Abstract
Pin1 is the only known peptidyl-prolyl cis–trans isomerase (PPIase) that specifically recognizes and isomerizes the phosphorylated Serine/Threonine-Proline (pSer/Thr-Pro) motif. The Pin1-mediated structural transformation posttranslationally regulates the biofunctions of multiple proteins. Pin1 is involved in many cellular processes, the aberrance of which lead to both degenerative and neoplastic diseases. Pin1 is highly expressed in the majority of cancers and its deficiency significantly suppresses cancer progression. According to the ground-breaking summaries by Hanahan D and Weinberg RA, the hallmarks of cancer comprise ten biological capabilities. Multiple researches illuminated that Pin1 contributes to these aberrant behaviors of cancer via promoting various cancer-driving pathways. This review summarized the detailed mechanisms of Pin1 in different cancer capabilities and certain Pin1-targeted small-molecule compounds that exhibit anticancer activities, expecting to facilitate anticancer therapies by targeting Pin1.
Similar content being viewed by others
Facts
-
Pin1 is the only known peptidyl-prolyl cis–trans isomerase (PPIase) that regulates the conformational transformation of phosphorylated Serine/Threonine-Proline (pSer/Thr-Pro) motif.
-
Pin1 is highly expressed in the majority of cancers and negatively related to the clinical prognosis.
-
Pin1 facilitates multiple cancer-driving pathways.
-
Pin1 is a potential target for cancer therapy.
Open Questions
-
What are the mechanisms for the high expression of Pin1 in cancer?
-
How does Pin1 upregulate the oncogenes and inhibit the cancer suppressors?
-
What are the molecular mechanisms of Pin1 that lead to cancer immune escape?
-
How does Pin1 facilitate the tumor-promoting inflammation?
Introduction
Proline (Pro)-directed Serine/Threonine (Ser/Thr) phosphorylation is a common modification of numerous signaling pathways. Many Pro-directed kinases, including mitogen-activated protein kinases and cyclin-dependent kinases (CDKs), are involved in this process1,2,3. Owing to the unique side-chain groups of proline, peptidyl-prolyl adopts an alterable cis or trans conformation4. The peptidyl-prolyl cis–trans isomerases (PPIases) accelerate the structural transformation of peptidyl-prolyl to regulate the folding, subcellular location, stability, activation, and interaction of multiple proteins5,6,7. The PPIase superfamily includes cyclophilins, FK506-binding proteins (FKBPs), and parvulins8 (Fig. 1). Cyclophilins and FKBPs can be inhibited by the immunosuppressants cyclosporin A (CyA) and FK506/rapamycin, respectively8. Pin1 belongs to parvulins and can be inhibited by juglone9. It is the only known PPIase that mediates the isomerization of phosphorylated Ser/Thr-Pro (pSer/Thr-Pro) motif10. Pin1 is comprised of an N-terminal WW domain and a C-terminal PPIase domain, which are connected by a flexible linker11,12.
Peptidyl-prolyl uniquely exhibits a cis or trans conformation. Different proline-directed kinases and phosphatases specifically recognize cis or trans Ser/Thr-Pro motif to modify the function of target proteins that highlight the PPIase-mediated structural transformation. The PPIases, including cyclophilins, FKBPs, and parvulins. Cyclophilins and FKBPs mediate the turnover of unphosphorylated substrates and Pin1 mediates the isomerization of phosphorylated substrates
The expression of Pin1 is immediately regulated by transcription factors E2F13 and NOTCH114. Beside, the CCAAT/enhancer binding protein-α (C/EBPα)-p30 increases Pin1 expression by recruiting E2F to the promoter of Pin115. After that, the mRNA level of Pin1 is reduced by some microRNAs (miRNAs), including the recently discovered miR-37016 and miR874-3p17. Moreover, the posttranslational modifications of Pin1, including phosphorylation18,19, sumoylation20, ubiquitination21, and oxidation22, regulate the stability, substrate-binding ability, PPIase activity, and subcellular localization of Pin1. These processes are always aberrant in cancer that contribute to the high expression and/or overactivation of Pin1 (Fig. 2).
Pin1 is involved in multiple cellular processes, including division23, differentiation24, senescence25, and apoptosis26. Pin1 is always deficient in degenerative disorders, including Parkinson’s disease (PD)27, Alzheimer’s disease (AD)28, and Huntington’s disease (HD)29. In contrast, it is highly expressed in most cancers, especially in cancer stem cells (CSCs), and negatively related to the clinical prognosis30,31,32. The depletion of Pin1 significantly inhibits tumorigenesis in the mice models that are derived by mutation of p5333, activation of HER2/RAS34 or constitutive expression of c-Myc35. Additionally, many Pin1-targeted inhibitors, including all trans retinoic acid (ATRA)36, juglone37, and KPT-656638, have showed cancer suppression ability in multiple researches (Table 1).
According to the existing research, Pin1 upregulates >50 oncogenes or proliferation-promoting factors while inhibits >20 tumor suppressors or proliferation-restraining factors8,30 (Fig. 2). However, what are the specific mechanisms of Pin1 in different cancer capabilities? Following the ground-breaking summaries of Hanahan and Weinberg, we have a clearer recognition regarding the hallmarks of cancer. The ten major cancer capabilities are sustaining proliferative signaling, evading growth suppressors, activating invasion and metastasis, enabling replicative immortality, inducing angiogenesis, resisting cell death, evading immune destruction, tumor-promoting inflammation, reprogramming of energy metabolism, and genome instability and mutation39,40. Mounts of researches indicated that Pin1 is an active participant of these aberrant processes8 (Fig. 3). In this review, we summarized the detailed mechanisms of Pin1 that contribute to these cancer capabilities and certain Pin1-targeted small-molecule compounds that exhibit anticancer activities, expecting to facilitate anticancer therapies by targeting Pin1.
Pin1 sustains the proliferative signaling
Cell proliferation is strictly regulated by the intracellular and extracellular signals41,42, but cancer cells utilize many pathways to sustain proliferation43,44. Pin1 was initially identified as a regulator of mitosis45 and many subsequent studies showed that it facilitates multiple proliferation-promoting pathways in cancer46 (Fig. 4).
Pin1 upregulates transcription factors, such as β-catenin, HIF1, ERα, NF-κB, JUN, FOS, NUR77, and co-activator PKM2 by enhancing their stability, nuclear translocation, transcriptional activity, and/or activating their upstream regulators in multiple cancers. These transcription factors induce the expression of target genes especially various cyclins to control the cell cycle. Besides, Pin1 is a target of the transcription factor E2F, which is inhibited by pRb. In many cancers, pRb can be phosphorylated and inactivated by CDKs, which facilitates the expression of Pin1. Moreover, Pin1 reverses the p53-mediated growth inhibition by stabilizing ΔNp63. In general, Pin1 facilitates the cancer proliferation through regulating different substrates at several levels
Estrogen receptor α (ERα) promotes the proliferation of cancer, especially breast cancer, by regulating the expression of estrogen response element (ERE)-containing genes47. Research indicated that Pin1 increases the transcriptional activity48, ERE binding affinity49, and inhibits the E3 ligase E6AP-induced degradation of ERα in breast cancer50. The garlic extract diallyl trisulfide-treated breast cancer cells exhibit a reduced expression of Pin1 along with reduced ERα activity and cell proliferation51. Besides, the high expression of Pin1 and HER2 are concurrent in most breast cancers. Pin1 activates HER2 by inhibiting its ubiquitination52 and destabilizing its transcriptional corepressor SMRT53.
Additionally, activation of the nuclear factor (NF)-κB pathway strongly induces cancer cell proliferation. Pin1 activates the NF-κb pathway by enhancing the nuclear accumulation of RelA/p65, c-Rel, and v-Rel54,55. Besides, Pin1 inhibits the E3 ligase SOCS-1-mediated ubiquitination of p6554. Pin1-mediated activation of the NF-κB pathway is involved in the proliferation of glioblastoma56, endometrial carcinoma57, acute myeloid leukemia (AML)58, and hepatocellular carcinoma (HCC)59.
Furthermore, ΔNp63, an isoform of p63 that lacks an intact N-terminal transactivation domain, is critical for tumorigenesis60. Pin1 inhibits the E3 ligase WWP1-induced ubiquitination of ΔNp63 to increase the proliferation of human oral squamous cell carcinoma61. Pin1 also stabilizes bromodomain-containing protein 4 (BRD4), a transactivator of multiple oncogenes, to promote the proliferation, migration, and invasion of gastric cancer62. Besides, Pin1 upregulates many other proliferation-inducing factors, including β-catenin63, FoxM164, XBP165, NUR7766, c-Jun67, and c-Myc68.
Moreover, Pin1 also induces the proliferation of non-tumorous cells, such as pancreatic β cells69, hepatic oval cells70, and spermatogonial stem cells71. The deficiency of Pin1 significantly suppresses the growth of multiple cell types, indicating that Pin1 is a potential target to treat hyperplastic diseases.
Pin1 downregulates numerous tumor suppressors
The tumor suppressors act as surveillant of multiple cellular processes to prevent cancerization and suppress cancer progression, but cancer cells utilize various mechanisms to surmount these barriers. Research indicated that Pin1 is guilty for the inactivation of numerous tumor suppressors.
When DNA is damaged, the tumor-suppressor retinoblastoma protein (pRb) directly inhibits the transcription factor E2F to arrest the cell cycle72. However, pRb is usually inactivated in cancer cells due to reduced expression and/or continuously hyperphosphorylation73,74, which partially attribute to Pin1. The insulin-like growth factor 1-stimulated wild-type mouse embryonic fibroblasts (MEFs) exhibit hyperphosphorylated pRb and highly expressed Pin1 simultaneously, but Pin1−/− MEFs show a considerably lower level of phosphorylated pRb75. Research illuminated that Pin1 promotes CDK-induced phosphorylation76 and inhibits PP2A-mediated dephosphorylation77 of pRb that subsequently activate E2F and trigger cells into S phase.
The promyelocytic leukemia protein (PML) is another powerful tumor suppressor but always mutant in cancer. Research indicated that Pin1 destabilizes PML to promote the survival and proliferation of breast cancer78. Pin1 enhances the E3 adapter KLHL20-induced ubiquitination of PML to promote the proliferation and angiogenesis of prostate cancer79. Besides, Pin1 also stabilizes the oncogenic fusion protein PML-RARα in AML81. Suppression of Pin1 significantly inhibits the proliferation of breast cancer cells and restores the expression of PML and SMRT80.
