Abstract
Osteoporosis caused by aging is characterized by reduced bone mass and accumulated adipocytes in the bone marrow cavity. How the balance between osteoblastogenesis and adipogenesis from bone marrow mesenchymal stem cells (BMSCs) is lost upon aging is still unclear. Here, we found that the RNA-binding protein Musashi2 (Msi2) regulates BMSC lineage commitment. Msi2 is commonly enriched in stem cells and tumor cells. We found that its expression was downregulated during adipogenic differentiation and upregulated during osteogenic differentiation of BMSCs. Msi2 knockout mice exhibited decreased bone mass with substantial accumulation of marrow adipocytes, similar to aging-induced osteoporosis. Depletion of Msi2 in BMSCs led to increased adipocyte commitment. Transcriptional profiling analysis revealed that Msi2 deficiency led to increased PPARγ signaling. RNA-interacting protein immunoprecipitation assays demonstrated that Msi2 could inhibit the translation of the key adipogenic factor Cebpα, thereby inhibiting PPAR signaling. Furthermore, the expression of Msi2 decreased significantly during the aging process of mice, indicating that decreased Msi2 function during aging contributes to abnormal accumulation of adipocytes in bone marrow and osteoporosis. Thus, our results provide a putative biochemical mechanism for aging-related osteoporosis, suggesting that modulating Msi2 function may benefit the treatment of bone aging.
Similar content being viewed by others
Introduction
Aging-induced osteoporosis is characterized by reduced bone formation and the accumulation of adipocytes in the bone marrow chamber.1,2 Both osteoblasts and adipocytes are differentiated from the same multipotent precursor bone marrow mesenchymal stem cells (BMSCs).3,4,5 Compared with young MSCs, MSCs in old organisms show enhanced senescence, have reduced self-renewal and mainly differentiate into adipocytes instead of osteoblasts.6 The dynamic balance of MSC differentiation between adipogenesis and osteoblastogenesis is controlled by the expression of key transcription factors, including PPARγ, C/EBPα, and RUNX2.7,8 The adipose tissue of mice lacking C/EBPα is underdeveloped, and endogenous PPARγ cannot be induced.9 The transcription factor PPARγ plays a crucial role in bone development by inducing adipogenesis and inhibiting osteoblastogenesis.10,11 In addition to transcriptional regulation, posttranscriptional mechanisms also play important roles in regulating cell fate determination.12
RNA-binding proteins have been shown to regulate multiple steps of post-translational regulatory processes, such as RNA stability, RNA polyadenylation and translation, and determine cell fate.13 Whether RNA-binding proteins can regulate the commitment of MSCs has not been fully explored. Furthermore, the relationship between RNA-binding proteins and master transcription factors has not been fully elucidated. Revealing the functions of more RNA-binding proteins will help us further understand the orchestrated regulation of cell fate determination.
The Musashi (Msi) family of RNA-binding proteins contains two members, namely, Msi1 and Msi2, in mammals.13 Msi1 and Msi2 are evolutionarily conserved, containing two tandem RNA recognition motifs and a carboxyl terminal poly-A–binding protein association domain.13,14,15 Msi proteins bind to r(G/A)U1–3AGU sequences (MSI binding elements, MBEs) at the 3′ untranslated region (UTR) of the target mRNA to prevent poly-A binding protein from entering the extension initiation complex to repress translation.14,16 Several studies have reported that MSI proteins act as translation repressors.17,18,19 MSI proteins contribute to the control of symmetric and asymmetric stem cell division, regulate stem cell function, and play a role in cell fate determination.15,20 Msi1 is mainly involved in regulating the self-renewal of neuronal stem cells, and Msi2 is mainly expressed in hematopoietic stem cells and regulates hematopoietic function.15,21 Msi2 plays an important role in hair regeneration, maintaining the resting state of hair follicle stem cells, translation of cancer stem cells, and self-renewal and differentiation of hematopoietic stem cells.22,23,24 Msi2 plays a critical role in the differentiation of osteoclasts in vitro, which are derived from HSCs. Loss of Msi2 inhibits Notch signaling during osteoclast differentiation and induces apoptosis in preosteoclasts.25 Studies have found that Hh signaling can negatively regulate osteogenic differentiation by inhibiting RNA binding to Msi1.26 The Msi family plays a role in a variety of stem cells. Whether Msi2 can regulate mesenchymal stem cells and whether it can regulate the fate determination of MSCs have not yet been reported. Whether Msi2 has a regulatory effect on bone homeostasis and bone aging in vivo and the downstream molecular mechanism of the regulation are still unclear. The link between the RNA-binding protein Msi2 and osteoporosis is worth exploring in depth.
Here, we revealed the functions of Msi2 in balancing the osteoblast/adipocyte lineage commitment of BMSCs and aging-induced osteoporosis. Msi2 knockout mice displayed accumulation of adipocytes in the bone marrow cavity and decreased bone mass, mimicking osteoporosis. Msi2 promotes the differentiation of BMSCs into osteoblasts and inhibits the differentiation of BMSCs into adipocytes. Msi2 specifically binds the 3′UTR of mRNA of the key adipogenesis-related transcription factor Cebpα to inhibit its translation, thereby inhibiting PPARγ signaling. Furthermore, we found that Msi2 expression was decreased in aged BMSCs, indicating that the decreased Msi2 expression during aging shifts the osteogenesis/adipogenesis balance toward adipogenesis and leads to osteoporosis. Overall, these results suggested that increasing Msi2 function may benefit the treatment of aging-related bone loss.
Results
The Msi2 expression level decreases during adipogenesis and increases during osteogenesis of BMSCs
BMSCs were able to differentiate into both osteoblasts and adipocytes. To explore the functions of Msi2 in BMSCs, we first surveyed the protein level of Msi2 during BMSC differentiation. When BMSCs were induced to differentiate into adipocytes that were stained with Oil Red O and BODIPY (Fig. 1a), both the mRNA and protein levels of Msi2 decreased, while the expression levels of adipocyte markers such as CCAAT/enhancer binding protein α (Cebpα), peroxisome proliferative activated receptor γ (Pparγ), fatty acid binding protein 4 (Fabp4) and perilipin increased, indicating efficient differentiation into adipocytes (Fig. 1b–d). In contrast, when BMSCs were induced to differentiate into osteoblasts that were stained with ALP and Alizarin red S (Fig. 1e), both the mRNA and protein levels of Msi2 increased during the differentiation process. Consistently, the expression levels of osteoblast markers, including Runt-related transcription factor 2 (Runx2), Sp7 transcription factor (Osterix), and Collagen type 1 alpha 1 (Col1α1), increased, suggesting efficient differentiation into osteoblasts (Fig. 1f–h). The dynamic changes in Msi2 expression levels during BMSC adipogenesis and osteogenesis indicate that Msi2 may play distinct roles in adipogenesis and osteoblastogenesis from BMSCs.
