Abstract
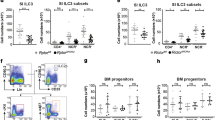
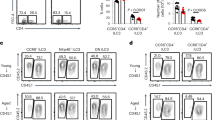
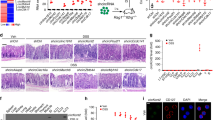
Group 3 innate lymphoid cells (ILC3s) are mediators of intestinal immunity and barrier function. Recent studies have investigated the role of the mammalian target of rapamycin complex (mTOR) in ILC3s, whereas the mTORC1-related mechanisms and crosstalk between mTORC1 and mTORC2 involved in regulating ILC3 homeostasis remain unknown. In this study, we found that mTORC1 but not mTORC2 was critical in ILC3 development, IL-22 production, and ILC3-mediated intestinal homeostasis. Single-cell RNA sequencing revealed that mTORC1 deficiency led to disruption of ILC3 heterogeneity, showing an increase in differentiation into ILC1-like phenotypes. Mechanistically, mTORC1 deficiency decreased the expression of NFIL3, which is a critical transcription factor responsible for ILC3 development. The activities of both mTORC1 and mTORC2 were increased in wild-type ILC3s after activation by IL-23, whereas inhibition of mTORC1 by Raptor deletion or rapamycin treatment resulted in increased mTORC2 activity. Previous studies have demonstrated that S6K, the main downstream target of mTORC1, can directly phosphorylate Rictor to dampen mTORC2 activity. Our data found that inhibition of mTORC1 activity by rapamycin reduced Rictor phosphorylation in ILC3s. Reversing the increased mTORC2 activity via heterozygous or homozygous knockout of Rictor in Raptor-deleted ILC3s resulted in severe ILC3 loss and complete susceptibility to intestinal infection in mice with mTORC1 deficiency (100% mortality). Thus, mTORC1 acts as a rheostat of ILC3 heterogeneity, and mTORC2 protects ILC3s from severe loss of cells and immune activity against intestinal infection when mTORC1 activity is diminished.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data are included in the manuscript. The datasets analyzed in the current study are available from the corresponding author on reasonable request.
References
Spits H, Artis D, Colonna M, Diefenbach A, Di Santo JP, Eberl G, et al. Innate lymphoid cells–a proposal for uniform nomenclature. Nat Rev Immunol. 2013;13:145–9.
Klose CSN, Artis D. Innate lymphoid cells control signaling circuits to regulate tissue-specific immunity. Cell Res. 2020;30:475–91.
Gury-BenAri M, Thaiss CA, Serafini N, Winter DR, Giladi A, Lara-Astiaso D, et al. The spectrum and regulatory landscape of intestinal innate lymphoid cells are shaped by the microbiome. Cell. 2016;166:1231–46.e13.
von Burg N, Chappaz S, Baerenwaldt A, Horvath E, Dasgupta SB, Ashok D, et al. Activated group 3 innate lymphoid cells promote T-cell-mediated immune responses. Proc Natl Acad Sci USA. 2014;111:12835–40.
Castro-Dopico T, Fleming A, Dennison TW, Ferdinand JR, Harcourt K, Stewart BJ, et al. GM-CSF calibrates macrophage defense and wound healing programs during intestinal infection and inflammation. Cell Rep. 2020;32:107857.
Pearson C, Thornton EE, McKenzie B, Schaupp AL, Huskens N, Griseri T, et al. ILC3 GM-CSF production and mobilisation orchestrate acute intestinal inflammation. eLife. 2016;5:e10066.
O’Neill LA, Kishton RJ, Rathmell J. A guide to immunometabolism for immunologists. Nat Rev Immunol. 2016;16:553–65.
Jones RG, Pearce EJ. MenTORing immunity: mTOR signaling in the development and function of tissue-resident immune cells. Immunity. 2017;46:730–42.
Saxton RA, Sabatini DM. mTOR signaling in growth, metabolism, and disease. Cell. 2017;168:960–76.
Pollizzi KN, Powell JD. Regulation of T cells by mTOR: the known knowns and the known unknowns. Trends Immunol. 2015;36:13–20.
Szwed A, Kim E, Jacinto E. Regulation and metabolic functions of mTORC1 and mTORC2. Physiol Rev. 2021;101:1371–426.
