Abstract
Ca2+ oscillation is a system-level property of the cellular Ca2+-handling machinery and encodes diverse physiological and pathological signals. The present study tests the hypothesis that Ca2+ oscillations play a vital role in maintaining the stemness of liver cancer stem cells (CSCs), which are postulated to be responsible for cancer initiation and progression. We found that niche factor-stimulated Ca2+ oscillation is a signature feature of CSC-enriched Hep-12 cells and purified α2δ1+ CSC fractions from hepatocellular carcinoma cell lines. In Hep-12 cells, the Ca2+ oscillation frequency positively correlated with the self-renewal potential. Using a newly developed high signal, endoplasmic reticulum (ER) localized Ca2+ sensor GCaMP-ER2, we demonstrated CSC-distinctive oscillatory ER Ca2+ release controlled by the type 2 inositol 1,4,5-trisphosphate receptor (IP3R2). Knockdown of IP3R2 severely suppressed the self-renewal capacity of liver CSCs. We propose that targeting the IP3R2-mediated Ca2+ oscillation in CSCs might afford a novel, physiologically inspired anti-tumor strategy for liver cancer.
Similar content being viewed by others
Introduction
Ca2+ signaling plays essential roles in the initiation and progression of diverse diseases including cancer. Recent studies have shown that dysregulation of Ca2+ homeostasis emerges as an important hallmark of tumor cells because remodeling of Ca2+ channels and transporters is common to tumor progression1,2 and Ca2+ oscillation orchestrates invadopodium formation and invasion during cancer metastasis3.
Cancer stem cells (CSCs) are thought to be the origin of cancer initiation and progression. It has been reported that Ca2+/calmodulin-dependent protein kinase IIγ prompts the stem-like properties of lung cancer cells4, while calcineurin, a Ca2+/calmodulin-dependent protein phosphatase, represses keratinocyte CSC potential5. Recently, we have shown that spontaneous Ca2+ oscillation occurs in the CSC-enriched liver cell line Hep-126, and modulates the efficiency of self-renewal6,7. Ca2+ oscillation decodes the stimulations not only by its amplitude, but also by its frequency and duration, thereby expanding the capabilities of the signaling pathway8,9,10.
The endoplasmic reticulum (ER), which extends over the entire cytoplasm as an elaborated nanotubular network, acts as the most important intracellular Ca2+ store. However, investigation of ER Ca2+ signaling has been limited by a lack of ER-localized, high-signal sensors that can report store Ca2+ dynamics in real time. Circularly permutated EGFP variants fused with calmodulin and M13 motif, termed GCaMPs or G-GECOs are robust Ca2+ sensors that have undergone progressive improvements11,12,13,14,15, and for which the structural basis of Ca2+-dependent fluorescence has been determined16. While GCaMPs have been optimized for the detection of rapid cytosolic Ca2+ signaling, there has been limited success in creating variants with high dynamic range within the micromolar Ca2+ endoplasmic/sarcoplasmic environment. Recent progress has been made in this area using new classes of fluorescent protein sensors17,18, or GCaMP variants with mutated calmodulin moieties18,19. However, the brightness and dynamic range of these proteins, have been limited.
In the present study, we aimed to test the hypothesis that Ca2+ oscillation might play a vital role in maintaining the stemness of CSCs. We investigated the Ca2+ phenotypes, their subcellular and molecular mechanisms, their significance in cancer biology and potential strategies for intervening CSCs through manipulating Ca2+ signaling in CSC-enriched Hep-12 cells. In addition, we developed GCaMP-ER2, a novel ER localized Ca2+ sensor, to probe the store Ca2+ dynamics simultaneously. Our comprehensive approaches allow us to propose that targeting the IP3R2-mediated Ca2+ oscillation in CSCs might afford a novel, physiologically inspired anti-tumor strategy for liver cancer.
Results
Ca2+ oscillation is a signature feature of liver CSCs
To test the hypothesis that Ca2+ oscillation is a hallmark of CSCs, we applied a panel of potential niche factors, including ATP, epidermal growth factor (EGF)/basic fibroblast growth factor (FGFb)/B27, and interleukin (IL)-6, and assessed the Ca2+ response in the liver CSC line Hep-12 and its matching hepatocellular cancer (HCC) line Hep-11. ATP is frequently released from cells to facilitate metastasis20. EGF is a well-accepted niche factor and EGF/FGFb/B27 are commonly used in media for in vitro spheroid formation assays, the most popular end-point experiment for self-renewal determination in solid tumors21,22. IL-6 has been reported to elevate or maintain the self-renewal of liver CSCs23. Besides these, we opted to include methacholine, a muscarinic receptor agonist capable of inducing dynamic Ca2+ changes in many types of cells24. All of these stimulants are linked to the inositol 1,4,5-trisphosphate (IP3) signaling pathway and thereby trigger intracellular Ca2+ release from the ER via IP3 receptors (IP3Rs)24,25,26,27.
We found that these stimulants each induced a large, slow Ca2+ transient that lasted several minutes in Hep-12 cells. Remarkably, Ca2+ oscillations of spiky appearance overlaid the evoked Ca2+ transient, and often continued even in the tail of the transient, when the cytosolic Ca2+ level returned to baseline (Fig. 1a, Movie S1). In contrast, typical Hep-11 cells did not show any Ca2+ change in response to most of the stimulants (EGF/FGFb/B27, IL-6, and methacholine), or exhibited only a smooth, monophasic Ca2+ transient in response to ATP (Fig. 1a, b, Fig. S1b, 2). That is, CSCs and HCCs exhibited distinct Ca2+ responses upon stimulation by self-renewal-related factors or IP3 pathway-related agonists.
