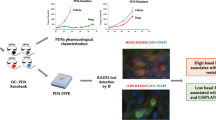
Abstract
Background
Olaparib (Lynparza™) is a PARP inhibitor approved for advanced BRCA-mutated (BRCAm) ovarian cancer. PARP inhibitors may benefit patients whose tumours are dysfunctional in DNA repair mechanisms unrelated to BRCA1/2. We report exploratory analyses, including the long-term outcome of candidate biomarkers of sensitivity to olaparib in BRCA wild-type (BRCAwt) tumours.
Methods
Tumour samples from an olaparib maintenance monotherapy trial (Study 19, D0810C00019; NCT00753545) were analysed. Analyses included classification of mutations in genes involved in homologous recombination repair (HRR), BRCA1 promoter methylation status, measurement of BRCA1 protein and Myriad HRD score.
Results
Patients with BRCAm tumours gained most benefit from olaparib; a similar treatment benefit was also observed in 21/95 patients whose tumours were BRCAwt but had loss-of-function HRR mutations compared to patients with no detectable HRR mutations (58/95). A higher median Myriad MyChoice® HRD score was observed in BRCAm and BRCAwt tumours with BRCA1 methylation. Patients without BRCAm tumours derived benefit from olaparib treatment vs placebo although to a lesser extent than BRCAm patients.
Conclusions
Ovarian cancer patients with tumours harbouring loss-of-function mutations in HRR genes other than BRCA1/2 may constitute a small, molecularly identifiable and clinically relevant population who derive treatment benefit from olaparib similar to patients with BRCAm.
Similar content being viewed by others
Introduction"
Poly(ADP-ribose) polymerase (PARP) plays a vital role in the repair of single-strand DNA breaks through the base excision repair pathway. PARP inhibitors are thought to become trapped at the sites of single-strand DNA breaks leading to double-strand DNA breaks when DNA replication is attempted.1 The double-strand DNA breaks would normally be repaired by the process of homologous recombination repair (HRR), which is a complex process including many proteins, notably BRCA1 and BRCA2.2 Tumours with an HRR deficiency, such as those found in BRCA-mutated cancers, cannot accurately repair DNA damage when PARP protein function is also disrupted and both the base excision and HRR DNA repair pathways are rendered inoperative. In these tumours, DNA repair by low fidelity repair mechanisms such as non-homologous end joining can cause the accumulation of genomic instability, ultimately resulting in cell death; a concept referred to as synthetic lethality.3 Additionally, preclinical data suggest that PARP inhibitors may also benefit patients whose tumours are sensitive to platinum-based chemotherapy and who have an HRR deficiency caused by mutations other than those in the BRCA1/2 genes.4,5 Hence, the clinical and molecular profiles of high-grade serous ovarian cancer (SOC) appeared well matched to PARP inhibitor biology, as this cancer is noted for genome instability thought to be driven by HRR deficiency, repeated and prolonged platinum sensitivity and a high frequency of deficiency in BRCA and other candidate HRR proteins.6
Maintenance treatment with the PARP inhibitor olaparib (Lynparza™) in an AstraZeneca-sponsored, randomised, phase II trial of 265 patients with platinum-sensitive relapsed high-grade SOC (Study 19, D0810C00019; NCT00753545) led to a significant increase in progression-free survival (PFS) vs placebo, which was greatest in the subgroup of 136 patients with BRCA1/2-mutated tumours (hazard ratio 0.18; 95% confidence interval [CI] 0.10–0.31; P < 0.001).7 This subgroup included patients with inherited and somatic (tumour only) mutations.7 Recent analyses strongly suggest that somatic loss-of-function mutations in BRCA1 and BRCA2 are an early event in the development of high-grade SOC and that tumours with somatic BRCA mutations phenocopy tumours in patients with inherited germline BRCA mutations in terms of genetic epidemiology, natural history and response to platinum chemotherapy, and response to olaparib and other PARP inhibitors.8,9,10,11,12,13
Of interest, in Study 19 a significant PFS benefit for olaparib vs placebo was also observed in the subgroup of 118 patients with wild-type BRCA tumours, although the treatment benefit was less (PFS hazard ratio 0.54; 95% CI 0.34–0.85; P < 0.01) than in those patients with BRCA-mutated tumours.7 A similar trend of broad activity in ovarian cancer with a stronger effect in tumours with BRCA mutations has been noted for other PARP inhibitors.11,12,13 Hence, there is considerable interest in understanding the molecular basis of sensitivity to PARP inhibitors in patients whose tumours do not have mutations in the BRCA genes and in tumour tests that may aid in the identification of patients who will benefit most from treatment. In particular, an important question to address is whether in BRCA wild-type tumours, mutations in other HRR genes account for the benefit observed. To further characterise genetic changes in SOC tumours, we conducted exploratory candidate biomarker analyses on BRCA wild-type tumour samples from Study 19 and investigated the possible relationship of HRR deficiencies and clinical benefit.
Materials and methods
Study design and population
Study 19 was a phase II, randomised, double-blind, multicentre trial, undertaken at 82 sites in 16 countries. The study design, patient population and statistical analyses have previously been published in detail.7,14 In brief, eligible patients were aged 18 years or older and had relapsed SOC (Grade 2 or 3) that was platinum sensitive. Patients entering the study were required to have received two or more previous courses of platinum-based chemotherapy and to have demonstrated an objective response (complete or partial) according to Response Evaluation Criteria in Solid Tumors (RECIST) or Gynecologic Cancer InterGroup criteria. Patients were randomised 1:1 to receive either olaparib 400 mg twice daily (b.i.d.) capsules or matching placebo. Study treatment was continued until progression in the absence of unacceptable toxicity. The primary endpoint was PFS, as determined by RECIST v1.0, and overall survival (OS) was a secondary endpoint.
