Abstract
The acquisition of radioresistance by breast cancer cells during radiotherapy may lead to cancer recurrence and poor survival. Signal transducer and activator of transcription 3 (Stat3) is activated in breast cancer cells and, therefore, may be an effective target for overcoming therapeutic resistance. Mesenchymal stem cells (MSCs) have been investigated for use in cancer treatment. Here, we investigated the potential of MSC conditioned medium (MSC-CM) in sensitizing breast cancer to radiotherapy. It was found that MSC-CM could inhibit the level of activated Stat3, suppress cancer growth, and exhibit synergetic effects with radiation treatment in vitro and in vivo. Furthermore, MSC-CM reduced the ALDH-positive cancer stem cells (CSCs) population, modulated several potential stem cell markers, and decreased tumor migration, as well as metastasis. These results demonstrate that MSC-CM suppresses breast cancer cells growth and sensitizes cancer cells to radiotherapy through inhibition of the Stat3 signaling pathway, thus, providing a novel strategy for breast cancer therapy by overcoming radioresistance.
Similar content being viewed by others
Introduction
Breast cancer is the most common malignancy and is the leading cause of cancer-related deaths in females worldwide1,2. Currently, the major clinical therapeutic methods for breast cancer include traditional surgical treatment, chemotherapy, and radiotherapy. Among them, radiotherapy is an important treatment modality to achieve local control and reduce the risk of recurrence. However, its curative effect is sometimes limited by radioresistance of cancer cells. Recently, the regulation of tumour radiosensitivity has attracted much attention, and identification of novel radiosensitizing agents that can increase the radiosensitivity of breast cancer has become an area of interest for radiation oncology investigators.
Several studies have shown that mesenchymal stem cells (MSCs) could be used to treat and enhance the radiosensitivity of cancer cells3,4. MSCs are multipotent cells that reside in various tissues and have the potential of multidirectional differentiation, which allows these cells to differentiate into multiple mesodermal cell lineages5,6,7,8. MSCs have been isolated from many different tissues, including bone marrow, adipose tissue, umbilical cord blood, peripheral blood, and skeletal muscle9,10 and are a promising source for cell therapy in regenerative medicine. While several studies have demonstrated that MSCs contribute to tumour progression and metastasis11,12, other reports have shown that MSCs could suppress tumour growth13,14. The different effects of MSCs on tumour growth depend on a variety of factors, including the type and origin of MSCs, the tumour models, and the dose and time of administration of cell treatments15. Therefore, it is necessary to explore the potential mechanisms of MSC-induced tumour inhibitory effects in breast cancer cells.
Signal transducer and activator of transcription 3 (Stat3) played a vital role in tumourigenesis16,17,18. An early research of human breast cancer cell lines demonstrated that Stat3 was activated in five of the nine cell lines19,20. Stat3 activation is found in all classes of breast cancers, but is most often associated with triple negative breast tumors. The Stat3 signaling pathway was recently reported to contribute to tumour progression and the survival of breast cancer-derived stem cells. Some studies have shown that the Stat3 signaling pathway is required for growth of CD44+CD24– stem cell–like breast cancer cells21, such as several basal-like breast cancer cells (MDA-MB-231, BT-549, HCC1937, Hs 578T, MDA-MB-468, and SUM159PT ), not expressed in luminal breast cancer cell lines (BT-474, MCF7, MDA-MB-453, SK-BR-3, T-47D, and ZR-75-1)22. However, whether the tumour inhibitory effect of MSCs is mediated by the Stat3 signaling pathway is unclear.
In this study, we used MSC-conditioned medium (MSC-CM) combined with radiation treatment and an imaging approach to explore how the aggressive breast cancer cells (MDA-MB-231) respond to the combination treatment and to investigate the possible underlying mechanisms. Our results indicated that MSC-CM reduces the growth of MDA-MB-231 cells and sensitises the cancer cells to radiation therapy through inhibition of Stat3 activation. This work identifies Stat3 as a potential therapeutic target that may radiosensitise cells prior to conventional radiation therapy and provides a basis for the clinical application of radiation combined with MSC therapy, thus suggesting a more effective treatment for breast cancer patients.
Results
Construction of optical imaging tumour cells
To evaluate the effect of the MSCs on cancer cells and track the transplanted cancer cells in vivo using imaging analysis, we constructed double imaging MDA-MB-231 cells (Fluc/GFP-pStat3/Rluc) with Fluc and eGFP reporter genes drived by a ubiquitin promoter, Rluc reporter gene drived by a seven-repeat Stat3-binding sequence (enhancer) and minimal TA (promoter) in response to the activated Stat3. The fluorescence images showed that the expression of eGFP was robust in MDA-MB-231 cells (Fig. 1a). FACS analysis indicated that GFP was expressed in >95% of cells after sorting (data not shown). A strong correlation (r2 = 0.9976) between the cell number and firefly signal intensity was observed in vitro using the Xenogen IVIS system, which quantified tumour cells by analysing firefly signal intensity (Fig. 1b). The Rluc expression was controlled by Stat3 activation. Once activated, the phosphorylated Stat3 underwent dimerization and entered the nucleus to bind the seven-repeat response elements inducing the expression of Rluc. When the cells were administrated with coelenterazine, photon signals could be detected. The signaling intensity of Rluc therefore indicated the activation of Stat3 signaling in cancer cells.
Tumour suppressor activity of MSC-CM and its combination with radiotherapy
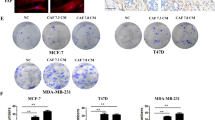
To investigate the effect of MSCs on breast cancer cells, we treated MDA-MB-231 cells with MSC-CM firstly. The cell proliferation and survival were decreased (Figure S1). Then we treated MDA-MB-231 cells with MSC-CM, radiation, and MSC-CM in combination with radiation to examine the effect of MSC-CM on tumour radiosensitivity. MDA-MB-231 cells in the MSC-CM-treated and combination groups lost their normal spindle shape and became thinner and longer after 48 h of treatment, the number of cells was sharply reduced (Fig. 2a). Furthermore, the proliferation rate of tumour cells in the MSC-CM-treated and combination groups grew slowly compared with the control groups. As shown in Fig. 2b, MSC-CM could inhibit the growth of tumour cells, however, there was no obvious difference between the control and the only radiation group. The imaging assay showed decreased Fluc activity in cells treated with MSC-CM and the combination groups (Fig. 2d). Quantitative analysis demonstrated that the Fluc signal intensity of tumour cells in the combination group was less than the control groups (Fig. 2e). The cell survival assay was performed using Trypan blue staining, which showed a decrease in the MSC-CM and combination groups (Fig. 2c). To determine the effect of MSCs on tumour cells radioresistance in vitro, we performed colony formation assays. A clonogenic survival assay showed that MSC-CM increased cell radiosensitivity compared with that of the control group (Fig. 2f).
