Abstract
Oxidative stress-induced DNA damage has been well acknowledged as a major cause leading to cell death, which is etiologically linked to ischemic injury and degenerative alterations. The most common oxidation product of DNA is base lesion 8-oxo-7,8-dihydroguanine (8-oxoG), which is repaired by 8-oxoG glycosylase1 (OGG1)-initiated baseexcision repair (BER) pathway (OGG1-BER); however, the role of OGG1-BER in oxidative stress-induced cell death is poorly investigated. DNA strand breaks and apurinic/apyrimidinic (AP) sites are effective substrates to activate DNA damage sensor poly(ADP-ribose) polymerase 1 (PARP1). Overactivation of PARP1 is associated with apoptosis-inducing factor (AIF)-mediated and caspase-independent cell death (parthanatos). We hypothesized that after an excessive oxidative insult, OGG1-BER-generated strand breaks result in hyperactivation of PARP1 and consequently cell death. To test, wild type, knockout, siRNA-depleted MEFs and neuroblastoma cells, or those expressing repair-deficient OGG1 mutants were oxidatively stressed and the role of OGG1 was examined. Results showed that OGG1-BER further increases the levels of ROS-induced DNA damage by generating repair intermediates, leading to PARP1 overactivation and cell death. Cells lacking or expressing repair-deficient OGG1 showed lower levels of DNA strand lesions, PARP1 activation, and nuclear translocation of apoptosis-inducing factor, resulting in the increased resistance to ROS-induced parthanatos. These results suggested that OGG1 guards genome integrity through either lesion repair or elimination of cells with malignant potential, to maintain the homeostasis of the host, which might depend on the magnitude of guanine oxidation.
Similar content being viewed by others
Introduction
Oxidative stress is referred to elevated intracellular level of reactive oxygen species (ROS) that inevitably derive from various endogenous physiological processes, which can be exacerbated by environmental exposures1. ROS cause oxidative damage of lipids, proteins, and DNA, and to maintain genome integrity, DNA lesions ought to be repaired1,2. In the genomic DNA, one of the most common oxidation products is base lesion 8-oxo-7,8-dihydroguanine (8-oxoG), which is one of the best biomarker of oxidative stress3,4. 8-OxoG is mutagenic, and the cognate enzyme that specifically recognizes and repairs 8-oxoG and its open-ring product 2,6-diamino-4-hydroxy-5-formamidopyrimidine (FapyG) is 8-oxoguanine DNA glycosylase 1 (OGG1), a functional homolog of Escherichia coli protein MutM5,6. Base excision repair (BER) is a multistep process, which is described as a “hand-off” model, including lesion recognition, base excision, and strand cleavage, followed by recruitment of apurinic/apyrimidinic (AP) endonuclease 1 (APE1), DNA pol β, and DNA ligase III to the presumptive scaffold protein X-ray repair cross-complementing 1 (XRCC1) to complete the repair process7,8,9.
Oxidative stress-induced DNA damage has been well acknowledged as a major cause leading to cell death, which is etiologically linked to ischemic injury and degenerative alterations10,11. However, the role of OGG1-BER in oxidative stress-induced cell death is poorly investigated. Dr. Dawson’s group documented a distinct mode of cell death, namely, parthanatos, which is refered to as PARP1-dependent, apoptosis-inducing factor (AIF)-mediated and caspase-independent cell death12,13. Unlike typical apoptosis, parthanatos does not cause apoptotic body formation or small-scale DNA fragmentation. When apoptosis causes phosphatidylserine flipping onto the outer plasma membrane and manifests propidium iodide (PI) exclusion, parthanatos exhibits both annexin-V labeling and PI staining as positive14. As a DNA damage sensor, PARP1 can be activated via binding to both DNA breaks and AP sites15,16, and upon activation, PARP1 catalyzes the formation of polymerized ADP-ribose (PAR) from nicotinamide adenine dinucleotide (NAD+), and subsequently the covalent attachment of PAR polymers to target proteins. In turn, PAR polymer is removed from the target proteins by successively activated PAR glycohydrolase (PARG), forming the free PAR17. Acting as death messenger, free PAR is mainly produced in the nucleus and then redistributed to the cytoplasm and mitochondria, where it is critical for the release of apoptosis-inducing factor (AIF) from mitochondria and then its translocation into the nucleus18. AIF induces chromatin condensation and large-scale DNA fragmentation (~50 kb) leading to cell death12,13,18.
Parthanatos is linked to diseases including stroke, Parkinson’s disease, heart attack, diabetes, and ischemia reperfusion injury19,20, where the intracellular context is well-acknowledged to be highly perturbed by ROS, and guanines are supposed to be excessively oxidized4,21. Studies have documented that excitotoxic activation of N-methyl-d-aspartate (NMDA) receptor results in formation of ROS, which causes DNA damage, thereby the excessive PARP1 activation and AIF-mediated cell death (parthanatos)14,22. It has been proposed that processing of oxidatively generated clustered DNA lesions may be prolonged, which leads to the increase in the persistent strand breaks23. Given that AP sites and strand breaks are the intermediates of OGG1-BER, we questioned whether OGG1-initiated base-excision repair plays a role in PARP1-driven parthanatos. Our study showed that OGG1 expression increased, while genetic or siRNA-mediated ablation of Ogg1, as well as expression of enzymatically deficient OGG1 decreased accumulation of DNA strand breaks, PARP1 overactivation, and AIF nuclear translocation, and consequently, cell death. We also documented OGG1’s role in NMDA-induced parthanatos in SH-SY5Y neuroblastoma cells. The present study indicated that excessive OGG1-BER could be lethal for cells subjected to a high level of oxidative insult.
Results
OGG1 exacerbates H2O2-induced PARP1-dependent AIF-induced cell death
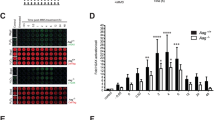
Although it is not a free radical, hydrogen peroxide (H2O2) has a relative long lifespan, may travel long distances within cells, and be efficiently converted into hydroxyl radical, efficacious to activate PARP1;24,25 thus we used H2O2 to mimic oxidative stress. Incubation of MEFs with H2O2 for 30 min induced protein PARylation in a dose-dependent manner. An overt increase in the level of PARylated proteins was observed upon 400 μM H2O2 exposure (Fig. 1a; Fig. S1A), which was diminished by PARP1 inhibitor PJ34 (Fig. 1b; Fig. S1B). Flow cytometry analysis showed that H2O2 exposure also induced cell death in a dose-dependent manner (Fig. 1c). Incubation with 400 μM H2O2 resulted in ~60% annexin V and PI dual-positive cells, which was markedly suppressed by PJ34 to <10% (Fig. 1d). As control, etoposide (ETO) was utilized. ETO is widely used for chemotherapy that introduces DNA double-strand breaks (DSBs) by inhibition of top II, leading to cells undergoing typical apoptosis26. Flow cytometry analysis showed that ETO exposure triggered cell death in a dose-dependent manner (Fig. S2A). However, incubation of MEFs with ETO for 30 min did not induce a typical PARP1 activation regarding the magnitude and the location of the PARylation signals (Fig 1e; Fig. S2B). While H2O2-induced PARylation majorly emerged from the nucleus, forming the signal foci, ETO-induced PARylation was weak and evenly distributed both in the cytoplasm and nucleus (Fig. 1e). Nevertheless, PJ34 could not suppress ETO-induced cell death, which primarily exhibited as annexin V positive and PI negative (Fig. 1f). In parallel, cleavage of caspase-3 and PARP1, the typical marks of apoptosis were induced by ETO but not H2O2 exposure (Fig. 1g), and caspase-3 activity was hardly detectable during the timescale of H2O2 exposure (Fig. S3). The broad-spectrum caspase inhibitor (caspase-inhibitor VI; Z-VAD-FMK) failed to block H2O2-induced increase in annexin V and PI dual-positive cells, but effectively inhibited ETO-induced increase in annexin-V-positive and PI-negative cells (Fig. 1h). To further prove that the H2O2-induced cell death is AIF-mediated, MEFs were transfected with siRNA against Aif or control, and then incubated with H2O2. Flow cytometry indicated that the percentage of annexin V and PI dual-positive cells was significantly decreased by siAIF (Fig. 1i). Data suggested that oxidative stress-induced cell death is parthanatos one.