Exploratory analyses
To identify patients that benefit from olaparib that do not have BRCA mutations, exploratory biomarker analyses were conducted on tumour samples from Study 19 including BRCA1 promoter methylation, BRCA1 protein expression, HRR gene mutation, and Myriad homologous recombination deficiency (HRD) testing (which includes a tumour test for BRCA mutations).
BRCA-mutated and BRCA wild-type subgroups in Study 19
Analysis of the BRCA mutation status of patients was prespecified in the Study 19 statistical analysis plan. Molecular analyses to define the BRCA status of patients in Study 19 were completed retrospectively and were blinded to clinical outcomes.7 In brief, the BRCA-mutated population comprised patients with a mutation in their tumour and/or blood sample. Germline BRCA mutation status was either reported on case report forms after local testing or was established retrospectively using the Integrated BRACAnalysis® assay (Myriad Genetics Laboratories, Salt Lake City, UT, USA), with DNA extracted from blood samples obtained prior to randomisation.15
The analyses presented here are based on BRCA mutation status subgroups defined retrospectively, which were blind to clinical outcomes but were not prespecified in the Study 19 statistical analysis plan.
Tumour samples
The provision of an archival tumour sample (blocks or sections) was mandatory for participation in Study 19. Samples received as blocks were made into tumour microarrays with two 0.6 mm cores from each tumour at the University of British Columbia.
HRR mutation status
Tumour mutation status in BRCA1, BRCA2 and other key HRR-related genes was established using the same DNA sequencing analysis performed to determine tumour BRCA mutation status as previously described.7 In brief, DNA was extracted from formalin-fixed, paraffin-embedded (FFPE) archival tumour samples using a cancer gene panel enrichment procedure and deep resequencing performed with Illumina technology. Specifically, analysis was not performed with the commercially available Foundation Focus diagnostic test but with the Foundation Medicine T5 panel (entire coding sequence of 287 cancer-related genes plus select introns from 27 genes and other genetic alterations, deletions and functional rearrangements) at Foundation Medicine (Cambridge, MA, USA).16 Tumour analysis was performed on coded tumour samples and results were returned blind to the original Study 19 data set. The classification of variants was based on the American College of Medical Genetics recommendations. Patients with no known BRCA mutation and patients with a BRCA mutation classed as a variant of unknown significance (VUS) were included in the BRCA wild-type group as previously described.7
Patients in the BRCA wild-type group were further subdivided into three groups: BRCA wild-type HRR-mutated, patients whose tumours had a loss-of-function mutation in a high-confidence HRR gene; HRR status unknown, patients with a potential loss-of-function mutation in any gene associated with DNA repair; and BRCA and HRR wild type, patients with no potential loss-of-function mutations in any gene involved in DNA repair. The HRR genes interrogated are listed in Supplementary Table 1.
Myriad MyChoice® HRR deficiency test and BRCA1 promoter methylation analysis of tumour samples
Tumour samples were analysed as described by Timms et al. (2014).17 Next-generation sequencing-based assays were used to generate genome-wide single-nucleotide polymorphism profiles, BRCA1/2 mutation screening and BRCA1 promoter methylation data. These analyses separately reported tumour BRCA mutation (tBRCAm) status, tumour BRCA1 methylation status and tumour HRD score. An HRD score ≥42 was considered positive. The reported results are based on a research assay performed at Myriad Genetics and not upon the commercially available test.18
Immunohistochemistry for BRCA1
FFPE sections of 4 µm thickness were cut and mounted on SuperFrost Plus™ electrostatically charged glass slides and dried overnight at 37 °C in an incubator. Sections were dewaxed in xylene, passed through a graded alcohol series and optimal antigen retrieval obtained using Dako pH 9 antigen retrieval buffer (Dako, Copenhagen, Denmark), heated in the RHS Microwave Histo-processor (Milestone, Italy) to 110 °C with full pressure sustained for 2 min. The pressure vessel was then cooled under running tap water, the lid removed, and the slides washed.
Immunohistochemical staining was performed at room temperature on a Lab Vision 720 Autostainer. Sections were first treated with 3% H2O2 to quench endogenous peroxidase activity and then blocked with serum-free protein block (Dako). Following blocking, sections were incubated for 60 min with anti-BRCA1 (Ab-1) mouse monoclonal antibody (MS110) at a concentration of 1 µg/ml (OP92; MerckMillipore, Watford, UK). Visualisation was achieved using Dako REAL™ Envision™, HRP rabbit/mouse (Dako), applied for 30 min, followed by 10 min incubation in 3,3'-diaminobenzidine. Sections were lightly counterstained in Carazzi’s haematoxylin, dehydrated through alcohol grades and mounted under a glass coverslip with Histomount (RA Lamb; Thermo Fisher Scientific, MA, USA).
Staining was negatively controlled by substituting mouse immunoglobulin fraction, diluted to the same concentration as that for the primary antibody. Immunostained slides were evaluated with light microscopy.
Statistical analysis
The primary endpoint for Study 19 was PFS, which was reported for the full analysis population following a data cutoff (DCO) of 30 June 2010. PFS data were not collected following this DCO. Collection of time to first subsequent therapy or death (TFST), time to second subsequent therapy or death (TSST) and OS data continued, and the interim OS data reported here have a DCO of May 2016.
Analyses of PFS, TFST, TSST and OS use the same methodology; namely a Cox proportional hazards model adjusted for treatment, ethnic descent (Jewish vs non-Jewish), time to progression on penultimate platinum therapy (6–12 vs >12 months), and response to platinum therapy before randomisation (complete response vs partial response). This methodology was used for the primary analyses of the study.7,14
Where there were fewer than 15 events formal statistical tests were not performed.