Additionally, the runt-related transcription factor 3 (RUNX3) acts as an ERα inhibitor in breast cancer82. Pin1 decreases the transcriptional activity and increases the ubiquitin-dependent degradation of RUNX383. The E3 ligase FBXW7 suppresses cancer by reducing multiple oncogenes, but Pin1 inactivates FBXW7 by disrupting its dimerization and promoting its self-ubiquitination84. Pin1 also downregulates other tumor suppressors, including Kruppel-like factor 1085, suppressor of variegation 3-9 homolog 186, and CDK1087.
Interestingly, a number of studies indicated that Pin1 increases p53-induced cell senescence and apoptosis88,89. However, Pin1 expression is higher in HCC cells with mutant p53 (p53M) compared to wild-type p53 (p53WT), and the deletion of Pin1 significantly reduces the proliferation of p53M HCC cells but not p53WT90. More research revealed that Pin1 facilitates the p53M-induced aggressiveness of cancers33,91,92, which contributes to a reasonable explanation for why p53 is aberrant in most cancers.
Pin1 promotes cancer invasion and metastasis
Cancer invasion and metastasis are the leading cause of death in cancer patients. Research revealed that the expression of Pin1 is much higher in the metastatic cancer compared with primary, which reduces the invasion- and metastasis-promoting function of Pin193,94.
The transforming growth factor (TGF)-β pathway inhibits the proliferation but promotes the metastasis of cancer95,96. The SMAD proteins are major downstream adapters of TGF-β signal and extensively recognized by WW domain-containing proteins, including Pin197,98. Initial research revealed that Pin1 induces the E3 ligase Smurf-2-mediated degradation of SMADs to suppress the TGF-β signal99. However, the later research indicated that Pin1 promotes the TGF-β-induced metastasis of prostate cancer cells100. Inhibiting the phosphorylation of SMAD3 reduces the interaction with Pin1 and remarkably suppresses the aggressiveness of breast cancer101. Therefore, the function of the TGF-β pathway is complex and Pin1-mediated TGF-β pathway in cancer requires a deeper investigation.
Pin1 also increases the invasion and metastasis of cancer by activating the NOTCH pathway. In breast cancer, Pin1 facilitates the transcriptional activity of NOTCH1 by potentiating its γ-secretase-mediated cleavage14. Meanwhile, NOTCH1 induces the expression of Pin1, which consequently form a positive loop to enhance cancer cell transformation14. Pin1 also promotes breast CSC self-renewal and metastasis by inhibiting FBXW7-mediated degradation of NOTCH1 and NOTCH432. Besides, Pin1 activates the NOTCH3 signal by enhancing its cleavage and stabilizing its intracellular domain in T cell acute lymphoblastic leukemia (T-ALL) cell lines and mouse models. The deletion of Pin1 markedly decreases the NOTCH-induced invasion of T-ALL cells102.
Moreover, PTP-PEST and FAK are two pivotal effectors of the RAS signal, which are involved in tumor metastasis103. Pin1 facilitates the interaction of PTP-PEST with FAK to accelerate the Tyr397 dephosphorylation of FAK, which consequently induce the metastasis of numerous cancers104,105. Pin1 also promotes the epithelial–mesenchymal transition of MCF-7 cells by increasing the transcriptional activity of signal transducer and activator of transcription factor 3 (STAT3) and recruiting its transcription coactivator p300106. Additionally, Pin1 promotes the invasion and metastasis of multiple cancers by activating NF-κB107, p53M108, β-catenin63, and BRD462.
Pin1 enables the replicative immortality of cancer
After a limited number of cycles, the majority of normal cells enter a nonproliferative but viable state, which called cellular senescence. The cells that continue to divide will face a fatal crisis, which causes the death of most cells, but the minority that passes this barrier will be immortal39,40. The mechanisms that control the proliferative barrier include telomere shortening, DNA damage, and mitochondria damage109,110,111. The antisenescence function of Pin1 is widely revealed in vascular smooth muscle cells112, cardiac progenitor cells25, tendon stem/progenitor cells113, fibroblasts114, and various cancer cells.
In most cancer, telomerase is reactivated to maintain the telomeric DNA, but telomeric repeat-binding factors (TRFs) prevent its elongation115. Research demonstrated that Pin1 elongates the telomere via promoting E3 ligase Fbx4-mediated degradation of TRF1 in multiple cancer cells116. Additionally, the early mitotic inhibitor 1 (Emi1) is an inhibitor of the DNA damage-induced senescence117. Pin1 promotes the proliferation and suppresses the senescence of several cancer cells by preventing the E3 ligase βTrCP-induced degradation of Emi1118. Pin1 also suppresses the senescence-inducing factors pRb and PML in multiple cancers. Pin1 enhances p53-induced senescence and apoptosis; however, inhibiting Pin1 leads to senescence in p53-interfered BJ cells and overexpression of Pin1 reverses the p53 responder BTG2-induced senescence114.
Furthermore, the Pin1 deficit contributes to many degenerative diseases, including AD28, HD29, and PD27, all of which are related to aberrant neuronal senescence and apoptosis. Considering the antisenescence function, Pin1 and its substrates are potential targets to treat both degenerative diseases and cancers.
Pin1 enhances cancer-induced angiogenesis
The angiogenesis is strictly controlled in vivo. However, the rapidly expanded cancer can induce continuous angiogenesis to maintain the sustenance of nutrients and oxygen, as well as the elimination of metabolic waste and carbon dioxide. Abundant evidence illuminated that Pin1 is involved in cancer-induced angiogenesis119.
The hypoxia-inducible factor 1α (HIF-1α) strongly induces angiogenesis by promoting the expression of vascular endothelial growth factor (VEGF) in the hypoxia cancer tissue120. Studies indicated that the high expression of Pin1, HIF-1α, and VEGF are positively related in TAM-resistant MCF-7 (TAMR-MCF-7) cells121,122. Pin1 increases the stability and transcriptional activity of HIF-1α in many cancers123,124. PML inhibits HIF-1α-induced angiogenesis both in clear cell renal cell carcinoma125 and human umbilical vein endothelial cells126, but Pin1 destabilizes PML in multiple cancers. HIF-1α also induces the expression of KLHL20, which cooperates with Pin1 to induce the ubiquitin-dependent degradation of PML79. Besides, Pin1 facilitates the NF-κB-induced expression of VEGF in HCC59. Additionally, the VEGF-promoting transcriptional factors, such as FoxM1127 and β-catenin128, are upregulated by Pin1 in numerous cancers.
Inhibition of Pin1 significantly reduces the cancer-induced angiogenesis. Directly suppressing the expression of Pin1 by RNAi inhibits both growth and angiogenesis of prostate cancer129. The phosphoinositide-3 kinase (PI3K)/p38 signals increase the expression of Pin1 via activating E2F1121. In TAMR-MCF-7 cells, the PI3K inhibitor quercetin122 and the E2F1 inhibitor amurensin G130 markedly reduced the expression of Pin1, secretion of VEGF, and angiogenesis. In conclusion, Pin1 enhances the angiogenesis of multiple cancers by promoting the expression of VEGF.
Pin1 facilitates the cell death resistance of cancer
Apoptosis is an important form of programmed cell death, which acts as a natural barrier to prevent cells from developing into cancers131,132. However, cancer cells can block the proapoptotic signals and activate antiapoptotic signals to make them survive in cytotoxic stress. Pin1 is a powerful “weapon” of cancer to against apoptosis133.
First, Pin1 inhibits the proapoptotic factors. Outer mitochondrial membrane located BAX and BAK induce apoptosis by enhancing the release of cytochrome c134. In human eosinophils (Eos), Pin1 inhibits the BAX-induced apoptosis by preventing its mitochondria translocation135. Besides, the death-associated proteins DAXX and FADD are two critical responders of CD95/Fas-induced apoptosis136,137. The Fas signal notably increases the activity and nuclear translocation of FADD by phosphorylating its Ser194 and inhibits Pin1 by phosphorylating its Ser16138. However, exogenetic expression of Pin1 maintains the cytoplasmic location of FADD by accelerating its dephosphorylation, which consequently blocks the Fas-FADD pathway138. Pin1 also isomerizes DAXX to promote its ubiquitin-dependent degradation in malignant human gliomas139. Additionally, DNA damage-induced apoptosis is mediated by many surveillance proteins, such as p53, PML, and pRb. However, PML and pRb are downregulated by Pin1 in numerous cancers. Pin1 enhances p53-induced apoptosis but facilitates the cancer-driving function of p53M.
Second, Pin1 upregulates the antiapoptosis factors. The B-cell lymphoma 2 (BCL-2) family proteins inhibit apoptosis via directly inactivating BAX and BAK depending on their shared BH3 domain140. Research indicated that Pin1 enhances the stabilization and cell death resistance ability of BCL-2 and myeloid cell leukemia-1 (MCL-1)141,142. Some anticancer drugs, such as sorafenib and amsacrine, induce the apoptosis of cancer cells by reversing the Pin1-mediated stability of MCL-1141,143. Pin1 also enhances the survival of cisplatin-treated cervical cancer cells by upregulating Wnt/β-catenin and FoxM1 pathways144. In addition, Pin1 upregulates LC-3 to induce protective autophagy, which consequently increases the tamoxifen resistance of breast cancer145.
Interestingly, Pin1 increases the antiapoptotic activity of Survivin in HCC146, which is opposite in neuroblastoma147. In addition to p53 and Survivin, Pin1 also induces cell apoptosis by activating and stabilizing the tumor-suppressor homeodomain interacting protein kinase 2148, increasing the mitochondrial translocation of p66Shc26, as well as inhibiting the activity of ataxia telangiectasia and rad3 related149. However, all of these results were observed in non-tumorous cells.
Pin1 helps cancer cells to evade immune destruction
With further research on cancer and the immune system, the traditional concept that the immune system prevents tumor initiation and development has been questioned150. Cancer cells arising from immunocompetent mice are much more aggressive than that arising from immunodeficient mice150. Multiple studies demonstrated that Pin1 participates in the regulation of immune response.
