Abstract
Chemo-resistance and metastasis of triple negative breast cancer (TNBC) contributed the most of treatment failure in the clinic. MicroRNAs (miRNAs) have been proved to be involved in many biological processes and diseases. In this study, we aimed to determine the role of miR-770 in the regulation of chemo-resistance and metastasis of TNBC. Clinically, miR-770 was highly expressed in chemo-sensitive tissues and predicted a better prognosis of TNBC. Functionally, ectopic expression of miR-770 suppressed the doxorubicin-resistance of TNBC cell lines via regulation of apoptosis and tumor microenvironment, which was mediated by exosomes. Moreover, miR-770 overexpression inhibited the migration and invasion. Rescue of STMN1 could partly reverse the effect of miR-770 in TNBC behaviors. Furthermore, we also demonstrated that overexpression of miR-770 inhibited DOX resistance and metastasis in vivo. Taken together, our results proved that miR-770 could suppress the doxorubicin-resistance and metastasis of TNBC cells, which broaden our insights into the underlying mechanisms in chemo-resistance and metastasis, and provided a new prognostic marker for TNBC cells.
Similar content being viewed by others
Introduction
Breast cancer is one of the most common tumors among women, and the second leading cause of cancer-related death in the world1. Approximately 1 to 1.3 million cases are diagnosed every year, and about 15-20% of patients belongs to the triple negative subtype (TNBC)2. The TNBC was defined as a subtype which lacks of estrogen receptor, progesterone receptor, and human epidermal growth factor receptor type 2 gene expression3. We have previously reported that patients with TNBC have a relatively poorer outcome for the rapid proliferation, early metastasis and lack of molecular targets for treatment4. For TNBC patients, surgery and radiotherapy are employed routinely in a similar way as other types of breast cancer, but adjuvant chemotherapy seemed to be more important for the lack of molecular targets, which became the only systematic treatment5. TNBC could be chemo-sensitive particularly to cytotoxic agents such as anthracyclines and taxanes, but once the chemo-resistance developed, the cells became more aggressive and metastatic6. The metastasis and chemo-resistance of TNBC were the most common causes leading to the treatment failure, disease recurrence and eventual death in clinic7.
Currently, anthracycline-based combination chemotherapy is one of the most important front-line chemotherapeutic agents, generally solely used or combined with other drugs to treat advanced or metastasis breast cancer8. TNBC has been reported to be more sensitive to anthracycline-based chemotherapy compared to endocrinal positive subtypes despite more than 70% of TNBC patients have residual invasive disease after chemotherapy, which partly result from the arisen of chemo-resistance, and only as few as half of the patients may experience the benefits from chemotherapy9,10,11,12. Moreover, studies have reported that the arisen of chemo-resistance may contribute to the metastasis of breast cancers, which further decreased the prognosis of patients6. Thus, it is of great significance to explore the mechanism of chemo-resistance and metastasis.
MicroRNAs (miRNAs) are a class of small non-coding regulatory RNAs that play an important role in various biological processes, including the proliferation, metastasis and chemo-resistance of triple negative breast cancer13,14. Recently, several studies reported miRNAs could play a role not only inside cells but also in the tumor microenvironment15,16. Exosomes are 30 to 100-nm vesicles containing miRNAs, lncRNAs, proteins etc, and released by most cell types, which have been reported to have great significance in the cell-to-cell communications17,18. Previous studies demonstrated that exosomes could influence the chemo-resistance and metastasis of breast cancer via transportation of miRNAs19,20. However, though a few miRNAs have been reported, the definite molecular mechanism of miRNAs function has not been well elucidated in TNBC.
In our study, we detected the miRNAs expression in chemo-sensitive and chemo-resistant tissues by miRNA microarray, and we found miR-770 was significantly decreased in chemo-resistant group. Further experiments proved miR-770 could antagonize the chemo-resistance and metastasis via targeting of STMN1, and modify the tumor microenvironment via transportation to tumor-associated macrophage.
Results
MiR-770 is a prognostic biomarker in triple negative breast cancer
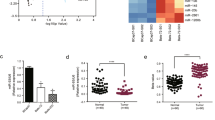
To identify miRNAs biomarker associated chemo-resistance of TNBC, we preformed miRNA expression array in two pairs of chemo-sensitive and chemo-resistant tissues. We identified 23 miRNAs with higher expression level and 27 with lower expression level in chemo-resistant tissues with the filter of 2 fold (Fig. 1a). Among miRNAs with different expression, we found miR-770 was significantly decreased in chemo-resistance tissues, which has not been well understood in TNBC, and we focused on this miRNA in our subsequent investigations.
MiR-770 is aberrantly expressed in chemo-sensitive and chemo-resistant breast tissues and is prognostic. a Heat map diagram depicting expression of 50 miRNAs dysregulated in chemo-sensitive compared with chemo-resistant breast tissues. b, c Kaplan-Meier and Cox-regression analysis of miR-770 levels and overall survival in all b or TNBC c patients. *P < 0.05, **P < 0.01
Next, to further determine the prognostic significance of miR-770, Kaplan-Meier survival plots and Cox regressions were computed using the datasets from GEO database (GSE20685)21. High expression of miR-770 contributed to a better overall survival in all type of breast cancers (n = 327, HR = 2.25, P < 0.01) and TNBC subtype (n = 49, HR = 4.87, P = 0.014) (Figs. 1b, c). These findings further demonstrated that miR-770 was a prognostic biomarker and up-regulation of miR-770 improved the prognosis of TNBC patients, which suggested that miR-770 play an important role in the regulation of chemo-resistance and progression of TNBC.
