Abstract
A potentially important aspect in the regulation of tumour metastasis is endocytosis. This process consists of internalisation of cell-surface receptors via pinocytosis, phagocytosis or receptor-mediated endocytosis, the latter of which includes clathrin-, caveolae- and non-clathrin or caveolae-mediated mechanisms. Endocytosis then progresses through several intracellular compartments for sorting and routing of cargo, ending in lysosomal degradation, recycling back to the cell surface or secretion. Multiple endocytic proteins are dysregulated in cancer and regulate tumour metastasis, particularly migration and invasion. Importantly, four metastasis suppressor genes function in part by regulating endocytosis, namely, the NME, KAI, MTSS1 and KISS1 pathways. Data on metastasis suppressors identify a new point of dysregulation operative in tumour metastasis, alterations in signalling through endocytosis. This review will focus on the multicomponent process of endocytosis affecting different steps of metastasis and how metastatic-suppressor genes use endocytosis to suppress metastasis.
Similar content being viewed by others
Background
Cancer is the second leading cause of global mortality.1 The spread of cancer cells from the primary tumour to distant organs and their subsequent progressive colonisation is referred to as metastasis. It is estimated that 90% of cancer-related deaths are due to metastatic disease rather than to the primary tumour growth. Typically, treatments for metastatic cancer are systemic therapy involving chemotherapy or molecular drugs, hormonal agents, immune checkpoint drugs, radiation therapy or surgery. Despite progress in extending cancer-survivorship rates,2 limited progress has been made in the treatment of metastatic cancer due to its complex nature and an inadequate understanding of the molecular and biochemical mechanisms involved.
Metastasis is a multistep process involving tumour cell invasion to neighbouring areas, intravasation into the bloodstream, arrest in the capillary bed of a secondary organ, extravasation from the circulatory system and colonisation at the secondary site.3 All of the above steps occur via complex interactions between cancer cells and their microenvironments. Despite the documented complexity and redundancy of the metastatic process, mutation or changes in the expression of single genes have been reported to alter metastatic ability. Genes that are involved in the promotion of metastasis at distant sites are referred to as metastasis promoting genes. Expression of these genes facilitates cancer cell establishment of appropriate interactions with changing microenvironments to promote continued survival and proliferation at secondary sites. Similarly, genes that inhibit the process of metastasis without affecting the growth of the primary tumour are referred to as metastasis suppressor genes and are described in detail in the later part of this review.
This review will highlight an often-overlooked aspect of metastasis, receptor endocytic pathways. Contributing to each step in metastasis is the distribution of multiple cell-surface receptors on tumour and microenvironmental cells. Receptor signalling is, in turn, modulated by endocytosis (internalisation, recycling or degradation). In recent years, there has been significant progress made towards understanding the mechanisms of the endocytosis pathway and its alterations that occur during metastasis. A growing body of literature suggests that receptor endocytosis affects metastasis and could be a tool for the functioning of metastasis suppressor or metastasis promoters. This review will focus on the role of endocytosis in metastasis and how these pathways are used by metastasis suppressors.
Endocytic pathways and metastasis
The term ‘endocytosis’ is derived from the Greek word ‘endon’, meaning within, ‘kytos’, meaning cell and ‘-osis’, meaning process. So, endocytosis is the process by which cells actively internalise molecules and surface proteins via an endocytic vesicle. Depending on the cargo type, internalisation route and scission mechanism, there are three general modes of vesicular endocytic trafficking that coexist in the cell and operate concurrently: phagocytosis, pinocytosis and receptor-mediated endocytosis. In phagocytosis, the cell’s plasma membrane surrounds a macromolecule (large solid particles > 0.5 μm) or even an entire cell from the extracellular environment and generates intracellular vesicles called phagosomes.4 Cellular pinocytosis/cellular drinking is a process in which fluids and nutrients are ingested by the cell, by pinching in and forming vesicles that are smaller than the phagosomes (0.5–5 μm).5 Both phagocytosis and pinocytosis are non-selective modes of taking up molecules. However, there are times when specific molecules are required by cells and are taken up more efficiently by the process of receptor-mediated endocytosis (RME). The endocytosis of specific cargoes via specific receptors can take place by clathrin-mediated (CME), caveolae-mediated (CavME), clathrin- and caveolae-independent endocytic (CLIC/GEEC) pathways. These endocytic pathways are briefly described below. Table 1 links selected endocytic proteins to in vitro components of the metastatic process and in vivo metastasis in cancer.
Clathrin-mediated endocytosis (CME)
The most studied endocytic mechanism is CME. It was first found to play an important role in low-density lipoprotein6 and transferrin uptake.7 It is known to be involved in internalisation and recycling of multiple receptors engaged in signal transduction (G-protein and tyrosine-kinase receptors), nutrient uptake and synaptic vesicle reformation.8 Clathrin-coated pits (CCP) are assemblies of cytosolic coat proteins, which are initiated by AP2 (assembly polypeptide 2) complexes that are recruited to a plasma membrane region enriched in phosphatidylinositol-(4,5)-bisphosphate lipid.9 AP2 acts as a principal cargo-recognition molecule and recognises internalised receptors through a short sequence motif in their cytoplasmic domains.10 As the nascent invagination grows, AP2 and other cargo-specific adaptor proteins recruit and concentrate the cargo, which is now facing the inside of the vesicle. Following cargo recognition/concentration, AP2 complexes along with other adaptor proteins to recruit clathrin. Clathrin recruitment stabilises the curvature of the growing CCP with the help of BAR (Bin-Amphiphysin-Rvs)-domain-containing proteins until the entire region invaginates to form a closed vesicle.11
Release of mature clathrin-coated vesicles from the plasma membrane is performed by the large multi-domain GTPase, Dynamin. Proteins such as amphiphysin, endophilin and sorting nexin 9 (BAR-domain-containing proteins) recruit Dynamin around the necks of budding vesicles.12 Similarly, other Dynamin partners (i.e., Grb2) also bind to Dynamin and increase its oligomerisation, which results in a higher GTPase activity.13 Oligomerised Dynamin assembles into collar-like structures encircling the necks of deeply invaginated pits and undergoes GTP hydrolysis to drive membrane fission.14 After a vesicle is detached from the plasma membrane, the clathrin coat is disassembled by the combined action of the ATPase HSC70 and the coat component auxilin.15,16 The released uncoated vesicle is ready to travel and fuse with its target endosome.