(a) Microscopic analysis showed morphological changes in cells treated with MSC-CM, 4 Gy radiation and MSC-CM combined with 4 Gy radiation. The scale bar represents 100 μm. (b) Growth curves made by counting cell numbers every day revealed a decreased proliferation rate of cells treated by MSC-CM and the MSC-CM combination with 4 Gy radiation. *P < 0.05 vs Ctrl, **P < 0.01 vs Ctrl, ***P < 0.001 vs Ctrl, #P < 0.05 vs MSC-CM, ##P < 0.01 vs MSC-CM, $$P < 0.01 vs 4 Gy, n = 3. (c) Trypan blue cell viability assays showed a low survival rate. **P < 0.01 compared to controls, n = 3. (d) Fluc imaging showed decreasing bioluminescence signals at 24, 48, and 72 h in the MSC-CM-treated and combination groups. (e) Quantitative analysis of Fluc imaging signals. The signal activity showed the suppressed growth in the combination group. *P < 0.05 vs Ctrl, #P < 0.05 vs MSC-CM, ##P < 0.01 vs MSC-CM, $$P < 0.01 vs 4 Gy, n = 3. (f) MDA-MB-231 cells treated with control medium and MSC-CM were exposed to ionizing radiation. Cell growth was analysed by colony formation assays as described in the Materials and Methods. Reported values are the mean±SD for three separate experiments
MSC-CM promotes apoptosis and inhibits the cell cycle of breast cancer cells
Irradiation primarily leads to the production of double-stranded DNA breaks (DSBs), which result in cell apoptosis. Radiation leads to cell apoptosis, which could be understood as radiosensitivity23,24. Here, we evaluated the apoptosis of tumour cells affected by MSC-CM. Flow cytometry apoptosis assays showed that MSC-CM increased apoptosis of tumour cells compared with that of the control groups, especially in cells exposed to 8 Gy radiation (Fig. 3a). Previous studies showed that most radioresistant cells were arrested in S phase, with fewer cells in G2-M phase, after irradiation compared with radiosensitive cells25. Accordingly, we further studied the cell cycle changes in response to radiation and MSC-CM using flow cytometry. As shown in Fig. 3b, no difference was observed in the G0-G1, S and G2-M phases in the control, CM or radiation only groups; however, compared with these groups, the combination group had fewer cells in S phase and more cells in G2-M phase, suggesting that combined treatment of MSC-CM with radiation could lead to the cell cycle block at G2-M phase, which is consistent with the typical radiosensitive phenotype. Taken together, these results demonstrated that MSC-CM could sensitise MDA-MB-231 cells to irradiation in vitro.
(a) MDA-MB-231 cells were treated with control medium and MSC-CM and then exposed to 8 Gy ionizing irradiation. The apoptosis of cells was assessed by FACS after 48 h of irradiation. (b) The cells were treated with control medium and MSC-CM and then at 24 h after exposed to 2 Gy or 4 Gy radiation, stained with PI and analysed using FACS. The proportion of DNA in the different cell phases was analysed. Data are shown as the mean±SD (error bars) from three independent experiments. *P < 0.05 vs Ctrl, **P < 0.01 vs Ctrl, #P < 0.05 vs MSC-CM, ##P < 0.01 vs MSC-CM, $$P < 0.01 vs 8 Gy, n = 3
MSC-CM inhibits the Stat3 pathway in breast cancer cells
Recent studies demonstrated that Stat3 plays critical roles in the initiation and progression of breast cancer19. The most common cause of disease relapse and radioresistance in breast cancer is the presence of stem cell-like cells (or CSCs) in tumour tissues26. Multiple signaling pathways, including the Stat3 signaling pathway, help maintain stem cell programmes in normal cells as well as in cancer cells27,28,29.
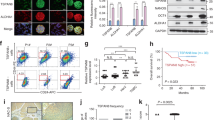
To explore the activation of the Stat3 signaling pathway in MDA-MB-231 cells in different groups, we performed Rluc imaging, which showed the activated Stat3 level. After 48 h and 72 h treated with MSC-CM, the Rluc activity was significantly decreased in the MSC-CM and the combination groups as shown by the Renilla signal intensity (Fig. 4a, b). Stat3 phosphorylated at Try-705 was substantially decreased in cells treated with MSC-CM and MSC-CM combined with radiation groups as shown by western blot and cell-based ELISA (Fig. 4c, d). Then, we studied the expression of stemness genes (Sox2, Oct4, Nanog, and c-Myc) and Stat3 signaling pathway-related genes (cyclin D1, Bcl-xl, p53) affected by MSC-CM; these genes were substantially down-regulated in the combination group compared with the control groups (Fig. 4g). These results indicated that MSC-CM suppressed Stat3 signaling activation. As MSC-CM exhibited an inhibitory effect on Stat3 activation, we further examined its anti-CSC properties. The CSCs in breast cancer cells were assessed using ALDEFLUOR and Mammosphere formation assays. ALDEFLUOR-positive cells (Fig. 4e, f) and the mammosphere formation efficiency (Figure S3) were reduced after treatment with MSC-CM for 48 h, especially when combined with the irradiation treatment.
(a) Renilla luciferase (Rluc) imaging of activated Stat3 in vitro. (b) Quantitative analysis of imaging signals. The signal activity showed the suppression in the combination group. *P < 0.05 vs Ctrl, **P < 0.01 vs Ctrl, n = 3. (c) The expression of Stat3, phosphorylated (Try-705) Stat3 measured with Western blotting. (d) Quantification of Stat3 phosphorylation; the phospho-Try705 Stat3 and total Stat3 protein levels in cells were quantitatively determined using a target specific primary antibody and HRP-conjugated secondary antibody detection agent, and the crystal violet provided whole cell staining that was used for cell number counts. The OD value was normalized by cell counts. *P < 0.05 vs Ctrl, n = 3. (e) Aldehyde dehydrogenase (ALDH) activities in MDA-MB-231 cells were assessed utilizing the ALDEFLUOR assay. Cells were incubated with 1.5 μl of activated ALDEFLUOR substrate at 1 × 106 cells per ml. Diethylaminobenzaldehyde (DEAB) was used to establish the baseline fluorescence of these cells and to define the ALDEFLUOR-positive region. Incubation of cells with ALDEFLUOR substrate in the absence of DEAB induces a shift in BAAA fluorescence, defining the ALDEFLUOR-positive population. (f) Quantitative analysis of ALDEFLUOR-positive cells. The number of ALDEFLUOR-positive cells was reduced after treatment with MSC-CM, especially when combined with the irradiation treatment. *P < 0.05 vs Ctrl, $P < 0.05 vs 4 Gy, n = 3. (g) PCR analysis of Stat3 signalling pathway-related gene expression. *P < 0.05 vs Ctrl, **P < 0.01 vs Ctrl, ***P < 0.001 vs Ctrl, #P < 0.05 vs MSC-CM, ##P < 0.01 vs MSC-CM, $P < 0.05 vs 4 Gy, $$P < 0.01 vs 4 Gy, n = 3
To further detect MSC-CM could sensitize cancer cells to radiotherapy via inhibiting Stat3 signaling pathway, we used Stat3 inhibitor Stattic to inhibit the Stat3 signaling pathway and then analyzed the radiosensitivity of cancer cells affected by MSC-CM. The cells survival, epithelial-mesenchymal transition, metastasis, and angiogenesis were inhibited in MSC-CM combination with radiation group after treated with 5 uM Stattic (Figure S4).
