Abstract
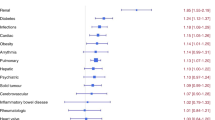
We analyzed micro-RNAs (miRs) as possible diagnostic biomarkers for relevant comorbidities prior to and prognostic biomarkers for mortality following hematopoietic cell transplantation (HCT). A randomly selected group of patients (n = 36) were divided into low-risk (HCT-comorbidity index [HCT-CI] score of 0 and survived HCT) and high-risk (HCT-CI scores ≥ 4 and deceased after HCT) groups. There were 654 miRs tested and 19 met the pre-specified significance level of p < 0.1. In subsequent models, only eight miRs maintained statistical significance in regression models after adjusting for baseline demographic factors; miRs-374b and -454 were underexpressed, whereas miRs-142-3p, -191, -424, -590-3p, -29c, and -15b were overexpressed among high-risk patients relative to low-risk patients. Areas under the curve for these eight miRs ranged between 0.74 and 0.81, suggesting strong predictive capacity. Consideration of miRs may improve risk assessment of mortality and should be further explored in larger future prospective studies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Sorror ML, Maris MB, Storb R, Baron F, Sandmaier BM, Maloney DG, et al. Hematopoietic cell transplantation (HCT)-specific comorbidity index: a new tool for risk assessment before allogeneic HCT. Blood. 2005;106:2912–9.
Lu M, Zhang Q, Deng M, Miao J, Guo Y, Gao W, et al. An analysis of human microRNA and disease associations. PLoS ONE. 2008;3:e3420.
Li M, Marin-Muller C, Bharadwaj U, Chow KH, Yao Q, Chen C. MicroRNAs: control and loss of control in human physiology and disease (Review). World J Surg. 2009;33:667–84.
Pan ZW, Lu YJ, Yang BF. MicroRNAs: a novel class of potential therapeutic targets for cardiovascular diseases (Review). Zhongguo Yao Li Xue Bao/Acta Pharmacol Sin. 2010;31:9.
Nakasa T, Miyaki S, Okubo A, Hashimoto M, Nishida K, Ochi M, et al. Expression of microRNA-146 in rheumatoid arthritis synovial tissue. Arthritis Rheumatism. 2008;58:1284–92.
Xiao B, Wang Y, Li W, Baker M, Guo J, Corbet K, et al. Plasma microRNA signature as a noninvasive biomarker for acute graft-versus-host disease. Blood. 2013;122:3365–75.
Ranganathan P, Heaphy CE, Costinean S, Stauffer N, Na C, Hamadani M, et al. Regulation of acute graft-versus-host disease by microRNA-155. Blood. 2012;119:4786–97.
Stickel N, Hanke K, Marschner D, Prinz G, Kohler M, Melchinger W, et al. MicroRNA-146a reduces MHC-II expression via targeting JAK/STAT signaling in dendritic cells after stem cell transplantation. Leukemia. 2017;31:2732–41.
Chen S, Smith BA, Iype J, Prestipino A, Pfeifer D, Grundmann S, et al. MicroRNA-155-deficient dendritic cells cause less severe GVHD through reduced migration and defective inflammasome activation. Blood. 2015;126:103–12.
Sorror M. How I assess comorbidities prior to hematopoietic cell transplantation. Blood. 2013;121:2854–63.
Xie LN, Zhou F, Liu XM, Fang Y, Yu Z, Song NX, et al. Serum microRNA155 is increased in patients with acute graft-versus-host disease. Clin Transplant. 2014;28:314–23.
Knouf EC, Wyman SK, Tewari M. The human TUT1 nucleotidyl transferase as a global regulator of microRNA abundance. PLoS ONE. 2013;8:e69630.
Geiss GK, Bumgarner RE, Birditt B, Dahl T, Dowidar N, Dunaway DL, et al. Direct multiplexed measurement of gene expression with color-coded probe pairs. Nat Biotechnol. 2008;26:317–25.
Pritchard CC, Cheng HH, Tewari M. MicroRNA profiling: approaches and considerations. Nat Rev Genet. 2012;13:358–69.
Derda AA, Thum S, Lorenzen JM, Bavendiek U, Heineke J, Keyser B. et al. Blood-based microRNA signatures differentiate various forms of cardiac hypertrophy. Int J Cardiol. 2015;196:115–22. https://doi.org/10.1016/j.ijcard.2015.05.185.
Liu Y, Cai Q, Bao PP, Su Y, Cai H, Wu J, et al. Tumor tissue microRNA expression in association with triple-negative breast cancer outcomes. Breast Cancer Res Treat. 2015;152:183–91.
Schaefer JS, Attumi T, Opekun AR, Abraham B, Hou J, Shelby H, et al. MicroRNA signatures differentiate Crohn’s disease from ulcerative colitis. BMC Immunol. 2015;16:5.
Sayed AS, Xia K, Li F, Deng X, Salma U, Li T, et al. The diagnostic value of circulating microRNAs for middle-aged (40-60-year-old) coronary artery disease patients. Clinics. 2015;70:257–63.
Zhu H, Leung SW. Identification of microRNA biomarkers in type 2 diabetes: a meta-analysis of controlled profiling studies. Diabetologia. 2015;58:900–11.
Steen SO, Iversen LV, Carlsen AL, Burton M, Nielsen CT, Jacobsen S, et al. The circulating cell-free microRNA profile in systemic sclerosis is distinct from both healthy controls and systemic lupus erythematosus. J Rheumatol. 2015;42:214–21.