Signalling through CME is critical in cancer and metastasis. Clathrin light-chain isoform (CLCb) is specifically upregulated in non-small-cell lung cancer (NSCLC) cells and is associated with poor prognosis. NSCLC cells expressing CLCb exhibit increased rates of CME through Dynamin 1. This leads to activation of a positive feedback loop involving enhanced epidermal growth factor receptor (EGFR)-dependent Akt/GSK-3β (glycogen synthase kinase 3β) phosphorylation, resulting in increased cell migration and metastasis.17 Dynamin 2 is crucial for the endocytosis of several proteins known to be involved in cancer motility and invasiveness (e.g., β-1 integrin and focal adhesion kinase). Dynamin 2 overexpression correlates with poor prognosis.18
The regulation of certain receptors that are known to affect cancer and metastasis (i.e., EGFR and transforming growth factor β receptor (TGFβR)) by clathrin- and non-clathrin-mediated internalisation pathways preferentially targets the receptors to different fates (i.e., recycling or degradation).19,20 Different fates of receptors determine the net signalling output in a cell and affect cancer progression. Interestingly, CME is known to skew EGFR fate towards recycling rather than degradation, leading to prolonged duration of signalling.20 Similarly, the internalised EGF–EGFR complex may maintain its ability to generate cell signalling from endosomes affecting multiple downstream pathways.21 This active endosomal EGFR is known to regulate oncogenic Ras activity by co-internalising its regulators including Grb2, SHC, GAP and Cbl.21,22
Caveolae-mediated endocytosis (CavME)
CavME is the second most studied pathway of endocytosis and has been shown to be important in transcytotic trafficking across cells and mechanosensing.23 The CavME process involves formation of a bulb-shaped, 50–60-nm plasma membrane invaginations called caveolae (little caves), which is driven by both integral membrane proteins called caveolins and peripheral membrane proteins called cavins (cytosolic coat proteins). Caveolins (encoded by CAV-1, 2 and 3 paralogues) are small integral membrane proteins that are inserted into the inner side of the membrane bilayer through its cytosolic N-terminal region that binds to cholesterol. About 50 cavin molecules associate with each caveolae and exist in a homo- or hetero-oligomeric form (using four cavin family members).24 CavME is triggered by ligand binding to cargo receptors concentrated in caveolae. Budding of caveolae from the plasma membrane is regulated by kinases and phosphatases, such as Src tyrosine kinases and serine/threonine protein phosphatases PP1 and PP2A.25 As with CME, Dynamin is required to pinch off caveolae vesicles from the plasma membrane.26
Components of CavME have a vital role in cell migration, invasion and metastasis. It is speculated that CAV-1 has a dual role in cancer progression and metastasis. In the early stages of the disease, it functions predominantly as a tumour suppressor, whereas at later stages, its expression is associated with tumour progression and metastasis.27,28,29 As with a tumour suppressor, CAV-1 is often deleted in human cancers and mechanistically known to act through the caveolin scaffolding domain (CSD) by inhibiting cytokine receptor signalling.28,30 The CAV-1 effect on the late-stage tumour progression and metastasis has been attributed to tyrosine (Tyr14) phosphorylation of its protein product by Src kinases, leading to increased Rho/ROCK signalling and subsequent focal adhesion turnover.31 Knockdown of CAV-1 in breast and prostate cancer cells reduced the velocity, directionality and persistency of cellular migration.31,32 Similarly, expression of CAV-1 has been used as a marker of prognosis and overall survival in various types of human cancer. In pancreatic adenocarcinoma, positive expression of CAV-1 was found to correlate with tumour diameter, histopathological grade and poor prognosis. In lung cancer, CAV-1 expression statistically correlates with poor differentiation, pathological stage, lymph-node metastasis and poor prognosis. However, in hepatocellular carcinoma tissues, low expression of CAV-1 is associated with poor prognosis.33
Clathrin-independent endocytosis (CIE)
As per the name, the endocytic vesicles involved in CIE have no distinct coat and were first discovered by their resistance to inhibitors that block CME and CavME.34 CIE encompasses several pathways. (i) An endophilin-, Dynamin- and RhoA-dependent pathway for endocytosis of interleukin-2 receptor.35 (ii) A clathrin- and Dynamin-independent (CLIC/GEEC) pathway in which the GTPases RAC1 and CDC42 lead to actin-dependent formation of clathrin-independent carriers (CLICs). This, in turn, forms the glycosylphosphatidylinositol (GPI)-AP-enriched endosomal compartments (GEECs).36,37 (iii) An ARF6-dependent pathway involving the small GTPase ARF6, to activate phosphatidylinositol-4-phosphate 5-kinase that produces phosphatidylinositol-(4,5)-bisphosphate, leading to stimulation of actin assembly and endocytosis.38 The CIE pathway has been shown to suppress cancer cell blebbing and invasion through GTPase-activating protein GRAF1 (GTPase regulator associated with focal adhesion kinase-1) (Table 1).39 Various receptors are endocytosed by the CIE pathway, including interleukin-2 receptor (IL-2R), T-cell receptor (TCR) and GPI-linked proteins.40
Downstream endosomal trafficking
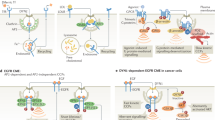
Internalised receptor–ligand cargoes can merge into a common endosomal network by undergoing multiple rounds of fusions. The first set of fusion leads to the formation of early endosomes where initial sorting routes are engaged, and the fate of the internalised receptors is decided (Fig. 1). Early endosomes are identified by the association of several proteins on their cytosolic surface, including RAB5, along with its effector VPS34/p150, a phosphatidylinositol 3-kinase complex. VPS34/p150 generates phosphatidylinositol 3-phosphate, which regulates the spatiotemporal and compartmentalisation aspects of endosomal functions.41,42 Structurally, early endosomes have tubular (membrane) and vacuolar (vacuoles) domains. Most of the membrane surface area lies in the tubules, while much of the volume is in the vacuoles. The membrane domains are enriched in proteins, including RAB5, RAB4, RAB11, ARF1/COPI, retromer and caveolin.43,44 These proteins are involved in multiple functions, including molecular sorting of early endosomes to distinct organelles, its recycling and maturation to late endosomes or to the trans-Golgi network (TGN) (Fig. 1). The role of these endocytic proteins in metastasis in vivo and their prognostic potential, if any, have been listed in Table 1.
A wide variety of receptors and their ligands are moved intracellularly by endocytosis. Clathrin-mediated endocytosis begins with initiation and maturation of clathrin-coated pits by AP2 complexes that are recruited to the plasma membrane and act as a principal cargo-recognition molecule. As the nascent invagination grows, AP2 and other cargo-specific adaptor proteins recruit and concentrate the cargo. AP2 complexes along with other adaptor proteins to recruit clathrin. Clathrin recruitment stabilises the curvature of the growing pit with the help of other BAR-domain-containing proteins. BAR-domain-containing proteins also recruit Dynamin to the neck of the budding vesicle, until the entire region invaginates to form a closed vesicle. Dynamin is a large GTPase, which forms a helical oligomer around the constricted neck and, upon GTP hydrolysis, mediates the fission of the vesicle to release it into the cytoplasm. Following vesicle detachment from the plasma membrane, the clathrin coat is disassembled. The released vesicle goes through a first set of fusion, leading to formation of early endosomes, where initial sorting decisions are made, and the fate of the internalised sorting proteins and lipids is decided. The RAB proteins primarily localised to the early endosome include RAB5 and RAB4, along with lesser-known RAB21 and RAB22. They regulate the motility of early endosome on actin and microtubule tracks, its homotypic fusion and specialised functions of sorting and trafficking. The internalised receptors can be sorted into recycling pathways through extensive tubulation of the early endosome membranes, wherein receptors that are sorted into the newly formed tubular membranes recycle back to the plasma membrane through recycling endosomes. Alternately, early endosome growth and maturation could lead to the trans-Golgi network (TGN) or to late endosomes. Mature late endosomes are approximately 250–1000 nm in diameter and are characterised by the generation of a RAB7 domain. Late endosomes undergo homotypic fusion reactions, grow in size and acquire more intralumenal vesicles (ILVs). ILVs containing late endosomes get enriched with RAB35 and RAB27 and their effectors that promote their fusion to plasma membrane to release exosomes (40–100 nm in diameter vesicles). Predominantly, late endosomes move to the perinuclear area of the cell where they undergo transient fusions with each other and eventually fuse with lysosomes for degradation of its content. Cellular proteins synthesised in the rough endoplasmic reticulum (ER) are constantly secreted from ER to the Golgi complex in mammals through an ER–Golgi intermediate compartment (ERGIC). Points where metastasis suppressors interact with the endocytic process are highlighted.