Effects of MSC-CM on breast cancer cells migration, metastasis, and angiogenesis
Epithelial-to-mesenchymal transition (EMT) plays an important role in metastasis and invasiveness of cancer cells and is associated with poor clinical outcome in lots of cancers30,31,32. EMT is an important process involved in progression of cancers, and previous studies have demonstrated that Stat3 inhibition leads to EMT characterised by up-regulation of epithelial cell-specific proteins (E-cadherin), and by down-regulation of the mesenchymal cell-specific protein N-cadherin33. The Stat3 pathway is tightly associated with tumour invasion, metastasis, and angiogenesis in cancers34,35,36.
Our data indicated that MDA-MB-231 cells treated with MSC-CM and radiation exhibited substantial inhibition of cell proliferation and Stat3 activation. Then, we examined the effects of MSC-CM on the migration, metastasis, and angiogenesis of cancer cells. Wound scratch assays were performed to test cell migration in different groups. After 48 h of MSC-CM, radiation or a combination of CM and radiation treatment, a significant difference was observed between the combination treatment and control groups. The results indicated that MSC-CM impaired tumour cells migration ability (Fig. 5a, b). As tumour cell migration is closely associated with EMT37, then we examined epithelial and mesenchymal biomarker expression. The RT-PCR results showed that treatment with CM and radiation decreased Vimentin and N-cadherin expression (mesenchymal markers) and increased E-cadherin expression (epithelial markers). Activated Stat3 regulates tumour invasion by regulating the gene transcription of matrix metalloproteinase 2 (MMP-2), MMP-9, TGF-β1 and β-catenin, which also decreased after treatment with MSC-CM (Fig. 5d). These results indicated that the inhibitory effect of MSC-CM in combination with radiotherapy on the tumour metastasis is probably through the EMT pathway.
(a) Representative photographs of wound-healing assays of MDA-MB-231 cells treated with MSC-CM and 4 Gy irradiation. The scale bar represents 100 μm. (b) Wound healing area per field was decreased after treatment with MSC-CM and MSC-CM+4 Gy. *P < 0.05 vs Ctrl, **P < 0.01 vs Ctrl, ***P < 0.001vs Ctrl, #P < 0.05 vs MSC-CM, $P < 0.05 vs 4 Gy, $$P < 0.01 vs 4 Gy, n = 3. (c) Analysis of angiogenic factor expression in MDA-MB-231 cells in different groups. *P < 0.05 vs Ctrl, **P < 0.01 vs Ctrl, #P < 0.05 vs MSC-CM, $P < 0.05 vs 4 Gy, n = 3. (d) Expression of metastasis and EMT-related genes was determined by RT-PCR. All experiments were performed at least in triplicate, and the data are presented as the mean±SD. *P < 0.05 vs Ctrl, #P < 0.05 vs MSC-CM, $P < 0.05 vs 4 Gy, $$P < 0.01 vs 4 Gy, n = 3. See also Table S1
Phosphorylated Stat3 was shown to bind to HIF-1α directly, and Stat3 has also been shown to enhance HIF-1α-mediated expression of VEGF38. Therefore, we examined tumour angiogenesis-related gene expression. The results indicated that MSC-CM maybe inhibit angiogenesis (Fig. 5c). All these data showed that the MSC-CM combination with radiotherapy could inhibit the migration, metastasis, and angiogenesis of breast cancer cells.
DNA repair is inhibited by decreased Stat3
We demonstrated that cells treated with MSC-CM exhibited strong down-regulation of Stat3 expression. As the Stat3 pathway has been shown to play an important role in the radiotherapy of cancers39 primarily through regulation of the radiation-induced DSBs40, we tested the possible impact of MSC-CM treatment on the regulation of DNA damage. Several factors involved in repair of DSBs were assessed, including BRCA1 (breast cancer 1, early onset), which is a well-known tumour suppressor with critical roles in DNA repair41; 53BP1, which is an important regulator of DSB repair protecting broken DNA ends from processing, a mechanism that is induced by BRCA142,43; and ATM, which is the master regulator of the DNA damage response44. Our data showed that the protein expression levels of p-BRCA1, 53BP1, and p-ATM were increased after treatment with MSC-CM, especially in the combination group (Fig. 6). We assessed γH2AX and 53BP1 foci as indicators of DNA DSBs by immunostaining (Figure S2). According to regular statistical standard for analyzing DNA damage repair protein foci45,46, cells with >10 foci were counted as positive cells. As expected, dsDNA damage was significantly increased in the combination group, and this damage persisted significantly longer (24 h) compared with that of the control group. The effect on regulators of cell cycle arrest was also evaluated in our current study by assessing the two transducers of CHK1 (checkpoint kinase-1) in the ATM pathway47 and the central stress protein p53 in the DNA damage response48,49. Our results showed that the protein levels of p-p53 and p-CHK1 were increased in the combination groups. These results indicated that MSC-CM enhanced DNA damage after radiation, which increased the radiosensitivity of tumour cells.
MSC-CM and the MSC-CM combination with radiation increased p-BRCA1, 53BP1, p-ATM, p-p53 and p-CHK1expression. Cells were treated with MSC-CM or treated with ionizing radiation. Six hours after ionizing radiation, cells were harvested, and the cell lysates were analysed by immunoblotting using the indicated antibodies. *P < 0.05 vs Ctrl, **P < 0.01 vs Ctrl, ***P < 0.001 vs Ctrl, #P < 0.05 vs MSC-CM, ##P < 0.01 vs MSC-CM, ###P < 0.001 vs MSC-CM, $$ P < 0.01 vs 4 Gy, $$$P < 0.001 vs 4 Gy, n = 3
The MSC-CM combination with radiotherapy inhibits Stat3 signaling activation in MDA-MB-231 cells in vivo
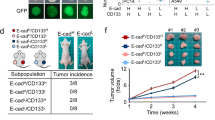
To determine whether the pre-treatment of MSC-CM combination with radiotherapy on cells could inhibit Stat3 activation in vivo, we subcutaneously implanted equal numbers of MDA-MB-231 cells (Fluc/GFP-pStat3/Rluc) that were treated with MSC-CM or control medium into the upper and lower right region of Nu/Nu Nude mice. Tumour cells pre-treated with MSC-CM combined with 4 Gy radiation or radiation alone were injected into the upper and lower right region of the mouse body. Bioluminescence imaging of Rluc was performed to assess Stat3 activation in the tumours. The imaging results demonstrated that the MSC-CM combination with radiotherapy pre-treatment could significantly inhibited the Stat3 signaling pathway in vivo (Fig. 7a, c).