a Microscopic assessment of protein PARylation in cells exposed to H2O2. MEFs were incubated with increasing concentrations of H2O2 for 30 min. Immunofluorescence microscopy was performed to visualize PAR signals. Nuclei were counterstained with DAPI. Scale bar: 10 μm. b PJ34 inhibits H2O2-induced PARP1 activation. MEFs were incubated with 400 μM H2O2 in the presence of 2.5 μM PJ34 or not. Immunofluorescence microscopy was conducted to analyze PARP1 activation. The lower row shows the three-dimensional plot of the intensity of PARylation shown in the upper panels, as determined by using Image J software. Scale bar: 10 μm. c H2O2 exposure induces cell death in a dose-dependent manner. MEFs were incubated with increasing concentrations of H2O2 for 24 h. Flow cytometry analysis of annexin V-FITC/PI staining was conducted to examine cell death. d PARP1 activation is involved in cell death induced by H2O2. MEFs were incubated with 400 μM H2O2 for 24 h in the presence of 2.5 μM PJ34 or not. Flow cytometry analysis of annexin V-FITC/PI staining was conducted to examine cell death. e, f ETO fails to elicit typical PARP1 activation. e MEFs were incubated with 400 μM H2O2 or 300 μM ETO for 30 min. Immunofluorescence microscopy was conducted to visualize PAR signals. Nuclei were counterstained with DAPI. The middle panel shows the fluorescence intensity of PARylation from line scans in the merged images, analyzed by Image Pro Plus software (red line: PARlytion; blue line: DAPI). The right panel shows the relative fluorescence intensity of Parylation with an average of 100 cells counted and analyzed by Image J software. Scale bar: 10 μm. **p < 0.01. And (f) MEFs were incubated with 300 μM ETO for 24 h in the presence of 2.5 μM PJ34 or not. Flow cytometry analysis of annexin V-FITC/PI staining was conducted to examine cell death. g Treatment of ETO but not H2O2 induces the cleavage of PARP1 and caspase-3. MEFs were incubated with 400 μM H2O2 or 300 μM ETO for 24 h. Whole-cell lysates were prepared, and western blotting was performed to determine the cleavage of PARP1 and caspase-3. h H2O2-induced cell death is caspase-independent. MEFs were incubated with 400 μM H2O2 or 300 μM ETO for 24 h in the presence of pan-caspase inhibitor (Z-VAD.FMK; 100 μM) or not. Flow cytometry analysis of annexin V-FITC/PI staining was conducted to examine cell death. i Knockdown of AIF decelerates cell death induced by H2O2 exposure. MEFs were transfected with siRNA targeting Aif or control for 36 h, and then incubated with H2O2 (400 μM) for 24 h. Flow cytometry analysis of annexin V-FITC/PI staining was conducted to examine cell death (left). The right panel shows the efficiency of AIF interference (upper) and the quantifications of annexin V and PI double-positive cells (lower). **p < 0.01, n ≥ 3
Next, we tested whether OGG1 expression affects H2O2-induced cell death. Wild-type (Ogg1+/+) and Ogg1 knockout (Ogg1–/–) MEFs cells were exposed to 400 μM H2O2 for 24 h. Flow cytometry showed that the percentage of annexin V and PI double-positive Ogg1+/+ MEFs was almost sixfold higher than that of Ogg1−/− MEFs (Fig. 2a). In parallel, knockout of endonuclease III-like protein I (Nth1), the BER enzyme specially repairing thymine glycol, reduced H2O2-induced parthanatos to ~50% (Fig. S4), suggesting that OGG1 expression is more decisive in oxidative stress-induced cell death. To further affirm the effect of OGG1, MEFs were transfected with siRNA against Ogg1 or control, and then incubated with H2O2. Flow cytometry indicated that the percentage of cell death was significantly decreased by siOgg1 (Fig. 2b). Furthermore, YFP-OGG1 and the YFP-vector plasmids were transfected into Ogg1−/− MEFs. The YFP-positive cells were gated out and H2O2-induced cell death was analyzed. Results showed that Ogg1−/− MEFs with YFP-OGG1 overexpression underwent more severe cell death than the cells expressing YFP alone did (Fig. 2c). The combined data demonstrated that OGG1 expression exacerbated oxidative stress-induced cell parthanatos.
a Ogg1+/+ MEFs are more vulnerable to undergo parthanatos than Ogg1−/− MEFs upon H2O2 exposure. Ogg1+/+ and Ogg1−/− MEFs were incubated with 400 μM H2O2 for 24 h. Flow cytometry analysis of annexin V-FITC/PI staining was conducted to examine cell death (upper). b Knockdown of Ogg1 ameliorates cell death induced by H2O2 exposure. MEFs were transfected with siRNA targeting Ogg1 or control for 36 h, and then incubated with H2O2 (400 μM) for 24 h. Flow cytometry analysis of annexin V-FITC/PI staining was conducted to examine cell death (upper). c Overexpression of OGG1 enhances H2O2-induced cell death. YFP-OGG1 and control YFP-vector plasmids were transfected into Ogg1–/– MEFs for 36 h, and then incubated with H2O2 (400 μM) for 24 h. The YFP-positive cells were gated out, and H2O2-induced cell death was analyzed by flow cytometry analysis of annexin V-APC/PI staining (upper). Lower panels show the quantifications of annexin V and PI double-positive cells shown in the upper panels (n ≥ 3) as well as the efficiency of Ogg1 interference. **p < 0.01, ***p < 0.001
OGG1 is implicated in the activation of parthanatos signal pathway in oxidatively stressed cells
To determine whether OGG1 is the causative factor for the increase in DNA strand breaks that enhances the activation of PARP1, TUNEL assay was conducted, and DNA breaks induced by increasing levels of H2O2 were examined. TUNEL-positive Ogg1-expressing and Ogg1-null cells showed no significant difference along with the increase in H2O2 concentrations up to 200 µM. However, when cells were challenged with 400 µM H2O2, the number of TUNEL-positive Ogg1-expressing cells was twofold more than that of Ogg1-null cells (Fig. 3a). Data suggested that if oxidative stress is moderate, OGG1 may repair the lesion effectively and efficiently, and DNA breaks may come from the direct attack of ROS to DNA backbone; whereas, OGG1-BER may unexpectedly become a major source of DNA strand breaks in response to an aggressive ROS insult. Next, we performed comet assays under different conditions. Alkaline comet assay, which detects DNA breaks including both DSBs and SSBs, revealed that H2O2-treated Ogg1+/+ MEFs have more DNA strand breaks compared with Ogg1−/− MEFs (Fig. 3b). Neutral comet assay, by which only DSBs are detectable, also showed that H2O2-treated Ogg1+/+ MEFs have more double DNA strand breaks (Fig. 3c). Phosphorylation of ser139 residue of H2AX (γH2AX) is a mark of DSBs. In support, western blotting indicated a rapid increase in γH2AX content in Ogg1+/+ MEFs, which retained at the high level from 2 to 6 h post H2O2 exposure; whereas, Ogg1−/− MEFs showed less H2AX phosphorylation and the level of γH2AX gradually decreased after 2 h of H2O2 addition (Fig. S5A).