Results
Classification of HRR gene mutation status
Patients with BRCA wild-type and HRR-mutated tumours or BRCA and HRR wild-type tumours
Archival tumour samples were available for 253/265 (96%) patients and tumour sequencing was completed for 209/265 (79%) patients. Forty-four samples failed due to insufficient tumour cells, DNA quantity, sequencing library quality or sequence read coverage. As previously described,7 111/209 patients were classified as having deleterious or suspected deleterious mutations present in BRCA1 and/or BRCA2 in their tumour samples and 98 patients had BRCA wild-type tumours; three further patients were known to have single exon indels in blood sample DNA but these were below the detection limit in tumour samples.7 Therefore, 95/209 patients (45.5%) were classified as having BRCA wild-type tumours. Twenty-one of these patients (22%) had at least one loss-of-function mutation in a candidate HRR gene, of whom 12 received olaparib and 9 received placebo. The specific HRR mutations identified in the 21 tumour samples were: BRIP1 (n = 5), CDK12 (n = 3), RAD54L (n = 2), RAD51B (n = 2), RAD54L rearr (n = 1), ATM rearr (n = 1), FANCA rearr (n = 1), FANCD2 (n = 1), FANCL rearr (n = 1), FANCL (n = 1), RAD51C (n = 1), RAD52 del (n = 1), XRCC3 rearr (n = 1) (Fig. 1).
For exploratory purposes, we defined a subset of patients as BRCA-mutated or HRR-mutated as a group containing a loss-of-function mutation in at least one pathway gene.
Of the remaining 74 patients, 16 were found to have mutations in genes that are involved more broadly in DNA damage repair (e.g., MSH2, MUTYH), or in genes where a postulated role in DNA repair has been the subject of significant controversy (e.g., EMSY); these patients were classified as HRR unknown. The remaining 58 patients’ tumours did not have a candidate loss-of-function mutation in a relevant gene. Twenty-five of these patients were randomised to olaparib and 33 to placebo.
PFS and OS in HRRm and HRRwt patients
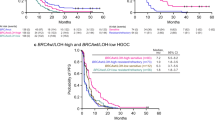
PFS was the primary endpoint of Study 19. Patients with a BRCA result from tumour testing at Foundation Medicine and BRCA wild-type subsets of Study 19 patients had comparable PFS hazard ratios to the overall BRCA-mutated (hazard ratio 0.16, 95% CI 0.08–0.3 vs 0.18, 0.10–0.31, respectively) and BRCA wild-type populations (hazard ratio 0.57, 0.34–0.94 vs 0.54, 0.34–0.85, respectively) (Fig. 2).7
PFS and OS by BRCA and HRR status determined at the time of Study 19 and in the exploratory subgroups defined by the Foundation Medicine analysis. *P < 0.05, **P < 0.01, ***P < 0.001. BRCAm BRCA mutation, BRCAwt BRCA wild type, CI confidence interval, HR hazard ratio, HRRm homologous recombination repair gene mutation, HRRwt homologous recombination repair genes wild type, n number of patients, OS overall survival, PFS progression-free survival, tBRCAm tumour BRCA mutation, tBRCAwt tumour BRCA wild type. Three patients who were classified as tBRCAwt following Foundation Medicine analyses were found to Have BRCAm following germline analysis
Data suggest that olaparib is associated with a greater PFS benefit in HRR-mutated patients without a BRCA mutation (hazard ratio 0.21, 0.04–0.86) than in patients with no detectable BRCA or HRR mutation (hazard ratio 0.71, 0.37–1.35) who received olaparib (Fig. 2). Furthermore, all study endpoints showed a consistent trend for a larger treatment effect in HRR-mutated patients and a less positive treatment effect in the patients with no detectable HRR mutations than in the overall BRCA wild-type population evaluable for HRR mutation analysis (Supplementary Table 2).
The PFS hazard ratio for the exploratory subgroup of patients with a mutation in a BRCA or other HRR gene (n = 157) was 0.20, 95% CI 0.12–0.33. It should be noted that PFS in the ITT population was the primary endpoint of the study and hence other analyses are exploratory subgroups of secondary or exploratory endpoints.
Classification of tumour BRCA mutation status, BRCA1 methylation and HRD score at Myriad
Myriad tumour testing, which comprised BRCA mutation testing, BRCA1 methylation testing and determining HRD score, was performed in 219 tumour samples.
Myriad tumour BRCA testing
Myriad tumour BRCA status was reported for 212 patients, of whom 118 (56%) had a BRCA mutation. In total, BRCA status was determined for tumours from 235 patients using either the Myriad and/or Foundation Medicine analyses. Of the 193 patients for whom tumour BRCA status was available at both laboratories, tumour samples from 106 patients were classified as mutated and 84 cases were classed as wt/VUS by both laboratories (VUS samples were predominantly missense variants: impact on gene function is unknown). The concordance of tumour BRCA testing between the two laboratories was 98.5% for sequence analysis (Table 1).
In the tumour samples from three patients, BRCA mutations were detected by Myriad testing but not by the Foundation Medicine test. Myriad testing classified the three BRCA mutations as a BRCA1 exon 24 deletion, a BRCA1 three-exon duplication and a BRCA2 32-bp deletion. The efficacy analyses for the Myriad BRCA-mutated and wild-type tumours were consistent with the previously published analysis (Fig. 3).7
PFS and OS by BRCA and HRR status determined at the time of Study 19 and in the exploratory subgroups defined by the Myriad MyChoice HRD score analysis. **P < 0.01, ***P < 0.001. BRCAm BRCA mutation, BRCAwt BRCA wild type, CI confidence interval, HR hazard ratio, HRD homologous recombination deficiency, ITT intention to treat, n number of patients, OS overall survival, PFS progression-free survival, tBRCAm tumour BRCA mutation, tBRCAwt tumour BRCA wild type
BRCA1 methylation
Eighteen of the 212 evaluable patients had BRCA1-methylated tumours, of which the tumours from 16 patients were BRCA wild type. Four of the 16 BRCA wild-type, BRCA1-methylated patients were treated with olaparib and 12 with placebo. At the DCO for PFS, 75% of patients in each arm had disease progression.
