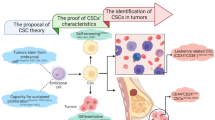
Abstract
Therapy-induced expansion of cancer stem cells (CSCs) has been identified as one of the most critical factors contributing to therapeutic resistance, but the mechanisms of this adaptation are not fully understood. UHRF1 is a key epigenetic regulator responsible for therapeutic resistance, and controls the self-renewal of stem cells. In the present study, taxane-resistant cancer cells were established and stem-like cancer cells were expanded. UHRF1 was overexpressed in the taxane-resistant cancer cells, which maintained CSC characteristics. UHRF1 depletion overcame taxane resistance in vitro and in vivo. Additionally, FOXM1 has been reported to play a role in therapeutic resistance and the self-renewal of CSCs. FOXM1 and UHRF1 are highly correlated in prostate cancer tissues and cells, FOXM1 regulates CSCs by regulating uhrf1 gene transcription in an E2F-independent manner, and FOXM1 protein directly binds to the FKH motifs at the uhrf1 gene promoter. This present study clarified a novel mechanism by which FOXM1 controls CSCs and taxane resistance through a UHRF1-mediated signaling pathway, and validated FOXM1 and UHRF1 as two potential therapeutic targets to overcome taxane resistance.
Similar content being viewed by others
Background
Taxane, including paclitaxel (Taxol), and docetaxel (Taxotere), has been widely used in cancer chemotherapy. Taxol has a significant role in the treatment of ovarian, breast, lung, head and neck, esophageal, prostate and bladder cancers, and Taxotere is effective in the treatment of breast, lung, head and neck, gastric, ovarian, and bladder cancers. Taxanes bind to β-tubulin, thereby reducing depolymerization. By stabilizing microtubules and dampening microtubule dynamics, taxanes prevent the formation of mitotic spindles, and chronically activate the spindle assembly checkpoint (SAC), which in turn leads to mitotic arrest and eventually induces cell death1,2. However, cancer cells develop resistance to taxanes. The molecular mechanisms by which cancer cells develop taxane resistance are not fully understood.
Taxane resistance is subclassified as innate resistance and acquired resistance. Acquired resistance results from the increased expression of drug efflux proteins such as ATP-binding cassette (ABC transporters)3, the altered expression and function of certain tubulin isotypes4, and the deregulation of Bcl-2 molecules5,6. Importantly, taxanes induced the expansion of stem-cell-like cancer cells, resulting in the development of taxane resistance and cancer relapse7.
FOXM1 is a cell proliferation-specific transcription factor that regulates the transcription of genes critical for the G1/S and G2/M cell cycle transition8,9,10. In addition to its roles as an oncogene11, FOXM1 overexpression is critical to the development of taxane resistance12,13. Several mechanisms have been reported for taxane resistance. FOXM1 increases drug efflux due to the upregulation of abcc5 gene transcription3, promotes DNA damage repair through the transcriptional regulation of DNA repair genes14, drives abnormal mitotic spindle formation and mitotic catastrophe5 and upregulates apoptosis-associated molecules such as XIAP and Survivin15. In addition, FOXM1 regulates the stemness and self-renewal of cancer stem cells (CSCs) by directly regulating the gene transcription of CSC-associated genes16, or the crosstalk with CSC signaling pathways such as Wnt/β-Catenin17,18. The regulation of CSC expansion by FOXM1 is crucial for the development of taxane-resistance.
Compelling evidence suggests that the ubiquitin-like PHD and RING finger domain containing 1 (UHRF1), a key epigenetic regulator of DNA methylation, also contributes to the development of therapeutic resistance, including chemoresistance19,20 and radioresistance21,22. UHRF1 promotes DNA damage repair by regulating multiple DNA damage repair pathways, such as homologous recombination and the nonhomologous end joining (NHEJ) double-strand DNA repair pathway23. Additionally, UHRF1 controls the self-renewal and differentiation of stem cells24. Recent studies suggest that UHRF1 controls the self-renewal versus differentiation of hematopoietic stem cells by epigenetically regulating the cell-division modes25. Targeted deletion of uhrf1 in epithelial basal stem cells results in premature cell senescence after injury without affecting cell survival or inducing premature differentiation26. However, no report is available about its functions in CSCs. RNA-seq data from recent studies indicated that UHRF1 might be regulated by FOXM1, and promoted the development of esophageal adenocarcinoma27. Whether FOXM1 regulates the maintenance and expansion of CSCs through a UHRF1-mediated signaling pathway is unknown.
In this study, we first established taxane-resistant cancer cells by long-term treatment with low doses of taxane. The stem-like cancer cells were expanded as taxane-resistant cancer cells. FOXM1 and UHRF1 were overexpressed in the taxane-resistant cancer cells, and positively regulated the maintenance of CSCs. FOXM1 and UHRF1 are also consistently expressed in prostate cancer tumor specimens and cells, with high correlation between the two molecules. Furthermore, we found that FOXM1 regulates CSCs and taxane resistance by directly regulating uhrf1 gene transcription.