a High level of H2O2 induces more DNA strand breaks in cells expressing Ogg1 than that in Ogg1 knockout cells. Ogg1+/+ and Ogg1−/− MEFs were incubated with increasing concentrations of H2O2 for 3 h, and TUNEL assay was conducted. Cells were stained for detection of 3′-OH DNA termini, as described under Materials and methods and analyzed by flow cytometry (left). Percentages of TUNEL-positive cells were quantified (right). n ≥ 3. b, c Analyses of DNA strand breaks in cells exposed to H2O2 by comet assays. Ogg1+/+ and Ogg1−/− MEFs were incubated with 400 μM H2O2 for 30 min. b Alkaline comet assay was conducted to analyze both SSBs and DSBs, and (c) neutral comet assay to analyze DSBs, scale bar: 10 μm. The olive tail moment was determined by using Cometscore software. One hundred individual comets were counted for each sample (left panels). d PARP1 activation is stronger in Ogg1-expressing cells upon exposure of H2O2. Ogg1+/+ and Ogg1−/− MEFs were exposed to H2O2 for the increasing lengths of time as indicated. Western blotting was conducted to analyze the levels of protein PARylation. e OGG1 promotes translocation of AIF to the nucleus in response to H2O2 exposure. Ogg1+/+ and Ogg1−/− MEFs were incubated with H2O2 for 12 h, and then, the cells were fixed and labeled with an antibody against AIF. The nuclei were counterstained with DAPI. The right column of the left panel shows magnified views of the boxed area in the AIF images. The profiles in the middle panel show the fluorescence intensity of AIF from line scans in the merged images, analyzed by Image Pro Plus software (red line: AIF; blue line: DAPI). Quantitative analysis of AIF nuclear transfer is shown in the right panel, with an average of 100 cells counted (for image analysis method see Materials and methods). Scale bar: 10 μm. **p < 0.01, ***p < 0.001
Accordingly, as shown by the formation of PAR polymers, PARP1 activation was elicited at 30 min, and reversed down from 2 h, and went back to the basal level at 6 h after addition of H2O2 in Ogg1+/+ MEFs; whereas in Ogg1−/− MEFs, the level of protein PARylation was markedly lower, and normalized after 3 h of H2O2 exposure (Fig. 3d). Moreover, immunofluorescence microscopy showed that nuclear AIF signals were overt in Ogg1+/+ MEFs compared with that in the Ogg1−/− MEFs (Fig. 3e). This result was confirmed by western blot (Fig. S5B). Taken together, these data suggested that OGG1 increases oxidative stress-induced DNA strand breaks, which led to the hyperactivation of PARP1 and the translocation of AIF to the nucleus.
OGG1’s BER activity accounts for oxidative stress-induced parthanatos
We explored whether OGG1-BER exacerbates oxidative stress-induced DNA strand break formation and PARP1 activation. To this end, nuclear ROS level and genomic 8-oxoG content were monitored first. The nuclear ROS sensor pHyPer-Nuc GFP was overexpressed in MEFs, and after addition of H2O2 in a medium, the ROS level presented by the intensity of GFP fluorescence reached the summit at 5 min and then gradually decreased to basal level by 60 min (Fig. 4a). Likewise, the kinetics of pHyper-Nuc GFP fluorescence in Ogg1−/− MEFs showed the same changes (data not shown). Accordingly, dot-blot analysis showed that H2O2 exposure led to a global increase in genomic 8-oxoG level in Ogg1+/+ MEF cells from 5 min onward, which peaked at 15 min, and then fell back progressively; whereas the increase in 8-oxoG level in Ogg1−/− MEF cells persisted from 5 min onward to over 90 min (Fig. 4b). The result not only indicated an increase in 8-oxoG formation upon the robust elevation of nuclear ROS level, as well as a permissible repair of the guanine lesions in the OGG1-expressing cells along with the restoration of the redox balance, but also implied that the high level of 8-oxoG itself may not increase the fatality.
a The kinetic changes in the nuclear ROS level in MEF cells after H2O2 exposure. Plasmids pHyper-Nuc were transfected into MEFs for 48 h, and then treated with H2O2 (400 μM). The sequential images were captured by spinning-disk confocal microscope. Scale bar: 10 μm. Upper: the kinetic changes in the nuclear ROS level; left of lower: the distribution of the nuclear ROS sensor pHyPer-Nuc GFP; right of lower: quantification of the kinetics of the nuclear ROS level by Volocity 5.5.1 software. b Impaired 8-oxoG repair in Ogg1-null cells upon H2O2 exposure. Ogg1+/+ and Ogg1−/− MEFs were incubated with H2O2 for increasing lengths of time as indicated. Genomic DNA was isolated and global 8-oxoG levels were analyzed by dot-blot assay using 8-oxoG antibody as described under Materials and methods. c Functional defect of OGG1 ameliorates DNA strand breaks in cells upon H2O2 exposure. YFP-OGG1, YFP-OGG1K249A, and YFP-OGG1K249Q plasmids were transfected into Ogg1−/− MEFs for 36 h, and then incubated with H2O2 (400 μM) for 6 h. Cells were stained for detection of 3′-OH DNA termini (TUNEL assay) and the YFP-positive cells were gated out and analyzed by flow cytometry, n ≥ 3; **p < 0.01, ***p < 0.001
To verify that OGG1-BER causes PARP1 overactivation and contributes to oxidative stress-induced parthanatos, YFP-OGG1 and the inactive mutants YFP-OGG1K249A, OGG1K249Q plasmids were constructed and transfected in Ogg1−/− MEFs. The residue lysine 249 in OGG1 active site is crucial for its DNA glycosylase and AP lyase activities27. Accordingly, TUNEL labeling showed that YFP-OGG1-expressing cells had a significantly higher level of DNA strand breaks (~40%) than YFP-OGG1K249A- (~12%) and OGG1K249Q-expressing (~24%) cells (Fig. 4c).
