Abstract
Background
Circulating cell-free DNA (cfDNA) may help understand the molecular response to pharmacologic treatment and provide information on dynamics of clonal heterogeneity. Therefore, this study evaluated the correlation between treatment outcome and activating EGFR mutations (act-EGFR) and T790M in cfDNA in patients with advanced NSCLC given osimertinib.
Methods
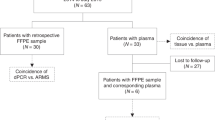
Thirty-four NSCLC patients resistant to first/second-generation EGFR-TKIs, positive for both act-EGFR and T790M in cfDNA at the time of progression were enrolled in this study. Plasma samples were obtained at osimertinib baseline and after 3 months of therapy; cfDNA was analyzed by droplet digital PCR and results were expressed as mutant allele frequency (MAF).
Results
At baseline, act-EGFR MAF was significantly higher than T790M (p < 0.0001). act-EGFR MAF and T790M/act-EGFR MAF ratio were significantly correlated with disease response (p = 0.02). Cut-off values of act-EGFR MAF and T790M/act-EGFR ratio of 2.6% and 0.22 were found, respectively. The PFS of patients with act-EGFR MAF of > 2.6% and < 2.6%, were 10 months vs. not reached, respectively (p = 0.03), whereas patients with T790M/act-EGFR ≤ 0.22 had poorer PFS than patients with a value of > 0.22 (6 months vs. not reached, respectively, p = 0.01).
Conclusion
act-EGFR MAF and T790M/act-EGFR MAF ratio are potential markers of outcome in patients treated with osimertinib.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 24 print issues and online access
$259.00 per year
only $10.79 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Gazdar, A. F. Activating and resistance mutations of EGFR in non-small-cell lung cancer: role in clinical response to EGFR tyrosine kinase inhibitors. Oncogene 28(Suppl 1), S24–S31 (2009).
Dixit, A. & Verkhivker, G. M. Hierarchical modeling of activation mechanisms in the ABL and EGFR kinase domains: thermodynamic and mechanistic catalysts of kinase activation by cancer mutations. PLoS Comput. Biol. 5, e1000487 (2009).
Lin, Y., Wang, X. & Jin, H. EGFR-TKI resistance in NSCLC patients: mechanisms and strategies. Am. J. Cancer Res. 4, 411–435 (2014).
Yun, C. H. et al. The T790M mutation in EGFR kinase causes drug resistance by increasing the affinity for ATP. Proc. Natl Acad. Sci. USA 105, 2070–2075 (2008).
Janne, P. A. et al. AZD9291 in EGFR inhibitor-resistant non-small-cell lung cancer. N. Engl. J. Med. 372, 1689–1699 (2015).
Mok, T. S. et al. Osimertinib or platinum-pemetrexed in EGFR T790M-positive lung cancer. N. Engl. J. Med. 376, 629–640 (2017).
Oxnard, G. R. et al. Association between plasma genotyping and outcomes of treatment with osimertinib (AZD9291) in advanced non-small-cell lung cancer. J. Clin. Oncol. 34, 3375–3382 (2016).
Ma, C., Wei, S. & Song, Y. T790M and acquired resistance of EGFR TKI: a literature review of clinical reports. J. Thorac. Dis. 3, 10–18 (2011).
Yu, H. A., Arcila, M. E., Hellmann, M. D., Kris, M. G., Ladanyi, M. & Riely, G. J. Poor response to erlotinib in patients with tumours containing baseline EGFR T790M mutations found by routine clinical molecular testing. Ann. Oncol. 25, 423–428 (2014).
Tan, D. S. et al. The International Association for the Study of Lung Cancer Consensus statement on optimizing management of EGFR mutation-positive non-small cell lung cancer: status in 2016. J. Thorac. Oncol. 11, 946–963 (2016).
Jovelet, C. et al. Circulating cell-free tumour DNA analysis of 50 genes by next-generation sequencing in the prospective MOSCATO trial. Clin. Cancer Res. 22, 2960–2968 (2016).
Del, Re. M. et al. Patients with NSCLC may display a low ratio of p.T790M vs. activating EGFR mutations in plasma at disease progression: implications for personalised treatment. Oncotarget 8, 86056–86065 (2017).
Del, Re. M. et al. Contribution of KRAS mutations and c.2369C>T (p.T790M) EGFR to acquired resistance to EGFR-TKIs in EGFR mutant NSCLC: a study on circulating tumour DNA. Oncotarget 8, 13611–13619 (2017).
Seki, Y. et al. Picoliter-droplet digital polymerase chain reaction-based analysis of cell-free plasma DNA to assess EGFR mutations in lung adenocarcinoma that confer resistance to tyrosine-kinase inhibitors. Oncologist 21, 156–164 (2016).
Yi, X., Ma, J., Guan, Y., Chen, R., Yang, L. & Xia, X. The feasibility of using mutation detection in ctDNA to assess tumour dynamics. Int. J. Cancer 140, 2642–2647 (2017).