Results
Cancer cells developed taxane-resistance after long-term and intermittent exposure
We previously developed a paclitaxel-resistant cell line, CNE2TR, by intermittently exposing CNE2 cells to low doses of paclitaxel over a long period3,28. In this study, we developed another docetaxel-resistant DU145 prostate cancer cell line (DU145-DR) using similar methods. We compared the drug sensitivity of DU145-DR cells to parental DU145 cells. The IC50 values of docetaxel in DU145-DR cells were significantly higher than DU145 (54.55 vs. 30.66 nM) (Fig. 1a). We tested the response of DU145 and DU145-DR cells to different doses of docetaxel (50 or 100 nM) over time. Cell viability significantly decreased with time. Comparatively, docetaxel killed more DU145 cells than DU145-DR cells 72 h after treatments (Fig. 1b). With docetaxel treatment in stepwise concentrations as shown in Fig. 1c for 48 h, more DU145-DR cell colonies formed than parental DU145 cells three weeks later. DU145-DR cells demonstrated much stronger resistance to docetaxel-induced cell killing (Fig. 1c).
Docetaxel-resistant DU145 cells (DU145-DR) were induced. a The IC50 values of docetaxel were compared between DU145 and DU145-DR cells. The cells were treated with docetaxel at the doses shown. MTS assays were used to test cell viability 72 h after treatment. After three repeats for each dose, the relative cell survival was calculated (treatment vs. control) to compare the IC50 values. b Cell response to different doses of docetaxel with time. DU145 and DU145-DR cells were treated with paclitaxel at 50 or 100 nM individually, and cell viability was tested by a MTS assay 24, 48, and 72 h after treatment. The relative cell viability represents a ratio of docetaxel vs. control. c Cell colony formation assay. DU145 and DU145-DR cells were treated with docetaxel at stepwise concentrations for 48 h. One thousand cells were re-seeded in 6-well plates, and the cell clones formed were stained with crystal violet. The number of colonies was counted and the relative colony formation ability was analyzed 21 days after cell seeding
Docetaxel-resistant cancer cells acquired CSC characteristics
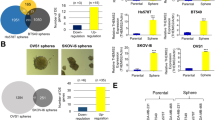
We first tested the proportion of CSCs among DU145-DR and DU145 cell populations. The percentage of CD44highCD133high cells in the DU145-DR population markedly increased compared to DU145 cells (8.82 vs. 5.49%, Fig. 2a). Cell spheres formed by DU145 cells were fewer and smaller than those formed by DU145-DR cells (Fig. 2b), and the protein levels of ALDH1, SOX2, and Sonic Hedgehog (SHH), typical stem cell markers in DU145-DR cells, were much higher than in the parental DU145 cells (Fig. 2c), indicating that the subgroup of docetaxel-resistant DU145-DR cells had acquired CSC characteristics. We also observed elevated protein levels of ALDH1, SOX2, and SHH in paclitaxel-resistant CNE2TR cells compared to CNE2 cells (Fig. 2d). These data indicated that the acquisition of CSC characteristics by a subpopulation of cancer cells contributes to taxane resistance.
a The CSC sub-population. DU145 and DU145-DR cells were labeled with fluorescent antibodies against CD44 (APC) and CD133 (PE). CD44highCD133high cells were detected by flow cytometry. b The same number of DU145 and DU145-DR cells were plated in soft agar, and cell sphere formation was observed 21 days after cell seeding. The spheres were counted in five randomly selected fields, and the difference of spheres in two groups was compared by statistical analysis. c The protein levels of CSC-associated molecules such as ALDH1, SOX2, and SHH were tested in DU145 and DU145-DR cells by Western blot. d The protein levels of ALDH1, SOX2, and SHH were tested in CNE2 and CNE2TR cells by Western blot
UHRF1 is overexpressed in taxane-resistant cancer cells, which contributes to maintenance of the CSC phenotype
UHRF1 has reported roles in the maintenance of both self-renewal and differentiation of hematopoietic stem cells25. We compared the protein levels of UHRF1 in CNE2 and CNE2TR cells, DU145 and DU145-DR cells, and SKOV3 and SKOV3R cells. Consistently in three pairs of cell lines, UHRF1 protein levels were significantly higher in docetaxel-resistant cells than the parental cancer cells (Fig. 3a). To validate the roles of UHRF1 in the regulation of CSCs, we depleted UHRF1 in DU145-DR and CNE2TR cells, and assessed the sub-population of CD44highCD133high cells. The depletion of UHRF1 significantly decreased the CD44highCD133high cell sub-population (Fig. 3b). UHRF1 depletion consistently and significantly decreased sphere formation by the two taxane-resistant cancer cell lines compared to the parental cancer cells (Fig. 3c). We also tested the protein levels of CSC-associated molecules ALDH1, NANOG, SOX2, and SHH with the depletion of UHRF1. Consistently in the DU145-DR and CNE2TR docetaxel-resistant cancer cells, protein levels of CSC-associated molecules declined with UHRF1 knockdown (Fig. 3d). To validate UHRF1 regulation of CSC-associated molecules, we tested protein levels when UHRF1 was overexpressed in the parental DU145 or CNE2 cells by transient transfection. The overexpression of UHRF1 significantly elevated the levels of CSC-associated molecules (Fig. 3e). These data identified the roles of UHRF1 in the maintenance of CSC characteristics.