MiR-770 promoted the chemo-sensitivity of TNBC to doxorubicin via induction of apoptosis
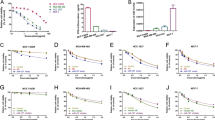
For the miRNAs expression array showed that miR-770 was decreased in chemo-resistant tissues, to further understand the mechanism of miR-770 in the regulation of chemo-resistance on DOX, we transfected MDA-MB-468 and MDA-MB-231 with miR-NC or miR-770. We firstly examined the effect of miR-770 on the IC50 of DOX in two TNBC cell lines. As shown in Fig. 2a, we found that overexpression of miR-770 could significantly decrease the IC50 of DOX in both cell lines, which suggested that miR-770 could promote the chemo-sensitivity of TNBC. Moreover, we also examined the cytotoxicity of specific concentrations of DOX on MDA-MB-468 (0.4 μM) and MDA-MB-231 (0.1 μM), and the cell viability was detected daily for 5 days. Our results proved that miR-770 could also promote the chemo-sensitivity in a time-dependent manner (Fig. 2b). What’s more, to eliminate the possibility of miR-770 may effect cell growth and cell cycle, we performed cell proliferation and cell cycle assay, and results showed that miR-770 overexpression alone do not affect cell proliferation and cell cycle (Supplementary Fig. S2).
Mir-770 sensitizes TNBC cells to DOX via inducing apoptosis. a Effect of miR-770 overexpression in MDA-MB-468 and MDA-MB-231 cells on the IC50 value. b The cytotoxicity assay of special concentrations of DOX on MDA-MB-468 (0.4 μM) and MDA-MB-231 cells (0.1 μM). c Representative flow cytometer images of TNBC cells transfected with miR-NC or miR-770 with or without DOX treatment. The y-axis quantifies the number of cells stained with propidium iodine and the x axis quantifies number of cells stained with Annexin V-FITC. d Quantification of apoptosis cells of Fig. 2.C. e Western blot analysis of apoptosis related markers in cells transfected with miR-NC and miR-770 treated with or without DOX. Data represent means ± S.D. of at least three independent experiments. *P < 0.05, **P < 0.01
Previous studies demonstrated that DOX could kill the TNBC cells via induction of apoptosis22,23. To further explore the mechanism of miR-770 on the regulation of DOX sensitivity, we detected the synergistic effect of miR-770 and DOX on the percentage of apoptosis. Results proved that the addition of miR-770 alone showed not significant effect on the apoptosis, while the combination use of miR-770 and DOX could significantly promote the DOX induced apoptosis (Figs. 2c, d). Moreover, we found that the combination use of miR-770 and DOX could also active the apoptosis pathways (Fig. 2e). These findings suggested that miR-770 could sensitize TNBC cells to DOX via activation of DOX induced apoptosis.
MiR-770 could be transferred by exosomes and regulate TAMs induced chemo-resistance
Based on above results, we proved that miR-770 could sensitize TNBC cells to DOX via its endogenous effect on apoptosis pathway. Macrophages are one of the most important components of the stroma which shift their functional phenotypes in response to various microenvironmental signals generated from tumor and stromal cells, and was defined as tumor associated macrophages (TAMs)24. Previous studies have reported that the aggregation of TAMs decreased the cytotoxicity of DOX to breast cancer cells25,26, to further explore the influence of miR-770 overexpression on DOX sensitivity, we examined whether miR-770 could be transferred by exosomes to modify the tumor microenvironment.
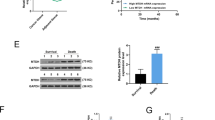
Now that we have proved that miR-770 could be transferred to TAMs, we explored the effect of miR-770 overexpression on TAMs phenotypes and TAMs induced chemo-resistance. After PMA treated for 24 h, THP-1 cells was transfected with miR-NC and miR-770, then IL-4 and IL-13 was added for 24 h for induction of M2 phenotype. For mRNA detection, M1 markers (MCP-1, iNOS, CD80) was significantly increased while M2 markers (CD206, ARG-1, MRC-2) was remarkable decreased in miR-770 group (Fig. 3e). Then the detection of protein levels proved the same result (Fig. 3f). These results suggested that miR-770 may participate in the process of macrophage polarization, which promoted M1 phenotype and inhibited M2 phenotype. We then determined the effect of miR-770 overexpression in THP-1 cells on TAMs induced chemo-resistance. As shown in Fig. 3g, the conditioned medium from THP-1 transfected with miR-770 (CM-miR-770) could antagonize the chemo-resistance induced by M2 phenotype THP-1 transfected with miR-NC (CM-miR-NC), the IC50 was significantly decreased in both TNBC cells treated with CM-miR-770. In conclusion, our results demonstrated that the overexpression of miR-770 could be transferred from TNBC cells to TAMs and regulate the polarization of macrophages, thus sensitizing TNBC cells to DOX.
Mir-770 could be packed in TNBC derived exosomes and transferred to influence to phenotype of TAMs. a The expression of miR-770 in exosomes that derived from TNBC cells transfected with miR-NC and miR-770. b The ideograph of co-culture system. c MDA-MB-468 and MDA-MB-231 was transiently transfected with FAM labeled miR770 or without transfection (Control) were co-cultured with PMA treated THP-1. Fluorescence microscopy was used to detect the green signals in THP-1 cells. d The relative miR-770 expression in THP-1 cells co-cultured with TNBC cells transfected with miR-NC or miR-770. e THP-1 cells were treated with 150 nM PMA for 24 h and were transfected with miR-NC/770, then 20 ng/ml IL-4 and 20 ng/ml were added for another 24 h. The mRNA of both cells were isolated and the relative mRNA expression of M1 and M2 markers was detected. f THP-1 cells were treated as above, the protein was isolated and iNOS and Arg-1 were detected. g THP-1 cells were treated as above, then the conditioned medium (CM) was collected. The IC50 of both MDA-MB-468 and MDA-MB-231 cells to DOX was detected when cultured with 468 CM-miR-NC/770. Data represent means ± S.D. of at least three independent experiments. *P < 0.05, **P < 0.01
MiR-770 inhibited the migration and invasion ability via modification of EMT pathway
Previous studies reported that the arisen of chemo-resistance may associated with the high metastatic ability of breast cancer cells27. To investigate the role of miR-770 on the metastatic ability of TNBC cells, we transfected MDA-MB-468 and MDA-MB-231 cells with miR-NC or miR-770, and transwell system was used to examine the change of migration and invasion ability. As shown in Figs. 4a–c, the cell counts of MDA-MB-468 and MDA-MB-231 were significantly decreased in both migration and invasion systems after transfection with miR-770 mimics, which suggested miR-770 could inhibit the metastatic ability of TNBC cells. Further exploration of the mechanism of miR-770, we found that miR-770 could modify the EMT pathway, which promoted epithelial phenotypes and inhibited mesenchymal phenotypes (Fig. 4d). In conclusion, miR-770 could inhibit the migration and invasion of TNBC cells via inhibition of EMT pathway.