(a) Renilla luciferase (Rluc) imaging of p-Stat3. Representative animals were injected with 2 × 106 MDA-MB-231 (Fluc/GFP-pStat3/Rluc) cells into the upper and lower right side of the mouse body after treatment with MSC-CM and control medium for 48 h. Quantitative analysis of Rluc signals indicated decreased p-Stat3 level. *P < 0.05 vs Ctrl, n = 6. (b) Firefly luciferase (Fluc) imaging of tumour growth. Representative animals injected with 2 × 106 MDA-MB-231 (Fluc/GFP-pStat3/Rluc) cells into the upper and lower right of the mouse body after treatment with MSC-CM and control medium for 48 h. Quantitative analysis of the Fluc signal. **P < 0.01 vs Ctrl, n = 6. (c) Renilla luciferase (Rluc) imaging of p-Stat3. Representative animals injected with 2 × 106 MDA-MB-231 (Fluc/GFP-pStat3/Rluc) cells into the upper and lower right side of the mouse body after treatment with MSC-CM combination with radiation or radiation alone. Quantitative analysis of Rluc signal. *P < 0.05 vs 4 Gy, n = 6. (d) Firefly luciferase (Fluc) imaging of tumour growth. Representative animals injected with 2 × 106 MDA-MB-231 (Fluc/GFP-pStat3/Rluc) cells into the upper and lower right side of the mouse body after treatment with MSC-CM combination with radiation or radiation alone. Quantitative analysis of Fluc signal. *P < 0.05 vs 4 Gy, n = 6
To investigate the inhibition effect of MSC-CM combination with radiotherapy on Stat3 signaling pathway, we also performed another group of animal experiment in which cells were injected subcutaneously into the right hind leg of animal first, and then the effect of radiotherapy to the animal tumour with or without MSC-CM treatment for the animal was observed. The Rluc activity increased more slowly in the MSC-CM combination with radiotherapy group than the radiotherapy group (Fig. 8a, c).
(a) Renilla luciferase (Rluc) imaging of p-Stat3. Representative animals were injected with 2 × 106 MDA-MB-231 (Fluc/GFP-pStat3/Rluc) cells into the right hind leg first, and then received radiotherapy to the animal tumour with or without MSC-CM treatment. (b) Firefly luciferase (Fluc) imaging of tumour growth. Representative animals injected with 2 × 106 MDA-MB-231 (Fluc/GFP-pStat3/Rluc) cells into the right hind leg first, and then received radiotherapy to the animal tumour with or without MSC-CM treatment. (c) Quantitative analysis of Rluc signals indicated decreased p-Stat3 level. *P < 0.05 vs IR, **P < 0.01 vs IR, n = 6. (d) Quantitative analysis of the Fluc signal. *P < 0.05 vs IR, **P < 0.01 vs IR, n = 6. (e) Proposed model for tumour-suppressive and radiosensitivity effects of the MSC-CM combination with irradiation. Some factors secreted by MSC cells could efficiently inhibit Stat3 activation in MDA-MB-231 cells. The Stat3 signalling pathway regulates the expression of c-Myc, Sox2, MMPs, VEGF, and E-cadherin and then is involved in cancer cell growth, stemness, invasion, and angiogenesis
MSC-CM combined with radiotherapy significantly delays xenograft tumour growth
To confirm whether the down-regulation of Stat3 leads to tumour growth inhibition, we performed Fluc imaging to detect the tumour growth in vivo. Cells were pre-treated with MSC-CM, 4Gy radiation or MSC-CM combination with 4Gy radiation before injecting subcutaneously into the mice. From the data, we discovered that the signal intensity of cells treated with CM combined with radiation increased more slowly than the cells in the control group (Fig. 7b, d). These data demonstrated that MSC-CM weakened the tumour formation ability and inhibited tumour growth by repressing the Stat3 signaling pathway.
To further explore the inhibition effect of MSC-CM combination with radiotherapy on tumour growth, we injected cells subcutaneously into the right hind leg of animal first, and then observed the effect of radiotherapy to the animal tumour with or without MSC-CM treatment through Fluc imaging. The results demonstrated that the tumours grew more slowly in MSC-CM combination with radiotherapy group than the only radiotherapy group (Fig. 8b, d).
Discussion
Radiotherapy is a critical therapeutic method for breast cancer. However, its therapeutic effect depends on the radioresistance of cancer cells. In this study, we report several meaningful findings. First, inhibitory growth of MDA-MB-231 breast cancer cells treated by MSC-CM was observed. Second, MSCs could enhance the therapeutic effect of radiotherapy by inhibiting the Stat3 signal pathway. To our knowledge, no other reports have indicated the molecular mechanisms underlying MSC-mediated radiosensitivity in MDA-MB-231 cells. In addition, given that MSCs have not only been investigated for the treatment of cancers50,51 but also for the management of cancer radiosensitivity3, it is noteworthy that we uncovered the interplay between the factors secreted by MSCs and the Stat3 pathway which influences cellular radiosensitivity, and this will help develop novel Stat3-targeting therapies.
Some reports have shown the MSC-mediated suppression of cancer cell proliferation4,52,53, indicating that MSCs inhibit the growth and promote the apoptosis of cancer cells. In addition, one report on breast cancer cells further showed that MSCs significantly down-regulated the expression of VEGF in tumour cells, which led to inhibition of angiogenesis in vitro and in vivo. Moreover, other studies showed that MSCs enhanced the radiotherapy effect on cancers likely through inhibition of tumour cell proliferation and enhancement cancer cell apoptosis3. Thus, inhibitory cell growth, low colony-forming efficiency and cell apoptosis were observed in our data (Fig. 2). However, the molecular mechanism of MSC-induced tumour suppression remains poorly understood. In the current study, we demonstrated that MSC-CM could inhibit MDA-MB-231 cell proliferation, migration, and angiogenesis and promote DNA damage, likely by inhibiting the Stat3 pathway as shown by bioluminescence imaging technology.
Stat3 is a member of the Stat protein family, in response to cytokines and growth factors. Stat3 becomes activated after phosphorylation of tyrosine residue (Y705)54. Recent studies suggest that Stat3 plays a central role in development of cancer radioresistance55,56. Stat3 has been shown to enhance expression of regulatory factors related to cancer cell proliferation, metastasis, angiogenesis, and resistance to apoptosis-related genes57. Therefore, inhibition of Stat3 may be a promising therapeutic strategy for the management of cancer radioresistance58,59. Indeed, as shown in Fig. 4a, c in our study, the phosphorylated Stat3 expression levels were decreased in the combination treatment groups. These results suggest that radiosensitivity of MDA-MB-231 cells can be regulated by the Stat3 pathway, a hypothesis supported by evidence demonstrating that inhibition of the Stat3 pathway sensitises cancer cells to radiation56,60,61,62.