a Cytosolic Ca2+ dynamics in different cell types. The liver cancer stem cell line Hep-12 was compared to another liver cancer cell line derived from the same patient (Hep-11), other cancer cell lines (LM3, MHCC97-L, and Huh7), and a nontumorigenic immortalized liver cell line (MIHA). Representative time courses are shown as the normalized Fluo-4 fluorescence signal (F/F0, where F0 refers to basal fluorescence). Arrows mark the onset of stimulation. b Statistics of Ca2+ oscillation frequencies. The number of Ca2+ oscillations on top of the large, slow Ca2+ transient was counted over a 15-min window starting at the onset of stimulation (n > 100 cells/group. ***P < 0.001 versus Hep-12 cells). c–f Differential Ca2+ responses to ATP or EGF/FGFb/B27 stimulation in α2δ1+ and α2δ1− cells from Hep-11 or Huh7 cells. Arrows in representative traces c, e mark the onset of stimulation. For statistics d, f, Hep-11: n = 40 (α2δ1+) and 86 cells (α2δ1−) for ATP stimulation; n = 36 (α2δ1+) and 69 cells (α2δ1−) for EGF/FGFb/B27 stimulation. Huh7: n = 120 (α2δ1+) and 137 cells (α2δ1−) for ATP stimulation; n = 109 (α2δ1+) and 94 (α2δ1−) cells for EGF/FGFb/B27 stimulation (***P < 0.001 versus α2δ1+ cells). All data were acquired with three independent experiments
We next extended the experiments to include multiple HCC lines, such as highly-metastatic LM3, poorly metastatic MHCC97-L, and Huh7 cells. ATP did elicit oscillatory Ca2+ responses in these cells, but their frequencies were significantly lower than those in Hep-12 cells (Fig. 1a, b). Further, none of the other three stimulants induced any oscillatory Ca2+ response in the vast majority of cells of each type (Fig. 1a, b). Similar results were found in the nontumorigenic immortalized liver cell line MIHA (Fig. 1a, b, Fig. S1b).
We have previously shown that α2δ1, a subunit associated with L-, P-, N-, and R-type Ca2+ channels, is a surface marker in liver CSCs6,7. We used the α2δ1 biomarker for sorting CSCs. More α2δ1+ Hep-11 cells than α2δ1− cells displayed Ca2+ oscillation and the ensemble-averaged oscillation frequency was 3.1- and 8.9-fold higher after stimulation with ATP and EGF/FGFb/B27, respectively (Fig. 1c, d). Likewise, α2δ1+ Huh7 cells also displayed higher Ca2+ oscillation frequency (Fig. 1e, f). Taken together, we conclude that Ca2+ oscillation is a signature that distinguishes liver CSCs from HCCs and nontumorigenic liver cells.
Development of high-signal ER-targeted Ca2+ sensor GCaMP-ER2
To measure ER Ca2+ dynamics, we developed GCaMP-ER2, a genetically encoded ER-targeting Ca2+ indicator with low Ca2+ affinity and high-dynamic range (Kd = 388 μM, ΔFmax/F0 = 40). Briefly, we screened a series of random and structure–based mutations in the four calmodulin EF-hand loops of G-GECO1.213, and identified a D129A mutant within the EF-hand of loop IV (GCaMP-L1, Fig. 2a), which has a dynamic range of 32 (∆Fmax/F0), and a Kd for Ca2+ of 106 μM. This variant was systematically mutated in EF-hand loops, resulting in an array of molecules with Ca2+ affinities ranging from 106 μM to 5.9 mM (Table S1). Figure 2a, b show the most promising ER sensor GCaMP-L2, with a dynamic range (∆Fmax/F0) of 40, a marked improvement over the previously highest signal strength ER reporter, GCaMPer/GCaMP3 (10.19)19 (∆Fmax/F0 = 14). As the Kd of GCaMP-L2 (388 μM) is well below the resting level of ER Ca2+, the indicator would be expected to operate near maximum brightness prior to ER Ca2+ release events.
a Schematic structure of low affinity GCaMPs (GCaMP-L1 and GCaMP-L2) and ER-targeting of GCaMP-L2 (GCaMP-ER2). b In vitro Ca2+ titration of G-GECO1.2 and low affinity mutants GCaMP-L1 and GCaMP-L2. The Kd of GCaMP-L2 is 388 µM and the dynamic range (∆Fmax/F0) is 40. c Thapsigargin (Tg) induced depletion of Ca2+ in the endoplasmic reticulum. The ER Ca2+ and cytosolic Ca2+ response were observed by co-expression of GCaMP-ER2 and R-CaMP1.01 in HeLa cells. The arrowhead indicated time of 2 µM Tg stimulation. n = 9 cells. d Subcellular distribution of GCaMP-ER2 and R-CaMP1.01 in HeLa cells. R-CaMP1.01 is localized to the cytoplasm and nucleus (scale bar, 10 μm)
Targeting of GCaMP-L2 to the ER was achieved by fusion of the N-terminal calreticulin ER targeting sequence MLLSVPLLLGLLGLAVA and the C-terminal ER retention signal KDEL. Addition of a KL(AP)6 linker between CaM and KDEL improved GCaMP-L2 brightness in HeLa cells, resulting in GCaMP-ER2 (Fig. 2a), which exhibited high fluorescence changes when thapsigargin was used to deplete ER Ca2+ (Fig. 2c). Figure 2d displays intracellularly compartmentalized signals in a HeLa cell recorded simultaneously with our newly developed GCaMP-ER2 and cytosol and nuclear localized RCaMP1.01.
Distinctive ER Ca2+ dynamics in liver CSCs
Ca2+ oscillation is an emergent property of the entire cellular Ca2+-handling system. To investigate the cellular mechanism underlying the distinctive Ca2+ response of CSCs, we measured the resting cytosolic Ca2+ concentration ([Ca2+]c) and the capacity of the releasable ER Ca2+ store. The latter was indexed as the peak cytosolic Ca2+ concentration during release stimulated by the Ca2+ ionophore A23187 (10 μM) ([Ca2+]release) (Fig. 3a). We found that both [Ca2+]c and [Ca2+]release were similar in the two types of cells (Fig. 3b, c), excluding the possibility that Ca2+ oscillation in Hep-12 cells is brought about by a cytosolic or ER Ca2+ overload. Then we sought to measure ER Ca2+ dynamics. With the newly developed GCaMP-ER2 and Rhod-4, a cytosol-retained Ca2+ indicator, we simultaneously assessed the dynamic responses in both ER and cytosol (Fig. S2b). In Hep-11 cells, the ER Ca2+ store declined precipitously upon ATP stimulation, mirroring the rapid rise of the cytosolic Ca2+ transient (Fig. 3e). Subsequently, the GCaMP-ER2 fluorescence remained stable at a low level (nadir ΔF/F0 = −0.77) (Fig. 3f) even though the cytosolic Ca2+ level was returning toward an elevated steady-state level (Fig. 3f). This result suggests that the ER Ca2+ recycling mechanism is ineffective, presumably because of continued opening of the ER Ca2+ release channels. In Hep-12 cells, however, the initial ER Ca2+ depletion was smaller (nadir ΔF/F0 = −0.34) (Fig. 3d, f), and strikingly, it was rapidly restored and often followed by a conspicuous overshoot (peak F/F0 = 1.16) (Fig. 3d, g). The peak [Ca2+]c, however, was comparable in both types of cells (F/F0 = 3.68 ± 0.18 for Hep-11 and 3.23 ± 0.16 for Hep-12) (Fig. S2c, d). Close observation further revealed that each spiky cytosolic Ca2+ oscillation was accompanied by an antiphasic ER Ca2+ oscillation (Fig. 3d inset). This finding indicates that the cytosolic Ca2+ oscillation originates from periodic ER release and recycling. ER store recovery and overshoot were also found in the subpopulation of MHCC97-L cells with stimulated Ca2+ oscillation, whereas persistent ER Ca2+ depletion was associated with the population of cells lacking the oscillatory behavior (Fig. S2e). Taken together, we concluded that distinctive regulation of ER Ca2+ dynamics underlies the Ca2+ oscillation phenotype characteristic of liver CSCs.