A recycling pathway returns endosomes to the cell surface either by a fast recycling route (via RAB4-positive endosomes) or by a slow recycling route (via RAB11-positive endosomes).45 Internalised receptors in early endosomes can be sorted into the recycling pathway through an extensive tubulation of the early endosome membranes in a process called ‘geometry-based sorting’ wherein receptors that are sorted into the newly formed tubular membranes of the early endosome are recycled back to the plasma membrane. Intralumenal vesicles (ILVs) also form in early endosomes, driven by clathrin and components of the endosomal sorting complex required for transport (ESCRT).46 ESCRT-mediated receptor sorting into ILVs is an evolutionarily conserved process that is required for multivesicular body (MVB) formation. ESCRT uses its various complexes for receptor recognition (ESCRT-0), inward budding (ESCRT-I and II) and final ESCRT-III-mediated abscission.47 This separates the cytoplasmic portion of the receptors from the rest of the cell, leading to abrogation of its signalling. Interestingly, depletion of ESCRT-0 and ESCRT-I subunits inhibits the degradation of EGFR and results in enhanced recycling and sustained activation of extracellular signal-regulated kinase (ERK) signalling.48,49
A role for endosomal acidification and ligand dissociation has also been established. Recycling of receptors to the plasma membrane takes place if the ligands are released in the early endosome (i.e., transferrin receptor), where the pH is maintained at ~6.5.50 Conversely, some signalling receptors (i.e., EGFR) often retain ligand binding and remain active even at low (~4.5) pH, leading to their continual signalling from endosomal compartments until they are sorted into ILVs and degraded in the lysosome.51
Some internalised receptors in early endosomes can be sorted to the TGN in a process called retrograde transport (i.e., mannose-6-phosphate receptors and several toxins such as Shiga, cholera and ricin). The TGN is a network of interconnected tubules and vesicles at the trans-face of the Golgi apparatus. It is essential for maintaining cellular homoeostasis and is known to play a crucial role in protein sorting or diverting proteins and lipids away from lysosomal degradation.
Mature late endosomes are approximately 250–1000 nm in diameter and are round/oval in shape. They are characterised by the presence of RAB7-GTPase, which is fundamental for the maturation of early-to-late endosomes and for the lysosomal biogenesis. Maturation of early-to-late endosomes depends on the formation of a hybrid RAB5/RAB7 endosome, wherein RAB7 is recruited to the early endosome by RAB5-GTP.52 Late endosomes undergo homotypic fusion reactions, grow in size and acquire more intraluminal vesicles. Once intralumenal vesicles containing late endosomes become enriched with RAB35, RAB27A, RAB27B and their effectors Slp4 and Slac2b, they fuse to plasma membrane to release exosomes.37 The released exosomes are small (40–100 nm in diameter), single membrane-bound vesicles that contain protein, DNA and RNA. Mostly, however, late endosomes move to the perinuclear area of the cell in the vicinity of lysosomes using dynein-dependent transport. Here, late endosomes undergo transient fusions with each other and eventually fuse with lysosomes to generate a transient hybrid organelle called the endolysosome. It is in the endolysosomes in which most of the hydrolysis of endocytosed cargo takes place.37 Following a further maturation process, the endolysosome is converted into a classical dense lysosome.
Cellular contents and organelles can also be delivered to lysosomes through a separate pathway called autophagy. Autophagy or self-eating is a unique membrane trafficking process whereby a newly formed isolation membrane can elongate and engulf part of the cytoplasm or organelles to form autophagosomes that are delivered to the lysosome for degradation. There are an increasing number of reports pointing to a mechanistic role for autophagy in the process of tumour metastasis, detailed in a recent review.53
An astonishing number of endosomal trafficking pathway proteins are known to be functionally important in tumour progression and metastasis (Table 1). Many have been validated in cancer cell motility and invasion, but a considerable number have been shown to modulate in vivo metastasis. The alterations identified include up- or down-regulation of expression, or mutation, and generally lead to an aberrant receptor trafficking/recycling/degradation/signal duration, which has a profound effect on cancer cell migration, invasion and/or proliferation. While most of these reports focus on a single signalling pathway, it is likely that multiple pathways are also affected. These mechanistic studies cover a wide range of cancer types. Additional details on different endosomal trafficking members and their role(s) in cancer and metastasis can be found in recent reviews.54,55,56
Integrin and extracellular matrix trafficking in metastasis
Cancer cells invade through the extracellular matrix (ECM) in part by producing matrix metalloproteinases (MMPs) and other proteinases that degrade the ECM, thereby creating paths for migration. Similarly, cells attach to the ECM by means of integrins that are key regulators of cell adhesion, migration and proliferation. The interplay between integrins and ECM remodelling proteases is a major regulator of tumour invasion.
In oral squamous cell carcinoma (SCC), increased αVβ6 integrin expression leads to the activation of MMP-3 and promotes oral SCC cell proliferation and metastasis in vivo.57 MMP-14 (membrane type 1 metalloprotease MT1-MMP), along with integrin αVβ3 co-localised to the protruding ends of invadopodia, and its high local concentration on the cell membrane promoted metastasis.58 Interestingly, WDFY2 (a cytosolic protein) controls the recycling of MT1-MMP to the membrane, and loss of WDFY2 leads to enhanced secretion of MT1-MMP leading to active invasion of cells.59
Recent studies highlight the importance of integrin trafficking (endocytosis and recycling) as a modulator of cancer cells’ fate. For example, rapid recycling of integrins from the leading edge of individual cells assists in efficient cell motility by providing a supply of fresh receptors that are internalised at the trailing edge. More details on the trafficking of MMPs and integrins and its role in metastasis can be found in recent reviews.60,61
Metastasis suppressors and endocytosis
Metastasis suppressors are a group of genes that suppress the metastatic potential of cancer cells without significantly affecting the size of primary tumour.62 So far, more than 20 metastasis suppressor genes (including miRNAs) have been identified in multiple cancer types with a wide range of biochemical activities.63 Some of the metastasis suppressor genes working through alterations in endocytosis are described below:
NME1 (NM23/NM23-H1, non-metastatic clone 23, isoform H1)
NME is a multifunctional protein that is highly conserved from yeast to humans. Its enforced expression suppressed metastasis in a variety of cancer cell lines without altering primary tumour growth.64 Apart from being a metastasis suppressor, it is also known to have a developmental function.