a The protein levels of UHRF1 were analyzed in CNE2 and CNE2TR cells, DU145 and DU145-DR cells, and SKOV3 and SKOV3R cells by Western blot. b The depletion of UHRF1 in DU145-DR and CNE2TR cells decreased the CSC sub-population. DU145-DR and CNE2TR cells were infected with lentivirus-delivered shRNA-UHRF1 or shRNA-CTR. Cells were labeled with fluorescent antibodies against CD44 (APC) and CD133 (PE). CD44highCD133high cells were detected by flow cytometry. c CNE2TR and DU145-DR cells were infected with lentivirus-delivered shRNA-UHRF1 or shRNA-CTR. One thousand cells were re-seeded in culture media-mixed agarose, and the cell spheres formed were stained with crystal violet and analyzed 21 days after cell seeding. The spheres were counted in five randomly selected fields, and the difference of spheres in two groups was compared by statistical analysis. d UHRF1 depletion decreased the expression of CSC-associated molecules ALDH1, NANOG, SOX2, and SHH in CNE2TR and DU145-DR cells. e UHRF1 overexpression elevated the levels of CSC-associated molecules ALDH1, NANOG, SOX2 and SHH in CNE2, and DU145 cells
UHRF1 depletion re-sensitized cancer cells to docetaxel
Since UHRF1 plays a critical role in docetaxel resistance and maintenance of CSC characteristics, we depleted UHRF1 in DU145-DR cells by infecting cells with lentivirus-delivered shRNA-UHRF1 or shRNA-CTR. DU145-DR cells with UHRF1 stable depletion were selected by puromycin. Compared to DU145-DR cells expressing the control shRNA, the cells expressing sh-UHRF1 were more sensitive to docetaxel-induced cell killing (Fig. 4a, IC50 65.89 vs. 44.58 nM).
a DU145-DR cells were infected with lentivirus-delivered shRNA-UHRF1 or shRNA-CTR, followed by puromycin drug selection. The IC50 values of docetaxel were compared between DU145-DR cells expressing the control shRNA and sh-UHRF1. b Tumor xenografts were established in immune-deficient nude mice with DU145-DR or UHRF1-depleted DU145-DR cells. Nude mice bearing tumor xenografts were treated with or without docetaxel (20 mg/kg) by i.p. injection once a week. The sizes of tumor xenografts were measured every 3 days. c The tumor masses were harvested from immune-deficient nude mice at the experimental endpoint. d The weight of tumors in each group was averaged and compared. e The tumor masses were immersed in embedding reagent OCT and frozen in liquid nitrogen. The frozen tissues were sectioned and stained with a One Step TUNEL Apoptosis Assay Kit. The nuclei were stained with DAPI. The TUNEL-positive red fluorescent cells were observed by fluorescent microscopy with 550 nm excitation and 570 nm emission wavelengths
To confirm the in vitro results, we tested whether UHRF1 depletion reversed docetaxel resistance in vivo. Tumor xenografts were established in immune-deficient nude mice by inoculating DU145-DR or UHRF1-depleted DU145-DR cells. The nude mice bearing tumor xenografts were treated with or without docetaxel (20 mg/kg) by i.p. injection. The sizes of tumor xenografts containing UHRF1-depleted DU145-DR cells and treated with docetaxel were much smaller than those containing UHRF1-depleted DU145-DR cells or treated with docetaxel alone (Fig. 4b). The tumor masses were harvested at the endpoint of the experiment, and tumor size and weight were compared. The tumor size and average weight of tumors treated with UHRF1 depletion and docetaxel were much less than with the single treatment (Fig. 4c, d). The harvested tumors were rapidly frozen and sectioned, and the apoptotic cells inside the tumors were labeled with TUNEL staining. There were many more TUNEL-positive apoptotic cells in the double treatment group than with single treatment (Fig. 4e). The results showed that UHRF1 depletion significantly re-sensitized tumor xenografts containing docetaxel-resistant cells to docetaxel treatment.
FOXM1 is overexpressed in docetaxel-resistant cancer cells, contributing to the maintenance of the CSC phenotype
FOXM1 has been reported to affect drug resistance, including docetaxel resistance29,30. Additionally, FOXM1 contributes to drug resistance through the maintenance of CSCs18,31. We first compared FOXM1 protein levels between the docetaxel-resistant and parental cancer cells. Consistently in three pairs of cell lines, FOXM1 protein levels were significantly higher in docetaxel-resistant cells than in parental cancer cells (Fig. 5a). To validate the roles of FOXM1 in the regulation of CSCs, we depleted FOXM1 in docetaxel-resistant CNE2TR and DU145-DR cells, and assessed the protein levels of CSC-associated molecules ALDH1, NANOG, SOX2, and SHH. Consistently in the taxane-resistant DU145-DR and CNE2TR cancer cells, the protein levels of CSC-associated molecules declined with the knockdown of FOXM1 (Fig. 5b). Conversely, we tested the protein levels of CSC-associated molecules when FOXM1 was ectopically overexpressed in DU145 and CNE2 cells by transient transfection. As shown in Fig. 5c, the protein levels of CSC-associated molecules were elevated after ectopic overexpression of FOXM1. These data validated the critical roles of FOXM1 in the maintenance of the CSC phenotype.