To investigate the role of OGG1-BER in parthanatos, plasmids carrying OGG1, OGG1K249A, and OGG1K249Q were transfected into Ogg1−/− MEFs, respectively. It is worthwhile to note that excessive PARylation occurred in YFP-OGG1-expressing cells compared with those without YFP signals; whereas, the levels of PARylation in the YFP-OGG1K249A- and YFP-OGG1K249Q-expressing cells were weak and similar as that in YFP-negative cells (Fig. 5a). Immunofluorescence detection also revealed that in response to H2O2 exposure, nuclear AIF signals were overt in Flag-OGG1-expressing cells but not in those absent from Flag signals. Moreover, less AIF translocated into nuclei in the Ogg1−/− MEFs expressing Flag-OGG1K249A or Flag-OGG1K249Q (Fig. 5b). Moreover, flow cytometry assay showed that incubation of H2O2 induced more YFP-OGG1-expressing cells undergoing parthanatos than the YFP-OGG1K249A- and YFP-OGG1K249Q-expressing ones (Fig. 5c). Taken together, these data indicated that OGG1-BER exacerbates oxidative stress-induced parthanatos.
a Cells expressing OGG1 mutants exhibit hampered protein PARylation in response to H2O2 exposure. YFP-OGG1, YFP-OGG1K249A, and YFP-OGG1K249Q plasmids were transfected into Ogg1−/− MEFs for 36 h. Cells were incubated with H2O2 (400 μM) for 30 min. Immunofluorescence microscopy was performed to examine protein PARylation. Nuclei were counterstained with DAPI. The graphs (in the right panel) show the fluorescence intensity profiles in three fluorescence channels along line scans in the merged images (in the left panel), analyzed by Image Pro Plus software (red line: PARylation; green line: YFP; blue line: DAPI). b, c Cells expressing OGG1 K249 mutants exhibit hampered nuclear translocation of AIF and weakened parthanatos in response to H2O2 exposure. Transfection was conducted as described in a, and then (b) cells were incubated with H2O2 for 12 h. Microscopy was performed to visualize AIF and Flag. Nuclei were counterstained with DAPI. The right panels show the fluorescence intensity profiles as described in a (red line: AIF; green line: Flag; blue line: DAPI); or c cells were incubated with H2O2 for 24 h, and the YFP-positive cells were gated out. Cell death was examined by flow cytometry analysis of annexin V-APC/PI staining (left). Percentages of annexin-V and PI double-positive cells (parthanatos) were quantified and shown at the right panel. Scale bar: 10 μm. n ≥ 3; **p < 0.01
OGG1 is implicated in NMDA-induced neuronal cell parthanatos
Parthanatos is a predominant form of neuron cell death triggered by the neurotoxicity of NMDA28. To verify the role of OGG1-BER in the induction of parthanatos, SH-SY5Y neuroblastoma cells were utilized. To determine the level of nuclear ROS induced by NMDA, pHyPer-Nuc GFP was overexpressed. Confocal microscopy detection revealed that NMDA exposure indeed increased the level of nuclear ROS, which was inhibited by NAC (Fig. 6a; Fig. S6). Thus, we proposed that OGG1-BER may enhance PARP1 overactivation and contribute to NMDA-induced neurons cell death. Immunofluorescence labeling showed that NMDA stimulation led to AIF translocation into the nucleus, which was hampered if the cells were pretreated with NAC or siRNA targeting OGG1 (Fig. 6b). Likewise, NMDA-induced cell death was also significantly lessened by administration of NAC or OGG1 silencing (Fig. 6c). The OGG1 interference efficiency was shown in Fig. 6d (left). These results taken together suggested that OGG1-BER may play an exacerbating role in NMDA-induced neurons parthanatos.
a NMDA exposure increases the level of intracellular ROS. Plasmids pHyper-Nuc were transfected into MEFs for 48 h, and cells were incubated with NMDA (500 μM) for the increasing lengths of times, as indicated in the presence or absence of 10 mM NAC. The average of GFP fluorescence intensity representing the nuclear ROS level was quantified using Image J software. Fifty cells were counted for each sample. b ROS scavenger or siRNA targeting OGG1 significantly hampered NMDA-induced nuclear translocation of AIF. SH-SY5Y cells were transfected with a siRNA against OGG1 or control for 36 h. Cells were incubated with 500 μM NMDA for 30 min in the presence or absence of 10 mM NAC, and after 12 h, they were fixed and labeled with an antibody against AIF. The nuclei were counterstained with DAPI. Scale bar: 10 μm. The right column of the left panel shows magnified views of the boxed area in the AIF images. The profiles in the right panel show the fluorescence intensity of AIF from line scans in the merged images, which were analyzed by Image Pro Plus software (red line: AIF; blue line: DAPI). c ROS scavenger or siRNA targeting OGG1 significantly inhibited NMDA-induced cell death. SH-SY5Y cells were transfected with siRNA against OGG1 or control for 36 h. Cells were incubated with NMDA for 30 min in the presence or absence of 10 mM NAC, and after 24 h, cell death was examined by flow cytometry analysis of annexin V-FITC/PI staining. d Western blot detection of siOGG1 interference efficiency (left). Western blot analyses of Sh-OGG1 and siAPE1 interference efficiency in SH-SY5Ycells (right). e APE1 silencing has a deleterious effect on NMDA-induced cell death only with OGG1-proficient cells. Sh-CTR and Sh-OGG1 SH-SY5Y cells were transfected with siRNA against APE1 or control for 36 h. Cells were incubated with NMDA for 30 min, and after 24 h, cell death was examined by flow cytometry analysis of annexin V-FITC/PI staining (left). Percentages of annexin-V and PI double-positive cells (parthanatos) were quantified (right). n ≥ 3; **p < 0.01, ***p < 0.001
To further prove that DNA breaks left unrepaired after OGG1-BER play a major role in cell death induction, we detected the effects of OGG1 and APE1 double silencing. Recently, APE1 conditional knockout mouse was developed. The induced deletion of APE1 dramatically enlarged the infarct volume along with activation of PARP1 after transient focal cerebral ischemia29. Since APE1 primarily acts after glycosylases in BER pathway, we tested the effect of OGG1 expression on the consequence of APE1 deletion in the present study. To do so, we established OGG1 stably silenced SH-SY5Y cells as described under Materials and methods (Fig. 6d, right). OGG1-proficient or OGG1-deficient cells were further subjected to APE1 deletion or not, and then exposed to NMDA. In OGG1-proficient cells, APE1 deletion resulted in ~70% cells undergoing parthanatos upon NMDA treatment, which was ~20% higher than that of APE1-expressing cells; while OGG1 was deficient, cell death rate was only ~15%, and APE1 silencing did not further increase the deleterious effect (Fig. 6d, e). This result suggested an epistatic effect of OGG1 in cell death induction, and once OGG1 initiates the lesion excision, the repair intermediates may lead to hyperactivation of PARP1 and result in increased cell parthanatos if the follow-up steps in BER pathway are uncoupled.
Discussion
The excessive ROS and resultant cell death have been well established as etiological factors involved in ischemic injury and degenerative alterations10,30. ROS cause damage to DNA, and the predominant form of DNA damages is base oxidation4. BER is a frontline defense to remove oxidized base lesions2. While it has been widely acknowledged that unrepaired DNA damages would lead to various deteriorated consequences31,32, our present study demonstrated that after oxidative insult, OGG1-initiated BER plays an exacerbating role in the induction of PARP1-dependent, AIF-mediated cell death, linking the process of base-excision repair to the pathogenesis of parthanatos.
Oxidized bases are the common oxidative damage along with other DNA lesions such as oxidized sugar fragments, abasic (AP) sites, and single-strand breaks (SSBs)32,33. Due to guanine’s lowest ionization potential among DNA bases3, the most predominant oxidative base product is 8-oxoG, which is primarily recognized and repaired by OGG15. Repair of oxidized bases could further indirectly result in SSBs, and closely spaced SSBs may convert to DSBs23. Processing of oxidatively generated clustered DNA lesions may be prolonged, resulting in the increase in the persistent strand breaks23, which may account for the hyperactivation of PARP1 under oxidative stress.