The Drosophila homologue of NME is awd (abnormal wing discs) and is known to regulate cell differentiation and motility of multiple organs in late embryogenesis by regulating growth factor receptor signalling through endocytosis. These studies identified a genetic interaction between awd and dynamin (shi).65 An aberrant endocytosis was associated with mutant awd phenotypes and complemented RAB5 or shi genes.65,66,67 It was also shown that awd regulated tracheal cell motility in development by modulating the fibroblast growth factor receptor (FGFR) levels through dynamin-mediated endocytosis.65,68 Interestingly, loss of awd gene also blocked Notch signalling by altering the receptor processing that leads to Notch accumulation in the early endosomes.67
Recent reports in mammalian cancer models have also highlighted the role of NME as an interacting partner of Dynamin in endocytosis.69,70 NME transfectants of multiple cell lines exhibited increased endocytosis of EGFR and transferrin in concert with motility suppression. Both the increased endocytic and motility-suppression phenotypes were blocked by inhibitors of Dynamin. In a lung-metastasis assay, NME1 overexpression failed to significantly suppress metastasis in cells in which Dynamin 2 was also knocked down. Using the EGF/EGFR signalling axis as an in vitro model, NME1 decreased the phospho-EGFR and phospho-Akt levels in a Dynamin 2-dependent manner, highlighting the relevance of this interaction for downstream signalling. It was speculated that NME acted as a GTP provider/oligomerising agent of Dynamin 2, leading to higher Dynamin 2 GTPase activity and increased endocytosis (Fig. 1).69,70 Our data identified another function of a NME–Dynamin interaction: in vitro, NME promoted the oligomerisation of Dynamin and its increased GTPase activity, which are needed for vesicle scission.69
KAI1 (CD82, cluster of differentiation 82)
KAI1/CD82 is a member of the evolutionarily conserved tetraspanin family, and was initially identified as a metastasis suppressor in prostate cancer.71 KAI1 has since been established as a metastasis suppressor in a variety of solid tumours. Its higher expression predicts a better prognosis,72,73,74 whereas reduced expression of KAI1 has been widely correlated with an aggressive cancer in several cancer types, including pancreatic, hepatocellular, bladder, breast and non-small-cell lung cancers.73,75,76
KAI1-mediated suppression of metastasis is thought to be achieved primarily by inhibiting cancer cell migration and invasion.77 This phenotype is the result of forming oligomeric complexes with binding partners such as integrins, EGFR and intracellular signalling proteins, such as protein kinase C (PKC). This complex generally leads to either redistribution or increased internalisation of multiple receptors. For example, overexpression of KAI1 leads to redistribution of urokinase-type plasminogen activator receptor (uPAR) into a stable complex with integrin α5β1 in focal adhesions.78 Focal adhesion binding of uPAR reduces its ability to bind the ligand uPA and consequently to cleave and activate plasminogen. Similarly, KAI1 also binds with EGFR, ErbB2 and ErbB3; for EGFR, this leads to accelerated endocytosis and desensitisation.79,80 KAI1 also specifically inhibits ligand-induced EGFR dimerisation and alters the distribution of EGFR in the plasma membrane, which consequently affects its activation.80
MTSS1 (metastasis suppressor protein 1 or MIM, missing in metastasis)
MTSS1/MIM, originally identified in bladder cancer cell lines, was present in non-metastatic but not metastatic bladder cancer cells.81 It is hypothesised that MTSS1 suppresses metastasis by acting as a scaffold protein to interact with actin-associated proteins to regulate cytoskeletal dynamics and lamellipodia formation, consequently affecting invasion and metastatic behaviour of cancer cells.82 In head and neck squamous cell carcinoma, MTSS1 augments EGF signalling by antagonising EGFR endocytosis at low cell densities and promotes cellular proliferation at early stages of primary head and neck squamous cell carcinoma tumour growth. However, at high cell densities, MTSS1 has a negative impact on EGF signalling and inhibits metastasis.83
KISS1 (kisspeptin-1)
The KISS1 gene produces a peptide product called kisspeptins (KP), which act as an endogenous ligand for a G-protein-coupled receptor, KISS1R (GPR54).84 KISS1 acts as a metastasis suppressor gene through its KP/KISS1R signalling in numerous human cancers (melanoma, pancreatic cancer and gastric carcinoma) by inhibiting cellular motility, proliferation, invasion, chemotaxis and metastasis.85 However, in breast cancer, KP stimulates invasion of cancer cells and high expression of KISS1; GPR54 mRNA levels positively correlated with shorter relapse-free survival. Interestingly, GPR54 directly complexes with EGFR, and stimulation of breast cancer cells by either EGF or KP-10 regulated the endocytosis of both GPR54 and EGFR.86 This signalling has an opposite effect on breast cancer cells, i.e., it is pro-migratory and pro-invasive in human breast cancer cells.
Metastasis suppressor genes, while often showing statistically significant inverse trends of tumour expression and patient survival, are not likely to become clinically used prognostic factors, in favour of more complex gene signatures. As with tumour suppressors, their translation to the clinic has also been problematic. Restoration of metastasis suppressor expression in every metastatic tumour cell would be needed for optimal activity, which is unrealistic. Our laboratory explored the transcriptional upregulation of NME by high-dose medroxyprogesterone acetate.87 A Phase 2 trial, conducted at Indiana University, was a technical failure, as serum levels of medroxyprogesterone acetate were not sufficiently elevated, although some long-term stable disease was observed.88 How the endocytic pathways can contribute to a metastatic-suppressor clinical–translational effort is currently unknown but of high interest. More research to identify the complex mechanisms underlying these processes is warranted.
Conclusions
Endocytosis is a process of internalisation of the plasma membrane along with its membrane proteins and lipids. Cells use endocytosis to regulate signalling and to sample the extracellular milieu for appropriate responses. It affects almost all of the steps of metastasis and is used as a tool for the functioning of metastasis suppressors. Based on the literature, endocytosis regulates receptor internalisation, recycling and degradation, or could affect cytoskeleton dynamics to alter cancer cell invasion or metastasis. However, the majority of the above conclusions have been made based on studies conducted on cancer cell lines. These studies would benefit from validation on patient-derived tissues. Other challenges in this field are a lack of high-resolution knowledge of the endosomal sorting complexes and their central regulators, and how signalling in cancer cells is altered at specific stages of endocytosis. These issues will undoubtedly be clarified as research progresses. Identification of these central regulators could serve as trafficking nodes that are amenable to therapeutic interception. A potential issue in translation is the effect of an inhibitor of an endocytic node on multiple signalling pathways that it engages, and how the cumulative effects modulate the metastatic phenotype. This issue is not unique to endocytosis and applies to DNA methylation and other cancer processes. In summary, targeting the endocytic machinery could be a viable and promising therapeutic strategy for cancer and metastasis.
References
Siegel, R., Naishadham, D. & Jemal, A. Cancer statistics, 2013. CA Cancer J. Clin. 63, 11–30 (2013).
Jemal, A., Ward, E. M., Johnson, C. J., Cronin, K. A., Ma, J., Ryerson, B. et al. Annual report to the nation on the status of cancer, 1975–2014, featuring survival. J. Natl Cancer Inst. 109, djx030 (2017).
Chambers, A. F., Groom, A. C. & MacDonald, I. C. Dissemination and growth of cancer cells in metastatic sites. Nat. Rev. Cancer 2, 563–572 (2002).
Underhill, D. M. & Ozinsky, A. Phagocytosis of microbes: complexity in action. Annu Rev. Immunol. 20, 825–852 (2002).
Haigler, H. T., McKanna, J. A. & Cohen, S. Rapid stimulation of pinocytosis in human carcinoma cells A-431 by epidermal growth factor. J. Cell Biol. 83, 82–90 (1979).
Carpentier, J. L., Gorden, P., Anderson, R. G., Goldstein, J. L., Brown, M. S., Cohen, S. et al. Co-localization of 125I-epidermal growth factor and ferritin-low density lipoprotein in coated pits: a quantitative electron microscopic study in normal and mutant human fibroblasts. J. Cell Biol. 95, 73–77 (1982).
Neutra, M. R., Ciechanover, A., Owen, L. S. & Lodish, H. F. Intracellular transport of transferrin- and asialoorosomucoid-colloidal gold conjugates to lysosomes after receptor-mediated endocytosis. J. Histochem Cytochem 33, 1134–1144 (1985).