MiR-770 inhibits aggressive behaviors of TNBC cells. a, b Effect of miR-770 overexpression on the metastatic a and invasive b ability of MDA-MB-468 and MDA-MB-231 cells. Cells were counted after staining with crystal violet. Representative images are shown. c Quantification of migration and invasion of cells transfected with miR-770 compared with control. d Western blot analysis of EMT related markers in cells transfected with miR-NC and miR-770. Data represent means ± S.D. of at least three independent experiments. *P < 0.05, **P < 0.01
Knockout of miR-770 promoted the chemoresistance and metastasis of TNBC cells
To further demonstrated the important role of miR-770 in the regulation of chemoresistance and metastasis, we knockout miR-770 with miR-770 inhibitor. In accordance with results above, we proved that knockdown of miR-770 could promote DOX resistance in both MDA-MB-468 and MDA-MB-231 cell lines, and increase the ability of migration and invasion. Results were shown as Supplementary Fig. S1 in supplementary materials.
STMN1 is a direct target of miR-770
To explore the underlying molecular mechanism of miR-770-mediated chemo-sensitivity and metastasis suppression, we performed in silico studies to search for potential gene targets of miR-770 using the bioinformatics algorithms: TargetScan, miRDB, and PICTAR5. All three algorithms predicted STMN1 as a target of miR-77028; the putative targets sequence are in base pairs 54-60, 267-273, 409-415 of the STMN1 3′ UTR (Fig. 5a). Homology searches revealed that this putative binding site is evolutionarily conserved (Fig. 5b).
STMN1 mRNA is a direct target of miR-770. a The three predicted targeting sequence of miR-770 of the STMN1 3′ UTR. b Sequence alignment of the predicted miR-770 seed region in the 3′ UTR of STMN1 from 5 organisms. c qRT-PCR and western blot assay revealed that overexpression of miR-770 level significantly reduced STMN1 expression in MDA-MB-468 and MDA-MB-231 cells. d The diagram illustrates the construction of the luciferase reporter plasmids. The substitutions of nucleotides within the three binding sites were shown and the three constructs named as mut1, mut2, and mut 3 separately (Wt: wild type; Mut: mutant type.). e Luciferase reporter assay in HEK293T cells revealed miR-770 suppressed STMN1 3′ UTR luciferase activity of all wide-type constructs. Firefly luciferase activity was normalized to the internal Renilla luciferase activity. Data represent means ± S.D. of at least three independent experiments. *P < 0.05, **P < 0.01
To further analysis the role of miR-770 on the expression of STMN1, the mRNA and protein level of STMN1 was examined after the transfection of miR-770. Our results showed that the overexpression of miR-770 could significantly decrease the STMN1 expression in both mRNA and protein level, our results suggested that miR-770 might regulate the STMN1 expression via targeting of 3’ UTR (Fig. 5c).
To determine whether miR-770 could bind directly to the 3′ UTR of target mRNA, we cloned both three wild type and mutant binding sites of miR-770 into the pmiRGLO vector, respectively, which was synthesized from Vigene Biosciences (Fig. 5d). For luciferase activity assays, miR-NC or miR-770 was co-transfected with pmiRGLO-3′ UTR vectors into HEK293T cells. As shown in Fig. 5e, the relative luciferase activity was remarkably decreased by miR-770 when the three wild-type 3′ UTR of STMN1 was present, which 3′ UTR1 and 3′ UTR2 exerted a stronger repression effect compared with 3′ UTR3. In addition, when transfected with mutant vectors which contained the STMN1 3′ UTR with mutations in the putative miR-770 binding sites, miR-770 mimic caused no obvious change in luciferase activity. Taken together, these findings indicated that STMN1 mRNA is a direct, down-stream target of miR-770 in breast cancer cells.
The tumor suppressor role of miR-770 is mediated by down-regulation of STMN1
STMN1 has been proved as the direct target of miR-770, to evaluate whether STMN1 was a functional target gene, we firstly interfered its expression via specific siRNA. As shown in Figs. 6a, b, knockdown of STMN1 significantly decreased the ability of both DOX resistance and metastasis. Based on this finding, we further determine whether miR-770-induced inhibition of chemo-resistance, migration and invasion could be reversed by restoration of STMN1 expression, and we performed a gain-of-function assay. Specifically, we transfected a vector expressing STMN1 without its 3′ UTR, which resulted in constitutive expression of STMN1 without the potential for miR-770-mediated degradation, and the overexpression of STMN1 was verified by qRT-PCR and western blot (Fig. 6c). Then the pcDNA3.1 or pcDNA3.1-STMN1 was co-transfected with miR-NC or miR-770. As shown in Fig. 6d, we also proved that STMN1 could antagonize miR-770 induced decrease of IC50 in both TNBC cell lines. Moreover, STMN1 could significantly reverse the miR-770 induced repression of chemo-resistance in a time dependent manner. (Fig. 6e).