a–c Measurements of the resting cytosolic Ca2+ level ([Ca2+]c) and the releasable ER Ca2+ store ([Ca2+]release) in Hep-12 and Hep-11 cells. a Representative trace and experimental protocol (details in Methods). b Scatter-plots of [Ca2+]c. c Scatter-plots of [Ca2+]release, an index of the Ca2+ content in the ER. n = 51 cells. d–g ER Ca2+ recovers and overshoots in Hep-12 cells displaying Ca2+ oscillation. d, e Representative time courses of cytosolic and ER Ca2+ dynamics in Hep-12 (d) and Hep-11 cells (e) upon 40 μM ATP stimulation. Dashed lines indicate the basal ER Ca2+ level. Inset shows an enlarged view of ER and cytosolic Ca2+ oscillations. f, g Statistics of the nadir f and maximal levels of ER Ca2+ during the 10 min of observation (g). Note the overshoot after initial depletion of ER Ca2+ in Hep-12 cells. n = 80 cells for Hep-12 and 54 for Hep-11. Data are shown as mean ± SEM (***P < 0.001 versus Hep-11 cells). All data were acquired with three independent experiments
Role of ER Ca2+ cycling in the genesis of Ca2+ oscillation
Store-operated Ca2+ entry (SOCE), the main Ca2+ entry pathway in nonexcitable cells, is activated upon depletion of the ER Ca2+ and plays pivotal roles in the timely replenishment of the internal Ca2+ store. In both cell types, inhibition of ER Ca2+ uptake by thapsigargin (4 μM) caused a biphasic cytosolic Ca2+ increase, with the second peak reflecting the SOCE capacity (Fig. 4a). It was clear that the Hep-12 cells displayed a greater SOCE component (Fig. 4b). In addition, faster ER Ca2+ uptake occurred in Hep-12 cells (Fig. 4c–e). To further delineate the specific role of SOCE, we measured cytosolic and ER Ca2+ dynamics in the absence of extracellular Ca2+. We found that the ATP-stimulated initial Ca2+ transients remained essentially unchanged; Ca2+ oscillation occurred in the early phase, but was quickly damped and eventually disappeared in 200 s (Fig. 4g). Concomitantly, the ER Ca2+ store showed a general trend of depletion without replenishment (Fig. 4g, l). Application of 100 μM 2-aminoethoxydiphenyl borate (2-APB), an inhibitor of both SOCE and IP3Rs at this concentration, greatly attenuated the Ca2+ oscillation and prevented the ER store from recovering (Fig. 4h, j, l). The initial Ca2+ transient was also markedly smaller and briefer (Fig. 4k), perhaps because of partial inhibition of release. Simultaneous inhibition of Ca2+ entry and ER Ca2+ recycling (0 Ca2+ and 4 μM thapsigargin) had additive effects (Fig. 4i, j, l). These results indicate that the persistence of Ca2+ oscillation is attributable to the rapid refilling and overshoot of the ER store due to enhanced SOCE and ER Ca2+ uptake. Nonetheless, they are not obligatory for the genesis of stimulated Ca2+ oscillation; rather, they play a permissive role in preventing it from damping.
a, b SOCE in Hep-11 and Hep-12 cells. a Time courses of averaged cytosolic and ER Ca2+ transients in cells in which ER Ca2+ reuptake was inhibited with 4 μM Tg. Arrows mark the second peaks of cytosolic Ca2+ transients, reflecting the magnitude of SOCE. b Amplitudes of the second peaks (n = 62 cells for Hep-12 and 53 for Hep-11). c–e ER Ca2+ uptake in Hep-11 and Hep-12 cells. c Time courses of averaged ER Ca2+ changes after addition of 1 μM Ca2+. Cells were first permeabilized by 50 μg/mL saponin and ER Ca2+ was depleted by 4 μM Tg. d ER Ca2+ uptake time. e Maximal ER Ca2+ level during the 5-min observation (n = 38 cells for Hep-12 and 110 for Hep-11). f–l Role of SOCE and SERCA in Ca2+ oscillation in Hep-12 cells. Representative time courses of cytosolic and ER Ca2+ responding to 40 μM ATP without other treatment f or in conditions of (1) Ca2+-free solution without 4 μM Tg g; (2) addition of 100 μM 2-APB h; and (3) Ca2+-free solution with 4 μM Tg i. j–l Statistics of oscillation frequency j, first peak amplitude k, and the ER Ca2+ nadir l (n = 79 cells for control, 84 for 0 Ca2+, 95 for 100 μM 2-APB, and 117 for 0 Ca2+ and 4 μM Tg). Data are shown as mean ± SEM (***P < 0.001 versus Hep-11 b, e or control j–l). All data were acquired with three independent experiments
IP3R2 as the intrinsic oscillator underlying Ca2+ dynamics in Hep-12 cells
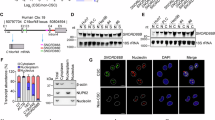
Next, we sought to identify the key molecular players underlying the Ca2+ oscillation phenotype. By RNA-sequencing analysis, we identified 1887 genes expressed differently in Hep-12 versus Hep-11 cells (Fig. 5a). Among them, 175 genes were found in Ca2+-related Gene Ontology (1093 genes included) (Fig. 5a). Interestingly, of the ten genes that directly participate in Ca2+ transport, all were significantly upregulated in Hep-12 cells (Fig. 5a). These genes code proteins for pore subunits of the N-type (α1B) and T-type (α1G) Ca2+ channels, subunits associated with most voltage-operated Ca2+ channels (VOCCs) (α2δ1, α2δ2, and γ5), ER Ca2+ release channels (IP3R1, IP3R2, and RyR2), and the ER Ca2+ uptake pump (SERCA3), as well as transient receptor potential TRPC4 channels. Notably, α2δ1, which helps to anchor VOCCs in the plasma membrane, has been previously reported by us as a functional liver CSC marker6,7.