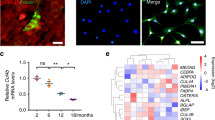
Changes in Msi2 expression levels in the adipogenesis and osteogenesis of BMSCs. a BMSCs isolated from 4-week-old wild-type mice. Cultures were stained with Oil Red O and BODIPY as shown. Scale bar = 200 μm. b Western blot analysis of Msi2 levels during adipogenesis for different durations. c qPCR analysis of Msi2 expression in BMSCs during adipogenesis for the indicated durations. Data represent the mean ± SD, n = 4. d qPCR analysis of the expression of adipocyte markers, including perilipin, Fabp4, Pparγ and Cebpα, in BMSCs during adipogenesis for the indicated durations. Data represent the mean ± SD, n = 4. e BMSCs isolated from 6-week-old wild-type mice. Cultures were stained with ALP and Alizarin red S as shown. Scale bar = 3 mm. f Western blot analysis of Msi2 levels during osteogenesis for different durations. g qPCR analysis of Msi2 expression in BMSCs during osteogenesis for the indicated durations. Data represent the mean ± SD, n = 4. h qPCR analysis of the expression of osteoblast markers, including Runx2, Col1a1 and Osterix, in BMSCs during osteogenesis for the indicated durations. Data represent the mean ± SD, n = 4. i Immunostaining of Msi2 (green), CD105 (red) and DAPI (blue) in tibia from 6-week-old WT mice. Scale bar = 200 μm (left). Scale bar =30 μm (right). j Immunostaining of Msi2 (green) and DAPI (blue) in femurs from 6-week-old Prx1-Cre Tdtomato mice. Scale bar = 1 000 μm (left). Scale bar = 200 μm (right)
We next determined the MSI2 expression level in long bone in vivo and found that MSI2 was highly expressed in the growth plate and trabecular bone (Fig. S1A). Interestingly, MSI2 was also expressed in the internal and external periostea but was barely expressed in cortical bone (Fig. S1B). CD105 is a marker of MSCs. Further research found that MSI2 and CD105 can be colocalized (Fig. 1i). Further research was performed to determine whether MSI2 is expressed in Prx1-positive cells, which are mainly MSCs. We found that MSI2 expression colocalized with Prx1-positive cells (Fig. 1j). These results further suggested that Msi2 may have functions in MSC commitment and bone formation.
Msi2-deficient mice display increased bone marrow adipocytes and decreased bone mass
To investigate the function of Msi2 in BMSC differentiation, we generated Msi2 knockout mice using CRISPR-Cas9 technology to introduce a frameshift in the first intron of Msi2 (Fig. 2a). Immunofluorescence staining and western blotting confirmed the knockout of Msi2 in bone and BMSCs (Fig. 2b and Fig. 6j). We tested the knockout efficiency of Msi2 in the main organs of the knockout mice. The results showed that Msi2 was almost completely eliminated in the Msi2 knockout mice (Fig. S2A, B). In addition, we tested whether Msi1, a homolog of Msi2, has a compensatory effect in knockout mice, and the results showed that Msi1 expression in the BMSCs of knockout mice was not significantly different from that in the control mice. (Fig. S2C). The Msi2−/− mice survived normally after birth and had normal fertility. However, the Msi2−/− mice exhibited short stature and skeletal dysplasia regardless of sex (Fig. 2c and Fig. S2D). Compared with the control mice, the Msi2 knockout mice had reduced body weight, body length, and femur length (Fig. S2E-G). BODIPY staining results of the Msi2−/− mouse tibia revealed increased adipocyte accumulation in the tibia bone marrow of the Msi2−/− mice, and older Msi2 knockout mice had more fat vacuoles in the bone marrow cavity (Fig. 2d). Immunofluorescence staining of perilipin A, a mature adipocyte marker, also confirmed adipocyte accumulation in the Msi2−/− mice (Fig. 2e, f). Both the number and the size of adipocytes in the bone marrow cavity increased with age in the Msi2−/− mice (Fig. 2g, h).
Msi2-deficient mice show increased bone marrow adipocytes. a Mouse construction strategy. b Immunostaining of MSI2 (green) and DAPI (blue) in tibiae from 6-week-old WT and Msi2−/− mice. Scale bar = 200 μm. c Representative view of the wild-type, Msi2+/−, and Msi2−/− 6-week-old mice. d BODIPY (green) staining of tibiae from the 6-week-old WT and Msi2−/− mice. Scale bar = 500 μm (top). Hematoxylin-eosin staining of femurs from the 30-week-old WT and Msi2−/− mice. Scale bar = 500 μm (middle). Scale bar = 500 μm (down). e Immunostaining of perilipin A/B (green) and OPN (red) of femurs from the 6-week-old WT and Msi2−/− mice. Scale bar = 50 μm. f Quantification of the relative areas of OPN and perilipin in (e). g Immunostaining of perilipin A/B (green) and OPN (red) of femurs from the 15-week-old WT and Msi2−/− mice. Scale bar = 50 μm. h Quantification of the relative areas of OPN and perilipin in (g)
We further investigated whether bone formation was affected. Microquantitative computed tomography (μ-CT) analysis was performed to compare the changes in bone-related elements in the long bones of the Msi2 knockout mice and the WT littermates. We found that the 6-week-old Msi2−/− mice showed significantly decreased bone mass (Fig. 3a). Trabecular bone per tissue volume (BV/TV) in the Msi2−/− mice was decreased compared to that in the age-matched WT littermates (Fig. 3c), accompanied by a reduction in trabecular number (Tb.N) (Fig. 3d), a reduction in trabecular bone thickness (Tb.Th) and an increase in trabecular bone spacing (Tb.Sp) (Fig. 3e, f). There was no significant difference in cortical bone thickness (Cor.Th) of the Msi2−/− mice compared with that of the WT mice, which is consistent with the observation that Msi2 is rarely expressed on cortical bone (Figs. 1j, 3b, g).