a The protein levels of FOXM1 were analyzed in CNE2 and CNE2TR cells, DU145 and DU145-DR cells, and SKOV3 and SKOV3R cells by Western blot. b FOXM1 depletion decreased the expression of CSC-associated molecules ALDH1, NANOG, SOX2 and SHH in CNE2TR, and DU145-DR cells. c The ectopic overexpression of FOXM1 in DU145 or CNE2 cells elevated the expression levels of CSC-associated molecules ALDH1, NANOG, SOX2, and SHH
FOXM1 and UHRF1 were consistently expressed in prostate cancer tumor tissues and cell lines, and FOXM1 regulates UHRF1 expression
We analyzed FOXM1 and UHRF1 protein expression levels in 546 prostate tumor tissues using the TCGA data. The prostate cancer tumors were classified in three grades by Gleason score. The patients in Gleason 1–4 were identified as low grade, 5–7 as middle grade, and 8–10 as high grade. The results showed that FOXM1 and UHRF1 expression levels increased along with the Gleason scores (Fig. 6a). FOXM1 and UHRF1 protein levels showed a strong positive correlation in prostate cancer tumor tissues (Fig. 6b, R = 0.6927). Additionally, we assessed the expression of FOXM1 and UHRF1 proteins in a panel of prostate cancer cell lines and non-malignant prostate epithelial cells. FOXM1 or UHRF1 expression levels showed high correlation in prostate cancer cells and non-malignant prostate epithelial cells (Fig. 6c).
FOXM1 and UHRF1 protein expression levels were analyzed in 546 prostate cancer tumor tissues using the TCGA data. The prostate cancer tumors were subclassified in three grades by Gleason score (low: 1–4, middle: 5–7, and high 8–10). a FOXM1 and UHRF1 expression levels were up-regulated as Gleason scores increased. b FOXM1 and UHRF1 protein levels showed a strong positive correlation in prostate cancer tissues (R = 0.6927). c FOXM1 and UHRF1 expression showed a high correlation in prostate cancer cells and normal prostate epithelial cells. d FOXM1 maintains CSC characteristics by regulating UHRF1. FOXM1 was depleted with a small molecule inhibitor, siomycin A, and UHRF1 was elevated by transient transfection. The expression levels of CSC-associated molecules ALDH1, SOX2, and SHH were tested by Western blot. e The depletion of FOXM1 with siRNA in CNE2TR and DU145-DR cells decreased the mRNA and protein levels of UHRF1. f The upregulation of FOXM1 in CNE2 and DU145 cells elevated the UHRF1 levels
Since both FOXM1 and UHRF1 regulate the CSC phenotype, we asked whether FOXM1 regulates the CSC phenotype through UHRF1-associated signaling pathways. We treated DU145-DR cells with a small molecule inhibitor of FOXM1 siomycin A. Consistent with FOXM1 siRNA, siomycin A significantly decreased the protein levels of CSC-associated molecules ALDH1, SOX2 and SHH, and the ectopic overexpression of UHRF1 reversed the siomycin A-induced decreases in CSC-associated molecules (Fig. 6d). The results showed that FOXM1 regulates the CSC phenotype through UHRF1. In DU145-DR and CNE2TR cells, we depleted FOXM1 with siRNA, and tested the protein expression of UHRF1. The depletion of FOXM1 significantly decreased the mRNA and protein levels of UHRF1 (Fig. 6e). Conversely, we tested the protein levels of UHRF1 in DU145 and CNE2 cells when FOXM1 was ectopically overexpressed by transient transfection. FOXM1 elevation increased the expression level of UHRF1 (Fig. 6f). The data suggested that FOXM1 regulates the gene expression of UHRF1.
FOXM1 regulates uhrf1 gene transcription by directly binding to the uhrf1 gene promoter
The gene transcription of uhrf1 is reported to be regulated by E2F molecules32. To clarify whether FOXM1 regulates uhrf1 gene transcription through the E2F pathway, we tested the expression of E2F1, E2F2, E2F3, and E2F8 when FOXM1 was depleted in DU145 and PC3 cells with siRNA. We did not observe a clear impact on E2F molecules at the mRNA and protein levels (Fig. 7a,b). We further tested the impact of FOXM1 on the promoter activity of uhrf1 gene. A plasmid using a 2000bp fragment of uhrf1 gene promoter controlling the luciferase gene was constructed. We tested the impact of FOXM1 on uhrf1 gene promoter activity in HEK-293 cells when FOXM1 was depleted with siRNA. The depletion of FOXM1 significantly decreased the uhrf1 gene promoter activity (Fig. 7c). We further tested the DNA binding of FOXM1 to uhrf1 gene promoter by ChIP-qPCR. The forkhead box (FKH) consensus motif was identified in the uhrf1 gene promoter, and the FOXM1 protein-bound DNA in HEK-293 and CNE2TR cells was purified by ChIP, and specific PCR primers spanning the FKH binding motif were designed. Consistently, FOXM1 directly bound to the uhrf1 gene promoter (Fig. 7d). These data verified that FOXM1 regulates uhrf1 gene transcription by direct DNA binding to the FKH motif at the uhrf1 gene promoter.
a FOXM1 regulates uhrf1 gene transcription, but not through the E2F pathway. FOXM1 was depleted with siRNA in DU145 cells, and the mRNA levels of uhrf1, e2f1, e2f2, e2f3, and e2f8 genes were tested by RT-PCR. The protein expression of E2F1 and E2F3 was tested when FOXM1 was depleted with siRNA. b FOXM1 was depleted with siRNA in PC-3 cells, and the mRNA levels of uhrf1, e2f1, e2f2, e2f3, and e2f8 genes were tested by RT-PCR. The protein expression of E2F1 and E2F3 was tested when FOXM1 was depleted with siRNA. c The depletion of FOXM1 with siRNA in HEK-293T cells decreased uhrf1 gene promoter activity. d FOXM1 directly binds to uhrf1 gene promoter. The FOXM1 protein-bound DNA in HEK-293T and CNE2TR cells was purified by ChIP, and primers spanning the FKH binding motif at the uhrf1 gene promoter were designed. FOXM1 protein binding to the uhrf1 gene promoter was detected by ChIP-PCR
Discussion
The expansion of CSCs after treatment has been identified as one of the most important factors responsible for acquired therapeutic resistance. To study the association of taxane resistance and CSCs, we generated taxane-resistant cancer cells by treating cells with a high dose of taxane, and maintaining low dose treatment over a long period (Fig. 1). The subpopulation of CSCs significantly increased along with the establishment of acquired taxane resistance, and CSC-associated molecules such as ALDH1, SOX2, and SHH were remarkably elevated (Fig. 2). These results showed that the expansion of the CSC subpopulation is indeed a critical factor in the development of taxane resistance.
