, China). GLP from the sporodum-broken spores of G. lucidum was extracted by hot water extraction method as described before11. Briefly, 5 g power of sporodum-broken spores of G. lucidum was placed in 100 ml of ultrapure water, lipid was first removed as described before76 and then oscillated (300 rpm) at 70 ℃ for 12 h. However, the proportion of residual lipid and protein contents were not determined at present. The solution was centrifuged at 4000 rpm for 15 min to remove insoluble materials, and further purified to remove protein using Sevage method as described before76, and centrifuged again to collect the supernatant. The supernatant was then concentrated and freeze-dried using H051 freeze dryer (LaboGene, Lynge, Denmark) to obtain GLP for subsequent experiments.
Cell viability assay
HT-29 and HCT116 cells were seeded in 96-well plates at a density of 1 × 104 cells in 200 μl of serum-containing DMEM medium until cells reach 40–50% confluence. To evaluate the cytotoxic effect of GLP on cells, HT-29 and HCT116 cells were treated with different concentrations (0, 2.5, 5, and 10 mg/ml) of GLP for 24, 48, and 72 h. For mechanistic studies, cells were either pre-treated or co-incubated with 3-MA (5 mM), PD98059 (20 μM), Rap (2 μM), or CQ (10 μM) for certain time in the presence or absence of GLP as indicated in figure legends. After incubation, MTT solution (5 mg/ml) was added to the DMEM and incubated for an additional 4 h. Then, supernatants were removed and the purple formazan crystals were dissolved in 150 μl DMSO and absorbance was measured at 490 nm using a microplate reader (Bio-Tek Instrument Inc., Winooski, VT, USA). The percentage of viability was expressed as (ODsample−ODblank)/(ODcontrol−ODblank) × 100%, where OD is the absorbance.
Quantification of GFP-LC3 puncta
HT-29 and HCT116 cells were seeded in CELLviewTM glass bottom dish (USA Scientific, Ocala, FL, USA) at a density of 1 × 103 cells in 1 ml of serum-containing DMEM until cells reach 30% confluence. The cells were transfected with adenovirus GFP-LC3 (Hanbio Biotech, Shanghai, China), and then treated with GLP or indicated chemicals for additional 24 h. Cells were observed under a laser scanning confocal microscope (LSM880, Carl Zeiss, Germany). The average number of GFP-LC3 puncta per cell was counted with at least 100 cells for each cell line.
Transmission electron microscope
HT-29 cells were seeded in 6-well plates at a density of 2 × 105 cells in 2 ml of serum-containing DMEM until cells reach 40–50% confluence, Cells were then treated with GLP (5 mg/ml), 3-MA (5 mM), CQ (10 μM), GLP/3-MA, or GLP/CQ for 24 h. The cells were washed with PBS and harvested by trypsinization, and then the cells were fixed with 2.5% glutaraldehyde for overnight at 4 ℃. Next day, glutaraldehyde was removed and washed three times in the PBS for 15 min at each step. Then, cells were first embedded in agarose gel and fixed with 1% OsO4 in PBS for 1 h, OsO4 was removed and washed three times in the PBS for 15 min at each step. The cells were then dehydrated through an ascending series of ethanol (30, 50, 70, 80, 90, 95%) for 15 min at each step, then dehydrated by 100% ethanol for 20 min. Next, the cells were transferred to absolute acetone for 20 min, and the cells were placed in 1:1 mixture of absolute acetone and Spurr resin for 1 h at room temperature, then transferred to 1:3 mixture of absolute acetone and the Spurr resin for 3 h and further transferred to Spurr resin mixture for overnight. Samples were placed in Eppendorf tube containing Spurr resin and heated at 70 ℃ for more than 9 h. The samples were sectioned with Leica EM UC7 ultramicrotome (Leica, Nussolch, Germany). After staining with uranyl acetate and alkaline lead citrate for 10 min, respectively, samples were observed with a Hitachi H-7650 transmission electron microscope (Hitachi, Tokyo, Japan). All images were acquired by a Gatan SC1000 (Model 832) CCD camera (Gatan Inc, Pleasanton, CA, USA).