Vinuesa CG, Rigby RJ, Yu D. Logic and extent of miRNA-mediated control of autoimmune gene expression (Review). Int Rev Immunol. 2009;28(4-Mar):112–38.
Villa C, Fenoglio C, De Riz M, Clerici F, Marcone A, Benussi L, et al. Role of hnRNP-A1 and miR-590-3p in neuronal death: genetics and expression analysis in patients with Alzheimer disease and frontotemporal lobar degeneration. Rejuvenation Res. 2011;14:275–81.
Ortega FJ, Mercader JM, Catalan V, Moreno-Navarrete JM, Pueyo N, Sabater M, et al. Targeting the circulating microRNA signature of obesity. Clin Chem. 2013;59:781–92.
Gidlof O, Smith JG, Miyazu K, Gilje P, Spencer A, Blomquist S, et al. Circulating cardio-enriched microRNAs are associated with long-term prognosis following myocardial infarction. BMC Cardiovasc Disord. 2013;13:12.
Adyshev DM, Moldobaeva N, Mapes B, Elangovan V, Garcia JG. MicroRNA regulation of nonmuscle myosin light chain kinase expression in human lung endothelium. Am J Respir Cell Mol Biol. 2013;49:58–66.
Gandhi R, Healy B, Gholipour T, Egorova S, Musallam A, Hussain MS, et al. Circulating microRNAs as biomarkers for disease staging in multiple sclerosis. Ann Neurol. 2013;73:729–40.
Ellis KL, Cameron VA, Troughton RW, Frampton CM, Ellmers LJ, Richards AM. Circulating microRNAs as candidate markers to distinguish heart failure in breathless patients. Eur J Heart Fail. 2013;15:1138–47.
Vuppalanchi R, Liang T, Goswami CP, Nalamasu R, Li L, Jones D, et al. Relationship between differential hepatic microRNA expression and decreased hepatic cytochrome P450 3A activity in cirrhosis. PLoS ONE. 2013;8:e74471.
Lin J, Welker NC, Zhao Z, Li Y, Zhang J, Reuss SA, et al. Novel specific microRNA biomarkers in idiopathic inflammatory bowel disease unrelated to disease activity. Mod Pathol. 2014;27:602–8.
Ren J, Zhang J, Xu N, Han G, Geng Q, Song J, et al. Signature of circulating microRNAs as potential biomarkers in vulnerable coronary artery disease. PLoS ONE. 2013;8:e80738.
Lu Y, Hou S, Huang D, Luo X, Zhang J, Chen J, et al. Expression profile analysis of circulating microRNAs and their effects on ion channels in Chinese atrial fibrillation patients. Int J Clin Exp Med. 2015;8:845–53.
Huang C, Zheng JM, Cheng Q, Yu KK, Ling QX, Chen MQ, et al. Serum microRNA-29 levels correlate with disease progression in patients with chronic hepatitis B virus infection. J Dig Dis. 2014;15:614–21.
Niu G, Li B, Sun J, Sun L. miR-454 is down-regulated in osteosarcomas and suppresses cell proliferation and invasion by directly targeting c-Met. Cell Prolif. 2015;48:348–55.
Ward JA, Esa N, Pidikiti R, Freedman JE, Keaney JF, Tanriverdi K, et al. Circulating cell and plasma microRNA profiles differ between non-ST-segment and ST-segment-elevation myocardial infarction. Fam Med Med Sci Res. 2013;2:108.
Mitchell PS, Parkin RK, Kroh EM, Fritz BR, Wyman SK, Pogosova-Agadjanyan EL, et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci USA. 2008;105:10513–8.
Hunter MP, Ismail N, Zhang X, Aguda BD, Lee EJ, Yu L, et al. Detection of microRNA expression in human peripheral blood microvesicles. PLoS ONE. 2008;3:e3694. [Erratum appears in PLoS ONE. 2010;5 https://doi.org/10.1371/annotation/b15ca816-7b62-4474-a568-6b60b8959742].
Funding
This study was supported by grants DK056465, HL099993, and HL088021 from the National Institutes of Health. MLS is also supported by a Research Scholar Grant No. RSG-13-084-01-CPHPS from the American Cancer Society (ACS) and a Contract No. CE-1304-7451 from the Patient-Centered Outcome Research Institute (PCORI). We are grateful to Gary Schoch for acquisition of data. We would like to thank Bonnie Larson, Helen Crawford, and Jennifer E. Nyland for their assistance with manuscript preparation.
Author contributions
MLS, BJT-S, and MT contributed to study design; MLS collected data and obtained funding for the study; MLS and JH contributed to sample analyses; TAG performed statistical analysis; MLS, KM, JH, MAM, BJT-S, and MT contributed to analysis and interpretation of results; MLS wrote the manuscript; MLS, MAM, BJT-S, and MT edited the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
KHM is employed by NanoString Technologies, which provided the assay to determine expressions of miRs. The remaining authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Sorror, M.L., Gooley, T.A., Maclean, K.H. et al. Pre-transplant expressions of microRNAs, comorbidities, and post-transplant mortality. Bone Marrow Transplant 54, 973–979 (2019). https://doi.org/10.1038/s41409-018-0352-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41409-018-0352-9
This article is cited by
-
Overexpression of miR-92a attenuates kidney ischemia–reperfusion injury and improves kidney preservation by inhibiting MEK4/JNK1-related autophagy
Cellular & Molecular Biology Letters (2023)