FOXM1 and UHRF1 both have reported roles in therapeutic resistance, including taxane resistance, as well as in the regulation of stem cell self-renewal and differentiation 5,13,17,19,21. In addition, FOXM1 and UHRF1 are highly correlated in tumor specimens and prostate cancer cell lines (Fig. 6). In our previous studies, we found that FOXM1 is overexpressed in paclitaxel-resistant cancer cells, and FOXM1 depletion overcame the paclitaxel-resistance by decreasing drug efflux3. In this study, FOXM1 is closely associated with the stemness of taxane-resistant cancer cells. FOXM1 is overexpressed in taxane-resistant cancer cells, and the depletion or elevation of FOXM1 accordingly changed the expression levels of CSC-associated molecules (Fig. 5). These results showed that FOXM1-regulated CSC stemness was responsible for the taxane-resistance. However, the roles of UHRF1 in the expansion of CSCs in taxane-resistance are not well known. In this study, UHRF1 overexpression was detected in taxane-resistant cancer cells, and the depletion of UHRF1 correspondingly lowered the expression of CSC-associated molecules and decreased the subpopulation of CSCs and sphere formation ability. Furthermore, UHRF1 depletion significantly promoted sensitivity to docetaxel in a prostate cancer xenograft model (Fig. 4). This is the first report that UHRF1 plays a critical role in the expansion of CSCs and development of acquired therapeutic resistance.
Gene transcription of uhrf1 is regulated by several identified transcription factors. E2F1 and E2F8 are two well-known transcription factors that control UHRF1 expression33,34. Specificity protein 1 (SP1) is a transcription factor directly binding to uhrf1 gene promoter, and 3,3′,5-Triiodo-L-thyronine (T3)/thyroid receptor (TR) downregulated UHRF1 in HepG2 cells by repressing SP1 binding35. Transcription factor Yingyang 1 (YY1) is a mediator for G9a recruitment binding to the uhrf1 gene promoter, and represses uhrf1 gene transcription in the H1299 lung cancer cell line36. A recent study indicated by ChIP-seq that UHRF1 may be a direct target of FOXM1 transcription factor in the esophageal adenocarcinoma-derived OE33 cell line27. However, the exact mechanism is elusive. Our present study addressed whether FOXM1 regulates taxane resistance and CSCs through a UHRF1-mediated signaling pathway. In our present report, the depletion or elevation of FOXM1 significantly influenced UHRF1 expression at both the mRNA and protein levels. Siomycin A, a small molecule inhibitor of FOXM1, decreased the expression of CSC-associated molecules, while UHRF1 overexpression reversed the decrease of CSC-associated molecules. The results suggested that FOXM1 regulates the stemness of CSCs through a UHRF1-mediated signaling pathway (Fig. 6). Our further studies clarified that FOXM1 regulates uhrf1 gene transcription in an E2F-independent manner, and by directly binding to the FKH motif at the promoter.
Altogether, our results in this study identified UHRF1 as a critical regulator of CSCs and taxane resistance, and demonstrated that FOXM1 is an important transcription factor regulating both the stemness of CSCs and taxane resistance by regulating uhrf1 gene transcription. FOXM1 and UHRF1 may both be therapeutic targets to overcome taxane resistance.
Materials and methods
Cell culture
Prostate cancer cell lines PC3 and DU145, and ovarian cancer cell line SKOV3 cells were purchased from ATCC (Manassas, MA, USA). Nasopharyngeal cancer (NPC) cell line CNE2 was purchased from the Cancer Research Institute of Central South University (Changsha, Hunan, China). The paclitaxel resistant cell line CNE2TR were generated by the methods as described in a previous publication37. The docetaxel resistant cell line DU145-DR was generated by treating cells with high doses of docetaxel (50 nM), and maintaining the residual colonies at low doses (20 nM) over 12 weeks. SKOV3R cells were kindly gifted from Dr. Yu Zhang from Xiangya Hospital Central South University. These cells were cultured in RPMI-1640 media supplemented with 10% FBS and 1% streptomycin/penicillin.