mRFP-GFP-LC3 adenovirus translocation and colocalization analysis
HT-29 and HCT116 cells were seeded in CELLviewTM glass bottom dish (USA Scientific) at a density of 1 × 103 cells in 1 ml of serum-containing DMEM until cells reach 30% confluence. Cells were then transfected with the mRFP-GFP-LC3 adenovirus (multiplicity of infection = 300) for 6 h in DMEM supplemented with 10% FBS and 1% p/s. DMEM were removed and washed twice with PBS. Cell were incubated for 24 h in serum-containing DMEM. Cells were treated with GLP (5 mg/ml), Rap (2 μM), or CQ (10 μM) for 24 h. After incubation, image acquisition was done using a laser scanning confocal microscope (LSM880, Carl Zeiss, Germany). mRFP and GFP expressed in mRFP-GFP-LC3 tandem fluorescent protein adenovirus were used to track LC3. As GFP fluorescent protein is sensitive to acidity, the GFP fluorescence was quenched after the fusion of autophagosome and lysosome. Therefore, the weaken of GFP could indicate the fusion of autophagosome and lysosome to form autolysosome. When red and green fluorescence merged, the yellow spots represent autophagosomes and red spots represent autolysosomes. When autophagy occurs, there will be more red spots than yellow spots, and when the fusion of autophagosomes-lysosomes is damaged, the yellow spots are dominant22,23.
Measurement of intralysosomal acidity
The LysoSensorTM Green DND-189 reagent (Yeasen Biotech, Shanghai, China) exhibits an acid-dependent increase in fluorescence intensity. Cell staining was performed according to the manufacturer’s protocol. Briefly, cells were treated with GLP (5 and 10 mg/ml) or CQ (10 μM) for 24 h. The cells were then washed twice with PBS and added a final concentration of 2 uM LysoSensor reagent, and further incubated for 20 min at room temperature (RT) in the dark. The fluorescence intensity was observed under an Ti-S inverted fluorescence microscope (Nikon, Tokyo, Japan).
RNA isolation and quantitative real-time PCR
Total RNA isolation was performed using EASYspin reagent Kit (Aidlab Biotech, Beijing, China), following the manufacturer’s protocol. Both the quality and quantity of total RNA were analyzed by the NanoDrop 2000c spectrophotometer (Thermo Scientific, Grand Island, NY, USA). First-strand cDNA was synthesized from 1 µg of total RNA using iScript Reverse transcription supermix (Bio-Rad, Hercules, CA, USA). qRT-PCR analysis was performed using SYBR Green master mix (Bio-Rad) on CFX96 Real-Time PCR system (Bio-Rad). Primers were designed with open-sourced software Primer3Plus (Cambridge, MA, USA). Beta-Actin was used as the reference gene. The relative expression of mRNA was calculated as follows: ΔCt = Ct (sample)−Ct (β-Actin), ΔΔCt (sample) = ΔCt (sample)−ΔCt (calibrator). The fold change of mRNA was calculated according to the relative quantification (2−ΔΔCt). Primer sequences of qRT-PCR reactions are shown in Supplementary Table 1.
Western blotting
Polyclonal LC3, Beclin-1, p62, MAPK/ERK, p-MAPK/ERK, mTOR, p-mTOR (Ser2448), AMPKα, p-AMPKα (Ser485), PARP, β-Actin, and monoclonal CTSB antibodies were purchased from Cell Signaling Technology (Danvers, MA, USA). Monoclonal CTSD and LAMP-2 antibodies were purchased from Abcam (Cambridge, England). Cells were lysed in ice-old RIPA buffer (PBS, 1% Igepal, 10% SDS, 0.5% Sodium Deoxycholate) supplemented with complete protease inhibitors and total protein concentrations were measured by bicinchoninic acid (BCA) protein assay kit (Pierce, Rockford, IL, UAS). An equal amount of total protein (40 μg) was resolved by SDS–PAGE and transferred to PVDF (polyvinylidene difluoride) membrane (MilliporeSigma, USA). After blocking with 5% nonfat milk in 1× TBST (Tris-buffered saline, 50 mM of Tris, 150 mM of NaCl, pH 7.5, 0.1% Tween-20) at RT for 1 h, the membranes were probed with the following antibodies: LC3, Beclin-1, p62, p-mTOR (Ser2448), t-mTOR, p-ERK, t-ERK, p-AMPKα (Ser485), t-AMPKα, CTSB, CTSD, PARP, LAMP-2, and β-Actin at indicated concentrations as listed in Supplementary Table 2. Membranes were incubated with primary antibody for overnight at 4 ℃. After washing with TBST, the membranes were incubated with secondary anti-rabbit antibody (1:2000) at RT for 1 h. Then signals were captured using western lightning Plus ECL enhanced chemiluminescence substrate from PerkinElmer (Waltham, MA, USA), and detected using Minichemi TM610 chemical imaging System (Beijing, China). The quantitative of optical density was analyzed using ImageJ 1.41 software (Bethesda, MD, USA).