The present study showed that the expression of OGG1 significantly augments cell death induced by a high level of H2O2 (Figs.1 and 2), whereas, lack of OGG1, even though resulting in sustained high level of 8-oxoG in genomic DNA (Fig. 4b), lowered H2O2-induced activation of PARP1 due to less level of DNA breaks including SSBs and DSBs (Fig. 3b–d). Likewise, less DNA breaks, decreased PARP1 activation, and AIF nuclear translocation were observed in OGG1K249A- and OGG1K294Q-expressing cells upon H2O2 exposure (Fig. 5a, b). On the contrary, while concentration of H2O2 was lower, OGG1 expression did not increase DNA strand breaks (Fig. 3a). Results collectively suggested that aggressive ROS may result in glycosylase-initiated BER overwhelming, and the successive strand break repair uncoupled; thus, the left-behind repair intermediates turn into major substrates for PARP1 activation.
The Ogg1−/− mice were developed in two laboratories independently34,35. However, despite the accumulation of supraphysiological levels of 8-oxoG in the nuclear and mitochondrial genomes, the null animals displayed no noticeable changes in phenotype, including no impaired embryonic development, no marked pathological changes, or tumor induction34,35,36. Interestingly, homozygous deletion of nearly all of the DNA glycosylases is compatible with life37,38. In contrast, homozygous deletion of the core component of BER (such as APE1, XRCC1, and POL β) leads to early-stage embryonic or postnatal lethality39,40. The phenotypes of these null mice and the data of the present study suggested that once BER initiated, the uncoupled repair intermediates may evoke parthanatos, which accounts for the lethality of glycosylase genes deletion, whereas, the existence of oxidized bases is tolerable. In support, a study utilizing a transient focal cerebral ischemia model showed the activation of PARP1 and enlarged infarct volume in APE1 conditional knockout mouse29. And our result revealed that the deleterious effect of APE1 deletion in parthanatos induction only displayed in OGG1-proficient but not OGG1-deficient cells (Fig. 6e).
Both OGG1 and PARP1 are responsible for DNA damage repair, and the cross talk between two types of enzymes has drawn attention41. Binding with and modification by PARP1 decreases the BER function of OGG1, which may suggest a feedback in turn ceasing OGG1-initiated BER-caused PARP1 overactivation41. There are studies documenting that no significant difference in cell survival was observed between wt and Ogg1−/− cells41, and cells deficient in NTH1 or OGG1 exhibited elevated cell death after exposure of H2O242. We deduced that the discrepancy of these conclusions from our studies may result from the mild extent of oxidative insults such as low concentrations or short exposure of H2O2, under which OGG1 expression does not increase DNA strand breaks (Fig. 3a), and glycosylase-initiated BER should be beneficial for cell survival (Fig. 7, left). But if base oxidation is excessive, OGG1-BER leads to strand break overwhelming and the repair uncoupled, which promotes PARP1 overactivation and cell death (Fig. 7, right).
OGG1-initiated BER prevents accumulation of mutagenic guanine base lesions, and is beneficial for cell survival when oxidative insult is mild, and the level of 8-oxoG is low (left). However, if base oxidation is excessive, OGG1-BER causes strand break overwhelming and the repair uncoupled, which promotes PARP1 overactivation, free PAR formation and release, AIF transfer to the nucleus, and chromatin hydrolysis, leading to cell parthanatos (right)
Since our and others’ data12,24,43,44 demonstrated that H2O2-induced DNA damage effectively elicits PARP1 overactivation, ROS and glycosylase-initiated BER are implicated in N-methyl-d-aspartate (NMDA) and N-methyl-N′-nitro-N′-nitrosoguanidine (MNNG)-triggered cell death that has been widely taken as parthanatos, which arises as a question. Neuronal damage following stroke or in neurodegenerative diseases is thought to stem in part from overexcitation of NMDA receptors by glutamate, which results in parthanatos22,45. Excitotoxic activation of NMDA receptor results in formation of ROS and peroxynitrite (ONOO–), which inhibits cellular respiration, in turn leading to the positive feedback of increase in ROS generation. ROS cause DNA damage, which leads to the excessive PARP1 activation and AIF-mediated cell death (parthanatos)14,22. Our data showed that NMDA indeed elevated the intracellular ROS level in SH-SY5Y neuroblastoma cells (Fig. S6A, Fig. 6a), ROS scavenger and OGG1 interference significantly blocked AIF nuclear translocation and cell death (Fig. 6b, c). The alkylating agent MNNG induced oxidative stress and also caused excess DNA strand breaks that led to PARP1 overactivation and parthanatos12,46. Further study showed that antioxidant NAC abrogated all phenomena caused by MNNG, including enhanced DNA damage and PARP-1 activation47. It is possible that glycosylases such as OGG1 have a role in MNNG-induced parthanatos. All the studies suggested that ROS generation is a relevant event in pathological contexts, leading to DNA damage and then PARP1 overactivation. Glycosylase-initiated uncoupled BER exacerbates DNA damage, and thus it is a driving force for cells undergoing parthanatos.
Previous studies from our and others' groups presented that OGG1-BER activity is redox regulated48,49,50. When the intracellular context is oxidized, binding of enzymatically inactive OGG1 with promoter-located guanine lesion may be utilized to assemble the transcriptional machinery to launch the transcription of “ROS-response-ome”49,51,52. While redox balance is reestablished, OGG1 resumes its BER function. In the present study, the coincident kinetics of nuclear ROS and genomic 8-oxoG levels reflected an initiation of guanine lesion repair along with the reestablishment of the redox balance (Fig. 4a, b); however, the effective completion of the multiple-stepped BER somehow failed. Taken together, data suggested that while the redox levels orchestrate OGG1 to play a role either in gene transcription or in lesion repair; once BER is initiated after redox permission, the magnitude of base lesions, predominantly of 8-oxoG, defines the fate of cells, survival, or death.
In summary, our data link the repair of oxidized DNA bases to cell death triggered by PARP1 overactivation, and indicate that the OGG1-BER could be lethal for cells after exorbitant oxidative stress. Thus, the assessment of the prompt repair of oxidized bases under different conditions should be discrete. The present study suggested that OGG1 guards genome integrity not only depending on its capability to repair the base lesions, but also through promoting cell death to eliminate the cells with malignant potential, thus to maintain the homeostasis and protect the host.
Materials and methods
Antibodies and reagents
Rabbit anti-PAR polyclonal antibody (Ab) (4336-BPC-100; WB, 1:2000; IF, 1:200) was purchased from Trevigen (Gaithersburg, MD, USA). Mouse anti-8-oxoG monoclonal Ab (MOG100P; 1:1000) was from JaICA (Institute for the Control of Aging, Japan). Rabbit anti-Ogg1 polyclonal Ab with the specificity for recognition of mouse Ogg1 (ab135940; 1:3000) and monoclonal Ab with the specificity for recognition of human OGG1 (ab124741; 1:3000) were purchased from Abcam (San Francisco, CA). Mouse monoclonal anti-AIF (sc-13116; WB, 1:1000; IF, 1:200) and anti-APE1 (sc-17774; 1:2000) Abs were from Santa Cruz Biotechnology (Santa Cruz, CA, USA). Anti-γH2AX (9718S; 1:3000), rabbit polyclonal anti-AIF (4642S; IF, 1:200), and anti-caspase-3 (9662S; 1:3000) Abs were from Cell Signaling Technology (Boston, MA, USA). Anti-β-tubulin (HC101; 1:8000), anti-β-actin (HC201; 1:8000) mouse, and anti-GAPDH (H301; 1:5000) mouse monoclonal Abs were purchased from TRANS (Beijing, China). The monoclonal antibody against FLAG (F1804; IF, 1:500) was from Sigma (Saint Louis, MO, USA). Rabbit anti-lamin B polyclonal (Ab) (12987-1-AP; 1:3000) was purchased from Proteintech (Chicago, USA). Hydrogen peroxide (H2O2, 216763), PARP1 inhibitor PJ34 (P4365), N-methyl-D-aspartic acid (NMDA, M3262), and N-acetyl cystenic (NAC, A7250) and caspase-inhibitor VI (Z-VAD-FMK, 219007) were purchased from Sigma-Aldrich (St. Louis, MO, USA).