Msi2-deficient mice show decreased bone mass. a Three-dimensional μ-CT images of trabecular bone of distal femurs isolated from the 6-week-old female WT and Msi2−/− mice (n = 6). b Three-dimensional μ-CT images of cortical bone of distal femurs isolated from the 6-week-old female WT and Msi2−/−mice (n = 6). c–g μ-CT analysis of distal femurs from the 6-week-old WT and Msi2−/− mice for trabecular bone volume per tissue volume (BV/TV) (c), trabecular number (Tb.N) (d), trabecular separation (Tb.Sp) (e), trabecular thickness (Tb.Th) (f) and cortical bone thickness (Cor.Th) (g). h Masson trichrome staining of the 6-week-old WT and Msi2−/− mice. Scale bar = 500 μm. i–n Histomorphometric analysis of distal femurs from the 5-week-old WT and Msi2−/− mice to determine the trabecular bone volume per tissue volume (BV/TV) (i), trabecular number (Tb.N) (j), trabecular thickness (Tb.Th) (k) trabecular separation (Tb.Sp) (l) and number of osteoblasts per bone perimeter (N.Ob/B.Pm) (m) and osteoblast surface per bone surface (Ob.S/BS) (n). Data represent the mean ± SD, n = 4. *P < 0.05, **P < 0.01, ns indicates no significance, unpaired Student’s t test. o Immunostaining of OPN (red) and DAPI (blue) in femurs from the 6-week-old WT and Msi2−/− mice. Scale bar = 500 μm. p Quantification of the relative area of OPN in (o). q Immunohistochemical staining of Col1α1 from the 6-week-old WT and Msi2−/− mice. Scale bar = 100 μm. r Quantification of the relative area of Col1α1 in (q)
To further explore the function of Msi2 in bone formation, we performed histomorphometric analysis to evaluate static and dynamic parameters of bone formation and resorption (Fig. 3h). Consistent with the μ-CT data, histomorphometric analysis also showed that the Msi2−/− mice had a significant decrease in both BV/TV and Tb.Th and also showed a significant increase in Tb.Sp but no changes in Tb.N (Fig. 3i–l). The numbers of osteoblasts per bone perimeter (N.Ob/B.Pm) and osteoblast surface per bone surface (Ob.S/BS) were decreased in the Msi2−/−mice compared to the WT control mice (Fig. 3m, n). Further immunofluorescence staining analysis of the distal femur of the Msi2−/− mice revealed decreased expression of the osteoblast markers osteopontin (OPN) and COL1α1 in the Msi2−/− mice (Fig. 3o–r).
Bone formation by osteoblasts and bone resorption by osteoclasts are essential for the maintenance of bone homeostasis. Our results showed that the osteoclast differentiation of the Msi2 knockout mice was weakened in vitro (Fig. S3A, B). Interestingly, no changes in the number of HSCs were detected in the bone marrow cells of the Msi2−/− mice (Fig. S3C). TRAP staining for osteoclast activity showed no significant difference between the WT and Msi2−/− mice in vivo (Fig. S3D, E). This finding indicates that the decrease in bone mass in the Msi2−/− mice is mainly due to decreased bone formation. Taken together, the above results suggested that Msi2 is required for proper bone formation.
Msi2 promotes osteoblastogenesis and inhibits BMSC adipogenesis
The accumulation of adipocytes and decreased bone formation in the bone of the Msi2−/− mice prompted us to further explore how Msi2 regulates BMSC differentiation. BMSCs were isolated from the WT or Msi2−/− mice and were differentiated in adipogenic medium for 7 days. Adipogenic differentiation was enhanced in the Msi2−/− BMSCs, as indicated by increased Oil Red O staining and BODIPY staining compared to that of the BMSCs from the WT mice (Fig. 4a, b). The expression levels of adipocyte markers such as Cebpα, Cebpβ, Fabp4, lipoprotein lipase (Lpl), perilipin and Pparγ increased in the Msi2 knockout BMSCs upon induction to adipogenesis compared to those of the WT BMSCs (Fig. 4c).
Msi2 deficiency promotes adipogenesis and inhibits osteoblastogenesis in BMSCs. a Oil Red O and BODIPY staining of BMSCs cultured with adipocyte differentiation medium for 6 days. Data are representative of three independent experiments. Scale bar = 40 μm. b Statistical analysis of the percentage of Oil Red O-positive area via ImageJ. Data are presented as the mean ± SD, n = 4 in each group. Data represent the mean ± SD, ***P < 0.005, unpaired Student’s t test. c qPCR analysis of Cebpα, Cebpβ, Fabp4, Lpl, perilipin and Pparγ expression in BMSCs from the WT and Msi2−/− mice after adipocyte differentiation for 6 days. Data represent the mean ± SD, n = 4. *P < 0.05, **P < 0.01, ***P < 0.005, unpaired Student’s t test. d ALP staining and Alizarin red S staining after osteoblast differentiation for 7 days (upper) and 14 days (lower). Data are representative of three independent experiments. Scale bar = 3 mm. e ALP activity was measured by phosphatase substrate assays. Data represent the mean ± SD, n = 3. **P < 0.01, unpaired Student’s t test. f qPCR analysis of Runx2, Alp, Bsp, Col1α1, Osterix, and ATF4 expression after osteoblast differentiation for 7 days; BMSCs were from the WT and Msi2−/− mice. Data represent the mean ± SD, n = 4. *P < 0.05, **P < 0.01, ns: no significance; unpaired Student’s t test
We next examined the role of Msi2 in the osteoblast differentiation of BMSCs. BMSCs were isolated from the WT or Msi2−/− mice and were induced to differentiate in osteogenic medium for 1 week and 2 weeks. Alkaline phosphatase (ALP) activity assays and Alizarin red histochemical staining revealed reduced osteoblast differentiation in BMSCs from the Msi2−/− mice (Fig. 4d, e). The expression levels of osteoblast markers, such as Alp, bone sialoprotein (Bsp), Col1α1, Osterix and Atf4, also decreased in the Msi2−/− BMSCs (Fig. 4f).
Taken together, the above results revealed that Msi2 regulates the balance of BMSC fate commitment by repressing adipocyte differentiation and enhancing osteoblast differentiation.
MSI2 inhibits PPAR signaling in BMSCs
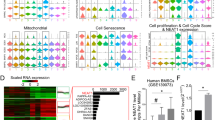
To explore the molecular mechanism by which Msi2 regulates osteoblast-adipocyte lineage commitment, we performed RNA sequencing analysis using BMSCs from the WT and Msi2−/− mice (7 days after osteoblast differentiation) and compared the gene expression profiles. Genes related to adipocyte differentiation showed upregulated expression, and genes related to osteoblast differentiation showed downregulated expression (Fig. 5a). Gene set enrichment analysis (GSEA) was then performed to identify significantly enriched Gene Ontology (GO) terms. Lipid localization or storage regulators and adipocyte differentiation markers showed upregulated expression in the Msi2−/− BMSCs (Fig. 5b). Ossification-, skeletal development- and bone development-related genes showed significantly downregulated expression (Fig. 5c). Kyoto Encyclopedia of Genes and Genomes pathway analysis indicated that the PPAR signaling pathway was significantly enhanced in the Msi2 knockout BMSCs (Fig. 5d). To further analyze the changes in the PPAR signaling pathway in the Msi2 knockout cells, we utilized GSEA to mine the RNA-seq data, and the results showed that Msi2 knockout increased the enrichment score for the PPAR signaling pathway module (Fig. 5e). Genes with upregulated expression that showed a significant difference in expression in the GSEA were visualized by a heatmap (Fig. 5f). The expression levels of the genes with upregulated and downregulated expression were further confirmed in the Msi2 knockout BMSCs by RT-PCR (Fig. 5g). As PPARγ is considered to be one of the major drivers of adipogenesis,10,11 these results suggested that Msi2 may regulate BMSC commitment by inhibiting the PPARγ signaling pathway.