Flow cytometric analysis of apoptosis
Flow cytometric analysis was used for apoptosis assay. Cells (2 × 105) were seeded in 6-well plate per well and treated with GLP (5 mg/ml), 3-MA (5 mM), CQ (10 μM), GLP/3-MA or GLP/CQ for 24 h. When analyzing the effect of PD98059 and Rap on GLP-induced apoptosis, cells were starved overnight, then pre-treated with PD98059 (20 μM) and Rap (2 μM) for 2 h, respectively, and then treated for 24 h in the absence or presence of GLP (5 mg/ml). The cells were washed with PBS, harvested by trypsinization and washed twice with PBS. The cells were then stained with propidium iodide (PI) and annexin V-FITC using FITC Annexin V Apoptosis Detection Kit (BD Pharmingen, San Diego, CA, USA) according to manufacturer’s instruction. Briefly, cells were re-suspended with 1× binding buffer at a density of 1 × 106 cells/ml. Then added PI and Annexin V-FITC (5 μl) to cell suspension (100 μl) before further incubation for 15 min at RT in the dark. Stained cells were diluted with 1× binding buffer and analyzed by Guava EasyCyte flow cytometer (Merck KGaA, Darmstadt, Germany). The percentage of Annexin V+/PI− (early apoptosis), Annexin V+/PI+ (late apoptosis), and Annexin V−/PI+ (necrosis) cells was analyzed on the basis of manufacture’s instruction (BD Pharmingen). Data was represented as rate of total apoptotic cells with both early and late apoptotic rate indicated.
Xenograft mouse model
All the experimental procedures were performed according to the Guide for the Care and Use of Laboratory Animals of the National Institutes of Health. All procedures of the experiment are in line with the Animal Welfare Act Regulations. This study was approved by the Committee on the Ethics of Animal Experiments of Zhejiang Chinese medical University (Permit Number: SYXK 2012-0002). Male BALB/C nude mice (four-week-old) were kept in specific pathogen free (SPF) environment at the experimental animal center at the Zhejiang Chinese Medical university. After 2 week of adaptation, the mice were randomly divided by weight into treatment and control groups (n = 10 per group): control group (saline), low dose group (150 mg/kg), and high dose group (300 mg/kg). HT-29 cells (5 × 106 cells in 200 ul PBS) were implanted into the right flank of nude mouse.
The day after implantation of cancer cells, the mice were treated with GLP by gavage per day in low and high dose groups, while the control group mice were treated with saline per day. Body weights and tumor volume were measured twice a week and distinct tumors were measured in length and width using a digital vernier caliper (0.01 mm). Tumor volume was assessed according to the formula V (mm3) = length × width2/2. Six weeks after injection, all mice were sacrificed. At the end of study, 1–3 mice of each group did not grow tumor: control (n = 1); 150 mg/kg of GLP (n = 2); 300 mg/kg of GLP (n = 3). In addition, there were 2 mice in control group and 1 mouse in low dose group developed large tumor (>1.8 cm) very early during the study, and thus, these mice were sacrificed early and removed from study. Therefore, data from 7 mice of each group were analyzed. After necropsy, xenograft tumors were carefully excised and weighed. The tumor tissues were snap frozen in liquid nitrogen and stored at −80 ℃ for subsequent analysis.