The Toll-like receptors (TLRs) recognize pathogen-associated molecular patterns to initiate the immune response151,152. Pin1 is involved in the regulation of the TLR signals. In plasmacytoid dendritic cells, engaged TLR7/TLR9 activate interleukin (IL)-1 receptor associated kinase-1 (IRAK1) and subsequently activated IRAK1 induces the secretion of type I interferon (IFN-αβ) by activating the transcription factor IFN-regulatory factor 7 (IRF7)153,154. IRAK1 is autophosphorylated within the activated TLR complex and then Pin1-mediated isomerization induces its liberation from the complex155. Pin1-induced overactivation of the TLR-7/TLR-9/IRAK-1/IRF-7 signal contributes to the autoimmune disease systemic lupus erythematosus156. In addition, IRF3 is a downstream adapter of TLR3/TLR4 signal that induces the expression of IFN-β in the antiviral response157. However, Pin1 reduces the transcriptional activity and promotes ubiquitin-dependent degradation of IRF3 leading to reduced production of IFN-β in poly(I)poly(C)- or RIG-I-stimulated immune cells158. Furthermore, the double-stranded RNA-induced expression of IFN-β is significantly lower and the replication of the invading virus is higher in Pin1+/+ mice compared with Pin1–/– mice158. Both tripartite motif-containing 21 and PML isoform IV stabilize IRF3 and enhance IRF3-mediated production of IFN-β by disturbing the interaction of Pin1 with IRF3159,160. Besides, the IRF3-mediated expression of IFN-λ1 is also decreased by the exogenous expressed Pin1161. In conclusion, the function of Pin1 in immune regulation is cell-type and pathogen dependent.
Pin1 is involved in the immune escape of cancer. Indoleamine-pyrrole 2,3-dioxygenase (IDO) exhausts local tryptophan to limit the function of T lymphocytes162. Research indicated that the cytotoxic T lymphocyte-associated protein 4 (CTLA-4)-stimulated dendritic cells (DCs) produce IDO upon simultaneously activating the PI3K and NOTCH pathways. Pin1 increases the enzyme activity of casein kinase II to abolish the PTEN-mediated suppression of PI3K163. Suppressing the NOTCH signal significantly reduces the expression of Pin1 and the CTLA-4-induced IDO production163. Besides, TGF-β powerfully inhibits the proliferation and function of multiple immune cells to disorganize the host immune surveillance164. In the lung, liver, and cardiac fibrosis mouse models and even in patients, Pin1 increases the expression of TGF-β by stabilizing its mRNA and protein165,166,167. It is clear that Pin1 enhances the TGF-β-induced invasion and migration of cancers, but the Pin1-mediated TGF-β signal in cancer immune escape requires a deeper investigation. In conclusion, targeting the immunoediting checkpoints is an effective strategy to treat cancer168 and Pin1 is a potential candidate.
Pin1 participates in the tumor-promoting inflammation
The release of necrotic cell content and microbe infection induce the inflammatory response by recruiting inflammatory cells into the local microenvironment169,170. Research indicated that chronic inflammation is correlative to cancer initiation and progression171.
Pin1 is a regulator of inflammatory response. In human neutrophils, NADPH oxidase catalyzes the production of superoxide which subsequently generates reactive oxygen species (ROS) to annihilate ingested microbes172. Pin1 promotes the formation of integrated NADPH oxidase by facilitating the membrane translocation of its cytosolic component, p47phox173,174. Besides, granulocyte macrophage colony-stimulating factor (GM-CSF) is essential for activation of circulating leukocytes175. The 3′-untranslated region of GM-CSF mRNA is abundant in AU-rich elements (AREs) and ARE-binding protein AUF1 negatively regulates its mRNA stability176. Pin1 stabilizes GM-CSF mRNA by decreasing the ARE-binding ability of AUF1 in Eos and T lymphocytes177,178. Inhibiting Pin1 significantly reduces the production of GM-CSF in allergen-treated rats179.
The aberrant Pin1-mediated inflammation contributes to numerous diseases, including cancer. Pin1 enhances the IL-22-induced proliferation and survival of breast cancer cells by activating mitogen-activated extracellular signal-regulated kinases, c-Jun, and STAT3180. Pin1 also increases the nuclear translocation of IRAK-M in DCs to enhance the IL-33-induced allergic airway inflammation181. Besides, Pin1 is correlated in other inflammatory diseases, including rheumatoid arthritis182, periodontitis183, diabetes-induced atherosclerosis184, nonalcoholic steatohepatitis185, and primary biliary cholangitis186.
However, research indicates that Pin1 also stabilizes and activates some anti-inflammatory proteins, such as NUR77187 and glucocorticoid receptor188. Similar to the immune regulation, Pin1 probability plays a dual role in inflammation regulation, but the cancer-promoting function of Pin1 is much more clarified.
Pin1 regulates the metabolic reprogramming of cancer
The metabolic pattern of glucose in mammalian cells is dependent on the oxygen environment. In aerobic conditions, the common intermediate pyruvate is mainly transferred to the mitochondria and efficiently produces massive ATP through oxidative phosphorylation, while in anaerobic conditions, pyruvate rapidly produces small amounts of ATP via glycolysis in the cytoplasm. The majority of normal cells are fueled by more efficient oxidative phosphorylation except for some special conditions, such as muscle cells in high-intensity exercise. However, the major metabolic pattern of cancer cells is glycolysis even in aerobic conditions, which called “aerobic glycolysis” or “Warburg effect” after its discoverer, Otto Warburg189. The “Warburg effect” is beneficial for cancer progression by supplying intermediates for rapid biosynthesis and avoiding cytostatic controls that are induced by activated oncogenes and/or mutant cancer suppressors190,191. The mechanisms that result in “Warburg effect” include increased glucose import, aberrant hypoxia response system, and the incapacitation of oxidative phosphorylation40. They are correlated with many aberrant signals, including PI3K/Akt/mammalian target of rapamycin192, Wnt/β-catenin193, HIF-1α194, and non-coding RNAs195, all of which finally regulate the expression and/or activation of metabolism-associated proteins.
Research indicates that Pin1 is a crucial regulator of the “Warburg effect” (Fig. 5). Pin1 increases the nuclear localization of the phosphorylated pyruvate kinase isozyme M2 (PKM2)196. The nuclear translocated PKM2 phosphorylates histone H3-T11 leading to the H3-K9 acetylation of target genes. Meanwhile, it also acts as a co-activator of β-catenin to increase the expression of CCDN1 and c-Myc. Subsequently, c-Myc induces the expression of glucose transporter 1 and lactate dehydrogenase A to promote the “Warburg effect”196,197. In addition, pyruvate dehydrogenase kinase 1 (PDHK1) suppresses the tricarboxylic cycle by phosphorylating and inactivating the pyruvate dehydrogenase (PDH) complex198. Pin1 increases the mitochondria translocation of PGK1 where it activates PDHK1 to facilitate the PDH-inhibited activity of PDHK1199. Moreover, Pin1 upregulates many other metabolic regulators directly, such as β-catenin, HIF-1α, and c-Myc. Targeting the metabolic reprogramming process is effective to suppress cancer progression200, and these studies revealed that Pin1 is a potential candidate to reverse these dysregulations.
“Warburg effect” that first observed by Otto Warburg is an aberrant characteristic of cancer. Pin1 facilitates the glucose metabolic reprogramming of cancer. On the one hand, Pin1 enhances the nuclear localization of PKM2. Nucleus PKM2 acts as a co-activator of β-catenin to promote the expression of LDH-A and GLUT1, thereby enhancing lactification and increasing glucose import, respectively. On the other hand, Pin1 increases the mitochondrial translocation of PGK1. Mitochondrial PGK1 phosphorylates and activates PDHK1, thus inactivating PDH and finally inhibits TCA cycle
Pin1 contributes to the genome instability and mutations of cancer
Cytotoxic factors, such as ionizing radiation and DNA topoisomerase II poisons, induce DNA double-strand breaks (DSBs) to trigger cellular senescence and apoptosis201. It is effective to suppress cancer by bringing in a mass of DNA damage202, but most cancer is insensitive to genome instability and mutations. Pin1 not only suppresses the DAN damage-induced senescence and apoptosis but also contributes the genome instability in cancers.
The mechanisms of DSBs repair include error-free homologous recombination (HR) and fallible nonhomologous end-joining (NHEJ)203. CtIP facilitates HR by promoting DNA-end resection204. However, Pin1 promotes the ubiquitin-dependent degradation of CtIP to attenuate HR and increase NHEJ, which consequently increase genome instability205. Pin1 also interacts with the DSB repair regulators 53BP1 and BRCA1, but there is no further study on them205. Besides, it is a promising direction that Pin1 modifies DSB repair through regulating the CDK-mediated DNA-end resection206. Meanwhile, rapid proliferation leads to replication-associated DNA damage207. Whether the Pin1-promoted proliferation induces DNA damage is also worth of deeper research.
The aberrant mitotic process also contributes to genome instability. Separase promotes sister chromatid paired segregation by cleaving ring-shaped cohesin208. Research indicated that the full function of separase requires the Pin1-mediated isomerization at its pSer1126-Pro motif209. The centrosome protein 55 kDa (Cep55) is crucial for the formation of midbodies in cytokinesis210. Pin1 increases the midbody translocation of Cep55 by facilitating its polo-like kinase 1-mediated phosphorylation at Ser436211. Pin1 is also involved in the SEPT9-mediated final separation of daughter cells212. Moreover, the overexpression of Pin1 promotes an abnormal centrosome duplication and chromosome instability in breast cancer213. Human papillomavirus (HPV) infection also induces genome instability and enhances the malignant phenotype of cervical cancers by promoting aberrant centrosome synthesis214. Research indicates that HPV-infected cervical lesions exhibit an elevated level of Pin1215, but the relationship of HPV-induced Pin1 and genome instability in cervical cancer requires a deeper investigation.