MSI2 inhibits PPAR signaling in BMSCs. a Heatmap of RNA sequencing data between the WT and Msi2−/− mouse BMSCs cultured in osteoblast differentiation medium for 7 days, n = 2 for each group. b Upregulated (red) GO analysis associated with significantly regulated genes (P < 0.05) in the Msi2 knockout versus WT control groups. c Downregulated (blue) GO analysis associated with significantly regulated genes (P < 0.05) in the Msi2 knockout versus WT control groups. d Upregulated (red) pathways associated with significantly regulated genes (P < 0.05) in the Msi2 knockout versus WT control groups. e GSEA of the enrichment of all genes in RNA sequencing. f Heatmap of genes with upregulated expression in the PPAR signaling pathway obtained by GSEA. g qPCR results of adipogenesis-related gene (Cebpα, Lpl, Perilipin, Pparγ) and osteogenesis-related gene (Alp, Bsp, Col1α1) expression in the WT and Msi2−/− mouse BMSCs
Msi2 inhibits Cebpα translation and PPARγ expression in BMSCs
Msi2 is an RNA-binding protein. Previous results demonstrated that three phenylalanine residues in Msi2 are essential for Msi2 RNA binding. To determine whether RNA binding is essential for the function of Msi2, we mutated three phenylalanine residues essential for Msi2 RNA binding to leucine (F64/66/69 L) to generate an RNA binding-deficient mutant of Msi2 (hereafter Msi2RBDmut) (Fig. 6a).16,27 We next compared the function of Msi2 with that of Msi2RBDmut. As shown in Fig. 6b, overexpression of Msi2 reduced the differentiation of BMSCs into adipocytes, but Msi2RBDmut overexpression did not reduce the differentiation of BMSCs into adipocytes (Fig. 6b). Moreover, overexpression of Msi2 enhanced the differentiation of BMSCs into osteoblasts, but Msi2RBDmut overexpression did not (Fig. 6c). These results suggest that the mRNA binding activity of Msi2 is required for BMSC commitment.
Msi2 inhibits Cebpα translation and PPARγ activation in BMSCs. a Schematic illustration of Msi2 and the Msi2RBDmut mutation. b BMSCs isolated from 4-week-old wild-type mice and treated with Msi2 and Msi2RBD lentivirus. Cultures were stained with Oil Red O and BODIPY as shown. Scale bar = 200 μm. c BMSCs isolated from 4-week-old wild-type mice, and treated with Msi2 and Msi2RBD lentivirus. Cultures were stained with ALP, and ALP activity was quantified as shown. Scale bar = 3 mm. Data represent the mean ± SD, *P < 0.05, one-way ANOVA. d Western blot analysis of PPARγ and perilipin protein levels in the C3H10 cells overexpressing Flag-tagged Msi2 and Msi2RBDmut protein; GAPDH was used as a reference protein. e Schematic of the mouse Cebpα transcript. Bars, the putative MBEs (r(G/A)U1–3AGU). Two MBEs were identified within the 3′ UTR of Cebpα. CDS, coding sequence for mC/EBPα protein. f RIP with anti-Flag antibody from C3H10 cells expressing empty vector, Flag-tagged Msi2 or Flag–Msi2RBDmut. Coimmunoprecipitated RNAs were analyzed for the enrichment of Cebpα transcripts. n = 3 each. Data represent the mean ± SD, ***P < 0.001, ****P < 0.000 1, one-way ANOVA. g RIP with anti-Msi2 antibody or a control rabbit IgG from BMSCs. Coimmunoprecipitated RNAs were analyzed for the enrichment of Cebpα transcripts. n = 3 each. Data represent the mean ± SD, ***P < 0.001, ordinary one-way ANOVA. h qPCR results of Cebpα in the C3H10 cells overexpressing Flag-tagged Msi2 and Msi2RBDmut proteins. Data represent the mean ± SD, ns: no significance, one-way ANOVA. i qPCR results of Pparγ in the C3H10 cells overexpressing Flag-tagged Msi2 and Msi2RBDmut proteins. Data represent the mean ± SD, ****P < 0.000 1, one-way ANOVA. j Western blot analysis of C/EBPα, PPARγ, FABP4, LPL, perilipin and Msi2 protein levels in the WT and Msi2−/− mouse BMSCs. GAPDH was used as a reference protein. k The model of Msi2 regulating PPAR signaling
Transcriptional profiling analysis suggested that Msi2 may regulate BMSC commitment by inhibiting the PPARγ signaling pathway. Msi2 is considered to be a translational repressor by binding the 3′ UTR of the target mRNA.13 We next explored whether Msi2 regulates the PPARγ signaling pathway by repressing the translation of key components of PPARγ signaling. As shown in Fig. 6d, overexpression of Msi2 reduced the protein levels of PPARγ and perilipin when BMSCs were induced to differentiate into adipocytes (Fig. 6d). In contrast, Msi2RBDmut overexpression abolished the inhibitory effect of Msi2 (Fig. 6d). These results indicated that mRNA binding activity is required for Msi2 to inhibit the PPARγ signaling pathway.
We next examined how Msi2 relies on the mRNA binding ability to regulate PPAR signaling. The C/EBP family has been reported to be closely related to the regulation of PPAR signaling, and the mRNA level of Cebp factors was not changed significantly in our RNA sequencing data. We then examined the putative MBEs in the 3′ UTR of different Cebps and found that only Cebpα’s 3′ UTR has two MBEs; the Cebpβ and Cebpδ 3′ UTRs did not (Fig. 6e). We then performed an RNA immunoprecipitation (RIP) assay using C3H10 cells transfected with plasmids expressing Flag-tagged Msi2 or Flag-tagged Msi2RBDmut. Interestingly, Cebpα transcripts were significantly enriched by Flag immunoprecipitation when Flag-Msi2 was expressed. In contrast, Cebpα transcripts were not enriched when Flag-tagged Msi2RBDmut was expressed (Fig. 6f). These results suggested that Msi2 binds to the mRNA of Cebpα. Consistently, RIP with an anti-Msi2 antibody also specifically enriched Cebpα transcripts relative to that of an immunoglobulin-G (IgG) control (Fig. 6g), further confirming the interaction between Msi2 and Cebpα mRNA. Msi2 overexpression in C3H10 cells did not change the RNA level of Cebpα (Fig. 6h), However, the RNA level of Pparγ, which is regulated by Cebpα, was significantly downregulated when Msi2 was overexpressed in C3H10 cells (Fig. 6i). The protein level of Cebpα was increased significantly in the MSI2 knockout BMSCs, and PPARγ signaling markers were also significantly increased in the MSI2 knockout BMSCs (Fig. 6j). These data indicated that binding of Msi2 to Cebpα transcripts negatively regulates the translation of Cebpα. Regulation of PPARγ signaling by Msi2 is essential for the dynamic balance of the commitment between osteoblasts and adipocytes (Fig. 6k).