Wang F, Meng M, Mo B, Yang Y, Ji Y, Huang P, et al. Crosstalks between mTORC1 and mTORC2 variagate cytokine signaling to control NK maturation and effector function. Nat Commun. 2018;9:4874.
Teufel C, Horvath E, Peter A, Ercan C, Piscuoglio S, Hall MN, et al. mTOR signaling mediates ILC3-driven immunopathology. Mucosal Immunol. 2021;14:1323–34.
Di Luccia B, Gilfillan S, Cella M, Colonna M, Huang SC. ILC3s integrate glycolysis and mitochondrial production of reactive oxygen species to fulfill activation demands. J Exp Med. 2019;216:2231–41.
Castellanos JG, Woo V, Viladomiu M, Putzel G, Lima S, Diehl GE, et al. Microbiota-induced TNF-like ligand 1A drives group 3 innate lymphoid cell-mediated barrier protection and intestinal T cell activation during colitis. Immunity. 2018;49:1077–89.e5.
Tumanov AV, Koroleva EP, Guo X, Wang Y, Kruglov A, Nedospasov S, et al. Lymphotoxin controls the IL-22 protection pathway in gut innate lymphoid cells during mucosal pathogen challenge. Cell Host Microbe. 2011;10:44–53.
Deng Y, Wu S, Yang Y, Meng M, Chen X, Chen S, et al. Unique phenotypes of heart resident type 2 innate lymphoid cells. Front Immunol. 2020;11:802.
Huang P, Wang F, Yang Y, Lai W, Meng M, Wu S, et al. Hematopoietic-specific deletion of Foxo1 promotes NK Cell specification and proliferation. Front Immunol. 2019;10:1016.
Hao Y, Hao S, Andersen-Nissen E, Mauck WM 3rd, Zheng S, Butler A, et al. Integrated analysis of multimodal single-cell data. Cell. 2021;184:3573–87.e29.
Aibar S, González-Blas CB, Moerman T, Huynh-Thu VA, Imrichova H, Hulselmans G, et al. SCENIC: single-cell regulatory network inference and clustering. Nat methods. 2017;14:1083–6.
Hänzelmann S, Castelo R, Guinney J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinforma. 2013;14:7.
Yao B, Yang Q, Yang Y, Li Y, Peng H, Wu S, et al. Screening for active compounds targeting human natural killer cell activation identifying daphnetin as an enhancer for IFN-γ production and direct cytotoxicity. Front Immunol. 2021;12:680611.
Buonocore S, Ahern PP, Uhlig HH, Ivanov II, Littman DR, Maloy KJ, et al. Innate lymphoid cells drive interleukin-23-dependent innate intestinal pathology. Nature. 2010;464:1371–5.
Ivanov II, McKenzie BS, Zhou L, Tadokoro CE, Lepelley A, Lafaille JJ, et al. The orphan nuclear receptor RORgammat directs the differentiation program of proinflammatory IL-17+ T helper cells. Cell. 2006;126:1121–33.
Bando JK, Gilfillan S, Song C, McDonald KG, Huang SC, Newberry RD, et al. The tumor necrosis factor superfamily member RANKL suppresses effector cytokine production in group 3 innate lymphoid cells. Immunity. 2018;48:1208–19.e4.
Chen L, He Z, Slinger E, Bongers G, Lapenda TLS, Pacer ME, et al. IL-23 activates innate lymphoid cells to promote neonatal intestinal pathology. Mucosal Immunol. 2015;8:390–402.
Liu B, Liu N, Zhu X, Yang L, Ye B, Li H, et al. Circular RNA circZbtb20 maintains ILC3 homeostasis and function via Alkbh5-dependent m(6)A demethylation of Nr4a1 mRNA. Cell Mol Immunol. 2021;18:1412–24.
Krzywinska E, Sobecki M, Nagarajan S, Zacharjasz J, Tambuwala MM, Pelletier A, et al. The transcription factor HIF-1α mediates plasticity of NKp46+ innate lymphoid cells in the gut. J Exp Med. 2022;219:e20210909.
Klose CS, Kiss EA, Schwierzeck V, Ebert K, Hoyler T, d’Hargues Y, et al. A T-bet gradient controls the fate and function of CCR6-RORgammat+ innate lymphoid cells. Nature. 2013;494:261–5.