Targeting Pin1 is effective to suppress cancer
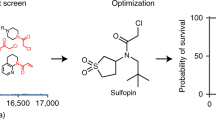
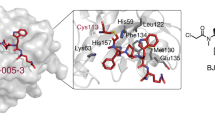
Targeting a single pathway to treat cancer is challenging because multiple aberrant pathways are involved in cancer progression39,40. Moreover, almost all of the current therapies are ineffective to treat CSCs216. Therefore, a strategy that simultaneously targets multiple cancer-driving pathways is urgently required. In cancers, Pin1 promotes and suppresses numerous oncogenes and tumor suppressors, respectively. Multiple studies demonstrated that inhibition of Pin1 is effective to suppress the progression of cancers. Aurora kinase A217, ribosomal S6 kinase 2218 and protein kinase A19 abolish the substrate interaction ability of Pin1 by phosphorylating its Ser16 residue in the WW domain. The death-associated protein kinase 1 inactivates Pin1 by phosphorylating Ser71 in the catalytic core of Pin1 that consequently inhibits the centrosome amplification of breast cancer cells18 but aggravates neurodegeneration219. Pin1-targeted microRNAs (miRNAs), including miR-200b220, miR-200c31, miR296-5p221, miR-37016, and miR874-3p17, inhibit human cancer progression via directly decreasing the mRNA level of Pin1. However, Pin1 can suppress the miRNA biogenesis by inhibiting exportin-5 (XPO5)222. In addition, researchers have discovered and synthesized many Pin1-targeted small-molecule compounds that exhibit anticancer activity (Table 1). ATRA, a currently used target drug for acute promyelocytic leukemia (APL), mechanically combines with the substrate-binding site of Pin1 to inhibit its activation in APL and breast cancer cells36. Yang et al. designed a novel slow-releasing, non-toxic, biodegradable, and biocompatible ATRA formulation. Their work showed that this novel formulation exhibits long-term inhibition of Pin1 and is more efficient than the traditional ATRA to suppress HCC cell growth223. Juglone, a compound extracted from walnut tree, and its derivative buparvaquone both inhibit Pin1 by covalently modifying its catalytic core9,224. Juglone is effective to suppress multiple cancer cells and universally used in Pin1 research. Cis-locked alkene peptidomimetics inhibit Pin1 by simulating a substrate of Pin1 and exhibit antiproliferation activities in an ovarian cancer cell line225. Imazamethabenz, an imidazoline ketone herbicide, inhibits migration and invasion and induces apoptosis in breast cancer cells via directly combining with Pin1226. The Pin1 inhibitor API-1 suppresses HCC development by restoring Pin1-inhibited miRNA biogenesis227. Recently, a novel Pin1 inhibitor, KPT-6566, inhibits the PPIase activity and induces the degradation of Pin1 by covalently interacting with its catalytic core38. Moreover, inhibiting Pin1 sensitizes many cancer cells to chemotherapy, including HCC to sorafenib228, breast cancer to trastuzumab229, rapamycin53, Taxol and 5-fluorouracil141, colon cancer to Taxol84, and AML to retinoic acid81. Increasing evidence showed that Pin1 is a potential target for cancer therapy. However, the majority of the existing inhibitors lack the required specificity, efficacy, and safety in clinical application.
Conclusions
The diversity and complexity of cancers are always challenging the treatment. Fortunately, after decades of research, we are uncovering the secret of cancer. Hanahan and Weinberg summarized the common hallmarks of cancer cells, which lets us recognize cancers more clearly. Currently, targeted therapies have applied to the treatment of multiple cancers, which have a higher efficacy and lower side effects than traditional therapies. However, the “smart” cancer cells, especially the CSCs, utilize multiple mechanisms to avoid being eradicated. Therefore, therapies that target common aberrant pathways to block the escape of tumor cells are urgently warranted. Pro-directed phosphorylation is an extensive modification of numerous pathways, which regulate multiple basic cellular processes, including proliferation, differentiation, metabolism, and death. Pin1 is the only known PPIase that mediates the cis–trans isomerization of pSer/Thr-Pro bond, which highlights its significance in the regulation of Pro-directed phosphorylation.
Pin1 is highly expressed in most cancers, especially CSCs. According to the existing discovery, some regulation loops, including at least Pin1/NOTCH/Pin1 axis14, Pin1/pRb/E2F/Pin1 axis13,72,76, and Pin1/XPO5/miRNAs/Pin1 axis31,222, contribute to the high expression of Pin1. Pin1 is involved in almost every cancer biocapability, suggesting that it is a potential common therapeutic target. There are many Pin1-targeted inhibitors, including the novel structure-based designed compounds, natural extracts, and their derivatives, that exhibit anticancer activity in in vitro, animal models and even in certain patients. Hence, the mechanisms of Pin1-induced cancer progression and targeting Pin1 for cancer therapy are worthy of further investigation. When matured, the extensive clinical applications would benefit many cancer patients.
References
Pearson, G. et al. Mitogen-activated protein (MAP) kinase pathways: regulation and physiological functions. Endocr. Rev. 22, 153–183 (2001).
Morgan, D. O. Cyclin-dependent kinases: engines, clocks, and microprocessors. Annu. Rev. Cell Dev. Biol. 13, 261–291 (1997).
Lu, K. P., Liou, Y. C. & Zhou, X. Z. Pinning down proline-directed phosphorylation signaling. Trends Cell Biol. 12, 164–172 (2002).
Reimer, U. et al. Side-chain effects on peptidyl-prolyl cis/trans isomerisation. J. Mol. Biol. 279, 449–460 (1998).
Lu, K. P., Finn, G., Lee, T. H. & Nicholson, L. K. Prolyl cis-trans isomerization as a molecular timer. Nat. Chem. Biol. 3, 619–629 (2007).
Takahashi, K., Uchida, C., Shin, R. W., Shimazaki, K. & Uchida, T. Prolyl isomerase, Pin1: new findings of post-translational modifications and physiological substrates in cancer, asthma and Alzheimer’s disease. Cell Mol. Life Sci. 65, 359–375 (2008).
Gothel, S. F. & Marahiel, M. A. Peptidyl-prolyl cis-trans isomerases, a superfamily of ubiquitous folding catalysts. Cell Mol. Life Sci. 55, 423–436 (1999).
Zhou, X. Z. & Lu, K. P. The isomerase PIN1 controls numerous cancer-driving pathways and is a unique drug target. Nat. Rev. Cancer 16, 463–478 (2016).
Hennig, L. et al. Selective inactivation of parvulin-like peptidyl-prolyl cis/trans isomerases by juglone. Biochemistry 37, 5953–5960 (1998).
Lu, K. P. & Zhou, X. Z. The prolyl isomerase PIN1: a pivotal new twist in phosphorylation signalling and disease. Nat. Rev. Mol. Cell Biol. 8, 904–916 (2007).
Verdecia, M. A., Bowman, M. E., Lu, K. P., Hunter, T. & Noel, J. P. Structural basis for phosphoserine-proline recognition by group IV WW domains. Nat. Struct. Biol. 7, 639–643 (2000).
Matena, A., Rehic, E., Hönig, D., Kamba, B. & Bayer, P. Structure and function of the human parvulins Pin1 and Par14/17. Biol. Chem. 399, 101–125 (2018).
Ryo, A. et al. PIN1 is an E2F target gene essential for Neu/Ras-induced transformation of mammary epithelial cells. Mol. Cell Biol. 22, 5281–5295 (2002).
Rustighi, A. et al. The prolyl-isomerase Pin1 is a Notch1 target that enhances Notch1 activation in cancer. Nat. Cell Biol. 11, 133–142 (2009).
Pulikkan, J. A. et al. Elevated PIN1 expression by C/EBPalpha-p30 blocks C/EBPalpha-induced granulocytic differentiation through c-Jun in AML. Leukemia 24, 914–923 (2010).
Chen, M. et al. Downregulation of microRNA-370 in esophageal squamous-cell carcinoma is associated with cancer progression and promotes cancer cell proliferation via upregulating PIN1. Gene 661, 68–77 (2018).
Leong, K. et al. miR-874-3p is down-regulated in hepatocellular carcinoma and negatively regulates PIN1 expression. Oncotarget 8, 11343–11355 (2017).
Lee, T. H. et al. Death-associated protein kinase 1 phosphorylates Pin1 and inhibits its prolyl isomerase activity and cellular function. Mol. Cell 42, 147–159 (2011).
Lu, P. J., Zhou, X. Z., Liou, Y. C., Noel, J. P. & Lu, K. P. Critical role of WW domain phosphorylation in regulating phosphoserine binding activity and Pin1 function. J. Biol. Chem. 277, 2381–2384 (2002).
Chen, C. H. et al. SENP1 deSUMOylates and regulates Pin1 protein activity and cellular function. Cancer Res. 73, 3951–3962 (2013).
Eckerdt, F. et al. Polo-like kinase 1-mediated phosphorylation stabilizes Pin1 by inhibiting its ubiquitination in human cells. J. Biol. Chem. 280, 36575–36583 (2005).
Chen, C. H. et al. Pin1 cysteine-113 oxidation inhibits its catalytic activity and cellular function in Alzheimer’s disease. Neurobiol. Dis. 76, 13–23 (2015).
Yaffe, M. B. et al. Sequence-specific and phosphorylation-dependent proline isomerization: a potential mitotic regulatory mechanism. Science 278, 1957–1960 (1997).
Hsu, T., McRackan, D., Vincent, T. S. & Gert de Couet, H. Drosophila Pin1 prolyl isomerase Dodo is a MAP kinase signal responder during oogenesis. Nat. Cell Biol. 3, 538–543 (2001).
Toko, H. et al. Differential regulation of cellular senescence and differentiation by prolyl isomerase Pin1 in cardiac progenitor cells. J. Biol. Chem. 289, 5348–5356 (2014).
Pinton, P. et al. Protein kinase C beta and prolyl isomerase 1 regulate mitochondrial effects of the life-span determinant p66Shc. Science 315, 659–663 (2007).
Ryo, A. et al. Prolyl-isomerase Pin1 accumulates in lewy bodies of parkinson disease and facilitates formation of alpha-synuclein inclusions. J. Biol. Chem. 281, 4117–4125 (2006).
Pastorino, L. et al. The prolyl isomerase Pin1 regulates amyloid precursor protein processing and amyloid-beta production. Nature 440, 528–534 (2006).
Carnemolla, A., Michelazzi, S. & Agostoni, E. PIN1 modulates Huntingtin levels and aggregate accumulation: an in vitro model. Front. Cell Neurosci. 11, 121 (2017).
Lu, Z. & Hunter, T. Prolyl isomerase Pin1 in cancer. Cell Res. 24, 1033–1049 (2014).
Luo, M. L. et al. Prolyl isomerase Pin1 acts downstream of miR200c to promote cancer stem-like cell traits in breast cancer. Cancer Res. 74, 3603–3616 (2014).
Rustighi, A. et al. Prolyl-isomerase Pin1 controls normal and cancer stem cells of the breast. EMBO Mol. Med. 6, 99–119 (2014).
Girardini, J. E. et al. A Pin1/mutant p53 axis promotes aggressiveness in breast cancer. Cancer Cell 20, 79–91 (2011).