Yung, T. K., Chan, K. C., Mok, T. S., Tong, J., To, K. F. & Lo, Y. M. Single-molecule detection of epidermal growth factor receptor mutations in plasma by microfluidics digital PCR in non-small cell lung cancer patients. Clin. Cancer Res. 15, 2076–2084 (2009).
Soria, J. C. et al. Osimertinib in untreated EGFR-mutated advanced non-small-cell lung cancer. N. Engl. J. Med. 378, 113–125 (2018).
Remon, J. et al. Osimertinib benefit in EGFR-mutant NSCLC patients with T790M-mutation detected by circulating tumour DNA. Ann. Oncol. 28, 784–790 (2017).
Chabon, J. J. et al. Circulating tumour DNA profiling reveals heterogeneity of EGFR inhibitor resistance mechanisms in lung cancer patients. Nat. Commun. 7, 11815 (2016).
Piotrowska, Z. et al. Heterogeneity underlies the emergence of EGFRT790 wild-type clones following treatment of T790M-positive cancers with a third-generation EGFR inhibitor. Cancer Discov. 5, 713–722 (2015).
Thress, K. S. et al. Complete clearance of plasma EGFR mutations as a predictor of outcome on osimertinib in the AURA trial. J. Clin. Oncol. 35(15_suppl), 9018–9018 (2017).
Chic, N., Mayo-de-Las-Casas, C. & Reguart, N. Successful treatment with gefitinib in advanced non-small cell lung cancer after acquired resistance to osimertinib. J. Thorac. Oncol. 12, e78–e80 (2017).
Knebel, F. H. et al. Sequential liquid biopsies reveal dynamic alterations of EGFR driver mutations and indicate EGFR amplification as a new mechanism of resistance to osimertinib in NSCLC. Lung Cancer 108, 238–241 (2017).
Ou, S. I. et al. Emergence of novel and dominant acquired EGFR solvent-front mutations at Gly796 (G796S/R) together with C797S/R and L792F/H mutations in one EGFR (L858R/T790M) NSCLC patient who progressed on osimertinib. Lung Cancer 108, 228–231 (2017).
Ou, S. I., Agarwal, N. & Ali, S. M. High MET amplification level as a resistance mechanism to osimertinib (AZD9291) in a patient that symptomatically responded to crizotinib treatment post-osimertinib progression. Lung Cancer 98, 59–61 (2016).
Minari, R. et al. Primary resistance to osimertinib due to SCLC transformation: Issue of T790M determination on liquid re-biopsy. Lung Cancer 115, 21–27 (2018).
Acknowledgements
This work was supported in part by research funding granted to Romano Danesi from Fondazione Cassa di Risparmio di Lucca (Lucca, Italy).
Availability of data and materials
Data and results are available at the Unit of Clinical Pharmacology and Pharmacogenetics, University Hospital of Pisa.
Author information
Authors and Affiliations
Contributions
Conception and design: M.D.R., M.T., I.P., and R.D. Development of methodology: M.D.R., E.R., G.R., S.C., and E.A. Clinical protocols/amendments: M.D.R., R.D., and M.T. Acquisition of data: M.D.R., P.B., E.R., G.R., S.C., E.A., and R.M. Analysis and interpretation of data: M.D.R., P.B., E.R., I.P., M.T., R.M., and R.D. Writing, review, and/or revision of the manuscript: all authors. Administrative, technical, or material support: M.D.R., P.B., E.R., and R.D. Study supervision: R.D., M.T., and I.P.
Corresponding author
Ethics declarations
Competing interests
T.M. received honoraria for participation in advisory boards and speakers’ bureau of Astra-Zeneca. R.D. received a unrestricted research grant from Astra-Zeneca. The other Authors declare no competing interests.
Ethics statement
This work was approved by the Ethics Committee of the Area Vasta Nord Ovest Toscana (Italy) under the reference number 612/2015 and was conducted in accordance with the principles of the Declaration of Helsinki. Patients were instructed about the experimental procedures of the study and enrolled after signature of the informed consent.
Consent for publication
No individual person’s data are reported.
Note
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution 4.0 International (CC BY 4.0).
Rights and permissions
About this article
Cite this article
Del Re, M., Bordi, P., Rofi, E. et al. The amount of activating EGFR mutations in circulating cell-free DNA is a marker to monitor osimertinib response. Br J Cancer 119, 1252–1258 (2018). https://doi.org/10.1038/s41416-018-0238-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-018-0238-z
This article is cited by
-
IDH1 mutation is detectable in plasma cell-free DNA and is associated with survival outcome in glioma patients
BMC Cancer (2024)
-
Predictive significance of circulating tumor DNA against patients with T790M-positive EGFR-mutant NSCLC receiving osimertinib
Scientific Reports (2023)
-
Liquid biopsy and non-small cell lung cancer: are we looking at the tip of the iceberg?
British Journal of Cancer (2022)
-
A comprehensive prognostic analysis of osimertinib treatment in advanced non-small cell lung cancer patients with acquired EGFR-T790M mutation: a real-world study
Journal of Cancer Research and Clinical Oncology (2022)
-
Current and Emerging Applications of Droplet Digital PCR in Oncology: An Updated Review
Molecular Diagnosis & Therapy (2022)