a Volcano plots of mRNA level differences in Hep-12 and Hep-11 cells. Light blue dots mark calcium regulators and Deep blue dots represent specific direct Ca2+-handling molecules. b, c Effects of knockdown of Ca2+ regulators identified in (a) on ATP (40 μM)-stimulated Ca2+ oscillation. b Representative Ca2+ dynamics after α2δ1 and IP3R2 knockdown. Arrows mark time of ATP stimulation. c Statistics of oscillation frequency (n > 100 cells; data are shown as mean ± SEM; *P < 0.05 versus scrambled control, **P < 0.01, ***P < 0.001). d, e During 40 μM ATP treatment, ER Ca2+ continued to drop in Hep-12 cells with IP3R2 knockdown. d Time courses of averaged cytosolic Ca2+ and ER Ca2+ of IP3R2-knockdown Hep-12 cells responding to 40 μM ATP. e Statistics of ER Ca2+ nadir (n = 80 cells in control, 37 for IP3R2 KD1, and 37 for IP3R2 KD2). All data were acquired with three independent experiments
To identify key Ca2+ regulators, we targeted the top five genes on the list, CACNA2D1, CACNA2D2, ITPR1, ITPR2, and ATP2A3, and determine the consequences of the Ca2+ oscillation phenotype. We used α2δ1 as a positive control on some occasions, and included ITPR3 (IP3R3 gene) alongside ITPR1 (IP3R1 gene) and ITPR2 (IP3R2 gene) to acquire a comprehensive view of the IP3R family. We established stable cell lines harboring shRNAs against these genes (Fig. S3a). Functional analysis revealed that Ca2+ oscillation frequency was severely suppressed after knockdown of α2δ1 and IP3R2, modestly decreased with knockdown of IP3R1 and SERCA3, and largely unchanged after knockdown of α2δ2 and IP3R3 (Fig. 5b, c, Fig. S3b). The amplitude of the first peak, which reflects the ER Ca2+ content as well as fractional release, was mildly decreased (knockdown of IP3R1, IP3R2, or IP3R3) or unchanged (knockdown of α2δ1, α2δ2, or SERCA3) (Fig. S3c). These results show that multiple Ca2+ regulators orchestrate the phenotype of Ca2+ oscillation. Nonetheless, other than α2δ1, IP3R2 emerged as the most important molecular determinants. Particularly, after IP3R2 knockdown, the ER Ca2+ phenotype was drastically altered, with greater initial release and without any recovery (Fig. 5d, e), resembling those in non-oscillatory HCCs. That is, IP3R2 may serve as an intrinsic oscillator essential for Ca2+ oscillation. Previous studies have also shown that, unlike IP3R1 and IP3R3, IP3R2 is prone to generate the Ca2+ oscillation behavior28,29,30.
Ca2+ oscillation frequency positively correlates with spheroid-forming efficiency in Hep-12 cells
To determine the functional significance of Ca2+ oscillation in CSC biology, we assessed the efficiency of spheroid formation, which reflects the self-renewal capacity, in relation to Ca2+ oscillation frequency over a broad range of experimental conditions. The experimental groups included: (i) knockdown of the VOCC subunits α2δ1 and α2δ2; (ii) knockdown of the ER release channels IP3R1, IP3R2, and IP3R3; (iii) removal of extracellular Ca2+; (iv) inhibition of SOCE; and (v) treatment with BAPTA-AM or EGTA-AM to retard Ca2+ changes. We found that, while inhibiting Ca2+ oscillation, both α2δ1 and α2δ2 knockdown significantly compromised the self-renewal (Fig. 6a, b, Fig. S4a), confirming and extending our previous report6. Spheroid formation was also reduced when extracellular Ca2+ was removed, SOCE was inhibited or cytosolic Ca2+ was buffered (Fig. 6a, c, Fig. S4a). Notably, of the three IP3R subtypes, knockdown of IP3R2, but not IP3R1 and IP3R3, greatly curtailed the spheroid formation (Fig. 6a, b, Fig. S4a).
a Phase-contrast images of spheroids formed by Hep-12 cells under different conditions (scale bar: 100 μm). b, c Statistics of spheroid-forming efficiency. b Scrambled control group and shRNA knockdown groups. c Groups with and without drug treatment. One hundred cells per well were plated. Spheroids (R > 100 μm) were counted under a stereomicroscope (n = 6–12; ***P < 0.001 versus control). d Scatter-plot of oscillation frequency versus spheroid-forming efficiency. Linear regression (solid line) yielded a positive correlation coefficient (r) of 0.76 (P < 0.001). Data are expressed as mean ± SEM, n = 18. All data were acquired with at least three independent experiments
From these data, a general pattern emerged in which lowering the Hep-12 Ca2+ oscillation frequency effectively decreased the efficiency of spheroid formation, regardless of whether the type of Ca2+ manipulation was genetic, biophysical, or pharmacological. For quantitative analysis, we summarized our data in a scatter-plot (Fig. 6d). Linear regression revealed a strong positive correlation between spheroid-forming efficiency and Ca2+ oscillation frequency (r = 0.76, p < 0.001) (Fig. 6d). This result strongly suggests that the signature Ca2+ oscillation in liver CSCs plays an important functional role in promoting their self-renewal.
Targeting IP3R2-mediated Ca2+ oscillation suppresses tumor formation of liver CSCs
In light of the newly established relationship between Ca2+ oscillation and spheroid formation, we hypothesized that targeting the Ca2+ oscillation machinery might provide an effective therapeutic strategy to limit CSC expansion and therefore retard or even prevent the genesis of tumors. Among the aforementioned molecular participants in the genesis and regulation of Ca2+ oscillation, IP3R2 is particularly noteworthy, not only because it is greatly upregulated in liver CSCs, but also its cyclic opening may serve as the intrinsic oscillator28,29,30. To further evaluate the impact of IP3R2 on the self-renewal of Hep-12 cells, we injected 103 or 102 scrambled control cells and IP3R2-knockdown cells subcutaneously into nonobese diabetic/severe combined immunodeficient (NOD/SCID) mice to test their tumorigenicity potential. Tumors were dissected and weighed 42 or 52 days after injection of 103 or 102 cells, respectively. As shown in Fig. 7a, b, the tumorigenicity of Hep-12 cells was significantly suppressed by IP3R2 knockdown. To further confirm the role of IP3R2, we purified α2δ1+ CSCs from Huh7 cells and assayed spheroid formation and tumorigenicity with IP3R2 knockdown. Both the efficiency of spheroid formation and the frequency of tumorigenic cells were markedly inhibited (Fig. 7c, d), substantiating the pivotal role of IP3R2 in the self-renewal of liver CSCs.