Msi2 expression is downregulated during aging
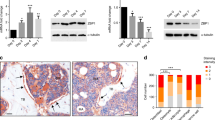
The depletion of Msi2 in mice led to decreased bone mass with increased marrow adipocytes, resembling aging-induced osteoporosis. We next examined whether Msi2 expression changed during aging. We isolated BMSCs from 2-month-old (young) or 24-month-old (old) mice and found that the Msi2 expression level was decreased in old BMSCs, as indicated by RT-qPCR assays (Fig. 7a). Immunohistochemical staining also showed that Msi2 protein expression levels were downregulated in the aged mice (Fig. 7b). μ-CT analysis confirmed that the bone mass of the old mice was significantly reduced (Fig. 7c, d), accompanied by increased Tb.Sp (Fig. 7e) and decreased Tb.N (Fig. 7f). Interestingly, compared with that in the young mice, cortical bone in the aging mice was thicker (Fig. S4A, B). Similar to the phenotype of the Msi2 knockout mice, abnormal accumulation of adipocytes in the bone marrow cavity of the aged mice was observed (Fig. 7g), suggesting the occurrence of aging-related osteoporosis. Immunofluorescence staining revealed decreased expression levels of the osteoblast marker OPN and increased expression levels of the adipocyte marker perilipin in the bone marrow cavity of the old mice (Fig. 7h, i). Similar to the scenario in Msi2−/− BMSCs, the RNA level of Cebpα remained unchanged in BMSCs isolated from the aged mice, and the RNA level of Pparγ increased in BMSCs isolated from the aged mice (Fig. 7j). Immunohistochemical staining also showed that PPARγ protein expression levels were upregulated in the aged mice (Fig. 7k, l). Consistent with previous reports,28 the mRNA level of the senescence marker p16 increased in the old BMSCs. In addition, the target genes of PPARγ increased significantly (Fig. S4C). The old BMSCs had a phenotype similar to that of the Msi2−/− BMSCs, which is consistent with the decreased expression level of Msi2 in the aged BMSCs.
Msi2 expression is downregulated during aging. a qPCR results of Msi2 expression in BMSCs from 8-week-old and 24-month-old mice (n = 4). Data represent the mean ± SD, ***P < 0.001, unpaired Student’s t test. b Immunohistochemistry staining of Msi2 from 8-week-old and 24-month-old mice. Scale bar = 50 μm. c Three-dimensional μ-CT images of trabecular bone of distal femurs isolated from 8-week-old female and 24-month-old female mice (n = 6). d μ-CT analysis of trabecular bone volume per tissue volume (BV/TV) in the distal femur of 8-week-old female and 24-month-old female mice. e μ-CT analysis of the distal femur of 8-week-old female and 24-month-old female mice for trabecular separation (Tb.Sp). f μ-CT analysis of the trabecular number (Tb.N) of the distal femur of 8-week-old female and 24-month-old female mice. g Hematoxylin-eosin staining of femurs from wild-type mice at 8 weeks and 24 months. Scale bar = 500 μm. h Immunostaining of perilipin A/B (green) and OPN (red) of femurs from 8-week-old and 24-month-old mice. Scale bar = 50 μm. i Quantification of the relative areas of OPN and Perilipin in (h). j qPCR results of Cebpα and Pparγ expression in BMSCs from 8-week-old and 24-month-old mice (n = 4). Data represent the mean ± SD, ***P < 0.01, ns: no significance, unpaired Student t test. k Immunohistochemistry staining of PPARγ from 8-week-old and 24-month-old mice. Scale bar = 100 μm. l Quantification of the relative area of PPARγ in (k)
These results suggest that Msi2 could be one of the contributors to aging-induced osteoporosis. In old BMSCs, the reduction in the Msi2 expression level leads to a shift in the differentiation balance of BMSCs. Adipogenesis is enhanced, and osteoblastogenesis declines, which results in aging-induced osteoporosis.
Discussion
RNA-binding proteins play an important role in cell fate determination through posttranscriptional regulation. Here, we found that the RNA-binding protein Msi2 controls the fate of BMSCs. By binding to the 3′ UTR of the mRNA of the key adipogenesis-related factor, Msi2 inhibits the adipogenic potential of BMSCs. In aged BMSCs, the Msi2 expression level decreased, and the balance of BMSC differentiation shifted toward adipogenesis, which led to osteoporosis indicated by abnormal accumulation of adipocytes in the bone marrow cavity and decreased bone mass. Our results revealed that Msi2 is an important contributor to osteoporosis by modulating protein translation.
MSI is an evolutionarily conserved family of RNA-binding proteins that play key roles in the maintenance of self-renewal of stem cells and HSC fate.15,29 Previous studies on Msi2 have mainly focused on its function in tumors.30,31,32 In this study, we reported for the first time that Msi2 regulates BMSC commitment. Msi2 plays a key role in maintaining the balance between osteoblastogenesis and adipogenesis. Although several previous studies have shown that there is a negative correlation between MSC osteogenesis and adipogenesis,3,4,33,34 the RNA-binding protein involved is the first to be discovered. Among the limited number of genes identified downstream of MSI2,13,19,27 our study found that MSI2 specifically binds to the 3′ UTR of Cebpα to regulate PPARγ signaling and control the differentiation of BMSCs. These results suggest that RNA-binding proteins such as Msi2 can be considered upstream of the PPARγ signaling pathway for drug targeting research.
Aging-related osteoporosis causes progressive fat accumulation and trabecular bone loss.35 Existing studies on osteoporosis have mainly focused on transcription factors, epigenetics and hormone metabolism.4,36,37 However, the mechanism of RNA-binding proteins involved in regulating osteoporosis remains to be discovered. In our study, it was confirmed that Msi2 greatly reduced both RNA and protein levels in aging mouse bone samples, and theMsi2-deficient mice showed an age-dependent osteoporosis-like phenotype. Our research established a link between osteoporosis and RNA-binding proteins, and we discovered a regulatory relationship between MSI2 and PPARγ signaling. This result may provide new ideas for future research on targeted therapy for osteoporosis.