Regarding the interplay between the radiosensitivity effect of MSCs on cancer cells and the inhibition of the Stat3 signaling pathway, to our knowledge, no report has demonstrated that MSC-CM inactivates the Stat3 pathway after radiation in MDA-MB-231 cells. To test our hypothesis, in addition to evaluating the Stat3 activation via bioluminescence imaging and cell-based ELISA, we also determined ALDH expression and the Mammosphere formation ability which are widely accepted to identify the CSCs. Numerous studies have found that Stat3 activation is involved in regulating radioresistance of cancers, which could be due to its central roles in CSC maintenance61,63,64. CSCs were reported to induce tumour growth, radioresistance, metastasis and angiogenesis in breast cancer65. Therefore, we found that the migration, invasion and angiogenesis abilities of tumour cells were weakened in combination groups (Fig. 5a–d).
Findings of this study indicate that suppressing the Stat3 signaling pathway combined with traditional radiotherapy could be a promising novel approach for overcoming radioresistance in MDA-MB-231 breast cancer cells. However, an inherent limitation of our study is that the factors in MSC-CM mediating the inactivation of Stat3 were unclear. A further study that specifically elucidates how MSC-CM results in the inhibition of Stat3 and thus regulates the radiosensitivity of cancer cells will be performed in the future.
In conclusion, we demonstrated that MSC-CM is a potent tumour suppressor of MDA-MB-231 cells by inhibiting Stat3 activation and down-regulating its downstream gene expression. Our data indicate that Stat3 could be a potential target and MSCs could provide a basis for a promising therapeutic strategy for radiotherapy of breast cancer.
Materials and methods
Reagents
Heat-inactivated fetal bovine serum (FBS) and cell culture medium DMEM, DMEM/F12 were purchased from Gibco (Invitrogen China Limited, China). A real-time PCR (RT-PCR) kit was purchased from TransGen Biotech (Beijing, China). D-Luciferin was purchased from Biosynth International (Naperville, USA). Coelenterazine was purchased from NanoLight Technology (Pinetop, AZ, USA). Antibodies against p-BRCA1 (Ser1524), 53BP1, p-ATM (Ser1981), p-p53 (Ser15), p53, p-CHK1 (Ser317), p-Stat3 (Tyr705), Stat3 were purchased from Abcam (Cambridge, MA, USA). Antibodies against β-actin, β-tubulin, and GADPH were obtained from ZSGB-Bio (China).
Cell culture
The human breast cancer cell line MDA-MB-231 was purchased from ATCC (Manassas, VA, USA), and human umbilical cord MSCs (hUC-MSCs) were isolated and cultured as described previously66. MDA-MB-231 cells were grown in DMEM medium supplemented with 10% foetal bovine serum (FBS), 1% penicillin streptomycin solution (Gibco, Rockville, MD), and 1% MEM non-essential amino acid solution (Gibco). The culture medium of hUC-MSCs was DMEM/F12 (Gibco), 10% FBS, penicillin (100 units/ml), streptomycin (100 mg/ml)(Gibco). For imaging, MDA-MB-231 cells were conducted with double imaging reporter genes. The reporter construct carried firefly luciferase (Fluc) and enhanced green fluorescence protein (eGFP) drived by an ubiquitin promoter, Renilla luciferase (Rluc) reporter gene drived by a 7-repeat of Stat3-recognition sites (enhancer) and a small TA promoter in response to phosphorylated Stat3.
Collection of conditioned medium
The hUC-MSC cells and MDA-MB-231 cells (purchased from ATCC) were cultured to 40% confluence in the previously described culture media, and the medium for hUC-MSCs cells was replaced with 7 ml DMEM/F12 medium. The medium for MDA-MB-231 cells was replaced with 7 ml DMEM. After 2 days, the supernatants were collected and stored at –80 °C.
Morphological observation
2.5 × 105 of MDA-MB-231 cells were seeded into six-well plate. The medium was replaced with hUC-MSC-CM and 231-CM mixed with the same amount of DMEM medium (containing 10% FBS) on the next day.
Cell proliferation and viability assays
MDA-MB-231 cells (3 × 104) were seeded in 12-well plates. The medium was replaced with different CMs and mixed with DMEM medium (containing 10% FBS) on the next day. Cells in plate were counted in three days. Cell viability was determined by Trypan blue dye exclusion assays.
Bioluminescence imaging
Bioluminescence imaging of the fate of transplanted cells in living mice was performed for 30 days using an IVIS cooled CCD optical Imaging System (Xenogen Corporation, Hopkinton, MA). Fluc imaging of MDA-MB-231 cells was used to assess tumour progression, and the expression of phosphorylated-Stat3 was determined by using Rluc imaging. Fluc imaging was performed with D-Luciferin (150 mg/kg; Biosynth International, Naperville, IL, USA), the reactive substrate that was intraperitoneally injected into mice. Coelenterazine (2.5 mg/kg; NanoLight Technology, Pinetop, AZ, USA) was used to evaluate Rluc expression51,67.
Cell irradiation
A Cs-137 (Gammacell-40) irradiator was purchased from Atomic Energy Co. (Atomic Energy of Canadian Inc., Mississauga, ON, Canada). Cell or animal samples were placed in the centre of the irradiation chamber and exposed to the radiation at a dose rate of 1.02 Gy/min.
Real-time PCR
To assess the mRNA expression levels of the genes, total RNA was extracted from MDA-MB-231 cells treated with TRIzol reagent according to instructions. 2 μg of total RNA was reverse transcribed using a Reverse Transcription Kit (TIANGEN Biotech), and real-time PCR was performed with the Opticon® System (Bio-Rad) using a TransStart Green qPCR Super Mix Kit (TransGen Biotech). The 2–ΔΔct method was used to assess the relative mRNA expression.Primers are listed Supplemental Table 1.
Western blot analysis
To determine the protein expression, western blotting was performed. MDA-MB-231 cells were lysed on ice in lysis buffer (Cwbiotech, Beijing, China) including protease inhibitor cocktail. The total protein was quantified with BCA Protein Assay Kit as described previously [21]. Antibodies, anti-p-BRCA1 (Ser1524), anti-53BP1, anti-p-ATM(Ser1981), anti-p-p53(Ser15), anti-p53, anti-p-CHK1(Ser317), anti-p-Stat3(Tyr 705), anti- Stat3, anti-β-actin, anti-β-tubulin and anti-GADPH.
Cell-based ELISA
The activated Stat3 expression in MDA-MB-231 cells was detected by Cell-Based Colorimetric ELISA Kit (ImmunoWay, USA). After treatment with formaldehyde, cells were incubated with phospho-Stat3 (Try705) and total Stat3 antibodies and then the secondary HRP-conjugated antibody. The Stat3 phosphorylation levels were normalised by both the levels of total Stat3 protein and total cell number in each well68.