a, b In vivo tumorigenicity of IP3R2-knockdown Hep-12 cells. One thousand (a) or one hundred cells (b) per mouse were injected (n = 7). Left: Images showing dissected tumors at termination of the experiment time (scale bars: 1 cm). Right: Tumor weight. Data are shown as mean ± SEM (*P < 0.05 versus control, ***P < 0.001). c IP3R2 knockdown suppresses self-renewal of α2δ1+ Huh7 cells. Left: Phase-contrast images of spheroid formation. Right: Statistics of spheroid-forming efficiency. One hundred cells per well were plated. Spheroids (R > 100 μm) were counted under a stereomicroscope (n = 9; **P < 0.01 versus control, ***P < 0.001. d Tumorigenic cell frequency in α2δ1+ Huh7 cells with IP3R2 knockdown in NOD/SCID mice
Discussion
In this study, we demonstrate that Ca2+ oscillation is a functional signature of liver CSCs, and provide evidence that targeting Ca2+ oscillation can effectively limit the self-renewal of liver CSCs and thereby tumor initiation and progression. In response to a panel of niche factors (ATP, EGF/FGFb/B27, and IL-6), robust Ca2+ oscillation occurred in liver CSCs, including CSC-enriched Hep-12 cells, the small portion of α2δ1+ Hep-11 cells, and α2δ1+ Huh7 cells. This Ca2+ phenotype is CSC-specific because the vast majority of matching Hep-11 and Huh7 cancer cells did not show similar Ca2+ oscillation, and neither did HCC LM3, MHCC97-L, and the immortalized liver cell line MIHA. This finding is in general agreement with previous reports that Ca2+ oscillation occurs in a portion of highly metastatic, but not in weakly/non-metastatic human prostate and breast cancer cells31. Such CSC Ca2+ oscillation plays important roles in cancer biology. The efficiency of spheroid formation was significantly decreased by suppressing Ca2+ oscillation by removing extracellular Ca2+, inhibiting SOCE, buffering cytosolic Ca2+, or knocking down key Ca2+ regulators (IP3R2 or α2δ1). Strikingly, there was a positive correlation between the Ca2+ oscillation frequency and spheroid formation efficiency under a broad range of conditions. More importantly, we showed that CSCs with defective Ca2+ oscillation, i.e., knockdown of IP3R2, exhibited a greatly compromised ability to form tumors in NOD/SCID mice.
Ca2+ oscillation reflects a system-level, emergent behavior of the complex Ca2+-handling machinery. To sustain a persistent cyclic Ca2+ oscillation, the ER store must be kept replenished. With our newly developed ER Ca2+ sensor GCaMP-ER2, we disclosed a surprising new insight at the cellular level. In CSCs undergoing niche factor-stimulated Ca2+ oscillation, initial ER depletion was followed by a complete restoration and oftentimes an overshoot of the ER Ca2+ content. In HCCs displaying no Ca2+ oscillation, however, stimulated ER Ca2+ release was followed by a deep ER depletion without any sign of recovery over the period of observation (15 min). The distinctive ER Ca2+ dynamics are in part attributable to higher activity of SOCE and ER Ca2+ recycling in CSCs (this study) as well as Ca2+ entry through VOCCs as we reported previously6. More importantly, these results strongly suggest that the ER release channels in CSCs undergo cyclic, brief openings, allowing for ER Ca2+ retention and supporting the Ca2+ oscillation behavior, whereas those in HCCs may be kept open after stimulation, preventing replenishment of the ER Ca2+. Through RNA-seq and bioinformatics analysis confirmed by functional validation, we identified the ER release channel IP3R2 as the intrinsic oscillator driving the CSC Ca2+ phenotype, while the other Ca2+ proteins examined serve largely permissive roles (e.g., α2δ1 and SERCA3).
The IP3R family consists of three subtypes, IP3R1, IP3R2, and IP3R3, which share 60–80% homology of amino acid sequence, and the subtype expression is differentially regulated in response to physiological and pathological stimuli. There is a bell-shaped Ca2+-dependence of the channel activity: activation via the CICR mechanism at low [Ca2+]i, inhibition via Ca2+-dependent inactivation at higher [Ca2+]i32. These properties lay the foundation for Ca2+ oscillation in nonexcitable cells. All three IP3R subtypes are similar in terms of permeability, but their high-Ca2+ inactivation properties are variable33. Functional experiments have shown that IP3R2 is required for long-lasting and regular Ca2+ oscillation28,29,30, IP3R1 is responsible for short and irregular Ca2+ oscillation28, and IP3R3 produces a monophasic Ca2+ spike28 and even has anti-oscillatory actions28,34. More interestingly, IP3R2 co-expression with IP3R1 or IP3R3 facilitates Ca2+ oscillation28. In this regard, we found that IP3R2 was specifically expressed in liver CSCs but not HCCs; and knockdown of IP3R2, but not IP3R1 and IP3R3, repressed Ca2+ oscillation (Fig. 5b, c, Fig. S3b), and hence the capacity for self-renewal in CSCs (Fig. 6a, b, Fig. S4a). Knockdown of IP3R2 also caused ER Ca2+ phenotypes similar to that in a typical Hep-11 cell. Collectively, these data indicate that IP3R2 is a target of choice for limiting CSC self-renewal through manipulating Ca2+ oscillation. Indeed, CSCs with knockdown of IP3R2 showed depressed sphere formation (Figs. 6a, b, 7c) and in vivo tumorigenesis (Fig. 7a, b, d).
CSCs rely on niches to maintain their stemness and prompt metastasis35. In this regard, we found that all niche factors examined were able to elicit robust Ca2+ oscillation in liver CSCs, supporting the hypothesis that Ca2+ signaling is at the root of CSC maintenance and survival. Ca2+ oscillation can mediate cellular signaling in a frequency-modulatory mode, with the frequency varying from tens of Hz to tens of mHz. Many protein kinases, protein phosphatases, and transcription factors are known to decipher Ca2+ oscillation, and each has a specific dependence on oscillation frequency, interestingly with little overlap36. Future investigations to determine the involvement of these decoder proteins in CSC Ca2+ oscillation to signal self-renewal are warranted.