Cell culture and treatments
Wt, Ogg1−/−, and Nth1−/− MEFs were kindly provided by Dr. Istvan Boldogh (University of Texas Medical Branch at Galveston, Texas). SH-SY5Y neuroblastoma cells were kindly provided by Dr. Pengfei Ge (Department of Neurosurgery, First Hospital of Jilin University, Changchun, China). Cells were grown in DMEM (Invitrogen) supplemented with 10% fetal bovine serum. For the induction of cell death, cells were incubated with indicated concentrations of H2O2 for 24 h. For detection of protein PARylation, cells were incubated with H2O2 for 30 min. Alternatively, cells were incubated with H2O2 for 30 min or 24 h in the presence or absence of the following inhibitors: PJ34 (2.5 μM) or NAC (10 mM).
Plasmids and transfection
YFP-OGG1, Flag-OGG1 plasmids were constructed by inserting the OGG1 coding sequence into pZsYellow1-C1, pcDNA3.1(+) vectors in our laboratory. Transfection was performed using Lipofectamine 3000 reagent (Invitrogen, Carlsbad, CA) according to the manufacturer’s instructions. Cells were used for the experiment 36 h post transfection.
SiRNA-mediated interference
For RNA interference experiments, scramble control, or smart-pool siRNA respectively against hOGG1 and mOgg1 (Dharmacon), APE1 and AIF were transfected into MEFs or SH-SY5Y cells using Lipofectamine 3000 according to the manufacturer’s instructions. siRNA duplex oligoribonucleotides were synthesized by GenePharma (Shanghai, China). The sense sequences were as follows: control siRNA (nonsense oligo), 5′-UUCUCCGAACGUGUCACGUTT-3′; AIF siRNA, 5′-GAAACUGACCACAUAGUGATT-3′; and APE1 siRNA, 5′-GUCUGGUACGACUGGAGUACC-3′. Cells were used for experiments after 48 h of interference, and the efficacy of target gene knockdown was validated by western blotting.
ShRNAs and lentiviral transduction
To stably knock down OGG1 in SH-SY5Y cells, the pLKO.1-TRC cloning vector (Addgene) was used. The OGG1 shRNA sequences were 5′-CCGGAAAGAGGTGGCTCAGAAATTCCTCGAGGAATTTCTGAGCCACCTCTTTTTTTTG-3′ and 5′- AATTCAAAAAAAAGAGGTGGCTCAGAAATTCCTCGAGGAATTTCTGAGCCACCTCTTT-3′. The shRNAs were synthesized and cloned into the vectors. The constructed plasmids and shCtrl plasmid were transfected into 293T cells, together with the packaging plasmid psPAX2 and the envelope plasmid pMD2.G (both from Addgene) by using the Lipofectamine 2000 reagent (Invitrogen). And the collection of supernatant and 5 mg/ml polybrene (Sigma) were used to infect the cells. The cells were selected in puromycin-containing medium 3 days after infection.
Immunofluorescence microscopy
Cells grown on glass coverslips were fixed with 10% (v/v) formaldehyde in PBST (PBS with 0.1% Triton X-100) for 10 min, permeabilized with 0.5% (v/v) Triton X-100 for 5 min, and then blocked with 10% (w/v) bovine serum albumin for 30 min. Thereafter, cells were incubated with the primary Abs recognizing PAR (1:200), AIF (1:200), Flag (1:500), and then the secondary Abs successively. The nuclei of the cells were stained with 4,6-diamidino-2-phenylindole (DAPI) for 5 min. Cells were visualized by using a confocal microscope (Nikon, Tokyo, Japan) equipped with a ×60 oil-immersion objective lens. The total protein PARylation was quantified by using Image J software. Fluorescence intensity of proteins (PARylation, AIF, YFP, Flag, and DAPI) was quantified by using IPP software.
To quantify the amount of nuclear AIF, Image J software was used. First, the total AIF amount was measured, and then measure the AIF amount of nuclear. The transfer of AIF was quantified by the nuclear divided by the total amount of AIF.
Cell fractionation
Collection of cell fractionations was performed as described previously53, with slight modifications. Briefly, for western blot, cells were harvested, washed with PBS, and lysed in RIPA buffer (50 mM Tris-HCl, pH 7.4, 150 mM NaCl, 1% NP-40, 1 mM EDTA, 1 mM Na3VO4, 2 mM NaF, 1 mM β-glycerphosphate, and 2.5 mM sodium pyrophosphate containing 1 mM PMSF and protease inhibitors) for 30 min on ice. Lysates were centrifuged at 12,000 × g for 25 min at 4 °C, and the supernatant was the whole-cell extract (WE). Cytoplasmic and nuclear fractions were prepared using the CelLytic NuCLEAR Extraction Kit (Sigma) following the manufacturer’s instruction. Briefly, cells were lysed with cytosolic lysis buffer for 20 min, lysates were centrifuged at 11,000 × g for 1 min at 4 °C, and the supernatant was the cytoplasmic extract (CE). The pellet was washed twice with cytosolic lysis buffer and lysed with nuclear extraction buffer on ice for 30 min. Nuclear lysates were centrifuged at 21,000 × g for 5 min at 4 °C and the supernatant was the nuclear extract (NE).
Western blotting
MEFs and SH-SY5Y cells were stimulated and lysed. WE, CE, and NE, prepared as described above, were mixed with SDS sample buffer, separated by 10% SDS-PAGE, and transferred onto nitrocellulose membranes. Membranes were blocked with TBS containing 0.1% Tween 20 (TBST) and 5% skim milk for 1 h and incubated with primary Abs overnight. After three times of washing with TBST, membranes were incubated with secondary Abs and detected by ECL Plus western blot detection reagents.
Flow cytometry
Annexin V-FITC (AO2001-10) and Annexin V-APC (AO2001-11) Apoptosis Analysis Kits (Sungene Biotech, Tianjin, China) were used to detect cell apoptosis. Cell death detection was performed following the manufacturer’s instruction. Briefly, cells were harvested, washed with cold PBS, suspended in 1× binding buffer, and then centrifuged at 300 × g for 10 min. Supernatants were removed, and cells were resuspended in 100 μL of binding buffer with addition of 5 μL of annexin V-FITC (APC), and after a gentle vortex, they were incubated for 10 min in room temperature, and then another 5 min in room temperature after addition of 5 μL of PI solution. Flow cytometry was performed by using a FACSSanto II (BD Biosciences), and data were analyzed with FACSDiva software (BD Biosciences).
Comet assay
Alkaline, neutral comet assays were performed using the Comet Assay Kit from Trevigen (Gaithersburg, MD) following the manufacturer’s guidance. Images were captured using the fluorescent microscope (ECLIPSE, 80i, Nikon, Japan). Tail moment was determined using Cometscore software (TriTek, Sumerduck, USA).