Tumor-suppressive effect of miR-770 is mediated by targeting of STMN1 mRNA. Both MDA-MB-468 and MDA-MB-231 cells were transfected with miR-NC or miR-770 together with either STMN1 or pcDNA3.1. a mRNA and protein level were detected for verification. And IC50 assay b and cytotoxicity test c were performed. The migration d and invasion ability of TNBC cells. e Quantification of migration and invasion of cells. Data represent means ± S.D. of at least three independent experiments. *P < 0.05, **P < 0.01
Then we explored the role of STMN1 on the inhibition of metastatic ability caused by miR-770. Results showed that the overexpression of STMN1 could reverse the miR-770 induced inhibition of both migration and invasion ability in MDA-MB-468 and MDA-MB-231 cell lines (Figs. 6f, g). Moreover, we further proved overexpression of STMN1 could reverse the miR-770 induced inhibition of EMT pathway (Fig. 6h). These results reinforced that STMN1 mRNA is a direct functional target of miR-770.
MiR-770 suppressed DOX resistance and metastasis in a xenograft model
Our results have proved that overexpression of miR-770 inhibited DOX resistance and metastasis of TNBC cell lines in vitro. To further validated the role of miR-770, MDA-MB-231 was chosen to be transfected with pMSCV-NC and pMSCV-770 for its highly tumorigenic and metastatic abilities. The chemoresistance and metastasis of both pMSCV-NC and pMSCV-770 cell lines were validated. As shown in Figs. 7a, b, both pMSCV-NC and pMSCV-770 cell lines were seed subcutaneously, and mice were injected with saline or DOX (1 mg•kg-1) at day 5, 10, 15, 20 and 25. Our results demonstrated that tumors in pMSCV-770 group were significantly shrunken compared with pMSCV-NC group. Previous studies have proved that lung metastasis was an indicator of tumor aggressiveness. As shown in Fig. 7c, the metastasis of pMSCV-770 overexpression group was significantly decreased than the pMSCV-NC group, which was also demonstrated by HE staining assay (Fig. 7d). Our results have approved that over-expression of miR-770 could inhibit chemo-resistance as well as metastasis of TNBC in vivo Fig 8.
MiR-770 overexpression inhibited DOX resistance and metastasis in xenograft models. a, b MiR-770 synergized with DOX to suppress the growth of tumor. c Overexpression of miR-770 inhibited lung metastasis of MDA-MB-231 cells. d Representative images of hematoxylin and eosin (HE)-staining of lungs isolated from mice that received tail vein injection of MDA-MB-231 pMSCV-NC cells and pMSCV-770 cells
Discussion
The TNBC was firstly reported in 2006, which was defined as a subtype with a more aggressive behavior and poorer prognosis4. The research on TNBC never stopped for the aggressive clinical behavior and lack of molecular target. The chemo-resistance and metastasis were two of the most important causes leading to the relapse and eventually death of TNBC patients, and numerous studies have focused on this aspect29. MiRNAs have been reported to take part in the biological process of tumor development including chemo-resistance and metastasis, and a lot of miRNAs have been found to play an important role in this process. Niu et al. found that miRNA-181a could function as a tumor promoter via direct suppression of BAX, and the overexpression of miRNA-181a contributed to the chemo-resistance and metastasis of TNBC cells30. Hong et al. reported that miR-125 could inhibit the EMT process and chemo-resistance and TNBC cells via targeting of MAP2K731. These studies suggested that miRNAs might play an important role in regulating of chemo-resistance and metastasis of TNBC.
Mir-770 has been found to be correlated with kinds of cancers, including gastric cardia adenocarcinoma, lung cancer and ovarian cancer32,33,34. Zhao et al. reported that miR-770 was down-regulated in cisplatin resistant OVC patients and could predict the response to cisplatin. Overexpression of miR-770 promoted chemo-sensitivity of cancer cells via inhibition of DNA repair34. Moreover, Lee et al. proved that miR-770 sensitized cancer cells to radiation via targeting of PDZ-binding kinase33. However, the role of miR-770 in TNBC still remained unknown. In this study, we explored the significance of miR-770 on the chemo-resistance and metastasis of TNBC and whether miR-770 could function via exosomes to modify the tumor microenvironment.
Based on miRNA microarrays, we demonstrated that some miRNAs were different expressed in chemo-resistant TNBC and chemo-sensitive patients, and miR-770 was one of the most significantly changed miRNAs which high expression predicted more sensitive to chemotherapy. What’ more, the data from TCGA suggested that miR-770 could also serve as a prognostic marker which down-regulation of miR-770 predicted a poorer prognosis. To understand the mechanism of miR-770 in the regulation of chemo-resistance of DOX, we performed cytotoxicity test and we found that overexpression of miR-770 could decrease the IC50 of both MDA-MB-468 and MDA-MB-231 cell lines, and promote the sensitivity to DOX in a dose dependent manner. Further studies we validated that miR-770 could promote DOX induced activation of apoptosis, while miR-770 alone showed no significant effect on the apoptosis pathway, which proved that miR-770 could sensitize TNBC cells to DOX through promoting apoptosis. The metastasis and chemo-resistance have been reported to be closely related, the arisen of chemo-resistance could promote metastasis of cancer cells and the metastasis contributed to chemo-resistance35,36,37. After transfection of miR-770 mimics, we found that the metastasis and invasion ability were significantly decreased in both TNBC cell lines, and western blotting showed the process of epithelial-mesenchymal transition was inhibited. One critically important, yet often overlooked, component of tumor progression is the tumor microenvironment, which is primarily composed of fibroblasts, extracellular matrix proteins, endothelial cells, macrophages and so on38. The tumor microenvironment has been reported to be correlated with the proliferation, metastasis, and chemo-resistance of cancer cells. One of the most important components tumor microenvironment was macrophages which could shift their phenotypes to influence the behavior of tumor cells39. And we evaluated the effect of miR-770 on TAMs. We found that miR-770 could be packed into exosomes, and transfection of miR-770 in MDA-MB-468 and MDA-B-231 cells increased its expression in tumor derived exosomes. Co-culture assay proved that miR-770 in tumor derived exosomes could be transferred into TAMs, and increased the expression of miR-770 in macrophages. Further experiments found that high expression of miR-770 influenced the polarization of macrophages which promote M1 phenotype and inhibit M2 phenotype. Finally we evaluated whether up-regulation of miR-770 in TAMs influencing TAMs induced chemo-resistance, and we found the IC50 was significantly decreased in both TNBC cell lines cultured with miR-770-CM and compared with miR-NC-CM. In conclusion, we proved that miR-770 suppresses the chemo-resistance and metastasis of triple negative breast cancer via regulation of apoptosis and EMT and modification of tumor microenvironment.