In summary, we have shown that niche factor-stimulated Ca2+ oscillation is a signature feature pivotal to self-renewal of liver CSCs. Further, through the development of a new genetically coded ER Ca2+ sensor GCaMP-ER2, we have identified IP3R2, which is enriched in CSCs, as the intrinsic oscillator driving such Ca2+ oscillation. Targeting Ca2+ oscillation in general and IP3R2 in particular might afford a physiologically inspired anti-tumor strategy by uprooting CSCs to limit tumor initiation, progression, recurrence, and drug resistance.
Materials and methods
Cell culture and establishment of stable target gene-knockdown cell lines
Hep-12, Hep-11, and Huh7 cells37 were cultured in RPMI 1640 medium (Invitrogen) supplemented with 10% fetal bovine serum (FBS) (Invitrogen) at 37 ℃ under 5% CO2. The LM3 and MHCC97-L cell lines were gifted by Dongqin Yang (Huashan Hospital, Fudan University), and the MIHA cell line was gifted by Jinying Ning (Crownbio Co., Beijing, China). The HeLa cell line was kept in our labs. All four cell lines were cultured in Dulbecco’s modified Eagle’s medium (Invitrogen) supplemented with 10% FBS. To generate stable Hep-12 cell lines with knockdown of target genes, packaging plasmids pLP1, pLP2, and pVSVG (Invitrogen) were transfected with expression plasmids harboring shRNA targeting the specific genes into 293FT cells at the ratio 1:1:1:3, then the lentivirus was collected and cells were infected. The puromycin-resistant cells were cultured and stable target gene-knockdown cell lines were acquired. ShRNAs (Sigma) were gifted by Guoqiang Bi (University of Science and Technology of China) and their sequences are listed in Table S2.
Calcium measurement and imaging
Fluo-4 AM (Invitrogen) was used to monitor cytosolic Ca2+ dynamics. When ER Ca2+ was measured simultaneously using GCaMP-ER2, Rhod-4 AM (AAT Bioquest) was used for cytosolic Ca2+ detection. Briefly, cells were incubated in the presence of Fluo-4 AM (5 μM) or Rhod-4 AM (5 μM) at room temperature for 30 min. After washing with Tyrode’s solution consisting of (in mM) 137 NaCl, 5.4 KCl, 1.2 MgCl2, 1.2 NaH2PO4, 1.8 CaCl2, 10 glucose, and 20 HEPES (pH 7.35, adjusted with NaOH), Fluo-4 or Rhod-4 fluorescence was measured with a Zeiss LSM 710 confocal microscope equipped with a 40×, 1.3 NA oil-immersion objective. GCaMP-ER2 and RCaMP1.01 plasmids were co-transfected into HeLa cells using Lipofectamine 2000 (Invitrogen) 48–72 h before imaging and measurement. For ER Ca2+ measurement in CSCs and HCCs, adenovirus expressing GCaMP-ER2 was used to infect the cells 48 h before measurement. For Fluo-4 fluorescence measurement, cells were excited at 488 nm and emission collected at 493–622 nm. For GCaMP-ER2 and Rhod-4 dual indicator measurement, images were captured in multi-track mode with excitation at 488 or 543 nm and emission collection at 490–516 or 556–733 nm, respectively.
The cytosolic Ca2+ level was quantified using the equation [Ca2+] = Kd (F − Fmin)/(Fmax − F). After measurement of fluorescence at rest (F0), cells were exposed to Tyrode’s solution (zero Ca2+) with 4 mM EGTA, 5 μM thapsigargin, and 10 μM A23187. Store Ca2+ release was induced and cytosolic Ca2+ increased abruptly and reached the peak in ~10 s (Frelease) followed by a gradual decline. When stabilized, 100 μM BAPTA-AM was added to further chelate cytosolic Ca2+ to measure the minimum fluorescence level (Fmin). Then, for measurement of the maximal level (Fmax), cells were exposed to Tyrode’s solution containing 10 mM Ca2+, 5 μM thapsigargin, 12 μM A23187, 3 μM FCCP, and 20 mM 2-DG. Assuming that Kd = 1 μM for Fluo-4 in intact cells38, the resting Ca2+ [Ca2+]c, and the releasable store Ca2+ indexed by [Ca2+]release were obtained with the above equation.
Synthesis of low-affinity GCaMP (GCaMP-L) mutants
Mutations of CaM were introduced into the G-GECO1.2 sequence by overlap extension PCR. The amplified product was inserted into the NheI/HindIII sites of pRSET-A expression vector (Invitrogen). The mutant proteins were expressed in E. coli BL21 Star (DE3) pLysS cells and purified using Ni-charged resins as previously described39. After elution, the buffer was changed to 30 mM MOPS (pH 7.2) with 100 mM KCl using an Amicon Ultra-4 filter unit (Millipore). Protein concentration was measured using BCA Protein Assay (Pierce).
In vitro characterization of purified proteins
Calcium titration of G-GECO1.2 was performed by Calcium Calibration Buffer Kit #1 (Invitrogen). For calcium titration of low affinity mutants, a series of zero to 10 mM [Ca2+]free buffer were made in 1 mM EGTA, 50 mM MOPS, and 100 mM KCl (pH 7.2) and [Ca2+]free concentrations were calculated using WEBMAXC EXTENDED program (maxchelator.stanford.edu). The fluorescence of 1 µM purified protein in various [Ca2+]free buffers were measured with excitation at 485/20 nm and emission at 516/20 nm using a Synergy 2 Microplate Reader (Biotek).
Construction of ER-targeted GCaMP-ER2
The GCaMP-L2 was targeted to and retained in the ER via the N-terminal calreticulin ER targeting sequence MLLSVPLLLGLLGLAVA and the C-terminal ER retention signal KDEL, respectively, with a linker KL(AP)6 between CaM and retention signal. The final construct was generated by PCR with primers containing described coding sequences and GCaMP-L2 template. The PCR product was cloned into the pEGFP-N1 mammalian expression vector (replacing EGFP) using BglII and NotI restriction sites.
Cell labeling and flow cytometry
The α2δ1 antibody (Abcam) was directly labeled with the Lightning-Link PE-Cy5 Labeling Kit following the vendor’s protocol (Innova Biosciences). For flow cytometry, cells were digested, dispersed, labeled, and analyzed as previously described40.
Sphere formation assay
Cells were plated in ultra low attachment 96-well plates (Corning) and cultured in Dulbecco’s modified Eagle’s medium/F12 (Invitrogen) supplemented with B27 (Invitrogen), 40 ng/ml epidermal growth factor (Invitrogen), 40 ng/ml basic fibroblast growth factor (Peprotech), and 1% methylcellulose (Sigma). Cells were incubated for 2–3 weeks, and spheres were counted under a stereomicroscope (Olympus).