Dot-blot analysis
Dot-blot analysis was performed as described previously54, with slight modifications. Genomic DNA was isolated using a TIANamp Genomic DNA Kit (TIANGEN, Beijing, China). RNase A (Sigma) digestion was included in the isolation procedure. Isolated genomic DNA (1 μg per sample) was denatured (0.4 M NaOH and 10 mM EDTA) at 95 °C for 5 min, and chilled immediately on ice for 10 min. Samples were serially twofold diluted and spotted on nitrocellulose membrane using the BioDot Microfiltration Apparatus (170-6545, Bio-Rad). The blotted membrane was cross-linked under UV for 4 min, and incubated in 2× SSC for 5 min. The membrane was blocked with TBS containing 0.1% Tween 20 (TBST) and 5% skim milk for 1 h and incubated with anti-8-oxoG (1:1000) Ab overnight, and after three times of washing with TBST, it was incubated with secondary Ab and detected by ECL Plus western blot detection reagents.
TUNEL labeling
The YF594 TUNEL Assay Apoptosis Detection Kit (US Everbright Inc.) was used to label 3′-OH of DNA breaks following the manufacturer’s guidance. Data were analyzed by FACSDiva (BD Biosciences) and FlowJo7.6.1 software.
Nuclear ROS assay
To assess the changes in nuclear ROS level, the hydrogen peroxide sensor pHyper-Nuc GFP (Evrogen, Axxora Inc., Farmingdale, NY) was used55. Cells grown in 35-mm dishes with 14-mm glass bottoms were tranfected with pHyPer-Nuc GFP (nuclear localization signal fused to the C-terminus of Hyper) for 48 h, treated with 400 μM H2O2, and placed on a 37 °C heated stage with 5% CO2 atmosphere chamber. The sequential images were captured by using an UltraVIEW Vox (PerkinElmer Inc., Waltham, MA) spinning-disk confocal microscope with a Ti-E microscope (Nikon, Japan) in fluorescence channel at 60-s intervals over a 60-min timecourse using a ×20 objective.
To detect nuclear ROS level in SH-SY5Y cells, pHyper-Nuc GFP was transfected into SH-SY5Y cells for 48 h. Cells were treated with NMDA (500 μM) for the increasing lengths of time (0, 15, 30, 45, and 60 min). Then they were fixed with 10% (v/v) formaldehyde in PBST (PBS with 0.1% Triton X-100) for 10 min, permeabilized with 0.5% (v/v) Triton X-100 for 5 min, and then blocked with 10% (w/v) bovine serum albumin for 30 min. Cells were visualized by using a confocal microscope (Nikon, Tokyo, Japan) equipped with a ×60 oil-immersion objective lens. The relative GFP fluorescence intensity were quantified by using Image J software.
Statistical analysis
The data were analyzed using Student’s t-test and presented as the means ± SEM. Quantifications are based on at least three independent experiments. The level of significance was accepted at *P < 0.05, **P < 0.01, and ***P < 0.001
References
Schieber, M. & Chandel, N. S. ROS function in redox signaling and oxidative stress. Curr. Biol. 24, R453–R462 (2014).
David, S. S., O’Shea, V. L. & Kundu, S. Base-excision repair of oxidative DNA damage. Nature 447, 941–950 (2007).
Barnes, D. E. & Lindahl, T. Repair and genetic consequences of endogenous DNA base damage in mammalian cells. Annu. Rev. Genet. 38, 445–476 (2004).
Lindahl, T. & Barnes, D. E. Repair of endogenous DNA damage. Cold Spring Harb. Symp. Quant. Biol. 65, 127–133 (2000).
Wallace, S. S. DNA glycosylases search for and remove oxidized DNA bases. Environ. Mol. Mutagen. 54, 691–704 (2013).
Dizdaroglu, M., Kirkali, G. & Jaruga, P. Formamidopyrimidines in DNA: mechanisms of formation, repair, and biological effects. Free Radic. Biol. Med. 45, 1610–1621 (2008).
Krokan, H. E. & Bjoras, M. Base excision repair. Cold Spring Harb. Perspect. Biol. 5, a012583 (2013).
Izumi, T. et al. Mammalian DNA base excision repair proteins: their interactions and role in repair of oxidative DNA damage. Toxicology 193, 43–65 (2003).
Odell, I. D., Wallace, S. S. & Pederson, D. S. Rules of engagement for base excision repair in chromatin. J. Cell. Physiol. 228, 258–266 (2013).
Cross, C. E. et al. Oxygen radicals and human disease. Ann. Intern. Med. 107, 526–545 (1987).
Fulda, S., Gorman, A. M., Hori, O. & Samali, A. Cellular stress responses: cell survival and cell death. Int. J. Cell Biol. 2010, 214074 (2010).
Yu, S. W. et al. Mediation of poly(ADP-ribose) polymerase-1-dependent cell death by apoptosis-inducing factor. Science 297, 259–263 (2002).
Yu, S. W. et al. Apoptosis-inducing factor mediates poly(ADP-ribose) (PAR) polymer-induced cell death. Proc. Natl Acad. Sci. USA 103, 18314–18319 (2006).
Wang, Y., Dawson, V. L. & Dawson, T. M. Poly(ADP-ribose) signals to mitochondrial AIF: a key event in parthanatos. Exp. Neurol. 218, 193–202 (2009).
Khodyreva, S. N. et al. Apurinic/apyrimidinic (AP) site recognition by the 5’-dRP/AP lyase in poly(ADP-ribose) polymerase-1 (PARP-1). Proc. Natl Acad. Sci. USA 107, 22090–22095 (2010).
Wang, Z., Wang, F., Tang, T. & Guo, C. The role of PARP1 in the DNA damage response and its application in tumor therapy. Front. Med. 6, 156–164 (2012).
Luo, X. & Kraus, W. L. On PAR with PARP: cellular stress signaling through poly(ADP-ribose) and PARP-1. Genes Dev. 26, 417–432 (2012).
Wang, Y. et al. Poly(ADP-ribose) (PAR) binding to apoptosis-inducing factor is critical for PAR polymerase-1-dependent cell death (parthanatos). Sci. Signal. 4, ra20 (2011).
Pacher, P. & Szabo, C. Role of the peroxynitrite-poly(ADP-ribose) polymerase pathway in human disease. Am. J. Pathol. 173, 2–13 (2008).
Wang, H., Shimoji, M., Yu, S. W., Dawson, T. M. & Dawson, V. L. Apoptosis inducing factor and PARP-mediated injury in the MPTP mouse model of Parkinson’s disease. Ann. N. Y. Acad. Sci. 991, 132–139 (2003).
Ba, X. et al. The role of 8-oxoguanine DNA glycosylase-1 in inflammation. Int. J. Mol. Sci. 15, 16975–16997 (2014).
Dawson, V. L. & Dawson, T. M. Deadly conversations: nuclear-mitochondrial cross-talk. J. Bioenerg. Biomembr. 36, 287–294 (2004).
Georgakilas, A. G. Processing of DNA damage clusters in human cells: current status of knowledge. Mol. Biosyst. 4, 30–35 (2008).
Batnasan, E. et al. 17-Beta estradiol inhibits oxidative stress-induced accumulation of AIF into nucleolus and PARP1-dependent cell death via estrogen receptor alpha. Toxicol. Lett. 232, 1–9 (2015).
Virag, L. & Szabo, C. The therapeutic potential of poly(ADP-ribose) polymerase inhibitors. Pharmacol. Rev. 54, 375–429 (2002).
Jamil, S., Lam, I., Majd, M., Tsai, S. H. & Duronio, V. Etoposide induces cell death via mitochondrial-dependent actions of p53. Cancer Cell Int. 15, 79 (2015).