There are several limitations in our study. The animal model used in this study is Msi2−/− mice, and the influence of other organs on bone cannot be ruled out. Although the expression level of Msi2 in the BMSCs was not the highest, we found that the expression level of Msi2 in the spleen was lower than that in the BMSCs, and Msi2 has been shown to play an important role in the spleen. Our knockout mice also have a similar phenotype of reduced spleen.19 We observed a certain difference between the detection of RNA levels and the detection of protein levels, which may be caused by slightly different posttranscriptional translation of proteins in different tissue environments. However, in vivo and in vitro experiments showed that Msi2 was knocked out in BMSCs, which resulted in an osteoporotic phenotype. Although we have established a connection between Msi2 and PPARγ signaling, the direct genetic evidence remains to be further examined.
Our work reveals the function of Msi2 in regulating the commitment of MSCs, thereby regulating bone homeostasis. We further tested the proportion of HSCs in the Msi2−/− mice and the control mice, and the results showed that the absence of Msi2 did not affect the proportion of HSCs in bone marrow cells (Fig. S3C). Although we also observed that Msi2 deletion inhibited osteoclast differentiation in vitro,25 there was no significant difference in TRAP staining in vivo. The effect of Msi2 on bone in HSCs through other methods of compensation cannot be ruled out. This issue also needs to be revealed in future research.
Because RNA sequencing showed the tight integration of the Msi2 and PPAR signaling pathways, Msi2 plays a key role in the regulation of the PPAR signaling pathway in mesenchymal stem cells. We focused on genes such as Cebpα, Cebpβ, Pparg, Fabp4, and Lpl. We excluded some genes by analyzing whether there are Msi2 binding sites on the 3′ UTR and then conducted RIP verification, but unfortunately, the results were not verified by the RIP experiment. In addition, we focused on Runx2, which is the core transcription factor for bone formation. However, we did not find a mouse skull closure disorder, and the results were not verified in the RIP experiment. We do not know whether Msi2 will bind to the 3′ UTR of other molecules and regulate protein expression, nor can we eliminate other ways of Msi2 molecular regulation. Future work should investigate these issues.
RNA-binding proteins are closely related to the occurrence and development of cancer. As a potential target for cancer treatment, small molecules have been developed to act as inhibitors of Msi2. We need to consider the effect of this medication on patients with osteoporosis and other skeletal degenerative diseases.38,39 Realizing tissue-specific and spatiotemporal specificity to restore the normal expression of Msi2 will be crucial for the occurrence and development of the disease. Exploring compounds and small molecules that regulate Msi2 will promote the treatment of cancer and osteoporosis.
Overall, our work demonstrated that Msi2 functions as a repressor of Cebpα to inhibit the activation of PPARγ signaling. This work defined the role of Msi2 in regulating MSC commitment and identified a new target for aging-induced osteoporosis treatment. It is not clear whether Msi2 is also involved in regulating the translation of other targets in the process of aging. If so, how Msi2 cooperates among different molecules will be another interesting question worthy of further study.
Materials and methods
Msi2 mice were constructed using the Crispr-Cas9 strategy. Msi2−/− mice were constructed using the Crispr-Cas9 strategy. At the end of the first exon of Msi2, 10 bases were deleted using Crispr-Cas9 technology, resulting in a gene frameshift. The deleted base sequence is AGCACGACCC. All mice analyzed had a C57BL/6 background. Animals were maintained under specific pathogen-free conditions in the institutional animal facility of the Shanghai Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences. All animal experiments were performed with a protocol approved by the Animal Care and Use Committee of Shanghai Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences.
Antibodies
Anti-Flag antibody (F-3165, 1:5 000, Sigma), rabbit IgG (SC-2027, Santa Cruz Biotechnology), anti-perilipin A/B (Sigma, P1873), and anti-OPN (R&D, AF808) were used. Anti-Col1a1 (Rockland, 600-400-103), anti-PPARγ (Santa Cruz, sc-7273), anti-LPL (R&D, AF7197) and anti-Msi2 (Abcam, ab76148) were obtained.
Cell culture
Cells were cultured at 37 °C in humidified incubators containing an atmosphere of 5% CO2. HEK-293T cells were maintained in DMEM (Corning, Corning, NY) supplemented with 10% fetal bovine serum (FBS) and 1% penicillin/streptomycin (Gibco) solution. C3H10T1/2 cells were maintained in α-MEM (Corning, Corning, NY) supplemented with 10% FBS and 1% penicillin/streptomycin (Gibco) solution.
Osteoblast differentiation and adipocyte differentiation
We collected femurs from the WT and Msi2−/− mice and flushed out the bone marrow cells with phosphate-buffered saline (PBS). All nuclear cells were seeded (2 × 106 cells per dish) in 100 mm culture dishes (Corning) and incubated at 37 °C under 5% CO2 conditions. After 24 h, the cells were supplemented with fresh medium. After 48 h, nonadherent cells were washed with PBS, and adherent cells were cultured in alpha minimum essential medium (Corning, Corning, NY) supplemented with 10% FBS and 1% penicillin/streptomycin (Gibco) solution for an additional 5 days.
For induction of the differentiation of BMSCs into adipocytes, cells were first cultured in adipogenic induction medium (α-MEM/10% FBS containing 1 μmol·L−1 dexamethasone, 0.1 mmol.L−1 rosiglitazone, 0.5 mmol·L−1 IBMX, 10 µg.mL−1 insulin) for 1 day, and then, adipogenic maintenance medium (α-MEM/10% FBS containing 10 µg.mL−1 insulin) was added for 2 days. After mature adipocyte formation, cells were stained with 2 mg.mL−1 Oil Red O solution or BODIPY 493/503.
For induction of the differentiation of BMSCs into osteoblasts, cells were cultured in α-MEM containing 10% FBS, 50 μg.mL−1 L-ascorbic acid, and 1080 mg.mL−1 β-glycerophosphate. The osteoblast differentiation assay was performed following a previously published method. For quantitative analysis of ALP activity, cells were incubated with Alamar Blue to calculate cell numbers and then incubated with phosphatase substrate (Sigma-Aldrich, St. Louis, MO) dissolved in 6.5 mmol.L−1 Na2CO3, 18.5 mmol.L−1 NaHCO3, and 2 mmol.L−1 MgCl2 after washing with PBS. ALP activity was then read with a luminometer (Envision). Bone nodule formation was stained with 1 mg.mL−1 Alizarin red S solution (pH 5.5) after 14 days of induction.
RIP assay
BMSCs isolated from the WT and Msi2−/− mice or C3H10 cells infected with Flag-tagged Msi2 or Flag-tagged Msi2RBDmut lentivirus were lysed in 50 mmol.L−1 Tris/HCl (pH 7.4) containing 100 mmol.L−1 NaCl, 5 mmol.L−1 EDTA, 1% NP-40, Protease Inhibitor Cocktail (HY-K0010, 1:100, MedChem Express), and RNase inhibitor (Thermo Scientific). We performed immunoprecipitations with anti-FLAG, anti-Msi2 or rabbit normal IgG and protein G magnetic beads for 2 h at 4 °C. The immunoprecipitated protein-RNA complexes were washed five times with wash buffers (25 mmol.L−1 Tris/HCl (pH 7.4), 20 mmol.L−1 MgCl2, 100 mmol.L−1 NaCl, 0.2% Tween-20, and 0.05% NP40). Total RNA was purified from the washed beads using TRIzol (T9424, Sigma) and subjected to RT-qPCR analysis for quantification. For each sample, we calculated the enrichment multiple of the transcript content in the RIP score relative to the amount (RIP/input) that was present before the RIP in the input sample.