Stathmin1 (STMN1), also known as oncoprotein 18, is an important cytoplasmic phosphoprotein that regulates cellular microtubule dynamics, which has been reported to take part in many types of cancers40. STMN1 promotes microtubule depolymerization by sequestering tubulin and stimulating catastrophes41,42. The overexpression of STMN1 has been reported to be negatively associated with the prognosis of many cancers including nasopharyngeal carcinoma43, hepatocellular carcinoma44, colorectal cancer45 and so on. What’ more, studies proved that up-regulation of STMN1 could promote the proliferation, metastasis, and chemo-resistance of cancers, which the expression of STMN1 alone could be a prognostic indicator46,47,48. In our study, we proved that miR-770, a potential tumor suppressor, could specifically target and down-regulate STMN1, thus inhibit the metastasis and chemo-resistance of TNBC cells. Moreover, studies about STMN1 also focused on its role in macrophage polarization, Xu et al. reported that down-regulation of STMN1 is required for the phenotypic changes and classical activation of macrophages. This study also confirmed our discovery that miR-770 overexpression is associated with M1 polarization of macrophages. In a word, we proved that the tumor suppression effect of miR-770 was mediated by targeting of STMN1, which has been demonstrated as a tumor promoter in kinds of cancers.
In conclusion, we demonstrated that miR-770 could significantly inhibit the DOX resistance and metastasis of triple negative breast cancer cells, which is mediated via regulation of apoptosis and EMT and modification of tumor microenvironment (Fig. 7). Conversely, overexpression of STMN1, which is a direct functional target of miR-770, promoted breast cancer cell DOX resistance and metastasis. The newly identified miR-770/STMN1 axis provides novel insight into the chemo-resistance and metastasis of triple negative breast cancer, and represents a potential therapeutic target for the treatment of triple negative breast cancer.
Materials and methods
Ethics statement and human tissue samples
All experimental procedures were approved by the Ethical Committee of Shandong University. Breast tissues were obtained at the time of surgery from patients admitted to Qilu hospital, and immediately stored at −80 °C until use. All patients provided written informed consent for the use of these clinical materials in research.
Cell lines and cell culture
All cell lines were obtained from the American Type Culture Collection (Manassas, VA), and were maintained using standard media and condition. Briefly, MDA-MB-231 and MDA-MB-468 were culture with Dulbecco’s modified Eagle’s medium (Invitrogen, Carlsbad, CA, USA) containing 10% fetal bovine serum (Hyclone). THP-1 were were maintained in Roswell Park Memorial Institute (RPMI) 1640 medium supplemented with 10% fetal bovine serum and 5 mg/ml insulin. All cell lines were grown at 37 °C in a 5% CO2 cell culture incubator.
Cell transfection with miRNA and plasmids
MiR-770 mimic, miR mimic control, miR-770 inhibitor and inhibitor control were purchased from GenePharma (Shanghai, China). For miR-770 overexpression, plasmids was constructed from Vigene Biosciences (Rockville, USA). For STMN1 interference, siRNA and its control were synthesized from GenePharma (Shanghai, China). For STMN1 overexpression, vector was constructed as described previously49. Briefly, STMN1 cDNA was cloned into the multiple cloning sites of vector pcDNA3.1 (Invitrogen, Carlsbad, CA, USA), and the resultant expression vector and empty vector were transfected into MDA-MB-468 and MDA-MB-231 cells to establish STMN1 overexpressing and control cell lines, respectively. Sequence of siRNAs and primers for plasmid construction are shown in the Supplementary Table S1. Cells were seeded in 60 mm dishes (6 × 105 cells/dish) and transfected using Lipofectamine 2000 (Invitrogen) for further assays.
Cell viability assay
Cell viability was determined by a 3-(4, 5-dimethylthiazol-2-yl)-2, 5-diphenyltetrazolium bromide (MTT) assay. MDA-MB-468 and MDA-MB-231 cells were cultured in 96-well plates, after incubation overnight, the medium was replaced with solutions containing different concentrations of DOX (0, 0.01, 0.05, 0.1, 0.2, 0.5, 1 μM). After incubation for the indicated time, 20 μl of MTT (5 mg/ml in PBS) was added into each well and incubated for 4 h. The supernatants were carefully aspirated, and 100 μl of dimethyl sulfoxide was added to each well. Absorbance values at 490 nm were measured on a Microplate Reader (Bio-Rad).
Migration and invasion assays
Migration and invasion assays were performed as described previously using the Transwell system (Corning Costar, Lowell, MA, USA)50. In the migration assay, 700 μL of medium with 20% Fetal bovine serum (FBS) was added to the lower well of each chamber and 1 × 105 cells suspended in serum-free medium were added to the upper inserts. After incubation for the indicated time, the total number of cells adhering to the lower surface of the membrane was quantified in six representative fields. The invasion assay was performed in the same way as the migration assay except that the membrane was coated with matrigel (BD Biosciences, Bedford, MA, USA).