Tumorigenicity assay in NOD/SCID mice
Cells were suspended in 50 μL of a 1:1 mixture of plain RPMI 1640 and Matrigel (BD Biosciences) and injected subcutaneously into the back of 4- to 6-week-old NOD/SCID mice (Vital River). Tumor formation was monitored weekly.
To assess the tumorigenicity detection of α2δ1+ Huh7 cells with IP3R2 knockdown, sorted cells were first infected with shRNA-IP3R2 or scrambled control lentivirus for 4 h at 37 ℃.
All animals were treated in compliance with the Guide for the Care and Use of Laboratory animals published by the US National Institutes of Health (NIH Publication No. 85-23, revised 1996) and approved by the Animal Care and Use Committee of Peking University (accredited by AAALAC international).
RNA sequencing (RNA-seq) analysis
Using TRIzol reagent (Invitrogen), total RNA was isolated from cells according to the manufacturer’s protocol. The quality of the extracted RNA was controlled using Agilent 2100. A sequencing library was prepared using the NEBNext®UltraTM RNA Library Prep Kit for Illumina® (New England Biolabs) following the manufacturer’s recommendations. Deep sequencing was then performed on an Illumina Hiseq 4000 platform and 150-bp paired-end reads were generated. To obtain the differentially-expressed genes, RNA-seq reads were first aligned to the human genome (hg19) with TopHat (v 1.2.0) and all the mapping results were evaluated as previously described41. Cuffdiff software (v 2.0.2) was used to obtain differentially expressed genes with a cutoff of fold-change > 2 and q value < 0.05. By searching Gene Ontology (http://www.geneontology.org/) we found Ca2+-related genes distributed in process, function, and component.
Western blotting
Cells lysates were obtained by incubating cells directly with sodium dodecyl sulphate polyacrylamide gel electrophoresis (SDS-PAGE) loading buffer. After ultrasonicating 5 times (5 s each), lysates were heated at 100 °C for 10 min. Proteins were separated on 6% SDS-PAGE gel (for IP3R expression) or 8% SDS-PAGE gel (for α2δ1, α2δ2, and SERCA3 expression) and transferred to a 0.45-μm polyvinylidene difluoride membrane (Millipore). Membranes were blocked with 5% bovine serum albumin (for IP3R expression) or 5% nonfat dry milk (for α2δ1, α2δ2, and SERCA3 expression) and incubated with primary antibody overnight at 4 °C. Primary antibodies against IP3R1 (Abcam, 1:500), IP3R2 (Millipore, 1:50), IP3R3 (BD Biosciences, 1:1000), α2δ1 (Abcam, 1:1000), α2δ2 (Sigma, 1:2000), SERCA3 (Abcam, 1:500), and tubulin (Sigma-Aldrich, 1:2000) were used.
Statistics
The data are expressed as the mean ± SEM and, when appropriate, Student’s t test was applied to determine statistical significance. P < 0.05 was considered statistically significant.
References
Roderick, H. L. & Cook, S. J. Ca2+ signalling checkpoints in cancer: remodelling Ca2+ for cancer cell proliferation and survival. Nat. Rev. Cancer 8, 361–375 (2008).
Prevarskaya, N., Skryma, R. & Shuba, Y. Ion channels and the hallmarks of cancer. Trends Mol. Med. 16, 107–121 (2010).
Sun, J. et al. STIM1- and Orai1-mediated Ca(2+) oscillation orchestrates invadopodium formation and melanoma invasion. J. Cell Biol. 207, 535–548 (2014).
Chai, S. et al. Ca2+/calmodulin-dependent protein kinase IIgamma enhances stem-like traits and tumorigenicity of lung cancer cells. Oncotarget 6, 16069–16083 (2015).
Wu, X. et al. Opposing roles for calcineurin and ATF3 in squamous skin cancer. Nature 465, 368–372 (2010).
Zhao, W. et al. 1B50-1, a mAb raised against recurrent tumor cells, targets liver tumor-initiating cells by binding to the calcium channel alpha2delta1 subunit. Cancer Cell 23, 541–556 (2013).
Han, H. et al. PBX3 is targeted by multiple miRNAs and is essential for liver tumour-initiating cells. Nat. Commun. 6, 8271 (2015).
Levine, J. H., Lin, Y. & Elowitz, M. B. Functional roles of pulsing in genetic circuits. Science 342, 1193–1200 (2013).
Dolmetsch, R. E., Xu, K. & Lewis, R. S. Calcium oscillations increase the efficiency and specificity of gene expression. Nature 392, 933–936 (1998).
Cai, L., Dalal, C. K. & Elowitz, M. B. Frequency-modulated nuclear localization bursts coordinate gene regulation. Nature 455, 485–490 (2008).
Tallini, Y. N. et al. Imaging cellular signals in the heart in vivo: cardiac expression of the high-signal Ca2+ indicator GCaMP2. Proc. Natl Acad. Sci. USA 103, 4753–4758 (2006).
Tian, L. et al. Imaging neural activity in worms, flies and mice with improved GCaMP calcium indicators. Nat. Methods 6, 875–881 (2009).
Zhao, Y. et al. An expanded palette of genetically encoded Ca(2)(+) indicators. Science 333, 1888–1891 (2011).
Ohkura, M. et al. Genetically encoded green fluorescent Ca2+ indicators with improved detectability for neuronal Ca2+ signals. PLoS ONE 7, e51286 (2012).
Chen, T. W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Wang, Q., Shui, B., Kotlikoff, M. I. & Sondermann, H. Structural basis for calcium sensing by GCaMP2. Structure 16, 1817–1827 (2008).
Tang, S. et al. Design and application of a class of sensors to monitor Ca2+ dynamics in high Ca2+ concentration cellular compartments. Proc. Natl. Acad. Sci. USA 108, 16265–16270 (2011).
Suzuki, J. et al. Imaging intraorganellar Ca2+ at subcellular resolution using CEPIA. Nat. Commun. 5, 4153 (2014).
Henderson, M. J. et al. A Low Affinity GCaMP3 Variant (GCaMPer) for Imaging the Endoplasmic Reticulum Calcium Store. PLoS ONE 10, e0139273 (2015).
Schneider, G. et al. Extracellular nucleotides as novel, underappreciated pro-metastatic factors that stimulate purinergic signaling in human lung cancer cells. Mol. Cancer 14, 201 (2015).