Myriad MyChoice homologous recombination deficiency score
Myriad MyChoice HRD scores were determined using an assay that calculates whole-genome tumour loss of heterozygosity, telomeric allelic imbalance and large-scale state transition scores.17 The HRD score is the sum of the three scores which have a bimodal distribution and a proposed threshold of ≥42.17 Analyses were performed blind to previous BRCA mutation data.
HRD scores were determined for 199 patients, of whom 139 (70%) were classified as HRD score high (HRD scores ≥42). Of the 139 tumours with high HRD scores, 101 patients (73%) had a BRCA mutation, 36 (26%) had no BRCA mutation and BRCA analysis failed in two patients (1%). Of the 60 patients (30%) with low HRD scores 8 patients (13%) had a BRCA mutation, 51 patients (85%) had no BRCA mutation and 1 (2%) had a failed analysis. Overall, 156 patients would be classified as Myriad HRD positive with a BRCA mutation in the tumour or a Myriad HRD score ≥42.
Relationships between HRD score, HRR gene mutations and BRCA1 methylation status
BRCA-mutated tumours had the highest median HRD score, HRR-mutated tumours had an intermediate median HRD score and tumours without mutations in BRCA or other HRR genes had the lowest median HRD score (Fig. 4).
Summary of Myriad MyChoice HRD scores. Patients must be evaluable for both Myriad HRD score and mutation/methylation status to be included. BRCAwt BRCA wild type, HRD homologous recombination deficiency, HRRm homologous recombination repair gene mutation, HRRwt homologous recombination repair genes wild type, n number of patients, tBRCAm tumour BRCA mutation
HRD scores and efficacy
Olaparib is currently approved for use in patients with platinum-sensitive relapsed ovarian cancer (PSROC), regardless of BRCA mutation status, in the US (tablet formulation) and in PSROC patients with a BRCA mutation in the EU (capsule formulation). We therefore performed exploratory analyses to examine separately the impact of HRD scores within patient groups with BRCA wild-type and BRCA-mutated tumours defined by Myriad tumour BRCA testing. There were only eight patients with BRCA mutations and HRD scores <42; this subgroup was not analysed further. When the prespecified threshold of ≥42 was used to define a BRCA wild-type HRD high group the PFS hazard ratio (hazard ratio 0.48; 95% CI 0.18–1.27; P = 0.14; n = 36; 20 olaparib, 16 placebo) was numerically superior to the PFS hazard ratio for the BRCA wild-type HRD low group (hazard ratio 0.60; 95% CI 0.31–1.17, P = 0.13; n = 51; 25 olaparib, 26 placebo) but was very similar to that for the overall BRCA wild-type population evaluable for HRD scores (hazard ratio 0.54; 95% CI 0.32–0.91; P = 0.02; n = 87) (Fig. 3). Further exploratory analyses did not show a trend for an improved hazard ratio for PFS with higher Myriad HRD scores in either the BRCA wild-type or mutated populations when the HRD cut-off was varied (data not shown). However, the relevant extremes of the HRD score distribution were only populated by small numbers of patients such that very few tumours with BRCA mutations had scores below 30 (n = 1) and conversely few tumours without BRCA mutations had scores greater than 50 (n = 30). Similar marginal differences in the hazard ratios observed for PFS between the BRCA wild-type HRD high group and the BRCA wild-type HRD low group were broadly observed for TFST, TSST and OS (Supplementary Table 3).
The PFS hazard ratio for the exploratory subgroup of patients with HRD-positive status (n = 156) was 0.24, 95% CI 0.15–0.39. It should be noted that PFS in the ITT population was the primary endpoint of the study and hence other analyses are exploratory subgroups of secondary or exploratory endpoints.
A recent study suggested patients with tumours with HRD scores <33 are less likely to benefit from platinum chemotherapy.19 In Study 19 this patient population, BRCAwt/HRD<33, had a worse PFS result (PFS hazard ratio 0.71, 95% CI 0.34–1.46, n = 41; 21 olaparib, 20 placebo) than the HRD-negative group (BRCAwt, HRD score <42) or the BRCA-mutated and/or HRD ≥33 group (PFS hazard ratio 0.25, 95% CI 0.16–0.38, n = 185, 97 olaparib, 88 placebo).
Markers associated with long-term treatment
In Study 19 there were 16 patients treated for 6 years or longer, of whom one was on placebo and had a germline mutation in BRCA1. Of the 15 patients on olaparib for more than 6 years, 8 were originally reported as having BRCA mutations.7 One further patient was found to have somatic mutations in both BRCA1 and BRCA2 on Myriad blood and tumour testing (no Foundation Medicine tumour testing data were available); hence, 9/15 patients are known to have a BRCA mutation (5 with BRCA2, 3 with BRCA1 and 1 with both) in their tumour or blood sample, with 3/9 being a somatic mutation. This enrichment for BRCA2 mutation and presence of somatic cases in long-term responders has been noted previously.20 One of the remaining six patients was not evaluable for tBRCA status at either Myriad or Foundation Medicine, hence 5/15 patients treated with olaparib for more than 6 years had no detectable BRCA mutation. Three out of five patients had Myriad HRD scores available (2x HRD positive and 1x HRD negative) and 5/5 had HRRm status available (1x HRRm, 2x HRR uncertain, 2x HRRwt). Of the two HRRwt tumours, one was Myriad HRD negative (Myriad HRD score 15) and the other was Myriad HRD status unknown. Therefore, patients without BRCA-mutated tumours and whose tumours tested negative for additional candidate biomarkers (HRRwt or BRCAwt HRD negative) were found among the patients who remained on olaparib for over 6 years.