Plasmids, siRNA, and shRNA
FOXM1 and UHRF1 cDNAs in pCMV-XL5 vector were purchased from Origene (Rockville, MD, USA). A 2000 bp uhrf1 gene promoter was obtained by the PCR method using genomic DNA as the template, and then subcloned to PCR2.1 vector (Thermo Fisher Scientific, Shanghai, China). pGL3-UHRF1-Luc was generated by cutting the uhrf1 gene promoter fragment with Kpn1 and HindIII, then inserting to the pGL3-basic vector. The sequences of siRNAs of FOXM1 and UHRF1 were as follows: the sense sequence of siRNA FOXM1: 5′-CUCUUCUCCCUCAGAUAUATT-3′, and the antisense sequence: 5′-UAUAUGAGGGAGAGTT-3′; the sense sequence of siRNA URHF1 #1: 5′-GCGCUGGCUCUCAACUGCU-3′, and the antisense sequence: AGCAGUUGAGCCAGCGC-3′; the sense sequence of siRNA UHRF1 #2: 5′-GCAUCUACAAGGUUGUGAA-3′, and the antisense sequence: 5′-UUCACAACCUUGUAGAUGC-3′. The primers 5′-CCGG–GCGCUGGCUCUCAACUGCU-CTCGAG-AGCAGTTGAGAGCCAGCGC-TTTTT-3′ were designed to synthesize UHRF1shRNA.
Transient transfection
cDNA transient transfection was conducted using lipofectamin 2000 (Thermo Fisher Scientific, Wilmington, DE, USA). siRNA transfection was conducted using DharmaFECT Transfection Reagent (Thermo Fisher Scientific). The experimental protocol was modified from the manufacturer’s manuals.
Antibodies and chemicals
The primary antibodies anti-FOXM1, anti-UHRF1, anti-ALDH1, anti-NANOG, anti-E2F1, anti-E2F3, anti-β-ACTIN were purchased from Santa Cruz Biotechnology (Dallas, UT, USA). Anti-SHH antibody was purchased from Cell Signal Transduction (CST, Danvers, MA, USA) and anti-SOX2 was purchased from ABclonal (Woburn, MA, USA). The small molecule inhibitor of FOXM1 Siomycin A was purchased from Sigma Aldrich (St. Louis, MO, USA) and Docetaxel was purchased from Selleck Chemicals (Houston, Texas, USA).
RT-PCR
The total RNA was extracted from cells using a RNAiso Plus kit (Takara Bio Inc, Shiga, Japan). The concentration of RNA was measured by spectrophotometer and the RNA was reverse transcripted to cDNA (Takara Bio Inc). Quantitative real-time PCR was performed using the Applied Biosystems 7500 Real Time PCR system (Thermo Fisher Scientific).
Flow cytometric analysis
Cells suspended with PBS were inoculated with anti-CD133 (PE-conjugated, MiltenyiBiotec, San Diego, CA, USA) and anti-CD44 (APC-conjugated, BD PharMingen, San Jose, CA, USA) at 37 °C for 20 min. The positive cells were analyzed by flow cytometry (Millipore, Temecula, CA, USA).
Luciferase reporter assay
HEK-293T cells were transfected with pGL3-UHRF1-Luc, together with siRNA-FOXM1 or siRNA-CTR (control) using the TransIT-X2 Dynamic Delivery System (Mirus, Madison, WI, USA), with pRL-SV40 as a transfection efficiency control. The cells were frozen/thawed for two cycles in lysis buffer (Genecopoeia, Rockville, MD, USA), and firefly/Renilla luciferase activities were tested by luminometer.
Cell proliferation assay (MTS)
The cells were plated on 96-well plates (5000 cells per well) and treated with 50 or 100 nM docetaxel. The cells were stained with MTS solution [3-(4, 5-dimethylthiazol-2-yl)-5-(3-carboxymethoxyphenyl)-2-(4-sulfophenyl)-2H-tetrazolium] (Promega, Madison, WI, USA) at 37 °C in 5% CO2 for 1 h. Cell viability was assessed by measuring the 490 nM absorbance in a spectrometer.
Colony formation assay
DU145 and DU145-DR cells were seeded in 6-well plates (500 cells per well) and exposed to docetaxel at different concentrations (0, 2, 4, 6, 8, 10 nM) for 48 h. Docetaxel was washed away and then the cells were maintained for another 2 weeks for colony formation. The cell colonies were fixed with 3.7% paraformaldehyde and stained with 1% crystal violet solution. The cell colonies were dissolved in 1% SDS, and cell survival was evaluated by measuring the absorbance at 570 nM.
Chromatin immunoprecipitation (ChIP)-PCR
HEK-293T or CNE2TR cells were plated in 100 mm culture dishes. The cells were cross-linked with 1% paraformaldehyde when cell confluence attained nearly 90%. The cells were lysed and the DNA fragment was sonicated to shear a mean DNA fragment size of about 500 bp. The chromatin–protein complex was incubated with FOXM1 antibodies on rotating platform overnight at 4 °C, with IgG as the negative control. Five percent of the total lysate was used for input control. The immune complexes were harvested with beads, and then de-crosslinked and the chromatin DNA fragments purified. The degree of FOXM binding to chromatin DNA was analyzed by semi-quantitative PCR and quantitative RT-PCR. The PCR primers spanning the FKH binding motif at the uhrf1 gene promoter were designed as follows: Primer 1 forward: AAAGACAGCAAACAAGCCCTG, and reverse: CTCGCACGCATTGACCAGTA; Primer 2 forward: CACTTGGTTGAGTTCCCCCG, reverse: GAAGGTCCAACCCATCCCTC. The binding efficiency of FOXM1 was calculated by following the calculation formula described in a previous publication38.