Viant C, Rankin LC, Girard-Madoux MJ, Seillet C, Shi W, Smyth MJ, et al. Transforming growth factor-β and Notch ligands act as opposing environmental cues in regulating the plasticity of type 3 innate lymphoid cells. Sci Signal. 2016;9:ra46.
Fiancette R, Finlay CM, Willis C, Bevington SL, Soley J, Ng STH, et al. Reciprocal transcription factor networks govern tissue-resident ILC3 subset function and identity. Nat Immunol. 2021;22:1245–55.
Cunningham JT, Rodgers JT, Arlow DH, Vazquez F, Mootha VK, Puigserver P. mTOR controls mitochondrial oxidative function through a YY1-PGC-1alpha transcriptional complex. Nature. 2007;450:736–40.
Saleiro D, Platanias LC. Intersection of mTOR and STAT signaling in immunity. Trends Immunol. 2015;36:21–9.
Guo X, Qiu J, Tu T, Yang X, Deng L, Anders RA, et al. Induction of innate lymphoid cell-derived interleukin-22 by the transcription factor STAT3 mediates protection against intestinal infection. Immunity. 2014;40:25–39.
Bauché D, Joyce-Shaikh B, Fong J, Villarino AV, Ku KS, Jain R, et al. IL-23 and IL-2 activation of STAT5 is required for optimal IL-22 production in ILC3s during colitis. Sci Immunol. 2020;5:eaav1080.
Geiger TL, Abt MC, Gasteiger G, Firth MA, O’Connor MH, Geary CD, et al. Nfil3 is crucial for development of innate lymphoid cells and host protection against intestinal pathogens. J Exp Med. 2014;211:1723–31.
Seillet C, Rankin LC, Groom JR, Mielke LA, Tellier J, Chopin M, et al. Nfil3 is required for the development of all innate lymphoid cell subsets. J Exp Med. 2014;211:1733–40.
Laplante M, Sabatini DM. mTOR signaling in growth control and disease. Cell. 2012;149:274–93.
Sarbassov DD, Ali SM, Sengupta S, Sheen JH, Hsu PP, Bagley AF, et al. Prolonged rapamycin treatment inhibits mTORC2 assembly and Akt/PKB. Mol cell. 2006;22:159–68.
Julien LA, Carriere A, Moreau J, Roux PP. mTORC1-activated S6K1 phosphorylates Rictor on threonine 1135 and regulates mTORC2 signaling. Mol Cell Biol. 2010;30:908–21.
Kim J, Guan KL. mTOR as a central hub of nutrient signalling and cell growth. Nat Cell Biol. 2019;21:63–71.
Delgoffe GM, Pollizzi KN, Waickman AT, Heikamp E, Meyers DJ, Horton MR, et al. The kinase mTOR regulates the differentiation of helper T cells through the selective activation of signaling by mTORC1 and mTORC2. Nat Immunol. 2011;12:295–U117.
Dibble CC, Asara JM, Manning BD. Characterization of Rictor phosphorylation sites reveals direct regulation of mTOR complex 2 by S6K1. Mol Cell Biol. 2009;29:5657–70.
Acknowledgements
This study was supported by funding from the National Natural Science Foundation of China (Nos. 81874313 and 81922068 to YCD; No. 81703521 to YFD; No. 81900055 to HYP; and No. 82070571 to YLW). This study was also supported by funding from the Natural Science Foundation of Hunan Province (No. 2023JJ30320 to YFD and No. 2021JJ40274 to HYP) and the Natural Science Foundation of Changsha, Hunan Province (No. kq2208090 to YFD) and the Project of Army Medical University (2109XQ04 to YCD). This work was also supported by funding from the Science Foundation of Hunan Children’s Hospital.
Author information
Authors and Affiliations
Contributions
The work presented was performed in collaboration by all authors. YFD designed and performed the experiments, analyzed the data, and wrote the manuscript. STW, HYP, YNL, YY, MM, LLH, PWX, SYL, QLY, LLW and XYL performed the experiments and analyzed the data. LPL, XLL and XHL designed the research and supervised the study. YLW, ZHX, JHY and YCD revised the concept, designed the research, supervised the study, and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Deng, Yf., Wu, St., Peng, Hy. et al. mTORC2 acts as a gatekeeper for mTORC1 deficiency-mediated impairments in ILC3 development. Acta Pharmacol Sin 44, 2243–2252 (2023). https://doi.org/10.1038/s41401-023-01120-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41401-023-01120-8