Takei, K. & Haucke, V. Clathrin-mediated endocytosis: membrane factors pull the trigger. Trends Cell Biol. 11, 385–391 (2001).
Traub, L. M. & Bonifacino, J. S. Cargo recognition in clathrin-mediated endocytosis. Cold Spring Harb. Perspect. Biol. 5, a016790 (2013).
Sorkin, A. Cargo recognition during clathrin-mediated endocytosis: a team effort. Curr. Opin. Cell Biol. 16, 392–399 (2004).
Qualmann, B., Koch, D. & Kessels, M. M. Let’s go bananas: revisiting the endocytic BAR code. EMBO J. 30, 3501–3515 (2011).
Ferguson, S. M., Raimondi, A., Paradise, S., Shen, H., Mesaki, K., Ferguson, A. et al. Coordinated actions of actin and BAR proteins upstream of dynamin at endocytic clathrin-coated pits. Dev. Cell 17, 811–822 (2009).
Barylko, B., Binns, D., Lin, K. M., Atkinson, M. A., Jameson, D. M., Yin, H. L. et al. Synergistic activation of dynamin GTPase by Grb2 and phosphoinositides. J. Biol. Chem. 273, 3791–3797 (1998).
Schmid, S. L. & Frolov, V. A. Dynamin: functional design of a membrane fission catalyst. Annu Rev. Cell Dev. Biol. 27, 79–105 (2011).
Newmyer, S. L. & Schmid, S. L. Dominant-interfering Hsc70 mutants disrupt multiple stages of the clathrin-coated vesicle cycle in vivo. J. Cell Biol. 152, 607–620 (2001).
Ungewickell, E., Ungewickell, H., Holstein, S. E., Lindner, R., Prasad, K., Barouch, W. et al. Role of auxilin in uncoating clathrin-coated vesicles. Nature 378, 632–635 (1995).
Chen, P. H., Bendris, N., Hsiao, Y. J., Reis, C. R., Mettlen, M., Chen, H. Y. et al. Crosstalk between CLCb/Dyn1-mediated adaptive clathrin-mediated endocytosis and epidermal growth factor receptor signaling increases metastasis. Dev. Cell 40, 278–288 e275 (2017).
Xu, B., Teng, L. H., Silva, S. D., Bijian, K., Al Bashir, S., Jie, S. et al. The significance of dynamin 2 expression for prostate cancer progression, prognostication, and therapeutic targeting. Cancer Med. 3, 14–24 (2014).
Di Guglielmo, G. M., Le Roy, C., Goodfellow, A. F. & Wrana, J. L. Distinct endocytic pathways regulate TGF-beta receptor signalling and turnover. Nat. Cell Biol. 5, 410–421 (2003).
Sigismund, S., Argenzio, E., Tosoni, D., Cavallaro, E., Polo, S. & Di Fiore, P. P. Clathrin-mediated internalization is essential for sustained EGFR signaling but dispensable for degradation. Dev. Cell 15, 209–219 (2008).
Wang, Y., Pennock, S., Chen, X. & Wang, Z. Endosomal signaling of epidermal growth factor receptor stimulates signal transduction pathways leading to cell survival. Mol. Cell Biol. 22, 7279–7290 (2002).
Levkowitz, G., Waterman, H., Zamir, E., Kam, Z., Oved, S., Langdon, W. Y. et al. c-Cbl/Sli-1 regulates endocytic sorting and ubiquitination of the epidermal growth factor receptor. Genes Dev. 12, 3663–3674 (1998).
Parton, R. G. & Simons, K. The multiple faces of caveolae. Nat. Rev. Mol. Cell Biol. 8, 185–194 (2007).
Gambin, Y., Ariotti, N., McMahon, K. A., Bastiani, M., Sierecki, E., Kovtun, O. et al. Single-molecule analysis reveals self assembly and nanoscale segregation of two distinct cavin subcomplexes on caveolae. Elife 3, e01434 (2013).
Kiss, A. L. Caveolae and the regulation of endocytosis. Adv. Exp. Med Biol. 729, 14–28 (2012).
Henley, J. R., Krueger, E. W., Oswald, B. J. & McNiven, M. A. Dynamin-mediated internalization of caveolae. J. Cell Biol. 141, 85–99 (1998).
Quest, A. F., Gutierrez-Pajares, J. L. & Torres, V. A. Caveolin-1: an ambiguous partner in cell signalling and cancer. J. Cell Mol. Med 12, 1130–1150 (2008).
Goetz, J. G., Lajoie, P., Wiseman, S. M. & Nabi, I. R. Caveolin-1 in tumor progression: the good, the bad and the ugly. Cancer Metastasis Rev. 27, 715–735 (2008).
Quest, A. F., Leyton, L. & Parraga, M. Caveolins, caveolae, and lipid rafts in cellular transport, signaling, and disease. Biochem Cell Biol. 82, 129–144 (2004).
Engelman, J. A., Zhang, X. L., Galbiati, F. & Lisanti, M. P. Chromosomal localization, genomic organization, and developmental expression of the murine caveolin gene family (Cav-1, -2, and -3). Cav-1 and Cav-2 genes map to a known tumor suppressor locus (6-A2/7q31). FEBS Lett. 429, 330–336 (1998).
Joshi, B., Strugnell, S. S., Goetz, J. G., Kojic, L. D., Cox, M. E., Griffith, O. L. et al. Phosphorylated caveolin-1 regulates Rho/ROCK-dependent focal adhesion dynamics and tumor cell migration and invasion. Cancer Res. 68, 8210–8220 (2008).
Urra, H., Torres, V. A., Ortiz, R. J., Lobos, L., Diaz, M. I., Diaz, N. et al. Caveolin-1-enhanced motility and focal adhesion turnover require tyrosine-14 but not accumulation to the rear in metastatic cancer cells. PLoS ONE 7, e33085 (2012).
Chen, D. & Che, G. Value of caveolin-1 in cancer progression and prognosis: emphasis on cancer-associated fibroblasts, human cancer cells and mechanism of caveolin-1 expression (Review). Oncol. Lett. 8, 1409–1421 (2014).
Moya, M., Dautry-Varsat, A., Goud, B., Louvard, D. & Boquet, P. Inhibition of coated pit formation in Hep2 cells blocks the cytotoxicity of diphtheria toxin but not that of ricin toxin. J. Cell Biol. 101, 548–559 (1985).
Lamaze, C., Dujeancourt, A., Baba, T., Lo, C. G., Benmerah, A. & Dautry-Varsat, A. Interleukin 2 receptors and detergent-resistant membrane domains define a clathrin-independent endocytic pathway. Mol. Cell 7, 661–671 (2001).
Kirkham, M., Fujita, A., Chadda, R., Nixon, S. J., Kurzchalia, T. V., Sharma, D. K. et al. Ultrastructural identification of uncoated caveolin-independent early endocytic vehicles. J. Cell Biol. 168, 465–476 (2005).
Huotari, J. & Helenius, A. Endosome maturation. EMBO J. 30, 3481–3500 (2011).
Donaldson, J. G. & Jackson, C. L. ARF family G proteins and their regulators: roles in membrane transport, development and disease. Nat. Rev. Mol. Cell Biol. 12, 362–375 (2011).