Immunohistochemistry for BRCA1
Staining was performed on duplicate cores on the tissue microarrays generated from the 145 patients submitting a tumour block (Supplementary Table 4). A trend for less staining in BRCA1-mutated cases was observed (Supplementary Table 5). However, there was no indication of a prognostic or predictive effect on PFS of BRCA1 protein levels determined by immunohistochemical staining using Cox regression analysis.
Discussion
Elegant preclinical experiments led to the concept of synthetic lethality, which suggested that tumours without functional BRCA1 or BRCA2 protein would be more susceptible to PARP inhibitors compared with non-tumour tissues in the same patients.21 In this model, PARP inhibition leads to an accumulation of unrepaired single-strand breaks in DNA, which are converted to double-strand breaks when the tumour cell attempts DNA replication in the S-phase. The resulting double-strand breaks are then repaired in non-tumour cells that are proficient in HRR but accumulate and lead to mitotic catastrophe in tumour cells that are HRR deficient. BRCA1 and BRCA2 play important roles in homologous recombination and, in germline carriers of mutations, loss of the remaining un-mutated copy of the gene and concomitant loss of function is thought to occur early in the development of ovarian and breast cancer.22 Hence, early clinical studies with the PARP inhibitor olaparib focused on tumours in germline-mutated BRCA carriers and confirmed activity in patients with breast and ovarian cancer.23,24 Preclinically, deficiencies in other proteins involved in homologous recombination conferred sensitivity to PARP inhibition, albeit with a lesser impact than BRCA deficiency5 and high-grade SOC was initially thought to represent a particularly promising tumour type for PARP inhibitor therapy because, in addition to a high frequency of BRCA mutations, it is characterised by large-scale genomic instability and repeated, durable platinum sensitivity (considered likely to be hallmarks of HRR deficiency). In support of this theory, a non-randomised study of olaparib monotherapy in ovarian cancer subsequently found response rates of 41% and 24% in the germline BRCA-mutated and non-mutated carrier patient populations, respectively.25 Therefore, it was expected that the non-BRCA-mutated carrier high-grade SOC patient population included a significant number of patients whose tumours could be susceptible to PARP inhibition and Study 19 was a placebo-controlled study in the maintenance setting designed to test this hypothesis.7 Study 19 was a positive study overall and in a preplanned subgroup analysis the magnitude of PFS effect was greater in patients who had a BRCA mutation than those without.7 Tumour testing in Study 19 successfully identified the vast majority of germline BRCA-mutated cases and identified a further subset of patients with somatic or tumour only mutations.7 Subsequent exploratory analyses suggested that tumours with somatic BRCA mutations phenocopy those from germline carriers, both in terms of tumour genetics and in clinical benefit on olaparib10 and a clinical trial is under way to confirm this observation.26 Hence, BRCA mutation testing of tumour samples is the clearest indicator of tumour responsiveness to olaparib treatment in current clinical practice. Importantly, the data presented here confirm a very high level of consistency in research testing for tumour BRCA mutations in two independent laboratories and therefore confirms our previous observation that tumour testing accurately detects germline mutations. The number of mutations not captured by the Foundation Medicine assay was relatively small (n = 3, all indels). Nonetheless, it is important that all tumour testing laboratories further develop their methodologies to accurately detect rare variants to allow tumour testing to detect all clinically relevant mutations.
We have shown that Study 19 contained a readily identifiable, biologically coherent subpopulation of patients with inherited or acquired mutations in BRCA1 or BRCA2 who gain a PFS benefit from olaparib treatment. It is important that we determine whether there is a molecularly identifiable subgroup of patients with BRCA wild-type tumours that is differentiated from patients solely selected on available clinical parameters of platinum sensitivity and high-grade serous histology used in Study 19. Furthermore, if such a subgroup exists it will be important to determine whether the differential in benefit is clinically meaningful. Several candidate molecular measures have been suggested to fulfil this role including a continuous measure of genomic instability or HRR deficiency ‘scar’, BRCA1 methylation and loss-of-function mutations in other homologous recombination pathway genes. The supporting data behind the candidate biomarkers investigated are summarised in Table 2.
Here, we report that the ovarian BRCA wild-type patient population may include a majority of patients whose tumours lack DNA repair gene loss-of-function mutations and who gain less benefit from PARP olaparib treatment than patients with candidate loss-of-function mutations in genes involved in HRR.
First, we attempted to identify patients with loss-of-function mutations in candidate HRR genes in their tumours. Clearly this approach is limited by the genes sequenced by the Foundation Medicine panel and by the relative absence of relevant preclinical and clinical data. We therefore chose a pragmatic approach to include all genes with a DNA repair pathway role and some supportive preclinical data. Subsequently, we defined a higher confidence patient population by differentiating between candidate genes in which the primary role is related to mismatch repair or where, despite a purported role in double-strand DNA repair, either the genetic mechanism is distinct and/or the phenotypic impact on platinum sensitivity and prognosis in ovarian cancer is directionally opposed to that of BRCA mutations (i.e., EMSY amplification and PTEN loss).8
We identified 37 patients with mutations in any gene involved in DNA repair, of these patients 21 had mutations in the (relatively) higher confidence gene set. Genes involved in more than one patient were BRIP1 (n = 5), CDK12 (n = 3), RAD54L (n = 2) and RAD51B (n = 2). BRIP1 (BRCA1 interacting protein C-terminal helicase 1) is a protein that works with BRCA1 to repair damaged DNA and the inheritance of two mutated copies of BRIP1 causes Fanconi anaemia. Women who inherit one mutated copy of BRIP1 are likely to have an increased risk of ovarian cancer; however, the implication of this for breast cancer risk is inconclusive.27,28,29
CDK12 is the least validated gene included in our HRR gene list. It is biallelically inactivated in ovarian cancer; hypothesis-directed experiments have found that mutations or depletion are associated with sensitivity to radiation, platinum or PARP inhibitor treatment and it was also highlighted as a candidate gene in a hypothesis-free synthetic lethal screen approach.30,31 However, mechanistically it has been proposed to have an indirect effect on DNA repair via transcriptional regulation32 and it was recently suggested to be associated with a different pattern of genome instability than that seen in patients with BRCA mutations or BRCA1 methylation.33
The number of patients with mutations in any HRR gene (other than BRCA1 or BRCA2) in Study 19 is too small to derive definitive conclusions. Furthermore, because of the low prevalence of mutations in any one gene a similarly sized trial could easily result in a different profile of mutated genes. However, it was possible to define a subpopulation of 58 patients without a candidate loss-of-function mutation in any measured relevant HRR gene. While it is acknowledged that mutations or alterations in other unmeasured genes may not have been detected in these 58 patients, it is notable that for all efficacy measures there was marked attenuation of the treatment benefit from olaparib. Our data are consistent with those of another study,8 suggesting that tumours with mutations in HRR genes may be associated with increased platinum sensitivity and prolonged survival.