Wulf, G., Garg, P., Liou, Y. C., Iglehart, D. & Lu, K. P. Modeling breast cancer in vivo and ex vivo reveals an essential role of Pin1 in tumorigenesis. EMBO J. 23, 3397–3407 (2004).
D’Artista, L. et al. Pin1 is required for sustained B cell proliferation upon oncogenic activation of Myc. Oncotarget 7, 21786–21798 (2016).
Wei, S. et al. Active Pin1 is a key target of all-trans retinoic acid in acute promyelocytic leukemia and breast cancer. Nat. Med. 21, 457–466 (2015).
Kim, M. R. et al. Enhancement of vascular endothelial growth factor-mediated angiogenesis in tamoxifen-resistant breast cancer cells: role of Pin1 overexpression. Mol. Cancer Ther. 8, 2163–2171 (2009).
Campaner, E. et al. A covalent PIN1 inhibitor selectively targets cancer cells by a dual mechanism of action. Nat. Commun. 8, 15772 (2017).
Hanahan, D. & Weinberg, R. A. The hallmarks of cancer. Cell 100, 57–70 (2000).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Wang, Y. et al. N6-methyladenosine RNA modification regulates embryonic neural stem cell self-renewal through histone modifications. Nat. Neurosci. 21, 195–206 (2018).
Gu, Y., Lindner, J., Kumar, A., Yuan, W. & Magnuson, M. A. Rictor/mTORC2 is essential for maintaining a balance between beta-cell proliferation and cell size. Diabetes 60, 827–837 (2011).
Espana-Agusti, J., Warren, A., Chew, S. K., Adams, D. J. & Matakidou, A. Loss of PBRM1 rescues VHL dependent replication stress to promote renal carcinogenesis. Nat. Commun. 8, 2026 (2017).
Raleigh, D. R. et al. Hedgehog signaling drives medulloblastoma growth via CDK6. J. Clin. Invest. 128, 120–124 (2018).
Lu, K. P., Hanes, S. D. & Hunter, T. A human peptidyl-prolyl isomerase essential for regulation of mitosis. Nature 380, 544–547 (1996).
Lin, C. H. et al. Landscape of Pin1 in the cell cycle. Exp. Biol. Med. (Maywood) 240, 403–408 (2015).
Anderson, E. The role of oestrogen and progesterone receptors in human mammary development and tumorigenesis. Breast Cancer Res. 4, 197–201 (2002).
Rajbhandari, P. et al. Regulation of estrogen receptor α N-terminus conformation and function by peptidyl prolyl isomerase Pin1. Mol. Cell Biol. 32, 445–457 (2012).
Rajbhandari, P., Ozers, M. S., Solodin, N. M., Warren, C. L. & Alarid, E. T. Peptidylprolyl isomerase Pin1 directly enhances the DNA binding functions of estrogen receptor α. J. Biol. Chem. 290, 13749–13762 (2015).
Rajbhandari, P. et al. Pin1 modulates ERα levels in breast cancer through inhibition of phosphorylation-dependent ubiquitination and degradation. Oncogene 33, 1438–1447 (2014).
Hahm, E. R. & Singh, S. V. Diallyl trisulfide inhibits estrogen receptor-α activity in human breast cancer cells. Breast Cancer Res. Treat. 144, 47–57 (2014).
Lam, P. B. et al. Prolyl isomerase Pin1 is highly expressed in Her2-positive breast cancer and regulates erbB2 protein stability. Mol. Cancer 7, 91 (2008).
Stanya, K. J., Liu, Y., Means, A. R. & Kao, H. Y. Cdk2 and Pin1 negatively regulate the transcriptional corepressor SMRT. J. Cell Biol. 183, 49–61 (2008).
Ryo, A. et al. Regulation of NF-kappaB signaling by Pin1-dependent prolyl isomerization and ubiquitin-mediated proteolysis of p65/RelA. Mol. Cell 12, 1413–1426 (2003).
Fan, G. et al. Peptidyl-prolyl isomerase Pin1 markedly enhances the oncogenic activity of the rel proteins in the nuclear factor-kappaB family. Cancer Res. 69, 4589–4597 (2009).
Atkinson, G. P. et al. The prolyl isomerase Pin1 regulates the NF-kappaB signaling pathway and interleukin-8 expression in glioblastoma. Oncogene 28, 3735–3745 (2009).
Saegusa, M., Hashimura, M. & Kuwata, T. Pin1 acts as a modulator of cell proliferation through alteration in NF-κB but not β-catenin/TCF4 signalling in a subset of endometrial carcinoma cells. J. Pathol. 222, 410–420 (2010).
Chen, J. Y. et al. The oncometabolite R-2-hydroxyglutarate activates NF-κB-dependent tumor-promoting stromal niche for acute myeloid leukemia cells. Sci. Rep. 6, 32428 (2016).
Shinoda, K. et al. Pin1 facilitates NF-κB activation and promotes tumour progression in human hepatocellular carcinoma. Br. J. Cancer 113, 1323–1331 (2015).
Murray-Zmijewski, F., Lane, D. P. & Bourdon, J. C. p53/p63/p73 isoforms: an orchestra of isoforms to harmonise cell differentiation and response to stress. Cell Death Differ. 13, 962–972 (2006).
Li, C. et al. Pin1 modulates p63α protein stability in regulation of cell survival, proliferation and tumor formation. Cell Death Dis. 4, e943 (2013).
Hu, X. et al. Prolyl isomerase PIN1 regulates the stability, transcriptional activity and oncogenic potential of BRD4. Oncogene 36, 5177–5188 (2017).
Zhu, Z. et al. Pin1 promotes prostate cancer cell proliferation and migration through activation of Wnt/β-catenin signaling. Clin. Transl. Oncol. 18, 792–797 (2016).
Kruiswijk, F. et al. Targeted inhibition of metastatic melanoma through interference with Pin1-FOXM1 signaling. Oncogene 35, 2166–2177 (2016).
Chae, U. et al. Critical role of XBP1 in cancer signalling is regulated by PIN1. Biochem. J. 473, 2603–2610 (2016).
Chen, H. Z. et al. Prolyl isomerase Pin1 stabilizes and activates orphan nuclear receptor TR3 to promote mitogenesis. Oncogene 31, 2876–2887 (2012).
Csizmok, V. et al. Multivalent interactions with Fbw7 and Pin1 facilitate recognition of c-Jun by the SCFFbw7 ubiquitin ligase. Structure 26, 28–39 (2018).
Helander, S. et al. Pre-anchoring of Pin1 to unphosphorylated c-Myc in a fuzzy complex regulates c-Myc activity. Structure 23, 2267–2279 (2015).
Nakatsu, Y. et al. The prolyl isomerase Pin1 increases β-cell proliferation and enhances insulin secretion. J. Biol. Chem. 292, 11886–11895 (2017).
Risal, P., Shrestha, N., Chand, L., Sylvester, K. G. & Jeong, Y. J. Involvement of prolyl isomerase PIN1 in the cell cycle progression and proliferation of hepatic oval cells. Pathol. Res. Pract. 213, 373–380 (2017).
Kanatsu-Shinohara, M., Onoyama, I., Nakayama, K. I. & Shinohara, T. Skp1-Cullin-F-box (SCF)-type ubiquitin ligase FBXW7 negatively regulates spermatogonial stem cell self-renewal. Proc. Natl Acad. Sci. USA 111, 8826–8831 (2014).
Avni, D. et al. Active localization of the retinoblastoma protein in chromatin and its response to S phase DNA damage. Mol. Cell 12, 735–746 (2003).
Valenciaga, A. et al. Reduced retinoblastoma protein expression is associated with decreased patient survival in medullary thyroid cancer. Thyroid 27, 1523–1533 (2017).
Ying, L. et al. Chronic inflammation promotes retinoblastoma protein hyperphosphorylation and E2F1 activation. Cancer Res. 65, 9132–9136 (2005).
You, H. et al. IGF-1 induces Pin1 expression in promoting cell cycle S-phase entry. J. Cell Biochem. 84, 211–216 (2002).
Rizzolio, F. et al. Retinoblastoma tumor-suppressor protein phosphorylation and inactivation depend on direct interaction with Pin1. Cell Death Differ. 19, 1152–1161 (2012).
Tong, Y. et al. Pin1 inhibits PP2A-mediated Rb dephosphorylation in regulation of cell cycle and S-phase DNA damage. Cell Death Dis. 6, e1640 (2015).
Lim, J. H., Liu, Y., Reineke, E. & Kao, H. Y. Mitogen-activated protein kinase extracellular signal-regulated kinase 2 phosphorylates and promotes Pin1 protein-dependent promyelocytic leukemia protein turnover. J. Biol. Chem. 286, 44403–44411 (2011).
Yuan, W. C. et al. A Cullin3-KLHL20 ubiquitin ligase-dependent pathway targets PML to potentiate HIF-1 signaling and prostate cancer progression. Cancer Cell. 20, 214–228 (2011).
Liu, T., Liu, Y., Kao, H. Y. & Pei, D. Membrane permeable cyclic peptidyl inhibitors against human peptidylprolyl isomerase Pin1. J. Med. Chem. 53, 2494–2501 (2010).
Gianni, M. et al. Inhibition of the peptidyl-prolyl-isomerase Pin1 enhances the responses of acute myeloid leukemia cells to retinoic acid via stabilization of RARalpha and PML-RARalpha. Cancer Res. 69, 1016–1026 (2009).
Huang, B. et al. RUNX3 acts as a tumor suppressor in breast cancer by targeting estrogen receptor α. Oncogene 31, 527–534 (2012).
Nicole Tsang, Y. H. et al. Prolyl isomerase Pin1 downregulates tumor suppressor RUNX3 in breast cancer. Oncogene 32, 1488–1496 (2013).
Min, S. H. et al. Negative regulation of the stability and tumor suppressor function of Fbw7 by the Pin1 prolyl isomerase. Mol. Cell 46, 771–783 (2012).
Hwang, Y. C. et al. Destabilization of KLF10, a tumor suppressor, relies on thr93 phosphorylation and isomerase association. Biochim. Biophys. Acta 1833, 3035–3045 (2013).
Khanal, P. et al. Prolyl isomerase Pin1 negatively regulates the stability of SUV39H1 to promote tumorigenesis in breast cancer. FASEB J. 27, 4606–4618 (2013).
Khanal, P. et al. Proyl isomerase Pin1 facilitates ubiquitin-mediated degradation of cyclin-dependent kinase 10 to induce tamoxifen resistance in breast cancer cells. Oncogene 31, 3845–3856 (2012).
