Abstract
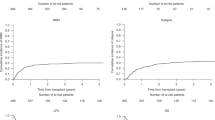
Although allogeneic hematopoietic stem-cell transplantation (HSCT) provides high cure rates for children with high-risk acute lymphoblastic leukaemia (ALL), relapses remain the main cause of treatment failure. Whereas donor killer cell immunoglobulin-like receptor (KIR) genotype was shown to impact on relapse incidence in adult myeloid leukaemia similar studies in paediatric ALL are largely missing. Effect of donor KIR genotype on transplant outcome was evaluated in 317 children receiving a first myeloablative HSCT from an HLA-matched unrelated donor or sibling within the prospective ALL-SCT-BFM-2003 trial. Analysis of donor KIR gene polymorphism revealed that centromeric presence and telomeric absence of KIR B haplotypes was associated with reduced relapse risk. A centromeric/telomeric KIR score (ct-KIR score) integrating these observations correlated with relapse risk (hazard ratio (HR) 0.58; P = 0.002) while it had no impact on graft-versus-host disease or non-relapse mortality. In multivariable analyses ct-KIR score was associated with reduced relapse risk (HR 0.58; P = 0.003) and a trend towards improved event-free survival (HR 0.76; P = 0.059). This effect proved independent of MRD level prior to HSCT. Our data suggest that in children with ALL undergoing HSCT after myeloablative conditioning, donor selection based on KIR genotyping holds promise to improve clinical outcome by decreasing relapse risk and prolonged event-free survival.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Peters C, Schrappe M, von Stackelberg A, Schrauder A, Bader P, Ebell W, et al. Stem-cell transplantation in children with acute lymphoblastic leukemia: A prospective international multicenter trial comparing sibling donors with matched unrelated donors-The ALL-SCT-BFM-2003 trial. J Clin Oncol. 2015;33:1265–74.
Babor F, Fischer JC, Uhrberg M. The role of KIR genes and ligands in leukemia surveillance. Front Immunol. 2013;4:27.
Manser AR, Weinhold S, Uhrberg M. Human KIR repertoires: shaped by genetic diversity and evolution. Immunol Rev. 2015;267:178–96.
Cooley S, Trachtenberg E, Bergemann TL, Saeteurn K, Klein J, Le CT, et al. Donors with group B KIR haplotypes improve relapse-free survival after unrelated hematopoietic cell transplantation for acute myelogenous leukemia. Blood. 2009;113:726–32.
Cooley S, Weisdorf DJ, Guethlein LA, Klein JP, Wang T, Le CT, et al. Donor selection for natural killer cell receptor genes leads to superior survival after unrelated transplantation for acute myelogenous leukemia. Blood. 2010;116:2411–9.
Cooley S, Weisdorf DJ, Guethlein LA, Klein JP, Wang T, Marsh SG, et al. Donor killer cell Ig-like receptor B haplotypes, recipient HLA-C1, and HLA-C mismatch enhance the clinical benefit of unrelated transplantation for acute myelogenous leukemia. J Immunol. 2014;192:4592–4600.
Oevermann L, Michaelis SU, Mezger M, Lang P, Toporski J, Bertaina A, et al. KIR B haplotype donors confer a reduced risk for relapse after haploidentical transplantation in children with ALL. Blood. 2014;124:2744–7.
Sobecks RM, Wang T, Askar M, Gallagher MM, Haagenson M, Spellman S, et al. Impact of KIR and HLA genotypes on outcomes after reduced-intensity conditioning hematopoietic cell transplantation. Biol Blood Marrow Transplant. 2015;21:1589–96.
Locatelli F, Merli P, Pagliara D, Li Pira G, Falco M, Pende D et al. Outcome of children with acute leukemia given HLA-haploidentical HSCT after alphabeta T-cell and B-cell depletion. Blood https://doi.org/10.1182/blood-2017-04-779769 2017.
Uhrberg M, Parham P, Wernet P. Definition of gene content for nine common group B haplotypes of the Caucasoid population: KIR haplotypes contain between seven and eleven KIR genes. Immunogenetics. 2002;54:221–9.
Vilches C, Castano J, Gomez-Lozano N, Estefania E. Facilitation of KIR genotyping by a PCR-SSP method that amplifies short DNA fragments. Tissue Antigens. 2007;70:415–22.