While these data are encouraging, significant challenges remain, due to the differing biological role of each gene and the low prevalence of mutations in any one gene. Hence, determining utility for clinical decision-making suffers from the analogous issues facing genetic teams with low prevalence/penetrance inherited gene mutations, although the risk–benefit profile for the treatment decisions in a cancer patient may be arguably different to that for prophylactic surgery. Further data are needed to confirm this observation and there is a need for the clinical, research and regulatory community to understand what evidence base can be required for individual or ‘buckets’ of low prevalence genes with high biological plausibility. Exploratory analyses of the recently reported ARIEL2, ARIEL3 and NOVA trials could significantly add to the present evidence base available.11,12,13
The ability to avoid these issues for causal genetic changes and use an HRD score or genomic read-out of the effects of HRD, regardless of causal mechanism, is inherently attractive in theory. In Study 19 there were only 36 patients with BRCA wild-type tumours with a high Myriad HRD score. In agreement with data presented from a preliminary analysis of the NOVA trial we note that HRR-mutated patients tend to have higher Myriad HRD scores in Study 19 but that BRIP mutations appear to have a weak correlation such that, in both Study 19 and the preliminary NOVA data, only half of BRIP-mutated tumours were HRD score positive. In Study 19 only half of all the HRD score evaluable HRR-mutated patients were HRD positive (scores ≥42), whereas 14 of the 15 patients with BRCA1 methylation who were evaluable for HRD score were HRD positive. BRCA1-methylated cases accounted for at least 39% of the BRCA wild-type HRD-positive population in Study 19. An exploratory analysis of the 36 BRCA wild-type Myriad HRD-positive cases suggested that these patients were only marginally more likely to benefit from olaparib than selection based on study entrance criteria (response to platinum and high-grade serous histology); it may be that this is largely because of an inability of BRCA1 methylation to phenocopy BRCA1 mutation in terms of olaparib sensitivity/ease of reversibility as has also been reported for platinum sensitivity.8 The BRCA wild-type HRD scores ≥42 analysis reported here is an exploratory analysis of a small number of patients (n = 36) and aligns with the recently reported data for the NOVA and ARIEL3 trials which used the Myriad HRD (42) score and Foundation Medicine LOH score, respectively.12,13 In further exploratory analysis of Study 19, we show that relaxing the HRD threshold to a value of 33 differentiates a potential population receiving enhanced benefit compared to patients below the new threshold (hazard ratio 0.71 in HRD score <33 vs hazard ratio = 0.25 in HRD score ≥33). Particularly in a setting where the majority of patients appear to derive some level of benefit from PARP inhibitors, considering a biomarker that aims to exclude those patients least likely to benefit, rather than identifying those that are most likely to benefit, might be an attractive strategy. Compared to the HRD threshold of 42, this modified HRD scar approach, using a threshold <33, may have the potential to better identify a patient population that would derive least benefit from PARP inhibitors, but requires further study.
Importantly, although we observed an enrichment for BRCA2 mutations in the long-term responders, neither an HRRm or HRD approach was able to exclude patients who remained on treatment for 6 years or more. It is likely that additional mechanisms, such as those relating to the immune system, are important in determining which patients may receive long-term benefit from olaparib and potentially also from chemotherapy.34,35
Conclusions
In summary, the analyses reported here demonstrate that next-generation sequence-based tumour testing for BRCA mutations in clinically available FFPE samples can be highly reproducible and may identify a subpopulation of patients with the greatest PFS benefit from olaparib. A measure of HRD or genomic instability (Myriad MyChoice HRD) identified most patients with BRCA mutations and nearly all patients with BRCA1 methylation; however, only ~50% of a small number of patients with mutations in other candidate HRR genes were identified as HRR deficient. In underpowered exploratory analyses (i) patients with BRCA wild-type HRD-positive tumours did not achieve as high a treatment benefit from olaparib as patients with BRCA-mutated tumours and (ii) patients with tumours without mutations in candidate HRR genes, which comprised the majority of the BRCA wild-type subpopulation, received the least benefit from olaparib treatment, an observation which may be particularly important in the maintenance setting in the absence of any acute indicator of tumour response to olaparib treatment.
Further clinical studies examining all candidate tumour DNA-based measures of cause and effect deficiencies in HRR pathways are required to confirm that these additional measurements can provide clinically relevant information and treatment benefits for patients with BRCA wild-type high-grade SOCs.
Availability of data and materials
AstraZeneca’s policy on clinical trial data, results and other information from or regarding AstraZeneca-sponsored clinical trials are described in full in the AstraZeneca website: https://astrazenecagrouptrials.pharmacm.com/ST/Submission/Search
References
Pommier, Y., O’Connor, M. J. & de Bono, J. Laying a trap to kill cancer cells: PARP inhibitors and their mechanisms of action. Sci. Transl. Med. 8, 362ps17 (2016).