Sorrentino, G. et al. The prolyl-isomerase Pin1 activates the mitochondrial death program of p53. Cell Death Differ. 20, 198–p208 (2013).
Follis, A. V. et al. Pin1-induced proline isomerization in cytosolic p53 mediates BAX activation and apoptosis. Mol. Cell 59, 677–684 (2015).
Bae, J. S. et al. PIN1 in hepatocellular carcinoma is associated with TP53 gene status. Oncol. Rep. 36, 2405–2411 (2016).
Liao, P. et al. Mutant p53 gains its function via c-Myc activation upon CDK4 phosphorylation at serine 249 and consequent PIN1 binding. Mol. Cell 68, 1134–1146 (2017).
Hu, H. & Wulf, G. M. The amplifier effect: how Pin1 empowers mutant p53. Breast Cancer Res. 13, 315 (2011).
Chen, S. Y. et al. Activation of beta-catenin signaling in prostate cancer by peptidyl-prolyl isomerase Pin1-mediated abrogation of the androgen receptor-beta-catenin interaction. Mol. Cell Biol. 26, 929–939 (2006).
He, J. et al. Overexpression of Pin1 in non-small cell lung cancer (NSCLC) and its correlation with lymph node metastases. Lung Cancer 56, 51–58 (2007).
Derynck, R., Akhurst, R. J. & Balmain, A. TGF-beta signaling in tumor suppression and cancer progression. Nat. Genet. 29, 117–129 (2001).
Massagué, J. TGFbeta in cancer. Cell 134, 215–230 (2008).
Shen, Z. J. et al. Pin1 protein regulates Smad protein signaling and pulmonary fibrosis. J. Biol. Chem. 287, 23294–23305 (2012).
Aragón, E. et al. A Smad action turnover switch operated by WW domain readers of a phosphoserine code. Genes Dev. 25, 1275–1288 (2011).
Nakano, A. et al. Pin1 down-regulates transforming growth factor-beta (TGF-beta) signaling by inducing degradation of Smad proteins. J. Biol. Chem. 284, 6109–6115 (2009).
Matsuura, I. et al. Pin1 promotes transforming growth factor-beta-induced migration and invasion. J. Biol. Chem. 285, 1754–1764 (2010).
Thomas, A. L. et al. Inhibition of CDK-mediated Smad3 phosphorylation reduces the Pin1-Smad3 interaction and aggressiveness of triple negative breast cancer cells. Cell Cycle 16, 1453–1464 (2017).
Franciosa, G. et al. Prolyl-isomerase Pin1 controls Notch3 protein expression and regulates T-ALL progression. Oncogene 35, 4741–4751 (2016).
Lee, K. J. et al. CD99-derived agonist ligands inhibit fibronectin-induced activation of β1 integrin through the protein kinase A/SHP2/extracellular signal-regulated kinase/PTPN12/focal adhesion kinase signaling pathway. Mol. Cell Biol. 37, e00675–16 (2017).
Zheng, Y. et al. FAK phosphorylation by ERK primes ras-induced tyrosine dephosphorylation of FAK mediated by PIN1 and PTP-PEST. Mol. Cell 35, 11–25 (2009).
Zheng, Y. et al. Ras-induced and extracellular signal-regulated kinase 1 and 2 phosphorylation-dependent isomerization of protein tyrosine phosphatase (PTP)-PEST by PIN1 promotes FAK dephosphorylation by PTP-PEST. Mol. Cell Biol. 31, 4258–4269 (2011).
Lufei, C., Koh, T. H., Uchida, T. & Cao, X. Pin1 is required for the Ser727 phosphorylation-dependent Stat3 activity. Oncogene 26, 7656–7664 (2007).
Wang, J. et al. FOXC1 regulates the functions of human basal-like breast cancer cells by activating NF-κB signaling. Oncogene 31, 4798–4802 (2012).
Muller, P. A. et al. Mutant p53 drives invasion by promoting integrin recycling. Cell 139, 1327–1341 (2009).
Frey, N., Venturelli, S., Zender, L. & Bitzer, M. Cellular senescence in gastrointestinal diseases: from pathogenesis to therapeutics. Nat. Rev. Gastroenterol. Hepatol. 15, 81–95 (2018).
Birch, J., Barnes, P. J. & Passos, J. F. Mitochondria, telomeres and cell senescence: Implications for lung ageing and disease. Pharmacol. Ther. 183, 34–49 (2018).
He, S. & Sharpless, N. E. Senescence in health and disease. Cell 169, 1000–1011 (2017).
Lv, L. et al. Downregulation of Pin1 in human atherosclerosis and its association with vascular smooth muscle cell senescence. J. Vasc. Surg. S0741-5214, 32213–32219 (2017).
Chen, L. et al. The role of Pin1 protein in aging of human tendon stem/progenitor cells. Biochem. Biophys. Res. Commun. 464, 487–492 (2015).
Wheaton, K., Muir, J., Ma, W. & Benchimol, S. BTG2 antagonizes Pin1 in response to mitogens and telomere disruption during replicative senescence. Aging Cell 9, 747–760 (2010).
van Steensel, B. & de Lange, T. Control of telomere length by the human telomeric protein TRF1. Nature 385, 740–743 (1997).
Lee, T. H. et al. Essential role of Pin1 in the regulation of TRF1 stability and telomere maintenance. Nat. Cell Biol. 11, 97–105 (2009).
Hsu, J. Y., Reimann, J. D., Sørensen, C. S., Lukas, J. & Jackson, P. K. E2F-dependent accumulation of hEmi1 regulates S phase entry by inhibiting APC(Cdh1). Nat. Cell Biol. 4, 358–366 (2002).
Bernis, C. et al. Pin1 stabilizes Emi1 during G2 phase by preventing its association with SCF(betatrcp). EMBO Rep. 8, 91–98 (2007).
Cheng, C. W., Leong, K. W. & Tse, E. Understanding the role of PIN1 in hepatocellular carcinoma. World J. Gastroenterol. 22, 9921–9932 (2016).
Pugh, C. W. & Ratcliffe, P. J. Regulation of angiogenesis by hypoxia: role of the HIF system. Nat. Med. 9, 677–684 (2003).
Lee, K. Y. et al. PI3-kinase/p38 kinase-dependent E2F1 activation is critical for Pin1 induction in tamoxifen-resistant breast cancer cells. Mol. Cells 32, 107–111 (2011).
Oh, S. J. et al. Inhibition of angiogenesis by quercetin in tamoxifen-resistant breast cancer cells. Food Chem. Toxicol. 48, 3227–3234 (2010).
Han, H. J. et al. Peptidyl prolyl isomerase PIN1 directly binds to and stabilizes hypoxia-inducible factor-1α. PLoS ONE 11, e147038 (2016).
Jalouli, M. et al. The prolyl isomerase Pin1 regulates hypoxia-inducible transcription factor (HIF) activity. Cell Signal. 26, 1649–1656 (2014).
Lin, Y. C. et al. SCP phosphatases suppress renal cell carcinoma by stabilizing PML and inhibiting mTOR/HIF signaling. Cancer Res. 74, 6935–6946 (2014).
Hsu, K. S. et al. Translational control of PML contributes to TNFα-induced apoptosis of MCF7 breast cancer cells and decreased angiogenesis in HUVECs. Cell Death Differ. 23, 469–483 (2016).
Wang, Z., Banerjee, S., Kong, D., Li, Y. & Sarkar, F. H. Down-regulation of Forkhead Box M1 transcription factor leads to the inhibition of invasion and angiogenesis of pancreatic cancer cells. Cancer Res. 67, 8293–8300 (2007).
Jiang, L. et al. Bach1 Represses Wnt/β-Catenin Signaling and Angiogenesis. Circ. Res. 117, 364–375 (2015).
Ryo, A. et al. Stable suppression of tumorigenicity by Pin1-targeted RNA interference in prostate cancer. Clin. Cancer Res. 11, 7523–7531 (2005).
Kim, J. A. et al. Amurensin G inhibits angiogenesis and tumor growth of tamoxifen-resistant breast cancer via Pin1 inhibition. Food Chem. Toxicol. 50, 3625–3634 (2012).
Evan, G. & Littlewood, T. A matter of life and cell death. Science 281, 1317–1322 (1998).
Lowe, S. W., Cepero, E. & Evan, G. Intrinsic tumour suppression. Nature 432, 307–315 (2004).
Polonio-Vallon, T., Krüger, D. & Hofmann, T. G. ShaPINg cell fate upon DNA damage: role of Pin1 isomerase in DNA damage-induced cell death and repair. Front. Oncol. 4, 148 (2014).
Shen, Z. J. & Malter, J. S. Determinants of eosinophil survival and apoptotic cell death. Apoptosis 20, 224–234 (2015).
Shen, Z. J., Esnault, S., Schinzel, A., Borner, C. & Malter, J. S. The peptidyl-prolyl isomerase Pin1 facilitates cytokine-induced survival of eosinophils by suppressing Bax activation. Nat. Immunol. 10, 257–265 (2009).
Yang, X., Khosravi-Far, R., Chang, H. Y. & Baltimore, D. Daxx, a novel Fas-binding protein that activates JNK and apoptosis. Cell 89, 1067–1076 (1997).
Helmke, C. et al. Ligand stimulation of CD95 induces activation of Plk3 followed by phosphorylation of caspase-8. Cell Res. 26, 914–934 (2016).
Oh, J. & Malter, J. S. Pin1-FADD interactions regulate Fas-mediated apoptosis in activated eosinophils. J. Immunol. 190, 4937–4945 (2013).
Ryo, A. et al. A suppressive role of the prolyl isomerase Pin1 in cellular apoptosis mediated by the death-associated protein Daxx. J. Biol. Chem. 282, 36671–36681 (2007).
Green, D. R. & Reed, J. C. Mitochondria and apoptosis. Science 281, 1309–1312 (1998).
Ding, Q. et al. Down-regulation of myeloid cell leukemia-1 through inhibiting Erk/Pin 1 pathway by sorafenib facilitates chemosensitization in breast cancer. Cancer Res. 68, 6109–6117 (2008).
Basu, A. & Haldar, S. Signal-induced site specific phosphorylation targets Bcl2 to the proteasome pathway. Int. J. Oncol. 21, 597–601 (2002).