Statistical analysis
All results presented are representatives from at least three independent experiments. Data are expressed as mean ± standard error (SE) of indicated number of experiments. For comparisons between two groups, a Student t-test was used. And for analysis with multiple comparisons, one-way analysis of variance (ANOVA) with Dunnett’s correction or Bonferroni’s correction for pairwise comparison was used. A value of *P < 0.05 was considered as statistically significant. All analyses were performed using Graphpad Prism 5.0 (GraphPad Software, Inc., La Jolla, CA, USA).
References
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer statistics, 2019. CA Cancer J. Clin. 69, 7–34 (2019).
Brenner, H., Kloor, M. & Pox, C. P. Colorectal cancer. Lancet 383, 1490–1502 (2014).
Batra, P., Sharma, A. K. & Khajuria, R. Probing Lingzhi or Reishi medicinal mushroom Ganoderma lucidum (higher Basidiomycetes): a bitter mushroom with amazing health benefits. Int. J. Med. Mushrooms 15, 127–143 (2013).
Loyd, A. L. et al. Identifying the “Mushroom of Immortality”: assessing the ganoderma species composition in commercial reishi products. Front. Microbiol. 9, 1557 (2018).
Bishop, K. S. et al. From 2000 years of Ganoderma lucidum to recent developments in nutraceuticals. Phytochemistry 114, 56–65 (2015).
Cheng, S. & Sliva, D. Ganoderma lucidum for cancer treatment: we are close but still not there. Integr. Cancer Ther. 14, 249–257 (2015).
Jiang, Y. et al. Overview of Ganoderma sinense polysaccharide-an adjunctive drug used during concurrent Chemo/Radiation therapy for cancer treatment in China. Biomed. Pharmacother. 96, 865–870 (2017).
Kladar, N. V., Gavaric, N. S. & Bozin, B. N. Ganoderma: insights into anticancer effects. Eur. J. Cancer Prev. 25, 462–471 (2016).
Ma, H. T., Hsieh, J. F. & Chen, S. T. Anti-diabetic effects of Ganoderma lucidum. Phytochemistry 114, 109–113 (2015).
Zeng, P. et al. Chemical, biochemical, preclinical and clinical studies of Ganoderma lucidum polysaccharide as an approved drug for treating myopathy and other diseases in China. J. Cell Mol. Med. 22, 3278–3297 (2018).
Na, K. et al. Anticarcinogenic effects of water extract of sporoderm-broken spores of Ganoderma lucidum on colorectal cancer in vitro and in vivo. Int. J. Oncol. 50, 1541–1554 (2017).
Wu, K. et al. Effects of non-steroidal anti-inflammatory drug-activated gene-1 on Ganoderma lucidum polysaccharides-induced apoptosis of human prostate cancer PC-3 cells. Int. J. Oncol. 53, 2356–2368 (2018).
Levine, B. & Kroemer, G. Autophagy in the pathogenesis of disease. Cell 132, 27–42 (2008).
Wilde, L., Tanson, K., Curry, J. & Martinez-Outschoorn, U. Autophagy in cancer: a complex relationship. Biochem. J. 475, 1939–1954 (2018).
Guo, J. Y., Xia, B. & White, E. Autophagy-mediated tumor promotion. Cell 155, 1216–1219 (2013).
Fuchs, Y. & Steller, H. Live to die another way: modes of programmed cell death and the signals emanating from dying cells. Nat. Rev. Mol. Cell Biol. 16, 329–344 (2015).
Marino, G., Niso-Santano, M., Baehrecke, E. H. & Kroemer, G. Self-consumption: the interplay of autophagy and apoptosis. Nat. Rev. Mol. Cell Biol. 15, 81–94 (2014).
Cooper, K. F. Till death do us part: the marriage of autophagy and apoptosis. Oxid. Med. Cell Longev. 2018, 4701275 (2018).
Klionsky, D. J. et al. Guidelines for the use and interpretation of assays for monitoring autophagy (3rd edition). Autophagy 12, 1–222 (2016).
Levine, B. & Klionsky, D. J. Development by self-digestion: molecular mechanisms and biological functions of autophagy. Dev. Cell 6, 463–477 (2004).
Lao, Y. et al. The natural compound oblongifolin C inhibits autophagic flux and enhances antitumor efficacy of nutrient deprivation. Autophagy 10, 736–749 (2014).