Roy, R., Chun, J. & Powell, S. N. BRCA1 and BRCA2: different roles in a common pathway of genome protection. Nat. Rev. Cancer 12, 68–78 (2011).
Farmer, H. et al. Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 434, 917–921 (2005).
O’Connor, M. J. et al. Generating preclinical models to assess bone marrow toxicity induced by the PARP inhibitor olaparib in combination with chemotherapy. Cancer Res. 73, 4420 (2013).
McCabe, N. et al. Deficiency in the repair of DNA damage by homologous recombination and sensitivity to poly(ADP-ribose) polymerase inhibition. Cancer Res. 66, 8109–8115 (2006).
Patch, A. M. et al. Whole-genome characterization of chemoresistant ovarian cancer. Nature 521, 489–494 (2015).
Ledermann, J. et al. Olaparib maintenance therapy in patients with platinum-sensitive relapsed serous ovarian cancer: a preplanned retrospective analysis of outcomes by BRCA status in a randomised phase 2 trial. Lancet Oncol. 15, 852–861 (2014).
Pennington, K. P. et al. Germline and somatic mutations in homologous recombination genes predict platinum response and survival in ovarian, fallopian tube, and peritoneal carcinomas. Clin. Cancer Res. 20, 764–775 (2014).
Lord, C. J. & Ashworth, A. Targeted therapy for cancer using PARP inhibitors. Curr. Opin. Pharmacol. 8, 363–369 (2008).
Dougherty, B. et al. Exploratory analyses suggest ovarian tumors with somatic or germline loss of function mutations in BRCA1 or BRCA2 are biologically similar and sensitive to PARP inhibition. Annual Meeting of the American Association for Cancer Research, Philadelphia, PA, USA, 18–22 April, abst 611 (2015).
Swisher, E. M. et al. Rucaparib in relapsed, platinum-sensitive high-grade ovarian carcinoma (ARIEL2 Part 1): an international, multicentre, open-label, phase 2 trial. Lancet Oncol. 18, 75–87 (2017).
Mirza, M. R. et al. Niraparib maintenance therapy in platinum-sensitive, recurrent ovarian cancer. N. Engl. J. Med 375, 2154–2164 (2016).
Coleman, R. L. et al. Rucaparib maintenance treatment for recurrent ovarian carcinoma after response to platinum therapy (ARIEL3): a randomised, double-blind, placebo-controlled, phase 3 trial. Lancet 390, 1949–1961 (2017).
Ledermann, J. et al. Olaparib maintenance therapy in platinum-sensitive relapsed ovarian cancer. N. Engl. J. Med 366, 1382–1392 (2012).
Myriad Genetic Laboratories. BRACAnalysis® technical specifications. https://www.myriad.com/lib/technical-specifications/BRACAnalysis-Technical-Specifications.pdf (accessed 24 Dec. 2013) (2012)
Frampton, G. M. et al. Development and validation of a clinical cancer genomic profiling test based on massively parallel DNA sequencing. Nat. Biotechnol. 31, 1023–1031 (2013).
Timms, K. M. et al. Association of BRCA1/2 defects with genomic scores predictive of DNA damage repair deficiency among breast cancer subtypes. Breast Cancer Res 16, 475 (2014).
Myriad. Tumor BRACAnalysis CDxTM. http://myriadgenetics.eu/products/tumor-bracanalysis-cdx-2/?lang=gb (1 January 2017).
Stronach, E. A. et al. Biomarker assessment of HR deficiency, tumor BRCA1/2 mutations, and CCNE1 copy number in ovarian cancer: associations with clinical outcome following platinum monotherapy. Mol. Cancer Res. 16, 1103–1111 (2018).
Lheureux, S. et al. Long-term responders on olaparib maintenance in high-grade serous ovarian cancer: clinical and molecular characterization. Clin. Cancer Res. 23, 4086–4094 (2017).
Lord, C. J., Tutt, A. N. & Ashworth, A. Synthetic lethality and cancer therapy: lessons learned from the development of PARP inhibitors. Annu Rev. Med. 66, 455–470 (2015).
George, S. H. & Shaw, P. BRCA and early events in the development of serous ovarian cancer. Front. Oncol. 4, 5 (2014).
Tutt, A. et al. Oral poly(ADP-ribose) polymerase inhibitor olaparib in patients with BRCA1 or BRCA2 mutations and advanced breast cancer: a proof-of-concept trial. Lancet 376, 235–244 (2010).
Audeh, M. W. et al. Oral poly(ADP-ribose) polymerase inhibitor olaparib in patients with BRCA1 or BRCA2 mutations and recurrent ovarian cancer: a proof-of-concept trial. Lancet 376, 245–251 (2010).
Gelmon, K. A. et al. Olaparib in patients with recurrent high-grade serous or poorly differentiated ovarian carcinoma or triple-negative breast cancer: a phase 2, multicentre, open-label, non-randomised study. Lancet Oncol. 12, 852–861 (2011).
Pignata, S. et al. ORZORA: open-label phase IV trial of olaparib in patients with BRCA-mutated ovarian cancer. Int. J. Gynaecol. Cancer 25, 1469–1470 (2015).
Rafnar, T. et al. Mutations in BRIP1 confer high risk of ovarian cancer. Nat. Genet. 43, 1104–1107 (2011).
Ramus, S. J. et al. Germline mutations in the BRIP1, BARD1, PALB2, and NBN genes in women with ovarian cancer. J. Natl. Cancer Inst. 107, djv214 (2015).
Easton, D. F. et al. No evidence that protein truncating variants in BRIP1 are associated with breast cancer risk: implications for gene panel testing. J. Med. Genet. 53, 298–309 (2016).