Cell apoptosis assay
Cell apoptosis was detected using an Annexin V-FITC/PI apoptosis detection kit (JingMei Biotech, Beijing, China) according to the manufacturer’s protocols. Briefly, cells were treated with EDTA-free trypsin), and resuspended with 400 μL Annexin binding buffer at a concentration of 106 cells/ml. Then, 5 ul FITC-conjugated AnnexinV and 10 ul PI were added to the cells and incubated at room temperature for 15 min in dark. Cell apoptosis assay was performed within 1 h post-staining on a flow cytometry.
Isolation of exosomes
After cell cultures reached 90% confluency, cells were washed with PBS and incubated with freshly prepared complete medium containing exosome-free FBS for 48 h. then exosomes were isolated with a polyethylene glycol-based method as previous described with little modification51. Vesicle-containing medium from cell culture was centrifuged at 500×g for 5 min and followed by 2000×g for 30 min to remove cellular debris and large apoptotic bodies. Then the media was added to an equal volume of a 2 × PEG solution and samples were mixed thoroughly by inversion, and incubated at 4 °C overnight (at least 12 h). The next day, samples were centrifuged in a tabletop centrifuge at 10,000×g for 1 hour at 4 °C to acquire the pellet of exosomes.
In vitro detection of miR-770 transfer
For the co-culture experiments, MDA-MB-468 and MDA-MB-231 cells were transfected with FAM-labeled miR-770 (GenePharma, Shanghai, China) or without transfection (Control) and were grown on the 0.4 μm pore size transwell (Thermo Fisher Scientific, Carlsbad, CA, USA), then THP-1 cells treated with 150 nM PMA and grown on the coverslips in the bottom well of the transwell. After 24 h of transfer, THP-1 cells were fixed with 4% paraformaldehyde and the nuclei were stained with 4,6-diamidino-2-phenylindole blue. Fluorescent microscopy was used to detect the green signals in the THP-1 cells.
Quantitative RT–PCR analysis
Total RNA was extracted from cells using the TRIzol reagent (Invitrogen, Carlsbad, CA, USA). PrimeScript reverse transcriptase (RT) reagent kit (TaKaRa, Shiga, Japan) was used to synthesize cDNA from total RNA. MiRNA from 1 μg of total RNA was reverse transcribed using the Prime-Script miRNA cDNA Synthesis Kit (TaKaRa, Shiga, Japan). Real-time PCR was performed on an Applied Biosystems StepOne plus Real-Time PCR System. Primer information used in the study is provided in the Supplementary Table S1.
Western blot analysis
Western blot analysis was performed as described previously [17]. In brief, cells were harvested and lysed in lysis buffer (1 × PBS, 1% NP40, 0.1% sodium dodecyl sulfate, 5 mM EDTA, 0.5% sodium deoxycholate and 1 mM sodium orthovanadate) containing protease inhibitors. Subsequently, 50 μg of total cellular protein from each sample were separated by 10% SDS-PAGE and electro-transferred onto polyvinylidene fluoride (PVDF) membrane using a semi-dry blotting apparatus (Bio-Rad, Hercules, CA, USA). The membranes were blocked with 5% nonfat milk at room temperature for 1 h, and then incubated with primary antibodies overnight at 4 °C. After incubation with the appropriate secondary antibodies, the protein bands were detected using the Pro-lighting HRP agent. Expression of β-actin was used as a loading control. The antibody used in this study was obtained from Cell Signaling Technology (Beverly, MA): E-Cadherin (24E10), N-Cadherin (D4R1H), ZEB1 (D80D3) Vimentin (D21H3), PARP (46D11), Caspase-3 (9662), Bax (D2E11), Bcl-2 (D17C4), STMN1 (3352); Santa Cruz Biotechnology (CA): ARG-1 (sc-4496), INOS (sc-649), Fibronectin (sc-29011).
Dua-luciferase reporter assay
The dual-luciferase miRNA Target Expression vector pmirGLO (Promega, Madison, WI, USA) was used to generate luciferase reporter constructs. Three 3′ UTR of STMN1, containing different putative miR-770-binding sites were amplified and cloned into pmirGLO. Mutant reporter plasmids were constructed as described previously52. Cells were seeded in 96-well plates and co-transfected with wild-type or mutated STMN1 3′ UTR constructs, and miR-770 mimic or negative control. Luciferase activity was measured with the dual-luciferase reporter assay system (Promega). Firefly luciferase activity was normalized against Renilla luciferase activity.
In vivo chemo-resistance and metastasis assay
The in vivo chemo-resistance and metastasis assay was performed as described previously50. Briefly, MDA-MB-231 vector and MDA-MB-231 miR-770 cells (1 × 107cells) in 200 μl of PBS: Matrigel (1:3, v/v)) were injected subcutaneously into left flank of 4-6 weeks-old BALB/c nu/nu female mice (five mice per group), and mice were injected with saline or DOX (1 mg•kg-1) at day 5, 10, 15, 20, and 25. Tumor growth rate was monitored by measuring tumor diameters every 5 days. Both maximum (L) and minimum (W) length of the tumor were measured using a slide caliper, and the tumor volume was calculated as ½LW2. When mice were killed, tumors were collected. To produce experimental lung metastasis, 5 × 105 cells were injected into the lateral tail veins of 4–6 weeks-old BALB/c nu/nu female mice (seven mice per group). After 2 weeks, all the mice were killed under anesthesia. The lungs were collected and fixed in 10% formalin. For tissue morphology evaluation, hematoxylin and eosin (HE) staining was performed on sections from embedded samples as previously described53. All animal experiments were performed with the approval of Shandong University Animal Care and Use Committee.
Statistical analysis
All data are presented as means ± S.D. from at least three independent experiments. The software SPSS V18.0 was used for statistical analysis. Statistical significance of differences between two groups was evaluated using Student’s t-test, and one-way ANOVA was used to determine the significance of differences among multiple groups. Differences with P < 0.05 were considered statistically significant.