Shan, J. et al. Nanog regulates self-renewal of cancer stem cells through the insulin-like growth factor pathway in human hepatocellular carcinoma. Hepatology 56, 1004–1014 (2012).
Valent, P. et al. Cancer stem cell definitions and terminology: the devil is in the details. Nat. Rev. Cancer 12, 767–775 (2012).
Iliopoulos, D., Hirsch, H. A. & Struhl, K. An epigenetic switch involving NF-kappaB, Lin28, Let-7 MicroRNA, and IL6 links inflammation to cell transformation. Cell 139, 693–706 (2009).
Aub, D. L. & Putney, J. W. Jr. Mobilization of intracellular calcium by methacholine and inositol 1,4,5-trisphosphate in rat parotid acinar cells. J. Dent. Res. 66, 547–551 (1987).
Salter, M. W. & Hicks, J. L. ATP causes release of intracellular Ca2+ via the phospholipase C beta/IP3 pathway in astrocytes from the dorsal spinal cord. J. Neurosci. 15, 2961–2971 (1995).
Mine, T., Kojima, I., Ogata, E. & Nakamura, T. Comparison of effects of HGF and EGF on cellular calcium in rat hepatocytes. Biochem. Biophys. Res. Commun. 181, 1173–1180 (1991).
Suzuki, T. et al. Interleukin-6 enhances glucose-stimulated insulin secretion from pancreatic beta-cells: potential involvement of the PLC-IP3-dependent pathway. Diabetes 60, 537–547 (2011).
Miyakawa, T. et al. Encoding of Ca2+ signals by differential expression of IP3 receptor subtypes. EMBO J. 18, 1303–1308 (1999).
Morel, J. L., Fritz, N., Lavie, J. L. & Mironneau, J. Crucial role of type 2 inositol 1,4,5-trisphosphate receptors for acetylcholine-induced Ca2+ oscillations in vascular myocytes. Arterioscler. Thromb. Vasc. Biol. 23, 1567–1575 (2003).
Fritz, N., Mironneau, J., Macrez, N. & Morel, J. L. Acetylcholine-induced Ca2+ oscillations are modulated by a Ca2+ regulation of InsP3R2 in rat portal vein myocytes. Pflugers Arch. 456, 277–283 (2008).
Rizaner, N. et al. Intracellular calcium oscillations in strongly metastatic human breast and prostate cancer cells: control by voltage-gated sodium channel activity. Eur. Biophys. J. 45, 735–748 (2016).
Mak, D. O., McBride, S. & Foskett, J. K. Inositol 1,4,5-trisphosphate [correction of tris-phosphate] activation of inositol trisphosphate [correction of tris-phosphate] receptor Ca2+ channel by ligand tuning of Ca2+ inhibition. Proc. Natl. Acad. Sci. USA 95, 15821–15825 (1998).
Foskett, J. K., White, C., Cheung, K. H. & Mak, D. O. Inositol trisphosphate receptor Ca2+ release channels. Physiol. Rev. 87, 593–658 (2007).
Hattori, M. et al. Distinct roles of inositol 1,4,5-trisphosphate receptor types 1 and 3 in Ca2+ signaling. J. Biol. Chem. 279, 11967–11975 (2004).
Plaks, V., Kong, N. & Werb, Z. The cancer stem cell niche: how essential is the niche in regulating stemness of tumor cells? Cell Stem Cell 16, 225–238 (2015).
Smedler, E. & Uhlen, P. Frequency decoding of calcium oscillations. Biochim. Biophys. Acta 1840, 964–969 (2014).
Xu, X. et al. Recurrent hepatocellular carcinoma cells with stem cell-like properties: possible targets for immunotherapy. Cytotherapy 12, 190–200 (2010).
Zhou, P. et al. Beta-adrenergic signaling accelerates and synchronizes cardiac ryanodine receptor response to a single L-type Ca2+ channel. Proc. Natl.Acad. Sci. USA 106, 18028–18033 (2009).
Shui, B. et al. Circular permutation of red fluorescent proteins. PLoS ONE 6, e20505 (2011).
Xu, X. L. et al. The properties of tumor-initiating cells from a hepatocellular carcinoma patient's primary and recurrent tumor. Carcinogenesis 31, 167–174 (2010).
Zhang, S. J. et al. Evolutionary interrogation of human biology in well-annotated genomic framework of rhesus macaque. Mol. Biol. Evol. 31, 1309–1324 (2014).
Acknowledgements
We thank Dr. Guoqiang Bi for providing the plasmids harboring shRNAs, Dr. Fujian Lu for packaging GCaMP-ER2 adenovirus, and Drs. Lain C. Bruce, Ruiping Xiao, Xiuwu Bian, and Ning Lu for valuable comments. This work was supported by the National Key Basic Research Program of China (2016YFA0500403 and 2016YFA0500303), the National Science Foundation of China (81730075, 91529104, 31821091 and 81330051), and the National Institutes of Health (R24-HL-120847 and RO1-HL-120323). GCaMP-ER2 and associated mouse strains are available through the Cornell Heart Lung Blood Resource of Optigenetic Mouse Signaling (CHROMusTM—https://chromus.vet.cornell.edu).
Author information
Authors and Affiliations
Contributions
H.C. and Z.Z. conceived and supervised the research and C.S., Z.Z. and H.C. designed the research; C.S. performed the experiment with contributions from W.Z., H.L. and W.L.; B.S., J.C.L., B.D., F.K.L., S.R. and M.I.K. developed the GCaMP-ER2 sensor; T.S. and Q.S. contributed analytical tools; C.S., H.L., X.W., Z.Z. and H.C. analyzed the data; and C.S., H.C., Z.Z., and M.I.K. wrote the paper with contributions from all other authors.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by G. Raschell
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sun, C., Shui, B., Zhao, W. et al. Central role of IP3R2-mediated Ca2+ oscillation in self-renewal of liver cancer stem cells elucidated by high-signal ER sensor. Cell Death Dis 10, 396 (2019). https://doi.org/10.1038/s41419-019-1613-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-019-1613-2
This article is cited by
-
Phospholipid scrambling induced by an ion channel/metabolite transporter complex
Nature Communications (2024)
-
IP3R2-mediated Ca2+ release promotes LPS-induced cardiomyocyte pyroptosis via the activation of NLRP3/Caspase-1/GSDMD pathway
Cell Death Discovery (2024)
-
Calcium channel α2δ1 subunit is a functional marker and therapeutic target for tumor-initiating cells in non-small cell lung cancer
Cell Death & Disease (2021)