Mizushima, N., Yoshimori, T. & Levine, B. Methods in mammalian autophagy research. Cell 140, 313–326 (2010).
Yoshii, S. R. & Mizushima, N. Monitoring and measuring autophagy. Int J Mol Sci 18, E1865 (2017).
Dikic, I. Proteasomal and autophagic degradation systems. Annu Rev Biochem 86, 193–224 (2017).
Fong, D., Calhoun, D. H., Hsieh, W. T., Lee, B. & Wells, R. D. Isolation of a cDNA clone for the human lysosomal proteinase cathepsin B. Proc Natl Acad Sci USA 83, 2909–2913 (1986).
Richo, G. R. & Conner, G. E. Structural requirements of procathepsin D activation and maturation. J. Biol. Chem. 269, 14806–14812 (1994).
Kim, J., Kundu, M., Viollet, B. & Guan, K. L. AMPK and mTOR regulate autophagy through direct phosphorylation of Ulk1. Nat. Cell Biol. 13, 132–141 (2011).
Sridharan, S., Jain, K. & Basu, A. Regulation of autophagy by kinases. Cancers 3, 2630–2654 (2011).
Kimmelman, A. C. The dynamic nature of autophagy in cancer. Genes Dev. 25, 1999–2010 (2011).
White, E., Karp, C., Strohecker, A. M., Guo, Y. & Mathew, R. Role of autophagy in suppression of inflammation and cancer. Curr. Opin. Cell Biol. 22, 212–217 (2010).
Livesey, K. M., Tang, D., Zeh, H. J. & Lotze, M. T. Autophagy inhibition in combination cancer treatment. Curr. Opin. Investig. Drugs 10, 1269–1279 (2009).
Zhou, J. et al. Andrographolide sensitizes cisplatin-induced apoptosis via suppression of autophagosome-lysosome fusion in human cancer cells. Autophagy 8, 338–349 (2012).
Carew, J. S., Kelly, K. R. & Nawrocki, S. T. Autophagy as a target for cancer therapy: new developments. Cancer Manag. Res. 4, 357–365 (2012).
Limpert, A. S. et al. Autophagy in cancer: regulation by small molecules. Trends Pharmacol. Sci. 39, 1021–1032 (2018).
Levy, J. M. M., Towers, C. G. & Thorburn, A. Targeting autophagy in cancer. Nat. Rev. Cancer 17, 528–542 (2017).
Doherty, J. & Baehrecke, E. H. Life, death and autophagy. Nat. Cell Biol. 20, 1110–1117 (2018).
Shen, J. et al. Autophagy inhibition induces enhanced proapoptotic effects of ZD6474 in glioblastoma. Br. J. Cancer 109, 164–171 (2013).
Masud Alam, M. et al. Inhibition of autophagy by chloroquine induces apoptosis in primary effusion lymphoma in vitro and in vivo through induction of endoplasmic reticulum stress. Apoptosis 21, 1191–1201 (2016).
Liu, L. et al. Lysosomal dysfunction and autophagy blockade contribute to IMB-6G-induced apoptosis in pancreatic cancer cells. Sci. Rep. 7, 41862 (2017).
Mittal, S. et al. Impaired lysosomal activity mediated autophagic flux disruption by graphite carbon nanofibers induce apoptosis in human lung epithelial cells through oxidative stress and energetic impairment. Part Fibre Toxicol. 14, 15 (2017).
Li, Z. et al. Exendin-4 impairs the autophagic flux to induce apoptosis in pancreatic acinar AR42J cells by down-regulating LAMP-2. Biochem. Biophys. Res. Commun. 496, 294–301 (2018).
Masuelli, L. et al. Chloroquine supplementation increases the cytotoxic effect of curcumin against Her2/neu overexpressing breast cancer cells in vitro and in vivo in nude mice while counteracts it in immune competent mice. Oncoimmunology 6, e1356151 (2017).
Bi, Y. et al. Inhibition of autophagy induced by quercetin at a late stage enhances cytotoxic effects on glioma cells. Tumour Biol. 37, 3549–3560 (2016).