Flow cytometric apoptosis assay
Apoptosis was measured by fluorescence-activated cell sorting (FACS) using the Annexin V-FITC Apoptosis Detection Kit. In brief, cells plated in 6-well plates were treated with MSC-CM for 48 h or in combination with radiotherapy. After 48 h of radiation, the cells were harvested for cell apoptosis assay and analysed by flow cytometry.
Colony formation assay
MDA-MB-231 cells were trypsinised, suspended in a single-cell suspension and plated into 6-well plates (800 cells/well). Then the next day, the medium in control group was replaced with 231-CM and DMEM (including 10% FBS), and the experimental group was MSC-CM and DMEM (including 10% FBS) at a ratio of 1:1. Cells were treated with the indicated doses of 137Cs–radiation (1.02 Gy/min). After 10 days culture, the cells were treated with crystal violet. Colonies containing 50 cells or more were counted as described previously69,70.
Cell-cycle analysis
After treatment with MSC-CM or a combination with radiotherapy, cells were harvested and washed with PBS. Then cells were treated with methanol and resuspended in PI-staining buffer (50 μl/ml PI, RNase A) for 15 min at 37 °C Fluorescence intensity was analyzed by flow cytometer.
Wound-healing assay
MDA-MB-231 cells (2 × 105) were plated in six-well plate. Then the medium was replaced with control medium or a mixture of MSC-CM with the same amount of DMEM (supplemented with 10% FBS). Then, cells were exposed to 4 Gy radiation. For scratch wound healing assay, cells were cultured in serum-free medium for 24 h and three separate wounds were made with sterile pipette tips (Corning). The percentage of wound closing was assessed using the following formula:
[1–(final area/initial area)] × 100%.
Immunofluorescence staining
A total of 1 × 104 cells were plated into each well of 12-well plates. After treatment with control medium or MSC-CM for 48 h, cells were exposed to 4 Gy radiation. The cells in four groups (Ctrl, CM, 4Gy, 4Gy+CM) were fixed with 4% paraformaldehyde (Sigma) for 15 min. Cells were first incubated with primary antibody, mouse anti-phospho-Histone H2AX (Ser139), rabbit anti-53BP1, followed by incubation with Cy3 goat anti-mouse IgG and goat anti-rabbit secondary antibodies. And then counter stained with DAPI. Images were acquired using an AMG EVOS microscope.
Mammosphere assay
Cells were grown in serum-free, growth factor-enriched conditions in low-attachment plates to generate mammospheres71,72,73. MDA-MB-231 cells were treated with control and MSC-CM for 48 h and then exposed to 4 Gy radiation. Then the cells were seeded into ultralow-attachment plates and cultured in serum-free DMEM medium containing basic fibroblast growth factor (bFGF), human epidermal growth factor (hEGF), and B27 (Invitrogen). The number of formed mammospheres was determined under an inverted microscope.
Aldefluor-positivity assay
The Aldefluor kit (Stem Cell Technologies) was used to assess ALDH activity with flow cytometry according to previous report71. Cells were suspended in ALDEFLUOR assay buffer containing ALDH substrate (Bodipy-Aminoacetaldehyde) and incubated at 37 °C for 45 min. As a negative control, 15 μM diethylaminobenzaldehyde (DEAB) was used. The fluorescent ALDH1-expressing cells were analyzed with flow cytometer.
In vivo xenograft studies
A group of 8–12-week-old female Nu/Nu nude mice were housed under standard laboratory conditions. A group of mice (n = 6 mice/group) were injected with 2 × 106 MDA-MB-231 (Fluc/GFP-pStat3/Rluc) cells into the upper and lower right region of the mouse body (day 0) after treatment with MSC-CM or control medium for 48 h36. Equal numbers of MDA-MB-231 (Fluc/GFP-pStat3/Rluc) cells after treatment with MSC-CM combined with radiation or radiation alone were injected into the upper and lower right of mice body to detect the tumour growth and the activated Stat3 expression level by imaging.
Another group of mice were injected with 2 × 106 MDA-MB-231 (Fluc/GFP-pStat3/Rluc) cells into the right hind leg of animal first (day 0), five days later, when the tumors were palpable, the mice were randomly divided into two groups (6 mice per group): a radiation group, which received γ-radiation radiotherapy of tumors in the legs twice weekly (2 Gy) for 2 weeks, and a combination group (CM + radiotherapy) treated with MSC-CM every two days for 2 weeks. For radiation treatment, mice were first anaesthetized and then placed onto a specially designed lead plate so as to radiate locally the tumors. The activated Stat3 expression level and tumour growth were detected by Rluc and Fluc imaging. All experimental procedures and protocols were conducted according to the guidelines of our local animal care and use committee.
Statistical analysis
All data were analyzed using GraphPad Prism 5.0 software. Data were analyzed by two-tailed Student’s t test or Two-way ANOVA. A P value less than 0.05 was considered statistically significant. Data were shown of three independent experiments.
Change history
20 January 2022
Editor's Note: Concerns have been raised about the integrity of the data reported in this article. This is currently being investigated. Further editorial action may be taken as appropriate once the investigation into the concerns is complete and all parties have been given an opportunity to respond in full.
References
Nelson, H. D. et al. Risk factors for breast cancer for women aged 40 to 49 years: a systematic review and meta-analysis. Ann. Intern. Med. 156, 635–648 (2012).
Jemal, A. et al. Global cancer statistics. CA Cancer J. Clin. 61, 69–90 (2011).
de Araujo Farias, V. et al. Human mesenchymal stem cells enhance the systemic effects of radiotherapy. Oncotarget 6, 31164–31180 (2015).
Clarke, M. R., Imhoff, F. M. & Baird, S. K. Mesenchymal stem cells inhibit breast cancer cell migration and invasion through secretion of tissue inhibitor of metalloproteinase-1 and -2. Mol. Carcinog. 54, 1214–1219 (2015).
Keating, A. Mesenchymal stromal cells: new directions. Cell. Stem. Cell. 10, 709–716 (2012).
Feisst, V., Brooks, A. E., Chen, C. J. & Dunbar, P. R. Characterization of mesenchymal progenitor cell populations directly derived from human dermis. Stem Cells Dev. 23, 631–642 (2014).
Tobita, M. & H. Mizuno Adipose-derived stem cells and periodontal tissue engineering. Int. J. Oral Maxillofac. Implants 28 (2013).
Pittenger, M. F. et al. Multilineage potential of adult human mesenchymal stem cells. Science 284, 143–147 (1999).
Campagnoli, C. et al. Identification of mesenchymal stem/progenitor cells in human first-trimester fetal blood, liver, and bone marrow. Blood 98, 2396–2402 (2001).
Erices, A., Conget, P. & Minguell, J. J. Mesenchymal progenitor cells in human umbilical cord blood. Br. J. Haematol. 109, 235–242 (2000).