RNA-seq
Isolation of total RNA with TRIzol comes from samples of osteoblastic differentiation induced by BMSCs from the WT and Msi2-deficient mice for 7 days. RNA library construction, sequencing and analysis are provided by Novogene. The top GO categories were selected according to the P values.
Real-time RT-PCR analysis
Total RNA was isolated from cells with TRIzol reagent (T9424, Sigma), and first-strand cDNA was synthesized from 500 ng of total RNA using the PrimeScript™ RT Reagent Kit (PR037A, TaKaRa). Real-time reverse transcriptase RT-PCR was performed with the Bio-Rad CFX96 system. Gene expression from RT-PCR was quantified relative to that of Hprt or Gapdh.
IP and immunoblotting
First, 293 T cells were seeded at 1 – 2 × 107 cells per 10 cm dish and cultured overnight. After transfection with PEI for 48 h, the cells were harvested and washed with cold PBS following experimental treatments. Then, the cells were lysed with EBC buffer (50 mmol.L−1 Tris, pH 7.5, 120 mmol.L−1 NaCl and 0.5% NP-40) containing protease inhibitor cocktail (HY-K0010, 1:100, MedChem Express). After ultrasonication (power: 25%, sonicate 5 s, stop 5 s, five times), lysates were subjected to IP with anti-Flag beads (M2, Sigma) at 4 °C for 4–6 h or overnight, followed by washing in lysis buffer, SDS–PAGE electrophoresis and immunoblotting with the indicated antibody.
μ-QCT analysis
Preparation of skeletal tissue and μ-QCT analysis were performed as previously described.3 The mouse femurs isolated from age- and sex-matched mice were skinned and fixed in 70% ethanol. Scanning was performed with the instrument μ-QCT system SkyScan1176 (Bruker Biospin). The mouse femurs were scanned at a 9 μm resolution for quantitative analysis. Three-dimensional images were reconstructed using a fixed threshold.
Histology and immunofluorescence
Tissues were fixed in 4% paraformaldehyde for 48 h, incubated in 15% DEPC-EDTA (pH 7.8) and ultrasonically decalcified. The specimens were embedded in paraffin or OCT and cut into 7 μm sections.
Immunofluorescence assay: Sections were blocked in PBS with 10% horse serum and 0.1% Triton for 1 h at room temperature. Then, the cells were stained overnight with rabbit anti-perilipin A/B (Sigma, P1873, 1:1 000, USA) and OPN (1:1 000; R&D, AF808). Donkey-anti-rabbit Alexa Fluor 488 (1:1 000; Molecular Probes, A21206) and donkey-anti-goat Cy3 (1:1 000; Jackson ImmunoResearch, 705–165–147) were used as secondary antibodies. DAPI (Sigma, D8417) was used for counterstaining. Slides were mounted with anti-fluorescence mounting medium (Dako, S3023), and images were acquired with an Olympus FV3000 and SP8 confocal microscope.
Immunohistochemical staining and Col1a1 (1:100; Rockland, 600–400–103) staining were performed as described by Dako.
Tissue sections were used for TRAP, BODIPY, and Oil Red O staining according to the standard protocol.
Statistics
Statistical analysis was performed by unpaired, two-tailed Student’s t test for comparison between two groups using GraphPad Prism Software. Through Prism software, one-way ANOVA was used to compare and analyze the three groups of data. A P value of <0.05 was considered statistically significant.
References
Letarouilly, J. G., Broux, O. & Clabaut, A. New insights into the epigenetics of osteoporosis. Genomics 111, 793–798 (2019).
Trajanoska, K. & Rivadeneira, F. The genetic architecture of osteoporosis and fracture risk. Bone 126, 2–10 (2019).
Wang, L. et al. H3K36 trimethylation mediated by SETD2 regulates the fate of bone marrow mesenchymal stem cells. PLoS Biol. 16, e2006522 (2018).
Li, H. et al. FOXP1 controls mesenchymal stem cell commitment and senescence during skeletal aging. J. Clin. Investig. 127, 1241–1253 (2017).
Pittenger, M. F. et al. Multilineage potential of adult human mesenchymal stem cells. Science 284, 143–147 (1999).
Chen, Q. et al. Fate decision of mesenchymal stem cells: adipocytes or osteoblasts? Cell Death Differ. 23, 1128–1139 (2016).
Farmer, S. R. Transcriptional control of adipocyte formation. Cell Metab. 4, 263–273 (2006).
Pierce, J. L., Begun, D. L., Westendorf, J. J. & McGee-Lawrence, M. E. Defining osteoblast and adipocyte lineages in the bone marrow. Bone 118, 2–7 (2019).
Wu, Z. et al. Cross-regulation of C/EBP alpha and PPAR gamma controls the transcriptional pathway of adipogenesis and insulin sensitivity. Mol. Cell 3, 151–158 (1999).
Takada, I., Kouzmenko, A. P. & Kato, S. Wnt and PPARgamma signaling in osteoblastogenesis and adipogenesis. Nat. Rev. Rheumatol. 5, 442–447 (2009).
Bai, M. et al. Targeted genetic screening in mice through haploid embryonic stem cells identifies critical genes in bone development. PLoS Biol. 17, e3000350 (2019).
Huang, X. et al. Insulin-like growth factor 2 mRNA-binding protein 1 (IGF2BP1) in cancer. J. Hematol. Oncol. 11, 88 (2018).
Fox, R. G., Park, F. D., Koechlein, C. S., Kritzik, M. & Reya, T. Musashi signaling in stem cells and cancer. Annu. Rev. Cell Dev. Biol. 31, 249–267 (2015).
Kawahara, H. et al. Neural RNA-binding protein Musashi1 inhibits translation initiation by competing with eIF4G for PABP. J. Cell Biol. 181, 639–653 (2008).
Park, S. M. et al. Musashi-2 controls cell fate, lineage bias, and TGF-β signaling in HSCs. J. Exp. Med. 211, 71–87 (2014).
Hattori, A. et al. Cancer progression by reprogrammed BCAA metabolism in myeloid leukaemia. Nature 545, 500–504 (2017).
Hemmati, H. D. et al. Cancerous stem cells can arise from pediatric brain tumors. Proc. Natl. Acad. Sci. USA 100, 15178–15183 (2003).