Joshi, P. M., Sutor, S. L., Huntoon, C. J. & Karnitz, L. M. Ovarian cancer-associated mutations disable catalytic activity of CDK12, a kinase that promotes homologous recombination repair and resistance to cisplatin and poly(ADP-ribose) polymerase inhibitors. J. Biol. Chem. 289, 9247–9253 (2014).
Bajrami, I. et al. Genome-wide profiling of genetic synthetic lethality identifies CDK12 as a novel determinant of PARP1/2 inhibitor sensitivity. Cancer Res. 74, 287–297 (2014).
Blazek, D. et al. The Cyclin K/Cdk12 complex maintains genomic stability via regulation of expression of DNA damage response genes. Genes Dev. 25, 2158–2172 (2011).
Popova, T. et al. Ovarian cancers harboring inactivating mutations in CDK12 display a distinct genomic instability pattern characterized by large tandem duplications. Cancer Res. 76, 1882–1891 (2016).
Weberpals, J. I. et al. Tumor molecular profiling to differentiate extreme responses to first-line platinum-based chemotherapy in suboptimally debulked serous ovarian cancer patients. J. Clin. Oncol. 36, abstr 5561 (2018).
Goode, E. L. et al. Dose-response association of CD8+tumor-infiltrating lymphocytes and survival time in high-grade serous ovarian cancer. JAMA Oncol. 3, e173290 (2017).
AstraZeneca. Global Policy: Bioethics. https://www.astrazeneca.com/content/dam/az/PDF/Bioethics_policy.pdf (2016)
Esteller, M. et al. Promoter hypermethylation and BRCA1 inactivation in sporadic breast and ovarian tumors. J. Natl. Cancer Inst. 92, 564–569 (2000).
Ruscito, I. et al. BRCA1 gene promoter methylation status in high-grade serous ovarian cancer patients – a study of the tumour bank ovarian cancer (TOC) and ovarian cancer diagnosis consortium (OVCAD). Eur. J. Cancer 50, 2090–2098 (2104).
Lesnock, J. L. et al. BRCA1 expression and improved survival in ovarian cancer patients treated with intraperitoneal cisplatin and paclitaxel: a Gynecologic Oncology Group Study. Br. J. Cancer 108, 1231–1237 (2013).
Nomura, H. et al. Expression of potential biomarkers associated with homologous recombination repair in patients with ovarian or triple-negative breast cancer. Cancer Biomark. 16, 145–152 (2016).
Milner, R. et al. Validation of the BRCA1 antibody MS110 and the utility of BRCA1 as a patient selection biomarker in immunohistochemical analysis of breast and ovarian tumours. Virchows Arch. 462, 269–279 (2013).
Abkevich, V. et al. Patterns of genomic loss of heterozygosity predict homologous recombination repair defects in epithelial ovarian cancer. Br. J. Cancer 107, 1776–1782 (2012).
Watkins, J. A. et al. Genomic scars as biomarkers of homologous recombination deficiency and drug response in breast and ovarian cancers. Breast Cancer Res. 16, 211 (2014).
Acknowledgements
This study was sponsored by AstraZeneca. We thank Martin Goulding from Mudskipper Business Ltd who provided medical writing assistance funded by AstraZeneca.
Authors’ contribution
D.R.H., B.A.D., Z.L., A.F., M.J.O.C., T.W.H., J.D.R, M.O., H.J., B.G., C.W., C.G. and J.L. contributed to the study design, conduct and collection of data. L.G. and S.S. analysed the data and provided statistical support. J.S.L., K.M.T. and A.G. performed genetic analyses. J.L. was the principal trial investigator. D.H. prepared the first draft of the manuscript. All authors made substantial contributions to interpretation of data, were involved in drafting the manuscript and/or revising it critically for important intellectual content and approved the final version for submission.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethics approval and consent to participate
The institutional review boards or independent ethics committees of all investigational sites approved the protocol and all patients provided written, informed consent. The study was conducted in accordance with the Declaration of Helsinki, Good Clinical Practice and the AstraZeneca policy on bioethics.36
Competing interests
D.R.H., B.A.D., Z.L., A.F., L.G., S.S., M.J.O.C., T.W.H., M.O., H.J. and J.C.B. are AstraZeneca employees and stockholders; J.D.R. and C.W. were AstraZeneca employees and stockholders at the time the study was conducted. J.S.L., K.M.T. and A.G. are Myriad Genetics Inc. employees and stockholders; C.G. has acted in an advisory role for AstraZeneca, Clovis, Roche and Nucana, received honoraria from AstraZeneca and Roche, and received research funding from AstraZeneca, Novartis and Aprea; J.L. has participated in an advisory board for AstraZeneca; B.G. declares no competing interests.
Funding
This study was funded by AstraZeneca.
Additional information
Note: This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution 4.0 International (CC BY 4.0).
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hodgson, D.R., Dougherty, B.A., Lai, Z. et al. Candidate biomarkers of PARP inhibitor sensitivity in ovarian cancer beyond the BRCA genes. Br J Cancer 119, 1401–1409 (2018). https://doi.org/10.1038/s41416-018-0274-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-018-0274-8
This article is cited by
-
Prognostic and Potential Therapeutic Roles of PRKDC Expression in Lung Cancer
Molecular Biotechnology (2024)
-
Heterogeneity and treatment landscape of ovarian carcinoma
Nature Reviews Clinical Oncology (2023)
-
A new wave of innovations within the DNA damage response
Signal Transduction and Targeted Therapy (2023)
-
A signal-seeking Phase 2 study of olaparib and durvalumab in advanced solid cancers with homologous recombination repair gene alterations
British Journal of Cancer (2023)
-
Performance of a RAD51-based functional HRD test on paraffin-embedded breast cancer tissue
Breast Cancer Research and Treatment (2023)