Ramasamy, R. et al. Mesenchymal stem cells inhibit proliferation and apoptosis of tumor cells: impact on in vivo tumor growth. Leukemia 21, 304–310 (2007).
Wong, R. S. Mesenchymal stem cells: angels or demons? J. Biomed. Biotechnol. 459510, 24 (2011).
Qiao, L. et al. Suppression of tumorigenesis by human mesenchymal stem cells in a hepatoma model. Cell Res. 18, 500–507 (2008).
Otsu, K. et al. Concentration-dependent inhibition of angiogenesis by mesenchymal stem cells. Blood 113, 4197–4205 (2009).
Klopp, A. H., Gupta, A., Spaeth, E., Andreeff, M. & Marini, F. 3rd Concise review: Dissecting a discrepancy in the literature: do mesenchymal stem cells support or suppress tumor growth? Stem Cells 29, 11–19 (2011).
Bowman, T., Garcia, R., Turkson, J. & Jove, R. STATs in oncogenesis. Oncogene 19, 2474–2488 (2000).
Darnell, J. E. Jr. STATs and gene regulation. Science 277, 1630–1635 (1997).
Clevenger, C. V. Roles and regulation of stat family transcription factors in human breast cancer. Am. J. Pathol. 165, 1449–1460 (2004).
Garcia, R. et al. Constitutive activation of Stat3 in fibroblasts transformed by diverse oncoproteins and in breast carcinoma cells. Cell Growth Differ. 8, 1267–1276 (1997).
Ling, X. et al. Mesenchymal Stem Cells Overexpressing IFN-beta Inhibit Breast Cancer Growth and Metastases through Stat3 Signaling in a Syngeneic Tumor Model. Cancer Microenviron. 3, 83–95 (2010).
Kaushik, N., et al. Low-dose radiation decreases tumor progression via the inhibition of the JAK1/STAT3 signaling axis in breast cancer cell lines. Sci. Rep. 7 (2017).
Mirzaei, H. et al. The therapeutic potential of human adipose-derived mesenchymal stem cells producing CXCL10 in a mouse melanoma lung metastasis model. Cancer Lett. 10, 029 (2018).
Gudkov, A. V. & Komarova, E. A. The role of p53 in determining sensitivity to radiotherapy. Nat. Rev. Cancer 3, 117–129 (2003).
Qu, J. Q. et al. MiR-23a sensitizes nasopharyngeal carcinoma to irradiation by targeting IL-8/Stat3 pathway. Oncotarget 6, 28341–28356 (2015).
Zhou, B. B. & Elledge, S. J. The DNA damage response: putting checkpoints in perspective. Nature 408, 433–439 (2000).
Al-Ejeh, F. et al. Breast cancer stem cells: treatment resistance and therapeutic opportunities. Carcinogenesis 32, 650–658 (2011).
Karamboulas, C. & Ailles, L. Developmental signaling pathways in cancer stem cells of solid tumors. Biochim. Biophys. Acta 2, 27 (2013).
Fouse, S. D. & Costello, J. F. Cancer Stem Cells Activate STAT3 the EZ Way. Cancer Cell. 23, 711–713 (2013).
Guan, J. L. Integrin signaling through FAK in the regulation of mammary stem cells and breast cancer. IUBMB Life 62, 268–276 (2010).
Norozi, F., Ahmadzadeh, A., Shahjahani, M., Shahrabi, S. & Saki, N. Twist as a new prognostic marker in hematological malignancies. Clin. Transl. Oncol. 18, 113–124 (2016).
Erta, M., Quintana, A. & Hidalgo, J. Interleukin-6, a major cytokine in the central nervous system. Int. J. Biol. Sci. 8, 1254–1266 (2012).
Kuai, W. X. et al. Interleukin-8 associates with adhesion, migration, invasion and chemosensitivity of human gastric cancer cells. World J. Gastroenterol. 18, 979–985 (2012).
Liu, J. et al. MiR-143 inhibits tumor cell proliferation and invasion by targeting STAT3 in esophageal squamous cell carcinoma. Cancer Lett. 373, 97–108 (2016).
Bishop, J. L., Thaper, D. & Zoubeidi, A. The Multifaceted Roles of STAT3 Signaling in the Progression of Prostate Cancer. Cancers 6, 829–859 (2014).
Xuan, X. et al. Stat3 promotes invasion of esophageal squamous cell carcinoma through up-regulation of MMP2. Mol. Biol. Rep. 42, 907–915 (2015).
He, N. et al. Embryonic stem cell preconditioned microenvironment suppresses tumorigenic properties in breast cancer. Stem Cell Res. Ther. 7, 95 (2016).
Foroni, C., Broggini, M., Generali, D. & Damia Epithelial-mesenchymal, G. transition and breast cancer: role, molecular mechanisms and clinical impact. Cancer Treat. Rev. 38, 689–697 (2012).
Xu, Q. et al. Targeting Stat3 blocks both HIF-1 and VEGF expression induced by multiple oncogenic growth signaling pathways. Oncogene 24, 5552–5560 (2005).
Wu, X. et al. Overcoming chemo/radio-resistance of pancreatic cancer by inhibiting STAT3 signaling. Oncotarget 7, 11708–11723 (2016).
Feng, L. et al. Cell cycle-dependent inhibition of 53BP1 signaling by BRCA1. Cell Discov. 1, 15019 (2015).
Huen, M. S., Sy, S. M. & Chen, J. BRCA1 and its toolbox for the maintenance of genome integrity. Nat. Rev. Mol. Cell Biol. 11, 138–148 (2010).
Bunting, S. F. et al. 53BP1 inhibits homologous recombination in Brca1-deficient cells by blocking resection of DNA breaks. Cell 141, 243–254 (2010).
Bouwman, P. et al. 53BP1 loss rescues BRCA1 deficiency and is associated with triple-negative and BRCA-mutated breast cancers. Nat. Struct. Mol. Biol. 17, 688–695 (2010).
Stracker, T. H., Roig, I., Knobel, P. A. & Marjanovic, M. The ATM signaling network in development and disease. Front Genet 4 (2013).
Patel, A. G., Sarkaria, J. N. & Kaufmann, S. H. Nonhomologous end joining drives poly(ADP-ribose) polymerase (PARP) inhibitor lethality in homologous recombination-deficient cells. Proc Natl Acad Sci USA 108, 3406–3411 (2011).
Raderschall, E., Golub, E. I. & Haaf, T. Nuclear foci of mammalian recombination proteins are located at single-stranded DNA regions formed after DNA damage. Proc Natl Acad Sci U S A 96, 1921–1926 (1999).
Zhou, B. B. & Bartek, J. Targeting the checkpoint kinases: chemosensitization versus chemoprotection. Nat. Rev. Cancer 4, 216–225 (2004).
Reinhardt, H. C. & Schumacher, B. The p53 network: cellular and systemic DNA damage responses in aging and cancer. Trends Genet. 28, 128–136 (2012).