References
Wahba, H. A. & El-Hadaad, H. A. Current approaches in treatment of triple-negative breast cancer. Cancer Biol Med 12, 106–116 (2015).
Anders, C. & Carey, L. A. Understanding and treating triple-negative breast cancer. Oncology. 22, 1233–1239 (2008). discussion1239-1240, 1243.
Bianchini, G., Balko, J. M., Mayer, I. A., Sanders, M. E. & Gianni, L. Triple-negative breast cancer: challenges and opportunities of a heterogeneous disease. Nat. Rev. Clin. Oncol. 13, 674–690 (2016).
Haffty, B. G. et al. Locoregional relapse and distant metastasis in conservatively managed triple negative early-stage breast cancer. J Clin Oncol 24, 5652–5657 (2006).
Nofech-Mozes, S. et al. Patterns of recurrence in the basal and non-basal subtypes of triple-negative breast cancers. Breast. Cancer. Res. Treat. 118, 131–137 (2009).
Nabholtz, J. M. et al. Multicentric neoadjuvant phase II study of panitumumab combined with an anthracycline/taxane-based chemotherapy in operable triple-negative breast cancer: identification of biologically defined signatures predicting treatment impact. Annal Oncol 25, 1570–1577 (2014).
Davuluri, G., Schiemann, W. P., Plow, E. F. & Sossey-Alaoui, K. Loss of WAVE3 sensitizes triple-negative breast cancers to chemotherapeutics by inhibiting the STAT-HIF-1alpha-mediated angiogenesis. Jak-Stat 3, e1009276 (2014).
Yao, H. et al. Triple-negative breast cancer: is there a treatment on the horizon? Oncotarget 8, 1913–1924 (2017).
Isakoff, S. J. Triple-negative breast cancer: role of specific chemotherapy agents. Cancer J 16, 53–61 (2010).
Liedtke, C. et al. Response to neoadjuvant therapy and long-term survival in patients with triple-negative breast cancer. J Clin Oncol 26, 1275–1281 (2008).
Carey, L. A. et al. The triple negative paradox: primary tumor chemosensitivity of breast cancer subtypes. Clin Cancer Res 13, 2329–2334 (2007).
Ahmad, A. Pathways to breast cancer recurrence. ISRN Oncol 2013, 290568 (2013).
Iorio, M. V. & Croce, C. M. MicroRNA dysregulation in cancer: diagnostics, monitoring and therapeutics. A comprehensive review. EMBO Mol Med 4, 143–159 (2012).
Bockhorn, J. et al. MicroRNA-30c inhibits human breast tumour chemotherapy resistance by regulating TWF1 and IL-11. Nat Commun 4, 1393 (2013).
Zhou, W. et al. Cancer-secreted miR-105 destroys vascular endothelial barriers to promote metastasis. Cancer. Cell. 25, 501–515 (2014).
Camacho, L., Guerrero, P. & Marchetti, D. MicroRNA and protein profiling of brain metastasis competent cell-derived exosomes. PLoS. ONE. 8, e73790 (2013).
Taylor, D. D. & Gercel-Taylor, C. MicroRNA signatures of tumor-derived exosomes as diagnostic biomarkers of ovarian cancer. Gynecol. Oncol. 110, 13–21 (2008).
Yang, C. & Robbins, P. D. The roles of tumor-derived exosomes in cancer pathogenesis. Clin. Dev. Immunol. 2011, 842849 (2011).
Di Modica, M. et al. Breast cancer-secreted miR-939 downregulates VE-cadherin and destroys the barrier function of endothelial monolayers. Cancer. Lett. 384, 94–100 (2017).
Santos, J. C., Ribeiro, M. L., Sarian, L. O., Ortega, M. M. & Derchain, S. F. Exosomes-mediate microRNAs transfer in breast cancer chemoresistance regulation. Am J Cancer Res 6, 2129–2139 (2016).
Kao, K. J., Chang, K. M., Hsu, H. C. & Huang, A. T. Correlation of microarray-based breast cancer molecular subtypes and clinical outcomes: implications for treatment optimization. BMC. Cancer. 11, 143 (2011).
Raiter, A., Yerushalmi, R. & Hardy, B. Pharmacological induction of cell surface GRP78 contributes to apoptosis in triple negative breast cancer cells. Oncotarget 5, 11452–11463 (2014).
Wang, H. et al. Next-generation proteasome inhibitor MLN9708 sensitizes breast cancer cells to doxorubicin-induced apoptosis. Sci Rep 6, 26456 (2016).
Qian, B. Z. & Pollard, J. W. Macrophage diversity enhances tumor progression and metastasis. Cell 141, 39–51 (2010).
Hughes, R. et al. Perivascular M2 Macrophages Stimulate Tumor Relapse after Chemotherapy. Cancer. Res. 75, 3479–3491 (2015).
Nakasone, E. S. et al. Imaging tumor-stroma interactions during chemotherapy reveals contributions of the microenvironment to resistance. Cancer. Cell. 21, 488–503 (2012).
Longley, D. B. & Johnston, P. G. Molecular mechanisms of drug resistance. J. Pathol. 205, 275–292 (2005).
Li, J. H., Liu, S., Zhou, H., Qu, L. H. & Yang, J. H. starBasev2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic. Acids. Res. 42, D92–D97 (2014).
Parvani, J. G. et al. Deptor enhances triple-negative breast cancer metastasis and chemoresistance through coupling to survivin expression. Neoplasia 17, 317–328 (2015).
Niu, J. et al. Induction of miRNA-181a by genotoxic treatments promotes chemotherapeutic resistance and metastasis in breast cancer. Oncogene. 35, 1302–1313 (2016).
Hong, L. et al. miR-125b inhibited epithelial-mesenchymal transition of triple-negative breast cancer by targeting MAP2K7. OncoTarget Ther 9, 2639–2648 (2016).