Animal experiments
The animal experiment protocol was approved by the Ethics Committee of Xiangya Hospital Central South University. DU145-DR or UHRF1-depleted DU145-DR cells were injected (5 × 106/100 μL) into the subcutaneous tissue of the left flank region of 6-week-old immune-deficient nude mice (BALB/C-nu/nu, SLAC Laboratory, Shanghai, China). The nude mice bearing tumor xenografts were treated with or without docetaxel (20 mg/kg) by i.p. injection once a week. The sizes of tumor xenografts were measured every 3 days, and volume (V) was calculated using the following formula: V = ab2/2 (a: the long diameter and b: the short diameter). The mice were sacrificed when tumor volume reached 1000 mm3 or the 28 days after treatments. The tumor masses were harvested at the experiment endpoint, and the size and weight of tumors in each group was averaged and compared.
TUNEL assay
The tumor masses were harvested from subcutaneous xenografts of nude mice at the experiment endpoint, and immediately immersed in OCT solution and frozen in liquid nitrogen. The frozen tissues were sectioned and stained using a One Step TUNEL Apoptosis Assay Kit (Beyotime, Shanghai, China) according to the manufacturer’s instruction. The nuclei were stained with DAPI. TUNEL-positive red fluorescent cells were observed by fluorescent microscopy with 550 nm excitation and 570 nm emission wavelengths.
TCGA data analysis
For mRNA sequencing data, the gene level and exon level quantification in Fragmants Per Kilobase of transcript per Million mapped reads (FPKM) were generated by Genomic Data Commons (GDC). These data can be downloaded from the Cancer Genome Atlas website (https://cancergenome.nih.gov/). UHRF1 and FOXM1 levels were analyzed according to the prostate cancer Gleason scores, and the correlation analysis of FOXM1 and UHRF1 molecules was calculated using GraphPad 6.0 (GraphPad Software, Inc. La Jolla, CA, USA).
Statistics
All in vitro experiments were done at least in triplicate. The data were presented as the mean ± SD. All statistical analysis was conducted with SPSS software. The statistical difference between two samples was analyzed by Students t test. The comparison of tumor sizes in four groups in the animal study was analyzed using one-way ANOVA. *P < 0.05; **P < 0.01 was considered statistically significant.
References
Jordan, M. A. & Wilson, L. Microtubules as a target for anticancer drugs. Nat. Rev. Cancer 4, 253–265 (2004).
Gascoigne, K. E. & Taylor, S. S. How do anti-mitotic drugs kill cancer cells? J. Cell Sci. 122(Pt 15), 2579–2585 (2009).
Hou, Y. et al. The FOXM1-ABCC5 axis contributes to paclitaxel resistance in nasopharyngeal carcinoma cells. Cell Death Dis. 8, e2659 (2017).
McGrogan, B. T., Gilmartin, B., Carney, D. N. & McCann, A. Taxanes, microtubules and chemoresistant breast cancer. Biochim. Biophys. Acta 1785, 96–132 (2008).
Khongkow, P. et al. Paclitaxel targets FOXM1 to regulate KIF20A in mitotic catastrophe and breast cancer paclitaxel resistance. Oncogene 35, 990–1002 (2016).
Frederiks, C. N., Lam, S. W., Guchelaar, H. J. & Boven, E. Genetic polymorphisms and paclitaxel- or docetaxel-induced toxicities: a systematic review. Cancer Treat. Rev. 41, 935–950 (2015).
Bosco, D. B., Kenworthy, R., Zorio, D. A. & Sang, Q. X. Human mesenchymal stem cells are resistant to Paclitaxel by adopting a non-proliferative fibroblastic state. PLoS ONE 10, e0128511 (2015).
Lam, E. W., Brosens, J. J., Gomes, A. R. & Koo, C. Y. Forkhead box proteins: tuning forks for transcriptional harmony. Nat. Rev. Cancer 13, 482–495 (2013).
Myatt, S. S. & Lam, E. W. Targeting FOXM1. Nat. Rev. Cancer 8, 242 (2008).
Liu, Y., Gong, Z., Sun, L. & Li, X. FOXM1 and androgen receptor co-regulate CDC6 gene transcription and DNA replication in prostate cancer cells. Biochim. Biophys. Acta 1839, 297–305 (2014).
Liu, Y., Hock, J. M., Van Beneden, R. J. & Li, X. Aberrant overexpression of FOXM1 transcription factor plays a critical role in lung carcinogenesis induced by low doses of arsenic. Mol. Carcinog. 53, 380–391 (2014).
Zhao, F. et al. Overexpression of forkhead box protein M1 (FOXM1) in ovarian cancer correlates with poor patient survival and contributes to paclitaxel resistance. PLoS ONE 9, e113478 (2014).
Carr, J. R., Park, H. J., Wang, Z., Kiefer, M. M. & Raychaudhuri, P. FoxM1 mediates resistance to herceptin and paclitaxel. Cancer Res. 70, 5054–5063 (2010).
Zona, S., Bella, L., Burton, M. J., Nestal de Moraes, G. & Lam, E. W. FOXM1: an emerging master regulator of DNA damage response and genotoxic agent resistance. Biochim. Biophys. Acta 1839, 1316–1322 (2014).
Nestal de Moraes, G. et al. targets XIAP and survivin to modulate breast cancer survival and chemoresistance. Cell Signal. 27, 2496–2505 (2015).
Xie, Z. et al. Foxm1 transcription factor is required for maintenance of pluripotency of P19 embryonal carcinoma cells. Nucleic Acids Res. 38, 8027–8038 (2010).
Gong, A. & Huang, S. FoxM. 1 and Wnt/beta-catenin signaling in glioma stem cells. Cancer Res. 72, 5658–5662 (2012).