Maier, P., L. Hartmann, F. Wenz, & C. Herskind Cellular Pathways in Response to Ionizing Radiation and Their Targetability for Tumor Radiosensitization. Int J Mol Sci 17 (2016).
Kim, S. M. et al. Irradiation enhances the tumor tropism and therapeutic potential of tumor necrosis factor-related apoptosis-inducing ligand-secreting human umbilical cord blood-derived mesenchymal stem cells in glioma therapy. Stem Cells 28, 2217–2228 (2010).
Leng, L. et al. Molecular imaging for assessment of mesenchymal stem cells mediated breast cancer therapy. Biomaterials. 35, 5162–5170 (2014).
Katakowski, M. et al. Exosomes from marrow stromal cells expressing miR-146b inhibit glioma growth. Cancer Lett. 335, 201–204 (2013).
Lee, J. K. et al. Exosomes derived from mesenchymal stem cells suppress angiogenesis by down-regulating VEGF expression in breast cancer cells. PLoS One 8, e84256 (2013).
Zhong, Z., Wen, Z. & Darnell, J. E. Jr. Stat3: a STAT family member activated by tyrosine phosphorylation in response to epidermal growth factor and interleukin-6. Science 264, 95–98 (1994).
Spitzner, M. et al. STAT3 inhibition sensitizes colorectal cancer to chemoradiotherapy in vitro and in vivo. Int. J. Cancer 134, 997–1007 (2014).
Zang, C. et al. IL-6/STAT3/TWIST inhibition reverses ionizing radiation-induced EMT and radioresistance in esophageal squamous carcinoma. Oncotarget 8, 11228–11238 (2017).
Deng, J. et al. STAT3 is associated with lymph node metastasis in gastric cancer. Tumour Biol. 34, 2791–2800 (2013).
Bu, X., Zhao, C., Wang, W. & Zhang, N. GRIM-19 inhibits the STAT3 signaling pathway and sensitizes gastric cancer cells to radiation. Gene 512, 198–205 (2013).
Han, Z. et al. Silencing of the STAT3 signaling pathway reverses the inherent and induced chemoresistance of human ovarian cancer cells. Biochem. Biophys. Res. Commun. 435, 188–194 (2013).
Zhang, Q. et al. STAT3 inhibitor stattic enhances radiosensitivity in esophageal squamous cell carcinoma. Tumour Biol. 36, 2135–2142 (2015).
Adachi, M., Cui, C., Dodge, C. T., Bhayani, M. K. & Lai, S. Y. Targeting STAT3 inhibits growth and enhances radiosensitivity in head and neck squamous cell carcinoma. Oral. Oncol. 48, 1220–1226 (2012).
Li, X., Wang, H., Lu, X. & Di, B. Silencing STAT3 with short hairpin RNA enhances radiosensitivity of human laryngeal squamous cell carcinoma xenografts in vivo. Exp. Ther. Med. 1, 947–953 (2010).
Ouedraogo, Z. G. et al. STAT3 Serine 727 Phosphorylation: A Relevant Target to Radiosensitize Human Glioblastoma. Brain. Pathol. 26, 18–30 (2016).
Agarwal, S. et al. G-CSF Promotes Neuroblastoma Tumorigenicity and Metastasis via STAT3-Dependent Cancer Stem Cell Activation. Cancer Res. 75, 2566–2579 (2015).
Thakur, R., Trivedi, R., Rastogi, N., Singh, M. & Mishra, D. P. Inhibition of STAT3, FAK and Src mediated signaling reduces cancer stem cell load, tumorigenic potential and metastasis in breast cancer. Sci. Rep. 5, 10194 (2015).
Yang, Z. X. et al. CD106 identifies a subpopulation of mesenchymal stem cells with unique immunomodulatory properties. PLoS One 8, 12 (2013).
Wang, L. et al. CD44 antibody-targeted liposomal nanoparticles for molecular imaging and therapy of hepatocellular carcinoma. Biomaterials. 33, 5107–5114 (2012).
Xu, S. et al. Salvianolic acid B inhibits platelets-mediated inflammatory response in vascular endothelial cells. Thromb. Res. 135, 137–145 (2015).
Liu, Y. et al. Rapamycin induces Bad phosphorylation in association with its resistance to human lung cancer cells. Mol. Cancer Ther. 11, 45–56 (2012).
Franken, N. A., Rodermond, H. M., Stap, J., Haveman, J. & van Bree, C. Clonogenic assay of cells in vitro. Nat. Protoc. 1, 2315–2319 (2006).
Charafe-Jauffret, E. et al. Breast cancer cell lines contain functional cancer stem cells with metastatic capacity and a distinct molecular signature. Cancer Res. 69, 1302–1313 (2009).
Gupta, P. B. et al. Identification of selective inhibitors of cancer stem cells by high-throughput screening. Cell 138, 645–659 (2009).
Fillmore, C. M. & Kuperwasser, C. Human breast cancer cell lines contain stem-like cells that self-renew, give rise to phenotypically diverse progeny and survive chemotherapy. Breast. Cancer Res. 10, 26 (2008).
Acknowledgements
This study was supported by the National Natural Science Foundation of China (No. 31670859, 31800703), CAMS Innovation Fund for Medical Science (No.2017-I2M-1-016), Natural Science Foundation of Tianjin (No.18JCQNJC12300, 18JCYBJC26800), the PUMC Youth Fund and the Fundamental Research Funds for the Central Universities (No.2017310028), China Postdoctoral Science Foundation (No.2018M630106), the Fundamental Research Funds for the Central Universities (10023201601602).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Y. Shi
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
He, N., Kong, Y., Lei, X. et al. MSCs inhibit tumor progression and enhance radiosensitivity of breast cancer cells by down-regulating Stat3 signaling pathway. Cell Death Dis 9, 1026 (2018). https://doi.org/10.1038/s41419-018-0949-3
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-018-0949-3
This article is cited by
-
Exosomes released by oxidative stress-induced mesenchymal stem cells promote murine mammary tumor progression through activating the STAT3 signaling pathway
Molecular and Cellular Biochemistry (2024)
-
Mesenchymal stem cell-released oncolytic virus: an innovative strategy for cancer treatment
Cell Communication and Signaling (2023)
-
Human umbilical cord mesenchymal stem cells conditioned medium exerts anti-tumor effects on KGN cells in a cell density-dependent manner through activation of the Hippo pathway
Stem Cell Research & Therapy (2023)
-
Mesenchymal Stem Cells and Selenium Nanoparticles Synergize with Low Dose of Gamma Radiation to Suppress Mammary Gland Carcinogenesis via Regulation of Tumor Microenvironment
Biological Trace Element Research (2023)
-
TA-MSCs, TA-MSCs-EVs, MIF: their crosstalk in immunosuppressive tumor microenvironment
Journal of Translational Medicine (2022)