Guo, W. et al. Promoter hypermethylation-mediated downregulation of miR-770 and its host gene MEG3, a long non-coding RNA, in the development of gastric cardia adenocarcinoma. Mol. Carcinog. 56, 1924–1934 (2017).
Lee, H. C. et al. Radiation-inducible miR-770-5p sensitizes tumors to radiation through direct targeting of PDZ-binding kinase. Cell Death Dis 8, e2693 (2017).
Zhao, H. et al. MiR-770-5p inhibits cisplatin chemoresistance in human ovarian cancer by targeting ERCC2. Oncotarget 7, 53254–53268 (2016).
Zheng, X. et al. Epithelial-to-mesenchymal transition is dispensable for metastasis but induces chemoresistance in pancreatic cancer. Nature. 527, 525–530 (2015).
Fischer, K. R. et al. Epithelial-to-mesenchymal transition is not required for lung metastasis but contributes to chemoresistance. Nature. 527, 472–476 (2015).
Geldof, A. A. & Rao, B. R. Doxorubicin treatment increases metastasis of prostate tumor (R3327-MatLyLu). Anticancer. Res. 8, 1335–1339 (1988).
Au Yeung, C. L. et al. Exosomal transfer of stroma-derived miR21 confers paclitaxel resistance in ovarian cancer cells through targeting APAF1. Nat Commun 7, 11150 (2016).
Niu, M., Valdes, S., Naguib, Y. W., Hursting, S. D. & Cui, Z. Tumor-Associated Macrophage-Mediated Targeted Therapy of Triple-Negative Breast Cancer. Mol. Pharm. 13, 1833–1842 (2016).
Bai, T. et al. High STMN1 level is associated with chemo-resistance and poor prognosis in gastric cancer patients. Br. J. Cancer. 116, 1177–1185 (2017).
Marklund, U., Larsson, N., Gradin, H. M., Brattsand, G. & Gullberg, M. Oncoprotein 18 is a phosphorylation-responsive regulator of microtubule dynamics. EMBO. J. 15, 5290–5298 (1996).
Rubin, C. I. & Atweh, G. F. The role of stathmin in the regulation of the cell cycle. J. Cell. Biochem. 93, 242–250 (2004).
Cheng, A. L. et al. Identification of novel nasopharyngeal carcinoma biomarkers by laser capture microdissection and proteomic analysis. Clin Cancer Res 14, 435–445 (2008).
Hsieh, S. Y. et al. Stathmin1 overexpression associated with polyploidy, tumor-cell invasion, early recurrence, and poor prognosis in human hepatoma. Mol. Carcinog. 49, 476–487 (2010).
Zhang, H. Q. et al. STMN1 in colon cancer: expression and prognosis in Chinese patients. Eur. Rev. Med. Pharmacol. Sci. 20, 2038–2044 (2016).
Wang, J. et al. Downregulation of stathmin 1 in human gallbladder carcinoma inhibits tumor growth in vitro and in vivo. Sci Rep 6, 28833 (2016).
Kuang, X. Y. et al. The phosphorylation-specific association of STMN1 with GRP78 promotes breast cancer metastasis. Cancer. Lett. 377, 87–96 (2016).
Kuang, X. Y. et al. Stathmin and phospho-stathmin protein signature is associated with survival outcomes of breast cancer patients. Oncotarget 6, 22227–22238 (2015).
Zhang, N. et al. MicroRNA-30a suppresses breast tumor growth and metastasis by targeting metadherin. Oncogene. 33, 3119–3128 (2014).
Li, Y. et al. Huaier extract suppresses breast cancer via regulating tumor-associated macrophages. Sci Rep 6, 20049 (2016).
Rider, M. A., Hurwitz, S. N. & Meckes, D. G. Jr. ExtraPEG: A Polyethylene Glycol-Based Method for Enrichment of Extracellular Vesicles. Sci Rep 6, 23978 (2016).
Heckman, K. L. & Pease, L. R. Gene splicing and mutagenesis by PCR-driven overlap extension. Nat. Protoc. 2, 924–932 (2007).
Liu, Y. et al. Transforming growth factor-beta1 upregulation triggers pulmonary artery smooth muscle cell proliferation and apoptosis imbalance in rats with hypoxic pulmonary hypertension via the PTEN/AKTpathways. Int. J. Biochem. Cell. Biol. 77(Pt A), 141–154 (2016).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No.81272903; No.81672613), Key Research and Development Program of Shandong Province (No.2016GGE2775), Shandong Science and Technology Development Plan (2016CYJS01A02) and Special Support Plan for National High Level Talents (“Ten Thousand Talents Program”) to Qifeng Yang. And the China Postdoctoral Science Foundation (no. 2015M572050) and the Key Research and Development Program of Shandong Province (no.2015GSF118035).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no conflict of interest.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, Y., Liang, Y., Sang, Y. et al. MiR-770 suppresses the chemo-resistance and metastasis of triple negative breast cancer via direct targeting of STMN1. Cell Death Dis 9, 14 (2018). https://doi.org/10.1038/s41419-017-0030-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-017-0030-7
This article is cited by
-
Exosomal miRNAs: the tumor's trojan horse in selective metastasis
Molecular Cancer (2024)
-
Drug resistance in breast cancer is based on the mechanism of exocrine non-coding RNA
Discover Oncology (2024)
-
Integrative analysis of homologous recombination repair patterns unveils prognostic signatures and immunotherapeutic insights in breast cancer
Journal of Applied Genetics (2024)
-
The role of non-coding RNAs in extracellular vesicles in breast cancer and their diagnostic implications
Oncogene (2023)
-
Relevance and mechanism of STAT3/miR-221-3p/Fascin-1 axis in EGFR TKI resistance of triple-negative breast cancer
Molecular and Cellular Biochemistry (2023)