Oskarsson, T. et al. Breast cancer cells produce tenascin C as a metastatic niche component to colonize the lungs. Nat. Med. 17, 867–874 (2011).
Park, S. M. et al. Musashi2 sustains the mixed-lineage leukemia-driven stem cell regulatory program. J. Clin. Investig. 125, 1286–1298 (2015).
Okano, H. et al. Function of RNA-binding protein Musashi-1 in stem cells. Exp. Cell Res. 306, 349–356 (2005).
de Andrés-Aguayo, L. et al. Musashi 2 is a regulator of the HSC compartment identified by a retroviral insertion screen and knockout mice. Blood 118, 554–564 (2011).
Ma, X. et al. Msi2 Maintains Quiescent State of Hair Follicle Stem Cells by Directly Repressing the Hh Signaling Pathway. J. Investig. Dermatol. 137, 1015–1024 (2017).
Vu, L. P. et al. Functional screen of MSI2 interactors identifies an essential role for SYNCRIP in myeloid leukemia stem cells. Nat. Genet 49, 866–875 (2017).
Hope, K. J. & Sauvageau, G. Roles for MSI2 and PROX1 in hematopoietic stem cell activity. Curr. Opin. Hematol. 18, 203–207 (2011).
Fujiwara, T., Zhou, J., Ye, S. & Zhao, H. RNA-binding protein Musashi2 induced by RANKL is critical for osteoclast survival. Cell Death Dis. 7, e2300 (2016).
Hong, I. S. et al. The effects of hedgehog on RNA binding protein Msi1 during the osteogenic differentiation of human cord blood-derived mesenchymal stem cells. Bone 56, 416–425 (2013).
Imai, T. et al. The neural RNA-binding protein Musashi1 translationally regulates mammalian numb gene expression by interacting with its mRNA. Mol. Cell. Biol. 21, 3888–3900 (2001).
Baker, D. J. et al. Clearance of p16Ink4a-positive senescent cells delays ageing-associated disorders. Nature 479, 232–236 (2011).
das Chagas, P. F., Baroni, M., Brassesco, M. S. & Tone, L. G. Interplay between the RNA binding-protein Musashi and developmental signaling pathways. J. Gene Med. 22, e3136 (2020).
Barbouti, A. et al. A novel gene, MSI2, encoding a putative RNA-binding protein is recurrently rearranged at disease progression of chronic myeloid leukemia and forms a fusion gene with HOXA9 as a result of the cryptic t(7;17)(p15;q23). Cancer Res. 63, 1202–1206 (2003).
Ito, T. et al. Regulation of myeloid leukaemia by the cell-fate determinant Musashi. Nature 466, 765–768 (2010).
Wang, S. et al. Transformation of the intestinal epithelium by the MSI2 RNA-binding protein. Nat. Commun. 6, 6517 (2015).
Zhao, X. et al. ZBP1 (DAI/DLM-1) promotes osteogenic differentiation while inhibiting adipogenic differentiation in mesenchymal stem cells through a positive feedback loop of Wnt/β-catenin signaling. Bone Res. 8, 12 (2020).
Wu, M. et al. Cbfβ governs osteoblast-adipocyte lineage commitment through enhancing β-catenin signaling and suppressing adipogenesis gene expression. Proc. Natl. Acad. Sci. USA 114, 10119–10124 (2017).
Moerman, E. J., Teng, K., Lipschitz, D. A. & Lecka-Czernik, B. Aging activates adipogenic and suppresses osteogenic programs in mesenchymal marrow stroma/stem cells: the role of PPAR-gamma2 transcription factor and TGF-beta/BMP signaling pathways. Aging Cell 3, 379–389 (2004).
Xu, Z. et al. SMURF2 regulates bone homeostasis by disrupting SMAD3 interaction with vitamin D receptor in osteoblasts. Nat. Commun. 8, 14570 (2017).
Adami, G., Rahn, E. J. & Saag, K. G. Glucocorticoid-induced osteoporosis: from clinical trials to clinical practice. Ther. Adv. Musculoskelet. Dis. 11, 1759720x19876468 (2019).
Wang, M. et al. Suppression of Musashi‑2 by the small compound largazole exerts inhibitory effects on malignant cells. Int J. Oncol. 56, 1274–1283 (2020).
Kudinov, A. E., Karanicolas, J., Golemis, E. A. & Boumber, Y. Musashi RNA-Binding Proteins as Cancer Drivers and Novel Therapeutic Targets. Clin. Cancer Res. 23, 2143–2153 (2017).
Acknowledgements
We thank the cell biology core facility and the animal core facility of Shanghai Institute of Biochemistry and Cell Biology for assistance.
Funding
This work was supported by the National Natural Science Foundation of China (NSFC) [81672119 and 81725010 to W.Z.], W Z is a scholar of ‘the National Science Fund for Distinguished Young Scholars’ (NSFC) [81725010], the Strategic Priority Research Program of the Chinese Academy of Science (XDA16020400 to P.H.), Ministry of Science and Technology of China (2017YFA0102700 to P.H.), National Natural Science Foundation of China (32170804 to PH) P.H.), the fellowship of China Postdoctoral Science Foundation (2021TQ0207, 2021M702184 to J.S.) and the ‘Basic research project of Shanghai Sixth People’s Hospital’ (ynqn202102 to J.S.).
Author information
Authors and Affiliations
Contributions
W.Z. P.H., Y.Z. and J.S. conceived and supervised the study. S.Z. and J.S. performed the experiments, analyzed the data and wrote the manuscript. L.Y. and P.H. provided genetic strains of mice. J.W. and B.J. made the constructs, C.L. performed RT-PCR experiments, S.Z., J.S., C.L. and Y.H. bred and analyzed Msi2-/- mice. J.S. and S.H. finished cell experiments. P.J.,Y.Z., P.H. and W.Z. contributed ideas and reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Suo, J., Zou, S., Wang, J. et al. The RNA-binding protein Musashi2 governs osteoblast-adipocyte lineage commitment by suppressing PPARγ signaling. Bone Res 10, 31 (2022). https://doi.org/10.1038/s41413-022-00202-3
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41413-022-00202-3
This article is cited by
-
Aging adipose tissue, insulin resistance, and type 2 diabetes
Biogerontology (2024)
-
Downregulation of the RNA-binding protein PUM2 facilitates MSC-driven bone regeneration and prevents OVX-induced bone loss
Journal of Biomedical Science (2023)
-
Accelerated aging in articular cartilage by ZMPSTE24 deficiency leads to osteoarthritis with impaired metabolic signaling and epigenetic regulation
Cell Death & Disease (2023)
-
Effect of RNA-binding proteins on osteogenic differentiation of bone marrow mesenchymal stem cells
Molecular and Cellular Biochemistry (2023)