Holst, M. R., Vidal-Quadras, M., Larsson, E., Song, J., Hubert, M., Blomberg, J. et al. Clathrin-independent endocytosis suppresses cancer cell blebbing and invasion. Cell Rep. 20, 1893–1905 (2017).
Mayor, S., Parton, R. G. & Donaldson, J. G. Clathrin-independent pathways of endocytosis. Cold Spring Harb. Perspect. Biol. 6, a016758 (2014).
Christoforidis, S., Miaczynska, M., Ashman, K., Wilm, M., Zhao, L., Yip, S. C. et al. Phosphatidylinositol-3-OH kinases are Rab5 effectors. Nat. Cell Biol. 1, 249–252 (1999).
Zerial, M. & McBride, H. Rab proteins as membrane organizers. Nat. Rev. Mol. Cell Biol. 2, 107–117 (2001).
Rojas, R., van Vlijmen, T., Mardones, G. A., Prabhu, Y., Rojas, A. L., Mohammed, S. et al. Regulation of retromer recruitment to endosomes by sequential action of Rab5 and Rab7. J. Cell Biol. 183, 513–526 (2008).
Hayer, A., Stoeber, M., Ritz, D., Engel, S., Meyer, H. H. & Helenius, A. Caveolin-1 is ubiquitinated and targeted to intralumenal vesicles in endolysosomes for degradation. J. Cell Biol. 191, 615–629 (2010).
Stenmark, H. Rab GTPases as coordinators of vesicle traffic. Nat. Rev. Mol. Cell Biol. 10, 513–525 (2009).
Raiborg, C., Bache, K. G., Gillooly, D. J., Madshus, I. H., Stang, E. & Stenmark, H. Hrs sorts ubiquitinated proteins into clathrin-coated microdomains of early endosomes. Nat. Cell Biol. 4, 394–398 (2002).
Raiborg, C. & Stenmark, H. The ESCRT machinery in endosomal sorting of ubiquitylated membrane proteins. Nature 458, 445–452 (2009).
Malerod, L., Stuffers, S., Brech, A. & Stenmark, H. Vps22/EAP30 in ESCRT-II mediates endosomal sorting of growth factor and chemokine receptors destined for lysosomal degradation. Traffic 8, 1617–1629 (2007).
Raiborg, C., Malerod, L., Pedersen, N. M. & Stenmark, H. Differential functions of Hrs and ESCRT proteins in endocytic membrane trafficking. Exp. Cell Res 314, 801–813 (2008).
Fuchs, R., Male, P. & Mellman, I. Acidification and ion permeabilities of highly purified rat liver endosomes. J. Biol. Chem. 264, 2212–2220 (1989).
Mellman, I. Endocytosis and molecular sorting. Annu Rev. Cell Dev. Biol. 12, 575–625 (1996).
Novick, P. & Zerial, M. The diversity of Rab proteins in vesicle transport. Curr. Opin. Cell Biol. 9, 496–504 (1997).
Mowers, E. E., Sharifi, M. N. & Macleod, K. F. Autophagy in cancer metastasis. Oncogene 36, 1619–1630 (2017).
Mellman, I. & Yarden, Y. Endocytosis and cancer. Cold Spring Harb. Perspect. Biol. 5, a016949 (2013).
Lanzetti, L. & Di Fiore, P. P. Endocytosis and cancer: an ‘insider’ network with dangerous liaisons. Traffic 9, 2011–2021 (2008).
Schmid, S. L. Reciprocal regulation of signaling and endocytosis: Implications for the evolving cancer cell. J. Cell Biol. 216, 2623–2632 (2017).
Li, X., Yang, Y., Hu, Y., Dang, D., Regezi, J., Schmidt, B. L. et al. Alphavbeta6-Fyn signaling promotes oral cancer progression. J. Biol. Chem. 278, 41646–41653 (2003).
Nakahara, H., Howard, L., Thompson, E. W., Sato, H., Seiki, M., Yeh, Y. et al. Transmembrane/cytoplasmic domain-mediated membrane type 1-matrix metalloprotease docking to invadopodia is required for cell invasion. Proc. Natl Acad. Sci. USA 94, 7959–7964 (1997).
Sneeggen, M., Pedersen, N. M., Campsteijn, C., Haugsten, E. M., Stenmark, H. & Schink, K. O. WDFY2 restrains matrix metalloproteinase secretion and cell invasion by controlling VAMP3-dependent recycling. Nat. Commun. 10, 2850 (2019).
Ramsay, A. G., Marshall, J. F. & Hart, I. R. Integrin trafficking and its role in cancer metastasis. Cancer Metastasis Rev. 26, 567–578 (2007).
Shay, G., Lynch, C. C. & Fingleton, B. Moving targets: emerging roles for MMPs in cancer progression and metastasis. Matrix Biol. 44-46, 200–206 (2015).
Khan, I. & Steeg, P. S. Metastasis suppressors: functional pathways. Lab. Invest. 98, 198–210 (2018).
Hurst, D. R. & Welch, D. R. Metastasis suppressor genes at the interface between the environment and tumor cell growth. Int Rev. Cell Mol. Biol. 286, 107–180 (2011).
Salerno, M., Ouatas, T., Palmieri, D. & Steeg, P. S. Inhibition of signal transduction by the nm23 metastasis suppressor: possible mechanisms. Clin. Exp. Metastasis 20, 3–10 (2003).
Dammai, V., Adryan, B., Lavenburg, K. R. & Hsu, T. Drosophila awd, the homolog of human nm23, regulates FGF receptor levels and functions synergistically with shi/dynamin during tracheal development. Genes Dev. 17, 2812–2824 (2003).
Woolworth, J. A., Nallamothu, G. & Hsu, T. The Drosophila metastasis suppressor gene Nm23 homolog, awd, regulates epithelial integrity during oogenesis. Mol. Cell Biol. 29, 4679–4690 (2009).
Ignesti, M., Barraco, M., Nallamothu, G., Woolworth, J. A., Duchi, S., Gargiulo, G. et al. Notch signaling during development requires the function of awd, the Drosophila homolog of human metastasis suppressor gene Nm23. BMC Biol. 12, 12 (2014).
Nallamothu, G., Woolworth, J. A., Dammai, V. & Hsu, T. Awd, the homolog of metastasis suppressor gene Nm23, regulates Drosophila epithelial cell invasion. Mol. Cell Biol. 28, 1964–1973 (2008).
Khan, I., Gril, B. & Steeg, P. S. Metastasis suppressors NME1 and NME2 promote dynamin 2 oligomerization and regulate tumor cell endocytosis, motility, and metastasis. Cancer Res. 79, 4689–4702 (2019).
Boissan, M., Montagnac, G., Shen, Q., Griparic, L., Guitton, J., Romao, M. et al. Membrane trafficking. Nucleoside diphosphate kinases fuel dynamin superfamily proteins with GTP for membrane remodeling. Science 344, 1510–1515 (2014).
Ichikawa, T., Ichikawa, Y. & Isaacs, J. T. Genetic factors and suppression of metastatic ability of prostatic cancer. Cancer Res 51, 3788–3792 (1991).
Dong, J. T., Isaacs, W. B. & Isaacs, J. T. Molecular advances in prostate cancer. Curr. Opin. Oncol. 9, 101–107 (1997).
Dong, J. T., Suzuki, H., Pin, S. S., Bova, G. S., Schalken, J. A., Isaacs, W. B. et al. Down-regulation of the KAI1 metastasis suppressor gene during the progression of human prostatic cancer infrequently involves gene mutation or allelic loss. Cancer Res. 56, 4387–4390 (1996).