Cheng, X. et al. Targeting autophagy enhances apatinib-induced apoptosis via endoplasmic reticulum stress for human colorectal cancer. Cancer Lett. 431, 105–114 (2018).
Button, R. W., Roberts, S. L., Willis, T. L., Hanemann, C. O. & Luo, S. Accumulation of autophagosomes confers cytotoxicity. J. Biol. Chem. 292, 13599–13614 (2017).
Mei, W. et al. Gambogenic acid kills lung cancer cells through aberrant autophagy. PLoS ONE 9, e83604 (2014).
Tammineni, P. & Cai, Q. Defective retrograde transport impairs autophagic clearance in Alzheimer disease neurons. Autophagy 13, 982–984 (2017).
Chen, X. et al. Trehalose, sucrose and raffinose are novel activators of autophagy in human keratinocytes through an mTOR-independent pathway. Sci. Rep. 6, 28423 (2016).
Hau, A. M. et al. Coibamide A induces mTOR-independent autophagy and cell death in human glioblastoma cells. PLoS ONE 8, e65250 (2013).
Manzoni, C. et al. mTOR independent regulation of macroautophagy by Leucine Rich Repeat Kinase 2 via Beclin-1. Sci. Rep. 6, 35106 (2016).
Williams, A. et al. Novel targets for Huntington’s disease in an mTOR-independent autophagy pathway. Nat. Chem. Biol. 4, 295–305 (2008).
Cina, D. P. et al. Inhibition of MTOR disrupts autophagic flux in podocytes. J. Am. Soc. Nephrol. 23, 412–420 (2012).
Button, R. W., Vincent, J. H., Strang, C. J. & Luo, S. Dual PI-3 kinase/mTOR inhibition impairs autophagy flux and induces cell death independent of apoptosis and necroptosis. Oncotarget 7, 5157–5175 (2016).
Grotemeier, A. et al. AMPK-independent induction of autophagy by cytosolic Ca2+increase. Cell Signal. 22, 914–925 (2010).
Yang, H. et al. Compound of Stout Camphor Medicinal Mushroom, Taiwanofungus camphoratus (Agaricomycetes), Induces Protective Autophagy in SPCA-1 Cells through AMPK Inhibition-Independent Blockade of the Akt/mTOR Pathway. Int. J. Med. Mushrooms 20, 727–738 (2018).
Vucicevic, L. et al. Compound C induces protective autophagy in cancer cells through AMPK inhibition-independent blockade of Akt/mTOR pathway. Autophagy 7, 40–50 (2011).
Elgendy, M. et al. Beclin 1 restrains tumorigenesis through Mcl-1 destabilization in an autophagy-independent reciprocal manner. Nat. Commun. 5, 5637 (2014).
Liu, C. et al. Autophagy-independent enhancing effects of Beclin 1 on cytotoxicity of ovarian cancer cells mediated by proteasome inhibitors. BMC Cancer 12, 622 (2012).
Elgendy, M. & Minucci, S. A novel autophagy-independent, oncosuppressive function of BECN1: degradation of MCL1. Autophagy 11, 581–582 (2015).
Athamneh, K. et al. Rhus coriaria increases protein ubiquitination, proteasomal degradation and triggers non-canonical Beclin-1-independent autophagy and apoptotic cell death in colon cancer cells. Sci. Rep. 7, 11633 (2017).
Smith, D. M. et al. Arsenic trioxide induces a beclin-1-independent autophagic pathway via modulation of SnoN/SkiL expression in ovarian carcinoma cells. Cell Death Differ. 17, 1867–1881 (2010).
Jakhar, R., Paul, S., Bhardwaj, M. & Kang, S. C. Astemizole-Histamine induces Beclin-1-independent autophagy by targeting p53-dependent crosstalk between autophagy and apoptosis. Cancer Lett. 372, 89–100 (2016).
Sun, L. et al. Beclin-1-independent autophagy mediates programmed cancer cell death through interplays with endoplasmic reticulum and/or mitochondria in colbat chloride-induced hypoxia. Am. J. Cancer Res. 5, 2626–2642 (2015).
Toton, E., Lisiak, N., Sawicka, P. & Rybczynska, M. Beclin-1 and its role as a target for anticancer therapy. J. Physiol. Pharmacol. 65, 459–467 (2014).