Prentice RL, Kalbfleisch JD, Peterson AV Jr., Flournoy N, Farewell VT, et al. The analysis of failure times in the presence of competing risks. Biometrics. 1978;34:541–54.
Gray R. A class of K-sample tests for comparing the cumulative incidence of a competing risk. Ann Stat. 1988;16:1141–54.
Fine J, Gray R. A proportional hazards model for the subdistribution of a competing risk. J Am Stat Assoc. 1999;94:496–509.
Bachanova V, Weisdorf DJ, Wang T, Marsh SG, Trachtenberg E, Haagenson MD, et al. Donor KIR B genotype improves progression-free survival of non-Hodgkin lymphoma patients receiving unrelated donor transplantation. Biol Blood Marrow Transplant. 2016;22:1602–7.
Lee SJ, Klein J, Haagenson M, Baxter-Lowe LA, Confer DL, Eapen M, et al. High-resolution donor-recipient HLA matching contributes to the success of unrelated donor marrow transplantation. Blood. 2007;110:4576–83.
Woolfrey A, Klein JP, Haagenson M, Spellman S, Petersdorf E, Oudshoorn M, et al. HLA-C antigen mismatch is associated with worse outcome in unrelated donor peripheral blood stem cell transplantation. Biol Blood Marrow Transplant. 2011;17:885–92.
Pidala J, Lee SJ, Ahn KW, Spellman S, Wang HL, Aljurf M, et al. Nonpermissive HLA-DPB1 mismatch increases mortality after myeloablative unrelated allogeneic hematopoietic cell transplantation. Blood. 2014;124:2596–606.
Kollman C, Spellman SR, Zhang MJ, Hassebroek A, Anasetti C, Antin JH, et al. The effect of donor characteristics on survival after unrelated donor transplantation for hematologic malignancy. Blood. 2016;127:260–7.
Ljungman P, Brand R, Hoek J, de la Camara R, Cordonnier C, Einsele H, et al. Donor cytomegalovirus status influences the outcome of allogeneic stem cell transplant: a study by the European group for blood and marrow transplantation. Clin Infect Dis. 2014;59:473–81.
Bader P, Kreyenberg H, von Stackelberg A, Eckert C, Salzmann-Manrique E, Meisel R, et al. Monitoring of minimal residual disease after allogeneic stem-cell transplantation in relapsed childhood acute lymphoblastic leukemia allows for the identification of impending relapse: results of the ALL-BFM-SCT 2003 trial. J Clin Oncol. 2015;33:1275–84.
Lee SJ, Klein JP, Barrett AJ, Ringden O, Antin JH, Cahn JY, et al. Severity of chronic graft-versus-host disease: association with treatment-related mortality and relapse. Blood. 2002;100:406–14.
Ruggeri L, Capanni M, Urbani E, Perruccio K, Shlomchik WD, Tosti A, et al. Effectiveness of donor natural killer cell alloreactivity in mismatched hematopoietic transplants. Science. 2002;295:2097–2100.
Ruggeri L, Mancusi A, Capanni M, Urbani E, Carotti A, Aloisi T, et al. Donor natural killer cell allorecognition of missing self in haploidentical hematopoietic transplantation for acute myeloid leukemia: challenging its predictive value. Blood. 2007;110:433–40.
Mehta RS, Rezvani K. Can we make a better match or mismatch with KIR genotyping? Hematology Am Soc Hematol Educ Program. 2016;2016:106–18.
Leung W. Use of NK cell activity in cure by transplant. Br J Haematol. 2011;155:14–29.
Leung W, Iyengar R, Turner V, Lang P, Bader P, Conn P, et al. Determinants of antileukemia effects of allogeneic NK cells. J Immunol. 2004;172:644–50.
Venstrom JM, Pittari G, Gooley TA, Chewning JH, Spellman S, Haagenson M, et al. HLA-C-dependent prevention of leukemia relapse by donor activating KIR2DS1. New Engl J Med. 2012;367:805–16.
Bari R, Rujkijyanont P, Sullivan E, Kang G, Turner V, Gan K, et al. Effect of donor KIR2DL1 allelic polymorphism on the outcome of pediatric allogeneic hematopoietic stem-cell transplantation. J Clin Oncol. 2013;31:3782–90.