Ow, K., Delprado, W., Fisher, R., Barrett, J., Yu, Y., Jackson, P. et al. Relationship between expression of the KAI1 metastasis suppressor and other markers of advanced bladder cancer. J. Pathol. 191, 39–47 (2000).
Kawana, Y., Komiya, A., Ueda, T., Nihei, N., Kuramochi, H., Suzuki, H. et al. Location of KAI1 on the short arm of human chromosome 11 and frequency of allelic loss in advanced human prostate cancer. Prostate 32, 205–213 (1997).
Uchida, S., Shimada, Y., Watanabe, G., Li, Z. G., Hong, T., Miyake, M. et al. Motility-related protein (MRP-1/CD9) and KAI1/CD82 expression inversely correlate with lymph node metastasis in oesophageal squamous cell carcinoma. Br. J. Cancer 79, 1168–1173 (1999).
Yang, X., Wei, L. L., Tang, C., Slack, R., Mueller, S. & Lippman, M. E. Overexpression of KAI1 suppresses in vitro invasiveness and in vivo metastasis in breast cancer cells. Cancer Res. 61, 5284–5288 (2001).
Bass, R., Werner, F., Odintsova, E., Sugiura, T., Berditchevski, F. & Ellis, V. Regulation of urokinase receptor proteolytic function by the tetraspanin CD82. J. Biol. Chem. 280, 14811–14818 (2005).
Odintsova, E., Sugiura, T. & Berditchevski, F. Attenuation of EGF receptor signaling by a metastasis suppressor, the tetraspanin CD82/KAI-1. Curr. Biol. 10, 1009–1012 (2000).
Odintsova, E., Voortman, J., Gilbert, E. & Berditchevski, F. Tetraspanin CD82 regulates compartmentalisation and ligand-induced dimerization of EGFR. J. Cell Sci. 116, 4557–4566 (2003).
Lee, Y. G., Macoska, J. A., Korenchuk, S. & Pienta, K. J. MIM, a potential metastasis suppressor gene in bladder cancer. Neoplasia 4, 291–294 (2002).
Woodings, J. A., Sharp, S. J. & Machesky, L. M. MIM-B, a putative metastasis suppressor protein, binds to actin and to protein tyrosine phosphatase delta. Biochem J. 371, 463–471 (2003).
Dawson, J. C., Timpson, P., Kalna, G. & Machesky, L. M. Mtss1 regulates epidermal growth factor signaling in head and neck squamous carcinoma cells. Oncogene 31, 1781–1793 (2012).
Cvetkovic, D., Babwah, A. V. & Bhattacharya, M. Kisspeptin/KISS1R system in breast cancer. J. Cancer 4, 653–661 (2013).
Beck, B. H. & Welch, D. R. The KISS1 metastasis suppressor: a good night kiss for disseminated cancer cells. Eur. J. Cancer 46, 1283–1289 (2010).
Zajac, M., Law, J., Cvetkovic, D. D., Pampillo, M., McColl, L., Pape, C. et al. GPR54 (KISS1R) transactivates EGFR to promote breast cancer cell invasiveness. PLoS ONE 6, e21599 (2011).
Palmieri, D., Halverson, D., Ouatas, T., Horak, C., Salerno, M., Johnson, J. et al. Medroxyprogesterone acetate elevation of Nm23-H1 metastasis suppressor expression in hormone receptor-negative breast cancer. J. Natl Cancer Inst. 97, 632–642 (2005).
Miller, K. D., Althouse, S. K., Nabell, L., Rugo, H., Carey, L., Kimmick, G. et al. A phase II study of medroxyprogesterone acetate in patients with hormone receptor negative metastatic breast cancer: translational breast cancer research consortium trial 007. Breast Cancer Res. Treat. 148, 99–106 (2014).
Azarnia Tehran, D., Lopez-Hernandez, T. & Maritzen, T. Endocytic adaptor proteins in health and disease: lessons from model organisms and human mutations. Cells 8, 1345 (2019).
Reis, C. R., Chen, P. H., Srinivasan, S., Aguet, F., Mettlen, M. & Schmid, S. L. Crosstalk between Akt/GSK3beta signaling and dynamin-1 regulates clathrin-mediated endocytosis. EMBO J. 34, 2132–2146 (2015).
Boulay, P. L., Schlienger, S., Lewis-Saravalli, S., Vitale, N., Ferbeyre, G. & Claing, A. ARF1 controls proliferation of breast cancer cells by regulating the retinoblastoma protein. Oncogene 30, 3846–3861 (2011).
Schlienger, S., Ramirez, R. A. & Claing, A. ARF1 regulates adhesion of MDA-MB-231 invasive breast cancer cells through formation of focal adhesions. Cell Signal 27, 403–415 (2015).
Rainero, E., Howe, J. D., Caswell, P. T., Jamieson, N. B., Anderson, K., Critchley, D. R. et al. Ligand-occupied Integrin Internalization Links Nutrient Signaling to Invasive Migration. Cell Rep. 10, 398–413 (2015).
Palacios, F., Schweitzer, J. K., Boshans, R. L. & D’Souza-Schorey, C. ARF6-GTP recruits Nm23-H1 to facilitate dynamin-mediated endocytosis during adherens junctions disassembly. Nat. Cell Biol. 4, 929–936 (2002).
Palacios, F., Price, L., Schweitzer, J., Collard, J. G. & D’Souza-Schorey, C. An essential role for ARF6-regulated membrane traffic in adherens junction turnover and epithelial cell migration. EMBO J. 20, 4973–4986 (2001).
Schlienger, S., Campbell, S., Pasquin, S., Gaboury, L. & Claing, A. ADP-ribosylation factor 1 expression regulates epithelial-mesenchymal transition and predicts poor clinical outcome in triple-negative breast cancer. Oncotarget 7, 15811–15827 (2016).
Miao, B., Skidan, I., Yang, J., You, Z., Fu, X., Famulok, M. et al. Inhibition of cell migration by PITENINs: the role of ARF6. Oncogene 31, 4317–4332 (2012).
Kalpana, G., Figy, C., Yeung, M. & Yeung, K. C. Reduced RhoA expression enhances breast cancer metastasis with a concomitant increase in CCR5 and CXCR4 chemokines signaling. Sci. Rep. 9, 16351 (2019).
Juin, A., Spence, H. J., Martin, K. J., McGhee, E., Neilson, M., Cutiongco, M. F. A. et al. N-WASP control of LPAR1 trafficking establishes response to self-generated LPA gradients to promote pancreatic cancer cell metastasis. Dev. Cell 51, 431–445 e437 (2019).
Ridley, A. J., Paterson, H. F., Johnston, C. L., Diekmann, D. & Hall, A. The small GTP-binding protein rac regulates growth factor-induced membrane ruffling. Cell 70, 401–410 (1992).
Morrison Joly, M., Williams, M. M., Hicks, D. J., Jones, B., Sanchez, V., Young, C. D. et al. Two distinct mTORC2-dependent pathways converge on Rac1 to drive breast cancer metastasis. Breast Cancer Res. 19, 74 (2017).
Yoon, C., Cho, S. J., Chang, K. K., Park, D. J., Ryeom, S. W. & Yoon, S. S. Role of Rac1 pathway in epithelial-to-mesenchymal transition and cancer stem-like cell phenotypes in gastric adenocarcinoma. Mol. Cancer Res. 15, 1106–1116 (2017).