Liu, L. et al. Effect of inhibiting Beclin-1 expression on autophagy, proliferation and apoptosis in colorectal cancer. Oncol. Lett. 14, 4319–4324 (2017).
Shen, H. et al. Knockdown of Beclin-1 impairs epithelial-mesenchymal transition of colon cancer cells. J. Cell Biochem. 119, 7022–7031 (2018).
Koukourakis, M. I. et al. Beclin 1 over- and underexpression in colorectal cancer: distinct patterns relate to prognosis and tumour hypoxia. Br. J. Cancer 103, 1209–1214 (2010).
Corcelle, E. et al. Disruption of autophagy at the maturation step by the carcinogen lindane is associated with the sustained mitogen-activated protein kinase/extracellular signal-regulated kinase activity. Cancer Res. 66, 6861–6870 (2006).
Kuo, H. H. et al. Derivatives of 6-cinnamamido-quinoline-4-carboxamide impair lysosome function and induce apoptosis. Oncotarget 7, 38078–38090 (2016).
Zhao, Y. et al. The axis of MAPK1/3-XBP1u-FOXO1 controls autophagic dynamics in cancer cells. Autophagy 9, 794–796 (2013).
He, W. et al. FOXO1, a potential therapeutic target, regulates autophagic flux, oxidative stress, mitochondrial dysfunction, and apoptosis in human cholangiocarcinoma QBC939 Cells. Cell Physiol. Biochem. 45, 1506–1514 (2018).
Zhang, J. et al. Histone deacetylase inhibitors induce autophagy through FOXO1-dependent pathways. Autophagy 11, 629–642 (2015).
Zhao, Y. et al. Cytosolic FoxO1 is essential for the induction of autophagy and tumour suppressor activity. Nat. Cell Biol. 12, 665–675 (2010).
Schaffner, I. et al. FoxO function is essential for maintenance of autophagic flux and neuronal morphogenesis in adult neurogenesis. Neuron 99, 1188–1203 e1186 (2018).
Janji, B., Berchem, G. & Chouaib, S. Targeting autophagy in the tumor microenvironment: new challenges and opportunities for regulating tumor immunity. Front. Immunol. 9, 887 (2018).
Zhao, X. et al. Structural characterization and immunomodulatory activity of a water soluble polysaccharide isolated from Botrychium ternatum. Carbohydrate Polymers 171, 136–142 (2017).
Acknowledgements
We thank Dr. Thomas E. Eling at the National Institute of Environmental Health Sciences (NIEHS) for proofreading this manuscript. We also thank Junying Li and Li Xie at the Bio-ultrastructure Analysis Laboratory of Agrobiology and Environmental Sciences of Zhejiang University for providing technique support for TEM analysis. This study was supported by the National Natural Science Foundation of China (grant no. 81473397), and the Science and Technology Department of Zhejiang Province Foundation (2019C02100).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Q. Chen
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pan, H., Wang, Y., Na, K. et al. Autophagic flux disruption contributes to Ganoderma lucidum polysaccharide-induced apoptosis in human colorectal cancer cells via MAPK/ERK activation. Cell Death Dis 10, 456 (2019). https://doi.org/10.1038/s41419-019-1653-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-019-1653-7
This article is cited by
-
Ganoderma lucidum polysaccharide ameliorates cholesterol gallstone formation by modulating cholesterol and bile acid metabolism in an FXR-dependent manner
Chinese Medicine (2024)
-
Identification of Fangjihuangqi Decoction as a late-stage autophagy inhibitor with an adjuvant anti-tumor effect against non-small cell lung cancer
Chinese Medicine (2023)
-
Exploring the anti-cancer potential of Ganoderma lucidum polysaccharides (GLPs) and their versatile role in enhancing drug delivery systems: a multifaceted approach to combat cancer
Cancer Cell International (2023)
-
A review of anti-tumour effects of Ganoderma lucidum in gastrointestinal cancer
Chinese Medicine (2023)
-
DDRGK1, a crucial player of ufmylation system, is indispensable for autophagic degradation by regulating lysosomal function
Cell Death & Disease (2021)