Lee, Y. C., Chen, Y. J., Huang, C. H. & Chang, L. S. Amsacrine-induced apoptosis of human leukemia U937 cells is mediated by the inhibition of AKT- and ERK-induced stabilization of MCL1. Apoptosis 22, 406–420 (2017).
Wang, T., Liu, Z., Shi, F. & Wang, J. Pin1 modulates chemo-resistance by up-regulating FoxM1 and the involvements of Wnt/β-catenin signaling pathway in cervical cancer. Mol. Cell Biochem. 413, 179–187 (2016).
Namgoong, G. M. et al. The prolyl isomerase Pin1 induces LC-3 expression and mediates tamoxifen resistance in breast cancer. J. Biol. Chem. 285, 23829–23841 (2010).
Cheng, C. W. et al. PIN1 inhibits apoptosis in hepatocellular carcinoma through modulation of the antiapoptotic function of survivin. Am. J. Pathol. 182, 765–775 (2013).
Dourlen, P. et al. The peptidyl prolyl cis/trans isomerase Pin1 downregulates the inhibitor of apoptosis protein Survivin. Biochim. Biophys. Acta 1773, 1428–1437 (2007).
Bitomsky, N. et al. Autophosphorylation and Pin1 binding coordinate DNA damage-induced HIPK2 activation and cell death. Proc. Natl Acad. Sci. USA 110, E4203–E4212 (2013).
Hilton, B. A. et al. ATR plays a direct antiapoptotic role at mitochondria, which is regulated by prolyl isomerase Pin1. Mol. Cell 60, 35–46 (2015).
Kim, R., Emi, M. & Tanabe, K. Cancer immunoediting from immune surveillance to immune escape. Immunology 121, 1–14 (2007).
Mahla, R. S., Reddy, M. C., Prasad, D. V. & Kumar, H. Sweeten PAMPs: role of sugar complexed PAMPs in innate immunity and vaccine biology. Front. Immunol. 4, 248 (2013).
Kwissa, M., Nakaya, H. I., Oluoch, H. & Pulendran, B. Distinct TLR adjuvants differentially stimulate systemic and local innate immune responses in nonhuman primates. Blood 119, 2044–2055 (2012).
Cao, Z., Henzel, W. J. & Gao, X. IRAK: a kinase associated with the interleukin-1 receptor. Science 271, 1128–1131 (1996).
Honda, K. et al. IRF-7 is the master regulator of type-I interferon-dependent immune responses. Nature 434, 772–777 (2005).
Tun-Kyi, A. et al. Essential role for the prolyl isomerase Pin1 in Toll-like receptor signaling and type I interferon-mediated immunity. Nat. Immunol. 12, 733–741 (2011).
Wei, S. et al. Pin1-targeted therapy for systemic lupus erythematosus. Arthritis Rheumatol. 68, 2503–2513 (2016).
Huai, W. et al. Mint3 potentiates TLR3/4- and RIG-I-induced IFN-β expression and antiviral immune responses. Proc. Natl Acad. Sci. USA 113, 11925–11930 (2016).
Saitoh, T. et al. Negative regulation of interferon-regulatory factor 3-dependent innate antiviral response by the prolyl isomerase Pin1. Nat. Immunol. 7, 598–605 (2006).
Yang, K. et al. TRIM21 is essential to sustain IFN regulatory factor 3 activation during antiviral response. J. Immunol. 182, 3782–3792 (2009).
El Asmi, F. et al. Implication of PMLIV in both intrinsic and innate immunity. PLoS Pathog. 10, e1003975 (2014).
Osterlund, P. I., Pietilä, T. E., Veckman, V., Kotenko, S. V. & Julkunen, I. IFN regulatory factor family members differentially regulate the expression of type III IFN (IFN-lambda) genes. J. Immunol. 179, 3434–3442 (2007).
Munn, D. H. & Mellor, A. L. Indoleamine 2,3 dioxygenase and metabolic control of immune responses. Trends Immunol. 34, 137–143 (2013).
Koorella, C. et al. Novel regulation of CD80/CD86-induced phosphatidylinositol 3-kinase signaling by NOTCH1 protein in interleukin-6 and indoleamine 2,3-dioxygenase production by dendritic cells. J. Biol. Chem. 289, 7747–7762 (2014).
Chen, W. & Ten Dijke, P. Immunoregulation by members of the TGFβ superfamily. Nat. Rev. Immunol. 16, 723–740 (2016).
Yang, J. W. et al. Pin1 induction in the fibrotic liver and its roles in TGF-β1 expression and Smad2/3 phosphorylation. J. Hepatol. 60, 1235–1241 (2014).
Shen, Z. J. et al. Pin1 regulates TGF-beta1 production by activated human and murine eosinophils and contributes to allergic lung fibrosis. J. Clin. Invest. 118, 479–490 (2008).
Liu, Y. et al. Manipulating PML SUMOylation via silencing UBC9 and RNF4 regulates cardiac fibrosis. Mol. Ther. 25, 666–678 (2017).
Efremova, M. et al. Targeting immune checkpoints potentiates immunoediting and changes the dynamics of tumor evolution. Nat. Commun. 9, 32 (2018).
Pasparakis, M. & Vandenabeele, P. Necroptosis and its role in inflammation. Nature 517, 311–320 (2015).
Sfanos, K. S., Yegnasubramanian, S., Nelson, W. G. & De Marzo, A. M. The inflammatory microenvironmentand microbiome in prostate cancer development. Nat. Rev. Urol. 15, 11–24 (2017).
Atsumi, T. et al. Inflammation amplifier, a new paradigm in cancer biology. Cancer Res. 74, 8–14 (2014).
Leto, T. L. & Geiszt, M. Role of Nox family NADPH oxidases in host defense. Antioxid. Redox Signal. 8, 1549–1561 (2006).
Makni-Maalej, K. et al. The TLR7/8 agonist CL097 primes N-formyl-methionyl-leucyl-phenylalanine-stimulated NADPH oxidase activation in human neutrophils: critical role of p47phox phosphorylation and the proline isomerase Pin1. J. Immunol. 189, 4657–4665 (2012).
Boussetta, T. et al. The prolyl isomerase Pin1 acts as a novel molecular switch for TNF-alpha-induced priming of the NADPH oxidase in human neutrophils. Blood 116, 5795–5802 (2010).
Xing, Z., Ohkawara, Y., Jordana, M., Graham, F. & Gauldie, J. Transfer of granulocyte-macrophage colony-stimulating factor gene to rat lung induces eosinophilia, monocytosis, and fibrotic reactions. J. Clin. Invest. 97, 1102–1110 (1996).
DeMaria, C. T. & Brewer, G. AUF1 binding affinity to A+U-rich elements correlates with rapid mRNA degradation. J. Biol. Chem. 271, 12179–12184 (1996).
Shen, Z. J., Esnault, S. & Malter, J. S. The peptidyl-prolyl isomerase Pin1 regulates the stability of granulocyte-macrophage colony-stimulating factor mRNA in activated eosinophils. Nat. Immunol. 6, 1280–1287 (2005).
Esnault, S., Shen, Z. J., Whitesel, E. & Malter, J. S. The peptidyl-prolyl isomerase Pin1 regulates granulocyte-macrophage colony-stimulating factor mRNA stability in T lymphocytes. J. Immunol. 177, 6999–7006 (2006).
Esnault, S. et al. A critical role for Pin1 in allergic pulmonary eosinophilia in rats. J. Allergy Clin. Immunol. 120, 1082–1088 (2007).
Kim, K. et al. Interleukin-22 promotes epithelial cell transformation and breast tumorigenesis via MAP3K8 activation. Carcinogenesis 35, 1352–1361 (2014).
Nechama, M. et al. The IL-33-PIN1-IRAK-M axis is critical for type 2 immunity in IL-33-induced allergic airway inflammation. Nat. Commun. 9, 1603 (2018).
Jeong, H. G. et al. Novel role of Pin1 induction in type II collagen-mediated rheumatoid arthritis. J. Immunol. 183, 6689–6697 (2009).
Cho, Y. A. et al. PIN1 inhibition suppresses osteoclast differentiation and inflammatory responses. J. Dent. Res. 94, 371–380 (2015).
Paneni, F. et al. Targeting prolyl-isomerase Pin1 prevents mitochondrial oxidative stress and vascular dysfunction: insights in patients with diabetes. Eur. Heart J. 36, 817–828 (2015).
Nakatsu, Y. et al. Role of Pin1 protein in the pathogenesis of nonalcoholic steatohepatitis in a rodent model. J. Biol. Chem. 287, 44526–44535 (2012).
Asuri, S. et al. Primary biliary cholangitis in British Columbia First Nations: clinical features and discovery of novel genetic susceptibility loci. Liver. Int. 38, 940–948 (2018).
van Tiel, C. M., Kurakula, K., Koenis, D. S., van der Wal, E. & de Vries, C. J. Dual function of Pin1 in NR4A nuclear receptor activation: enhanced activity of NR4As and increased Nur77 protein stability. Biochim. Biophys. Acta 1823, 1894–1904 (2012).
Poolman, T. M., Farrow, S. N., Matthews, L., Loudon, A. S. & Ray, D. W. Pin1 promotes GR transactivation by enhancing recruitment to target genes. Nucleic Acids Res. 41, 8515–8525 (2013).
Yu, L. et al. Modeling the genetic regulation of cancer metabolism: interplay between glycolysis and oxidative phosphorylation. Cancer Res. 77, 1564–1574 (2017).
Vander Heiden, M. G., Cantley, L. C. & Thompson, C. B. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324, 1029–1033 (2009).
Lee, N. & Kim, D. Cancer metabolism: fueling more than just growth. Mol. Cells 39, 847–854 (2016).
Wang, C. W., Purkayastha, A., Jones, K. T., Thaker, S. K. & Banerjee, U. In vivo genetic dissection of tumor growth and the Warburg effect. eLife 5, e18126 (2016).
Pate, K. T. et al. Wnt signaling directs a metabolic program of glycolysis and angiogenesis in colon cancer. EMBO J. 33, 1454–1473 (2014).
Liu, Z. et al. Nutrient deprivation-related OXPHOS/glycolysis interconversion via HIF-1α/C-MYC pathway in U251 cells. Tumour Biol. 37, 6661–6671 (2016).
Fan, C. et al. Role of long non-coding RNAs in glucose metabolism in cancer. Mol. Cancer 16, 130 (2017).
Yang, W. et al. ERK1/2-dependent phosphorylation and nuclear translocation of PKM2 promotes the Warburg effect. Nat. Cell Biol. 14, 1295–1304 (2012).