Stringaris K, Adams S, Uribe M, Eniafe R, Wu CO, Savani BN, et al. Donor KIR Genes 2DL5A, 2DS1 and 3DS1 are associated with a reduced rate of leukemia relapse after HLA-identical sibling stem cell transplantation for acute myeloid leukemia but not other hematologic malignancies. Biol Blood Marrow Transplant. 2010;16:1257–64.
Kroger N, Binder T, Zabelina T, Wolschke C, Schieder H, Renges H, et al. Low number of donor activating killer immunoglobulin-like receptors (KIR) genes but not KIR-ligand mismatch prevents relapse and improves disease-free survival in leukemia patients after in vivo T-cell depleted unrelated stem cell transplantation. Transplantation. 2006;82:1024–30.
Cooley S, Miller JS. “Self”-reflection by KIR. Blood. 2009;114:2–3.
Pfeiffer M, Schumm M, Feuchtinger T, Dietz K, Handgretinger R, Lang P. Intensity of HLA class I expression and KIR-mismatch determine NK-cell mediated lysis of leukaemic blasts from children with acute lymphatic leukaemia. Br J Haematol. 2007;138:97–100.
Chen C, Busson M, Rocha V, Appert ML, Lepage V, Dulphy N, et al. Activating KIR genes are associated with CMV reactivation and survival after non-T-cell depleted HLA-identical sibling bone marrow transplantation for malignant disorders. Bone Marrow Transplant. 2006;38:437–44.
Symons HJ, Leffell MS, Rossiter ND, Zahurak M, Jones RJ, Fuchs EJ. Improved survival with inhibitory killer immunoglobulin receptor (KIR) gene mismatches and KIR haplotype B donors after nonmyeloablative, HLA-haploidentical bone marrow transplantation. Biol Blood Marrow Transplant. 2010;16:533–42.
Furst D, Muller C, Vucinic V, Bunjes D, Herr W, Gramatzki M, et al. High-resolution HLA matching in hematopoietic stem cell transplantation: a retrospective collaborative analysis. Blood. 2013;122:3220–9.
Horan J, Wang T, Haagenson M, Spellman SR, Dehn J, Eapen M, et al. Evaluation of HLA matching in unrelated hematopoietic stem cell transplantation for nonmalignant disorders. Blood. 2012;120:2918–24.
Wilson MJ, Torkar M, Haude A, Milne S, Jones T, Sheer D, et al. Plasticity in the organization and sequences of human KIR/ILT gene families. Proc Natl Acad Sci USA. 2000;97:4778–83.
Martin AM, Kulski JK, Gaudieri S, Witt CS, Freitas EM, Trowsdale J, et al. Comparative genomic analysis, diversity and evolution of two KIR haplotypes A and B. Gene. 2004;335:121–31.
van Bergen J, Trowsdale J. Ligand specificity of Killer cell Immunoglobulin-like receptors: a brief history of KIR. Front Immunol. 2012;3:394.
Acknowledgements
We thank all parents who gave their consent to use the biological material from their minors. This work was supported by the Deutsche Krebshilfe e.V. (M.U., R.M., and A.B.) (project 110351), the Forschungskommission of the Medical Faculty of the Heinrich-Heine University Düsseldorf (F.B.), and the TRANSAID - Stiftung fuer krebskranke Kinder (F.B.). The current affiliation of M.Si. is Department of Pediatrics and Mother and Child Center, Hospital Neuwerk, Mönchengladbach, Germany.
Author contributions
F.B., A.M., J.M., C.E. and N.S. performed the experiments; M.S., M.A., G.C., T.F., B.G., T.G., P.A.H., B.K., P.L., M.M., I.M., J.M., L.O., H.P., F.R.S., M.S., D.S., B.S., W.W., G.E., M.Z., M.S., A.B. and P.B. provided samples and conducted data the analysis and interpretation, and participated in patient care; E.G. and U.P. designed and performed the bioinformatic analyses; F.B., C.P., M.U. and R.M. wrote the manuscript, designed and directed the study; and all authors contributed to the research and approved the final manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Babor, F., Peters, C., Manser, A.R. et al. Presence of centromeric but absence of telomeric group B KIR haplotypes in stem cell donors improve leukaemia control after HSCT for childhood ALL. Bone Marrow Transplant 54, 1847–1858 (2019). https://doi.org/10.1038/s41409-019-0543-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41409-019-0543-z
This article is cited by
-
Natural killer cell therapy for hematologic malignancies: successes, challenges, and the future
Stem Cell Research & Therapy (2021)