Yang, N. et al. FOXM1 recruits nuclear Aurora kinase A to participate in a positive feedback loop essential for the self-renewal of breast cancer stem cells. Oncogene 36, 3428–3440 (2017).
Jin, W. et al. UHRF1 inhibits MDR1 gene transcription and sensitizes breast cancer cells to anticancer drugs. Breast Cancer Res. Treat. 124, 39–48 (2010).
Wang, J. et al. Down-regulation of ICBP90 contributes to doxorubicin resistance. Eur. J. Pharmacol. 656, 33–38 (2011).
Li, X., Meng, Q., Rosen, E. M. & Fan, S. UHRF1 confers radioresistance to human breast cancer cells. Int. J. Radiat. Biol. 87, 263–273 (2011).
Yang, C. et al. Inhibiting UHRF1 expression enhances radiosensitivity in human esophageal squamous cell carcinoma. Mol. Biol. Rep. 40, 5225–5235 (2013).
Zhang, H. et al. A cell cycle-dependent BRCA1-UHRF1 cascade regulates DNA double-strand break repair pathway choice. Nat. Commun. 7, 10201 (2016).
Sen, G. L., Reuter, J. A., Webster, D. E., Zhu, L. & Khavari, P. A. DNMT1 maintains progenitor function in self-renewing somatic tissue. Nature 463, 563–567 (2010).
Zhao, J. et al. Uhrf1 controls the self-renewal versus differentiation of hematopoietic stem cells by epigenetically regulating the cell-division modes. Proc. Natl Acad. Sci. USA 114, E142–E151 (2017).
Xiang, H. et al. UHRF1 is required for basal stem cell proliferation in response to airway injury. Cell Discov. 3, 17019 (2017).
Wiseman, E. F. et al. Deregulation of the FOXM1 target gene network and its coregulatory partners in oesophageal adenocarcinoma. Mol. Cancer 14, 69 (2015).
Zhou, Z. et al. FOXC2 promotes chemoresistance in nasopharyngeal carcinomas via induction of epithelial mesenchymal transition. Cancer Lett. 363, 137–145 (2015).
Li, X. et al. FOXM1 mediates resistance to docetaxel in gastric cancer via up-regulating Stathmin. J. Cell. Mol. Med. 18, 811–823 (2014).
Wang, K., Zhu, X., Zhang, K., Zhu, L. & Zhou, F. FoxM1 inhibition enhances chemosensitivity of docetaxel-resistant A549 cells to docetaxel via activation of JNK/mitochondrial pathway. Acta Biochim. Biophys. Sin. 48, 804–809 (2016).
Zhang, N. et al. FoxM1 promotes beta-catenin nuclear localization and controls Wnt target-gene expression and glioma tumorigenesis. Cancer Cell 20, 427–442 (2011).
Kofunato, Y. et al. UHRF1 expression is upregulated and associated with cellular proliferation in colorectal cancer. Oncol. Rep. 28, 1997–2002 (2012).
Park S. A. et al. E2F8 as a novel therapeutic target for lung cancer. J. Natl Cancer Inst. 107, djv151(2015).
Unoki, M., Nishidate, T. & Nakamura, Y. ICBP90, an E2F-1 target, recruits HDAC1 and binds to methyl-CpG through its SRA domain. Oncogene 23, 7601–7610 (2004).
Wu, S. M. et al. Negative modulation of the epigenetic regulator, UHRF1, by thyroid hormone receptors suppresses liver cancer cell growth. Int. J. Cancer 137, 37–49 (2015).
Kim, K. B. et al. H3K9 methyltransferase G9a negatively regulates UHRF1 transcription during leukemia cell differentiation. Nucleic Acids Res. 43, 3509–3523 (2015).
Zhang, X. et al. Genomic methylation profiling combined with gene expression microarray reveals the aberrant methylation mechanism involved in nasopharyngeal carcinoma taxol resistance. Anti-Cancer Drugs 23, 856–864 (2012).
Qi, J. et al. The E3 ubiquitin ligase Siah2 contributes to castration-resistant prostate cancer by regulation of androgen receptor transcriptional activity. Cancer Cell 23, 332–346 (2013).
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (NSFC, 81572542), the Doctoral Fund of Ministry of Education of China (RFDP, 20130162110029), and Scientific Research Funds for Returned Overseas Chinese Scholars, Ministry of Education of China (SRF for ROCS, SEM, 2014-1685).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yuan, B., Liu, Y., Yu, X. et al. FOXM1 contributes to taxane resistance by regulating UHRF1-controlled cancer cell stemness. Cell Death Dis 9, 562 (2018). https://doi.org/10.1038/s41419-018-0631-9
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-018-0631-9
This article is cited by
-
AKT1 regulates UHRF1 protein stability and promotes the resistance to abiraterone in prostate cancer
Oncogenesis (2023)
-
UHRF1 inhibition epigenetically reprograms cancer stem cells to suppress the tumorigenic phenotype of hepatocellular carcinoma
Cell Death & Disease (2023)
-
Small molecule targeting FOXM1 DNA binding domain exhibits anti-tumor activity in ovarian cancer
Cell Death Discovery (2022)
-
Measuring expression heterogeneity of single-cell cytoskeletal protein complexes
Nature Communications (2021)
-
Response prediction biomarkers and drug combinations of PARP inhibitors in prostate cancer
Acta Pharmacologica Sinica (2021)