Yang, W. et al. Nuclear PKM2 regulates β-catenin transactivation upon EGFR activation. Nature 480, 118–122 (2011).
Hitosugi, T. et al. Tyrosine phosphorylation of mitochondrial pyruvate dehydrogenase kinase 1 is important for cancer metabolism. Mol. Cell 44, 864–877 (2011).
Li, X. et al. Mitochondria-translocated PGK1 functions as a protein kinase to coordinate glycolysis and the TCA cycle in tumorigenesis. Mol. Cell 61, 705–719 (2016).
van der Mijn, J. C., Panka, D. J., Geissler, A. K., Verheul, H. M. & Mier, J. W. Novel drugs that target the metabolic reprogramming in renal cell cancer. Cancer Metab. 4, 14 (2016).
Jackson, S. P. & Bartek, J. The DNA-damage response in human biology and disease. Nature 461, 1071–1078 (2009).
Matt, S. & Hofmann, T. G. The DNA damage-induced cell death response: a roadmap to kill cancer cells. Cell Mol. Life Sci. 73, 2829–2850 (2016).
Heyer, W. D., Ehmsen, K. T. & Liu, J. Regulation of homologous recombination in eukaryotes. Annu. Rev. Genet. 44, 113–139 (2010).
Sartori, A. A. et al. Human CtIP promotes DNA end resection. Nature 450, 509–514 (2007).
Steger, M. et al. Prolyl isomerase PIN1 regulates DNA double-strand break repair by counteracting DNA end resection. Mol. Cell 50, 333–343 (2013).
Ferretti, L. P., Lafranchi, L. & Sartori, A. A. Controlling DNA-end resection: a new task for CDKs. Front. Genet. 4, 99 (2013).
Burrell, R. A. et al. Replication stress links structural and numerical cancer chromosomal instability. Nature 494, 492–496 (2013).
Waizenegger, I. C., Hauf, S., Meinke, A. & Peters, J. M. Two distinct pathways remove mammalian cohesin from chromosome arms in prophase and from centromeres in anaphase. Cell 103, 399–410 (2000).
Hellmuth, S. et al. Human chromosome segregation involves multi-layered regulation of separase by the peptidyl-prolyl-isomerase Pin1. Mol. Cell 58, 495–506 (2015).
Fabbro, M. et al. Cdk1/Erk2- and Plk1-dependent phosphorylation of a centrosome protein, Cep55, is required for its recruitment to midbody and cytokinesis. Dev. Cell 9, 477–488 (2005).
van der Horst, A. & Khanna, K. K. The peptidyl-prolyl isomerase Pin1 regulates cytokinesis through Cep55. Cancer Res. 69, 6651–6659 (2009).
Estey, M. P. et al. Mitotic regulation of SEPT9 protein by cyclin-dependent kinase 1 (Cdk1) and Pin1 protein is important for the completion of cytokinesis. J. Biol. Chem. 288, 30075–30086 (2013).
Suizu, F., Ryo, A., Wulf, G., Lim, J. & Lu, K. P. Pin1 regulates centrosome duplication, and its overexpression induces centrosome amplification, chromosome instability, and oncogenesis. Mol. Cell Biol. 26, 1463–1479 (2006).
Duensing, S., Duensing, A., Crum, C. P. & Münger, K. Human papillomavirus type 16 E7 oncoprotein-induced abnormal centrosome synthesis is an early event in the evolving malignant phenotype. Cancer Res. 61, 2356–2360 (2001).
Prabhavathy, D., Vijayalakshmi, R., Kanchana, M. P. & Karunagaran, D. HPV16 E2 enhances the expression of NF-κB and STAT3 target genes and potentiates NF-κB activation by inflammatory mediators. Cell Immunol. 292, 70–77 (2014).
Dean, M., Fojo, T. & Bates, S. Tumour stem cells and drug resistance. Nat. Rev. Cancer 5, 275–284 (2005).
Lee, Y. C. et al. Pin1 acts as a negative regulator of the G2/M transition by interacting with the Aurora-A-Bora complex. J. Cell Sci. 126, 4862–4872 (2013).
Cho, Y. S. et al. TPA-induced cell transformation provokes a complex formation between Pin1 and 90 kDa ribosomal protein S6 kinase 2. Mol. Cell Biochem. 367, 85–92 (2012).
Del Rosario, J. S. et al. Death associated protein kinase (DAPK)-mediated neurodegenerative mechanisms in nematode excitotoxicity. BMC Neurosci. 16, 25 (2015).
Zhang, X., Zhang, B., Gao, J., Wang, X. & Liu, Z. Regulation of the microRNA 200b (miRNA-200b) by transcriptional regulators PEA3 and ELK-1 protein affects expression of Pin1 protein to control anoikis. J. Biol. Chem. 288, 32742–32752 (2013).
Lee, K. H. et al. MicroRNA-296-5p (miR-296-5p) functions as a tumor suppressor in prostate cancer by directly targeting Pin1. Biochim. Biophys. Acta 1843, 2055–2066 (2014).
Li, J., et al. Pin1 impairs microRNA biogenesis by mediating conformation change of XPO5 in hepatocellular carcinoma. Cell Death Differ. https://doi.org/10.1038/s41418-018-0065-z (2018).
Yang, D. et al. A novel controlled release formulation of the Pin1 inhibitor ATRA to improve liver cancer therapy by simultaneously blocking multiple cancer pathways. J. Control Release 269, 405–422 (2018).
Marsolier, J. et al. Theileria parasites secrete a prolyl isomerase to maintain host leukocyte transformation. Nature 520, 378–382 (2015).
Wang, X. J., Xu, B., Mullins, A. B., Neiler, F. K. & Etzkorn, F. A. Conformationally locked isostere of phosphoSer-cis-Pro inhibits Pin1 23-fold better than phosphoSer-trans-Pro isostere. J. Am. Chem. Soc. 126, 15533–15542 (2004).
Liu, C., Mu, C., Li, Z. & Xu, L. Imazamethabenz inhibits human breast cancer cell proliferation, migration and invasion via combination with Pin1. Mol. Med. Rep. 15, 3210–3214 (2017).
Pu, W., et al. Targeting Pin1 by inhibitor API-1 regulates microRNA biogenesis and suppresses hepatocellular carcinoma development. Hepatology https://doi.org/10.1002/hep.29819 (2018)
Zheng, M. et al. Inhibition of the prolyl isomerase Pin1 enhances the ability of sorafenib to induce cell death and inhibit tumor growth in hepatocellular carcinoma. Oncotarget 8, 29771–29784 (2017).
Sajadimajd, S. & Yazdanparast, R. Sensitizing effect of juglone is mediated by down regulation of Notch1 signaling pathway in trastuzumab-resistant SKBR3 cells. Apoptosis 22, 135–144 (2017).
Uchida, T. et al. Pin1 and Par14 peptidyl prolyl isomerase inhibitors block cell proliferation. Chem. Biol. 10, 15–24 (2003).
Guo, C. et al. Structure-based design of novel human Pin1 inhibitors (I). Bioorg. Med. Chem. Lett. 19, 5613–5616 (2009).
Dong, L. et al. Structure-based design of novel human Pin1 inhibitors (II). Bioorg. Med. Chem. Lett. 20, 2210–2214 (2010).
Zhang, Y. et al. Structural basis for high-affinity peptide inhibition of human Pin1. ACS Chem. Biol. 2, 320–328 (2007).
Potter, A. J. et al. Structure-guided design of alpha-amino acid-derived Pin1 inhibitors. Bioorg. Med. Chem. Lett. 20, 586–590 (2010).
Urusova, D. V. et al. Epigallocatechin-gallate suppresses tumorigenesis by directly targeting Pin1. Cancer Prev. Res. (Phila.) 4, 1366–1377 (2011).
Cui, G. et al. Synthesis and biological evaluation of pyrimidine derivatives as novel human Pin1 inhibitors. Bioorg. Med. Chem. 26, 2186–2197 (2018).
Bedewy, W. et al. Generation of a cell-permeable cycloheptapeptidyl inhibitor against the peptidyl-prolyl isomerase Pin1. Org. Biomol. Chem. 15, 4540–4543 (2017).
Lim, T. G. et al. The prolyl isomerase Pin1 is a novel target of 6,7,4′-trihydroxyisoflavone for suppressing esophageal cancer growth. Cancer Prev. Res. (Phila.) 10, 308–318 (2017).
Cho, J. H., Chae, J. I. & Shim, J. H. Rhein exhibits antitumorigenic effects by interfering with the interaction between prolyl isomerase Pin1 and c-Jun. Oncol. Rep. 37, 1865–1872 (2017).
Zhao, H., Cui, G., Jin, J., Chen, X. & Xu, B. Synthesis and Pin1 inhibitory activity of thiazole derivatives. Bioorg. Med. Chem. 24, 5911–5920 (2016).
Wu, K. J. et al. Small molecule Pin1 inhibitor blocking NF-κB signaling in prostate cancer cells. Chem. Asian J. 13, 275–279 (2017).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 81702441).
Author information
Authors and Affiliations
Contributions
Y.-b.Y. and S.-m.Y. conceived and designed this work and gave their professional criticism and suggestion. Y.-r.W., X.-z.L., and C.-j.H. edited the manuscript and figures. Y.C. and Y.-r.W. drafted the manuscript. H.-y.Y., M.-m.J., and Y.-y.W. collected data. X.-z.L. and C.-j.H. revised the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by I. Amelio
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, Y., Wu, Yr., Yang, Hy. et al. Prolyl isomerase Pin1: a promoter of cancer and a target for therapy. Cell Death Dis 9, 883 (2018). https://doi.org/10.1038/s41419-018-0844-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-018-0844-y
This article is cited by
-
Non-POU Domain-Containing Octomer-Binding (NONO) protein expression and stability promotes the tumorigenicity and activation of Akt/MAPK/β-catenin pathways in human breast cancer cells
Cell Communication and Signaling (2023)
-
METTL3 stabilization by PIN1 promotes breast tumorigenesis via enhanced m6A-dependent translation
Oncogene (2023)
-
Expression, Purification, Structural and Functional Characterization of Recombinant Human Parvulin 17
Molecular Biotechnology (2023)
-
Unconventional protein post-translational modifications: the helmsmen in breast cancer
Cell & Bioscience (2022)
-
Rationale for MYC imaging and targeting in pancreatic cancer
EJNMMI Research (2021)