Bruner, S. D., Norman, D. P. & Verdine, G. L. Structural basis for recognition and repair of the endogenous mutagen 8-oxoguanine in DNA. Nature 403, 859–866 (2000).
Yu, S. W., Wang, H., Dawson, T. M. & Dawson, V. L. Poly(ADP-ribose) polymerase-1 and apoptosis inducing factor in neurotoxicity. Neurobiol. Dis. 14, 303–317 (2003).
Stetler, R. A. et al. APE1/Ref-1 facilitates recovery of gray and white matter and neurological function after mild stroke injury. Proc. Natl Acad. Sci. USA 113, E3558–E3567 (2016).
Gorenkova, N., Robinson, E., Grieve, D. J. & Galkin, A. Conformational change of mitochondrial complex I increases ROS sensitivity during ischemia. Antioxid. Redox Signal. 19, 1459–1468 (2013).
Kidane, D. et al. Interplay between DNA repair and inflammation, and the link to cancer. Crit. Rev. Biochem. Mol. Biol. 49, 116–139 (2014).
Hegde, M. L., Izumi, T. & Mitra, S. Oxidized base damage and single-strand break repair in mammalian genomes: role of disordered regions and posttranslational modifications in early enzymes. Prog. Mol. Biol. Transl. Sci. 110, 123–153 (2012).
Hegde, M. L., Hazra, T. K. & Mitra, S. Early steps in the DNA base excision/single-strand interruption repair pathway in mammalian cells. Cell Res. 18, 27–47 (2008).
Klungland, A. et al. Accumulation of premutagenic DNA lesions in mice defective in removal of oxidative base damage. Proc. Natl Acad. Sci. USA 96, 13300–13305 (1999).
Minowa, O. et al. Mmh/Ogg1 gene inactivation results in accumulation of 8-hydroxyguanine in mice. Proc. Natl Acad. Sci. USA 97, 4156–4161 (2000).
Arai, T., Kelly, V. P., Minowa, O., Noda, T. & Nishimura, S. The study using wild-type and Ogg1 knockout mice exposed to potassium bromate shows no tumor induction despite an extensive accumulation of 8-hydroxyguanine in kidney DNA. Toxicology 221, 179–186 (2006).
Cabelof, D. C. Haploinsufficiency in mouse models of DNA repair deficiency: modifiers of penetrance. Cell. Mol. Life Sci. 69, 727–740 (2012).
Sampath, H. Oxidative DNA damage in disease--insights gained from base excision repair glycosylase-deficient mouse models. Environ. Mol. Mutagen. 55, 689–703 (2014).
Brenerman, B. M., Illuzzi, J. L. & Wilson, D. M. 3rd Base excision repair capacity in informing healthspan. Carcinogenesis 35, 2643–2652 (2014).
Abbotts, R. & Wilson, D. M. 3rd Coordination of DNA single strand break repair. Free Radic. Biol. Med. 107, 228–244 (2017).
Noren Hooten, N., Kompaniez, K., Barnes, J., Lohani, A. & Evans, M. K. Poly(ADP-ribose) polymerase 1 (PARP-1) binds to 8-oxoguanine-DNA glycosylase (OGG1). J. Biol. Chem. 286, 44679–44690 (2011).
Yang, N., Chaudhry, M. A. & Wallace, S. S. Base excision repair by hNTH1 and hOGG1: a two edged sword in the processing of DNA damage in gamma-irradiated human cells. DNA Repair (Amst.) 5, 43–51 (2006).
Xu, H. et al. Iduna protects HT22 cells from hydrogen peroxide-induced oxidative stress through interfering poly(ADP-ribose) polymerase-1-induced cell death (parthanatos). Cell. Signal. 25, 1018–1026 (2013).
Zheng, L. et al. JNK activation contributes to oxidative stress-induced parthanatos in glioma cells via increase of intracellular ROS production. Mol. Neurobiol. 54, 3492–3505 (2017).
Wang, Y. et al. A nuclease that mediates cell death induced by DNA damage and poly(ADP-ribose) polymerase-1. Science 354, aad6872 (2016).
Artus, C. et al. AIF promotes chromatinolysis and caspase-independent programmed necrosis by interacting with histone H2AX. EMBO J. 29, 1585–1599 (2010).
Chiu, L. Y., Ho, F. M., Shiah, S. G., Chang, Y. & Lin, W. W. Oxidative stress initiates DNA damager MNNG-induced poly(ADP-ribose)polymerase-1-dependent parthanatos cell death. Biochem. Pharmacol. 81, 459–470 (2011).
Bravard, A. et al. Redox regulation of human OGG1 activity in response to cellular oxidative stress. Mol. Cell. Biol. 26, 7430–7436 (2006).
Pan, L. et al. Oxidized guanine base lesions function in 8-oxoguanine DNA glycosylase-1-mediated epigenetic regulation of nuclear factor kappaB-driven gene expression. J. Biol. Chem. 291, 25553–25566 (2016).
Bravard, A. et al. Inactivation by oxidation and recruitment into stress granules of hOGG1 but not APE1 in human cells exposed to sub-lethal concentrations of cadmium. Mutat. Res. 685, 61–69 (2010).
Ba, X. et al. 8-Oxoguanine DNA glycosylase-1 augments proinflammatory gene expression by facilitating the recruitment of site-specific transcription factors. J. Immunol. 192, 2384–2394 (2014).
Ba, X. & Boldogh, I. 8-Oxoguanine DNA glycosylase 1: beyond repair of the oxidatively modified base lesions. Redox Biol. 14, 669–678 (2018).
Ke, Y. et al. PARP1 promotes gene expression at the post-transcriptiona level by modulating the RNA-binding protein HuR. Nat. Commun. 8, 14632 (2017).
Pan, L. et al. OGG1-DNA interactions facilitate NF-kappaB binding to DNA targets. Sci. Rep. 7, 43297 (2017).
Hajas, G. et al. 8-Oxoguanine DNA glycosylase-1 links DNA repair to cellular signaling via the activation of the small GTPase Rac1. Free Radic. Biol. Med. 61, 384–394 (2013).
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (grant nos.: 31571339 and 31371293 to X.B.), the Program for Introducing Talent to Universities (grant no. B07017 to X.B.), The State Scholarship Fund of China Scholarship Council (no. 201706625004 to X.B.), United States National Institute of Environmental Health and Sciences (grant no.: RO1 ES018948 to I.B.), and United States National Institute of Allergic and Infectious Diseases (grant no.: AI062885 to I.B.). We thank Dr. Lijuan Zhu (School of Foregin Language, Northeast Normal University) for her critically editing the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by M. Malewicz
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, R., Li, C., Qiao, P. et al. OGG1-initiated base excision repair exacerbates oxidative stress-induced parthanatos. Cell Death Dis 9, 628 (2018). https://doi.org/10.1038/s41419-018-0680-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-018-0680-0
This article is cited by
-
Covalent PARylation of DNA base excision repair proteins regulates DNA demethylation
Nature Communications (2024)
-
Aggregability of the SQSTM1/p62-based aggresome-like induced structures determines the sensitivity to parthanatos
Cell Death Discovery (2024)
-
NINJ1 is activated by cell swelling to regulate plasma membrane permeabilization during regulated necrosis
Cell Death & Disease (2023)
-
The natural product dehydrocurvularin induces apoptosis of gastric cancer cells by activating PARP-1 and caspase-3
Apoptosis (2023)
-
Inhibition of AKT induces p53/SIRT6/PARP1-dependent parthanatos to suppress tumor growth
Cell Communication and Signaling (2022)