Xiao, X. H., Lv, L. C., Duan, J., Wu, Y. M., He, S. J., Hu, Z. Z. et al. Regulating Cdc42 and Its signaling pathways in cancer: small molecules and microRNA as new treatment candidates. Molecules 23, 787 (2018).
Jiang, H. L., Sun, H. F., Gao, S. P., Li, L. D., Hu, X., Wu, J. et al. Loss of RAB1B promotes triple-negative breast cancer metastasis by activating TGF-beta/SMAD signaling. Oncotarget 6, 16352–16365 (2015).
Yang, X. Z., Li, X. X., Zhang, Y. J., Rodriguez-Rodriguez, L., Xiang, M. Q., Wang, H. Y. et al. Rab1 in cell signaling, cancer and other diseases. Oncogene 35, 5699–5704 (2016).
Kajiho, H., Kajiho, Y., Frittoli, E., Confalonieri, S., Bertalot, G., Viale, G. et al. RAB2A controls MT1-MMP endocytic and E-cadherin polarized Golgi trafficking to promote invasive breast cancer programs. EMBO Rep. 17, 1061–1080 (2016).
Chang, Y. C., Su, C. Y., Chen, M. H., Chen, W. S., Chen, C. L. & Hsiao, M. Secretory RAB GTPase 3C modulates IL6-STAT3 pathway to promote colon cancer metastasis and is associated with poor prognosis. Mol. Cancer 16, 135 (2017).
Yang, J., Liu, W., Lu, X., Fu, Y., Li, L. & Luo, Y. High expression of small GTPase Rab3D promotes cancer progression and metastasis. Oncotarget 6, 11125–11138 (2015).
Do, M. T., Chai, T. F., Casey, P. J. & Wang, M. Isoprenylcysteine carboxylmethyltransferase function is essential for RAB4A-mediated integrin beta3 recycling, cell migration and cancer metastasis. Oncogene 36, 5757–5767 (2017).
Mendoza, P., Ortiz, R., Diaz, J., Quest, A. F., Leyton, L., Stupack, D. et al. Rab5 activation promotes focal adhesion disassembly, migration and invasiveness in tumor cells. J. Cell Sci. 126, 3835–3847 (2013).
Diaz, J., Mendoza, P., Ortiz, R., Diaz, N., Leyton, L., Stupack, D. et al. Rab5 is required in metastatic cancer cells for Caveolin-1-enhanced Rac1 activation, migration and invasion. J. Cell Sci. 127, 2401–2406 (2014).
Silva, P., Mendoza, P., Rivas, S., Diaz, J., Moraga, C., Quest, A. F. et al. Hypoxia promotes Rab5 activation, leading to tumor cell migration, invasion and metastasis. Oncotarget 7, 29548–29562 (2016).
Pellinen, T., Arjonen, A., Vuoriluoto, K., Kallio, K., Fransen, J. A. & Ivaska, J. Small GTPase Rab21 regulates cell adhesion and controls endosomal traffic of beta1-integrins. J. Cell Biol. 173, 767–780 (2006).
Mizuta, R., LaSalle, J. M., Cheng, H. L., Shinohara, A., Ogawa, H., Copeland, N. et al. RAB22 and RAB163/mouse BRCA2: proteins that specifically interact with the RAD51 protein. Proc. Natl Acad. Sci. USA 94, 6927–6932 (1997).
Manshouri, R., Coyaud, E., Kundu, S. T., Peng, D. H., Stratton, S. A., Alton, K. et al. ZEB1/NuRD complex suppresses TBC1D2b to stimulate E-cadherin internalization and promote metastasis in lung cancer. Nat. Commun. 10, 5125 (2019).
Yu, M. H., Luo, Y., Qin, S. L. & Zhong, M. Increased expression of Rab5A predicts metastasis and poor prognosis in colorectal cancer patients. Int J. Clin. Exp. Pathol. 8, 6974–6980 (2015).
Xu, W., Shi, Q., Qian, X., Zhou, B., Xu, J., Zhu, L. et al. Rab5a suppresses autophagy to promote drug resistance in cancer cells. Am. J. Transl. Res 10, 1229–1236 (2018).
Vestre, K., Kjos, I., Guadagno, N. A., Borg Distefano, M., Kohler, F., Fenaroli, F. et al. Rab6 regulates cell migration and invasion by recruiting Cdc42 and modulating its activity. Cell Mol. Life Sci. 76, 2593–2614 (2019).
Alonso-Curbelo, D., Riveiro-Falkenbach, E., Perez-Guijarro, E., Cifdaloz, M., Karras, P., Osterloh, L. et al. RAB7 controls melanoma progression by exploiting a lineage-specific wiring of the endolysosomal pathway. Cancer Cell 26, 61–76 (2014).
Yoon, S. O., Shin, S. & Mercurio, A. M. Hypoxia stimulates carcinoma invasion by stabilizing microtubules and promoting the Rab11 trafficking of the alpha6beta4 integrin. Cancer Res. 65, 2761–2769 (2005).
Gundry, C., Marco, S., Rainero, E., Miller, B., Dornier, E., Mitchell, L. et al. Phosphorylation of Rab-coupling protein by LMTK3 controls Rab14-dependent EphA2 trafficking to promote cell:cell repulsion. Nat. Commun. 8, 14646 (2017).
Hendrix, A., Maynard, D., Pauwels, P., Braems, G., Denys, H., Van den Broecke, R. et al. Effect of the secretory small GTPase Rab27B on breast cancer growth, invasion, and metastasis. J. Natl Cancer Inst. 102, 866–880 (2010).
Ye, B., Duan, B., Deng, W., Wang, Y., Chen, Y., Cui, J. et al. EGF Stimulates Rab35 activation and gastric cancer cell migration by regulating DENND1A-Grb2 complex formation. Front. Pharm. 9, 1343 (2018).
Novo, D., Heath, N., Mitchell, L., Caligiuri, G., MacFarlane, A., Reijmer, D. et al. Mutant p53s generate pro-invasive niches by influencing exosome podocalyxin levels. Nat. Commun. 9, 5069 (2018).
Author information
Authors and Affiliations
Contributions
I.K. and P.S.S. reviewed the literature, drafted and revised the paper.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent to publish
Not applicable.
Data availability
Not applicable.
Competing interests
The authors declare no competing interests.
Funding information
This work is supported by the NIH Intramural program.
Additional information
Note This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution 4.0 International (CC BY 4.0).
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Khan, I., Steeg, P.S. Endocytosis: a pivotal pathway for regulating metastasis. Br J Cancer 124, 66–75 (2021). https://doi.org/10.1038/s41416-020-01179-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-020-01179-8
This article is cited by
-
Golgi-associated retrograde protein (GARP) complex-dependent endosomes to trans Golgi network retrograde trafficking is controlled by Rab4b
Cellular & Molecular Biology Letters (2024)
-
CPMI: comprehensive neighborhood-based perturbed mutual information for identifying critical states of complex biological processes
BMC Bioinformatics (2024)
-
Proteomic characterization of hUC-MSC extracellular vesicles and evaluation of its therapeutic potential to treat Alzheimer’s disease
Scientific Reports (2024)
-
PRUNE1 and NME/NDPK family proteins influence energy metabolism and signaling in cancer metastases
Cancer and Metastasis Reviews (2024)
-
The battle within: cell death by phagocytosis in cancer
Clinical and Translational Oncology (2024)